Abstract
Recently, imaging investigation of brain development has increasingly captured the attention of researchers and clinicians in an attempt to understand the link between the brain and behavioral changes. Although high-field MR imaging of infants is feasible, the necessary customizations have limited its accessibility, affordability, and reproducibility. Low-field MR, as an emerging solution for scrutinizing developing brain, has exhibited its unique advantages in safety, portability, and cost-effectiveness. The presented low-field infant structural MR data aims to manifest the feasibility of using low-field MR image to exam brain structural changes during early life in infants. The dataset comprises 100 T2 weighed MR images from infants with in-plane resolution of ~0.85 mm and ~6 mm slice thickness. To demonstrate the potential utility, we conducted atlas-based whole brain segmentations and volumetric quantifications to analyze brain development features in first 10 week in postnatal life. This dataset addresses the scarcity of a large, extended-span infant brain dataset that restricts the further tracking of infant brain development trajectories and the development of routine low-field MR imaging pipelines.
Similar content being viewed by others
Background & Summary
The study of the developing brain has become a critical field as early brain development lays the foundation for motor, cognitive, emotional, and social functioning throughout life1,2,3. Perturbations during this crucial yet vulnerable period can impede normal development and lead to a spectrum of psychiatric disorders with lifelong impacts4,5,6,7. Exploring these intricate changes in developing brain enhances our understanding of the mechanisms driving brain maturation, the origins of neurodevelopmental disorders, and the appropriate methods and timing for interventions, thereby advancing both brain science and infants’ healthcare.
Magnetic Resonance Imaging (MRI), as a powerful non-invasive imaging technique, is extensively utilized to visualize structural changes in the developing brain. Structural MRI has revealed significant differences between the infant brain and that of adults8. Longitudinal studies have quantified substantial volume growth in the whole brain and various brain regions, along with notable sexual dimorphism and hemispheric asymmetry9,10. Furthermore, more advanced MRI sequences disclose distinct asymmetry patterns across different brain regions11. In follow-up studies of preterm infants, lesions and reduced white matter volume are commonly observed12. Although conducting high-field MRI scans on infants is feasible, it requires specialized modifications to the coils and sequences, alongside additional monitoring to ensure imaging safety and quality13,14,15. These requirements pose challenges to patient accessibility and limit the reproductions across different institutions.
Recently, low-field MRI has gained renewed interest for imaging the developing brain with improved accessibility affordability and coziness. Low-field MRI systems with permanent magnets design offer smaller footprint, significantly lowering the prerequisite for cost, space, power supply and facilitating installation within neonatal intensive care units (NICUs)16,17. The low-field MRI also minimizes the risks related to noise, specific absorption rate (SAR), and transportation, providing a more favorable environment for infant, especially for those born preterm14,18,19. A study focusing on pediatric neuroimaging with low-field MRI has been conducted on children aged 6 weeks to 16 years and yielded volumetric results comparable to those from 3 T counterparts20. Another low-field MR deployed within NICU successfully imaged 14 neonatal inpatients aged 1–122 days with visible intracranial pathology21. The sound applications of low-field MRI in pediatric have illustrated its potential as a viable tool to advance our knowledge in neurodevelopment and enhance clinical diagnosis.
The largest MRI dataset for investigating brain development across the perinatal period is from Developing Human Connectome Project (dHCP)22,23.The dataset includes 3 T MRI scans of neonatal and fetal brains, ranging from 26 to 45 weeks post-menstrual age (PMA). For low-field MRI, efforts have been made to gather dataset to study brain injuries in newborn infants24, and comparison of clinical performance of paired low-field and high-field MR25. However, we found that the majority of available data is concentrated within the first three weeks post-delivery, with limited data beyond this period, as most infants being discharged from NICU after the first month26. This paucity of large, extended-span datasets without signs of abnormalities restricts further tracking of brain development trajectories and the development of routine pipeline of low-field MR.
