Abstract
The process of developing new drugs is arduous and costly, particularly for targets classified as “difficult-to-drug.” Macrocycles show a particular ability to modulate difficult-to-drug targets, including protein-protein interactions, while still allowing oral administration. However, the determination of membrane permeability, critical for reaching intracellular targets and for oral bioavailability, is laborious and expensive. In silico methods are a cost-effective alternative, enabling predictions prior to compound synthesis. Here, we present a comprehensive online database (https://swemacrocycledb.com/), housing 5638 membrane permeability datapoints for 4216 nonpeptidic macrocycles, curated from the literature, patents, and bioactivity repositories. In addition, we present a new descriptor, the “amide ratio” (AR), that quantifies the peptidic nature of macrocyclic compounds, enabling the classification of peptidic, semipeptidic, and nonpeptidic macrocycles. Overall, this resource fills a gap among existing databases, offering valuable insights into the membrane permeability of nonpeptidic and semipeptidic macrocycles, and facilitating predictions for drug discovery projects.
Similar content being viewed by others
Background & Summary
Developing a new drug from discovery to market is an expensive and time-consuming process1. Approximately half of the targets associated with human diseases are classified as “difficult-to-drug” with traditional molecules following Lipinski’s Rule of 5 (Ro5)2, which outlines limits for molecular weight (MW ≤ 500 Da), calculated lipophilicity (cLogP ≤ 5), as well as hydrogen bond donors and acceptors (HBD ≤ 5, HBA ≤ 10). Although biologics may be suitable for difficult-to-drug targets, their lack of cell permeability hinders access to intracellular targets and renders them unsuitable for oral administration. Recent research has shed light on the opportunities provided by compounds that reside outside the Ro5 boundaries, i.e. in the beyond Rule of 5 (bRo5) chemical space3,4,5. Among these compounds, macrocycles, characterized by a ring of at least 12 atoms, exhibit the capability to modulate difficult-to-drug targets, including those with tunnel, flat, or groove-shaped binding sites, as well as protein-protein interactions (PPIs), while still allowing for oral administration5,6,7.
Independent of chemical space, solubility, cell permeability and a not too high metabolism in the liver are the three most important determinants of the oral bioavailability of drugs. Optimizing this triad of drug properties becomes increasingly difficult as compounds grow in size, putting macrocycles and other compounds in the bRo5 space at higher risk. Despite the recent emergence of macrocyclic peptides as a promising chemical class in drug discovery8,9,10, they often suffer from issues with solubility, cell permeability and metabolic instability11. This originates from the high polarity of amide bonds in the peptide backbone12, and any polar groups in their side chains. In contrast, nonpeptidic macrocycles do not carry the burden of a polar backbone and more often display both cell permeability and oral bioavailability6.
Measurement of cell membrane permeability of drugs is not only crucial to assess their ability to reaching intracellular targets, regardless of their location in the central nervous system (CNS) or peripheral sites, but is also utilized as a model system for estimating oral absorption13. Various in vitro assays are employed to measure cell permeability, including the human colorectal adenocarcinoma cell line (Caco-2), Madin–Darby canine kidney (MDCK) cells, and the low-efflux MDCK clone Ralph Russ canine kidney (RRCK). The parallel artificial membrane permeability assay (PAMPA) provides a cost-effective assessment of passive membrane permeability in a cell free system, while the cell-based assays provide data that is more relevant for permeability and oral bioavailability in an in vivo setting. However, generating experimental permeability data is both time-consuming and expensive, in particular in cell-based systems. Alternatively, in silico methods are not only cost-effective but also sufficiently accurate and fast enough to be used as high-throughput filter in the drug discovery projects, enabling predictions before compound synthesis and testing14.
To facilitate the development of accurate and efficient computational predictions, it is crucial to collect and curate experimental data with structural information, making it available to scientific communities as per the FAIR guideline (Findable, Accessible, Interoperable, and Reusable)15. In this study, we report the construction of a membrane permeability database for 4216 macrocycles, ranging from nonpeptidic to semipeptidic, which has been collected and curated from the scientific literature, patents, and various bioactivity data repositories. This comprehensive online resource comprises structures annotated with molecular descriptors and permeability data obtained from different assays and endpoints. It is readily accessible and downloadable through the webserver (https://swemacrocycledb.com/). Our database is complementary to the CycPeptMPDB16, a comprehensive database of membrane permeability for more than 7000 cyclic peptides.
Methods
Data collection and curation
Macrocycles exhibiting membrane permeability were gathered from three different sources: 1) the scientific literature, 2) patents, and 3) public repositories and then incorporated in the database (Fig. 1, Supplementary Table 1). PubMed17 and Google Scholar were used to search the literature and identify macrocycles for which permeability data has been published. Keywords like “macrocycle” were combined with either the general term “permeability” or specific assay names (Caco-2, PAMPA, MDCK, RRCK) to query scientific journals. A similar search was done in Google Patents to collect approved patents which disclosed nonpeptidic macrocycle membrane permeability data. In a further attempt to gather data comprehensively, the ChEMBL database18 was mined using its Python web resource client; the query python code can be found in GitHub. The RDKit Molecule Substructure module19 was employed to filter macrocycles, defined as having a ring with at least 12 heavy atoms. Subsequently, all structures (SMILES) and cell permeability data were imported into Molecular Operating Environment (version 2022.02)20.
The workflow for construction of the membrane permeability database for nonpeptidic macrocycles and its functions. Structures and permeability data were retrieved from the literature, patents, and scientific databases, followed by manual curation. The webserver provides readily downloadable datasets for macrocycles evaluated in various membrane permeability assays often with different endpoints, as well as the structures and molecular descriptors of the macrocycles.
The dataset underwent manual curation, involving the removal of mixtures, inorganics, salts, solvent molecules, and also structural normalization. Descriptors for polarity (HBA and HBD) are highly influenced by the protonation state of the molecules6. Since predictions by different tools often yields different charge states for the same molecule21, we treated molecules as ‘uncharged’ for calculation of their descriptors. To allow analysis and model building permeability values were standardized, first by conversion to the unit cm/s and then by calculation of their logarithmic values. For permeability values reported with a “>” or “<” sign, this was retained in both the original and standardised values. There are 36 compounds having such undefined values in the dataset. Overall, the collection and curation resulted in a database containing 4216 diverse and unique macrocycles and 5638 permeability datapoints. In the future, the database will be updated on a biannual basis.
