Abstract
The design and optimization of antibodies and nanobodies using deep generative models hold transformative potential for therapeutic and diagnostic applications, which are hindered by the fragmented and inconsistent nature of existing datasets. To address these limitations, we introduce the Antibody and Nanobody Design Dataset (ANDD), a unified dataset that integrates sequence, structure, antigen, and affinity data from 15 diverse sources. ANDD is a comprehensive resource comprising 48,683 antibody/nanobody sequences, with structural data for 24,941 entries, and antigen sequences for 12,575 entries. We further augmented the affinity data with 2,271 predicted affinity values using ANTIPASTI, a robust model for binding affinity prediction. Consequently, ANDD includes 9,557 affinity values, making it the largest dataset to date for antibody/nanobody and antigen pairs with affinity data. By addressing challenges of data fragmentation and inconsistency, ANDD provides a robust foundation for training deep generative models. With ANDD, the models can better model antibody/nanobody-antigen interactions, while design novel antibodies and nanobodies with improved specificity and efficacy, paving the way for development of targeted therapeutics.
Similar content being viewed by others
Background & Summary
Antibodies are large, Y-shaped glycoproteins (~150 kDa) produced by the immune system to identify and neutralize foreign objects like pathogens. A typical IgG antibody is composed of two identical heavy chains and two identical light chains, held together by disulfide bonds. Nanobodies are a novel class of therapeutic fragments derived from the heavy-chain-only antibodies found in camelids (e.g., llamas, alpacas), they are the smallest antigen-binding fragments to date (~15 kDa, 2.5 nm in diameter), consisting of a monomeric variable domain (VHH), shown as Fig. 1.
In therapeutic applications, antibodies and nanobodies are critical components due to their ability to target specific pathogens1,2, cancer cells3,4, or disease-related proteins5,6. Antibodies and nanobodies are widely applied in treatments for cancer7, autoimmune disorders8, and infectious diseases9. Recently, nanobody, also known as VHH, has gained prominence as the next-generation scaffold for therapeutics development due to its favorable bio-physical properties, including compact size, high solubility, and exceptional thermo-stability.
Traditionally, scientists utilized wet-lab screening to find out new antibodies and nanobodies. As the demand for targeted and personalized therapies increases, the limitations of these traditional antibody/nanobody discovery methods, such as their time-intensive nature and high development costs, have been increasingly recognized10. Computational models for de novo antibody/nanobody design aim to address these challenges11,12, which include previous models based on molecular dynamics simulations and the latest works utilizing deep generative models. These models improve the speed and precision of therapeutic development13 by directly generating antibodies and nanobodies with high-fidelity and high-specificity14. However, the development of these deep generative models is highly dependent on the availability of a well-curated dataset covering sequence, structural, antigen, and binding affinity data11, which is still fragmented and incomplete to date, badly hinder the development of deep generative models for antibody/nanobody design.
For the dataset curation, we identified three key challenges:
The first major issue is data fragmentation: most current datasets provide only a limited perspective on antibody or nanobody data. For instance, the Observed Antibody Space (OAS)15 contains numerous antibody sequences but lacks structural and affinity data. Similarly, the Protein Data Bank (PDB)16,17 offers structural information for general proteins but omits specific antigen and affinity details for antibodies and nanobodies.
The second challenge is format inconsistency: data from different databases often differs in format and structure, which complicates efforts to integrate them into a unified dataset. For example, sequence data is well-structured in UNIPROT18, while sequence data in SAbDab19 follows a different organization. This inconsistency necessitates extensive preprocessing to standardize the data format.
The third issue is missing binding values: many antibody and nanobody databases either lack binding affinity data or include only a limited number of binding affinity entries. Affinity data is essential for training models aimed at optimizing antibody-antigen interactions20. Without sufficient binding affinity data, these models could not effectively generalize across antibody-antigen interactions.
To address these challenges, we introduce the Antibody and Nanobody Design Dataset (ANDD), the largest dataset to date for antibody/nanobody and antigen pairs with binding data, which compiles data mainly from 15 sources. ANDD is a comprehensive dataset of antibodies and nanobodies, integrating sequence, structural, antigen, and affinity data into a unified resource, which includes data from databases and publicly available patents21,22,23,24. Specifically, it contains sequence data for 48,683 antibodies/nanobodies, structural information for 24,941 entries, antigen sequences for 12,575 entries, and binding affinity data for 9,557 antibody/nanobody-antigen pairs. Moreover, ANDD augments affinity data with a predictive model, ANTIPASTI25, which supplements binding affinity values for 2,271 antibody/nanobody-antigen pairs based on the structural data, greatly improving the dataset’s completeness. These data are basically constituted of antibody/nanobody-specific data resources (including INDI26, sdAb-DB27, PLABDAB28, SAbDab-nano19, and abYbank29 for nanobody, and SAbDab19, OAS15, AB-Bind30, Paddlepaddle31, and abYbank29 for antibody), and six general protein databases, including PDB16, UNIPROT18, PDBbind32, SKEMPI 2.033, DACUM34, and MpdPPI35.
In this paper, we present the structure and content of the ANDD dataset, describe the data collection and curation process, and explore its application in training deep generative models for antibody and nanobody design. By integrating various databases, ANDD provides a robust foundation for training accurate and generalizable models, facilitating the in silico antibody/nanobody design, Fig. 2 demonstrates the pipeline.
Detailly, the ANDD dataset is systematically organized into two primary categories: antibody data and nanobody (VHH) data. Each category follows a hierarchical classification based on the type and level of details, this organization provides progressively detailed sub-datasets, where each sub-dataset with more comprehensive information is the subset of those with simpler data (as Figs. 3, 4).
For the antibody data (Fig. 3), the entries are progressively categorized based on four levels of data integration. The broadest level, includes 18,464 entries containing only antibody sequence information. A more detailed subset of equally 18,464 entries combines sequence and structural data of antibody, providing foundational insights for structural studies. Within this, 8,190 entries further incorporate antigen sequence information alongside the antibody sequence and structure, supporting detailed analyses of antibody-antigen binding mechanisms. Finally, the most detailed subset includes 7,737 entries with sequence, structure, antigen, and affinity data. This graded dataset represents a highly valuable resource for antibody design and optimization, enabling predictive modeling and in-depth analysis.
For the nanobody (VHH) data (Fig. 4), the classification similarly progresses through four levels of details. The broadest level includes 30,119 entries with sequence data only. A subset of 6,477 entries provides both sequence and structural data, facilitating structural analysis of nanobodies. Among these, 4,385 entries add antigen sequence information, supporting investigations into nanobody-antigen binding interactions. The most comprehensive category includes 1,817 entries with sequence, structure, antigen, and affinity data, making it invaluable for high-precision applications in predicting binding affinities and training nanobody design models.
This layered data organization ensures that each entry in the ANDD dataset is tailored to the specific need of antibody and nanobody research, from sequence-only data for broad studies to highly detailed data for specialized modeling, de novo designing, and affinity prediction.
