Abstract
The characterization of biodiversity is a crucial element of ecological investigations as well as environmental assessment and monitoring activities. Increasingly, amplicon-based environmental DNA metabarcoding (alternatively, marker gene metagenomics) is used for such studies given its ability to provide biodiversity data from various groups of organisms simply from analysis of bulk environmental samples such as water, soil or sediments. The Illumina MiSeq is currently the most popular tool for carrying out this work, but we set out to determine whether typical studies were reading enough DNA to detect rare organisms (i.e., those that may be of greatest interest such as endangered or invasive species) present in the environment. We collected sea water samples along two transects in Conception Bay, Newfoundland and analyzed them on the MiSeq with a sequencing depth of 100,000 reads per sample (exceeding the 60,000 per sample that is typical of similar studies). We then analyzed these same samples on Illumina’s newest high-capacity platform, the NovaSeq, at a depth of 7 million reads per sample. Not surprisingly, the NovaSeq detected many more taxa than the MiSeq thanks to its much greater sequencing depth. However, contrary to our expectations this pattern was true even in depth-for-depth comparisons. In other words, the NovaSeq can detect more DNA sequence diversity within samples than the MiSeq, even at the exact same sequencing depth. Even when samples were reanalyzed on the MiSeq with a sequencing depth of 1 million reads each, the MiSeq’s ability to detect new sequences plateaued while the NovaSeq continued to detect new sequence variants. These results have important biological implications. The NovaSeq found 40% more metazoan families in this environment than the MiSeq, including some of interest such as marine mammals and bony fish so the real-world implications of these findings are significant. These results are most likely associated to the advances incorporated in the NovaSeq, especially a patterned flow cell, which prevents similar sequences that are neighbours on the flow cell (common in metabarcoding studies) from being erroneously merged into single spots by the sequencing instrument. This study sets the stage for incorporating eDNA metabarcoding in comprehensive analysis of oceanic samples in a wide range of ecological and environmental investigations.
Similar content being viewed by others
Introduction
The inventorying and monitoring of biological diversity is a fundamental component of ecological and environmental studies. Additionally, characterizing biodiversity is part of the environmental impact assessments and ongoing environmental monitoring that are required by industry operating in environmentally sensitive locations1. Stakeholders are increasingly becoming more concerned about environmental stewardship, and this applies equally to the terrestrial2, freshwater3, and marine environments4 and covers all major taxonomic groups. A recent United Nations conference (UN Biodiversity Conference, Egypt, November 2018), highlighted the need for monitoring and protecting biodiversity with the key message of “investing in biodiversity for people and planet”. Despite the extreme importance of these efforts, the technology for carrying out biodiversity assessments has remained static for decades, relying heavily on observational data and capturing whole organisms from their environment for morphological analysis. Unfortunately, these procedures are error-prone, time-consuming, expensive, and tend to ignore small but ecologically important flora and fauna simply because they are difficult to identify visually2,5.
Over the past decade, increasing attention has been paid to the analysis of environmental DNA (eDNA)—a combination of DNA from whole cellular material or that is shed from organisms as they move through their environment. The existence of large reference databases (especially the common “DNA barcode” marker, cytochrome oxidase c subunit 1, or COI6,7) with the power of modern DNA sequencing instruments, enables environmental metabarcoding—the identification of many individual species from a simple water or sediment sample. Environmental metabarcoding is much faster than conventional techniques, is less labour-intensive, does not rely on the expertise of taxonomists, and produces orders of magnitude more information8.
Many side-by-side comparisons have been made between traditional morphological assessments and eDNA-based assessments. In all cases, eDNA is capable of detecting far more taxa overall. However, many of these studies also find that some organisms detected by traditional methods in the environment fail to be detected through metabarcoding1,4,9,10,11,12,13. There are a number of potential reasons for this discrepancy: the use of “universal” primers that don’t amplify some taxa as well as others5; employing markers that have biased representation in reference databases14; or an inadequate depth of sequencing to detect eDNA that is in low abundance. These factors are especially important when eDNA analyses are performed to track specific target organisms that might be present in low abundance in complex settings such as the oceans (e.g., endangered species or invasive species).
The goal of the present study was to investigate the influence of sequencing depth and more advanced workflow, including a patterned flowcell, offered by illumina’s NovaSeq platform on the ability to detect biological diversity present in a sample. To carry out this work, we analyzed samples on an Illumina MiSeq instrument at a sequencing depth that is typical of similar studies (Table 1), then analyzed those same PCR products on an Illumina NovaSeq 6000—the most advanced HTS instrument available today—with a sequencing depth approximately 700 times greater than that of the MiSeq. Not surprisingly, the NovaSeq detected many more taxa than the MiSeq: specifically, with one marker the NovaSeq detected 200% more metazoan families than the MiSeq. Contrary to our expectations, the NovaSeq still outperformed the MiSeq even when we subsampled the data to make depth-for-depth comparisons, suggesting that the NovaSeq has superior qualities beyond its much greater sequencing capacity.
Results
The Illumina MiSeq is currently the most popular metabarcoding platform
Twenty of the most-recently indexed papers in Google Scholar featuring the “metabarcoding” keyword were obtained in early November 2018 to perform a mini-metanalysis of the instrument most such studies are favouriting at the moment, as well as the depth of sequencing per sample that is generally employed. As shown in Table 1, the Illumina MiSeq is by far the instrument of choice presently, having been used by 14 (70%) of these studies. Sampling depth was not always reported clearly but was inferred where possible. Among these studies there was an extremely wide variance in sequencing depth, ranging from less than 10,000 reads per sample to nearly 900,000. However, the median was 60,000 (with a median absolute deviation of 55,000).
The NovaSeq finds more ESVs per sample than the MiSeq, even at the same sequencing depth
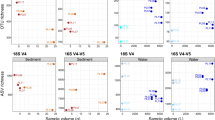
Based on our literature survey (Table 1), we decided to analyze our own samples on the MiSeq with a targeted sequencing depth of 100,000 reads per amplicon per sample—approximately 50% greater than the median sequencing depth of those studies. Two amplicons were analyzed, FishE and F230, both located with the standard barcode region of the mitochondrial gene cytochrome oxidase C subunit 1 (COI). Post-filtering, the mean depth per sample was 118,290 reads for the FishE marker and 84,500 for the F230 marker. We then analyzed these same PCR products on the Illumina NovaSeq 6000 at much greater depth, averaging 7 million reads per amplicon per sample. The resulting reads were processed using the DADA2 pipeline as described in the Methods. Perhaps not surprisingly given the ~700x greater sequencing depth, the NovaSeq was able to find more exact sequence variants (ESVs) in each sample than the MiSeq. To our surprise, however, even after rarefying the NovaSeq data to match the sequencing depth of the MiSeq, we still found greater diversity (i.e., more ESVs) within the NovaSeq data for the FishE (Fig. 1) and F230 (Supplementary Fig. 1) amplicons. Moreover, while there was substantial overlap between the ESVs found between the two platforms, the MiSeq had very few ESVs unique to itself while the NovaSeq found many ESVs that the MiSeq missed. We highlight that the exact same PCR products were used for both instruments, so these results cannot be the consequence of stochastic PCR biases. The two sites with higher diversity—7 and 8—are located close to shore and site 7 in particular is close to a wastewater outlet (see Methods).
For each biological replicate (A–C) within each sampling site (1 to 8), the NovaSeq (light bars) was able to find a greater number of ESVs than the MiSeq (dark bars) even when the NovaSeq data are subsampled to match the sequencing depth of the MiSeq. Sites 7 and 8 are very close to shore, and site 7 in particular is near a wastewater outlet.
