Abstract
The Qaidam Basin on the northern Tibetan Plateau represents a terrestrial Mars-analog desert characterized by hyperaridity, low temperatures, intense ultraviolet radiation, and high-salinity soils. To unveil the largely unexplored genomic diversity of microbes and viruses in this extreme environment, we collected 58 soil samples from various landforms and depths for metagenomic sequencing and analysis. We reconstructed 1,773 microbial metagenome-assembled genomes (mMAGs) and 2,060 viral MAGs (vMAGs), the vast majority (>94%) of which represent novel taxa. Among these, 327 mMAGs (completeness ≥ 90% and contamination ≤ 5%) and 325 vMAGs (completeness ≥ 90%) were classified as high-quality genomes. Taxonomic classification revealed that the bacterial, archaeal, and viral phyla with the largest numbers of genomes were Actinomycetota (n = 565), Halobacteriota (n = 111), and Uroviricota (n = 836), respectively. This metagenomic and genomic dataset provides valuable reference data for advancing our understanding of the diversity and function of microbial and viral communities across global desert ecosystems. Furthermore, these data offer astrobiological insights for research on life in Mars-analog extreme environments.
Similar content being viewed by others
Backgroud & Summary
The Qaidam Basin, located in the northeast part of the Tibetan Plateau, is one of the driest (average annual precipitation < 45 mm) and highest (average elevation ~2800 m) deserts on Earth1,2. It is characterized by polyextreme conditions, including hyperaridity, low temperatures, intense solar radiation, and hypersaline soils, making it a unique and representative Mars-analog site3,4,5. The western Qaidam Basin (arid index: 0.01–0.05) is the most arid region of the desert1. Mars-like geomorphological features (e.g., yardang, gully, playa, and dune) and evaporites (e.g., sulfate and chloride) are widespread in this region due to prolonged drought and intense aeolian erosion processes3,5.
In hyperarid deserts with low biomass and biodiversity, such as the Qaidam Basin6,7, the Antarctic Desert8, and the Atacama Desert9, microbial life plays an essential role in biogeochemical cycling and biogeological processes10. The ability of microorganisms to survive and thrive in these extreme environments expands our understanding of the limits of life and provides implications for potential extraterrestrial life11. Moreover, extremophiles in desert ecosystems represent largely unexplored reservoirs of biological and genetic resources12,13. However, the vast majority of microorganisms in hyperarid deserts remain uncultured and uncharacterized, commonly referred to as “microbial dark matter”10. In contrast to microbial research, studies on desert soil viruses and their interactions with microbiomes are nascent despite viruses playing vital roles in shaping microbial community structure and function14. Previous studies have revealed that viral auxiliary metabolic genes (AMGs) related to stress tolerance may enhance host adaptation and resilience in extreme environments such as the Atacama Desert15 and the Antarctic Desert16. Due to the challenges in isolating and culturing viruses, the viral communities in hyperarid deserts remain poorly characterized17,18. Nonetheless, the rapid progress of metagenomics, along with the development of bioinformatics tools, standardized protocols, and reference databases, have enabled the recovery of metagenome-assembled genomes, facilitating the identification of uncultured microbes and viruses and offering insights into their diversity and ecological roles in desert ecosystems.
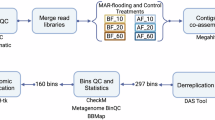
In this study, we reconstructed 1,773 mMAGs and 2,060 vMAGs from 58 soil metagenomes collected from the Qaidam Basin desert across different landforms and depths (Fig. 1a,b and Table S1). All of the 1,773 mMAGs were medium-quality with completeness > 50% and contamination < 10%. Among them, 327 were classified as high-quality genomes with completeness ≥ 90% and contamination ≤ 5% (Fig. 2a)19,20. Bin length of mMAGs ranged from 0.6 Mb to 8.8 Mb, and 74.5% (n = 1,326) had GC content ≥ 60% (Table S2). Based on GTDB R22021, 94.5% (n = 1,675) of the mMAGs represent novel taxa. These novel taxa comprise 4 orders, 29 families, 501 genera, and 1,141 species (Fig. 2b). The recovered mMAGs were assigned to 31 phyla, including 27 bacterial phyla (n = 1,630) and 4 archaeal phyla (n = 143) (Fig. 2c). The bacterial and archaeal phyla with the largest numbers of MAGs were Actinomycetota (n = 565) and Halobacteriota (n = 111), respectively (Figs. 2c, 4). Among the 2,060 vMAGs, 325 were classified as high-quality (completeness ≥ 90%), and 552 as medium-quality (50% ≤ completeness < 90%) (Fig. 3a). Viral bin length ranged from 10.2 kb to 363.0 kb (Table S3). Only 43.1% (n = 887) of the vMAGs could be taxonomically classified, among which two were assigned to unknown phyla (Fig. 3b, c). Notably, the vast majority (n = 853, 96.2%) of the classified vMAGs could not be assigned at species level. The viral phylum with the largest number of MAGs was Uroviricota (n = 836) within the realm of Duplodnaviria. Other classified vMAGs belonged to Nucleocytoviricota (n = 32), Dividoviricota (n = 7), Saleviricota (n = 5), Preplasmiviricota (n = 4), and Hofneiviricota (n = 1).
