Abstract
In renewing tissues, mutations conferring selective advantage may result in clonal expansions1,2,3,4. In contrast to somatic tissues, mutations driving clonal expansions in spermatogonia (CES) are also transmitted to the next generation. This results in an effective increase of de novo mutation rate for CES drivers5,6,7,8. CES was originally discovered through extreme recurrence of de novo mutations causing Apert syndrome5. Here, we develop a systematic approach to discover CES drivers as hotspots of human de novo mutation. Our analysis of 54,715 trios ascertained for rare conditions9,10,11,12,13, 6,065 control trios12,14,15,16,17,18,19 and population variation from 807,162 mostly healthy individuals20 identifies genes manifesting rates of de novo mutations inconsistent with plausible models of disease ascertainment. We propose 23 genes hypermutable at loss-of-function (LoF) sites as candidate CES drivers. An extra 17 genes feature hypermutable missense mutations at individual positions, suggesting CES acting through gain of function. CES increases the average mutation rate roughly 17-fold for LoF genes in both control trios and sperm and roughly 500-fold for pooled gain-of-function sites in sperm21. Positive selection in the male germline elevates the prevalence of genetic disorders and increases polymorphism levels, masking the effect of negative selection in human populations. Despite the excess of mutations in disease cohorts for 19 LoF CES driver candidates, only 9 show clear evidence of disease causality22, suggesting that CES may lead to false-positive disease associations.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All data used in this study, including intermediate processed datasets, are provided in Supplementary Tables 1–20. Associated summary files are also available on Zenodo (https://zenodo.org/records/15660433)61.
Code availability
The full analysis pipeline, including code to reproduce figures and statistical analyses, is available on GitHub at https://github.com/mikemoldovan/CES_Discovery.
References
Lawson, A. R. J. et al. Somatic mutation and selection at population scale. Nature https://doi.org/10.1038/s41586-025-09584-w (2025).
Maeda, H. & Kakiuchi, N. Clonal expansion in normal tissues. Cancer Sci. 115, 2117–2124 (2024).
Bernstein, N. et al. Analysis of somatic mutations in whole blood from 200,618 individuals identifies pervasive positive selection and novel drivers of clonal hematopoiesis. Nat. Genet. 56, 1147–1155 (2024).
Fowler, J. C. & Jones, P. H. Somatic mutation: what shapes the mutational landscape of normal epithelia? Cancer Discov. 12, 1642–1655 (2022).
Goriely, A., McVean, G. A. T., Röjmyr, M., Ingemarsson, B. & Wilkie, A. O. M. Evidence for selective advantage of pathogenic FGFR2 mutations in the male germ line. Science 301, 643–646 (2003).
Tiemann-Boege, I. et al. The observed human sperm mutation frequency cannot explain the achondroplasia paternal age effect. Proc. Natl Acad. Sci. USA 99, 14952–14957 (2002).
Maher, G. J. et al. Selfish mutations dysregulating RAS-MAPK signaling are pervasive in aged human testes. Genome Res. 28, 1779–1790 (2018).
Wood, K. A. et al. SMAD4 mutations causing Myhre syndrome are under positive selection in the male germline. Am. J. Hum. Genet. 111, 1953–1969 (2024).
Kaplanis, J. et al. Evidence for 28 genetic disorders discovered by combining healthcare and research data. Nature 586, 757–762 (2020).
Hamdan, F. F. et al. High rate of recurrent de novo mutations in developmental and epileptic encephalopathies. Am. J. Hum. Genet. 101, 664–685 (2017).
Jin, S. C. et al. Contribution of rare inherited and de novo variants in 2,871 congenital heart disease probands. Nat. Genet. 49, 1593–1601 (2017).
Zhou, X. et al. Integrating de novo and inherited variants in 42,607 autism cases identifies mutations in new moderate-risk genes. Nat. Genet. 54, 1305–1319 (2022).
Richter, F. et al. Genomic analyses implicate noncoding de novo variants in congenital heart disease. Nat. Genet. 52, 769–777 (2020).
Francioli, L. C. et al. Genome-wide patterns and properties of de novo mutations in humans. Nat. Genet. 47, 822–826 (2015).
Iossifov, I. et al. The contribution of de novo coding mutations to autism spectrum disorder. Nature 515, 216–221 (2014).
Genome of the Netherlands Consortium. Whole-genome sequence variation, population structure and demographic history of the Dutch population. Nat. Genet. 46, 818–825 (2014).