Quantification of the peptidic nature of macrocycles
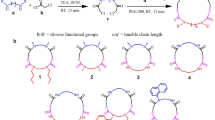
No standardized and quantitative definition exists for the peptidic nature of macrocycles. We propose that the amide ratio (AR, Eq. 1) is a relevant and intuitive descriptor of the peptidic nature of macrocycles. Calculation of the AR is based on the number of amide bonds (nAB), including both NH and N-alkylated ones, within the macrocyclic ring, multiplied by three to account for the number atoms (-C-N-Cα-) forming each amide bond. Division by the macrocycle ring size (MRS), i.e. the total number of atoms in the macrocyclic ring, then provides the AR.
AR returns values between 0 and 1, a value of 0 represents a completely nonpeptidic macrocycle, and 1 represents a full cyclic peptide. We also propose that macrocycles having an AR of from 0 to 0.3 are classified as nonpeptidic, those with an AR between 0.3 and 0.7 as semipeptidic, while an AR > 0.7 characterizes macrocycles which are mainly peptidic. The AR is identical to the recently reported the ‘Peptide Character Index’22. However, we have proposed thresholds to distinguish between nonpeptidic, semipeptidic, and peptidic macrocycles, which have been thoroughly validated using known datasets (cf. Quantification of peptide and nonpeptide macrocycles, below). Additionally, the code for calculating the AR metric is freely available.
Webserver implementation
The webserver implemented in this study was built on the Django web framework (version 3.2.23). The development of the web interface involved the use of standard web technologies, including HTML5, CSS, and JavaScript, with all data within the web server stored and managed using SQLite, a lightweight and efficient relational database management system. RDKit (version 2023.9.5)19 was employed for molecule visualization. Specifically, RDKit was used to generate structures that include the stereochemistry of the macrocycles23, and convert the resulting isomeric SMILES into PNG and SDF files within the web interface. ECharts (version v5.5.0) was utilized to support online data visualization24. The functionality for table sorting and filtering was implemented using DataTables (https://www.datatables.net/), a JavaScript library for enhancing HTML tables. The website has been thoroughly tested to ensure functionality across multiple operating systems and web browsers. Most of the codes used in this work are open-source and properly acknowledged. The code for the final version of the web server is provided on GitHub.
Data Records
The structures of the 4216 unique macrocycles, their molecular descriptors and the 5638 permeability datapoints reported for them are available on the https://swemacrocycledb.com/ web server (Fig. 2a). A unique molecule ID identifies each macrocycle, for which multiple permeability measurements may have been reported and included in the database. Three categories of information is provided for each permeability measurement: (i) a Representation, where the structure, InChI Key, isomeric SMILES for the overall macrocycle and the SMILES for the macrocyclic ring is shown; (ii) Permeability information containing the type of permeability assay, the endpoint, value, and unit; and (iii) key Molecular Descriptors for the macrocycle, including the descriptors of Lipinski´s2 and Veber´s25 rules, as well as other descriptors of macrocycle flexibility and structure.
Schematic representations of (a) a report which is available for each permeability entry in the database and (b) the ‘Search’ and ‘Sort’ functions. Abbreviations: MW: Molecular weight; HBA: Hydrogen bond acceptor; HBD: Hydrogen bond donor; cLogP: Calculated lipophilicity; TPSA: Topological polar surface area; NRotB: Number of rotatable bonds; Φ: Kier flexibility Index; AR: amide ratio; Fsp3: fraction of sp3 carbon atoms; MRS: macrocyclic ring size.
Due to the high computational cost and uncertainty in the conformational sampling of macrocycles26,27, this database does not provide conformations. Instead, we provide isomeric SMILES (contains chirality information) and descriptors for each macrocycle, as described above. The original sources from which the structure and permeability data were extracted are also available for the user. In order to provide ready-to-use datasets for QSAR modelling, all membrane permeability values were standardized into logarithmic values. All data records incorporated in the database are ready to download. The browse menu also offers multiple search and sorting options, primarily by unique ID, common name, permeability assay, endpoint, permeability value, unit and standardized permeability value (Fig. 2b).
In addition to the web server resource (https://swemacrocycledb.com/), the peer-reviewed version 1.0 of this database has been archived as a static repository on Figshare (https://doi.org/10.6084/m9.figshare.26964259)28. The repository is organized into two main directories, “Data” and “Code,” with a README file to guide users through the directory structure and contents. The Data directory is divided into three subdirectories, each containing membrane permeability data for specific endpoints in comma-separated values (.csv) format. A 2D representation of each compound is also available as an image file (.png) and in structured data file (.sdf) format, identified by a unique macrocyclic ID for consistency and easy reference. The Code directory includes a Jupyter Notebook documenting the step-by-step processing and data analysis workflow. This notebook allows users to directly access and run the code used for data extraction and preprocessing of macrocycles with membrane permeability data, enhancing reproducibility.
Technical Validation
Membrane permeability database statistics
Sources
The dataset reported herein consists of 5638 permeability datapoints for 4612 macrocycles, collected from 103 scientific articles and 9 patents published during 2006–2023 (last updated July 2023) as well as data from the ChEMBL database. Out of the 5638 datapoint records, 84%, 4%, and 11% are from scientific articles, patents, and the ChEMBL repository, respectively. Assays: The dataset has been divided into five membrane permeability assay categories (Fig. 3a and Table 1), namely, PAMPA, Caco-2, MDCK, RRCK, and others. PAMPA-based passive permeability records account for 67% (n = 3767), among which 91% of the datapoints (n = 3462) are from one publication29. This publication contains log Peff data measured under consistent experimental conditions, making it the largest source of consistent macrocycle permeability data available in the public domain. The next highest number of datapoint records originates from the Caco-2 assay, comprising 26% (n = 1502) of the datapoints. The largest categories of entries from the Caco-2 assay have Log Papp AB (permeability in the apical to basolateral direction) and Log Papp BA (permeability in the basolateral to apical direction), together with their efflux inhibited versions Log Papp AB + Inh and Log Papp BA + Inh, determined in the presence of a cocktail of efflux inhibitors, as endpoints. The efflux ratio (ER = Papp AB/Papp BA) reveals whether a compound undergoes active efflux, i.e. if it is actively transported out of the cells and how fast this transport is compared to passive uptake into the cells, while the ER + Inh shows to what extent the transporter mediated efflux can be blocked by inhibitors. Another commonly used cell-based permeability assay, using MDCK cells, had 264 datapoints with Log Papp AB and ER as the two major endpoints reported. Not many macrocycles have RRCK data (n = 7), while 98 cell permeability datapoints originate from other types of assays.