This trend is even more pronounced when plotted as an accumulation curve
When we combined all samples and then performed subsampling experiments to generate accumulation curves, the ability of the NovaSeq to detect new ESVs becomes even more stark: at each simulated sequencing depth, the NovaSeq detects greater biological diversity (i.e., ESVs) than the MiSeq (Fig. 2 for the FishE amplicon; see Supplementary Fig. 2 for the F230 amplicon). Curiously, while greater depth seems to reveal increasing numbers of ESVs on the NovaSeq (even beyond 2.5 million reads/sample), it is not clear that greater depth adds any new information for the MiSeq: the number of ESVs detected appears to level off at approximately 5,000. This strongly suggests that the NovaSeq outperforms the MiSeq in a depth-independent manner.
This trend is not an artefact of the DADA2 error-correcting model
DADA2 generates ESVs by applying an error correction model to raw FASTQ files, attempting to fix errors that were introduced through PCR or sequencing15. However, while the MiSeq reports base call qualities using pseudo-continuous Phred scores that can range from 0–40, the NovaSeq’s FASTQ files bin qualities into just four levels16. We therefore suspected that the phenomenon we were observing might be an artefact of the DADA2 program. Specifically, we hypothesized that the algorithm might be under-correcting errors in the NovaSeq data leading to a spurious increase in the number of ESVs. For this reason we repeated our analysis with simple OTU clustering at a 97% similarity threshold (described in greater detail in the Methods). OTU clustering applies no error correction model at all and is simply based on sequence similarity measures, and should therefore have the same performance on NovaSeq data as it does on MiSeq data. To our surprise, when accumulation curves were generated to compare the two instruments depth-for-depth, the NovaSeq once again outperformed the MiSeq (Fig. 3).
We note that Fig. 2 and Fig. 3 look quite different from each other in two ways: (1) the number of OTUs detected is far greater than the number of ESVs; and (2) while the number of new ESVs levels out for the MiSeq in Fig. 2, the trajectory continues upward for the OTUs in Fig. 3. This is due to the very different methodologies employed to generate ESVs versus OTUs. Because OTU clustering makes no attempt to model and correct for PCR and sequencing errors, the raw number of OTUs is expected to be much greater than the number of ESVs detected—many OTUs are simply the product of the accumulation of errors. By similar reasoning, both the MiSeq and NovaSeq OTU curves continue to rise with greater sequencing depth because additional sequencing errors will be encountered with that greater depth.
Greater sequencing depth on the MiSeq cannot achieve the level of diversity detected on the NovaSeq
The MiSeq’s accumulation curve in Fig. 2 suggests that additional sequencing depth would not increase the number of ESVs detected, but to thoroughly test this point we re-ran three samples (sites 1, 3, and 6, each with three biological replicates for a total of 9 samples) on the MiSeq at much greater sequencing depth—approximately 1 million reads per amplicon per sample—and then compared these data to the NovaSeq data. As illustrated for the FishE amplicon in Fig. 4, adding this greater sequencing depth to the MiSeq only marginally improves its detection of diversity from the samples. Conversely, the NovaSeq continues to detect new ESVs with greater sequencing depth. Note that the total number of ESVs detected is lower than that of Fig. 2, but this is because of the smaller number of samples analyzed in this experiment (three sites versus eight). Again, we repeated this experiment with the F230 amplicon and found the same trend (see Supplementary Fig. 3).
Figure 4 shows that even beyond 5 million reads the NovaSeq was still finding new ESVs with no sign of plateauing. As before, we suspected this might be the result of the DADA2 algorithm under-correcting sequencing errors in the NovaSeq data. To examine this possibility, we ran the accumulation curve out to its maximum and found that the curve does indeed hit a plateau of just over 9,000 ESVs at a sequencing depth of approximately 10 million reads (Fig. 5). This result indicates that the pattern observed for NovaSeq data are not likely to be an artefact of the DADA2 analysis. Moreover, it indicates that extremely deep sequencing is required if one wants to have a comprehensive survey of biodiversity in a region.
The MiSeq is less capable of sequencing low-abundance eDNA
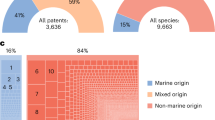
Our NovaSeq results indicate that in the locations in which we sampled, approximately 9,300 FishE ESVs are present (Fig. 5). However, the MiSeq was only able to obtain approximately 3,500 ESVs even at an unrealistically-high sequencing depth (Fig. 4). This suggests that the MiSeq could not identify approximately 60% of the diversity present in the environment. In order to determine taxonomic/biological breadth of these ESVs we performed taxonomic assignment on all the ESVs from both instruments—i.e., we combined the results of both MiSeq runs to give that platform the best possible chance of finding all the taxa present—and found that the MiSeq data contained 80 identifiable families. The NovaSeq also identified these same 80 families but was also able to identify an additional 32—a 40% increase. Those families unique to the NovaSeq analysis are listed in Table 2. Some of the taxa missing from the MiSeq data are of significant interest, including marine mammals (Delphinidae) and several fish. Other taxa include those that are clearly not marine organisms (e.g., cow and moose) but this is not surprising given the sampling sites’ close proximity to a human-populated shore, and still demonstrates that organisms with presumably low-abundance eDNA are less likely to be detected by the MiSeq than the NovaSeq.
Interestingly, when these taxa are plotted on a circular dendrogram we do not observe any obvious phylogenetic pattern to the distribution of missing taxa on the MiSeq (Fig. 6). Rather, it appears that the NovaSeq was generally able to detect more families within each order than the MiSeq.
Although we have no quantitative measurements of the abundance of taxa present in the locations we sampled, we note that many of the taxa missing from the MiSeq analysis are likely to have a very low abundance of eDNA (e.g., marine mammals, terrestrial organisms) compared to taxa where whole organisms or gametes may be present in the water samples (e.g., zooplankton). We can approximate this by looking at read abundances (Fig. 7). If we assume that read abundances roughly correlate with the original biomass present17 then it does indeed seem that the MiSeq is less capable of sequencing this low abundance eDNA than the NovaSeq, even at very high sequencing depths.
Families detected on the NovaSeq ranked by read count (note the y-axis has a logarithmic scale). The white bars indicate taxa that were detected by both the NovaSeq and the MiSeq, while black bars indicate taxa detected solely by the NovaSeq. We note that most of the taxa missed by the MiSeq have low read abundance.
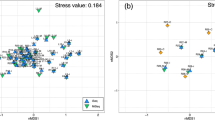
Whether or not this phenomenon will have a significant impact on a particular experiment depends strongly on the purpose of the experiment. Those that are attempting to detect or catalogue rare or endangered species may be strongly impacted, since these are precisely the organisms that the MiSeq is likely to miss. On the other hand, general biodiversity assays or comparative studies in community composition will be less impacted by missing these low-abundance taxa. To illustrate the point, we generated NMDS plots for genera identified within the 9 samples (3 sites) that were deep-sequenced on the MiSeq and NovaSeq. Even though the NovaSeq detected more genera overall, qualitatively both instruments pick up a gradient change from coastal to deeper waters along the primary axis (Fig. 8).
Discussion
Most environmental metabarcoding studies are not sequencing deep enough
Our results suggest that using seawater as the source of environmental DNA at a typical sequencing depth of 60,000 reads per sample, only half of the diversity detectable by the MiSeq will be captured (Fig. 2). To reach the MiSeq’s detection limits for analysis of seawater one would have to aim for 0.8–1 million reads per sample per marker—more than ten times the typical depth of sequencing currently performed in most metabarcoding studies. We further note that the samples from our study were obtained from the North Atlantic Ocean where biodiversity is presumably far less than that which might be found in tropical regions. For this reason, even deeper sequencing may be required in regions or substrates that have very high biodiversity.
Even when matched for depth, the NovaSeq can detect greater diversity than the MiSeq
The most remarkable finding in this study is that the NovaSeq can detect many taxa that the MiSeq cannot—even when the depth of sequencing is matched. This is true on a PCR -by-PCR basis (Fig. 1), and even greater depth of sequencing on the MiSeq cannot overcome this obstacle (Fig. 4). The outcome is that there may be a great deal of missing biodiversity in MiSeq analyses (Fig. 6).
