Abstract
Butterflies serve as key indicators of climate change impacts such as shifts in emergence timing and shifts in geographic range and distribution. However, the development of commonly used ecological forecasts based on butterfly physiological tolerance of temperature change has lagged behind that of other taxonomic groups. Here, we provide a series of related datasets comprising butterfly thermal physiological traits to enable such forecasts. We compiled data from the literature on butterfly heat and cold tolerance (critical thermal maxima and minima) for 117 species as well as heat resistance (knockdown time) for 45 species. We also present a new dataset comprising heat and cold tolerance and thermal sensitivity of metabolic rate of 28 common North American butterfly species. We envision these data to not only provide foundations for contemporary ecological forecasts of vulnerability to recent climate change, but also to aid in our understanding of butterfly ecology and evolution over historical timescales.
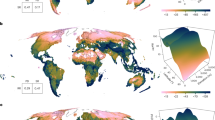
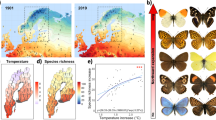
Similar content being viewed by others
Background & Summary
Datasets of heat and cold tolerance traits have been compiled across large spatial and taxonomic scales1. These datasets have contributed usefully to our understanding of the ecology and evolution of thermal physiology2. They have likewise provided the foundations for forecasts of vulnerability to ongoing climate change3. However, there are substantial biases in taxonomic coverage of thermal tolerance trait data. While butterflies (Papilionoidea) have been widely used to assess the impacts of climate change on natural systems4,5, including as indicators of overall ecosystem health6, there are vanishingly few butterfly records in these datasets of thermal tolerance traits. For example, of the 2133 species in GlobTherm1, none of these species are butterflies (in fact, none are Lepidopterans, i.e. are in the order containing moths and butterflies). Of the 102 species in a dataset of thermal tolerance acclimation capacity, none are butterflies (seven species belong to Lepidoptera, but all are moths)7. Indeed, although there are several trait-based syntheses for butterflies, such as LepTraits8 and others9,10, they tend to comprise resource breadth, morphology, and life history traits, rather than physiological traits related to climate change resilience.
Recently, butterfly-specific datasets of thermal tolerance traits have begun to accrue, either as a study of a particular assemblage11,12 or as cross-study trait compilations13. These data will allow for an improved understanding of historical and contemporary drivers of butterfly responses to climatic change. Yet even with these advances, limitations remain. In particular, cross-study comparisons of thermal tolerance traits are often complicated by methodological variation among studies13,14. These differences often have ecologically meaningful effects, as they can lead to qualitatively different conclusions regarding macrophysiological patterns and climate vulnerability15. Conditions prior to the assessment of tolerance such as developmental acclimation temperature regimes, acute temperature treatments (e.g. heat or cold hardening, initial assay temperatures), and developmental variation such as sex, age, and size, can all affect the estimation of tolerance16. In addition, whether tolerance is assessed via static or dynamic temperature assays, including differences in the rate of temperature change for dynamic assays, can likewise affect the estimation of tolerance (though some have argued for interoperability between static and dynamic tolerance estimates using thermal tolerance landscapes)17. On the one hand, these sources of variation in tolerance estimation can be considered a nuisance in that they introduce noise into comparative analyses. On the other hand, they represent the range of conditions over which natural populations interact with climatic variation and therefore can be important for understanding the range of possible responses to climate change. Indeed, intraspecific trait variation is increasingly recognized for its importance in the ability to forecast responses to climate change (especially across climatic gradients)18,19, but is often not captured in large cross-study trait compilations that typically focus on species mean trait values. Relatedly, covariation between multiple traits - not only tolerance, but also metrics of sub-lethal performance and life history traits - are also increasingly recognized as important quantities for forecasting responses to climate change20. However, at present, studies that quantify multiple thermal tolerance and performance traits tend to be focused on single species21,22.
Here we present two related datasets of butterfly physiological traits. Recognizing the tradeoff between enhanced data coverage (spatial and taxonomic) and diminished interoperability of trait values measured with different methods, we have developed a cross-study compilation of butterfly heat and cold tolerance traits from the available literature. These data comprise 279 total individual records among 117 species. As a supplement to these data, we compiled data on butterfly heat resistance (the length of time an organism can withstand a given high temperature environment until loss of motor function), which we converted into heat tolerance (the highest temperature an individual can withstand for a given length of time until loss of motor function); these data comprise 151 total individual records among 45 species. Likewise recognizing the potential importance of intraspecific trait variation, we experimentally assessed heat and cold tolerance traits as well as sub-lethal thermal performance (metabolic rate) and body mass for common North American butterflies. These data comprise 638 total individual records for thermal tolerance and 264 total individual records for metabolic rate among 28 species. Although these data are not spatially diverse, as they are obtained from several locations near the range centroids of these species, they uniquely combine heat and cold tolerance data with metabolic rate assessed at two acute test temperatures and are measured under standardized conditions and protocols. We envision these complementary datasets to be of use to ecological and evolutionary thermal physiologists and researchers interested in forecasting vulnerability to ongoing climate change.
