Abstract
Autotrophic CO₂ fixation is a fundamental metabolic process that enables microorganisms to inhabit carbon-limited environments. Multiple pathways mediate this process, with variants distributed across diverse taxa and some genes shared among pathways, making their identification from genomic data challenging. Here, we present a curated resource comprising pathway-specific KEGG Orthology marker genes and a lightweight, rule-based tool AutoFixMark for predicting the presence of seven known CO₂ fixation pathways in microbial genomes. To support marker gene identification and benchmarking, we compiled two reference datasets: (i) 347 manually curated chemolithoautotrophic genomes from 16 phyla, and (ii) a set of 15 well-characterized chemolithoautotrophic genomes used for defining pathway-specific marker genes. Using these marker genes, we developed AutoFixMark and evaluated its performance against two existing tools, METABOLIC and gapseq. Benchmarking results show that AutoFixMark achieves high precision and recall, particularly for pathways that are underrepresented in current tools. All curated gene sets, prediction rules, the AutoFixMark program, and benchmark datasets are publicly available, providing valuable resources for assessing autotrophic carbon fixation potential in microbial genomes.
Similar content being viewed by others
Background & Summary
Understanding carbon fixation in microbes is essential for elucidating global carbon cycling, developing sustainable biotechnological applications, and interpreting microbial contributions to diverse ecosystems. Autotrophic microorganisms employ diverse biochemical strategies to fix carbon. To date, seven distinct natural CO₂ fixation pathways have been characterized: the Calvin–Benson–Bassham (CBB) cycle, the reductive tricarboxylic acid (rTCA) cycle, the Wood–Ljungdahl (WL) pathway, the 3-hydroxypropionate (3HP) bicycle, the 3-hydroxypropionate/4-hydroxybutyrate (3HP/4HB) cycle, the dicarboxylate/4-hydroxybutyrate (DC/4HB) cycle, and the reductive glycine (rGly) pathway1. While the taxonomic distribution of these pathways has been summarized at higher ranks (e.g., phylum or class)2, a systematic assessment of their presence at the strain level remains largely unexplored. These CO₂ fixation pathways are diverse, with pathway variants occurring in different taxa and certain genes shared across multiple pathways2. Consequently, distinguishing these pathways based on gene content alone is often difficult, underscoring the need for dedicated tools capable of inferring CO₂ fixation pathways directly from genomic data. Existing tools such as METABOLIC and gapseq offer general metabolic pathway predictions3,4, yet their accuracy in detecting specific CO₂ fixation pathways has not been comprehensively evaluated. Moreover, these tools rely on KEGG Orthology (KO) and other reference databases for inference but lack clearly defined marker enzymes that can robustly distinguish among the seven known pathways. A key challenge arises from the evolutionary diversity of enzymes: even within a single pathway, phylogenetically distinct enzymes may catalyze equivalent reactions5, complicating straightforward rule-based predictions. In addition, the specific marker enzymes required to resolve pathway presence with high confidence have not been systematically defined or curated.
To address these limitations, we first compiled a set of 15 well-characterized chemolithoautotrophic genomes to define pathway-specific marker enzymes and their corresponding KO identifiers (IDs) for all seven CO₂ fixation pathways. Based on this curated marker set, we developed AutoFixMark, a lightweight, rule-based tool that predicts the presence or absence of each pathway from a genome’s KO profile. To evaluate the performance of AutoFixMark, we constructed a separate benchmark dataset comprising 347 genomes from 16 phyla, each manually annotated with literature-based evidence of CO₂ fixation pathway presence. We then compared AutoFixMark’s prediction accuracy with that of two existing tools, METABOLIC and gapseq. Our results indicate that AutoFixMark achieves high sensitivity and specificity across all seven CO₂ fixation pathways. By enabling accurate and interpretable predictions, AutoFixMark facilitates deeper insights into the functional potential of autotrophic microorganisms in diverse environments, including those investigated through metagenomic approaches.