Whether or not this has a significant impact on a study will depend on the nature of that study. The inability to detect low-abundance taxa is unlikely to have a large impact on comparative community composition analyses. On the other hand, studies that have an interest in low-abundance taxa (e.g., those that are rare or endangered) could be very significantly impacted. We also note that many studies comparing eDNA-based approaches to traditional morphological methods frequently show eDNA-based methods missing taxa that were observed visually. We wonder if some of these cases could be explained through the lower sensitivity of the MiSeq to low-abundance eDNA.
Possible causes for the difference between platforms
The NovaSeq has many differences from the MiSeq, including: a 2-colour chemistry rather than 4-colour used by the MiSeq; greatly improved hardware (presumably including better image capture abilities); different signal processing software; and a flow cell that has pre-defined binding spots for target DNA instead of the random lawn used by previous Illumina instruments. We suspect but cannot be certain that this last factor—the flow cell—is a significant cause for the NovaSeq’s superior performance. In the MiSeq and most previous Illumina instruments, DNA binds to the flow cell in a random fashion. Therefore, to distinguish one spot on the flow cell where DNA has bound from another, the spots are observed by the instrument for the first 25 rounds of sequencing and at that point clusters are determined18. This works well when performing shotgun sequencing (the primary use of Illumina’s instruments) because the spots can be clearly distinguished from each other thanks to the high level of sequence diversity. However, when performing amplicon-based sequencing the variability from one spot to the next—especially within the first 25 bases which likely covers primer regions—is minimal and this can cause two distinct spots to be merged together. To prevent this from happening, Illumina recommends spiking in PhiX genome18, but unless the proportion of PhiX is very high it’s nearly impossible to prevent similar sequences from sitting near each other on the flow cell. Conversely, the spots that DNA anneal to on the NovaSeq flow cell are pre-defined and known by the instrument’s base calling software, so inferring their location is not necessary and this largely prevents the “over-clustering” of low diversity reads.
Notably, the MiSeq runs that we performed for this project used the recommended levels of PhiX and the sequencing run statistics all matched Illumina recommendations. Nevertheless, we still suspect that over-clustering is at least partially responsible for the MiSeq missing out on diversity that the NovaSeq was able to detect.
Our results suggest that the NovaSeq 6000 may be a superior instrument for environmental metabarcoding studies especially in complex biodiversity-rich substrates where heterogenous abundance of taxonomic groups may confound detection. However, while the MiSeq is a relatively inexpensive instrument that could plausibly be obtained by many labs, the NovaSeq is expensive—roughly ten times the cost of the MiSeq. Moreover, while it produces hundreds of times more data per run, each run is also significantly more expensive: the smallest currently available NovaSeq flow cell and sequencing kit is approximately ten times more expensive than the popular MiSeq v3 600-cycle kit. For these reasons, the NovaSeq may be out of reach for many laboratories in the near term and we therefore suggest a few approaches that may aid obtaining more comprehensive biodiversity from the MiSeq.
Multiplex different markers together on the same flow cell
This is already quite a common practice, albeit frequently for money-saving purposes rather than to prevent over-clustering. In theory, multiplexing several markers (while still maintaining adequate sequencing depth per sample), will lead to greater sequence diversity on the flow cell and will reduce the probability of over-clustering.
Use dual-indices on the sequencing primers
It is common practice to add short (e.g., 8mer) oligonucleotide indices to the sequencing primers for the purposes of multiplexing, but this has the added benefit of contributing to additional base composition diversity on the flow cell. It should be noted that we did employ a dual-index strategy in the present paper (see the Methods), so this strategy alone does not seem sufficient to close the gap between the MiSeq and NovaSeq instruments.
Use large amounts of PhiX spike-in
PhiX genome fragments also serve to increase the complexity on the flow cell, but many labs try to minimize the amount of PhiX they use because it takes up precious sequencing capacity. Paradoxically though, increasing PhiX may in fact increase the number of quality reads generated. Illumina’s own recommendations range from 5–50%19 although in practice most experiments end up at the lower end of this range.
Use phased amplicons
Another approach was suggested by20, who designed overlapping 16 s amplicons that they described as “phased amplicon sequencing”. Despite covering the same region of interest, the different sequence composition at the 5′ end of the read reduced over-clustering by the MiSeq—so much so that the number of reads passing quality filters increased by 9–47% in their experiments.
Conclusions
Biodiversity analysis through genomics has enabled widespread applications from human microbiome studies to environmental assessment and monitoring. With rapid advances in sequencing hardware and computational approaches for data analysis, it is important to determine the impact that sequencing technology and strategy have on the data generated, especially where it may influence biological interpretations and their socio-economic implications. Here, we tackled the issue of sequencing technology and depth on an analysis of biodiversity in seawater through eDNA metabarcoding. Our analysis provides direct evidence of the superior utility of the newly introduced NovaSeq platform for elucidating a more comprehensive biodiversity measurement as compared to the current workhorse, the MiSeq platform. Our results strongly suggest that comprehensive detection of biota from eDNA in a complex environment such as the ocean is possible and will aid supporting scientific/societal endeavours for enhanced biodiversity analysis for people and the planet.
Methods
Sample collection
Triplicate 250 mL water samples were taken from surface water simultaneously. Samples were taken from eight locations along two transects in Conception Bay, Newfoundland and Labrador, Canada, on October 13–14, 2017 (Fig. 9). These samples cover a range from near-shore to approximately 10 km offshore (with a sea bottom depth ranging from a few metres nearshore to approximately 200 metres in the middle of the bay).
Laboratory procedures
DNA extraction
Filtration and DNA extraction was done in PCR clean laminar flow hoods (AirClean Systems) thoroughly decontaminated with ELIMINase (Decon Labs) and 70% EtOH prior to each set of three sample replicates. Water samples were thawed at 4 °C and immediately filtered with 0.22 µm PVDF Sterivex filters (MilliporeSigma). The DNeasy PowerWater Kit (Qiagen) was used to extract DNA with the automated QIAcube platform (Qiagen), following the DNeasy PowerWater IRT protocol. For lysis, bead tubes were heated for five minutes at 65 °C and then vortexed for ten minutes. Negative controls were generated during filtration and extraction to screen for contamination and cross-contamination. Filtration and extraction were done in a pre-PCR room isolated from post-PCR rooms.