Methods
Multicontinental datasets
We performed a literature search to identify studies with heat and cold tolerance data in butterflies. We performed searches in both Web of Science and Google Scholar. The last access date was 12 June 2024. We included peer-reviewed publications and dissertations, but not pre-printed manuscripts. In our main dataset, we focused on estimates of the critical thermal maximum, CTmax, and the critical thermal minimum, CTmin, as these were the most broadly available estimates measured in a consistent manner across different studies. The estimates include those from thermal tolerance assays where the temperature was dynamically ramped over time and those from thermal death time curves fitted to data from static temperature treatments (four estimates among three species and three studies)23,24,25. We used the following search terms: “butterfly”, “Papilionoidea”, “Lepidoptera”, “heat tolerance”, “cold tolerance”, “CTmax”, “CTmin”, “critical thermal maximum”, “critical thermal minimum”, “upper thermal limit”, “lower thermal limit”. These search criteria returned several studies that contained data on mortality across different temperatures. However, we ultimately excluded these studies as they would require fitting thermal performance curves and extracting heat and cold tolerance parameters, and most studies lacked sufficient sampling towards high and low extreme temperatures that would allow for robust curve fitting.
We likewise performed a literature search to identify studies with heat resistance data in butterflies. In this companion dataset, we focused on estimates of knockdown time at stressfully high temperatures. Heat knockdown time quantifies the length of time an organism can withstand a high temperature challenge before the onset of heat coma, i.e. when they are knocked down. We used the following search terms: “butterfly”, “Papilionoidea”, “Lepidoptera”, “heat resistance”, “heat knockdown time”, “HKD”, “HKDT”.
To be able to compare the heat resistance data beyond the conditions of a given study, we used thermal death time curves to estimate CTmax from knockdown time at a particular temperature26. We emphasize caution here in the interpretation of these CTmax values as they rely on the choice of temperature sensitivity coefficient. While this coefficient can be estimated for a particular species or population with multiple static temperature measures of heat knockdown time, this value must be assumed in cases of heat knockdown time at a single temperature. We computed CTmax from static single estimates of heat knockdown time based on the median temperature sensitivity coefficient among other butterfly species (median z = 5.20; Bicyclus anynana, range z = 4.62–5.72, n = 5; Pieris napi, range z = 4.10–4.11, n = 2; Pieris rapae, z = 5.10, n = 1)23,24,25. These sensitivity coefficients are generally within the range of values reported for other ectothermic species, range z = 0.8–5.7, n = 4127. Importantly, while there is clearly inter- and intra-specific variation in the temperature sensitivity coefficient, we lack this information for most observations and species in the heat resistance dataset.
Estimates of tolerance or resistance on wild-caught organisms were generally presented as species means and associated error measurements, occasionally with the data reported separately by sex. However, laboratory-based studies generally had multiple subgroups of data including different sexes, life stages, and treatment conditions (e.g. temperature regimes, food resource regimes). In these cases, we followed the presentation of the data by the authors and report means and errors associated with each group. We likewise followed the authors with respect to naming conventions for latin binomial descriptions of genera, species, and subspecies.
To extract data from articles, we used a combination of summary statistics from raw data included as a supplementary file to the article, extraction of values reported in the article text or table display elements, and extraction of values from article figure display elements using WebPlotDigitizer28.
As a minimum criterion for entry into the multicontinental dataset, we required a mean tolerance or resistance value. In a limited number of cases, only medians were available and were used as a summary statistic of central tendency. When possible, we included standard errors associated with mean tolerance or resistance values, either by reporting these directly or by calculating standard errors from other measures of spread such as from standard deviations, interquartile ranges, and sample sizes. We also did not require the tolerance or resistance data to be georeferenced, though again, we included this information when possible. In some cases, we were able to match an individual tolerance or resistance value to a single geographic location. In other cases, we were only able to match an individual tolerance or resistance value to a group of sites. In these cases, we report the geographic coordinates of all potential collection sites or each individual tolerance or resistance value. Although the combined multicontinental dataset of thermal tolerance and resistance traits contains records from each continent where butterflies are found except Australia, the data are nonetheless sparse in terms of spatial coverage (Fig. 1).