Methods
Definition of pathway-specific marker KOs for AutoFixMark
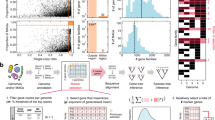
An overview of the study workflow is shown in Fig. 1. To define pathway-specific marker genes for the seven known natural CO₂ fixation pathways, we first reviewed the biochemical architecture of each pathway based on previously published literature, including pathway-focused reviews and primary studies describing enzymatic components indicative of specific pathway types1,6,7,8,9. While these pathways have been comprehensively described in those reviews, we do not repeat the mechanistic details here. CO₂ fixation pathways often exhibit multiple enzymatic variants that differ in specific reactions, substrates, or cofactors. These variants, observed across diverse microbial taxa, may utilize evolutionarily distinct enzymes that are assigned different KO IDs. We first compiled the genes and corresponding KO IDs involved in each CO₂ fixation pathway using genome information from representative autotrophic microbes. We then selected marker enzymes to identify each pathway. We systematically examined both canonical and non-canonical variants and compiled enzyme-coding genes for each, based on representative strains listed in Table 1. Variants lacking gene-level evidence were excluded from consideration. For each selected marker enzyme, KO IDs were assigned using KofamScan with default parameters10 on the RefSeq-derived protein sequences of representative genomes. Hits with scores exceeding the predefined adaptive threshold for each KOfam profile were assigned to the corresponding KO ID. If no hit exceeded the threshold for a given query, the KO ID of the top-scoring hit was assigned instead. When multiple KO IDs exceeded the threshold but clearly included enzymes unrelated to the target pathway, those KO IDs were excluded from further analysis. To account for the functional and evolutionary diversity of enzymes, we implemented a flexible rule-based framework to define the presence of each CO₂ fixation pathway. Three logical rules were used to accommodate pathway-specific features. The “one_of” rule was applied when multiple alternative enzymes (assigned to different KO IDs) could catalyze the same biochemical reaction, indicating that the presence of any one KO is sufficient. The “all_of” rule was used for reactions requiring a multi-subunit enzyme complex, requiring all listed KO IDs to be present. For the rGly pathway, where the essentiality of individual glycine cleavage system subunits remains uncertain, we introduced an “at_least” rule to require a minimum number of subunit KO IDs for prediction. These marker definitions and logical rules are encoded in a machine-readable JSON file, which is used as a core reference by AutoFixMark during prediction. Given a KO profile of a genome, AutoFixMark determines the presence or absence of each of the seven CO₂ fixation pathways based on these predefined rules. The JSON file is publicly available within the AutoFixMark source code repository11 and summarized in Table 2.
Marker enzyme genes for the Calvin–Benson–Bassham (CBB) cycle
For the Calvin–Benson–Bassham (CBB) cycle, we selected three marker enzyme genes: ribulose-1,5-bisphosphate carboxylase/oxygenase (RubisCO) large subunit [K01601] and phosphoribulokinase (PRK) [K00855], which synthesizes ribulose-1,5-bisphosphate12, and ribulose-phosphate 3-epimerase (RPE) [K01783]. Although RubisCO typically functions as a hetero-oligomer that includes a small subunit [K01602], some organisms are known to possess functional CBB cycles in the absence of the small subunit13; therefore, K01602 was not included as a required marker. Both PRK and the RubisCO large subunit have also been identified in certain archaeal species14, where they are hypothesized to function as part of the reductive hexulose-phosphate (RHP) pathway. The pathway was considered as a primitive prototype of the CBB cycle15. To distinguish between the CBB cycle and the RHP pathway, we included RPE, one of the enzymes that regenerates ribulose 5-phosphate.
Marker enzyme genes for the reductive tricarboxylic acid (rTCA) cycle
The reductive tricarboxylic acid (rTCA) cycle can be broadly divided into two mechanistic types: an asymmetric type (rTCA-I) and a symmetric type (rTCA-II), each characterized by distinct enzymatic steps16. In AutoFixMark, we considered the presence of either type to be indicative of the rTCA pathway. For rTCA-I, we selected ATP citrate lyase [K15230 and K15231] as the marker enzyme. For rTCA-II, we used a combination of citrate-CoA ligase [K15233 and K15232] and citryl-CoA lyase [K15234] as marker enzymes7. It is important to note that in some organisms, the conversion of citrate to acetyl-CoA is carried out by citrate synthase (an enzyme shared with the oxidative TCA cycle) resulting in a bidirectional or reversible TCA cycle17,18. In such cases, the genome lacks unique marker enzymes that distinguish rTCA from oxidative TCA, and therefore pathway prediction based on marker genes is not feasible. Such discrimination is beyond the scope of the current study and is left for future work.
