Abstract
The green algae Auxenochlorella pyrenoidosa is capable of synthesizing substantial amounts of protein, containing all essential and non-essential amino acids necessary for sustainable human nutrition. A. pyrenoidosa RLXCh3 is a novel germplasm isolated from soil, with 63.65% protein content. To achieve biofortification based on RLXCh3, PacBio II/IIe sequencing, alongside Hi-C sequencing and full-length transcriptome analysis, were performed to construct an improved chromosome-level haplotypic genome assembly consisting of 12 chromosomes and a genome size of 53.07 Mbp. Scaffold N50 (4.94 Mbp) and contig N50 (1.74 Mbp) were obtained, with a total of 12,091 protein-coding genes predicted. In summary, we acquired excellent A. pyrenoidosa isolates and reported the first chromosome-scale map of A. pyrenoidosa, providing a valuable resource for in-depth commercial protein bioproduction and nutrient biofortification.
Similar content being viewed by others
Background & Summary
The global human population will reach 9.7 billion by 20501. To support this population while meeting the United Nations Sustainable Development Goals (UN-SDGs) of #2-zero hunger, #3-good health and wellbeing, and #10-reduced inequalities, dietary protein demand is forecast to increase 32–78% compared to 2017, requiring significantly higher planetary resources2. However, the protein acquisition relies heavily on traditional farming and aquaculture at present. Traditional agricultural protein production methods face three major risk challenges: high land consumption, severe environmental pollution, and zoonotic diseases. Therefore, there is an urgent need to explore a new path for high-quality protein synthesis to replace traditional “soy protein, poultry protein, and livestock protein”.
Auxenochlorella pyrenoidosa has been widely used in commercial food and feed production on a large scale as early as the 1960s, and it was listed as a novel food resource by National Health Commission of China in 20123. Its high protein content, comprehensive amino acid profile and cost-effectiveness make it an attractive source of novel protein4. Apart from this, it is rich in carotenoids, lipids, polysaccharides, vitamins, and has shown antioxidant, anti-inflammatory, anti-tumor, and anti-bacterial activities and immune-enhancing properties3. A. pyrenoidosa has fast growth rate, short growth cycle and large productivity per unit area5. It can both photosynthesize like a plant (phototrophy) and ferment at a high density like a microbe (heterotrophy), and can also be cultivated as mixotrophy. It is estimated that through phototrophic cultivation, the protein production per unit area/year of microalgae is about 20 times of that produced by soybean6, with higher production using heterotrophic cultivation.
A draft genome assembly of A. pyrenoidosa FACHB-9 was generated with the assistance of Roche 454 sequencing system by Fan et al.7, and previous study with transcriptomics have identified important genes related to starch-lipid switch process. The assembly is 56.6-Mbp in length, with 9 Mbp of scaffold N50, 1.265 Mbp of contig N50, and 10,284 of protein coding genes. Due to limited genome resources, details of molecular clues to high value metabolites and vegetative growth remain elusive, resulting in a lack of molecular fortification techniques. Here, we performed long-read PacBio II/IIe, Illumina NovaSeq PE150, and high-resolution chromosome conformation capture (Hi-C) sequencing, with the main aim to obtain a high-quality and chromosome-level genome assembly of A. pyrenoidosa. Whole-genome sequencing, assembly, and annotation of this economically important microalgae were fulfilled with a great improvement. In addition, genes involved in the starch metabolism, fatty acid and glycerolipid metabolism, carotenoid biosynthesis, carbon fixation in photosynthetic organisms, chlorophyll metabolism, selenocompound metabolism, and TOR signaling pathway were discovered. In the coming future, these valuable genomic resources will facilitate breeding of novel A. pyrenoidosa strains to obtain higher valuable metabolites.
