Abstract
Bats are the only mammals capable of powered flight and their body temperature can reach up to 42 °C during flight. Additionally, bats display robust type I IFN interferon (IFN-I) responses and some species constitutively express IFN-α. Reference genes with stable expression under temperature oscillations and IFN-I release are therefore critical for normalization of quantitative reverse-transcription polymerase chain reaction (qRT-PCR) data in bats. The expression stability of reference genes in Rousettus aegyptiacus remains elusive, although this species is frequently used in the infection research. We selected ACTB, EEF1A1, GAPDH and PGK1 as candidate reference genes and evaluated their expression stability in various tissues and cells from this model bat species upon IFN-I treatment at 35 °C, 37 °C and 40 °C by qRT-PCR. We employed two statistical algorithms, BestKeeper and NormFinder, and found that EEF1A1 exhibited the highest expression stability under all tested conditions. ACTB and GAPDH displayed unstable expression upon temperature change and IFN-I treatment, respectively. By normalizing to EEF1A1, we uncovered that GAPDH expression was significantly induced by IFN-I in R. aegyptiacus. Our study identifies EEF1A1 as the most suitable reference gene for qRT-PCR studies upon temperature changes and IFN-I treatment and unveils the induction of GAPDH expression by IFN-I in R. aegyptiacus. These findings are pertinent to other bat species and may be relevant for non-volant mammals that show physiological fluctuations of core body temperature.
Similar content being viewed by others
Introduction
Bats are increasingly recognized as reservoir hosts of highly-virulent pathogens, such as Filoviruses, Lyssaviruses, Paramyxoviruses and Coronaviruses, including severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), which causes the current global pandemic1. Egyptian fruit bats, Rousettus aegyptiacus, have been identified as putative reservoir hosts of Marburg virus2, Kasokero virus3 and Sosuga virus4, and were shown to be susceptible to experimental challenge with SARS-CoV-25 and Rift valley fever phlebovirus6. These viruses may cause severe diseases with high mortality rates in humans. However, bats show minimal and often even no clinical manifestation upon natural infection. Under experimental conditions, Rousettus bats infected with high doses of Ebola7, SARS-CoV8 or SARS‐CoV‐25 only support transient viral replication and display limited pathology. Several hypotheses could explain the reservoir potential of bats. As the only mammals capable of powered flight, their body temperature can reach 42 °C during flight9. Hence, ‘flight-as-fever’ has been postulated as a unique mechanism conferring effective immune defence6. Further, the black flying fox, Pteropus alecto, constitutively expresses IFN-α10, and gene loci of type I interferons (IFN-I) in R. aegyptiacus, are markedly expanded11, indicating a potential contribution of boosted IFN-I signalling to antiviral immunity in bats. In addition to the enhanced antiviral immunity, bats can suppress excessive inflammation which may explain their lack of symptoms during viral infection1,12. For example, TNFα expression in Eptesicus fuscus cells is abolished upon poly(I:C) stimulation13, NLRP3-mediated inflammasome activation is impaired in P. alecto14 and Myotis davidii12, and R. aegyptiacus does not upregulate pro-inflammatory genes (CCL8, FAS and IL6) upon Marburg virus infection15. Enhanced antiviral immunity along with reduced inflammation likely explains the ability of bats to harbour high-impact pathogens in absence of clinical disease. Confirmation of these findings in distinct bat species as well as elucidation of novel immune mechanisms contributing to the reservoir potential of bats require accurate monitoring of their immune responses.