In conclusion, ANDD addresses the key challenges of fragmentation and incompleteness in antibody/nanobody dataset, providing a solid foundation for the development of more accurate and reliable design models. This dataset would play a key role in advancing antibody/nanobody research and therapeutic development. The dataset structure is shown as Fig. 5.
Methods
Overview of data curation
For those antibody/nanobody-specific data resources (including INDI, sdAb-DB, PLABDAB, SAbDab-nano, and abYbank for nanobody, and SAbDab, OAS, AB-Bind, Paddlepaddle, and abYbank for antibody), we directly adopt them into ANDD. This specific process involves: first, organizing them into a consensus data format; then, supplementing key affinity values; and finally, reorganizing them into a graded dataset based on a hierarchical classification system.
In contrast, for antibodies and nanobodies from those general protein databases (including PDB, UNIPROT, PDBbind, SKEMPI 2.0, DACUM, and MpdPPI.), we primarily re-process and filter them before integrate them into ANDD, shown as Fig. 6.
The schematic overview of data curation. This flowchart systematically illustrates the comprehensive methodology for constructing the ANDD. The process begins by integrating data from specialized antibody/nanobody databases and general protein databases. Data from specialized sources are directly incorporated, while entries from general databases undergo rigorous filtering to ensure relevance.
Noticing that ANDD lack large-scale antigen-specific nanobody sub-datasets for case-specific training, we supplement ANDD with publicly available nanobody patents targeting 4 therapeutically relevant antigens, including HER222, IL-621, CD4524, and the receptor binding domain (RBD) of the SARS-CoV-2 spike protein23, which are significantly helpful to nanobody design models for fine-grained capability. Detailed instructions refer to following methods.
Data processing and filtering of entries from general protein databases
To ensure our dataset was specific to antibodies and nanobodies, we first collected entries from nine authoritative antibody/nanobody databases (INDI, sdAb-DB, PLABDAB, SAbDab-nano, SAbDab, OAS, AB-Bind, Paddlepaddle, and abYbank), which curate antibody-focused and nanobody-focused data. These databases contained 7,757 unique PDB entries as the early-stage ANDD. We then compared these ANDD entries with data from general protein sources (mainly from PDB and UNIPROT, little from PDBbind, SKEMPI 2.0, DACUM, and MpdPPI). For entries from general protein sources, we primarily retain those entries that matched the identifiers from early-stage ANDD, after that, any PDB entry outside this intersection was manually validated for relevance to antibody and nanobody, and verified entries were brought into our prototype ANDD, shown as Fig. 6.
PDB data curation for antibody/nanobody structures
Structural data is critical in ANDD, often stored as pdb files in the RCSB Protein Data Bank (PDB), which archives both experimentally derived and computationally predicted protein structures. To download the structural data from PDB to ANDD, we used the “Advanced Search” function to filter specific antibody/nanobody-related identity, where we specified search terms relevant to immunoglobulins and excluded T-cell receptors to eliminate irrelevant entries. Besides downloading the pdb files, we also downloaded the structural property data in CSV format by selecting “Structure” as the “Tabular Report”, which includes items such as “Experimental Method” and “Structure Title”. Since the PDB download limit is 2,500 entries per batch, larger datasets were retrieved using the “Search and Data APIs”.
After processing, these PDB entries were manually filtered again to confirm any identifier refers to an antibody or a nanobody, which come up with 8,214 PDBs stored in the folder named “All_structures”.
UNIPROT data-filtering for antibody/nanobody sequences
We also utilized UNIPROT, which contains protein sequences and functional information. To introduce UNIPROT into ANDD, we used the keywords such as “antibody” or “nanobody”. To exclude T-cell receptors, we applied filters such as “NOT T-cell receptor.” Additional keywords such as “immunoglobulin” also ensured the high specificity. The refined UNIPROT entries were then matched with the prototype ANDD, while any unmatched entry was applied manual validation.
Other general protein sources follow the same way of pre-processing before been integrated into ANDD, shown as Fig. 6.
Augmentation and integration of affinity data
To address the lack of experimental affinity data, we utilized the ANTIPASTI. ANTIPASTI is a deep learning model designed to predict binding affinities by leveraging structural features of antibody/nanobody-antigen interactions. Specifically, we applied ANTIPASTI on ANDD entries containing structure data but missing affinity data. Besides, PDBbind contains binding affinity data of a vast range of complex structures, which was instrumental in providing binding affinity information to ANDD. Therfore, we downloaded the PDBbind v2020 dataset36 and introduced its affinity data into our ANDD. Similar procedures were applied to DACUM, MpdPPI, and SKEMPI 2.0. All affinity data were standardized to molarity (M) to ensure consistency across the dataset.
Data integration strategies of ANDD
To address data fragmentation, we implemented an integration strategy that consolidates data from multiple sources. This process involved extracting relevant information from antibody/nanobody-specific databases and general protein databases, then consolidating them into a consensus format. Our approach ensures that all relevant data is consolidated, enhancing the consistency and accuracy of the ANDD dataset.
We began at collecting antibody/nanobody-specific databases which contains structures, sequences, antigen information, and affinity data. Specifically, we utilized the following nine databases:
-
Database containing both antibody and nanobody: abYbank (latest public release, accessed 2023-06).
-
Databases for antibody: SAbDab (release dated 20250814), OAS (snapshot 202110), AB-bind (version 1.0), Paddlepaddle (official published release31).
-
Databases for nanobody: INDI (v1.0), SAbDab-nano (release dated 20250814), sdAb-DB (v1.0), PLAbDab (curated release, 202401).
In addition to antibody/nanobody-specific databases, we also sourced related information from six general protein databases: PDB (archive snapshot 2025-06-01), UNIPROT (release 2025_06), PDBbind (v2020), SKEMPI 2.0, DACUM (original published release34), and MpdPPI (original published release35).
All third-party sources used in this work are openly available. Data reuse and redistribution were conducted in accordance with the licenses and terms of use specified by the original data providers (e.g., CC BY, CC0, or equivalent open licenses). No access-restricted or proprietary datasets was included.
After that, we evaluated the data items to form a consensus format that is essential to the integration of ANDD.
As for general description, the consensus format includes 5 items: Source, Update_Date, Complex_Structure, Ab_or_Nano, and Source_Organism.
As for structural data, it includes 4 items: PDB_ID, PDB_ID_Changed, Experimental_Method, and Structure_Title. Besides, ANDD also provides 8,214 PDB files for these antibody and nanobody structures.
As for heavy chain data, it includes 8 items: H_Chain Entity ID, H_Chain Asym ID, H_Chain Auth Asym ID, H_Chain Database Name, H_Chain Accession Code (s), H_Chain Sequence Cluster ID, H_Chain Sequence Cluster Identity Threshold, and H_Chain Macromolecule Name.