Marker enzyme genes for the Wood–Ljungdahl (WL) pathway
The Wood–Ljungdahl (WL) pathway exists in two phylogenetically distinct forms: the bacterial type (WL-I), represented by acetogenic bacteria, and the archaeal type (WL-II), represented by methanogenic archaea7. In both cases, the key marker enzymes are anaerobic carbon monoxide dehydrogenase (CODH) and CO-methylating acetyl-CoA synthase (ACS)7, although the assigned KO IDs differ by lineage. For WL-I, the marker KO IDs are CODH [K00198] and ACS [K14138]. For WL-II, the corresponding marker set includes CODH [K00192, K00195] and ACS [K00193, K00194, K00197]. Importantly, the WL pathway is reversible, and even the complete presence of marker enzymes and associated genes does not necessarily indicate the capability for autotrophic growth19. In addition, hybrid variants that combine bacterial- and archaeal-type CODH/ACS subunits have been reported20, including our previous study5. These cases may require the development of separate classification rules in future versions of the tool.
Marker enzyme genes for the 3-hydroxypropionate (3HP) bicycle
The key enzyme for the 3-hydroxypropionate (3HP) bicycle is malonyl-CoA reductase/3-hydroxypropionate dehydrogenase [K14468], which catalyzes the formation of 3-hydroxypropionate from malonyl-CoA21. To differentiate this pathway from the 3HP/4HB cycle, we also included succinyl-CoA:L-malate CoA-transferase [K14471 and K14472] as a marker enzyme, as it is absent in the 3HP/4HB pathway.
Marker enzyme genes for the 3-hydroxypropionate/4-hydroxybutyrate (3HP/4HB) cycle
As with the 3HP bicycle, the first marker enzyme for the 3-hydroxypropionate/4-hydroxybutyrate (3HP/4HB) cycle is malonyl-CoA reductase [K15017], though it is associated with a different KO ID than that used for the 3HP bicycle. To distinguish this pathway from the DC/4HB cycle, we also included 3-hydroxypropionate dehydrogenase / malonic semialdehyde reductase [K15039], which catalyzes the subsequent step.
Marker enzyme genes for the dicarboxylate/4-hydroxybutyrate (DC/4HB) cycle
The dicarboxylate/4-hydroxybutyrate (DC/4HB) cycle shares partial routes with both the rTCA cycle and the 3HP/4HB cycle6, making it difficult to detect without a broader set of marker enzymes. We selected 4-hydroxybutyrate-CoA ligase [K18861 or K14467] and 4-hydroxybutanoyl-CoA dehydratase [K14534] as marker enzymes in the 4HB part. To differentiate this pathway from the 3HP/4HB cycle, we also included phosphoenolpyruvate carboxylase [K01595] and fumarate hydratase [K01676, K01677, or K01678] as marker enzymes. However, we note that certain strains assigned to the 3HP/4HB cycle possess most of the gene set for the DC/4HB cycle, making differentiation difficult based solely on gene presence. Therefore, further evidence from metabolic studies or experimental validation may be required to distinguish the functional use of these pathways.
Marker enzyme genes for the reductive glycine (rGly) pathway
The reductive glycine (rGly) pathway is a recently discovered CO₂ fixation pathway, first described in 202022, and its phylogenetic distribution remains largely unknown. According to the original study, the glycine reductase complex in which comprising seven subunits [K10671, K10672, K00384, K21577, K21576, K03671, K10670], was identified as a unique and characteristic feature of the pathway. In addition, the glycine cleavage system in which involving four enzymes [K00283, K00282, K02437, K00605], plays a central role in glycine metabolism. Therefore, both complexes were designated as marker enzyme sets. Following the detection criteria described in the original publication, we considered the presence of at least 5 out of 7 glycine reductase subunits and at least 3 out of 4 glycine cleavage system components as markers for the pathway. The conversion of glycine to pyruvate in this pathway can proceed either via serine or via acetyl-CoA. As the latter route was proposed to be dominant under autotrophic growth conditions, we selected enzymes specific to the acetyl-CoA–mediated route as additional markers.