The knowledge about the exceptional immune system of bats has significantly advanced during the past decade16. However, experimental tools to systematically investigate bat immune responses, such as species-specific or cross-reactive antibodies, are largely missing17,18,19. Accordingly, investigations on host immunity heavily rely on gene transcription profiling by qRT-PCR. This method is sensitive, specific, highly reproducible and accurate20,21. A critical step in qRT-PCR setup is selection of several stable reference genes. The inclusion of such reference genes is crucial for gene expression normalization and subsequent data interpretation. The ideal reference genes should maintain stable expression levels across diverse tissues and cell types as well as under different experimental conditions22. Considering that bats display unique physiological features, notably oscillating metabolic rates and core body temperature depending on flying and roosting phases23,24,25,26, the expression stability of reference genes must be evaluated in context of these physiologically relevant conditions. Multiple reference genes, including glyceraldehyde-3-phosphate dehydrogenase (GAPDH), actin-beta (ACTB), small nuclear ribonucleoprotein Sm D3 (SNRPD3) and 18S ribosomal RNA (18S rRNA) have been employed in gene expression studies of P. alecto, E. fuscus, M. davidii and other bats13,14,27,28,29. However, a comprehensive analysis of reference genes, particularly their expression stability under oscillating temperatures, has not been performed in bats, including the model bat R. aegyptiacus.
Here, we provide a first in-depth validation of four reference gene candidates, including ACTB, GAPDH, eukaryotic translation elongation factor 1 alpha 1 (EEF1A1) and phosphoglycerate kinase 1 (PGK1) for R. aegyptiacus. Our findings support EEF1A1 as an appropriate reference gene for normalization of qRT-PCR data in the Egyptian fruit bat and call for caution when using other candidate genes, i.e. GAPDH and ACTB, due to their instability under specific conditions.
Results
Performance of PCR primers targeting reference gene candidates
To evaluate the performance of the qRT-PCR assay, we first examined the specificity of the primer pairs with melting curve analysis, agarose gel electrophoresis and sequencing. Melting curve analysis revealed single peaks for all primer pairs (Fig. S1). Agarose gel electrophoresis further demonstrated single bands for all the PCR products with the predicted sizes, indicating high specificity of all primer pairs (Fig. S1). PCR products were sequenced and the specificity of the primers was confirmed (Fig. S2). To evaluate the amplification efficiency, standard curves with ten-fold dilution steps were generated (Fig. S3) and subsequently the linear dynamic range (LDR) and precision of each primer pair were assessed following the MIQE guideline30. Amplification efficiencies of all tested primers met the validation criteria, notably 100.51% for ACTB, 99.53% for EEF1A1, 106.52% for GAPDH and 106.44% for PGK1 (Table 1). The correlation coefficient (R2) of all candidates was above 0.99, suggesting excellent linearity of the standard curves. The LDR values of all primers were in the range of 3 to 300,000 copies and precision values varied from 0.31 (EEF1A1) to 1.27 (ACTB) (Table 1). Thus, all the primers for reference gene candidates demonstrated satisfactory specificity and efficiency in qRT-PCR.
Expression profile of reference gene candidates in various tissues from R. aegyptiacus
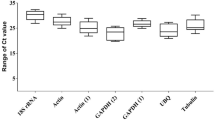
To investigate the expression of reference gene candidates in tissues from R. aegyptiacus, qRT-PCR was performed with pooled cDNA from nose (nasal epithelium), trachea, lung, blood, spleen and duodenum. Threshold cycle (Ct) values were employed to determine the expression levels of the candidate reference genes. Their expression levels varied and EEF1A1 displayed the highest expression levels across different tissues as indicated by the lowest Ct values. The overall Ct values of EEF1A1, ACTB, GAPDH and PGK1 were 23 ± 1.5, 33 ± 3, 27 ± 2 and 27 ± 1.5, respectively (Fig. 1). EEF1A1 and PGK1 showed the lowest variability in Ct values, suggesting that these two genes display the most stable expression across diverse tissues from R. aegyptiacus.