To address this need, we introduce a low-field brain MR dataset of infants aged 1 to 70 days. This dataset comprises brain scans from 100 infants, with each scan containing 14 to 16 axial slices in T2-weighted contrast. All images were obtained using a 0.35 T low-field MRI system specifically designed for infant brain imaging. This study aims to explore the feasibility of using low-field MRI to elucidate the trajectory of infant brain development during early life through week-by-week volumetric analysis. Together, we outline the methods used in analysis and highlight its potential value in tracking longitudinal changes in rapidly evolving infant brains.
Methods
Demographic information
This study was approved by the Medical Ethics Committee of Children’s Hospital (Approval Letter of IRB/EC, 2023-IRB-0287-P-01) and waived the need for written informed consent from patients, as long as the data of the patient remained anonymous. Ethics approval for the publication of this dataset has been obtained. Subjects accepted 0.35 T MRI as part of differential diagnosis to rule out cerebral complication, consequently, sedation was used during imaging for clinical assessment purpose27,28. The inclusion criteria for the study were:(a) subject’s chronological age between 0 to 70 days; (b) Radiologist’s assessment indicating no visible abnormalities on MRI; (c) Radiologist’s diagnosis of mild encephalopathy of prematurity (ESS). The exclusion criteria were:(d) Presence of congenital malformations or syndromes; (e) Brain disorders attributable to various etiologies; (f) Acquired punctate or focal lesions; (g) MRI artifacts too severe to be corrected; (h) Diagnosis suggesting a genetic or hereditary disorder; (i) Absence of T2-weighted imaging scans; (j) Preterm birth.
In this dataset, 54 female and 46 male Chinese infants without visible pathological alteration were scanned between Nov 9, 2022 and Sept 28, 2023 in Children’s Hospital of Zhejiang University School of Medicine. The chronological age of included subjects ranged from 1 to 70 days (Male subjects’ chronological age 36.72 \(\pm \) 19.28, female subjects’ chronological age 33.90 \(\pm \) 20.59). The detailed demographic information is presented in Table 1.
Image acquisition
All MR images were acquired by a 0.35 Tesla open bore permanent magnet MR scanner specifically optimized for neonatal/postnatal brain imaging(NIDO, Jiangsu Lici). High resolution 2D axial T2 weighted images were acquired by an optimized fast spin echo(FSE) sequence with TR = 5,021 ms, TE = 152 ms, echo train length = 16. High resolution planes were acquired along axial axis with in-plane resolution equal to = 0.82 × 0.82 \(m{m}^{2}\) or 0.89 × 0.89 \(m{m}^{2}\), matrix size = 256 × 256, FOV = 210 × 210 \(m{m}^{2}\) or 230 × 230 \(m{m}^{2}\). Due to the increasing brain size in first ten weeks of postnatal life, 14 to 16 axial slices were scanned to ensure the coverage of infant brain, with slice thickness being 6 to 7 mm. The total scan time of T2 FSE sequence was within 2 minutes. Subjects accepted 0.35 T MRI as part of differential diagnosis to rule out cerebral complication. All neonates received 50 mg/kg oral or enema chloral hydrate 30 min before scanning. Ear protectors were used for protection. The noise level during FSE T2W sequence was below 65 dB. The specific absorption rate was closely monitored during the scan to ensure a maximum limit of 0.026 watts per kilogram.
Data processing
The original DICOM file, exported from the scanner console, was de-identified, and converted to NIFTI format in the neurological orientation, with left side of image corresponding to left side of patient body. Then, the N4 bias field correction algorithm29 was applied to normalize non-uniform intensity within the FOV.
Manual segmentation on each image was conducted by experienced radiologist to segment the brain mask, using the Medical Imaging Interaction Toolkit (MITK)30. Individual brain mask maintains the same matrix size and spatial orientation information as the T2w images.