Benchmark genome dataset construction
To construct a benchmark dataset for CO₂ fixation pathway prediction, we first conducted a PubMed literature search using the keyword “CO₂ fixation” and similar words to identify candidate microbial species. Retrieved literatures were manually screened to extract microbial species names and their associations with CO₂ fixation activity. Since our focus was on chemolithoautotrophs, photosynthetic organisms were excluded. However, a subset of phototrophic species harboring the 3HP bicycle was retained to ensure representation of this pathway21. For each identified species, we searched for available genome sequences in the NCBI RefSeq and GenBank databases23. Genome selection followed a prioritized scheme: (i) type strains in RefSeq, (ii) non-type strains in RefSeq, and (iii) strains in GenBank. This process yielded an initial set of 460 genomes. We subsequently excluded organisms that exhibited only anaplerotic CO₂ fixation or lacked clear evidence of autotrophic growth24. The final curated dataset comprised 347 genomes representing confirmed chemolithoautotrophs25. Pathway presence for each species was manually identified through literature review. The level of supporting evidence varied across species and included biochemical enzyme assays, gene expression data, and inferences based on partial gene sets. In cases where direct evidence was unavailable, pathway presence was inferred based on closely related species with experimentally validated pathway presence.
Benchmarking of pathway prediction tools
We evaluated the performance of three tools (i.e., AutoFixMark, METABOLIC, and gapseq) in predicting CO₂ fixation pathways across the 347 benchmark genomes. To eliminate differences arising from protein-coding gene prediction, we used the RefSeq-annotated protein sequences from each genome in all three tools. For AutoFixMark, KO IDs were assigned to each protein sequence using the KofamScan with default parameters. Hits with scores exceeding the predefined adaptive threshold for each KOfam profile were assigned to the corresponding KO ID. If no hit exceeded the threshold for a given query, the KO ID of the top-scoring hit was assigned instead. METABOLIC version 4.0 was performed using default parameters. gapseq version 1.4 was executed with a custom pathway list (LWP-GS, CALVIN-PWY, P23-PWY, P42-PWY, PWY-5392, CODH-PWY, PWY-778, PWY-8303, PWY-5743, and PWY-5789). For each genome, the presence or absence of each pathway was predicted by all three tools and summarized into a binary presence/absence matrix. The predictions were then compared against the manually curated reference dataset to evaluate accuracy. For each pathway, we computed precision, recall, and F1 statistics based on the number of true positive, false positive, and false negative predictions across all genomes.
Data Records
The rule-based definitions of the marker KO ID combinations for the seven CO₂ fixation pathways are provided in a JSON file (kegg_key_enzymes.json), which is publicly available in the definitions folder of the AutoFixMark GitHub repository11. The rule-based pathway prediction tool AutoFixMark is implemented in Python and can be run as a standalone script without requiring external library dependencies. AutoFixMark Python script (predict_pathways.py) is distributed under the MIT license and is available in the app folder of the repository. Since AutoFixMark requires a genome-derived KO list as input, a format conversion Python script (kofamscan_parser.py) is also provided in the app folder to convert the TSV output from the KO assignment software KofamScan10 into the simple KO list.
The benchmark dataset consists of 347 genomes representing confirmed CO₂-fixing microbes and is available via Zenodo25. The dataset includes: (i) an Excel file containing the genome ID (INSDC GCA/GCF ID), species name, phylum name, CO₂ fixation pathway, rationale for pathway identification, and references (PubMed ID or DOI) for both pathway identification and experimental evidence of CO₂ fixation; and (ii) protein sequence FASTA files for the 347 genomes, with each file containing all protein sequences of a genome. The Excel file also lists 12 reference genomes of CO₂ fixation pathways; however, these were not included in the benchmark analysis and therefore their protein sequence FASTA files are not provided.