Methods
Sample materials: isolation, purification, and identification of Chlorella
Sampling soil crust with a distinctly green surface, along with seawater, river water, or lake water from Sichuan, Guangdong, Gansu, Tibet, and so on (Table S1). For soil samples, they were soaked and suspended in equal volume of sterile water after grinding, following by culturing at 28 °C and 150 rpm for 3 h to fully mix the samples. After natural settlement for 1 h, the suspension was gradient diluted into 100, 10-1, 10-2, 10-3 concentration. For water samples, microscopic examination first to determine whether any microalgae were in it. If yes, it was centrifuged at 5,000 rpm for 5 min, and poured off supernatant to concentrate into 10 times; the sample were gradient diluted into 100, 10-1, 10-2 concentration at the same time. 200 μL of each sample were coated on BG11 solid medium containing 100 mg/L ampicillin, 50 mg/L kanamycin and 250 mg/L cephalosporin. Each group was repeated 3 times. The BG11 plates with 20 g/L glucose were cultured upside down at 25 °C for 7–15 days with 16 h light /8 h dark until single algae colonies emerged. The purified colony was cultured in BG11 liquid medium with 20 g/L glucose. Scanning and transmission electron microscopy were used for both morphological observations and microscopic characteristics. For molecular identification, the ITS region, translation elongation factor Tu (tufA), and ribulose-1,5-bisphosphate carboxylase/oxygenase large subunit (RcbLZ) were sequenced by the three sets of primers: N5: TGGTGCCAGCAGCCGCG GTA/N11R: CTCAGTAAGCTTGATCCTTCCGCAGGTTCACC; tufA-F: TGAAACAGAAMAWC GTCATTATGC/tufA-R: CCTTCNCGAATMGCRAAWCGC; RcbLZ-F: CAACCAGGTGTTCCAS CTGAAG/RcbLZ-R: CT AAAGCTGGCATGTGCCATAC8. The isolated microalgae species were preserved in BG11 medium at 4 °C.
Further gradient dilution method and monoclonal purification, over 1000 strains were collected. Further molecular sequencing of ITS region, tufA and RcbLZ, fourty-three Chlorella species were identified after filtering same strains, which were mainly divided into Auxenochlorella pyrenoidosa (10), Chlorella sorokiniana (23), Parachlorella kessleri (5), Chlorella vulgaris (1), and Chlorella variabilis (4) through phylogenetic tree analysis with the combination of ITS-tufA-RcbLZ (Fig. 1). Among them, G2-1-1, T3-1-1, and T3-3-4 were isolated from saline-alkali soil, and the Chlorella with names beginning with SZ- and gd- were isolated from seawater, which were potential for saline-alkali treatment. Apart from this, four special Scenedesmus sp. including Desmodesmus abundans (2), Coelastrella tenuitheca (1), and Scenedesmus obtusus (1), were isolated.
Detection of nutritional components of Chlorella
To obtain high-quality of A. pyrenoidosa suitable for industrialization, the nutritional ingredient and content of isolates were quantified following the National Standard for Food Safety9, the lipid content was determined by gravimetric method, and the fatty acid composition was determined by gas-mass spectrometry10. The standard commercial A. pyrenoidosa strain (FACHB-9), and representative isolates of C. sorokiniana, P. kessleri, C. vulgaris, and C. variabilis were as comparable samples. Results showed that among the test isolates, the protein content of J3 reached 63.65 g/100 g (DW), which was significantly higher than FACHB-9 (46.54 g/100 g (DW)) and other isolates (Fig. 2A, Table S2). Additionally, the total amino acid of J3 was 303.96 g/100 g (DW), which contained all the essential and non-essential amino acid. The lipid content was over 10 g/100 g (DW) in all tested Chlorella; for most Chlorella isolates, C16:0 (palmitic acid) is the main composition of fatty acid, C18:2 (linoleic acid) is the second fatty acid composition. The C18:1 (oleic acid) and C18:3 (linolenic acid) were also rich in J3. All the tested isolates contained abundant macroelement, microelement, chlorophyll, Vc, starch, and reducing sugar (Table S2). Results indicated that the isolated Chlorella species were potential resources for protein production, lipid synthesis, and nutrition enrichment. Further growth curve determination by OD680 in BG11 medium with 20 g/L glucose at 150 rpm, 28 °C in darkness indicated that the growth rate of J3 was the fastest, with the highest OD680 value and dry weight (12.64 g/L) (Fig. 2B and S1). Optical and transmission electron microscopy showed that J3 contained typical cupped chloroplasts, pyrenoid, and mitochondria as A. pyrenoidosa (Fig. 2C,D), which was deposited in China Center for Type Culture Collection (CCTCC NO: M 2022648) as the name of RLXCh3.
The protein content, growth, and morphology of Auxenochlorella pyrenoidosa RLXCh3. (A) The protein content of isolated A. pyrenoidosa species from different environments. (B) The growth curves of the top four A. pyrenoidosa species with high protein content indicated by OD680 value. J3 is the A. pyrenoidosa species with the highest protein content and fastest growth rate, which was subsequently named RLXCh3. (C) Cell morphology of RLXCh3 under an optical microscope. (D) Single cell morphology of RLXCh3 under the scanning electron microscope (SEM) and transmission electron microscope (TEM). CW: cell wall; Py: pyrenoid; Chl, chloroplast; M, mitochondria; N: nucleus; V: vacuole. Each experiment had three biological replicates. Error bars indicate standard deviation (SD). Asterisks indicate significant differences (*P ≤ 0.05; ***P ≤ 0.001).