DNA library preparation
Two fragments were amplified by PCR from the 5′ end of the standard COI barcode region: the 235 bp F230 fragment10, and the 232 bp Mini_SH-E fragment21. Illumina-tailed PCR primers (tails underlined) were used to amplify targets: The F230 forward primer (LCO1490; 5′-TCG TCG GCA GCG TCA GAT GTG TAT AAG AGA CAG GGT CAA CAA ATC ATA AAG ATA TTG G-3′;22), the Mini_SH-E forward primer (Fish_miniE_F_t; 5′-TCG TCG GCA GCG TCA GAT GTG TAT AAG AGA CAG ACY AAI CAY AAA GAY ATI GGC AC-3′), and the reverse primer (230_R/Fish_miniE_R_t; 5′-GTC TCG TGG GCT CGG AGA TGT GTA TAA GAG ACA GCT TAT RTT RTT TAT ICG IGG RAA IGC-3′) was used for both F230 and Mini_SH-E fragments. Each amplification reaction contained 0.6 µL DNA, 1.5 µL 10X reaction buffer, 0.6 µL MgCl2 (50 mM), 0.3 µL dNTPs mix (10 mM), 0.3 µL of each Illumina-tailed primer (10 µM), and 0.3 µL Platinum Taq (0.5 U/µL; Invitrogen) in a total volume of 15 µL. PCR conditions were initiated with a heated lid at 95 °C for 3 mins, followed by 35 cycles of 94 °C for 30 s, 46 °C for 40 s, and 72 °C for 1 min, and a final extension at 72 °C for 10 mins. Three PCR replicates were amplified from each sample with the ProFlex thermocycler (Thermo Fisher) and then pooled for a single PCR cleanup with the QIAquick 96 PCR purification kit (Qiagen; 60 µL elution volume). Agarose (1.5% w/v) gel electrophoresis was used to verify amplification of samples, and for quality control of negative controls from PCR, extraction, filtration, and field collection. Each indexing reaction contained 2 µL amplicon DNA, 2.5 µL 10X reaction buffer, 1 µL MgCl2 (50 mM), 0.5 µL dNTPs mix (10 mM), 1 µL of F indexing primer (5 µM), 1 µL R indexing primer (5 µM), and 0.5 µL Platinum Taq (0.5U/µL; Invitrogen) in a total volume of 25 µL. Unique dual Nextera indexes were used to mitigate index misassignment (IDT; 8-bp index codes). PCR conditions were initiated with a heated lid at 95 °C for 3 mins, followed by 12 cycles of 95 °C for 30 s, 55 °C for 30 s, and 72 °C for 30 s, and a final extension at 72 °C for 5 mins. Indexing success was verified on the Bioanalyzer (Agilent) with the DNA 7500 kit. Samples were quantified with Quant-iT PicoGreen dsDNA assay with a Synergy HTX plate fluorometer (BioTek) and pooled to normalize DNA concentration. Libraries were cleaned with three successive AMPure XP cleanups: Left side selection with bead:DNA ratios of 1×, then 0.9×, and a right-side selection with 0.5×. Libraries were quantified with a Qubit fluorometer (Thermo Fisher) and the size distribution was checked with the DNA 7500 kit on the Agilent 2100 Bioanalyzer. Two libraries containing F230 or FishE amplicons from nine samples were sequenced on the Illumina MiSeq with two 600-cycle v3 kits. Two libraries containing F230 or FishE amplicons from 24 samples were pooled with other libraries and sequenced with two MiSeq. 600-cycle v3 kits. Field, filtration and extraction negatives were also sequenced in these MiSeq runs. Two libraries containing F230 and FishE amplicons were sequenced with a 300-cycle S4 kit on the NovaSeq 6000 following the NovaSeq XP workflow.
Bioinformatics
We employed two different workflows to analyze our data in order to reduce the possibility that our results were an artefact of the method used. In both workflows, base calling and demultiplexing were performed using Illumina’s bcl2fastq software (version 2.20.0.422). Primers were then trimmed from the forward and reverse reads using cutadapt v1.1623 with the default error tolerance and a minimum overlap equal to half the primer length. We discarded read pairs in which the primer was missing from either the forward or reverse read. We note that both amplicons studied in this paper are very short (~230 bases) and therefore there is ample overlap between the forward and reverse reads in both the MiSeq (300 cycle forward and reverse) and NovaSeq (150 cycle forward and reverse) data. We saw no significant difference in the rate of successful paired-end joining between the two instruments.
After this stage the two methodologies diverged and are described separately below.
DADA2 workflow: DADA2 v1.8.015 was used to perform quality filtering and joining of paired reads (maxEE = 2, minQ = 2, truncQ = 2, maxN = 0), and denoising (using default parameters) to produce exact sequence variants (ESVs). This was performed independently on the MiSeq and NovaSeq data since their error patters are presumed to be different and therefore they require different models to be trained. Singleton ESVs were discarded. To rapidly evaluate the overlap in ESVs between the two instruments, MD5 hashes24 were generated for each of the ESV sequences and then these sets of hashes were compared between the MiSeq and the NovaSeq.
OTU clustering workflow: Vsearch v2.8.225 was first used to join the paired ends of the reads (using default parameters), and perform quality filtering (using default parameters). The reads for the NovaSeq and MiSeq were then dereplicated, and these reads were combined into a single file so that OTU clustering (using the cluster_fast setting) could be performed on the entire set using an identity threshold of 97%. As with the ESVs, singleton OTUs were discarded.
Taxonomic assignment: NCBI’s blastn tool v2.6.026 was used to compare ESV sequences against the nt database (downloaded August 2018), using an e-value cut-off of 0.001. We filtered the resulting hits with the requirement of having at least 90% identities across at least 90% of the query sequence. In cases where there was not a single unambiguous best hit, we used a majority consensus threshold of 80% to assign taxonomy27.
Accumulation curves: Original read memberships were tracked through the various analytical steps: dereplication followed by OTU clustering, or ESV generation using DADA2. Subsamples were then generated using sampling proportional to the original read abundances with the “choices” function within the Python programming language’s “random” module28. These reads were then mapped to their respective ESVs/OTUs for comparison between the two DNA sequencing platforms.
NMDS plots: NMDS plots were generated using the default settings of the metaMDS function, part of the vegan library29 in the statistical package R30. Data were based on genera in both rarefied NovaSeq and whole MiSeq data that could be identified with 95% or better identity across 95% or more of the read.
Data Availability
All data have been deposited into NCBI’s Sequence Read Archive under accession number PRJNA513845.
References
Aylagas, E., Borja, Á. & Rodríguez-Ezpeleta, N. Environmental Status Assessment Using DNA Metabarcoding: Towards a Genetics Based Marine Biotic Index (gAMBI). PLOS ONE 9, e90529 (2014).
Baird, D. J. & Hajibabaei, M. Biomonitoring 2.0: a new paradigm in ecosystem assessment made possible by next-generation DNA sequencing. Mol. Ecol. 21, 2039–2044 (2012).
Veldhoen, N. et al. Implementation of Novel Design Features for qPCR-Based eDNA Assessment. PLOS ONE 11, e0164907 (2016).
Aylagas, E., Borja, Á., Muxika, I. & Rodríguez-Ezpeleta, N. Adapting metabarcoding-based benthic biomonitoring into routine marine ecological status assessment networks. Ecol. Indic. 95, 194–202 (2018).
Taberlet, P., Coissac, E., Pompanon, F., Brochmann, C. & Willerslev, E. Towards next-generation biodiversity assessment using DNA metabarcoding. Mol. Ecol. 21, 2045–2050 (2012).
Hebert, P. D. N., Cywinska, A., Ball, S. L. & deWaard, J. R. Biological identifications through DNA barcodes. Proc. Biol. Sci. 270, 313–321 (2003).
Ratnasingham, S. & Hebert, P. D. N. bold: The Barcode of Life Data System (http://www.barcodinglife.org). Mol. Ecol. Notes 7, 355–364 (2007).
Hajibabaei, M., Shokralla, S., Zhou, X., Singer, G. A. C. & Baird, D. J. Environmental barcoding: a next-generation sequencing approach for biomonitoring applications using river benthos. PloS One 6, e17497 (2011).
Ji, Y. et al. Reliable, verifiable and efficient monitoring of biodiversity via metabarcoding. Ecol. Lett. 16, 1245–1257 (2013).
Gibson, J. F. et al. Large-Scale Biomonitoring of Remote and Threatened Ecosystems via High-Throughput Sequencing. PloS One 10, e0138432 (2015).
Shaw, J. L. A. et al. Comparison of environmental DNA metabarcoding and conventional fish survey methods in a river system. Biol. Conserv. 197, 131–138 (2016).
DiBattista, J. D. et al. Assessing the utility of eDNA as a tool to survey reef-fish communities in the Red Sea. Coral Reefs 36, 1245–1252 (2017).
Cahill, A. E. et al. A comparative analysis of metabarcoding and morphology-based identification of benthic communities across different regional seas. Ecol. Evol. 8, 8908–8920 (2018).
Hajibabaei, M., Baird, D. J., Fahner, N. A., Beiko, R. & Golding, G. B. A new way to contemplate Darwin’s tangled bank: how DNA barcodes are reconnecting biodiversity science and biomonitoring. Phil Trans R Soc B 371, 20150330 (2016).