Sample locations for species in the multicontinental dataset. Each point represents an individual sampling location, with points colored by the natural log transformed number of species measured for thermal tolerance or resistance at each site. Thermal tolerance data are denoted by circles, and heat resistance data are denoted by triangles. Points have been strongly jittered to avoid overlap in some areas, and consequently, sampling locations are approximate on this map. Fourteen species in the heat and cold tolerance dataset and one species in the heat resistance dataset lacked geographic information and are not represented on this map.
Finally, we incorporated species means and standard errors for CTmin and CTmax among the species comprising our North American dataset (see below) into our multicontinental thermal tolerance dataset. These data are fully georeferenced at the observation level. To include them on a comparable scale to those in the multicontinental dataset, we computed species trait means for each collection location. Sex could not be determined for all individuals, so data were pooled across sexes.
The combined multicontinental heat and cold tolerance trait dataset includes representatives from all six butterfly families (Fig. 2). By contrast, the heat resistance trait dataset includes representatives from only three butterfly families (Lycaenidae, Pieridae, and Nymphalidae; Fig. 3). In our datasets, and as others have pointed out, research on butterfly responses to temperature tends to be biased towards the Nymphalidae family, though this is also the most globally diverse butterfly family29.
Heat and cold tolerance trait values mapped onto the NCBI taxonomy. Clade labels highlight the six butterfly families. For species with multiple entries in the multicontinental thermal tolerance dataset, species mean trait value are presented here. One species had no matches in the NCBI database and is represented only at the genus level (Opoptera sulcius). For records where individuals could not be identified to species in the original study, we include only the genus name.
Heat tolerance estimated from heat resistance (i.e. knockdown time using the median thermal sensitivity coefficient for butterflies of z = 5.20) mapped onto the NCBI taxonomy. Note that the color scale for CTmax is matched to that of Fig. 2 for comparison. For species with multiple entries in the heat resistance dataset, species mean trait value are presented here. Five species with sub-species designations had no matches in the NCBI database and were promoted to the level of species to be able to extract their taxonomic information. These included Heliconius clysonymus clysonymus, Heliconius elevatus willmotti, Heliconius sara sprucei, Heliconius xanthocles napoensis, and Pieris mannii alpigena.
North American dataset
Although the North American dataset contributes species (by location) means for CTmin and CTmax to the multicontinental dataset described above, we also provide access to the individual-level data for these traits and others including metabolic rate and body mass. Because thermal tolerance, metabolic rate, and body mass measurements were taken on the same individuals (excepting heat and cold tolerance measurements on the same individual, as these assays were destructive), they can be used to explore intraspecific associations between traits (Fig. 4a,b) in addition to interspecific comparisons of trait means (Fig. 4c).
Metabolic rate as a function of body mass at 20 and 30 °C for Cupido comyntas (panels a and b, respectively); both variables are natural log transformed. Panel c characterizes the thermal sensitivity of metabolic rate (as Q10 values) for species with at least 5 individuals measured at both 20 and 30 °C. Means, 95% confidence intervals (non-parametric bootstrap), and raw data are shown.
Collection information
Adult butterflies were collected in northeastern Ohio, USA from 2021 to 2023. Collection sites included Case Western Reserve University (latitude, longitude: 41.5043, −81.6084), Cuyahoga Valley National Park (41.2808, −81.5678), Holden Forests and Gardens (41.6027, −81.3061), Squire Valleevue Farm (41.4979, −81.4240), and private property in the city of Independence (41.3687, −81.6379), though we note that over 95% of the data came from the Squire Valleevue Farm site. Seasonal collections began in May and continued through late September or early October each year.
We used aerial nets (BioQuip, 38 cm diameter) to collect butterflies. For transportation from the field sites to the laboratory, butterflies were placed individually into 90–946 mL containers (with holes for airflow), depending on the size of the butterfly. At the laboratory, butterflies were transferred and housed individually in mesh cages (28 × 28 × 28 cm BugDorms from BioQuip). Butterflies were provided with a paper towel moistened with tap water (aged at least 24 hours) and a petri dish (60 × 15 mm) containing a sponge and a 10% sugar-water solution.
The butterflies were held for a 24 to 48-hour acclimation period in Percival growth chambers (36-VL) programmed to a constant 25 °C and a 14:10 L:D photoperiod cycle. This acclimation period was intended to minimize acute acclimation responses to field environmental conditions prior to the assessment of physiological traits.