Whole genome sequencing and genome assembly
RLXCh3 was cultured in BG11 liquid medium for 5 days, and the genomic DNA was extracted using GP1 method following the procedure of Plant Genomic DNA Kit (cat# DP305, TIANGEN BIOTECH CO. LTD), which was further detected by agarose gel electrophoresis and quantified by Qubit2.0 (ThermoFisher Scientific, Inc., USA). Long SMRT Bell libraries were constructed by SMRT bell TM Template kit (version 2.0) and then for the HiFi sequencing through PacBio Sequel II System (Pacific Biosciences, Inc., USA) with 50× sequencing depth. The sequenced HiFi reads were initially assembled to be contigs by hifiasm (version 0.14.2-r315)11, and were error corrected using Racon (version 1.4.13) and Pilon (version: 1.22). For Hi-C sequencing, the genomic DNA was fixed with formaldehyde, and digested by restriction endonuclease DPNII to build a Hi-C library. The concentration and fragment size of Hi-C library were detected by Qubit2.0 and Agilent2100, and were quantified by qPCR. The Hi-C sequencing was performed by Novaseq 6000 platform (Illumina, Inc., USA), with a PE150 sequencing read length and 100× sequencing depth. The Hi-C data was subsequently integrated to construct a high-quality de novo assembly at the chromosome level. Briefly, readfq (version 10) was used to filter the Hi-C raw reads, and then the unique mapped paired-end reads were generated by HiCUP (version 0.8.0). The LACHESIS (version 201701) was employed to link the contig sequences into chromosome-level sequences, and obtained the final genome.
To further get a high-quality genome annotation, full-length transcriptome sequencing was performed. The SMRTbellTM libraries were prepared according to the Isoform Sequencing Protocol (Iso-Seq) using the Clontech SMARTer PCR cDNA Synthesis Kit, and then sequenced on the PacBio Sequel II System. Aligning polished full-length isoforms reads to reference using GMAP with parameters-min-trimmed-coverage 0.85 and -min-identity 0.9 against Hi-C assembled genome. A routine whole-genome functional analysis strategy was applied. In brief, the coding genes was predicted by PASA12 (version 2024). Repeat sequences of the assembled A. pyrenoidosa genome were identified by employing RepeatMasker13 (Version open-4.0.5), and the tandem repeats were analyzed by Tandem Repeats Finder14 (Version 4.07b). The tRNA and rRNA were predicted by tRNAscan-SE15 (Version 1.3.1) and rRNAmmer (Version 1.2), respectively. The sRNA, snRNA, and miRNA were predicted by blasting in Rfam16 database. For gene functional analysis, the genome-wide blast was performed in GO17 (Gene Ontology), KEGG18 (Kyoto Encyclopedia of Genesand Genomes), KOG (Clusters of Orthologous Groups), NR (Non-Redundant Protein Database), TCDB19 (Transporter Classification Database), P450 and Swiss-Prot20 databases using optimized parameters (e-value < 1e-5, minimal alignment length percentage > 40%). SignalP (Version4.1) and TMHMM (Version2.0c) were used to predict secretory proteins; antiSMASH21 was for analyzing gene clusters of secondary metabolites; Diamond (v2.0.4.142) was for predicting the carbohydrate activity enzymes and pathogenicity through blasting CAZy (Carbohydrate-Active enzymes Database) and PHI (Pathogen Host Interactions Database). All the sequencing and annotation were performed in Allwegene Co., LTD.