Methods
Sample collection and soil physicochemical characteristics
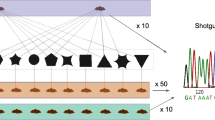
A total of 58 desert soil samples were collected from the Qaidam Basin. Sampling sites were primarily distributed in the western Qaidam Basin, the driest region in the basin. These samples comprised 27 surface soil samples (0–5 cm depth) and 31 subsurface soil samples (Table S1). Subsurface samples (n = 31) were collected from five vertical profiles, with 3–10 depth intervals per profile, to a maximum depth of 50 cm. At each sampling site, three replicated soil samples were aseptically collected with sterilized instruments and transferred into clean 50-mL centrifuge tubes. All samples were transported to the laboratory and stored at −80 °C until further processing. For each sample, three replicates were combined for further analysis. A Mettler Toledo DELTA 320 PH Meter (Mettler-Toledo, Switzerland) was used to measure soil pH in a 1:2 (w/v) soil-to-water slurry. The electrical conductivity (EC) of soil was measured in a 1:5 (w/v) soil-to-water mixture with a HACH HQ40D meter (HACH, USA). Soil water content was calculated based on the weight loss after drying at 105 °C for 24 hours. Total organic carbon (TOC) was measured by an Elemental Analyzer ECS 4024 (NC Technologies, Italy) using powdered soil pretreated with 3 M HCl to remove inorganic carbon. The metadata of soil physicochemical characteristics is included in Table S1.
Metagenomic sequencing and assembly
Metagenomic DNA was extracted from soil samples using a modified protocol as described in previous studies6,7. Briefly, DNA was extracted from approximately 30 g of soil using the PowerMax Soil DNA Isolation Kit (Qiagen, Hilden, Germany). The quality of extracted DNA was evaluated by agarose gel electrophoresis. Library preparation was conducted using the TruSeqTM DNA PCR-free library Prep Kit (Illumina, USA). The length of the inserted fragments was approximately 400 bp. Paired-end sequencing (Illumina NovaSeq 6000 platform, 2 × 150 bp) was conducted at Shanghai Majorbio Bio-pharm Technology (Majorbio, Shanghai, China). Raw metagenomic data were processed using fastp v1.0.122 to remove the adapter, short reads (<50 bp), and low-quality reads (quality scores < 20). Subsequently, clean metagenomic reads were additionally trimmed and quality-controlled using the “Read_qc” module of MetaWRAP v1.3.223. Filtered reads were de novo assembled by MEGAHIT v1.1.324 and contigs < 2000 bp were removed.
Microbial genome binning, taxonomic assignment, and phylogenetic analysis
Assembled contigs were binned using the “Binning module” of MetaWrap23 with MetaBAT2 v2.12.125, MaxBin2 v2.2.626, and CONCOCT v1.0.027. The resulting bins were refined using MetaWRAP’s “Bin_refinement” module (parameters: -c 50 -x 10). All bins were subsequently combined and dereplicated at 99% average nucleotide identity (ANI) using dRep v3.4.028 (parameters: -pa 0.9 -sa 0.99). The quality of MAGs was evaluated using CheckM v1.2.329 with the “lineage_wf” function, and MAGs with completeness > 50% and contamination < 10% were retained. The coverage of mMAGs was calculated by CoverM v0.7.030. Taxonomic classification of the 1,773 MAGs was performed using GTDB-Tk v2.4.021 with the GTDB Release R220 and the “classify_wf” function. Phylogenetic analysis was conducted using PhyloPhlAn v3.0.5831 with 400 universal marker genes (parameters: -d phylophlan–diversity low -f supermatrix_aa.cfg). During the phylogenetic analysis, 21 MAGs were excluded due to the detection of fewer than 100 universal marker genes, including genomes from Patescibacteria (n = 12), Thermoproteota (n = 4), Nanohaloarchaeota (n = 2), Proteobacteria (n = 2), and Thermoplasmatota (n = 1). The final phylogenetic tree was visualized and modified using the interactive Tree Of Life (iTOL v6)32.