Metabolic rate
To assess the thermal sensitivity of routine metabolic rate, we measured the metabolic rate of individuals at two constant acute test temperatures, 20 and 30 °C. Individuals were tested at 20 and 30 °C in a random order. To quantify metabolic rate at each test temperature, we used a CO2/H2O gas analyzer (Licor-7000) in push mode that pushed air from the environment (scrubbed of CO2 and H2O with soda lime and Drierite) through two flow control meters (Alicat Scientific, MC-200SCCM until July 2021 and MC-1SLPM from July 2021 through October 2023) and then a respirometry flow multiplexer (RM-8). The flow controllers were calibrated (Gilian Gilibrator-2 NIOSH) at both 20 and 30 °C. The respirometry equipment was held within a dark growth chamber (MIR 154 PHCBI for trials performed 2021-2022, and a MIR 153 for trials performed in 2023) set to 20 or 30 °C. The dark environment helped to limit movement during the trials. Butterflies were generally resting, although we could not entirely exclude the possibility of movement during the trials (e.g. walking around the testing chambers). Once placed inside the respirometry chambers, animals were allowed to acclimate at the given test temperature for 15 minutes prior to recording of metabolic rate.
We tested butterflies of small body mass inside 30 mL glass chambers, and butterflies of large body mass in 650 mL chambers (RC and RC-1, Sable Systems International). In the multiplexer, each animal chamber contained one individual, and CO2 was recorded for 10 minutes, with a two-minute flush, at a constant flow rate, adjusted separately for each test temperature to achieve a volumetric flow rate of 100 mL min−1. Note that there were a total of eight chambers, with one empty chamber to assess the baseline and up to seven animal chambers per trial. For butterflies of large body mass, the larger respirometry chambers required a higher flow rate, adjusted to achieve a target volumetric flow rate of 500 mL min−1, and a longer flush of 14 minutes. CO2 concentration (ppm) from the animal chambers was compared to the CO2 concentration (ppm) from the returning control line and recorded by the Licor-7000.
The metabolic rate data were processed through a UI-3 data acquisition interface and ExpeData software (Sable Systems International). Once all calibrations were complete, we then converted the raw CO2 values to the rate of CO2 production (VCO2 mL min−1) by dividing the values by 1,000,000 to get the fractional CO2 value, and multiplied this value by the flow rate. We considered the first five minutes as a settling-in period and used the last five minutes of recording to compute the mean metabolic rate over this period. These butterflies breathe continuously and so the five-minute interval is sufficient to get an estimation of metabolic rate, unlike species with discontinuous breathing30. We performed a complementary method to detect the flattest part of the trace using a rolling window analysis (each five-minute interval possible over the 10 minutes of recording) and identifying the lowest slope value. Because calculations of metabolic rate based on this rolling window method were nearly identical to the last five minutes recording approach, we elected to use the latter method for simplicity. While we report the mean metabolic rate for each butterfly at each temperature, we also computed thermal sensitivity of metabolic rate. We standardized the change in metabolic rate across the two test temperatures as Q10 values, which describe the increase in metabolic rate for every 10 °C increase in temperature.
Heat and cold tolerance
Following assessment of metabolic rate at both the 20 and 30 °C test temperatures, individuals were allowed to recover for a 15-minute period on the laboratory bench (~23 °C) prior to assessment of thermal tolerance. Individuals were randomly assigned to be assessed for either heat tolerance or cold tolerance. Heat tolerance was assessed by the critical thermal maximum, CTmax, and cold tolerance by the critical thermal minimum, CTmin. For both tolerance estimates, the loss of wing movement was used to determine the critical temperature.
The CTmax and CTmin trials were performed using water baths (A40 ARCTIC / SC150, A10 ARCTIC / SC150, or S13 SAHARA / SC150, Thermo Scientific) with a dynamic temperature ramping protocol of 1 °C min−1. The temperature ramped upward starting at 35 °C for CTmax. The temperature was ramped downward starting at 15 °C for CTmin. At the start of the trial, all individuals were placed in plastic containers plugged with a sponge. We used three different sizes of container based on the size of the butterfly. The containers needed to be large enough to allow the butterfly to fully extend its wings. For small butterflies we used 200 mL containers. For large butterflies, we used 473 mL or 946 mL containers. Butterflies were allowed to acclimate to the starting water bath temperature for 15 minutes prior to the start of temperature ramping. After thermal tolerance trials, butterflies were placed into glassine envelopes and frozen at −20 °C.
Body mass
Body mass of each individual was recorded multiple times. First, body mass was recorded following collection from the field and prior to being housed in the lab for pre-trial acclimation. Body mass was also recorded just prior to and immediately following the assessment of metabolic rate, and just prior to and immediately following the assessment of thermal tolerance (either CTmax or CTmin). Body mass was recorded to a precision of 0.0001 g (Sartorius MSE124S-100-DA).
Data Records
The multicontinental datasets and North American dataset, collectively referred to as “PapilioTherm” (version 1.0), are hosted on the Open Science Framework31 (https://osf.io/fguxa/). All data are stored as comma separated value (.csv) files. Metadata are contained in a single text file. Owing to the multicontinental and North American datasets being summarized at different levels, two distinct sets of files house the multicontinental (species or subgroup level) and North American (observation level) datasets. Note that the species-by-site mean trait values of the North American dataset are included in the multicontinental thermal tolerance dataset. Furthermore, the heat resistance (knockdown time) dataset is provided separately from the thermal tolerance dataset owing to the different types of information reported for each trait type.