Expression of candidate reference genes in primary bat fibroblasts upon IFN-I stimulation and incubation at various temperatures
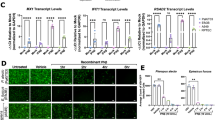
To investigate the expression stability of candidate reference genes, bat primary fibroblasts were incubated at 35 °C, 37 °C or 40 °C in the presence or absence of universal type I interferon (uIFN) for 4 h. We employed 35 °C, 37 °C and 40 °C to mimic the physiological daily oscillation of body temperature in R. aegyptiacus 9. The expression levels of all candidate reference genes under these conditions were assessed by qRT-PCR. ACTB showed a broad variation in Ct values, ranging from 29.7 to 39.01 at 40 °C. The Ct values of ACTB were lower at 35 °C, 31.5 ± 0.7 compared to 33.5 ± 1 at 37 °C, suggesting an unstable expression of ACTB upon temperature changes (Fig. 2). The expression levels of EEF1A1, GAPDH or PGK1 were comparable at 35 °C, 37 °C and 40 °C, demonstrating stable expression of these potential reference genes under temperature oscillations. Since uIFN activates IFN-I pathway in bats31, we employed this cytokine to further investigate the expression of the candidate reference genes upon IFN-I stimulation. The mean Ct values of GAPDH decreased upon IFN-I stimulation at 35 °C, 37 °C and 40 °C, indicating increased expression of GAPDH by IFN-I. The Ct values of EEF1A1 and PGK1 remained unchanged following stimulation with IFN-I at 35 °C, 37 °C or 40 °C, suggesting their stable expression under these experimental conditions (Fig. 2). These findings indicate that expression stabilities of ACTB and GAPDH are impaired by temperature oscillations and IFN-I, respectively.
Expression stability of candidate reference genes under different conditions. Bat primary fibroblasts were incubated at 35 °C, 37 °C or 40 °C in the presence or absence of 1,000 U/ml universal type I interferon (uIFN). Gene expression levels of ACTB, EEF1A1, GAPDH and PGK1 were determined by qRT-PCR. Data show mean ± SD from 3 independent experiments.
Expression stability analysis of reference gene candidates under different conditions
To quantitatively evaluate the expression stability of candidate reference genes, qPCR results were analysed with the statistical algorithms BestKeeper and NormFinder. BestKeeper enables pairwise correlation, regression analysis32,33 and calculations of standard deviation (SD) of all Ct values [SD (± Ct)] as well as of the standard deviation of absolute regulation coefficients [SD (± x-fold)]. Both parameters are indicators of expression variability. A suitable reference gene should display values of < 1 for SD (± Ct), < 2 for SD (± x-fold), and have a coefficient of correlation (r) close to 1 34. According to this algorithm, the [SD (± Ct)] values for ACTB, EEF1A1, GAPDH and PGK1 were 1.40, 0.34, 0.76 and 0.36, along with their corresponding r-values of 0.52, 0.96, 0.78 and 0.88. Thus, expression stability for candidate reference genes ranks as following: EEF1A1, PGK1, GAPDH, and ACTB (Table 2).
NormFinder was employed as the second algorithm to determine gene stability, since it allows evaluation of the overall stability as well as the individual stability for each condition35. The most stable reference genes display stability values close to 0 according to this algorithm. Based on this method, we calculated the overall and individual stability values. The total stability values for ACTB, EEF1A1, GAPDH and PGK1 were 0.097, 0.013, 0.05 and 0.015, respectively, suggesting EEF1A1 as the most stable reference gene under all tested conditions. EEF1A1 also displayed the highest expression stability under high temperature or upon IFN-I treatment (Table 3). Altogether, both algorithms suggest that EEF1A1 is the reference gene with the highest expression stability under the selected conditions, followed by PGK1, GAPDH, whereas ACTB has the lowest stability.