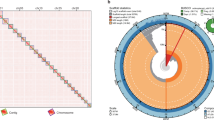
Through Hi-C genome sequencing and genome assembly, 12 chromosomes with a total length of 52.77 Mb were built, which account for about 99.45% of the whole genome assembly (53.07 Mb), individually ranging from 2.02 Mb (Chr12) to 6.59 Mb (Chr1) in length (Tables 1 and 2). Four contigs were not mapped to the chromosome. The graphical genomic maps exhibiting the genome structure and functions of RLXCh3 are presented in Fig. 3, of which the average GC content is 67% (Table 1). The final gene set is composed of 12,091 protein-coding genes (CDs), with a total of 17.41 Mb in length. In addition to CDs, the genome constituted 63 tRNA, 6 18 s rRNA, and 7 28 s rRNA (Table S3). It is worth noting that the gene internal length is 35.66 Mb, with 0.81 Mb repeated sequence, and 0.1058% of transposable elements (Table 1). A total of 2028 functional genes of the RLXCh3 genome were categorized using the Clusters of Orthologous Groups of proteins (KOG) database (Fig. S2 and Table S3). Most of the genes were related to posttranslational modification, protein turnover, chaperones (296), followed by translation, ribosomal structure and biogenesis (233), amino acid transport and metabolism (147), energy production and conversion (138), carbohydrate transport and metabolism (133), RNA processing and modification (122), signal transduction mechanisms (113), intracellular trafficking, secretion, and vesicular transport (101), lipid transport and metabolism (93). However, more than a half of genes were not predicted by the KOG database, of which specific functions need to be further verified. As a supplementary analysis, 2761 functional genes were annotated to 369 different KEGG pathway maps, the majority of which were associated with biosynthesis of secondary metabolites (349), biosynthesis of antibiotics (244), microbial metabolism in diverse environments (222), carbon metabolism (146), biosynthesis of amino acids (126), and ribosome (112) (Fig. S3 and Table S5). In addition, 7244, 2876, 1974, 7244, and 430 genes were annotated through GO, Swiss-Prot, NR, Pfam, and TCDB databases (Fig. S4 and Table S6-8). As a superfamily of ferrous heme-mercaptan proteins, 8 cytochrome P450 (CYP450) were discovered in the genome, while 4 of them are undeterminded (Fig. S5). At the same time, 181 CAZymes were identified, including 33 carbohydrate-binding modules, 4 carbohydrate esterases, 71 glycoside hydrolases, 64 glycosyl transferases, and 9 auxiliary activities, which are important for the complex carbohydrate metabolism (Fig. S6 and Table S9). However, no gene clusters of secondary metabolites were predicted by AntiSMASH.
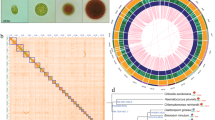
The genome of Auxenochlorella pyrenoidosa RLXCh3. (A) The heatmap view of Hi-C result. (B) The circos view of RLXCh3 genome. (C) Phylogeny analysis based on genome sequence of ten representative microalgae species, indicating that RLXCh3 was most closely related to and formed a clade with A. pyrenoidosa.
Data Records
The RLXCh3 was preserved at China Center for Type Culture Collection (CCTCC) under isolate number No. M2022648. The whole genome sequence of A. pyrenoidosa strain RLXCh3 from Hi-C library has been deposited in GenBank under the accession number of SRR3516822122, from PacBio HiFi library deposited in the Sequence Read Archive (SRA) was SRR3589591523. The raw data of full-length transcriptome sequencing data was deposited in the Genome Sequence Archive24 in National Genomics Data Center25 under the accession number of CRA03307826 that are publicly accessible at https://ngdc.cncb.ac.cn/gsa/s/BZqp2IiO. The accession number of assemble genome is GCA_047663505.127. For the 12 chromosomes, the GenBank accession number are CM105675.1-CM105686.1. The annotation data and protein sequences have been deposited at Figshare28.
Technical Validation
The quality of the extracted DNA was assessed using agarose gel electrophoresis, with DNA spectrophotometer ratios (260/280) greater than 1.8. The purified RNA quality was verified using the Nanodrop ND-8000 spectrophotometer (RIN > 8.0; Thermo Scientific). To ensure reads reliability and minimize artificial bias, raw reads were initially processed through an in-house quality control (QC) pipeline implemented in C scripts. QC criteria were as follows: (1) Removing reads with ≥10% unidentified nucleotides (N); (2) Removing reads with >50% bases having Phred quality score < 5. The longest scaffold of 6.59 Mb, contig N50 size of 1.74 Mb, scaffold N50 size of 4.94 Mb, and 67 contigs assembled to 12 chromosomes validated the high quality and reliability of our genome assembly.
Code availability
All software and pipelines utilized in this study adhered rigorously to the manuals and protocols of the established bioinformatic tools. The specific versions of the software are detailed in the Methods section. If parameters were not specified, default settings were used. No custom code was used in this study.
Data availability
References
United Nations Department of Economic and Social Affairs. Population Division (Ed.). World Population 9, 1–54 (2022).
Williamson, E., Ross, I. L., Wall, B. T. & Hankamer, B. Microalgae: Potential novel protein for sustainable human nutrition. Trends Plant Sci. 29, 370–382 (2024).
Torres-Tiji, Y., Fields, F. J. & Mayfield, S. P. Microalgae as a future food source. Biotechnol Adv. 41, 107536 (2020).
Becker, E. W. Micro-algae as a source of protein. Biotechnol Adv. 25, 207–210 (2007).
Krienitz, L., Huss, V. A. & Bock, C. Chlorella: 125 years of the green survivalist. Trends Plant Sci. 20, 67–69 (2015).