To extract segmentations of brain structures, an atlas-based segmentation method with a three-stage multi-scale registration process in ANTs(default configuration, including rigid, affine, and SyN)31,32 was used. First, the individual upsampled image was registered to the atlas space and the transformation profile between the individual and atlas space was estimated. Next, the atlas and tissue probabilities were inverse transformed to the individual space, aligning them with the T2w images. Finally, an expectation-maximization algorithm33,34 was employed to derive whole brain segmentations from the inverse-transformed tissue probability maps. Quantitative volumetric analysis was condcuted for the selected ROIs, including: white matter, gray matter, lateral ventricle, thalamus, caudate, putamen, pallidum, hippocampus, amygadala and accumbens.
Atlas generation
To build spatiotemporal developmental brain atlas for each week, a symmetric group-wise registration method was utilized to map populations to a common space35. In each week, 7–10 subjects were used as the input to create week-wise templates.
Linear regression and statistical comparison
The volume of interested ROIs were calculated based on the whole brain segmentation. To examine brain symmetry, corresponding ROIs in the left and right hemispheres were measured separately and the linear regressions were compared. To investigate sex dimorphism in brain volume growth over time, linear regression analyses were performed for male and female subjects independently.
Data Records
The dataset is available at the Science Data Bank36. The raw data of each subjects are stored in NIFTI format in neurological orientation and organized following brain imaging data structure(BIDS)37. The manual segmentation of brain mask and derived whole brain segmentation files were stored in separate folders under /derivates. The N4 bias field corrected image and estimated bias field were stored in /derivates/N4. For each subject, a good initial transform file between individual space and M-CRIB atals38 were provided in /derivates/initial_transformed to facilitate registration. The subjects’ gender and age information was documented in participaint.tsv.
Technical Validation
Image quality assessment
To demonstrate the signal quality of the low-field MRI data, we calculated the signal-to-noise ratio and contrast-to-noise ratio in each subject. The SNR of images were measured by calculating the mean signal divided by the standard deviation of noise39 (Fig. 1(a)). ROI of size 80 × 80 was selected at the center of each image, while four noise ROIs of size 30 × 30 were selected at the corners. The resulting SNR distribution is plotted in (Fig. 1(b)), with average SNR of dataset being 25.29 ± 6.42. The CNR of images were measured by calculating the mean differences between gray matter and white matter divided by the standard deviation of noise40 (Fig. 1(c)). Masks of gray matter and white matter were conducted by the atlas-based segmentation, same four noise ROIs as those in SNR calculation were used. The resulting SNR distribution is plotted in (Fig. 1(d)), with average SNR of dataset being 2.77 ± 1.33.
Image quality assessment of dataset. (a) SNR) analysis of the 0.35 T low-field MRI data. Images were presented in neurological orientation with R and L indicate right and left side of patient body position. (b) Distribution of SNR. (c) CNR analysis of the 0.35 T low-field MRI data. (d) Distribution of CNR.
Demonstration of preprocessing and atlas-based segmentation
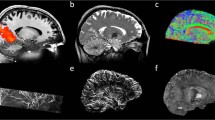
To demonstrate the 0.35 T neonatal brain MRI data have sufficient data quality to support the automatic atlas-based segmentation, we presented a step-by-step illustration of a representative patient in workflow (Fig. 2). The low-field T2w image still provided enough tissue contrast and structural information for robust registration using mutual information metric (Fig. 2(f–h)), despite the reduced SNR and image resolution due to the nature of low-field MRI scanner. To obtain the whole brain segmentation for volumetric analysis, the tissue probability maps attached with template were inverse transformed from template space to individual space respectively (Fig. 2(i–j)), followed by the expectation-maximization step to estimate the segmentations (Fig. 2(k)). All registration and inverse transform steps were visually inspected to ensure the individual space images were registered to template correctly.
Preprocesing steps and atlas based brain tissue segmentation. (a) Raw 0.35 T T2WI from a representative patient. (b) N4 bias field corrected image. (c) manual segmentation of brain parenchyma and segmented image in (d). (f–h) Registration from patient individual space to template space in three steps: rigid (h), affine (g), SyN (f). Red dashed line depicts the contour of template (e). The inverse transformed template (i) and corresponding tissue probability maps (j). (k) Whole brain segmentation estimated by EM.