(a) Nested-pie chart of the permeability data for the macrocycles in the database. The different types of permeability assays are indicated in the inner ring, endpoints in the outer ring. (b) TMAP visualization of the structural diversity of the nonpeptidic macrocyclic dataset (n = 4216). The different types of permeability assays are highlighted on the tree. (c) Molecular property distribution of the macrocycles included in the database, as described by 2D molecular descriptors representing size, polarity, a lipophilicity, flexibility, and amide ratio. The upper limits of the descriptors of the Ro5 and Veber´s rule are indicated by grey shading. Abbreviations: MW: Molecular weight; MRS: macrocyclic ring size; HBA: Hydrogen bond acceptor; HBD: Hydrogen bond donor; TPSA: Topological polar surface area; cLogP: Calculated lipophilicity; Φ: Kier flexibility Index; AR: amide ratio.
Macrocycle diversity
TMAP30, a tree-based high-dimensional visualization tool, which provides both local and distant structural cluster information, was used to characterize the structural diversity of the macrocycles in the dataset (Fig. 3b). TMAP clearly illustrates that the permeability data provided in the web server originates from a structurally very diverse set of macrocycles. In addition, mapping of the membrane permeability assays on the TMAP tree reveals that the data from the three major assays (PAMPA, Caco-2 and MDCK) has been generated for macrocycles that show a large structural diversity. The dataset consists of both nonpeptidic and semipeptidic macrocycles. The semipeptides are situated on the right side of the TMAP tree, exhibiting a higher fraction of sp3 carbons and larger macrocyclic rings compared to the nonpeptide macrocycles, which predominantly originate from the dataset reported by Rzepiela, et al. (n = 3462)29.
Molecular property analysis
To assess the diversity of the molecular properties of the macrocycles in the database, we analysed the distribution of key 2D molecular descriptors representing size [molecular weight (MW), macrocycle ring size (MRS)], polarity [hydrogen bond acceptors (HBA), hydrogen bond donors (HBD), topological polar surface area (TPSA)], lipophilicity (cLogP), flexibility [Kier flexibility index (Phi), number of rotatable bonds (NRotB)], and the peptide nature [amide ratio (AR)]. The molecular descriptors of a large number of the macrocycles in the dataset adhere to the cut-offs of Lipinski’s2 and Veber’s25 rules for drug-likeness (Fig. 3c). This is particularly true for polarity (HBA, HBD, TPSA) and lipophilicity (cLogP), while close to half of the macrocycles have a MW above the 500 Da cutoff. More than 350 compounds, accounting for 9% of the macrocycles in the dataset, reside in the beyond the rule of five space (bRo5) as defined by Doak, et al.5. These compounds have the potential to modulate difficult-to-drug targets, including those with extensive, flat, or groove-shaped binding sites, as well as protein-protein interactions, while still allowing for oral administration3.
Quantification of peptide and nonpeptide macrocycles
Since no metric that quantifies whether a macrocycle is nonpeptidic, semipeptidic or peptidic has been generally accepted, we proposed the amide ratio (AR) of the macrocyclic ring as a simple descriptor for quantification of the peptidic nature of macrocycles (see Methods section). Combination of the macrocycles from the nonpeptidic database reported herein and the cyclic peptide database (CycPeptMPDB)16 revealed that the three classes were well differentiated by the proposed AR cut offs and also validated that the cut offs reflect the terminology used in the literature (Fig. 4a). For instance, the vast majority of the compounds in the CycPeptMPDB including the drug cyclosporin A are classified as peptidic by the AR, while semipeptides31,32,33, are also classified in agreement with the original publications.
(a) Distribution of the amide ratio (AR) of macrocycles obtained by combination of the macrocycles from the nonpeptidic database reported herein (blue bars, n = 4216) and the cyclic peptide database (CycPeptMPDB16, red bars, n = 7849). The figure has been made so that the macrocycles from each database make up 100%. Classification of macrocycles by AR is shown above the figure. Representative examples of nonpeptidic (1 and 2), semipeptidic (3) and peptidic (4) macrocycles are shown for low to high AR values. Amide bonds within the macrocyclic ring have been shaded in blue. (b) Principal component analysis (PCA) comparing the chemical space of cyclic peptides (CycPeptMPDB) and nonpeptides and semipeptides from the database reported herein, with descriptor contributions highlighted by arrows. The first two principal components explain 88.7% of the variance in the dataset. (c) The chemical space of macrocycles reported in this study depicted using the first two principal components, which explain 75.1% of the variance in the dataset. Macrocycles are colored according to their amide ratio (AR) with blue to red circles in the two PCAs. The PCAs were constructed using the 10 descriptors provided for each macrocycle in the database.
A principal component analysis (PCA) of the combined set of the macrocycles from the nonpeptidic database reported herein and the cyclic peptide database (CycPeptMPDB)16 confirmed that nonpeptidic and peptidic macrocycles populated different parts of chemical space (Fig. 4b). Semipeptides were found in several regions, with most being found in between the nonpeptide and peptide classes. As expected most cyclic peptides were larger, more polar (higher TPSA and HBD count) and more flexible (higher Kier index, Φ) than the nonpeptidic macrocycles, which were somewhat more lipophilic. A separate PCA of only the nonpeptidic macrocycles from this database, but with cyclosporin A included as reference, showed a similar trend of semipeptides being more polar than the nonpeptides (Fig. 4c). Cyclosporin A was located in a chemical space far from the two other classes.
Usage Notes
The complete dataset consisting of the 4216 unique macrocycles, their molecular descriptors and the 5638 permeability datapoints available for them can be accessed at a webserver located at https://swemacrocycledb.com/28. The webserver offers three primary options for accessing and handling macrocyclic cell permeability data in the Browse, Download, and Statistic sections.
In the Browse section users can select permeability datasets for macrocycles they judge to be of interest. Users can select macrocycles by unique ID, name, assay type, molecular weight, endpoints, or a combination thereof and download the data as a CSV file. Clicking on each unique molecule ID in the selected set opens a separate window displaying the name and structure, permeability data, and molecular descriptors for the selected macrocycle. Additionally, any other permeability endpoints available for the macrocycle are provided, just as a list of similar macrocycles based on the same ‘macrocyclic ring’. These functionalities of the webserver help users to find all permeability endpoints reported for a macrocycle, and directs the user to neighbouring compounds and their molecular characteristics. The Download section allows users to download the full dataset or subsets selected by the user as a CSV file which includes the structure, cell permeability, and molecular descriptors of the macrocycles, including their peptidic nature and the original source of the permeability data. In the Statistics section users can analyse both cell permeability data and molecular descriptors for the overall dataset, and the three major subsets by permeability endpoint.