As for light chain data, it includes 8 items: L_Chain Entity ID, L_Chain Asym ID, L_Chain Auth Asym ID, L_Chain Database Name, L_Chain Accession Code (s), L_Chain Sequence Cluster ID, L_Chain Sequence Cluster Identity Threshold, and L_Chain Macromolecule Name.
As for antigen data, it includes 8 items: Ag_Entity ID, Ag_Asym ID, Ag_Auth Asym ID, Ag_Database Name, Ag_Accession Code (s), Ag_Name, Ag_Seq, and Ag_Source Organism.
As for mutation data, it refers to an item noted Ab/Nano_Mutation.
As for sequence data, it includes 9 items: Ab/Nano H_Chain AA, Ab/Nano L_Chain AA, Ab/Nano_CDR H1, Ab/Nano_CDR H2, Ab/Nano_CDR H3, Ab/Nano_CDR L1, Ab/Nano_CDR L2, Ab/Nano_CDR L3, and CDR Nomenclature.
As for affinity data, it includes 3 items: Affinity_Kd (M), ∆Gbinding, and Affinity_Method.
As for the quality control, it includes 3 items: Reason_Code, Predicted_or_Not, and Provenance.
What mentioned above constitutes the largest currently available dataset for antibody/nanobody and antigen pairs with binding data, which could facilitate the development of antibody/nanobody design model. These items would be detailedly described in Data Record.
However, we also encountered issues in some antibody-specific databases, such as the inclusion of T-cell receptor (TCR) data, which structurally differs from antibody but would be mistakenly mixed in our ANDD during the integration process. To ensure accuracy and relevance, we manually screened all entries to exclude TCR and other non-antibody or non-nanobody data. This rigorous and automated curation process guarantees that our ANDD table exclusively contains antibody and nanobody data.
Categorizing and organizing antibody-related and nanobody-related data
To resolve data inconsistencies, we developed a systematic approach to categorizing and organizing antibody/nanobody-related data of ANDD. This approach organized sequence data, structural data, antigen information, and affinity data into a standardized and interoperable structure, it reconstructed the dataset into a graded dataset based on a hierarchical classification system, which ensures consistency across the ANDD and establishes a reliable foundation for analysis and model training.
The graded dataset is organized into 4 levels: the broadest level refers to entries containing antibody/nanobody sequence information; a more detailed level refers to entries combining sequence and structural data of antibody or nanobody; the tertiary level indicates entries further incorporate antigen sequence information alongside the antibody/nanobody sequence and structure data; the most detailed level indicates entries with sequence, structure, antigen, and affinity data. This graded dataset represents a highly valuable resource for antibody/nanobody design and optimization, enabling ANDD for generative modeling and in-depth analysis.
For sequence data, we gathered them mainly from databases that support direct download, including OAS, INDI, Paddlepaddle, abYbank, PDB, and UNIPROT, we also supplemented it by manually taking down sequence data from those web-based databases and publicly available patents, including sdAB-DB and SAbDab. To enhance consistency, we standardized the sequence format and categorized the sequence data into light chain sequence, heavy chain sequence, antigen chain sequence, and sequences of complementarity-determining regions (CDRs).
For structural data, we mainly curated them from PDB, UNIPROT (partially), PDBbind, SKEMPI 2.0, DACUM, MpdPPI, SAbDab, AB-bind, abYbank, and INDI_structure. For each structure entry, we provided standardized annotations, including the structure title, the structure status (complex or monomer), and alternative ID. These details ensure that the structural data is both accurate and comprehensive, facilitating precise downstream applications. More detailed information, exhibited as 8,214 PDB files, were downloaded to the folder named All_structures from PDB database.
For affinity data, databases with explicit affinity measurements, such as AB-bind, SAbDab, and Paddlepaddle, were directly utilized. We annotated the ANDD table with the affinity value (Kd standardized to M, and ∆Gbinding standardized to kJ/mol) and the affinity measure method.
Addressing missing affinity data
For entries with sequence and structural data but lacking affinity data, we applied ANTIPASTI to estimate binding affinities, which added 2,271 predicted affinity values. This approach substantially expanded the ANDD, bridging significant gaps that experimental affinity data was scarce.
Manual curation from web-based databases
In situations where sequence, structure, and affinity data were unavailable to be directly downloaded but could only be accessed from the websites, we manually took down these data. For example, we manually collected sequence data from SAbDab, while manually collecting sequence, structure, and affinity data from sdAB-DB. This step was essential to compile a comprehensive dataset, and made ANDD a more accessible resource compared to those web-based databases.
For entries lacking specific types of data, we manually supplemented missing information when possible. For instance, we manually supplemented sequence data from SAbDab with corresponding structural and affinity data from PDBbind. Similarly, in cases like PDBbind and AB-bind, whose sequence data were missing, we supplemented the missing sequence data from SAbDab. For databases, such as UNIPROT, whose sequence data did not labeled with light chain and heavy chain, we manually utilized sequence annotations in PDB to mark the light chain ID and heavy chain ID. Additionally, we manually annotated the entries with mutation site from mutation-specific databases like SKEMPI 2.0, DACUM, and MpdPPI.
Through this manual curation, we ensured that the final ANDD is comprehensive and accurate, enhancing its potential for model training and analysis. This systematic approach of data collection, integration, and supplementation has enabled ANDD become the most comprehensive dataset on antibody and nanobody data, which is designed to support deep learning-based antibody/nanobody de novo design.
Provenance annotation
To ensure full traceability of data origin and processing history, a dedicated provenance field was added to each record. This field encodes both the contributing data sources and the deterministic processing steps applied during dataset construction in a delimiter-separated format. Source identifiers precede transformation tags, allowing each entry to be traced back to its original databases and to the specific harmonisation, filtering, de-duplication, standardisation, or prediction steps applied. Provenance strings follow a fixed, rule-based syntax and do not involve any free-text or manual annotations.
Fully scripted and reproducible workflow
An end-to-end, command-line-driven workflow that enables rebuilding the ANDD dataset entirely from raw public sources.
The pipeline covers: (i) automated data retrieval from all source databases, where raw data were programmatically retrieved from antibody/nanobody-specific databases and general protein databases using official APIs, bulk downloads, or archived releases.(ii) parsing and normalization into a consensus schema, (iii) controlled-vocabulary harmonisation, (iv) de-duplication and cross-source conflict resolution, (v) affinity unit standardization and provenance annotation, and (vi) optional affinity prediction using ANTIPASTI.
The workflow was implemented in Python and executed in a controlled computational environment with fixed software versions. Each processing step was encapsulated as an independent script, enabling modular execution and transparent inspection of intermediate outputs. Configuration files were used to define data sources, schema definitions, and controlled vocabularies, allowing the pipeline to be rerun or extended without modifying core code. As a result, the complete dataset can be regenerated in a clean environment by sequentially executing the provided scripts.
Utilizing this workflow, no step in the construction of ANDD requires manual web interaction; all operations are reproducible via scripts and configuration files, enabling deterministic regeneration of the dataset from raw public sources.