Multicontinental datasets
The main multicontinental dataset includes CTmax and CTmin trait data from 14 studies11,12,21,22,23,24,25,32,33,34,35,36,37 including the new data presented here31. At a minimum, species or group-level means are reported with standard errors reported when available. Groups can be defined according to combinations of sex, developmental stage, geographic location, and the type of assay performed. We included details of conditions prior to and during the collection of these trait values, including developmental acclimation conditions, initial temperatures for the assessment of heat or cold tolerance, and the rate of temperature change for dynamic ramping assays. We also included details of data extraction and processing to bring them into a standard format of trait means and standard errors. We reported the geographic coordinates of the sampling locations for all but 14 species that were obtained from laboratory stock populations with unknown provenance (Table 1).
The companion multicontinental dataset includes heat resistance (knockdown time) from 13 studies38,39,40,41,42,43,44,45,46,47,48,49,50. We used the same inclusion criteria, grouping structure, and georeferencing as described above for our main multicontinental dataset of thermal tolerance traits. We included details on developmental acclimation conditions and the temperature at which knockdown time was assessed. We also estimated heat tolerance (CTmax) using thermal death time curves (Table 2).
North American dataset
Physiological trait data
The North American physiological trait dataset contains individual-level, temporally and spatially referenced data on CTmax, CTmin, metabolic rate at 20 and 30 °C, and body mass at capture and prior to and following each physiological trait assay (Table 3). Summary statistics of these individual-level data were incorporated into the multicontinental dataset of heat and cold tolerance.
Physical specimens
Butterfly specimens are stored in the Diamond lab at Case Western Reserve University. Specimens are stored individually within glassine envelopes at −20 °C. Whole specimen loans or other sample requests (e.g. leg for DNA extraction) should be sent to the corresponding author and will be considered on a case-by-case basis.
Technical Validation
Multicontinental datasets
All extracted data were rechecked at least once following the initial extraction. Geographic coordinates were verified against collection locality names and by visualization of collection coordinates on a global map.
North American dataset
Specimens were identified to species at the time of collection, prior to performance of the physiological trait assays. Species identity for each individual was confirmed or updated on these specimens following the physiological trait assays and subsequent storage at −20 °C. Genus and species (including subspecies) names followed the Integrated Taxonomic Information System (ITIS).
Electronic data comprising heat and cold tolerances were checked against the original paper records and either confirmed or amended. Electronic data comprising metabolic rates were visually inspected on an individual basis. In cases of obvious equipment failures, i.e. leaks in tubing connections that resulted in negative metabolic rates, we excluded those trials from further consideration.
Usage Notes
The PapilioTherm datasets provide a first step in synthesizing information on butterfly thermal physiological traits. As data continue to accumulate in this area, PapilioTherm version 1.0 will be updated with subsequent versions on the Open Science Framework31. Ideally, the inclusion of new data will help to fill existing gaps in the dataset, both in terms of taxonomic representation and geographical distribution, which are currently limited (Figs. 1–3). Specifically, greater representation of species in families other than the Nymphalidae are needed, as are data from the tropics and the Global South.
Although there are many potential applications of these data, which we enumerate below, we first point out a few areas of caution when using these data. In an analogous manner to other such syntheses of thermal physiological traits, there are limitations on comparing across traits assessed under different conditions and with different protocols51. Furthermore, converting single-value heat knockdown time values to estimates of heat tolerance likewise require caution owing to inter- and intra-specific variation in the thermal sensitivity coefficient determining the relationship between knockdown time and intensity of thermal exposure27. Finally, we emphasize caution in the interpretation of tolerance and resistance traits measured on laboratory stock populations, as inadvertent selection in these environments could bias physiological trait values relative to those of natural populations52.
That said, we highlight several areas for potential use of these data:
-
(1)
These data could be used to answer fundamental research questions on drivers of biogeographic patterns of occurrence in butterflies. For example, physiological trait data can be used to understand extant relationships between species (potential) climatic niches and species range and distribution53.
-
(2)
These data can likewise be important for understanding phenotypic correlations between traits and the ecological and evolutionary constraints that might shape those relationships54. For example, in many ectothermic species, heat tolerance is more evolutionarily constrained compared with cold tolerance, which is more strongly shaped by current climatic conditions2, but this pattern has not yet been explored in butterflies.