Code availability
All the data connected to this article is available without restriction on the https://swemacrocycledb.com/ webserver. All source code is available on the GitHub (https://github.com/Macrocycle-Cell-Permeability/NPMMP-DB) and https://doi.org/10.6084/m9.figshare.26964259 with no restrictions to access.
References
Wouters, O. J., McKee, M. & Luyten, J. Research and Development Costs of New Drugs-Reply. JAMA 324, 518 (2020).
Lipinski, C. A., Lombardo, F., Dominy, B. W. & Feeney, P. J. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Advanced Drug Delivery Reviews 46, 3–26 (2001).
Doak, B. C., Over, B., Giordanetto, F. & Kihlberg, J. Oral druggable space beyond the rule of 5: insights from drugs and clinical candidates. Chemistry Biology 21, 1115–1142 (2014).
Poongavanam, V., Doak, B. C. & Kihlberg, J. Opportunities and guidelines for discovery of orally absorbed drugs in beyond rule of 5 space. Current Opinion in Chemical Biology 44, 23–29 (2018).
Doak, B. C., Zheng, J., Dobritzsch, D. & Kihlberg, J. How Beyond Rule of 5 Drugs and Clinical Candidates Bind to Their Targets. Journal of Medicinal Chemistry 59, 2312–2327 (2016).
Garcia Jimenez, D., Poongavanam, V. & Kihlberg, J. Macrocycles in Drug Discovery─Learning from the Past for the Future. Journal of Medicinal Chemistry 66, 5377–5396 (2023).
Over, B. et al. Structural and conformational determinants of macrocycle cell permeability. Nat Chem Biol 12, 1065–1074 (2016).
Merz, M. L. et al. De novo development of small cyclic peptides that are orally bioavailable. Nat Chem Biol, (2023).
Matsson, P., Doak, B. C., Over, B. & Kihlberg, J. Cell permeability beyond the rule of 5. Advanced Drug Delivery Reviews 101, 42–61 (2016).
Ohta, A. et al. Validation of a New Methodology to Create Oral Drugs beyond the Rule of 5 for Intracellular Tough Targets. Journal of the American Chemical Society 145, 24035–24051 (2023).
Dougherty, P. G., Sahni, A. & Pei, D. Understanding Cell Penetration of Cyclic Peptides. Chemical Reviews 119, 10241–10287 (2019).
Kenny, P. W. Hydrogen-Bond Donors in Drug Design. Journal of Medicinal Chemistry 65, 14261–14275 (2022).
Di, L. et al. The Critical Role of Passive Permeability in Designing Successful Drugs. ChemMedChem 15, 1862–1874 (2020).
Williams-Noonan, B. J. et al. Membrane Permeating Macrocycles: Design Guidelines from Machine Learning. Journal of Chemical Information and Modeling 62, 4605–4619 (2022).
Wilkinson, M. D. et al. The FAIR Guiding Principles for scientific data management and stewardship. Scintific Data 3, 160018 (2016).
Li, J. et al. CycPeptMPDB: A Comprehensive Database of Membrane Permeability of Cyclic Peptides. Journal of Chemical Information and Modeling 63, 2240–2250 (2023).
Kim, S. et al. PubChem 2023 update. Nucleic Acids Res 51, D1373–D1380 (2023).
Davies, M. et al. ChEMBL web services: streamlining access to drug discovery data and utilities. Nucleic Acids Research 43, W612–620 (2015).
RDKit. Open-source cheminformatics, https://www.rdkit.org.
Molecular Operating Environment (MOE), Chemical Computing Group ULC, 910-1010 Sherbrooke St. W., Montreal, QC H3A 2R7, Canada, 2024).
Bergazin, T. D. et al. Evaluation of log P, pK(a), and log D predictions from the SAMPL7 blind challenge. Journal of Computer-Aided Molecular Design 35, 771–802 (2021).
Viarengo-Baker, L. A., Brown, L. E., Rzepiela, A. A. & Whitty, A. Defining and navigating macrocycle chemical space. Chem Sci 12, 4309–4328 (2021).
Scalfani, V. F., Patel, V. D. & Fernandez, A. M. Visualizing chemical space networks with RDKit and NetworkX. Journal of Cheminformatics 14, 87 (2022).
Li, D. et al. ECharts: A declarative framework for rapid construction of web-based visualization. Visual Informatics 2, 136–146 (2018).
Veber, D. F. et al. Molecular Properties That Influence the Oral Bioavailability of Drug Candidates. Journal of Medicinal Chemistry 45, 2615–2623 (2002).
Poongavanam, V. et al. Predicting the Permeability of Macrocycles from Conformational Sampling - Limitations of Molecular Flexibility. Journal of Pharmaceutical Sciences 110, 301–313 (2021).
Poongavanam, V. et al. Predictive Modeling of PROTAC Cell Permeability with Machine Learning. ACS Omega 8, 5901–5916 (2023).
Feng, Q., Chavez, D. D., Kihlberg, J. & Poongavanam, V. in https://doi.org/10.6084/m9.figshare.26964259 (Figshare, 2024).
Rzepiela, A. A., Viarengo-Baker, L. A., Tatarskii, V., Kombarov, R. & Whitty, A. Conformational Effects on the Passive Membrane Permeability of Synthetic Macrocycles. Journal of Medicinal Chemistry 65, 10300–10317 (2022).
Probst, D. & Reymond, J.-L. Visualization of very large high-dimensional data sets as minimum spanning trees. Journal of Cheminformatics 12, 12 (2020).
L’Exact, M. et al. Beyond Rule-of-five: Permeability Assessment of Semipeptidic Macrocycles. Biochimica et Biophysica Acta (BBA) - Biomembranes 1865, 184196 (2023).
Le Roux, A. et al. Structure–Permeability Relationship of Semipeptidic Macrocycles—Understanding and Optimizing Passive Permeability and Efflux Ratio. Journal of Medicinal Chemistry 63, 6774–6783 (2020).