Data Records
The ANDD dataset37 is available at Zenodo on https://zenodo.org/records/18151718 with this section being the primary source of information on the availability and content of the data being described. The dataset is now accessible under the Creative Commons Attribution 4.0 International, which supports its use for educational and research purposes. Users should cite this paper when they incorporate the dataset into their projects. The presented ANDD consists of four parts. The first part is a spreadsheet (ANDD.csv) that summarizes the sequence, structure, antigen, and affinity information of all entries in the ANDD. The second part is a folder (/All_structures) containing the crystal structures (8,214 PDB files) of entries in the ANDD, shown as the Fig. 5. The third one is a data quality control report containing a comprehensive summary of dataset composition, and the forth one is a data dictionary describing all fields and controlled terms.
Data format
Source: The source of the data.
Update_Date: The latest date of the update.
PDB_ID: The PDB identifier of the entry, which can be used to retrieve the corresponding protein structure file from the PDB database and the All_structures folder. It should be noted that some antibodies only have sequence information without 3D structural data, and most nanobodies lack structural data, which is an issue that might be addressed by structural prediction model, such as Alphafold338.
PDB_ID_Changed: This item indicates whether the PDB ID has any alternative ID. If the PDB ID has been updated, this column provides the updated PDB ID, otherwise it would be noted as “No”, indicating that this entry still uses the original PDB ID.
Experimental_Method: This item specifies the technique used to obtain the structure of each entry, which could be one of these techniques: “ELECTRON CRYSTALLOGRAPHY”, “ELECTRON MICROSCOPY”, “SOLID-STATE NMR”, “SOLUTION NMR”, “SOLUTION SCATTERING”, and “X-RAY DIFFRACTION”.
Structure_Title: This item indicates the structure title retrieved from the PDB database. The title helps identify the protein type, ensuring it is an antibody/nanobody, and deciding whether the structure is a complex.
Complex_Structure: This item indicates whether the structure is a complex, marked as “TRUE” if it is, and “FALSE” otherwise.
Ab_or_Nano: This item indicates whether the entry is an Antibody, a Nanobody (VHH), a light-chain dimer (BJ), or a single-chain variable fragments (scFv), specifying the type of molecule in each entry.
Source_Organism: This item indicates the originating organism of the antibody/nanobody, specifying the biological source of each structure.
H/L_Chain Entity ID: The Entity ID identifies distinct chains within the entry, often distinguishing between heavy chain and light chain.
H/L_Chain Asym ID: This ID identifies the asymmetric units within the antibody/nanobody structure. In antibodies, this ID typically represents different chains, such as the heavy chain (H) or light chain (L). In single-chain nanobodies, there is only one Asym ID because a nanobody contains only a heavy chain.
H/L_Chain Auth Asym ID: This is the author’s specific ID for the asymmetric units, which is a specific notation used by the researchers. For example, an author might label the heavy chain with “H” and the light chain with “L,” which could differ from the official Asym IDs.
H/L_Chain Database Name: This item indicates the source database of the entry, such as GenBank, which refers to the external reference database of the antibody or nanobody.
H/L_Chain Accession Code (s): This is a unique ID specifically used in the external databases (e.g., GenBank), enabling users to trace this antibody or nanobody in its external reference databases.
H/L_Chain Sequence Cluster ID: The Sequence Cluster IDs refer to clusters created by grouping highly similar sequences together. This ID is useful to identify the similarity between different entities, it also indicates evolutionary relationships and functional similarities.
H/L_Chain Sequence Cluster Identity Threshold: This item indicates the similarity threshold used to group sequences into the clusters. For antibodies, high identity thresholds (e.g., 90% or 100%) might cluster sequences with nearly identical variable regions, which are critical for antigen binding specificity.
H/L_Chain Macromolecule Name: This item indicates the type of antibody/nanobody entity, such as “IGG1KAPPA 2E8 FAB (LIGHT CHAIN)” or “HEAVY CHAIN.” For antibodies, it distinguishes between heavy chains and light chains or indicates specific regions (e.g., Fab or Fc fragment).
Ag_Entity ID: The unique ID of antigen in each entity.
Ag_Asym ID: This item is the asymmetric unit ID of the antigen, indicating a specific chain or structural unit in the antigen.
Ag_Auth Asym ID: This item is author-assigned asymmetric unit ID of the antigen. It is often specifically used by researchers in their publications to label the antigen chains.
Ag_Database Name: This item is the name of the external database where the antigen originates, such as UNIPROT. It provides a reference database to retrieve additional information about the antigen.
Ag_Accession Code (s): The specific identifier or accession code of the antigen in the specified database (e.g., UNIPROT ID), which allows users to retrieve more information of the antigen from this external database.
Ag_Name: The name of the antigen, which often includes detailed information about its biological function or subunit identity.
Ag_Seq: The sequence of the antigen.
Ag_Source Organism: The organism of the antigen, from which the antigen was derived.
Ab/Nano_mutation: This item indicates whether there is an amino acid mutation in the antibody/nanobody sequence, formatted as “H A001B”, which means that the amino acid at position 001 of the H chain has mutated from A to B.
Ab/Nano_H_Chain AA: The amino acid sequence of the heavy chain.
Ab/Nano_L_Chain AA: The amino acid sequence of the light chain. For nanobodies, this column is absent.
Ab/Nano_CDR H1: The amino acid sequence of the first CDR in the heavy chain.
Ab/Nano_CDR H2: The amino acid sequence of the second CDR in the heavy chain.
Ab/Nano_CDR H3: The amino acid sequence of the third CDR in the heavy chain.
Ab/Nano_CDR L1: The amino acid sequence of the first CDR in the light chain.
Ab/Nano_CDR L2: The amino acid sequence of the second CDR in the light chain.
Ab/Nano_CDR L3: The amino acid sequence of the third CDR in the light chain.
CDR Nomenclature: The nomenclature of CDRs (Complementarity-Determining Regions).
Affinity_K_d (M): The Dissociation Constant (Kd) between the antibody/nanobody and the antigen, measured in mole (M). The Kd was collected only when measured using established biophysical or immunochemical assays widely accepted in the field, such as surface plasmon resonance (SPR), isothermal titration calorimetry (ITC), and related equilibrium binding techniques, which were performed under near-physiological conditions, typically at ambient or controlled temperatures (20–25 °C or 37 °C) and in buffered aqueous systems.
∆Gbinding (kJ/mol): The binding free energy was included only when directly derived from thermodynamic measurements, most commonly via ITC under near-equilibrium conditions with constant temperatures (20–25 °C), which can be converted to corresponding Kd value using the formula:
in this equation, ∆Gbinding represents the binding free energy, which quantifies the binding affinity between two molecules, such as an antibody/nanobody and its antigen. The symbol R stands for the universal gas constant, valued at 8.314 J · mol−1 · K−1, which is a key factor in thermodynamic calculations. The variable T represents the absolute temperature, measured in Kelvin (K), typically around 298 K (or 25 °C). Kd is the dissociation constant, which is a key measure of the affinity between binding molecules, a lower Kd indicates stronger binding affinity. Finally, the natural logarithm (ln) is used to convert the dissociation constant to the binding free energy, establishing the link between molecular interactions and thermodynamic principles.