Callahan, B. J. et al. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
NovaSeq. 6000 System Quality Scores and RTA3 Software Available at: https://www.illumina.com/content/dam/illumina-marketing/documents/products/appnotes/novaseq-hiseq-q30-app-note-770-2017-010.pdf (2017).
Evans, N. T. et al. Quantification of mesocosm fish and amphibian species diversity via environmental DNA metabarcoding. Mol. Ecol. Resour. 16, 29–41 (2016).
Optimizing Cluster Density on Illumina Sequencing Systems Available at: https://www.illumina.com/content/dam/illumina-marketing/documents/products/other/miseq-overclustering-primer-770-2014-038.pdf (2016).
How much PhiX spike-in is recommended when sequencing low diversity libraries on Illumina platforms? Available at https://support.illumina.com/bulletins/2017/02/how-much-phix-spike-in-is-recommended-when-sequencing-low-divers.html. (Accessed: 14th November 2018)
Wu, L. et al. Phasing amplicon sequencing on Illumina Miseq for robust environmental microbial community analysis. BMC Microbiol. 15, (2015).
Shokralla, S., Hellberg, R. S., Handy, S. M., King, I. & Hajibabaei, M. A. DNA Mini-Barcoding System for Authentication of Processed Fish Products. Sci. Rep. 5, 15894 (2015).
Folmer, O., Black, M., Hoeh, W., Lutz, R. & Vrijenhoek, R. DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Mol. Mar. Biol. Biotechnol. 3, 294–299 (1994).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal 17, 10–12 (2011).
Rivest, R. RFC1321: The MD5 Message-Digest Algorithm. Available at: https://www.ietf.org/rfc/rfc1321.txt (1992).
Rognes, T., Flouri, T., Nichols, B., Quince, C. & Mahé, F. VSEARCH: a versatile open source tool for metagenomics. PeerJ 4, e2584 (2016).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Schloss, P. D. & Westcott, S. L. Assessing and Improving Methods Used in Operational Taxonomic Unit-Based Approaches for 16S rRNA Gene Sequence Analysis. Appl. Environ. Microbiol. 77, 3219–3226 (2011).
Van Rossum, G. Python Programming Language. In USENIX Annual Technical Conference (2007).
Oksanen, J. et al. vegan: Community Ecology Package Available at: https://CRAN.R-project.org/package=vegan (2018).
R Core Team. R: A Language and Environment for Statistical Computing. (2018). Available at: https://www.R-project.org/.
Abrams, J. F. et al. Shifting up a gear with iDNA: from mammal detection events to standardized surveys. bioRxiv 449165 https://doi.org/10.1101/449165 (2018).
Ando, H. et al. Evaluation of plant contamination in metabarcoding diet analysis of a herbivore. Sci. Rep. 8, 15563 (2018).
Burgess, T. I., McDougall, K. L., Scott, P. M., Hardy, G. E. S. & Garnas, J. Predictors of Phytophthora diversity and community composition in natural areas across diverse Australian ecoregions. Ecography 42, 565–577 (2019).
Cahoon, A. B., Huffman, A. G., Krager, M. M. & Crowell, R. M. A meta-barcoding census of freshwater planktonic protists in Appalachia – Natural Tunnel State Park, Virginia, USA. Metabarcoding Metagenomics 2, e26939 (2018).
Egeter, B. et al. Challenges for assessing vertebrate diversity in turbid Saharan water-bodies using environmental DNA. Genome 61, 807–814 (2018).
Gran‐Stadniczeñko, S. et al. Protist Diversity and Seasonal Dynamics in Skagerrak Plankton Communities as Revealed by Metabarcoding and Microscopy. Journal of Eukaryotic Microbiology Preview available at: https://onlinelibrary.wiley.com/doi/abs/ https://doi.org/10.1111/jeu.12700.
Holman, L. E. et al. The detection of novel and resident marine non-indigenous species using environmental DNA metabarcoding of seawater and sediment. bioRxiv 440768 https://doi.org/10.1101/440768 (2018).
Hugoni, M. et al. Spatiotemporal variations in microbial diversity across the three domains of life in a tropical thalassohaline lake (Dziani Dzaha, Mayotte Island). Mol. Ecol. 27, 4775–4786 (2018).
Kerdraon, L., Balesdent, M.-H., Barret, M., Laval, V. & Suffert, F. Crop residues in wheat-oilseed rape rotation system: a pivotal, shifting platform for microbial meetings. bioRxiv 456178 https://doi.org/10.1101/456178 (2018).
Nuske, S. J. et al. The endangered northern bettong, Bettongia tropica, performs a unique and potentially irreplaceable dispersal function for ectomycorrhizal truffle fungi. Mol. Ecol. 27, 4960–4971 (2018).
Phan, H. C., Wade, S. A. & Blackall, L. L. Is marine sediment the source of microbes associated with accelerated low water corrosion? Appl. Microbiol. Biotechnol. 103, 449–459 (2019).
Pochon, X., Wecker, P., Stat, M., Berteaux-Lecellier, V. & Lecellier, G. Towards an in-depth characterization of Symbiodiniaceae in tropical giant clams via metabarcoding of pooled multi-gene amplicons https://doi.org/10.7287/peerj.preprints.27313v2 (2019).
Qian, X. et al. Shifts in community composition and co-occurrence patterns of phyllosphere fungi inhabiting Mussaenda shikokiana along an elevation gradient. PeerJ 6, e5767 (2018).
Shahraki, A. H., Chaganti, S. R. & Heath, D. Assessing high-throughput environmental DNA extraction methods for meta-barcode characterization of aquatic microbial communities. J. Water Health 17, 37–49 (2019).
Siegenthaler, A. et al. Metabarcoding of shrimp stomach content: Harnessing a natural sampler for fish biodiversity monitoring. Mol. Ecol. Resour. 19, 206–220 (2019).
Too, C. C., Keller, A., Sickel, W., Lee, S. M. & Yule, C. M. Microbial Community Structure in a Malaysian Tropical Peat Swamp Forest: The Influence of Tree Species and Depth. Front. Microbiol. 9 (2018).
Vesterinen, E. J., Puisto, A. I. E., Blomberg, A. S. & Lilley, T. M. Table for five, please: Dietary partitioning in boreal bats. Ecol. Evol. 8, 10914–10937 (2018).
Voulgari-Kokota, A., Grimmer, G., Steffan-Dewenter, I. & Keller, A. Bacterial community structure and succession in nests of two megachilid bee genera. FEMS Microbiol. Ecol. 95 (2019).
Wohlrab, S. et al. Metatranscriptome Profiling Indicates Size-Dependent Differentiation in Plastic and Conserved Community Traits and Functional Diversification in Dinoflagellate Communities. Front. Mar. Sci. 5 (2018).
Zinger, L. et al. Body size determines soil community assembly in a tropical forest. Mol. Ecol. 28, 528–543 (2019).
Acknowledgements
The authors would like to thank Kirk Rees, Captain of the Abrigo, for assisting with sample collection. This work was partly funded through a Petroleum R&D Grant from InnovateNL (contract number 5405.2121.101), an award from the Atlantic Canada Opportunities Agency’s Atlantic Innovation Fund (project number 781-37749-207993), and a grant from Petroleum Research Newfoundland and Labrador. Any opinions, findings, and conclusions or recommendations expressed in this publication are those of the authors and do not necessarily reflect the views of Petroleum Research or its members.