Comeau, C. et al. Modulation of the Passive Permeability of Semipeptidic Macrocycles: N- and C-Methylations Fine-Tune Conformation and Properties. J Med Chem 64, 5365–5383 (2021).
Ahlbach, C. L. et al. Beyond Cyclosporine A: Conformation-Dependent Passive Membrane Permeabilities of Cyclic Peptide Natural Products. Future Medicinal Chemistry 7, 2121–2130 (2015).
Giroud, M. et al. Repurposing a Library of Human Cathepsin L Ligands: Identification of Macrocyclic Lactams as Potent Rhodesain and Trypanosoma brucei Inhibitors. Journal of Medicinal Chemistry 61, 3350–3369 (2018).
Pasero, C. et al. Alkyl-guanidine Compounds as Potent Broad-Spectrum Antibacterial Agents: Chemical Library Extension and Biological Characterization. Journal of Medicinal Chemistry 61, 9162–9176 (2018).
Nožinić, D. et al. Assessment of Macrolide Transport Using PAMPA, Caco-2 and MDCKII-hMDR1 Assays. Croatica Chemica Acta 83, 323–331 (2010).
DeGoey, D. A., Chen, H.-J., Cox, P. B. & Wendt, M. D. Beyond the Rule of 5: Lessons Learned from AbbVie’s Drugs and Compound Collection. Journal of Medicinal Chemistry 61, 2636–2651 (2018).
Spencer, J. A. et al. Design and Development of a Macrocyclic Series Targeting Phosphoinositide 3-Kinase δ. ACS Med. Chem. Lett. 11, 1386–1391 (2020).
Richter, J. M. et al. Design and Synthesis of Novel Meta-Linked Phenylglycine Macrocyclic FVIIa Inhibitors. ACS Med. Chem. Lett. 8, 67–72 (2017).
Zhang, H. et al. Design, Synthesis, and Optimization of Macrocyclic Peptides as Species-Selective Antimalaria Proteasome Inhibitors. Journal of Medicinal Chemistry 65, 9350–9375 (2022).
Zhang, X. et al. Discovery of a Highly Potent, Selective, and Orally Bioavailable Macrocyclic Inhibitor of Blood Coagulation Factor VIIa–Tissue Factor Complex. Journal of Medicinal Chemistry 59, 7125–7137 (2016).
Yamaguchi-Sasaki, T. et al. Discovery of a potent dual inhibitor of wild-type and mutant respiratory syncytial virus fusion proteins through the modulation of atropisomer interconversion properties. Bioorganic & Medicinal Chemistry 28, 115818 (2020).
Zheng, M. et al. Discovery of Cyclic Peptidomimetic Ligands Targeting the Extracellular Domain of EGFR. Journal of Medicinal Chemistry 64, 11219–11228 (2021).
Duan, M. et al. Discovery of novel P3-oxo inhibitor of hepatitis C virus NS3/4A serine protease. Bioorganic & Medicinal Chemistry Letters 22, 2993–2996 (2012).
William, A. D. et al. Discovery of the Macrocycle 11-(2-Pyrrolidin-1-yl-ethoxy)-14,19-dioxa-5,7,26-triaza-tetracyclo[19.3.1.1(2,6).1(8,12)]heptacosa-1(25),2(26),3,5,8,10,12(27),16,21,23-decaene (SB1518), a Potent Janus Kinase 2/Fms-Like Tyrosine Kinase-3 (JAK2/FLT3) Inhibitor for the Treatment of Myelofibrosis and Lymphoma. Journal of Medicinal Chemistry 54, 4638–4658 (2011).
Wang, L. et al. Fragment-Based, Structure-Enabled Discovery of Novel Pyridones and Pyridone Macrocycles as Potent Bromodomain and Extra-Terminal Domain (BET) Family Bromodomain Inhibitors. Journal of Medicinal Chemistry 60, 3828–3850 (2017).
Hostalkova, A. et al. Isoquinoline Alkaloids from Berberis vulgaris as Potential Lead Compounds for the Treatment of Alzheimer’s Disease. J. Nat. Prod. 82, 239–248 (2019).
Zhang, H. et al. Macrocyclic Peptides that Selectively Inhibit the Mycobacterium tuberculosis Proteasome. Journal of Medicinal Chemistry 64, 6262–6272 (2021).
Wurtz, N. R. et al. Neutral macrocyclic factor VIIa inhibitors. Bioorganic & Medicinal Chemistry Letters 27, 2650–2654 (2017).
Randolph, J. T. et al. Prodrug Strategies to Improve the Solubility of the HCV NS5A Inhibitor Pibrentasvir (ABT-530). Journal of Medicinal Chemistry 63, 11034–11044 (2020).
Saunders, G. J. & Yudin, A. K. Property-Driven Development of Passively Permeable Macrocyclic Scaffolds Using Heterocycles. Angewandte Chemie International Edition 61, e202206866 (2022).
Engelhardt, H. et al. Start Selective and Rigidify: The Discovery Path toward a Next Generation of EGFR Tyrosine Kinase Inhibitors. Journal of Medicinal Chemistry 62, 10272–10293 (2019).
Kaneda, M., Kawaguchi, S., Fujii, N., Ohno, H. & Oishi, S. Structure–Activity Relationship Study on Odoamide: Insights into the Bioactivities of Aurilide-Family Hybrid Peptide–Polyketides. ACS Med. Chem. Lett. 9, 365–369 (2018).
Miyachi, H. et al. Structure, solubility, and permeability relationships in a diverse middle molecule library. Bioorganic & Medicinal Chemistry Letters 37, 127847 (2021).
Xiao, T. et al. Synthesis and structural characterization of a monocarboxylic inhibitor for GRB2 SH2 domain. Bioorganic & Medicinal Chemistry Letters 51, 128354 (2021).
Maccari, G. et al. Synthesis of linear and cyclic guazatine derivatives endowed with antibacterial activity. Bioorganic & Medicinal Chemistry Letters 24, 5525–5529 (2014).
Ladziata, V. U. et al. Synthesis and P1’ SAR exploration of potent macrocyclic tissue factor-factor VIIa inhibitors. Bioorganic & Medicinal Chemistry Letters 26, 5051–5057 (2016).
Du-Cuny, L. et al. Computational modeling of novel inhibitors targeting the Akt pleckstrin homology domain. Bioorganic & Medicinal Chemistry 17, 6983–6992 (2009).