An, J.-Y. et al. Genome-wide de novo risk score implicates promoter variation in autism spectrum disorder. Science 362, eaat6576 (2018).
Gulsuner, S. et al. Spatial and temporal mapping of de novo mutations in schizophrenia to a fetal prefrontal cortical network. Cell 154, 518–529 (2013).
Zhao, G. et al. Gene4Denovo: an integrated database and analytic platform for de novo mutations in humans. Nucleic Acids Res. 48, D913–D926 (2020).
Karczewski, K. J. et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 581, 434–443 (2020).
Neville, M. D. C. et al. Sperm sequencing reveals extensive positive selection in the male germline. Nature https://doi.org/10.1038/s41586-025-09448-3 (2025).
Amberger, J. S., Bocchini, C. A., Scott, A. F. & Hamosh, A. OMIM.org: leveraging knowledge across phenotype–gene relationships. Nucleic Acids Res. 47, D1038–D1043 (2019).
Halldorsson, B. V. et al. Characterizing mutagenic effects of recombination through a sequence-level genetic map. Science 363, eaau1043 (2019).
Seplyarskiy, V. B. & Sunyaev, S. The origin of human mutation in light of genomic data. Nat. Rev. Genet. 22, 672–686 (2021).
Seplyarskiy, V. et al. A mutation rate model at the basepair resolution identifies the mutagenic effect of polymerase III transcription. Nat. Genet. 55, 2235–2242 (2023).
Totsika, V., Liew, A., Absoud, M., Adnams, C. & Emerson, E. Mental health problems in children with intellectual disability. Lancet Child Adolesc. Health 6, 432–444 (2022).
Raudvere, U. et al. g:Profiler: a web server for functional enrichment analysis and conversions of gene lists (2019 update). Nucleic Acids Res. 47, W191–W198 (2019).
Tate, J. G. et al. COSMIC: the catalogue of somatic mutations in cancer. Nucleic Acids Res. 47, D941–D947 (2019).
Boßelmann, C. M. et al. Analysis of 1386 epileptogenic brain lesions reveals association with DYRK1A and EGFR. Nat. Commun. 15, 10429 (2024).
Gogate, A. et al. The genetic landscape of autism spectrum disorder in an ancestrally diverse cohort. NPJ Genomic Med. 9, 62 (2024).
Gallego-Martinez, A. et al. Using coding and non-coding rare variants to target candidate genes in patients with severe tinnitus. NPJ Genomic Med. 7, 70 (2022).
Wainberg, M. et al. Deletion of loss-of-function-intolerant genes and risk of 5 psychiatric disorders. JAMA Psychiatry 79, 78–81 (2022).
Rodan, L. H. et al. Gain-of-function variants in the ODC1 gene cause a syndromic neurodevelopmental disorder associated with macrocephaly, alopecia, dysmorphic features, and neuroimaging abnormalities. Am. J. Med. Genet. A. 176, 2554–2560 (2018).
Jansen, S. et al. De novo truncating mutations in the last and penultimate exons of PPM1D cause an intellectual disability syndrome. Am. J. Hum. Genet. 100, 650–658 (2017).
Harpak, A., Bhaskar, A. & Pritchard, J. K. Mutation rate variation is a primary determinant of the distribution of allele frequencies in humans. PLoS Genet. 12, e1006489 (2016).
Schraiber, J. G., Spence, J. P. & Edge, M. D. Estimation of demography and mutation rates from one million haploid genomes. Am. J. Hum. Genet. 112, 2152–2166 (2025).
Wakeley, J., Fan, W.-T. L., Koch, E. & Sunyaev, S. Recurrent mutation in the ancestry of a rare variant. Genetics 224, iyad049 (2023).
Nei, M. The frequency distribution of lethal chromosomes in finite populations. Proc. Natl Acad. Sci. USA 60, 517–524 (1968).
Zeng, T., Spence, J. P., Mostafavi, H. & Pritchard, J. K. Bayesian estimation of gene constraint from an evolutionary model with gene features. Nat. Genet. 56, 1632–1643 (2024).
Szklarczyk, D. et al. The STRING database in 2021: customizable protein-protein networks, and functional characterization of user-uploaded gene/measurement sets. Nucleic Acids Res. 49, D605–D612 (2021).
Schuurs-Hoeijmakers, J. H. M. et al. Recurrent de novo mutations in PACS1 cause defective cranial-neural-crest migration and define a recognizable intellectual-disability syndrome. Am. J. Hum. Genet. 91, 1122–1127 (2012).