Author information
Authors and Affiliations
Contributions
M.H. conceived the idea for this project and provided scientific oversight in experimental design, data analysis and interpretation and helped with writing the manuscript; N.A.F. designed and executed the sampling protocols; J.G.B. assisted with sampling and aided the bioinformatics analyses; A.M. led the laboratory work; G.S. performed the data analyses, figure generation, and wrote the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Singer, G.A.C., Fahner, N.A., Barnes, J.G. et al. Comprehensive biodiversity analysis via ultra-deep patterned flow cell technology: a case study of eDNA metabarcoding seawater. Sci Rep 9, 5991 (2019). https://doi.org/10.1038/s41598-019-42455-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-019-42455-9
This article is cited by
-
Metatranscriptomics-based metabolic modeling of patient-specific urinary microbiome during infection
npj Biofilms and Microbiomes (2025)
-
Transforming gastrointestinal helminth parasite identification in vertebrate hosts with metabarcoding: a systematic review
Parasites & Vectors (2024)
-
Mock community experiments can inform on the reliability of eDNA metabarcoding data: a case study on marine phytoplankton
Scientific Reports (2023)
-
River benthic macroinvertebrates and environmental DNA metabarcoding: a scoping review of eDNA sampling, extraction, amplification and sequencing methods
Biodiversity and Conservation (2023)
-
Impacts of dietary exposure to pesticides on faecal microbiome metabolism in adult twins
Environmental Health (2022)











Charles Warden
I am still waiting to be able to post something on Biostars (in terms of QC questions for barcoding), but I think this Twitter conversation currently represents some concerns that I would have:
https://twitter.com/pathoge...
Charles Warden Replied to Charles Warden
If anyone is interested, the barcode post is up on Biostars now:
https://www.biostars.org/p/...
You can also view more details from the IGC here, but you will importantly be missing out on the interactive discussion at Biostars.
The topic is slightly different, but I believe your methods indicate the samples for both machines were dual-barcoded.
If you don't mind, would it be possible for you to please go back and check the index quality scores (particularly for the second index, described as either I2 or i5) for the samples?
a) Is there increased diversity for samples with lower index quality scores?
b) For individual read assignments that may be unexpected (such as the cow and moose assignments that Nick pointed out), do those reads individually have lower quality scores (if separate FASTQ files are generated for those indices)? If not, that could add confidence that this relates to your observation that site 7 was close to a wastewater outlet (although the increased diversity seems to be more true for FishE than F230).
In the Biostars post, I don't go into the specifics for any particular projects, and I focus more on read loss when calling single-barcode samples as dual-barcode samples. Also, to be clear, I have not found an example of cross-contamination that could be explained from index quality scores. However, I would be interested to see if index quality scores had any influence on this study.
Charles Warden Replied to Charles Warden
Also, after having a chance to read the paper more carefully, I feel relatively more comfortable with the steps taken to critically assess the results. I also apologize for any confusion because I also trimmed down my response as I spent more time reviewing the paper.
Nevertheless, if anything can be done to better explain depth-matched difference between machines (MiSeq versus NovaSeq), then I think that would help (and, as much as possible, I will also try to continue to look into the paper and data more).
Thank you very much for putting together this paper for public discussion.
nickloman
Could the data be made available? The supplied NCBI BioProject accession does not currently work.
P SM Replied to nickloman
After publication, a reader can just ask genbank to release the data. It is up now:
https://www.ncbi.nlm.nih.go...
Charles Warden
I have aligned reads to the PhiX genome (used by Illumina). While I don’t know the details of the barcode design, there were no PhiX reads in the MiSeq samples but there were always some PhiX reads in the NovaSeq samples:
https://uploads.disquscdn.c...
In the interests of the fairness, I have plotted the percent PhiX on the left as well as the log10(absolute PhiX read counts) on the right. The counts are from samtools idxstats, so I have divided them by 2 (as an approximation from the paired-end alignments, having run Bowtie2 with the
--no-unalparameter).There is no barcode for the PhiX spike-ins, so there theoretically shouldn’t be any PhiX reads in the de-multiplexed samples (although the difference between machines is what I am more concerned about here).
You can also view IGV screenshots of the 3 samples with the most PhiX reads below:
https://uploads.disquscdn.c...
So, can you (or another post-publication reviewer), please check if the following affects the diversity metrics that you report?
1) The direct increase in PhiX sequences (if they weren’t filtered).
2) If higher diversity remains after filtering the PhiX sequences, do samples with relatively higher PhiX percentages tend to have higher diversity of other samples (which, in that situation, may be technical rather than biological)?
I also have some additional analysis in this discussion, but the impact on the PhiX percentages from the FASTQ files (instead of InterOp) has a different focus.
Charles Warden Replied to Charles Warden
To be fair, from what I understand, the MiSeq COI alignment rate seems low (but not all zero, like PhiX):
>EU273284_COI_CDS
atggcaatcacacgctgatttttctcaaccaatcacaaagatattggcaccctttatttagtatttggtgcttgagctggaatggtaggaacggctttaagcctcctaatccgagcagaattaagtcaaccaggcgcccttctaggggatgaccaaatttataatgtaattgttacagcacatgcatttgtaataattttctttatagtaatgccaattatgattggtgggtttggaaattgactaatcccattaatgattggagcccccgacatagcattcccacgaatgaataatataagcttttgacttctaccaccatccttcctgctactgcttgcatcctctgcagtagagtcaggtgctggaacaggatgaacagtctacccccctctggctggcaatctagcacatgcaggagcatctgtagatcttactattttctccctacaccttgcaggtatctcttcaattcttggggctattaactttattacaacaattatcaacataaaaccccccgctatctctcaatatcagacccctctatttgtctgagcagtattaattactgctgtcctactcctcctctcacttcctgtccttgctgcgggtattacaatgcttcttacagatcgaaacttaaatactaccttcttcgatccggcaggcggaggagaccccattttatatcagcacttattctgattcttcggccacccggaagtatacattcttatcttaccaggattcggaatgatttcacatatcgtcgcctactactcaggtaaaaaagaacctttcggctacataggaatagtatgggctataatagcaattggcttactaggatttatcgtatgagcccaccatatgttcacagtcggaatggacgtagacacacgtgcctacttcacatctgccactatgattattgcaattcctactggcgtaaaagtcttcagctgactggccacccttcatggaggatctatcaaatgagaaaccccactattatgagccctgggctttattttcttatttacagtaggaggcctaacagggattgttcttgccaattcttctttagacattgttctacacgacacatattacgtagtagcccacttccactatgtcctatctataggagctgtatttgctattgttgccgccttcgtccattgattcccactattctctggctacactctacacagtacctgaacaaaaattcacttcggaattatgtttgttggtgtaaacttaaccttcttcccacaacacttccttggtttagccggaatacctcgacggtactcagactacccagacgcctataccctgtgaaacactatctcttccattggctctctaatctccctagtagctgtaatcatgttcttatttattatctgagaagcattcgccgctaagcgtgaagtaatgtcagttgaactaacagcaactaacgtagaatgactccacggctgccctcccccttatcacacatttgaagaacctgcatacgtccaagtccagctaaatta
So, perhaps someone with more eDNA experience should look into this more closely?
For example, by that metric, the NovaSeq COI alignment would also relatively low (but higher than the MiSeq COI alignment). I am not following the pre-processing steps described (like merging read pairs), but it should also be noted if a large fraction of reads are being filtered (assuming those steps would not also help with a Bowtie2 PE and/or SE-merged alignment).
Charles Warden Replied to Charles Warden
Also, the sequence length distribution varies in the FASTQ files (even if pre-processing is necessary, the raw reads should be uploaded to the SRA, and it looks the reads were at least trimmed before upload).
Greg Singer Replied to Charles Warden
Hi Charles, the reads uploaded to SRA are just post-bcl2fastq (necessary for demultiplexing from other projects in the same run). This removes the adapters and indices from the reads, and (in theory) also removes PhiX since those don't have any index tags. There are two additional steps that should eliminate PhiX from the final results: (1) cutadapt was used to remove primer sequences, which won't be present in the PhiX spike-in, and (2) DADA2 also has a PhiX removal step. These steps were identical for the two platforms, so the final ASV-based comparison should not be influenced by PhiX.