Data Overview
The pathway-specific marker KO IDs and AutoFixMark tool description
In AutoFixMark version 1, pathway-specific KO IDs were defined for each of the seven CO₂ fixation pathways as follows: the CBB cycle: 3 KOs; the rTCA cycle: 5 KOs; the WL pathway: 7 KOs; the 3HP bicycle: 3 KOs; the 3HP/4HB cycle: 7 KOs; the DC/4HB cycle: 7 KOs; and the rGly pathway: 11 KOs. The rule-based definitions of these KO IDs combinations for each pathway are implemented in a JSON format file and are publicly available11. AutoFixMark is implemented in Python and can be installed as a standalone script without the need for external library dependencies. AutoFixMark requires a genome-derived KO list as input. KofamScan TSV outputs can be converted into the appropriate KO list format using our conversion Python program.
Benchmark genome dataset description
The benchmark genome dataset comprises 347 genomes representing confirmed CO₂-fixing microbes25. These genomes span 16 bacterial and archaeal phyla. The number of genomes predicted to possess each of the seven CO₂ fixation pathways is summarized in Fig. 2. Notably, due to the limited number of sequenced genomes and available pathway annotations for certain pathways, particularly the 3HP bicycle, the DC/4HB cycle, and the rGly pathway, the benchmark performance results for these pathways should be interpreted with caution. The total count of 352 pathways across 347 genomes reflects the presence of strains harboring multiple CO₂ fixation pathways. Specifically, three strains in the phylum Bacillota possess both the CBB and WL pathways, while two strains in the phylum Bacillota exhibit a combination of the WL and rGly pathways.
Technical Validation
We evaluated the predictive performance of AutoFixMark against two existing tools, METABOLIC and gapseq, using a benchmark dataset of 347 manually curated chemolithoautotrophic genomes. METABOLIC predicts metabolic pathways by identifying homologous proteins of marker genes across KEGG modules or custom-defined gene collections through HMMER-based searches against curated HMM profiles and assigning KO IDs based on sequence similarity3. gapseq annotates metabolic genes through BLAST- and HMM-based similarity searches and gapseq’s pathway inference relies on gene evidence scoring, network-consistency checks, and gap-filling heuristics to account for incomplete genomes or missing annotations4. In contrast, AutoFixMark takes as input the KO composition of a genome’s protein set, typically generated using KofamScan, and predicts the presence of the seven known CO₂ fixation pathways by evaluating explicitly defined combinations of pathway-specific marker KO IDs. The precision, recall, and F1 scores for each of the seven known CO₂ fixation pathways are summarized in Fig. 3. For the well-characterized pathways (i.e., the CBB cycle, the rTCA cycle, and the WL pathway), all three tools achieved high prediction accuracy. For the 3HP bicycle, pathway annotation remains extremely limited; only one genome (Roseiflexus castenholzii) in the benchmark set is annotated with this pathway, precluding meaningful precision or recall analysis at this stage. For the less-studied or more recently discovered pathways (i.e., 3HP/4HB, DC/4HB, and rGly pathway), AutoFixMark outperformed existing tools, some of which do not include these pathways in their prediction scope. In particular, AutoFixMark was able to make predictions for pathways entirely unsupported by METABOLIC or gapseq. The DC/4HB pathway was predicted exclusively by AutoFixMark, as the other two tools do not support prediction of this pathway. In addition, METABOLIC lacks a prediction function for the rGly pathway. The lower prediction accuracy of METABOLIC and gapseq for the 3HP, 3HP/4HB, and rGly pathways (in the case of gapseq) is likely due to insufficient marker gene definitions in their reference databases. The rGly pathway, first described in 202022, remains poorly characterized in terms of its taxonomic distribution. While AutoFixMark showed relatively low precision for rGly, it is possible that these predictions reflect true but as-yet unconfirmed pathway presence. Further experimental validation and the accumulation of supporting literature are needed to clarify the biological relevance of these predictions.
AutoFixMark enables accurate and interpretable prediction of CO₂ fixation potential from microbial genomes, including metagenome-assembled genomes. Its high accuracy primarily stems from the carefully curated, pathway-specific marker gene sets and the rule-based framework that allows precise discrimination among CO₂ fixation pathways that share homologous enzymes. By facilitating the identification of autotrophic microbes in carbon-limited environments, it contributes to a deeper understanding of microbial carbon metabolism and the ecological roles of autotrophs across diverse ecosystems. Importantly, AutoFixMark is intended as a first-pass screening tool to identify organisms with genetic potential for autotrophic CO₂ fixation, not to infer active metabolism or growth phenotype. The presence of marker genes alone does not guarantee pathway expression or functional activity, especially given that many CO₂ fixation pathways are biochemically reversible and may operate in either direction depending on the organism’s metabolic and energetic context. Accordingly, AutoFixMark does not infer flux directionality. As new studies continue to uncover alternative enzymes or novel variants of known pathways, we plan to update and expand the marker gene definitions in AutoFixMark to reflect the evolving understanding of microbial autotrophy.