Affinity_Method: This item specifies the technique used to obtain the affinity data of each entry.
Reason_Code: This item documents the cause of missingness for NA in affinity data, including not_reported, conflicting, and inferred.
Predicted_or_Not: This item indicates whether the affinity value was predicted by the ANTIPASTI or derived from experimental measurements.
Experimentally measured and predicted affinity values are explicitly separated using the Predicted_or_Not field. Records annotated as real correspond to experimentally measured affinities, whereas records annotated as predicted indicate affinities predicted by ANTIPASTI (version 2023, the latest version), which is fully consistent with the Reason_Code and Provenance fields.
Importantly, only records labeled as real should be interpreted into training data when modeling antibody/nanobody and antigen interaction, users could filter the real affinities from ANDD37 to form a training data snapshot.
Provenance: This item documents the data sources and processing history, enabling users to trace individual entries back to their original databases and applied transformation rules, which includes two parts: (i) identifiers of all contributing source databases, followed by (ii) a sequence of predefined transformation tags corresponding to deterministic processing steps. The transformation tags used in this dataset are summarised in Table 1.
For example, an entry derived from SAbDab and OAS, and subsequently harmonised is annotated as abYbank_ab|PDB|schema_harmonisation.
An example of these items is shown in Fig. 7, as a single, representative row from ANDD.
Data quality control report
In addition to full-length antibodies, there also existed other antibody-derived molecules, including light-chain dimer (BJ), nanobody (VHH) and scFv (single-chain variable fragments). To ensure consistent and biologically accurate annotation across heterogeneous data sources, we defined a controlled vocabulary for immunoglobulin formats using IMGT as the standard authority in Table 239.
ANDD entries were classified into four categories, including antibodies, scFv, VHH, and BJ, based on chain composition, domain organization, and biological origin. The H2L2 antibody is a full-length immunoglobulin with two heavy and two light chains; the scFv is a single-chain variable fragment as VH-linker-VL or VL-linker-VH, present of one heavy-chain variable region and one light-chain variable region; the VHH is a single-domain antibody derived from camelids, present of a single variable domain homologous to the heavy-chain variable region; the BJ is a Bence Jones-type or engineered light-chain dimer, present of only light-chain domains forming a homodimer. Formal definitions, required inclusion criteria, and exclusion rules for each format are summarized in Table 2. This controlled vocabulary was applied to harmonizing across all records following cross-source consistency checks.
To ensure annotation consistency and transparency, we performed a systematic audit after controlled vocabulary harmonization. A quality control analysis quantifies the distribution of entries across different formats and source organisms, as summarized in Fig. 8 and Fig. 9.
The quality control of entries’ source organisms. Species-level annotation of host/source organisms revealed broad taxonomic coverage, with the majority of entries originating from Homo sapiens (8,163 records), and Mus musculus (6,910 records). Camelidae mixed libraries may be generated through experimental cloning or synthetic approaches, and may comprise VHH-only repertoires or mixed VH/VHH constructs.
We conducted cross-source consistency checks to verify the appropriateness of nanobody (VHH) annotations. A total of 217 entries originally labeled as VHH were relabeled, including 96 entries reassigned to BJ and 121 entries reassigned to scFv, due to incompatible chain composition or non-camelid origin. These corrections were applied uniformly across the dataset and recorded in the provenance field.
Following relabeling, we quantified the post-QC distribution of entries by immunoglobulin format and species. Figure 8 demonstrates the distribution across different formats, with the majority of entries belonging to antibodies or nanobodies. Figure 9 summarizes the counts stratified by source organisms, showing that VHH annotations are exclusively associated with camelid species, whereas Fab/scFv/BJ formats predominantly originate from human and murine sources. Among these source organisms, “Camelidae mixed library” (NCBI Taxon ID: 1579311) refers to sequences derived from pooled camelid antibody libraries combining immunoglobulin repertoires from multiple camelid species (including VHH-only libraries or mixed VH/VHH libraries), for which a single donor species cannot be unambiguously assigned, and therefore represents a library-level origin rather than a biological species.
No unresolved ambiguities remained after this audit; all records could be confidently assigned to a specific antibody format. Status flags are provided in the provenance field in the released ANDD dataset to ensure transparency and traceability.
We uploaded the data quality control report to ANDD repository in Zenodo at https://zenodo.org/records/18151718, and distilled a concise QC checklist that captures the key QC measures applied to the ANDD dataset, shown in Table 3.
Data comparison with existing resources
To contextualise the scope and characteristics of ANDD37, we performed a systematic comparison with representative antibody-, nanobody-, and general protein databases (Table 4). The comparison covers data scale, modality coverage, schema design, update strategy, and data organisation.
Existing antibody-focused resources such as SAbDab, AB-bind, PaddlePaddle, and abYbank-ab provide valuable sequence- or structure-level annotations, yet typically focus on a single modality and lack explicit cross-links between sequence, structure, affinity, and antigen information. Nanobody-specific databases, including INDI, sdAb-DB, SAbDab-nano, and PLAbDab-nano, show similar limitations. General protein resources such as PDB, UniProt, PDBbind, SKEMPI 2.0, DACUM, and MpdPPI provide valuable structural or interaction data, but are not exhibited as a dedicated or hierarchical manner, which leads to data fragmentation and confused.
In contrast, ANDD integrates antibody and nanobody data within a unified, harmonised, and hierarchical dataset. Specifically, ANDD comprises 30,119 antibody sequences and 18,464 nanobody sequences, linked to 6,477 and 18,464 corresponding structures, respectively. The database further includes 7,737 antibody-antigen and 1,817 nanobody-antigen affinity values, together covering 12,474 unique antigen sequences and 12,617 antigen names, ANDD is now the largest dataset in this field.
Importantly, ANDD is the only resource that simultaneously supports harmonised schema design, systematic de-duplication, explicit cross-modality links, and hierarchical organisation across sequence, structure, antigen, and affinity levels. These features enable consistent downstream analysis and benchmarking, while avoiding redundancy and ambiguity, which demonstrates that the scope and novelty of ANDD is further beyond simple aggregation of existing databases.
Data Overview
As Fig. 10 shows, ANDD37 contains sequence data for 48,683 entries, structural information for 24,941 entries, antigen sequences for 12,575 entries, and binding affinity data for 9,557 antibody/nanobody-antigen pairs, these data is collected from different sources, which include antibody/nanobody-specific databases, general protein databases, and publicly available patents. Affinity data is also augmented by ANTIPASTI. As we can see, nanobody data dominates in the sequence information, while antibody owns a richer structural and antigen data. Affinity data of ANDD offers the most extensive collection currently available.