Olson, H. E. et al. A recurrent de novo PACS2 heterozygous missense variant causes neonatal-onset developmental epileptic encephalopathy, facial dysmorphism, and cerebellar dysgenesis. Am. J. Hum. Genet. 102, 995–1007 (2018).
Goriely, A. et al. Activating mutations in FGFR3 and HRAS reveal a shared genetic origin for congenital disorders and testicular tumors. Nat. Genet. 41, 1247–1252 (2009).
Ichimura, K. et al. Recurrent neomorphic mutations of MTOR in central nervous system and testicular germ cell tumors may be targeted for therapy. Acta Neuropathol. 131, 889–901 (2016).
Kimura, T. et al. Conditional loss of PTEN leads to testicular teratoma and enhances embryonic germ cell production. Development 130, 1691–1700 (2003).
Sommerer, F. et al. Mutations of BRAF and RAS are rare events in germ cell tumours. Int. J. Cancer 113, 329–335 (2005).
Knudson Hypothesis—an overview. ScienceDirect Topics https://www.sciencedirect.com/topics/medicine-and-dentistry/knudson-hypothesis (2002).
Goriely, A. & Wilkie, A. O. M. Paternal age effect mutations and selfish spermatogonial selection: causes and consequences for human disease. Am. J. Hum. Genet. 90, 175–200 (2012).
Penrose, L. S. Parental age and mutation. Lancet 269, 312–313 (1955).
EuroEPINOMICS-RES Consortium, Epilepsy Phenome/Genome Project, & Epi4K Consortium. De novo mutations in synaptic transmission genes including DNM1 cause epileptic encephalopathies. Am. J. Hum. Genet. 95, 360–370 (2014).
Helbig, K. L. et al. Diagnostic exome sequencing provides a molecular diagnosis for a significant proportion of patients with epilepsy. Genet. Med. 18, 898–905 (2016).
Heyne, H. O. et al. De novo variants in neurodevelopmental disorders with epilepsy. Nat. Genet. 50, 1048–1053 (2018).
Klöckner, C. et al. De novo variants in SNAP25 cause an early-onset developmental and epileptic encephalopathy. Genet. Med. 23, 653–660 (2021).
Allen, A. S. et al. De novo mutations in epileptic encephalopathies. Nature 501, 217–221 (2013).
Perez, G. et al. The UCSC Genome Browser database: 2025 update. Nucleic Acids Res. 53, D1243–D1249 (2025).
Karlsson, M. et al. A single–cell type transcriptomics map of human tissues. Sci. Adv. 7, eabh2169 (2021).
Xu, Y. et al. A single-cell transcriptome atlas profiles early organogenesis in human embryos. Nat. Cell Biol. 25, 604–615 (2023).
Agrawal, A. et al. WikiPathways 2024: next generation pathway database. Nucleic Acids Res. 52, D679–D689 (2023).
McLaren, W. et al. The Ensembl variant effect predictor. Genome Biol. 17, 122 (2016).
Cheng, J. et al. Accurate proteome-wide missense variant effect prediction with AlphaMissense. Science 381, eadg7492 (2023).
Moldovan, M. Reproducibility data for the paper ‘Cohort-level analysis of human de novo mutations points to drivers of clonal expansion in spermatogonia’. Zenodo https://doi.org/10.5281/zenodo.15660433 (2025).
Acknowledgements
We thank C. Boix, C. Chiang and R. Stana as well as A. Quinlan and J. Kunisaki for valuable discussions and helpful suggestions that improved the analyses presented in this study. This work was supported by the National Institutes of Health through grant nos. R35GM12713, R01MH101244 and U01HG012009.
Author information
Authors and Affiliations
Contributions
V.S., M.A.M. and S.S. jointly conceived the study and developed the methodological framework. V.S., M.A.M., E.K., P.K. and M.D.C.N. performed data analysis and interpreted results. V.S., M.A.M., E.K., P.K., M.D.C.N., R.R. and S.S. collaboratively drafted and revised the paper. S.S., R.R., V.S. and M.A.M. jointly supervised the project. All authors reviewed and approved the final version of the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Ziyue Gao, Mikkel Schierup and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
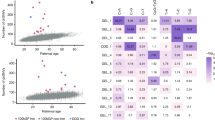
Extended Data Fig. 1 Counts of de novo variants in the NDD cohort.