Demonstration of neonatal brain tissue segmentation
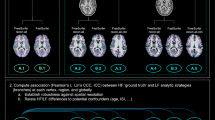
We presented representative brain tissue segmentations from 10 subjects in each age group (Fig. 3). The quality of the automatic brain tissue segmentations was visually inspected and confirmed by experienced radiologist. The results demonstrated that the data quality from the 0.35 T scanner is sufficient to support stable and convincing whole brain segmentations for volumetric analysis, even though the atlas-based segmentation methods and templates were initially developed using 1.5 T/3 T MR images.
Demonstration of neonatal brain developmental atlas
Next, we presented the week-wise spatiotemporal atlas in the first 10 weeks of postnatal life in (Fig. 4). All week-wise atlases were rigidly registered to atlas at week 1 to assure the 2D demonstration of atlases displayed the axial slices at the same orientation and height.
Demonstration of brain asymmetry validation
We further quantified volumetric asymmetry in brain regions to assess whether 0.35 T MRI provides measurements comparable to those from 3 T MRI. The selected ROIs included white matter, gray matter, lateral ventricle, hippocampus, amygdala, thalamus, caudate, putamen, pallidum, and accumbens. Consistent with prior studies9, the results showed that the left lateral ventricle was significantly larger than the right at birth, maintaining a similar growth rate during the first ten weeks of postnatal life (Fig. 5(c)). In addition to the lateral ventricle, notable left-right asymmetry was observed in the hippocampus, amygdala, caudate, and pallidum, with the left hemisphere being larger than the right (Fig. 5(d,e,g,i)).
Plots show the volumetric asymmetry of developmental trajectories in gray matter (a), white matter (b), lateral ventricle (c), hippocampus (d), amygdala (e), thalamus (f), caudate (g), putamen (h), pallidum (i), and accumbens (j). Each scatter represents the volume of selected brain region from on subject. Linear regressions and 95% confidence intervals were plotted for volume measurement of two hemispheres separately. Measurements, regressions, and confidence ranges from right hemisphere are in blue, and those from left hemisphere are in red.
Demonstration of sexual dimorphism validation
To validate whether the 0.35 T data could reflect similar gender-wise differences aligned with those reported in published studies, we performed linear regression separately for male and female groups in selected ROIs, including white matter, gray matter, lateral ventricle, hippocampus, amygdala, thalamus, caudate, putamen, pallidum, and accumbens. Consistent with findings from 3 T MRI studies9,10,41,42,43, our analysis revealed gender dimorphism in volumetric measurements from the 0.35 T data, with male brains generally appearing larger than female brains at the group level (Fig. 6).
Plots show the sex dimorphism of interested ROIs: gray matter (a), white matter (b), lateral ventricle (c), hippocampus (d), amygdala (e), thalamus (f), caudate (g), putamen (h), pallidum (i), and accumbens (j). Each scatter represents the volume of selected brain region from on subject. Linear regressions and 95% confidence intervals were plotted for volume measurement of two hemispheres separately. Measurements, regressions, and confidence ranges from female subject are in red, and those from male subjects are in blue.
Usage Notes
To access the data, users must complete the shared Data Use Agreement (DUA) form44 by Chinese Color Nest Project at https://doi.org/10.57760/sciencedb.o00133.00020, and submit it for review. Users can access both the original and derivative data at https://doi.org/10.57760/sciencedb.10690 after registration on the Science Data Bank website and completion of the DUA. We encourage researchers to incorporate this dataset into their publications, provided this article is cited, and welcome inquiries for further data sharing or collaboration opportunities.
The dataset encompasses the characteristics of low-field MRI data while retaining similarities to existing high-field datasets. Hence, it can serve as a general test dataset for a wide range of brain structural analysis methods, as well as for developing innovative methods optimized for infant brains. It could be an valuable resource for assessing the potential of 0.35 T low-field MR data in the volumetric analysis of postnatal brain growth.