Viral identification, genome binning, and classification
Viral sequences were identified from metagenomic assembly following the ViWrap v1.3.133 pipeline (parameters:–identify_method vb-vs–input_length_limit 5000). The intersection of the results of VIBRANT v1.2.134 and VirSorter2 v2.2.335 was retained to generate an accurate viral scaffold collection and viral scaffolds < 5,000 bp were removed. A total of 20,270 viral scaffolds were obtained from 58 samples. Subsequently, vRhyme v1.1.036 was used to bin vMAGs for each sample. A total of 2,060 vMAGs were recovered. The quality of viral genomes was evaluated using CheckV v1.0.137. The genus-level clusters were classified using vConTACT2 v0.11.038 (parameters:–rel-mode Diamond–pcs-mode MCL–vcs-mode ClusterONE), and species-level clusters were classified using dRep v3.4.028 (parameters: -pa 0.8 -sa 0.95 -nc 0.85). Viral taxonomy was assigned by ViWrap v1.3.133 against the NCBI RefSeq viral protein database39, the VOG HMM marker protein database40, and IMG/VR v4.1 high-quality vOTU representative proteins41.
Data Records
The metagenomic sequencing data have been deposited in NCBI Sequence Read Archive SRP56618042 under accession numbers SRR32481248-SRR32481298 (n = 51) and SRP33629043 under accession numbers SRR15827331-SRR15827333 (n = 3) and SRR24765285-SRR24765288 (n = 4). The retrieved mMAGs are available through NCBI BioProject PRJNA121334244 under accession numbers SAMN49651471-SAMN49653500. The mMAGs and vMAGs are also available at https://doi.org/10.5281/zenodo.1574321045.
Technical Validation
The metagenomic data were evaluated for quality using fastp22 and further quality-controlled using the “Read_qc” module of MetaWRAP v1.3.223. The binning of mMAGs and vMAGs was performed according to the pipelines of MetaWRAP v1.3.223 and ViWrap v1.3.133, respectively. The quality of mMAGs and vMAGs was assessed using CheckM v1.2.329 and CheckV v1.0.137, respectively. The mMAGs with completeness > 50% and contamination < 10% were retained for further analysis.
Data availability
The raw reads for 58 metagenomes are available at NCBI Sequence Read Archive SRP56618042 and SRP33629043. The retrieved mMAGs are available at NCBI BioProject PRJNA121334244. The mMAGs and vMAGs are also available at https://doi.org/10.5281/zenodo.1574321045.
Code availability
The versions and non-default parameters of all softwares are described in the Methods section. No custom code was used in this study.
References
Kong, F. et al. Dalangtan saline playa in a hyperarid region on Tibet Plateau-I: evolution and environments. Astrobiology 18, 1243–1253, https://doi.org/10.1089/ast.2018.1830 (2018).
Zheng, X., Zhang, M., Xu, Y. & Li, B. Salt lakes of China. Science Press (2002).
Anglés, A. & Li, Y. The western Qaidam Basin as a potential Martian environmental analogue: An overview. J Geophys Res-Planet 122, 856–888, https://doi.org/10.1002/2017je005293 (2017).
Shen, J. et al. Detection of biosignatures in terrestrial Mars analogs: Strategical and technical assessments. Earth Planet Phys 6, 431–450, https://doi.org/10.26464/epp2022042 (2022).
Xiao, L. et al. A new terrestrial analogue site for Mars research: The Qaidam Basin, Tibetan Plateau (NW China). Earth-Sci Rev 164, 84–101, https://doi.org/10.1016/j.earscirev.2016.11.003 (2017).
Liu, L., Chen, Y., Shen, J., Pan, Y. & Lin, W. Metabolic versatility of soil microbial communities below the rocks of the hyperarid Dalangtan Playa. Appl Environ Microbiol 89, e0107223, https://doi.org/10.1128/aem.01072-23 (2023).
Liu, L. et al. Microbial diversity and adaptive strategies in the Mars-like Qaidam Basin, North Tibetan Plateau, China. Env Microbiol Rep 14, 873–885, https://doi.org/10.1111/1758-2229.13111 (2022).