Data availability
The rule-based definitions of KO marker combinations and the AutoFixMark Python program are available at the AutoFixMark GitHub repository (https://github.com/h-mori/AutoFixMark). The benchmark genome dataset is available from Zenodo (https://doi.org/10.5281/zenodo.16956127).
Code availability
The AutoFixMark program and the marker gene definition JSON file are publicly available within the AutoFixMark source code repository (https://github.com/h-mori/AutoFixMark).
References
Santos Correa, S., Schultz, J., Lauersen, K. J. & Soares Rosado, A. Natural carbon fixation and advances in synthetic engineering for redesigning and creating new fixation pathways. J Adv Res. 47, 75–92, https://doi.org/10.1016/j.jare.2022.07.011 (2023).
Garritano, A. N., Song, W. & Thomas, T. Carbon fixation pathways across the bacterial and archaeal tree of life. PNAS Nexus. 1(5), pgac226, https://doi.org/10.1093/pnasnexus/pgac226 (2022).
Zhou, Z. et al. METABOLIC: high-throughput profiling of microbial genomes for functional traits, metabolism, biogeochemistry, and community-scale functional networks. Microbiome. 10(1), 33, https://doi.org/10.1186/s40168-021-01213-8 (2022).
Zimmermann, J., Kaleta, C. & Waschina, S. gapseq: informed prediction of bacterial metabolic pathways and reconstruction of accurate metabolic models. Genome Biol. 22(1), 81, https://doi.org/10.1186/s13059-021-02295-1 (2021).
Nobu, M. K. et al. Unique H2-utilizing lithotrophy in serpentinite-hosted systems. ISME J. 17(1), 95–104, https://doi.org/10.1038/s41396-022-01197-9 (2023).
Bährle, R., Böhnke, S., Englhard, J., Bachmann, J. & Perner, M. Current status of carbon monoxide dehydrogenases (CODH) and their potential for electrochemical applications. Bioresour Bioprocess. 10(1), 84, https://doi.org/10.1186/s40643-023-00705-9 (2023).
Berg, I. A. Ecological aspects of the distribution of different autotrophic CO2 fixation pathways. Appl Environ Microbiol. 77(6), 1925–36, https://doi.org/10.1128/AEM.02473-10 (2011).
Fuchs, G. Alternative pathways of carbon dioxide fixation: insights into the early evolution of life? Annu Rev Microbiol. 65, 631–58, https://doi.org/10.1146/annurev-micro-090110-102801 (2011).
Liu, Z., Wang, K., Chen, Y., Tan, T. & Nielsen, J. Third-generation biorefineries as the means to produce fuels and chemicals from CO2. Nat Catal. 3, 274–88, https://doi.org/10.1038/s41929-019-0421-5 (2020).
Aramaki, T. et al. KofamKOALA: KEGG Ortholog assignment based on profile HMM and adaptive score threshold. Bioinformatics. 36(7), 2251–2, https://doi.org/10.1093/bioinformatics/btz859 (2020).
AutoFixMark marker definition. https://github.com/h-mori/AutoFixMark/tree/main/definitions (2025).
Hügler, M. & Sievert, S. M. Beyond the Calvin cycle: autotrophic carbon fixation in the ocean. Ann Rev Mar Sci. 3, 261–89, https://doi.org/10.1146/annurev-marine-120709-142712 (2011).
Badger, M. R. & Bek, E. J. Multiple Rubisco forms in proteobacteria: their functional significance in relation to CO2 acquisition by the CBB cycle. J Exp Bot. 59(7), 1525–41, https://doi.org/10.1093/jxb/erm297 (2008).
Mueller-Cajar, O. & Badger, M. R. New roads lead to Rubisco in archaebacteria. Bioessays 29(8), 722–4, https://doi.org/10.1002/bies.20616 (2007).