Apart from brain structural analysis, one potential research application is image enhancement and image style transfer from low-field to high-field. Low-field MR images not only exhibit lower image quality, but the differing field strengths also affect tissue T1 and T2 relaxation times, leading to variations in tissue contrasts45,46. Using image enhancement and style transform methods to harmonize images obtained under different field strength could streamline radiologists’ and physicians’ interpretation of the images. One limitation of this low-field dataset is the insufficient resolution along the axial axis may compromise more detailed surface analysis from this dataset. Linear interpolation may not be the optimal solution for enhancing axial resolution; therefore, super-resolution techniques represent a promising research application for this dataset.
Head motion is another important concern in neonatal MR scans. Sedation was used for all subjects to minimize motion artifacts. However, motion artifacts could still be observed in some slices for certain subjects. For the purposes of atlas-based segmentation and volumetric quantification, the motion artifacts in this dataset are within an acceptable range and should not compromise the registration process. Researchers may choose to exclude these subjects based on whether the motion artifacts would affect their specific analysis. Conversely, the subjects with motion artifacts can be valuable for validating motion correction techniques in low-field MRI.
In this retrospective study, it is important to note that the majority of subjects were inpatients who received brain MRI scans as part of the clinical assessment. Hence, the subjects were not entirely healthy, and sedation was utilized for diagnostic rather than research purposes. Although radiologists confirmed that no visible alterations were found in the scans, researchers could exclude any subjects they deemed abnormal.
Code availability
Manual segmentation was conducted in MITK v2024.06. N4 bias filed correction, registration, and week-wise atlas is conducted by ANTs(Advanced Normalization Tools)31,32 commands N4biasfieldcorrestion, antsRegistrationSyN and antsMultivariateTemplateConstruction. Expectation maximization tool is implemented in DrawEM33,34.
References
Bear M., Connors B., Paradiso M. A. Neuroscience: exploring the brain, enhanced edition: exploring the brain. Jones & Bartlett Learning; 2020.
Weil, Z. M. & Karelina, K. Lifelong consequences of brain injuries during development: From risk to resilience. Front Neuroendocrinol. 55, 100793 (2019).
Anand, K. & Scalzo, F. M. Can adverse neonatal experiences alter brain development and subsequent behavior? Neonatology. 77(2), 69–82 (2000).
Narad, M. E. et al. Impact of secondary ADHD on long-term outcomes after early childhood traumatic brain injury. J Head Trauma Rehabil. 35(3), E271–9 (2020).
Kujabi, M. L., Petersen, J. P., Pedersen, M. V., Parner, E. T. & Henriksen, T. B. Neonatal jaundice and autism spectrum disorder: a systematic review and meta-analysis. Pediatr Res. 90(5), 934–49 (2021).
Omizzolo, C. et al. Neonatal brain abnormalities and memory and learning outcomes at 7 years in children born very preterm. Memory. 22(6), 605–15 (2013).
Garner, A. S. et al. Early Childhood Adversity, Toxic Stress, and the Role of the Pediatrician: Translating Developmental Science Into Lifelong Health. Pediatrics. 129(1), e224–31 (2012).
Sampaio R., Truwit C., Nelson C., Luciana M. Handbook of developmental cognitive neuroscience. MIT Press; 2001.
Holland, D. et al. Structural Growth Trajectories and Rates of Change in the First 3 Months of Infant Brain Development. JAMA Neurol. 71(10), 1266 (2014).
Peterson, M. R. et al. Normal childhood brain growth and a universal sex and anthropomorphic relationship to cerebrospinal fluid. J Neurosurg Pediatr. 28(4), 458–68 (2021).
Liu, T. et al. Diffusion MRI of the infant brain reveals unique asymmetry patterns during the first-half-year of development. NeuroImage. 242, 118465 (2021).