Technical Validation
Manual validation of all data
The ANDD37 database consists of antibody and nanobody entries, which has gone through rigorous quality control. We firstly manually proofread and validated the main data of ANDD. For the antibody/nanobody data, we re-validated the quantity of each level in the graded dataset. We also manually checked if the sequence data corresponded to the structural data of every entry. After that, we checked the accuracy of information by random selection, and proofread the sequence data, structural data, affinity data, and antigen data of the selected entries, for every 10 entries, at least one of them was selected to be manually validated. All data is available on the https://zenodo.org/records/18151718, and we will continuously inspect and update, while listing the details of each update in the metadata.
ANDD was built upon 15 different source datasets and 4 nanobody patents, any duplicate was removed and extra records were obtained that would otherwise have been overlooked, which came up with the largest currently available dataset for antibody/nanobody and antigen pairs with affinity data.
AlphaBind validation of affinity data
We also validated the Kd values in ANDD37 with a binding affinity proxy, AlphaBind40. AlphaBind is a pre-trained model predicting enrichment ratio (ER) between an antigen and an antibody/nanobody, enrichment ratio is a quantitative metric, which is defined as the ratio of bound to unbound fractions, a higher ER denotes a stronger binding affinity, ER could be converted to Kd using the formula:
in which, the natural logarithm (ln) is used to convert the dissociation constant to the enrichment ratio, and the L denotes the concentration of free ligands, measured in mole (M).
To evaluate the accuracy of affinity data in our ANDD, we filtered all 4030 entries with experiment-obtained Kd from ANDD, and predicted their ER utilizing AlphaBind. The association between predicted ER values and experimental affinities was evaluated by comparing −ln(Kd) and ln(ER). Pearson correlation analysis revealed a significant positive correlation (PCC = 0.750, p < 0.001, n = 4030), while Spearman’s rank correlation coefficient (SCC = 0.691, n = 4030) further supported a strong positive relationship. The coefficient of determination was R² = 0.563, indicating that a substantial proportion of the variance in experimental affinities is explained by the predicted values. The 95% confidence interval for the Pearson correlation coefficient is shown as the shaded region in Fig. 11, most of the data points fall within the 95% CI, indicating experiment-obtained Kd values from ANDD are highly precise.
In addition to correlation-based metrics, we quantified absolute error using mean squared error (MSE = 0.426, n = 4030, corresponding to RMSE = 0.653). We also analysed the residual error distribution (predicted values minus experimental values), shown in Fig. 12. The residual errors are approximately centred around zero, suggesting no pronounced systematic bias in the Kd data, while further reflecting the accuracy and reliability of Kd values in ANDD.
In conclusion, because the AlphaBind could not achieve state-of-the-art on binding affinity prediction, it is only possible to roughly determine the ranking and general trend of the binding affinity. The observed statistically significant correlations, together with absolute discrepancy metrics and residual analyses, support the reliability and accuracy of the experimentally measured affinity data integrated into ANDD, confirming their suitability for downstream analyses and benchmarking tasks.
We must clarify that these predicted affinity values are only used for technical validation, are never used as ground truth in any benchmark, evaluation, or downstream analysis.
Cross-mapping validation of affinity data
Mathematically related items could be validated with the cross-mapping. Two independent items were selected for a correlation analysis, which are Kd and ∆Gbinding, they both evaluate binding affinity, and could be converted to each other by the Eq. (1).
We filter all 1352 entries with both Kd and ∆Gbinding, and calculate the correlation between Kd and ∆Gbinding with Pearson’s correlation, and came up with a correlation coefficient of 1.000 with high significance (p < 0.01), the correlation map between ln (Kd) and ∆Gbinding is shown as Fig. 11.
In conclusion, the Kd and ∆Gbinding in ANDD are of high accuracy, which also highlights the reliability of the cross-mapping method.
Example usage in optimizing generative models
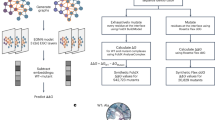
To illustrate the downstream usability of ANDD37, we conducted a minimal proof-of-concept fine-tuning experiment using DiffAb41, an existing diffusion-based generative model for antibody and nanobody sequence-structure co-design. This experiment is presented solely as an illustrative example to validate data usability and internal consistency, and is not intended as a benchmark of generative model performance. A subset of 12,617 ANDD entries containing paired sequence, structure, and antigen information was used to fine-tune the pretrained DiffAb model. Generated structures were evaluated using standard external metrics, including predicted affinity (predicted Kd by AlphaBind40 and binding affinity rank by Nanobinder42), structural diversity (RMSD and TM-score43), and developability properties (SASA and β-sheet content for stability; and humanness is measured by OASis44). Across all evaluated metrics, the DiffAb fine-tuned on ANDD showed consistent improvements relative to the vanilla baseline (p < 0.05), proving the high quality of ANDD, demonstrating the practical usability of ANDD as a coherent dataset for downstream antibody and nanobody modelling tasks. Detailed scripts and results were uploaded to our code repository at https://github.com/Wu6623/ANDD_workflow/tree/main/DiffAb_related.
Summary of validation pipeline
To ensure data quality and usability, we implemented a multi-step validation pipeline through qualifying field-level completeness for all core metadata fields, identifying duplicate records, and resolving cross-source conflicts.
The validation summary (Table 5) provides a high-level overview of dataset quality. Core metadata fields exhibit high completeness, ranging from 92.4% to 100%. Duplicate records were systematically removed based on identical identifiers and sequence similarity. Cross-source inconsistencies were resolved through a combination of manual curation and source prioritization based on source reliability, priority was given to experimentally validated annotations, followed by curated databases, and finally automated predictions. In addition, affinity values were validated by AlphaBind and with explicit evidence provenance annotation.
Table 6 summarizes field-level completeness together with the corresponding validation rules. Core annotation fields show high coverage, including Ab_or_Nano (100.0%), Reason_Code (100.0%), Predicted_or_Not (100.0%), and Ab/Nano H_Chain AA (99.26%). Structural identifiers derived from PDB records, such as PDB_ID and chain entity identifiers, exhibit moderate completeness, ranging from 33.45% to 44.68%. Affinity-related fields include quantitative Kd values available for 16.48% of entries, ΔG values reported for only 5.91%, and affinity measurements available for 25.65% of records.
Notably, ΔG values exhibit the lowest completeness, as binding free energies were included only when measured under equilibrium conditions with explicit temperature and assay information. Similarly, Kd values show limited completeness because only quantitative dissociation constants derived from established equilibrium binding assays with standardized units and traceable experimental metadata were retained.
This pattern reflects the true state of data availability and reporting practices, to preserve data integrity and avoid introducing artificial bias, no missing values were fabricated or inferred, instead, missing values were systematically documented using predefined validation rules and evidence annotations. This distribution reflects a deliberate design choice prioritizing data reliability and traceability over numerical completeness.