(a) Numbers of de novo missense variants stratified by recurrence. Genes harboring variants occurring >10 times in the cohort are shown in blue. (b) Numbers of de novo loss-of-function variants aggregated by gene stratified by recurrence. (c) Scatter plot of observed vs. expected de novo loss-of-function variant counts in the NDD cohort. LoF-1 set genes are shown in red; LoF-2 set genes are shown in purple. Upper bound of disease ascertainment in Eq. (1) given by the lower bound of prevalence of 1% is shown as a dashed line.
Extended Data Fig. 2 Expression of the identified CES drivers in germline tissues.
(a) nTPM values for spermatogonia reported in The Human Protein Atlas single-cell dataset. (b) nTPM values normalized by the maximal expression across all tissues for each gene. (c) nTPM values in oocytes. (d) Normalized nTPM values in oocytes.
Extended Data Fig. 3 Stability of LoF-2 set with respect to the metric of loss-of-function constraint.
(a) Observed-to-expected variant count ratio (o/e) for de novo LoFs in genes with FDR < 0.1 in the neurodevelopmental disorder cohort (NDD) merged with the autism spectrum disorder (ASD) cohort plotted against the Loss-of-function Observed/Expected Lower-bound Fraction (LOELF) scores. The dashed violet line indicates the minimal LOELF value across LoF-2 genes of 0.23. LoF-2 genes are shown in violet, LoF-1 genes are shown in red, genes above the chosen LOELF threshold but not included in the LoF-2 set (SHANK3 and ZMYM2) are shown in black. (b) Same as in (a), but for the Loss-of-function Observed/Expected (LOE) metric. The upper bound for LOE (shown as violet dashed line) is 0.355.
Extended Data Fig. 4 Ratio of observed-to-expected counts of LoF de novo mutations in a cohort of control trios for LoF-2 set genes.
The ratio is shown as a function of the LOEUF threshold: we aggregate all genes with LOEUF values lower than the value indicated on the x-axis and calculate the cumulative observed-to-expected ratio. The shaded grey area represents the 95% confidence interval obtained by permuting the LOEUF labels.
Extended Data Fig. 5 Validation of LoF-2 genes with a non-LOE metric.
Prior of the GeneBayes shet calculated using biological features of genes (x-axis) and the shet values updated with LoF polymorphism data from gnomAD-v4 (y-axis). See section ‘GeneBayes update and CES’ for details.
Extended Data Fig. 6 Expected LoF rate in NanoSeq data for LoF-1 and LoF-2 genes.
Rates are shown separately for genes overlapping with those significant in NanoSeq and for private LoF-1/2 genes. An asterisk (*) indicates p < 0.05 from the Mann–Whitney U test.
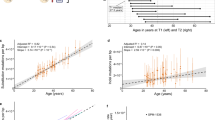
Extended Data Fig. 7 Power analysis of paternal transmission.
Statistical power (i.e., the probability of correctly detecting a signal when it exists calculated as the complement of type-2 error rate ß) of the Binomial test for paternal overtransmission relative to the baseline of 0.75 is shown across the range of CES-related mutation rate inflations κ and counts of observed variants. Results are presented for three significance levels: 0.05, 0.01, and 0.001.
Supplementary information
Supplementary Text
This file contains four Supplementary Notes 1–4. Note 1, Equivalence of de novo mutation rate and mutation probability under low mutation rates: justification for using the Bernoulli approximation to the Poisson distribution when the expected count is low. Note 2, CHIP leads to misinterpretation of LOEUF: rationale for, and description of, further filtering of results based on the involvement of genes in clonal haematopoiesis. Note 3, Misannotations of protein-truncating variants (PTVs) lead to misinterpretations of LOEUF: rationale for, and description of, further filtering of results to account for misannotation of predicted LoF variants. Note 4, Ascertainment of disease-causing variants under arbitrary trait architectures: derivation of equation (1) in the main text and discussion of its applicability to traits with non-strictly monogenic architecture.
Supplementary Fig. 1
Heatmap of expression of identified CES drivers in fetal single-cell expression clusters identified by Xu et al. (ref. 57).
Supplementary Tables
This file contains Supplementary Tables 1–20.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Seplyarskiy, V., Moldovan, M.A., Koch, E. et al. Hotspots of human mutation point to clonal expansions in spermatogonia. Nature (2025). https://doi.org/10.1038/s41586-025-09579-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41586-025-09579-7