Ortiz, M. et al. Multiple energy sources and metabolic strategies sustain microbial diversity in Antarctic desert soils. P Natl Acad Sci USA 118, e2025322118, https://doi.org/10.1073/pnas.2025322118 (2021).
Crits-Christoph, A. et al. Colonization patterns of soil microbial communities in the Atacama Desert. Microbiome 1, 28, https://doi.org/10.1186/2049-2618-1-28 (2013).
Pointing, S. B. & Belnap, J. Microbial colonization and controls in dryland systems. Nat Rev Microbiol 10, 551–562, https://doi.org/10.1038/nrmicro2831 (2012).
Merino, N. et al. Living at the extremes: extremophiles and the limits of life in a planetary context. Front Microbiol 10, 780, https://doi.org/10.3389/fmicb.2019.00780 (2019).
Alsharif, W., Saad, M. M. & Hirt, H. Desert Microbes for Boosting Sustainable Agriculture in Extreme Environments. Front Microbiol 11, 1666, https://doi.org/10.3389/fmicb.2020.01666 (2020).
Silva, L. J. et al. Actinobacteria from Antarctica as a source for anticancer discovery. Sci Rep 10, 13870, https://doi.org/10.1038/s41598-020-69786-2 (2020).
Chevallereau, A., Pons, B. J., van Houte, S. & Westra, E. R. Interactions between bacterial and phage communities in natural environments. Nat Rev Microbiol 20, 49–62, https://doi.org/10.1038/s41579-021-00602-y (2022).
Hwang, Y., Rahlff, J., Schulze-Makuch, D., Schloter, M. & Probst, A. J. Diverse Viruses Carrying Genes for Microbial Extremotolerance in the Atacama Desert Hyperarid Soil. mSystems 6, https://doi.org/10.1128/mSystems.00385-21 (2021).
Ettinger, C. L. et al. Highly diverse and unknown viruses may enhance Antarctic endoliths’ adaptability. Microbiome 11, 103, https://doi.org/10.1186/s40168-023-01554-6 (2023).
Zablocki, O., Adriaenssens, E. M. & Cowan, D. Diversity and Ecology of Viruses in Hyperarid Desert Soils. Appl Environ Microbiol 82, 770–777, https://doi.org/10.1128/aem.02651-15 (2016).
Jansson, J. K. Soil viruses: Understudied agents of soil ecology. Environ Microbiol 25, 143–146, https://doi.org/10.1111/1462-2920.16258 (2023).
Bowers, R. M. et al. Minimum information about a single amplified genome (MISAG) and a metagenome-assembled genome (MIMAG) of bacteria and archaea. Nat Biotechnol 35, 725–731, https://doi.org/10.1038/nbt.3893 (2017).
Parks, D. H. et al. Recovery of nearly 8,000 metagenome-assembled genomes substantially expands the tree of life. Nat Microbiol 2, 1533–1542, https://doi.org/10.1038/s41564-017-0012-7 (2017).
Chaumeil, P. A., Mussig, A. J., Hugenholtz, P. & Parks, D. H. GTDB-Tk v2: memory friendly classification with the genome taxonomy database. Bioinformatics 38, 5315–5316, https://doi.org/10.1093/bioinformatics/btac672 (2022).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890, https://doi.org/10.1093/bioinformatics/bty560 (2018).
Uritskiy, G. V., DiRuggiero, J. & Taylor, J. MetaWRAP-a flexible pipeline for genome-resolved metagenomic data analysis. Microbiome 6, 158, https://doi.org/10.1186/s40168-018-0541-1 (2018).
Li, D., Liu, C. M., Luo, R., Sadakane, K. & Lam, T. W. MEGAHIT: an ultra-fast single-node solution for large and complex metagenomics assembly via succinct de Bruijn graph. Bioinformatics 31, 1674–1676, https://doi.org/10.1093/bioinformatics/btv033 (2015).
Kang, D. D. et al. MetaBAT 2: an adaptive binning algorithm for robust and efficient genome reconstruction from metagenome assemblies. PeerJ 7, e7359, https://doi.org/10.7717/peerj.7359 (2019).
Wu, Y.-W., Simmons, B. A. & Singer, S. W. MaxBin 2.0: an automated binning algorithm to recover genomes from multiple metagenomic datasets. Bioinformatics 32, 605–607, https://doi.org/10.1093/bioinformatics/btv638 (2016).
Alneberg, J. et al. Binning metagenomic contigs by coverage and composition. Nat Methods 11, 1144–1146, https://doi.org/10.1038/nmeth.3103 (2014).