Usage Notes
All data included in ANDD37 are publicly available in Zenodo at https://zenodo.org/records/18151718 under a Creative Commons Attribution 4.0 International (CC-BY) license (https://creativecommons.org/licenses/by/4.0/). This database adheres to the FAIR principles45, allowing researchers to find, access, understand, and reuse data from ANDD. Authors may freely use our database under the condition that this paper is cited.
Potential uses of the dataset
We hope that this fine-grained dataset will help researchers, and deep generative algorithm engineers in the field of protein drug discovery to accelerate the development of targeted therapeutics.
The goal of ANDD is to enhance generative models’ capabilities in antibody/nanobody design by providing a broader, fine-grained dataset than currently available ones. Existing generative models are often constrained by the limitations of their training data. For instance, many models are trained exclusively on sequence-only databases, which may produce biologically plausible sequences that lack the structural fidelity and binding efficacy. By integrating ANDD into the training dataset, which offers sequence, structural, antigen, and affinity information, generative models could produce entities with improved biological and structural relevance.
ANDD is composed of PDB files as structural data stored in All_structures folder, and sequence data with affinity labels saved in Antibody and Nanobody Design Dataset (ANDD) table. When ANDD is used to fine-tune structure design models, the 8,214 pdb files could be randomly divided into a training subset, a test subset, and a validation subset. When ANDD is used in training sequence design models or affinity prediction models, 48,683 sequences and 9,557 affinity labels could be directly loaded from the csv table. Besides, ANDD also provides fine-grained items of nanobody/antibody, including source organism, chain ID, experimental method, and so on, contributing to its function as a powerful query tool, enabling researchers to uncover detailed mechanisms of antigen-antibody/nanobody interactions. ANDD fills the gap of data fragmentation, format inconsistency, and data incompleteness, which owns high potential in optimizing the performance of deep generative models and developing the targeted therapeutics.
Limitations of datasets
A key limitation of ANDD is that only a subset of affinity annotations has been experimentally validated under standardized conditions, as most values are compiled from heterogeneous data sources. In addition, although ANDD covers a broad range of antibody and nanobody interactions, the dataset can be further expanded to improve coverage across targets, formats, and species, which is essential for enhancing generalization in downstream modeling and engineering applications. Future work will prioritize systematic experimental validation and continued expansion of data diversity to strengthen the robustness and translational relevance of the dataset.
Data update
All data are available on the https://zenodo.org/records/18151718, and we will continuously inspect and update, while listing the details of each update in the metadata. In addition to our regular updates, any suggestion or update from any individual is appreciated, and please feel free to contact the author, we encourage researchers to participate in ANDD data updates as contributors. With the deployment of the ANDD, we plan to update the data about every three months. In addition, when major changes that require version control occur, the production version of the database will be updated regularly.
Data availability
The Antibody and Nanobody Design Dataset (ANDD)37 is publicly available under a Creative Commons Attribution 4.0 International (CC BY 4.0) license on Zenodo:
• Repository: Zenodo
• Resource Type: Dataset
• https://doi.org/10.5281/zenodo.18151718
• https://zenodo.org/records/18151718
• Publication Year: 2025
The dataset consists of four main components:
• The primary table file, Antibody and Nanobody Design Dataset (ANDD).csv, which integrates all sequence data, structure data, affinity data, and antigen data.
• The All_structures folder containing the corresponding 8,214 PDB files for structural entries.
• The data quality control report containing a comprehensive summary of dataset composition, field-level completeness, relabeled entries, and unresolved ambiguities identified during validation.
• The data dictionary describing all fields, controlled terms, units, and allowed values.
The Zenodo deposit corresponds exactly to the artefact used in the preparation of this manuscript. Specifically, the released ANDD represents the same tagged dataset used for all analyses, figures, and tables reported in the paper, with no post-deposition modifications.
Code availability
We utilized ANTIPASTI (a binding affinity proxy proposed by Michalewicz et al. in 202325) to augment affinity data, which is publicly available under the MIT License at https://github.com/kevinmicha/ANTIPASTI.git, and we utilized Diffab (an antibody/nanobody design model proposed by Luo et al. in 202240) for technical validation, which is publicly available under the Apache-2.0 license at https://github.com/luost26/diffab.git.
We constructed an end-to-end, command-line-driven workflow that enables rebuilding the ANDD37 dataset entirely from raw public sources. All scripts required to rebuild the ANDD dataset from raw public sources are publicly available at https://github.com/Wu6623/ANDD_workflow under the Apache-2.0 license, which also contains the environment.yml containing a frozen computational environment. The released workflow replaces all manual web-based operations with scripted equivalents and supports automated data retrieval, parsing, harmonisation, de-duplication, filtering, affinity standardisation, and validation. Detailed execution instructions and environment specifications are provided in the repository, enabling full reproducibility of the dataset. It also contains the scripts used for DiffAb fine-tuning and evaluation in the DiffAb_related folder.
References
Jin, B.K., Odongo, S., Radwanska, M. & Magez, S. NANOBODIES (R): A Review of Diagnostic and Therapeutic Applications. Int J Mol Sci 24 (2023).
Lu, R. M. et al. Development of therapeutic antibodies for the treatment of diseases. J Biomed Sci 27, 1 (2020).
Abdolvahab, M. H., Karimi, P., Mohajeri, N., Abedini, M. & Zare, H. Targeted drug delivery using nanobodies to deliver effective molecules to breast cancer cells: the most attractive application of nanobodies. Cancer Cell Int 24, 67 (2024).
Jovcevska, I. & Muyldermans, S. The Therapeutic Potential of Nanobodies. BioDrugs 34, 11–26 (2020).
Haddad, F., Dokmak, G., Kanwal, S. & Karaman, R. A Comprehensive Review on Therapeutic Potential of Nanobodies. Preprint at https://www.preprints.org/frontend/manuscript/8fc19cbc184efbf3f0096555d235be6f/download_pub (2023).
Panwar, U., Khan, M.A., Selvaraj, C. & Singh, S.K. Therapeutic antibodies against cancer—A step toward the treatment. in Resistance to Anti-CD20 Antibodies and Approaches for Their Reversal 3–29 (2024).
Sroga, P., Safronetz, D. & Stein, D. R. Nanobodies: A New Approach for the Diagnosis and Treatment of Viral Infectious Diseases. Future Virology 15, 195–205 (2020).
Felten, R., Mertz, P., Sebbag, E., Scherlinger, M. & Arnaud, L. Novel therapeutic strategies for autoimmune and inflammatory rheumatic diseases. Drug Discov Today 28, 103612 (2023).
Rizk, S. S., Moustafa, D. M., ElBanna, S. A., Nour El-Din, H. T. & Attia, A. S. Nanobodies in the fight against infectious diseases: repurposing nature’s tiny weapons. World J Microbiol Biotechnol 40, 209 (2024).
de Brito, P. M., Saruga, A., Cardoso, M. & Goncalves, J. Methods and cell-based strategies to produce antibody libraries: current state. Appl Microbiol Biotechnol 105, 7215–7224 (2021).