Halland, N., Blum, H., Buning, C., Kohlmann, M. & Lindenschmidt, A. Small Macrocycles As Highly Active Integrin α2β1 Antagonists. ACS Med. Chem. Lett. 5, 193–198 (2014).
Houštecká, R. et al. Biomimetic Macrocyclic Inhibitors of Human Cathepsin D: Structure–Activity Relationship and Binding Mode Analysis. Journal of Medicinal Chemistry 63, 1576–1596 (2020).
Joshi, A. et al. Design and Synthesis of P1–P3 Macrocyclic Tertiary-Alcohol-Comprising HIV-1 Protease Inhibitors. Journal of Medicinal Chemistry 56, 8999–9007 (2013).
Kock, H. A. D. et al. Macrocyclic inhibitors of hepatitis c virus. WO2007014918A1 (2007).
McCoull, W. et al. Discovery of Pyrazolo[1,5-a]pyrimidine B-Cell Lymphoma 6 (BCL6) Binders and Optimization to High Affinity Macrocyclic Inhibitors. Journal of Medicinal Chemistry 60, 4386–4402 (2017).
Naylor, M. R. et al. Lipophilic Permeability Efficiency Reconciles the Opposing Roles of Lipophilicity in Membrane Permeability and Aqueous Solubility. Journal of Medicinal Chemistry 61, 11169–11182 (2018).
Nilsson, M. et al. Synthesis and SAR of potent inhibitors of the Hepatitis C virus NS3/4A protease: Exploration of P2 quinazoline substituents. Bioorganic & Medicinal Chemistry Letters 20, 4004–4011 (2010).
Parsy, C. C. et al. Discovery and structural diversity of the hepatitis C virus NS3/4A serine protease inhibitor series leading to clinical candidate IDX320. Bioorganic & Medicinal Chemistry Letters 25, 5427–5436 (2015).
Parsy, C. C. et al. Macrocyclic serine protease inhibitors. WO2010118078A1 (2010).
Sund, C. et al. Design and synthesis of potent macrocyclic renin inhibitors. Bioorganic & Medicinal Chemistry Letters 21, 358–362 (2011).
Hopkins, B. A. et al. Development of a Platform To Enable Efficient Permeability Evaluation of Novel Organo-Peptide Macrocycles. ACS Med. Chem. Lett. 10, 874–879 (2019).
Li, Z. et al. Structure-Guided Design of Novel, Potent, and Selective Macrocyclic Plasma Kallikrein Inhibitors. ACS Med. Chem. Lett. 8, 185–190 (2017).
Peng, J. et al. Structure-activity relationship and mechanism of action studies of manzamine analogues for the control of neuroinflammation and cerebral infections. Journal of Medicinal Chemistry 53, 61–76 (2010).
Tess, D. A. et al. Predicting the Human Hepatic Clearance of Acidic and Zwitterionic Drugs. Journal of Medicinal Chemistry 63, 11831–11844 (2020).
Cee, V. J. et al. Discovery and Optimization of Macrocyclic Quinoxaline-pyrrolo-dihydropiperidinones as Potent Pim-1/2 Kinase Inhibitors. ACS Med. Chem. Lett. 7, 408–412 (2016).
Hess, S. et al. Effect of structural and conformation modifications, including backbone cyclization, of hydrophilic hexapeptides on their intestinal permeability and enzymatic stability. Journal of Medicinal Chemistry 50, 6201–6211 (2007).
Lindsley, S. R. et al. Design, synthesis, and SAR of macrocyclic tertiary carbinamine BACE-1 inhibitors. Bioorganic & Medicinal Chemistry Letters 17, 4057–4061 (2007).
Moore, K. P. et al. Strategies toward improving the brain penetration of macrocyclic tertiary carbinamine BACE-1 inhibitors. Bioorganic & Medicinal Chemistry Letters 17, 5831–5835 (2007).
Rescourio, G. et al. Discovery and in Vivo Evaluation of Macrocyclic Mcl-1 Inhibitors Featuring an α-Hydroxy Phenylacetic Acid Pharmacophore or Bioisostere. Journal of Medicinal Chemistry 62, 10258–10271 (2019).
Stachel, S. J. et al. Macrocyclic inhibitors of beta-secretase: functional activity in an animal model. Journal of Medicinal Chemistry 49, 6147–6150 (2006).
Wuelfing, W. P. et al. Dose Number as a Tool to Guide Lead Optimization for Orally Bioavailable Compounds in Drug Discovery. Journal of Medicinal Chemistry 65, 1685–1694 (2022).
Xu, H. et al. Quantitative measurement of intracellular HDAC1/2 drug occupancy using a trans-cyclooctene largazole thiol probe. Medchemcomm 8, 767–770 (2017).
Begnini, F. et al. Importance of Binding Site Hydration and Flexibility Revealed When Optimizing a Macrocyclic Inhibitor of the Keap1-Nrf2 Protein-Protein Interaction. J Med Chem 65, 3473–3517 (2022).
Adebesin, A. M. et al. Development of Robust 17(R),18(S)-Epoxyeicosatetraenoic Acid (17,18-EEQ) Analogues as Potential Clinical Antiarrhythmic Agents. Journal of Medicinal Chemistry 62, 10124–10143 (2019).
Andersson, V. et al. Macrocyclic Prodrugs of a Selective Nonpeptidic Direct Thrombin Inhibitor Display High Permeability, Efficient Bioconversion but Low Bioavailability. Journal of Medicinal Chemistry 59, 6658–6670 (2016).
Begnini, F. et al. Cell Permeability of Isomeric Macrocycles: Predictions and NMR Studies. ACS Med. Chem. Lett. 12, 983–990 (2021).
Begnini, F. et al. Mining Natural Products for Macrocycles to Drug Difficult Targets. Journal of Medicinal Chemistry 64, 1054–1072 (2021).
Boy, K. M. et al. Macrocyclic prolinyl acyl guanidines as inhibitors of β-secretase (BACE). Bioorganic & Medicinal Chemistry Letters 25, 5040–5047 (2015).
Clark, C. G. et al. Structure based design of macrocyclic factor XIa inhibitors: Discovery of cyclic P1 linker moieties with improved oral bioavailability. Bioorganic & Medicinal Chemistry Letters 29, 126604 (2019).
Corte, J. R. et al. Structure-Based Design of Macrocyclic Factor XIa Inhibitors: Discovery of the Macrocyclic Amide Linker. Journal of Medicinal Chemistry 60, 1060–1075 (2017).