Charles Warden Replied to Greg Singer
Hi Greg,
Thank you very much for your response.
Even though PhiX doesn’t have a barcode, you can definitely see PhiX sequence hopping into samples.
In that Biostars discussion, the plots are for the InterOp alignment percentages. However, I also checked within the de-multiplexed samples for one of those runs: “You can check for PhiX alignments with your samples (such as a Bowtie2 alignment to a RefSeq PhiX sequence). However, that number was noticeably different (lower) than the InterOp PhiX percentage, when I checked the lane with the highest InterOp percentage (1.08-1.47% for sample FASTQ, versus average of 5.10% for InterOp R1 and average of 4.95% for InterOp R2).” [emphasis changed to show that you will get PhiX in the de-multiplexed sequences, even though PhiX doesn’t have a barcode]
For HiSeq2500, using dual-barcode libraries reduced the presence of PhiX within the de-multiplexed samples, but other types of index hopping still occur. I thought NovaSeq libraries tended to be dual-barcode, but there would still somehow clearly be extra PhiX sequence in those samples. Plus, even with the HiSeq2500 reads the PhiX counts didn’t go to 0 with the dual-barcode base calling (for single-barcode samples). For example, the PhiX reads could go down from 402,463 PhiX reads to 13 PhiX reads when calling a single-barcode sample like a dual-barcode sample (by using the adapter sequence for the 2nd index, in a run with mixed library types).
Your MiSeq PhiX counts were all 0, but I have included your NovaSeq PhiX counts at the end of this response (ordered by number of PhiX reads). With the exception of ~4 samples, I think the counts for your samples look somewhere between the single-barcode and dual-barcode HiSeq2500 counts (in that run that used a higher PhiX percentage, which I believe should be similar or lower than you used for these samples, if you followed the recommendations cited in your paper).
Maybe we are already on the same page. However, in the SRA, there are also definitely PhiX reads in your NovaSeq samples.
So, I have the following comments and questions:
Response / Comment / Question 1) I am familiar with using cutadapt to trim out adapters/primers, but I apologize that I don’t see how this would resolve the issue that you are describing.
If there is an expected sequence that you trim out (such as a specific primer), are you saying you only use sequences that become shorter after using cutadapt?
If the primers could bluntly ligate to off-target sequence (though whatever mechanism causes normal barcode hopping?), maybe this could still be an issue. However, I think I should first make sure I understand exactly what you are saying, and/or see if you can help point out where I can find the sequence results for a question that I have at the end.
Response / Comment 2) Thank you - I will learn more about the effect of using the R DADA2 package. For example, it may take me a little while to perform some additional analysis, but I will look into i) are the PhiX reads successfully removed in your SRA data and ii) does removing the PhiX reads before running DADA2 affect the diversity of the other sequences?
Question a) There is some place where the filtered/corrected FASTQ files can be downloaded? I am wondering how the same analysis that I performed on the SRA data compares with that set of samples.
Question b) Just out of curiosity, can you please provide the barcode sequences that you used to generate FASTQ files using bcl2fastq? For example, did you use dual-barcode libraries for both MiSeq and NovaSeq? Or, are there any single-barcode samples?
Thank You,
Charles
NovaSeq PhiX Counts:
sampleIDphiX_counts
SRR842387319
SRR842382822
SRR842383253.5
SRR842387156
SRR8423841108.5
SRR8423870110.5
SRR8423842121
SRR8423827122.5
SRR8423833137
SRR8423867140.5
SRR8423843173.5
SRR8423875189.5
SRR8423830205.5
SRR8423840235
SRR8423831253.5
SRR8423866263
SRR8423829279.5
SRR8423874464.5
SRR8423835586
SRR8423834784
SRR8423872872.5
SRR84238374446
SRR84238367978
SRR842381610158
SRR842382611213
SRR842382414255.5
SRR842385515455
SRR842384915704.5
SRR842382316847.5
SRR842385316873
SRR842381821213
SRR842385021648
SRR842381725402
SRR842382527498
SRR842385129627.5
SRR842382130006
SRR842388533201.5
SRR842384636790.5
SRR842388238403
SRR842388341478
SRR842384857138.5
SRR842385458636.5
SRR842388464737
SRR842385273828
SRR8423820106146
SRR8423847335271
SRR8423819524132
SRR84238221547526
Read counts are sometimes fractions because they are calculated using samtools idxstats, summarized using the following R code:
#alignmentFolder="MiSeq_PhiX_Alignment"
#summaryFile="PhiX_counts_MiSeq_PhiX_Alignment.txt"
alignmentFolder="NovaSeq_PhiX_Alignment"
summaryFile="PhiX_counts_NovaSeq_PhiX_Alignment.txt"
sample.folders = list.dirs(alignmentFolder, full.names = FALSE)
sample.folders = sample.folders[sample.folders!=""]
sampleID = sample.folders
input.files = paste(alignmentFolder, sample.folders,"idxstats.txt", sep="/")
phiX_counts = c()
phiX_fraction = c()
phiX_percent = c()
total_reads = c()
for (i in 1:length(sampleID)){
print(sampleID[i])
input.table = read.table(input.files[i],head=F,sep="\t")
aligned.PE.counts = input.table$V3[1]
unaligned.PE.counts = input.table$V4[1]
no_alignments = input.table$V4[input.table$V1 == "*"]
fragment.counts = aligned.PE.counts-unaligned.PE.counts
fragment.counts[fragment.counts < 0]=0
phiX_counts[i]=fragment.counts / 2#correct for PE
}#end for (i in 1:length(sampleID))
output.table = data.frame(sampleID, phiX_counts)
write.table(output.table, summaryFile, quote=F, sep="\t", row.names=F)
Charles Warden Replied to Charles Warden
Hi Again Greg,
I confirmed what you said for Response / Comment 2i), where DADA2 filtered all of the reads that I identified as PhiX (making those filtered counts more like the MiSeq reads).
There are some other things that I think are probably good to look into, but I am glad that DADA2 can do this.
However, I think we agree that PhiX doesn't have a barcode, so this could be considered cross-contamination. If other cross-contamination exists that was harder to identify (but proportional to the PhiX sequence), then there could still be a problem.
This confirmation is without the cutadapt filter, but I realize that could matter for the other sequences.
Also, just for everybody's information, I am including the DADA2 filtered read percentages and code below (for DADA2 - the Bowtie2 alignment for confirmation just has 0 aligned reads for each sample).
Thanks Again!