Arulkumaran, S. et al. MRI Findings at Term-Corrected Age and Neurodevelopmental Outcomes in a Large Cohort of Very Preterm Infants. Am J Neuroradiol. 41(8), 1509–16 (2020).
Barkovich, M. J., Williams, C. & Barkovich, A. J. Technical and practical tips for performing brain magnetic resonance imaging in premature neonates. Semin Perinatol. 45(7), 151468 (2021).
King, R. et al. Practical Stepwise Approach to Performing Neonatal Brain MR Imaging in the Research Setting. Children. 10(11), 1759 (2023).
Hughes, E. J. et al. A dedicated neonatal brain imaging system: A Dedicated Neonatal Brain Imaging System. Magn Reson Med. 78(2), 794–804 (2016).
Arnold, T. C., Freeman, C. W., Litt, B. & Stein, J. M. Low‐field MRI: Clinical promise and challenges. J Magn Reson Imaging. 57(1), 25–44 (2022).
Kimberly, W. T. et al. Brain imaging with portable low-field MRI. Nat Rev Bioeng. 1(9), 617–30 (2023).
Graven, S. N. & Browne, J. V. Auditory development in the fetus and infant. Newborn Infant Nurs Rev. 8(4), 187–93 (2008).
Bryan, Y. F., Templeton, T. W., Nick, T. G., Szafran, M. & Tung, A. Brain magnetic resonance imaging increases core body temperature in sedated children. Anesth Analg. 102(6), 1674–9 (2006).
Deoni, S. C. et al. Accessible pediatric neuroimaging using a low field strength MRI scanner. Neuroimage. 238, 118273 (2021).
Sien, M. E. et al. Feasibility of and experience using a portable MRI scanner in the neonatal intensive care unit. Arch Dis Child-Fetal Neonatal Ed. 108(1), 45–50 (2023).
Edwards A. D. et al. The Developing Human Connectome Project Neonatal Data Release. Front Neurosci [Internet]. 16, https://doi.org/10.3389/fnins.2022.886772 (2022).
Findlay, E., Cullen, E. & Abernethy, C. A. study to assess the length of hospital stay in pre-term babies under 34 weeks gestation. Arch Dis Child. 107(Suppl 2), A136–7 (2022).
Bao R. et al. BOston Neonatal Brain Injury Dataset for Hypoxic Ischemic Encephalopathy (BONBID-HIE): Part I. MRI and Manual Lesion Annotation. https://doi.org/10.1101/2023.06.30.546841 (2023).
Cawley, P. et al. Development of neonatal-specific sequences for portable ultralow field magnetic resonance brain imaging: a prospective, single-centre, cohort study. eClinicalMedicine. 65, 102253 (2023).
Seaton, S. E. et al. Estimating neonatal length of stay for babies born very preterm. Arch Dis Child - Fetal Neonatal Ed. 104(2), F182–6 (2018).
Tocchio S., Kline-Fath B., Kanal E., Schmithorst V. J., Panigrahy A. MRI evaluation and safety in the developing brain. In: Seminars in perinatology. Elsevier; p. 73–104 2015.
Edwards, A. D. & Arthurs, O. J. Paediatric MRI under sedation: is it necessary? What is the evidence for the alternatives? Pediatr Radiol. 41, 1353–64 (2011).
Tustison, N. J. et al. N4ITK: Improved N3 Bias Correction. IEEE Trans Med Imaging. 29(6), 1310–20 (2010).
Klemm, M. et al. MITK-OpenIGTLink for combining open-source toolkits in real-time computer-assisted interventions. Int J Comput Assist Radiol Surg. 12(3), 351–61 (2016).
Avants, B., Epstein, C., Grossman, M. & Gee, J. Symmetric diffeomorphic image registration with cross-correlation: Evaluating automated labeling of elderly and neurodegenerative brain. Med Image Anal. 12(1), 26–41 (2008).
Avants B., Tustison N., Song G. Advanced Normalization Tools: V1.0. http://hdl.handle.net/10380/3113 (2009).