Barbosa, M. J., Janssen, M., Sudfeld, C., D’Adamo, S. & Wijffels, R. H. Hypes, hopes, and the way forward for microalgal biotechnology. Trends Biotechnol. 41, 452–471 (2023).
Fan, J. et al. Genomic foundation of starch-to-lipid switch in oleaginous Chlorella spp. Plant Physiol. 169, 2444–2461 (2015).
Zou, S. et al. Combining and comparing coalescent, distance and character-based approaches for barcoding microalgaes: A test with Chlorella-like species (Chlorophyta). PLoS One 11, e0153833 (2016).
Zhu, T. et al. Transcriptomic and metabolomic analysis reveal the effects of light quality on the growth and lipid biosynthesis in Chlorella pyrenoidosa. Biomolecules. 14, 1144 (2024).
Chen, L., Zhang, L. & Liu, T. Concurrent production of carotenoids and lipid by a filamentous microalga Trentepohlia arborum. Bioresource Technol. 214, 567–573 (2016).
Cheng, H., Concepcion, G. T., Feng, X., Zhang, H. & Li, H. Haplotype-resolved de novo assembly using phased assembly graphs with hifiasm. Nat Methods. 18, 170–175 (2021).
Do, V. H. et al. Pasa: leveraging population pangenome graph to scaffold prokaryote genome assemblies. Nucleic Acids Res. 52, e15 (2024).
Chen, N. Using RepeatMasker to identify repetitive elements in genomic sequences. Current protocols in bioinformatics Chapter 4, Unit 4.10 (2004).
Benson, G. Tandem repeats finder: a program to analyze DNA sequences. Nucleic Acids Res. 27, 573–580 (1999).
Chan, P. P., Lin, B. Y., Mak, A. J. & Lowe, T. M. tRNAscan-SE 2.0: improved detection and functional classification of transfer RNA genes. Nucleic Acids Res. 49, 9077–9096 (2021).
Kalvari, I. et al. Rfam 14: expanded coverage of metagenomic, viral and microRNA families. Nucleic Acids Res. 49, D192–D200 (2021).
Gene Ontology Consortium. The Gene Ontology Resource: 20 years and still GOing strong. Nucleic Acids Res. 47, D330–D338 (2019).
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Saier, M. H. et al. The Transporter Classification Database (TCDB): 2021 update. Nucleic Acids Res. 49, D461–D467 (2021).
Boeckmann, B. et al. The SWISS-PROT protein knowledgebase and its supplement TrEMBL in 2003. Nucleic Acids Res. 31, 365–70 (2003).
Blin, K. et al. antiSMASH 7.0: new and improved predictions for detection, regulation, chemical structures and visualisation. Nucleic Acids Res. 51, W46–W50 (2023).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRP613830 (2025).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRP637498 (2025).
Zhang, S. et al. The GSA Family in 2025: A broadened sharing platform for multi-omics and multimodal data. Genom Proteom Bioinf. 23, qzaf072 (2025).
CNCB-NGDC Members and Partners. Database resources of the National Genomics Data Center, China National Center for Bioinformation in 2025. Nucleic Acids Res. 53, D30–D44 (2025).
CNCB Genome Sequence Archive https://ngdc.cncb.ac.cn/gsa/browse/CRA033078 (2025).
Luo, X. M. GenBank https://identifiers.org/ncbi/insdc.gca:GCA_047663505.1 (2025).
Luo, X. M. Auxenochlorella pyrenoidosa genome annotation and protein sequences. figshare https://doi.org/10.6084/m9.figshare.28498355 (2025).
Acknowledgements
This work was supported by the National Key R and D Program of China (2025YFE0199700, 2023YFE0199400), Agricultural Science and Technology Innovation Program (ASTIP) (No. Y2024QC33), Chengdu Science and Technology Program (No. 2024-YF06-00116-HZ, 2024-YF06-00124-HZ), the Science and Technology Innovation Project of the Chinese Academy of Agricultural Sciences (No. 34-IUA-02).
Author information
Authors and Affiliations
Contributions
X.L.: Funding acquisition, Conceptualization, Methodology/Study design, Resources, Investigation, Formal analysis, Writing-original draft. H.S.: Formal analysis, Data curation, Investigation, Writing-original draft. G.G.: Formal analysis, Data curation, Investigation, Writing-original draft. M.R.: Funding acquisition, Data curation, Conceptualization, Supervision, Writing-review and editing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Luo, X., Su, H., Guan, G. et al. Chromosome-level genome assembly for an edible protein microalgae Auxenochlorella pyrenoidosa. Sci Data 13, 60 (2026). https://doi.org/10.1038/s41597-025-06358-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-025-06358-x