Bai, G. et al. Accelerating antibody discovery and design with artificial intelligence: Recent advances and prospects. Semin Cancer Biol 95, 13–24 (2023).
Hummer, A. M., Abanades, B. & Deane, C. M. Advances in computational structure-based antibody design. Curr Opin Struct Biol 74, 102379 (2022).
Mason, D. M. et al. Optimization of therapeutic antibodies by predicting antigen specificity from antibody sequence via deep learning. Nat Biomed Eng 5, 600–612 (2021).
Callaway, E. How generative AI is building better antibodies. Nature 617, 235–235 (2023).
Olsen, T. H., Boyles, F. & Deane, C. M. Observed Antibody Space: A diverse database of cleaned, annotated, and translated unpaired and paired antibody sequences. Protein Sci 31, 141–146 (2022).
Berman, H. M. et al. The protein data bank. Nucleic acids research 28, 235–242 (2000).
Burley, S. K. et al. RCSB Protein Data Bank (RCSB. org): delivery of experimentally-determined PDB structures alongside one million computed structure models of proteins from artificial intelligence/machine learning. Nucleic acids research 51, D488–D508 (2023).
The UniProt, C. UniProt: the universal protein knowledgebase. Nucleic Acids Res 45, D158–D169 (2017).
Schneider, C., Raybould, M. I. J. & Deane, C. M. SAbDab in the age of biotherapeutics: updates including SAbDab-nano, the nanobody structure tracker. Nucleic Acids Res 50, D1368–D1372 (2022).
Miller, N. L., Clark, T., Raman, R. & Sasisekharan, R. Learned features of antibody-antigen binding affinity. Front Mol Biosci 10, 1112738 (2023).
Tsuruta, H. et al. AVIDa-hIL6: a large-scale VHH dataset produced from an immunized alpaca for predicting antigen-antibody interactions. Advances in Neural Information Processing Systems 36, 42077–42096 (2023).
Shanehsazzadeh, A. et al. Unlocking de novo antibody design with generative artificial intelligence. BioRxiv, 2023.01. 08.523187 (2023).
Swanson, K., Wu, W., Bulaong, N.L., Pak, J.E. & Zou, J. The Virtual Lab of AI agents designs new SARS-CoV-2 nanobodies. Nature 1–3 (2025).
Perron, M. D. et al. Allosteric noncompetitive small molecule selective inhibitors of CD45 tyrosine phosphatase suppress T-cell receptor signals and inflammation in vivo. Molecular pharmacology 85, 553–563 (2014).
Michalewicz, K., Barahona, M. & Bravi, B. ANTIPASTI: Interpretable prediction of antibody binding affinity exploiting normal modes and deep learning. Structure 32, 2422–2434. e5 (2024).
Gottlieb, A., Stein, G. Y., Oron, Y., Ruppin, E. & Sharan, R. INDI: a computational framework for inferring drug interactions and their associated recommendations. Molecular systems biology 8, 592 (2012).
Wilton, E. E., Opyr, M. P., Kailasam, S., Kothe, R. F. & Wieden, H.-J. sdAb-DB: the single domain antibody database. ACS Synthetic Biology 7, 2480–2484 (2018).
Abanades, B. et al. The Patent and Literature Antibody Database (PLAbDab): an evolving reference set of functionally diverse, literature-annotated antibody sequences and structures. Nucleic Acids Research 52, D545–D551 (2024).
Ferdous, S. & Martin, A. C. AbDb: antibody structure database—a database of PDB-derived antibody structures. Database 2018, bay040 (2018).
Sirin, S., Apgar, J. R., Bennett, E. M. & Keating, A. E. AB-Bind: Antibody binding mutational database for computational affinity predictions. Protein Science 25, 393–409 (2016).
Ma, Y., Yu, D., Wu, T. & Wang, H. PaddlePaddle: An open-source deep learning platform from industrial practice. Frontiers of Data and Domputing 1, 105–115 (2019).
Wang, R., Fang, X., Lu, Y., Yang, C.-Y. & Wang, S. The PDBbind database: methodologies and updates. Journal of medicinal chemistry 48, 4111–4119 (2005).
Jankauskaitė, J., Jiménez-García, B., Dapkūnas, J., Fernández-Recio, J. & Moal, I. H. SKEMPI 2.0: an updated benchmark of changes in protein–protein binding energy, kinetics and thermodynamics upon mutation. Bioinformatics 35, 462–469 (2019).
DeOnna, J. DACUM: A versatile competency-based framework for staff development. Journal for Nurses in Professional Development 18, 5–11 (2002).
Yue, Y. et al. MpbPPI: a multi-task pre-training-based equivariant approach for the prediction of the effect of amino acid mutations on protein–protein interactions. Briefings in Bioinformatics 24, bbad310 (2023).
Su, M. et al. Comparative assessment of scoring functions: the CASF-2016 update. Journal of chemical information and modeling 59, 895–913 (2018).
Wu, Y. Antibody and Nanobody Design Dataset (ANDD). Zenodo https://doi.org/10.5281/zenodo.18151718 (2025).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Giudicelli, V. & Lefranc, M. P. IMGT-ONTOLOGY 2012. Front. Genet. 3, 79 (2012).
Agarwal, A.A. et al. AlphaBind, a domain-specific model to predict and optimize antibody–antigen binding affinity. in Mabs 17 2534626 (Taylor & Francis, 2025).
Luo, S. et al. Antigen-specific antibody design and optimization with diffusion-based generative models for protein structures. Advances in Neural Information Processing Systems 35, 9754–9767 (2022).
Shrestha, P. et al. NanoBinder: a machine learning assisted nanobody binding prediction tool using Rosetta energy scores. Journal of Cheminformatics 17, 96 (2025).
Baek, M. et al. Accurate prediction of protein structures and interactions using a three-track neural network. Science 373, 871–876 (2021).
Prihoda, D. et al. BioPhi: A platform for antibody design, humanization, and humanness evaluation based on natural antibody repertoires and deep learning. in MAbs 14 2020203 (Taylor & Francis, 2022).
Boeckhout, M., Zielhuis, G. A. & Bredenoord, A. L. The FAIR guiding principles for data stewardship: fair enough? European journal of human genetics 26, 931–936 (2018).
Author information
Authors and Affiliations
Contributions
Conceptualization: Yikai Wu; methodology: Yikai Wu; investigation, formal analysis, writing, and visualization: Yikai Wu and Xuejiao Liu; review and editing: Yikai Wu, Karin Hrovatin, Dezhi Wu, and Stephanie Linker; supervision: Mathias Winkel and Feng Tan. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wu, Y., Liu, X., Hrovatin, K. et al. A Unified Dataset for Antibody and Nanobody Design Including Sequence, Structure, and Binding Affinity Data. Sci Data 13, 295 (2026). https://doi.org/10.1038/s41597-026-06878-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-026-06878-0