Makropoulos, A. et al. Automatic Whole Brain MRI Segmentation of the Developing Neonatal Brain. IEEE Trans Med Imaging. 33(9), 1818–31 (2014).
Makropoulos, A. et al. The developing human connectome project: A minimal processing pipeline for neonatal cortical surface reconstruction. NeuroImage. 173, 88–112 (2018).
Avants, B. B. et al. A reproducible evaluation of ANTs similarity metric performance in brain image registration. NeuroImage. 54(3), 2033–44 (2011).
Sun, Z. & Huang, J. A. Low-Field MRI Dataset For Spatiotemporal Analysis of Developing Brain [Internet]. Science Data Bank https://doi.org/10.57760/sciencedb.o00133.00006 (2024).
Gorgolewski K. J. et al. The brain imaging data structure, a format for organizing and describing outputs of neuroimaging experiments. Sci Data [Internet]. 3(1), https://doi.org/10.1038/sdata.2016.44 (2016).
Alexander, B. et al. A new neonatal cortical and subcortical brain atlas: the Melbourne Children’s Regional Infant Brain (M-CRIB) atlas. NeuroImage. 147, 841–51 (2017).
Dietrich, O., Raya, J. G., Reeder, S. B., Reiser, M. F. & Schoenberg, S. O. Measurement of signal‐to‐noise ratios in MR images: Influence of multichannel coils, parallel imaging, and reconstruction filters. J Magn Reson Imaging. 26(2), 375–85 (2007).
Magnotta, V. A. & Friedman, L. Measurement of Signal-to-Noise and Contrast-to-Noise in the fBIRN Multicenter Imaging Study. J Digit Imaging. 19(2), 140–7 (2006).
Bethlehem, R. A. I. et al. Brain charts for the human lifespan. Nature. 604(7906), 525–33 (2022).
Torgerson C., et al Sex, gender diversity, and brain structure in early adolescence. Hum Brain Mapp [Internet]. 45(5), https://doi.org/10.1002/hbm.26671 (2024).
Kaczkurkin, A. N., Raznahan, A. & Satterthwaite, T. D. Sex differences in the developing brain: insights from multimodal neuroimaging. Neuropsychopharmacology. 44(1), 71–85 (2018).
Gao P., X.-N. Zuo. Chinese Color Nest Project Data Access RequestChinese Color Nest Project Data Access Request [Internet]. Science Data Bank; [cited 2024 Dec 11]. p. 803960 bytes, 2 files. https://www.scidb.cn/en/detail?dataSetId=efd2eeccd4ab41cf8e8b9cdef5a7dedb (2024).
Marques, J. P., Simonis, F. F. J. & Webb, A. G. Low‐field MRI: An MR physics perspective. J Magn Reson Imaging. 49(6), 1528–42 (2019).
Rutt, B. K. & Lee, D. H. The impact of field strength on image quality in MRI. J Magn Reson Imaging. 6(1), 57–62 (1996).
Acknowledgements
This work was partially supported by the National Key R&D Program of China (grant number 2023YFC2706400) and National Natural Science Foundation of China (grant number 62076218).
Author information
Authors and Affiliations
Contributions
Z.S.: Conceptualization, Software, Formal analysis, Writing – original draft, Writing – review & editing. J.H.: Data curation, Resources, Investigation, Writing – review & editing. L.H.: Software, Data Quality Inspection. X.M.: Data curation, Manual segmentation, Conceptualization. J.L.: Data curation, Manual segmentation. C.S.: Data curation, Data Quality Inspection. H.H.: Software, Conceptualization, Supervision, Writing – review & editing. G.Y.: Conceptualization, Supervision, Writing – review & editing, Funding acquisition, Project administration.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Sun, Z., Huang, J., Ma, X. et al. A Low-Field MRI Dataset For Spatiotemporal Analysis of Developing Brain. Sci Data 12, 109 (2025). https://doi.org/10.1038/s41597-025-04450-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-025-04450-w