Kono, T. et al. A RuBisCO-mediated carbon metabolic pathway in methanogenic archaea. Nat Commun. 8, 14007, https://doi.org/10.1038/ncomms14007 (2017).
Giovannelli, D. et al. Insight into the evolution of microbial metabolism from the deep-branching bacterium, Thermovibrio ammonificans. Elife. 6, e18990, https://doi.org/10.7554/eLife.18990 (2017).
Mall, A. et al. Reversibility of citrate synthase allows autotrophic growth of a thermophilic bacterium. Science 359(6375), 563–7, https://doi.org/10.1126/science.aao2410 (2018).
Nunoura, T. et al. A primordial and reversible TCA cycle in a facultatively chemolithoautotrophic thermophile. Science 359(6375), 559–63, https://doi.org/10.1126/science.aao3407 (2018).
Borrel, G., Adam, P. S. & Gribaldo, S. Methanogenesis and the Wood-Ljungdahl Pathway: An Ancient, Versatile, and Fragile Association. Genome Biol Evol. 8(6), 1706–11, https://doi.org/10.1093/gbe/evw114 (2016).
Adam, P. S., Borrel, G. & Gribaldo, S. Evolutionary history of carbon monoxide dehydrogenase/acetyl-CoA synthase, one of the oldest enzymatic complexes. Proc Natl Acad Sci USA 115(6), E1166–73, https://doi.org/10.1073/pnas.1716667115 (2018).
Hügler, M., Menendez, C., Schägger, H. & Fuchs, G. Malonyl-coenzyme A reductase from Chloroflexus aurantiacus, a key enzyme of the 3-hydroxypropionate cycle for autotrophic CO(2) fixation. J Bacteriol. 184(9), 2404–10, https://doi.org/10.1128/JB.184.9.2404-2410.2002 (2002).
Sánchez-Andrea, I. et al. The reductive glycine pathway allows autotrophic growth of Desulfovibrio desulfuricans. Nat Commun. 11(1), 5090, https://doi.org/10.1038/s41467-020-18906-7 (2020).
O’Leary, N. A. et al. Exploring and retrieving sequence and metadata for species across the tree of life with NCBI Datasets. Sci Data. 11(1), 732, https://doi.org/10.1038/s41597-024-03571-y (2024).
Braun, A. et al. Reviews and syntheses: heterotrophic fixation of inorganic carbon–significant but invisible flux in environmental carbon cycling. Biogeosciences. 18(12), 3689–700, https://doi.org/10.5194/bg-18-3689-2021 (2021).
Kawashima, S. et al. The 347 curated chemolithoautotrophs genome list. Zenodo https://doi.org/10.5281/zenodo.16956127 (2025).
Acknowledgements
We thank Dr. Koji Mori, Dr. Wakao Fukuda, Dr. Shin-ichiro Kato, and Dr. Takuro Nunoura for their valuable discussions on the metabolism of anaerobic bacteria. We also gratefully acknowledge Dr. Sumiko Yamamoto, Dr. Hiroko Maita, and Dr. Shinobu Okamoto for their support in pathway annotation. This work was primarily supported by the Green Innovation Fund Project (JPNP22010), commissioned by the New Energy and Industrial Technology Development Organization (NEDO), and partially supported by NBDC the Database Integration and Coordination Program (DICP), administered by the Japan Science and Technology Agency (JST) (Grant Number: JPMJND2206).
Author information
Authors and Affiliations
Contributions
Conceptualization: S.K., Y.O., N.I., H.M. Data collection and curation: S.K., Y.O., S.M., N.I., Y.N., T.N. Tool development: S.K., Y.O., H.N., H.M. Supervision: S.G., K.K., M.I., H.M. Writing - original draft: S.K., Y.O., N.I., H.M. Writing - review & editing: S.M., H.N., Y.N., T.N., S.G., K.K., M.I.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kawashima, S., Okabeppu, Y., Miyazawa, S. et al. A curated resource of chemolithoautotrophic genomes and marker genes for CO₂ fixation pathway prediction. Sci Data 13, 121 (2026). https://doi.org/10.1038/s41597-026-06655-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-026-06655-z