Corte, J. R. et al. Potent, Orally Bioavailable, and Efficacious Macrocyclic Inhibitors of Factor XIa. Discovery of Pyridine-Based Macrocycles Possessing Phenylazole Carboxamide P1 Groups. Journal of Medicinal Chemistry 63, 784–803 (2020).
Corte, J. R. et al. Macrocyclic inhibitors of Factor XIa: Discovery of alkyl-substituted macrocyclic amide linkers with improved potency. Bioorganic & Medicinal Chemistry Letters 27, 3833–3839 (2017).
Dilger, A. K. et al. Discovery of Milvexian, a High-Affinity, Orally Bioavailable Inhibitor of Factor XIa in Clinical Studies for Antithrombotic Therapy. Journal of Medicinal Chemistry 65, 1770–1785 (2022).
Do, H. T., Li, H., Chreifi, G., Poulos, T. L. & Silverman, R. B. Optimization of Blood-Brain Barrier Permeability with Potent and Selective Human Neuronal Nitric Oxide Synthase Inhibitors Having a 2-Aminopyridine Scaffold. Journal of Medicinal Chemistry 62, 2690–2707 (2019).
Fang, T. et al. Orally bioavailable amine-linked macrocyclic inhibitors of factor XIa. Bioorganic & Medicinal Chemistry Letters 30, 126949 (2020).
Farand, J. et al. Selectivity switch between FAK and Pyk2: Macrocyclization of FAK inhibitors improves Pyk2 potency. Bioorganic & Medicinal Chemistry Letters 26, 5926–5930 (2016).
Giordanetto, F. & Kihlberg, J. Macrocyclic drugs and clinical candidates: what can medicinal chemists learn from their properties? Journal of Medicinal Chemistry 57, 278–295 (2014).
Gozalbes, R., Jacewicz, M., Annand, R., Tsaioun, K. & Pineda-Lucena, A. QSAR-based permeability model for drug-like compounds. Bioorganic & Medicinal Chemistry 19, 2615–2624 (2011).
Granger, B. A. & Brown, D. G. Design and synthesis of peptide-based macrocyclic cyclophilin inhibitors. Bioorganic & Medicinal Chemistry Letters 26, 5304–5307 (2016).
Hoveyda, H. et al. Macrocyclic ghrelin receptor modulators and methods of using the same. WO2008130464A1 (2008).
Hoveyda, H. et al. Macrocyclic ghrelin receptor antagonists and inverse agonists and methods of using the same. WO2011053821A1 (2011).
Hoveyda, H. R. et al. Optimization of the potency and pharmacokinetic properties of a macrocyclic ghrelin receptor agonist (Part I): Development of ulimorelin (TZP-101) from hit to clinic. Journal of Medicinal Chemistry 54, 8305–8320 (2011).
Jing, Y.-R., Zhou, W., Li, W.-L., Zhao, L.-X. & Wang, Y.-F. The synthesis of novel taxoids for oral administration. Bioorganic & Medicinal Chemistry 22, 194–203 (2014).
Kettle, J. G. et al. Potent and Selective Inhibitors of MTH1 Probe Its Role in Cancer Cell Survival. Journal of Medicinal Chemistry 59, 2346–2361 (2016).
Lampa, A. et al. Vinylated linear P2 pyrimidinyloxyphenylglycine based inhibitors of the HCV NS3/4A protease and corresponding macrocycles. Bioorganic & Medicinal Chemistry 22, 6595–6615 (2014).
Li, D. et al. Development of Macrocyclic Peptides Containing Epoxyketone with Oral Availability as Proteasome Inhibitors. Journal of Medicinal Chemistry 61, 9177–9204 (2018).
Liederer, B. M., Fuchs, T., Vander Velde, D., Siahaan, T. J. & Borchardt, R. T. Effects of amino acid chirality and the chemical linker on the cell permeation characteristics of cyclic prodrugs of opioid peptides. Journal of Medicinal Chemistry 49, 1261–1270 (2006).
Lin, X., Skolnik, S., Chen, X. & Wang, J. Attenuation of intestinal absorption by major efflux transporters: quantitative tools and strategies using a Caco-2 model. Drug Metab Dispos 39, 265–274 (2011).
Liu, Z. et al. Discovery of the Next-Generation Pan-TRK Kinase Inhibitors for the Treatment of Cancer. Journal of Medicinal Chemistry 64, 10286–10296 (2021).
Lücking, U. et al. Novel macrocyclic sulfondiimine compounds. WO2017055196A1 (2017).
Lücking, U. et al. Novel macrocyclic compounds. WO2015155197A1 (2015).
Mackman, R. L. et al. Discovery of a Potent and Orally Bioavailable Cyclophilin Inhibitor Derived from the Sanglifehrin Macrocycle. Journal of Medicinal Chemistry 61, 9473–9499 (2018).
Marsault, É. et al. Macrocyclic antagonists of the motilin receptor for treatment of gastrointestinal dysmotility disorders. EP2054429B1, (2013).
McGowan, D. et al. Finger-loop inhibitors of the HCV NS5b polymerase. Part 1: Discovery and optimization of novel 1,6- and 2,6-macrocyclic indole series. Bioorganic & Medicinal Chemistry Letters 22, 4431–4436 (2012).
Moreau, B. et al. Discovery of hepatitis C virus NS3-4A protease inhibitors with improved barrier to resistance and favorable liver distribution. Journal of Medicinal Chemistry 57, 1770–1776 (2014).
Raboisson, P. et al. Structure-activity relationship study on a novel series of cyclopentane-containing macrocyclic inhibitors of the hepatitis C virus NS3/4A protease leading to the discovery of TMC435350. Bioorganic & Medicinal Chemistry Letters 18, 4853–4858 (2008).
Raboisson, P. et al. Discovery of novel potent and selective dipeptide hepatitis C virus NS3/4A serine protease inhibitors. Bioorganic & Medicinal Chemistry Letters 18, 5095–5100 (2008).
Rosenquist, Å. et al. Discovery and development of simeprevir (TMC435), a HCV NS3/4A protease inhibitor. Journal of Medicinal Chemistry 57, 1673–1693 (2014).
Rossi Sebastiano, M. et al. Impact of Dynamically Exposed Polarity on Permeability and Solubility of Chameleonic Drugs Beyond the Rule of 5. Journal of Medicinal Chemistry 61, 4189–4202 (2018).