-
(3)
These data could be used to answer more applied research questions on species conservation in the face of global change. With many butterfly communities declining in abundance and species diversity in association with climate change and land-use change55,56,57, forecasting their vulnerability to ongoing change can provide much needed aid in targeting species for conservation and land management interventions58. Heat and cold tolerance data can be used to inform the degree to which species might be buffered under future climatic warming in place59, and the capacity to track suitable habitat in space and time60,61. Both the trait data and physical specimens provide a baseline against which future sampling can be compared. Such resampling studies have provided valuable insights into capacities of populations to cope with changes in their environment62,63.
-
(4)
Finally, the datasets and meta-data themselves might provide useful templates for future data collection in this area, highlighting the importance of information on collection sites, collection timing, pre-assay conditions, and details on the physiological trait assays themselves.
References
Bennett, J. M. et al. GlobTherm, a global database on thermal tolerances for aquatic and terrestrial organisms. Sci. Data 5, 180022 (2018).
Bennett, J. M. et al. The evolution of critical thermal limits of life on Earth. Nat. Commun. 12, 1198 (2021).
da Silva, C. R. B., Beaman, J. E., Youngblood, J. P., Kellermann, V. & Diamond, S. E. Vulnerability to climate change increases with trophic level in terrestrial organisms. Sci. Total Environ. 865, 161049 (2023).
Parmesan, C. et al. Poleward shifts in geographical ranges of butterfly species associated with regional warming. Nature 399, 579–583 (1999).
Diamond, S. E., Frame, A. M., Martin, R. A. & Buckley, L. B. Species’ traits predict phenological responses to climate change in butterflies. Ecology 92, 1005–1012 (2011).
Thomas, J. A. Monitoring change in the abundance and distribution of insects using butterflies and other indicator groups. Philos. Trans. R. Soc. B Biol. Sci. 360, 339–357 (2005).
Weaving, H., Terblanche, J. S., Pottier, P. & English, S. Meta-analysis reveals weak but pervasive plasticity in insect thermal limits. Nat. Commun. 13, 5292 (2022).
Shirey, V. et al. LepTraits 1.0 A globally comprehensive dataset of butterfly traits. Sci. Data 9, 1–7 (2022).
Stelbrink, P. et al. Colour lightness of butterfly assemblages across North America and Europe. Sci. Rep. 9, 1–10 (2019).
Middleton-Welling, J. et al. A new comprehensive trait database of European and Maghreb butterflies, Papilionoidea. Sci. Data 7, 1–10 (2020).
Ashe-Jepson, E. et al. Tropical butterflies use thermal buffering and thermal tolerance as alternative strategies to cope with temperature increase. J. Anim. Ecol. 92, 1759–1770 (2023).
Diniz E Silva, V., Beirão, M. V. & Cardoso, D. C. Thermal tolerance of fruit-feeding butterflies (Lepidoptera: Nymphalidae) in contrasting mountaintop environments. Insects 11, 278 (2020).
Mutamiswa, R., Mbande, A., Nyamukondiwa, C. & Chidawanyika, F. Thermal adaptation in Lepidoptera under shifting environments: mechanisms, patterns, and consequences. Phytoparasitica 51, 929–955 (2023).
Clusella-Trullas, S., Garcia, R. A., Terblanche, J. S. & Hoffmann, A. A. How useful are thermal vulnerability indices? Trends Ecol. Evol. 36, 1000–1010 (2021).
Allen, J. L., Chown, S. L., Janion-Scheepers, C. & Clusella-Trullas, S. Interactions between rates of temperature change and acclimation affect latitudinal patterns of warming tolerance. Conserv. Physiol. 4, cow053 (2016).
Terblanche, J. S., Deere, J. A., Clusella-Trullas, S., Janion, C. & Chown, S. L. Critical thermal limits depend on methodological context. Proc. R. Soc. Lond. B Biol. Sci. 274, 2935–2943 (2007).
Jørgensen, L. B., Malte, H. & Overgaard, J. How to assess Drosophila heat tolerance: Unifying static and dynamic tolerance assays to predict heat distribution limits. Funct. Ecol. 33, 629–642 (2019).
Moran, E. V., Hartig, F. & Bell, D. M. Intraspecific trait variation across scales: implications for understanding global change responses. Glob. Change Biol. 22, 137–150 (2016).
Palacio, R. D. & Clark, J. S. Incorporating intraspecific variation into species responses reveals both their resilience and vulnerability to future climate change. Ecography 2023, e06769 (2023).
da Silva, C. R. et al. Physiological traits and their relationships vary along an elevational gradient within and among Fijian bee species. Preprint at https://doi.org/10.1101/2022.07.27.501487 (2024).
Dongmo, M. A. K. et al. Local adaptation in thermal tolerance for a tropical butterfly across ecotone and rainforest habitats. Biol. Open 10 (2021).