Sincerely,
Charles
NovaSeq Samples with Decreasing Filtered Percent:
Samplereads.inpercent.filtered
SRR8423822_1.fastq.gz1287589514.2515%
SRR8423819_1.fastq.gz101842457.9138%
SRR8423847_1.fastq.gz140927773.0122%
SRR8423820_1.fastq.gz156464031.5593%
SRR8423852_1.fastq.gz157220690.9085%
SRR8423884_1.fastq.gz182959800.7896%
SRR8423854_1.fastq.gz152045250.7064%
SRR8423846_1.fastq.gz131272310.6967%
SRR8423883_1.fastq.gz173693770.5847%
SRR8423848_1.fastq.gz202541560.5695%
SRR8423821_1.fastq.gz200201970.5011%
SRR8423824_1.fastq.gz120489290.4925%
SRR8423885_1.fastq.gz142204660.4812%
SRR8423882_1.fastq.gz203409440.4756%
SRR8423825_1.fastq.gz149050010.4429%
SRR8423817_1.fastq.gz113568230.4351%
SRR8423818_1.fastq.gz99158940.4152%
SRR8423849_1.fastq.gz107245370.3974%
SRR8423851_1.fastq.gz162553060.3802%
SRR8423823_1.fastq.gz127234720.3798%
SRR8423855_1.fastq.gz142963190.3384%
SRR8423850_1.fastq.gz160103830.3263%
SRR8423853_1.fastq.gz149023110.2943%
SRR8423816_1.fastq.gz135771900.2765%
SRR8423836_1.fastq.gz140479920.1289%
SRR8423826_1.fastq.gz98469490.1260%
SRR8423837_1.fastq.gz200260710.0296%
SRR8423872_1.fastq.gz140887670.0135%
SRR8423840_1.fastq.gz122673020.0102%
SRR8423866_1.fastq.gz134599580.0092%
SRR8423874_1.fastq.gz179037910.0092%
SRR8423843_1.fastq.gz142760390.0091%
SRR8423834_1.fastq.gz177243590.0082%
SRR8423829_1.fastq.gz129781810.0079%
SRR8423835_1.fastq.gz189024410.0077%
SRR8423830_1.fastq.gz137975730.0076%
SRR8423841_1.fastq.gz95418640.0072%
SRR8423827_1.fastq.gz134913430.0060%
SRR8423867_1.fastq.gz141825110.0059%
SRR8423831_1.fastq.gz180143230.0052%
SRR8423842_1.fastq.gz115976760.0052%
SRR8423875_1.fastq.gz222800170.0049%
SRR8423870_1.fastq.gz208683110.0046%
SRR8423832_1.fastq.gz280692230.0043%
SRR8423833_1.fastq.gz260444900.0043%
SRR8423828_1.fastq.gz205192410.0041%
SRR8423873_1.fastq.gz163084570.0040%
SRR8423871_1.fastq.gz171119570.0027%
Code:
#use code example from https://benjjneb.github.io/dada2/
library(dada2)
read.folder = "../Reads"
sequencer="Illumina NovaSeq 6000"
output.folder = "NovaSeq_DADA2_Processed_Reads"
mapping.file = "../Reads/PRJNA513845.txt"
mapping.table = read.table(mapping.file, head=T, sep="\t")
print(dim(mapping.table))
mapping.table = mapping.table[mapping.table$instrument_model == sequencer,]
print(dim(mapping.table))
reads = list.files(read.folder)
fnFs = paste(read.folder,"/",mapping.table$run_accession,"_1.fastq.gz",sep="")
fnRs = paste(read.folder,"/",mapping.table$run_accession,"_2.fastq.gz",sep="")
filtFs = paste(output.folder,"/",mapping.table$run_accession,"_1.fastq.gz",sep="")
filtRs = paste(output.folder,"/",mapping.table$run_accession,"_2.fastq.gz",sep="")
sample.names = mapping.table$run_accession
out = filterAndTrim(fnFs, filtFs, fnRs, filtRs,
rm.phix=TRUE, compress=TRUE)
summary.file = paste(plot_prefix,"_filter_summary.txt",sep="")
write.table(data.frame(Sample=rownames(out),out),
summary.file, quote=F, sep="\t", row.names=F)
######################################
### percent calculation was separate script ###
######################################
input.file = "NovaSeq_filter_summary.txt"
output.file = "NovaSeq_filter_summary-updated.txt"
input.table = read.table(input.file, head=T, sep="\t")
percent.filtered = paste(round(100 * (1 - input.table$reads.out / input.table$reads.in), digits=4),"%", sep="")
output.table = data.frame(input.table, percent.filtered)
write.table(output.table, output.file, quote=F, sep="\t", row.names=F)
Charles Warden Replied to Charles Warden
Thank you again for your responses!
These are some examples where I found your responses to be nicely helpful:
a) In the methods, it says “Unique dual Nextera indexes were used to mitigate index misassignment (IDT; 8-bp index codes)”. I think I have only seen dual-barcode NovaSeq samples, so I think this makes sense. To me, that seems a bit odd because the number of PhiX reads sharply dropped when single-barcode samples were called as dual-barcode samples on the HiSeq2500 referenced in the Biostars discussion (because the overall run had a mix of library types). However, even if something was done differently for base calling of the NovaSeq samples, I agree that this usually should help.
So, I apologize for not looking back into the paper earlier to see this.
b) I don’t think I completely understand what is going on, but running cutadapt to remove the Nextera transposase adapters was successful in removing a lot of the PhiX reads (although I didn’t test the PhiX outliers).
So, having this discussion has been very helpful in allowing me to understand the project and the methods better, and I think that is a very positive thing for science!
Importantly, I think you may be right to shift the focus from the PhiX reads (at least for variation within NovaSeq samples), even though I think that represents something important in the broader discussion (for the reasons that I previously described).
Anne Chenuil
Can you answer me to this point ?
Which platform reflects more accurately the relative abundances of the different sequences in the amplicon ? From your explanations on the flow-cells, if I understood well, the MIseq probably favours abundant sequences with respect to rare ones. (1) Does Novaseq also favour abundant sequences relative to abundant ones (but less than the MISEQ), (2) does it , on the contrary favour rare ones, or (3) is it, a priori, accurate with respect to this ?
Is there a study investigating sequencing platform biases which used MOCK samples with known abundances in amplicons (PCR-MOCK rather than DNA-MOCK) or do you plan to do so in near future ?
Greg Singer Replied to Anne Chenuil
Hi Anne, unfortunately we can't say with certainty because we don't know the actual relative concentrations of eDNA in the samples we started with. In general, other studies have shown only a weak correlation between eDNA concentrations and read abundance, although low-abundance reads probably do reflect low copy number eDNA in the samples.
Charles Warden
Thank you again for your responses.
I have some more observations and questions, which I will try to make clear with 2 separate comments.
In general, I have uploaded code and some content on GitHub:
https://github.com/cwarden45/PRJNA513845-eDNA_reanalysis
And I have also uploaded some processed sequences on Zenodo:
https://zenodo.org/record/4546559
Charles Warden Replied to Charles Warden
Raw Illumina reads from a run should all have the same read length. However, the uploaded reads have variable lengths.
For example, you can see this in the SRA run browser (summarized in this file). In that file, it looks like the FishE samples tend to have more variable lengths in the SRA, but I realize that separate analysis for the different amplicons was provided in the paper.
Is it possible to upload the raw reads in order to test re-processing all of the samples in a given way (even if that may or may not require re-running base calling)? Or, for me, I am interested in checking how many other valid Illumina barcodes can be found when you check the sequence near the adapters in the smaller read fragments (as I did for this unrelated sample).
Charles Warden Replied to Charles Warden
I think showing the variation by sample can be important.
I ran DADA2 with results on the GitHub page, but I ran DADA2 in a different way than described in this paper.
However, you also did some extra analysis with OTUs since you wanted to see if DADA2 could be introducing artifacts. I performed analysis that I think should be comparable to that (which I believe is also mostly comparable to the DADA2 analysis that I performed).
You can see the results for both FLASH and PEAR merged reads here, but I am showing an image for subset of the FLASH analysis (with all reads) below:
https://uploads.disquscdn.c...
Maybe there are a few outliers above the trendline that could affect the reported average for each group, but I think this fits relatively well for both sequencers.
If anything, I think the trend may even be reversed if you require at least 2 reads in at least 1 sample for each unique sequence/OTU:
https://raw.githubuserconte...
It might be a bit much to go over here, but I have extra questions that could be asked towards the bottom of the GitHub page. For example, there are 2 NovaSeq outliers (SRR8423848 and SRR8423854) that have merged read counts within the same range as the MiSeq samples. I also believe that there is some evidence that the MiSeq may be better at capturing some slightly larger amplicons.
In other words, I think the various plots showing the variably per sample may not match one of the conclusions mentioned in the abstract (such as “the NovaSeq can detect more DNA sequence diversity within samples than the MiSeq, even at the exact same sequencing depth”), at least not consistently across all or most of the samples per group.
So, this is my question: If you use the DADA2 and OTU analysis in the paper, what does that look like if you show the results for each sample separately (instead of combined per sequencer) as a function of coverage (perhaps with and without the minimum of 2 reads in at least 1 sample)?