GAPDH expression is induced by IFN-I in R. aegyptiacus cells
To evaluate the relative expression levels of ACTB, GAPDH and PGK1 under different conditions, we normalized each candidate to the most stable reference gene, EEF1A1 (Fig. 3A–C). The relative expression levels of PGK1 remained stable at all temperatures evaluated, irrespective of the IFN-I treatment. ACTB expression was also stable at 35 °C and 37 °C in the presence of IFN-I. However, the expression level was significantly reduced following a 2 h IFN-I stimulation at 40 °C (Fig. 3C). Intriguingly, the expression of GAPDH was significantly increased in cells treated with IFN-I at all selected temperatures, suggesting that the induction of GAPDH by IFN-I is temperature independent in R. aegyptiacus (Fig. 3A–C). To investigate whether GAPDH induction by IFN-I is specific to R. aegyptiacus, we stimulated human fibroblasts with IFN-I at 37 °C and 40 °C (Fig. 3D, E). Both human GAPDH and ACTB have been previously used as reference genes in several qRT-PCR studies36,37. Indeed, both genes displayed good expression stability at 37 °C, yet the variation of Ct values at 40 °C argued against stability of human GAPDH and ACTB at 40 °C. The relative expression of human GAPDH normalized against ACTB remained unchanged upon IFN-I treatment either at 37 °C or 40 °C, revealing that human GAPDH expression in fibroblasts is not modulated by IFN-I (Fig. 3D, E). Overall, we conclude that IFN-I triggers GAPDH expression specifically in R. aegyptiacus.
GAPDH expression is induced by IFN-I in R. aegyptiacus. Relative gene expression of ACTB, GAPDH and PGK1 normalized to EEF1A1 upon uIFN treatment at 35 °C (A), 37 °C (B) or 40 °C (C). (D, E) Ct values of human ACTB and GAPDH (D) and relative expression of human GAPDH normalized to ACTB (E) in human fibroblasts upon uIFN treatment and incubation at 37 °C or 40 °C. Bat (A, B, C) or human (D, E) primary fibroblasts were stimulated with PBS (ctrl) or 1000U/ml universal type I interferon (uIFN) for indicated time at 35 °C, 37 °C or 40 °C. Gene expression levels of ACTB, GAPDH and PGK1 were determined with qRT-PCR. Data show mean ± SD from 3 independent experiments. The statistical significance was calculated using one-way ANOVA with Holm-Šidák’s post-hoc test. (*) P ≤ 0.05, (**) P ≤ 0.01, (***) P ≤ 0.001, (****) P ≤ 0.0001.
Discussion
Accurate gene transcription measurements require selection of reference genes that maintain high stability under various experimental conditions. In this study, we selected ACTB, GAPDH, EEF1A1 and PGK1 as reference gene candidates due to their wide applications in other species14,32,36,39,42,43,44,45,46,47,48 and evaluated the suitability of these candidates in R. aegyptiacus under the physiological relevant conditions, notably temperature oscillation and IFN-I stimulation. By employing BestKeeper and NormFinder, we calculated the stability and validated EEF1A1 as the most stable reference gene in R. aegyptiacus under conditions relevant for the biology of this species36,38,39,40,41. By normalizing the qRT-PCR data to EEF1A1 we observed that expression of GAPDH was significantly induced by IFN-I, suggesting unsuitableness of using this commonly used reference gene in R. aegyptiacus. Consistent with our results, many reports have demonstrated that the expression of GAPDH and ACTB is unstable in various tissues or cells from mice and humans and upon certain stimulations, such as IL-2 and hypoxia 37,49,50,51,52. Whether GAPDH expression is induced by IFN-I, and whether GAPDH and ACTB are suitable reference genes in other bat species, require further investigations.