Medina-Báez, O. A., Lenard, A., Muzychuk, R. A., da Silva, C. R. B. & Diamond, S. E. Life cycle complexity and body mass drive erratic changes in climate vulnerability across ontogeny in a seasonally migrating butterfly. Conserv. Physiol. 11, coad058 (2023).
Carnicer, J. et al. Phenotypic biomarkers of climatic impacts on declining insect populations: A key role for decadal drought, thermal buffering and amplification effects and host plant dynamics. J. Anim. Ecol. 88, 376–391 (2019).
Vives-Ingla, M. et al. Interspecific differences in microhabitat use expose insects to contrasting thermal mortality. Ecol. Monogr. 93, e1561 (2023).
Klockmann, M., Günter, F. & Fischer, K. Heat resistance throughout ontogeny: body size constrains thermal tolerance. Glob. Change Biol. 23, 686–696 (2017).
Rezende, E. L., Castañeda, L. E. & Santos, M. Tolerance landscapes in thermal ecology. Funct. Ecol. 28, 799–809 (2014).
Jørgensen, L. B., Malte, H., Ørsted, M., Klahn, N. A. & Overgaard, J. A unifying model to estimate thermal tolerance limits in ectotherms across static, dynamic and fluctuating exposures to thermal stress. Sci. Rep. 11, 12840 (2021).
Rohatgi, A. Webplotdigitizer: Version 4.6 (2022).
Ashe-Jepson, E. et al. Hot topics in butterfly research: Current knowledge and gaps in understanding of the impacts of temperature on butterflies. Insect Conserv. Divers. 17, 1–15 (2024).
Winwood-Smith, H. S. & White, C. R. Short-duration respirometry underestimates metabolic rate for discontinuous breathers. J. Exp. Biol. 221, jeb175752 (2018).
Diamond, S. E. PapilioTherm (version 1.0). OSF, https://doi.org/10.17605/OSF.IO/FGUXA (2024).
Andersen, M. K., Willot, Q. & MacMillan, H. A. A neurophysiological limit and its biogeographic correlations: cold-induced spreading depolarization in tropical butterflies. J. Exp. Biol. 226, jeb246313 (2023).
Au, T. F. & Bonebrake, T. C. Increased suitability of poleward climate for a tropical butterfly (Euripus nyctelius) (Lepidoptera: Nymphalidae) accompanies its successful range expansion. J. Insect Sci. 19, 2 (2019).
Lehmann, P., Javal, M. & Terblanche, J. S. Oxygen limitation is not the cause of death during lethal heat exposure in an insect. Biol. Lett. 15, 20180701 (2019).
Ling, Y. F. & Bonebrake, T. C. Consistent heat tolerance under starvation across seasonal morphs in Mycalesis mineus (Lepidoptera: Nymphalidae). Comp. Biochem. Physiol. A. Mol. Integr. Physiol. 271, 111261 (2022).
Tremblay, P., MacMillan, H. A. & Kharouba, H. M. Autumn larval cold tolerance does not predict the northern range limit of a widespread butterfly species. Ecol. Evol. 11, 8332–8346 (2021).
Lenard, A. & Diamond, S. E. Evidence of plasticity, but not evolutionary divergence, in the thermal limits of a highly successful urban butterfly. J. Insect Physiol. 155, 104648 (2024).
Fischer, K. et al. Environmental effects on temperature stress resistance in the tropical butterfly Bicyclus anynana. PLOS ONE 5, e15284 (2010).
Fischer, K., Kölzow, N., Höltje, H. & Karl, I. Assay conditions in laboratory experiments: is the use of constant rather than fluctuating temperatures justified when investigating temperature-induced plasticity? Oecologia 166, 23–33 (2011).
Bauerfeind, S. S. & Fischer, K. Simulating climate change: temperature extremes but not means diminish performance in a widespread butterfly. Popul. Ecol. 56, 239–250 (2014).
Günter, F. et al. Genotype‐environment interactions rule the response of a widespread butterfly to temperature variation. J. Evol. Biol. 33, 920–929 (2020).
Karl, I., Janowitz, S. A. & Fischer, K. Altitudinal life-history variation and thermal adaptation in the copper butterfly Lycaena tityrus. Oikos 117, 778–788 (2008).
Karl, I., Schmitt, T. & Fischer, K. Phosphoglucose isomerase genotype affects life-history traits and cold stress resistance in a copper butterfly. Funct. Ecol. 22, 887–894 (2008).
Neu, A. & Fischer, K. Indications for rapid evolution of trait means and thermal plasticity in range‐expanding populations of a butterfly. J. Evol. Biol. 35, 124–133 (2022).