Olm, M. R., Brown, C. T., Brooks, B. & Banfield, J. F. dRep: a tool for fast and accurate genomic comparisons that enables improved genome recovery from metagenomes through de-replication. ISME J 11, 2864–2868, https://doi.org/10.1038/ismej.2017.126 (2017).
Parks, D. H., Imelfort, M., Skennerton, C. T., Hugenholtz, P. & Tyson, G. W. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res 25, 1043–1055, https://doi.org/10.1101/gr.186072.114 (2015).
Aroney, S. T. et al. CoverM: read alignment statistics for metagenomics. Bioinformatics 41, btaf147 (2025).
Asnicar, F. et al. Precise phylogenetic analysis of microbial isolates and genomes from metagenomes using PhyloPhlAn 3.0. Nat Commun 11, 2500, https://doi.org/10.1038/s41467-020-16366-7 (2020).
Letunic, I. & Bork, P. Interactive Tree of Life (iTOL) v6: recent updates to the phylogenetic tree display and annotation tool. Nucleic Acids Res 52, W78–w82, https://doi.org/10.1093/nar/gkae268 (2024).
Zhou, Z., Martin, C., Kosmopoulos, J. C. & Anantharaman, K. ViWrap: A modular pipeline to identify, bin, classify, and predict viral-host relationships for viruses from metagenomes. iMeta 2 https://doi.org/10.1002/imt2.118 (2023).
Kieft, K., Zhou, Z. & Anantharaman, K. VIBRANT: automated recovery, annotation and curation of microbial viruses, and evaluation of viral community function from genomic sequences. Microbiome 8, 90, https://doi.org/10.1186/s40168-020-00867-0 (2020).
Guo, J. et al. VirSorter2: a multi-classifier, expert-guided approach to detect diverse DNA and RNA viruses. Microbiome 9, 37, https://doi.org/10.1186/s40168-020-00990-y (2021).
Kieft, K., Adams, A., Salamzade, R., Kalan, L. & Anantharaman, K. vRhyme enables binning of viral genomes from metagenomes. Nucleic Acids Res 50, e83, https://doi.org/10.1093/nar/gkac341 (2022).
Nayfach, S. et al. CheckV assesses the quality and completeness of metagenome-assembled viral genomes. Nat Biotechnol 39, 578–585, https://doi.org/10.1038/s41587-020-00774-7 (2021).
Bin Jang, H. et al. Taxonomic assignment of uncultivated prokaryotic virus genomes is enabled by gene-sharing networks. Nat Biotechnol 37, 632–639, https://doi.org/10.1038/s41587-019-0100-8 (2019).
O’Leary, N. A. et al. Reference sequence (RefSeq) database at NCBI: current status, taxonomic expansion, and functional annotation. Nucleic Acids Res 44, D733–745, https://doi.org/10.1093/nar/gkv1189 (2016).
Grazziotin, A. L., Koonin, E. V. & Kristensen, D. M. Prokaryotic Virus Orthologous Groups (pVOGs): a resource for comparative genomics and protein family annotation. Nucleic Acids Res 45, D491–d498, https://doi.org/10.1093/nar/gkw975 (2017).
Camargo, A. P. et al. IMG/VR v4: an expanded database of uncultivated virus genomes within a framework of extensive functional, taxonomic, and ecological metadata. Nucleic Acids Res 51, D733–d743, https://doi.org/10.1093/nar/gkac1037 (2023).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRP566180 (2025).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRP336290 (2023).
NCBI BioProject https://identifiers.org/ncbi/bioproject:PRJNA1213342 (2025).
Liu, L. & Lin, W. 1,773 mMAGs and 2,060 vMAGs from the Qaidam Basin desert. Zenodo https://doi.org/10.5281/zenodo.15743210 (2025).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (grant number T2225011). We would like to thank Runjia Ji and Yan Chen for their assistance in bioinformatic analysis and sample collection, respectively.
Author information
Authors and Affiliations
Contributions
W.L. supervised the project. L.L and W.Z. conducted sampling. L.L. and Z.W. performed DNA extraction for metagenomic sequencing. L.L. conducted geochemical analyses and bioinformatic analyses. L.L. wrote the manuscript draft with input from all co-authors. All authors reviewed and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liu, L., Wang, Z., Zhang, W. et al. Recovery of 1,773 microbial genomes and 2,060 viral genomes from the Mars-analog Qaidam Basin desert. Sci Data 12, 1795 (2025). https://doi.org/10.1038/s41597-025-06085-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-025-06085-3