Vendeville, S. et al. Discovery of novel, potent and bioavailable proline-urea based macrocyclic HCV NS3/4A protease inhibitors. Bioorganic & Medicinal Chemistry Letters 18, 6189–6193 (2008).
Wang, C. et al. Discovery of D6808, a Highly Selective and Potent Macrocyclic c-Met Inhibitor for Gastric Cancer Harboring MET Gene Alteration Treatment. Journal of Medicinal Chemistry 65, 15140–15164 (2022).
Wieske, L. H. E., Atilaw, Y., Poongavanam, V., Erdélyi, M. & Kihlberg, J. Going Viral: An Investigation into the Chameleonic Behaviour of Antiviral Compounds. Chemistry 29, e202202798 (2023).
William, A. D. et al. Discovery of kinase spectrum selective macrocycle (16E)-14-methyl-20-oxa-5,7,14,26-tetraazatetracyclo[19.3.1.1(2,6).1(8,12)]heptacosa-1(25),2(26),3,5,8(27),9,11,16,21,23-decaene (SB1317/TG02), a potent inhibitor of cyclin dependent kinases (CDKs), Janus kinase 2 (JAK2), and fms-like tyrosine kinase-3 (FLT3) for the treatment of cancer. Journal of Medicinal Chemistry 55, 169–196 (2012).
William, A. D. et al. Discovery of the macrocycle (9E)-15-(2-(pyrrolidin-1-yl)ethoxy)-7,12,25-trioxa-19,21,24-triaza-tetracyclo[18.3.1.1(2,5).1(14,18)]hexacosa-1(24),2,4,9,14(26),15,17,20,22-nonaene (SB1578), a potent inhibitor of janus kinase 2/fms-like tyrosine kinase-3 (JAK2/FLT3) for the treatment of rheumatoid arthritis. Journal of Medicinal Chemistry 55, 2623–2640 (2012).
Xue, S.-T. et al. Substituted benzothiophene and benzofuran derivatives as a novel class of bone morphogenetic Protein-2 upregulators: Synthesis, anti-osteoporosis efficacies in ovariectomized rats and a zebrafish model, and ADME properties. Eur J Med Chem 200, 112465 (2020).
Yang, W. et al. Discovery of a High Affinity, Orally Bioavailable Macrocyclic FXIa Inhibitor with Antithrombotic Activity in Preclinical Species. Journal of Medicinal Chemistry 63, 7226–7242 (2020).
Himmelbauer, M. K. et al. Rational Design and Optimization of a Novel Class of Macrocyclic Apoptosis Signal-Regulating Kinase 1 Inhibitors. Journal of Medicinal Chemistry 62, 10740–10756 (2019).
Koštrun, S. et al. Macrolide Inspired Macrocycles as Modulators of the IL-17A/IL-17RA Interaction. Journal of Medicinal Chemistry 64, 8354–8383 (2021).
Lerchner, A. et al. Macrocyclic BACE-1 inhibitors acutely reduce Abeta in brain after po application. Bioorganic & Medicinal Chemistry Letters 20, 603–607 (2010).
Pennington, L. D. et al. Substituted macrocyclic compounds and related methods of treatment. WO2021108628A1 (2021).
Q, et al. Dihedral Angle-Based Sampling of Natural Product Polyketide Conformations: Application to Permeability Prediction. Journal of chemical information and modeling 56 (2016).
Xin, Z. et al. Discovery of CNS-Penetrant Apoptosis Signal-Regulating Kinase 1 (ASK1) Inhibitors. ACS Med. Chem. Lett. 11, 485–490 (2020).
Yang, A. et al. Fluorine-containing heterocyclic derivatives with macrocyclic structure and use thereof. EP4074715A1 (2022).
Basit, S., Ashraf, Z., Lee, K. & Latif, M. First macrocyclic 3rd-generation ALK inhibitor for treatment of ALK/ROS1 cancer: Clinical and designing strategy update of lorlatinib. Eur J Med Chem 134, 348–356 (2017).
Tyagi, M. et al. Toward the Design of Molecular Chameleons: Flexible Shielding of an Amide Bond Enhances Macrocycle Cell Permeability. Org Lett 20, 5737–5742 (2018).
Jiang, Y. et al. Discovery of danoprevir (ITMN-191/R7227), a highly selective and potent inhibitor of hepatitis C virus (HCV) NS3/4A protease. Journal of Medicinal Chemistry 57, 1753–1769 (2014).
Johnson, T. W. et al. Discovery of (10R)-7-amino-12-fluoro-2,10,16-trimethyl-15-oxo-10,15,16,17-tetrahydro-2H-8,4-(metheno)pyrazolo[4,3-h][2,5,11]-benzoxadiazacyclotetradecine-3-carbonitrile (PF-06463922), a macrocyclic inhibitor of anaplastic lymphoma kinase (ALK) and c-ros oncogene 1 (ROS1) with preclinical brain exposure and broad-spectrum potency against ALK-resistant mutations. Journal of Medicinal Chemistry 57, 4720–4744 (2014).
Machauer, R. et al. Macrocyclic peptidomimetic beta-secretase (BACE-1) inhibitors with activity in vivo. Bioorganic & Medicinal Chemistry Letters 19, 1366–1370 (2009).
Giroud, M. et al. 2 H-1,2,3-Triazole-Based Dipeptidyl Nitriles: Potent, Selective, and Trypanocidal Rhodesain Inhibitors by Structure-Based Design. Journal of Medicinal Chemistry 61, 3370–3388 (2018).
Acknowledgements
The work was funded by grants from the Swedish Research Council (Grant No. 2021-04747) and Olle Engkvist Stiftelse (Grants No. 214-0339 and 211-0019). The authors are grateful to Saw Simeon for assisting with the generation of the TMAP figure. We thank OpenEye scientific software and ChemAxon for providing academic licenses.
Funding
Open access funding provided by Uppsala University.
Author information
Authors and Affiliations
Contributions
Q.F.: Methodology, data curation, validation, formal analysis; D.D.C.: methodology, validation, formal analysis; J.K.: conceptualization, writing – original draft & review, project supervision, funding acquisition; V.P.: conceptualization, methodology, formal analysis, data curation, visualization, writing – original draft & review, and project supervision.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Feng, Q., De Chavez, D., Kihlberg, J. et al. A membrane permeability database for nonpeptidic macrocycles. Sci Data 12, 10 (2025). https://doi.org/10.1038/s41597-024-04302-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-024-04302-z