Ashoff, M. & Schmitt, T. Are different allozyme genotypes of the butterfly Polyommatus coridon adapted to resist cold and heat shocks? Ann. Zool. Fenn. 51, 413–422 (2014).
Klockmann, M., Wallmeyer, L. & Fischer, K. Variation in adult stress resistance does not explain vulnerability to climate change in copper butterflies. Insect Sci. 25, 894–904 (2018).
Montejo-Kovacevich, G. et al. Microclimate buffering and thermal tolerance across elevations in a tropical butterfly. J. Exp. Biol. 223, jeb220426 (2020).
Advani, N. K. Thermal ecology of the Glanville fritillary butterfly (Melitaea cinxia). (University of Texas at Austin, 2012).
Ossola, V. Thermal adaptation in tropical clearwing butterflies. (Université Paris Cité, 2023).
Luo, S. et al. Phenotypic plasticity in thermal tolerance in the Glanville fritillary butterfly. J. Therm. Biol. 42, 33–39 (2014).
Diamond, S. E. & Yilmaz, A. R. The role of tolerance variation in vulnerability forecasting of insects. Curr. Opin. Insect Sci. 29, 85–92 (2018).
Hoffmann, A. A. & Ross, P. A. Rates and patterns of laboratory adaptation in (mostly) insects. J. Econ. Entomol. 111, 501–509 (2018).
Sunday, J. et al. Thermal tolerance patterns across latitude and elevation. Philos. Trans. R. Soc. B 374, 20190036 (2019).
White, C. R., Alton, L. A., Bywater, C. L., Lombardi, E. J. & Marshall, D. J. Metabolic scaling is the product of life-history optimization. Science 377, 834–839 (2022).
Dirzo, R. et al. Defaunation in the Anthropocene. Science 345, 401–406 (2014).
Wepprich, T., Adrion, J. R., Ries, L., Wiedmann, J. & Haddad, N. M. Butterfly abundance declines over 20 years of systematic monitoring in Ohio, USA. PLOS ONE 14, e0216270 (2019).
Diamond, S. E., Bellino, G. & Deme, G. G. Urban insect bioarks of the 21st century. Curr. Opin. Insect Sci. 57, 101028 (2023).
Hill, G. M., Kawahara, A. Y., Daniels, J. C., Bateman, C. C. & Scheffers, B. R. Climate change effects on animal ecology: butterflies and moths as a case study. Biol. Rev. Camb. Philos. Soc. 96, 2113–2126 (2021).
Deutsch, C. A. et al. Impacts of climate warming on terrestrial ectotherms across latitude. Proc. Natl. Acad. Sci. 105, 6668–6672 (2008).
Kellermann, V. & van Heerwaarden, B. Terrestrial insects and climate change: adaptive responses in key traits. Physiol. Entomol. 44, 99–115 (2019).
Comte, L. et al. Bringing traits back into the equation: A roadmap to understand species redistribution. Glob. Change Biol. 30, e17271 (2024).
MacLean, H. J., Kingsolver, J. G. & Buckley, L. B. Historical changes in thermoregulatory traits of alpine butterflies reveal complex ecological and evolutionary responses to recent climate change. Clim. Change Responses 3, 13 (2016).
Buckley, L. B., Graham, S. I. & Nufio, C. R. Grasshopper species’ seasonal timing underlies shifts in phenological overlap in response to climate gradients, variability and change. J. Anim. Ecol. 90, 1252–1263 (2021).
R Core Team. R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing. (R Foundation for Statistical Computing, Vienna, Austria. http://www.R-project.org/ 2023).
Acknowledgements
We thank Ryan Martin for helpful comments on a previous version of this manuscript. We thank Melina Gabele and Jessica Furlough for assistance with animal husbandry, Gideon Deme for assistance with field collections, and Rut Muzychuk for assistance with animal husbandry, field collections, and physiological trait assays. We also thank Case Western Reserve University, Squire Valleevue Farm, Holden Forests and Gardens, Cuyahoga Valley National Park, and private landowner Wayne Benos for access to field sites. An NSF CAREER grant (DEB-1845126) to SED provided funding.
Author information
Authors and Affiliations
Contributions
S.E.D. conceived of the idea for data collection, developed protocols, compiled data (North American and multicontinental datasets), and wrote the first draft of the manuscript. C.R.B.dS. provided advice on respirometry data collection and data processing for the North American dataset and reviewed entries for the multicontinental dataset. O.M.B. refined protocols and performed field collections and physiological trait assays (metabolic rate, thermal tolerance) for the North American dataset. All authors contributed to revising the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Diamond, S.E., da Silva, C.R.B. & Medina-Báez, O.A. A multicontinental dataset of butterfly thermal physiological traits. Sci Data 11, 1348 (2024). https://doi.org/10.1038/s41597-024-04191-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-024-04191-2