Our finding that the expression of GAPDH, one of the key enzymes in glycolysis, is induced by IFN-I in R. aegyptiacus may have implications for the immunometabolism of bats. It is well established that metabolic reprogramming in cells controls their immune responses53,54. Glycolysis is commonly utilized by various immune cells to enable prompt responses to infections. In bone marrow derived macrophages, aerobic glycolysis promotes IL-1β production upon LPS stimulation or Bordetella pertussis infection 55. In CD4+ and CD8+ T-cells56,57,58,59,60, as well as in NK cells61,62, glycolysis is required for their effector functions, such as IFN-γ production and cytotoxicity. In human plasmacytoid dendritic cells (pDCs) and monocyte-derived DCs (moDC), glycolysis promotes IFN-I production upon TLR9 or RIG-I activation, respectively63. Further, IFN-I induces a metabolic shift towards glycolysis, contributing to the antiviral activity in fibroblasts and antigen presentation in DCs64,65. On the other hand, lactate, the end metabolite of glycolysis, directly binds to mitochondrial antiviral-signaling protein (MAVS), and consequently inhibits its activation and IFN-I production66. Thus, outcomes of such metabolic shift vary in diverse cells or under different stimulations. The nectarivore bat Glossophaga soricina employs high rates of glycolysis to generate ATP during flight67. As a fruit bat, R. aegyptiacus could also directly utilize dietary sugars to fuel both roosting and flight metabolism68. Whether induction of GAPDH expression in this species impacts on metabolic reprogramming towards glycolysis, needs to be clarified. Moreover, whether and how such metabolic shifts affect IFN-I signalling or other immune pathways in R. aegyptiacus remains to be uncovered.
In addition to its roles in glycolysis, GAPDH also modulates cell death, RNA export and cytoskeleton dynamics69. Upon serum deprivation and DNA damage, GAPDH translocates to the mitochondria and interacts with voltage-dependent anion channel (VDAC), leading to apoptosis70. It can also bind to the 3' untranslated region of TNFα mRNA and represses TNFα expression in human monocytes and macrophages71. Hence, GAPDH upregulation by IFN-I may contribute to apoptosis induction and TNFα repression in R. aegyptiacus, which could represent novel mechanisms for the prevention of excessive inflammation during viral infections.
Overall, our study provides an extensive analysis of reference genes and identifies EEF1A1 as the most stable reference gene in R. aegyptiacus under temperature changes and IFN-I stimulation, which allows us to perform accurate gene transcription studies in this species. Our findings also open new investigation avenues by showing that GAPDH is regulated by IFN-I which has a broad relevance in context of immunometabolism.
Material and methods
Selection of reference gene candidates and design of primer pairs
Reference gene candidates (ACTB, EEF1A1, GAPDH and PGK1) for R. aegyptiacus gene expression studies were selected based on their utilization in other bat species13,14,17. Primers against these reference genes were designed using the PrimerQuest tool (Integrated DNA Technologies, Inc.). The criteria for primer design were as follows: primer lengths around 17–30 bp, GC content of 40–55%, optimal melting temperature at 62 °C, and amplicon lengths within a range of 100–250 bp. Derived primer pairs were evaluated using the OligoAnalyzer tool to exclude primers with hairpin structures and homo- and/or heterodimer formation (Integrated DNA Technologies, Inc.). Primer sequences were also blasted using the NCBI BLAST tool to ensure their specificity for R. aegyptiacus. The primer pairs meeting all criteria were selected for further experiments. The characteristics of these primers are shown in Table 4.
Cells, tissues and stimulation experiments
Bat fibroblasts were derived from the lung of a female R. aegyptiacus bat from the R. aegyptiacus breeding colony at the Friedrich-Loeffler-Institut. Sampling was performed in accordance with current European and National Animal Welfare regulations, after ethical review and approval by the authority of the Federal State of Mecklenburg-Western Pomerania, Germany (file number 7221.3–2-042/17) and the experiments were carried out according to ARRIVE guidelines (https://arriveguidelines.org). The lung tissue was dissected into small pieces and digested with trypsin overnight at 4 °C. Dissociated cells were seeded in cell culture dishes in DMEM medium (DMEM high glucose medium, 10% fetal bovine serum (FBS), 2 mM glutamine and 100U/mL of Penicillin–Streptomycin) for 2 h, and only adherent cells were propagated. Confirmation of bat fibroblast identity was carried out by examining fibroblast activation protein (FAP) expression via PCR (data not shown).
Tissue samples from 12 individual bats were obtained from an animal experiment published before76. For IFN-I stimulation, bat fibroblasts and human dermal fibroblasts (#C0045C, Thermo Fisher Scientific) were incubated with 1000U/ml universal IFN-I (uIFN) (#11200-1, PBL Assay Science) for 1 h, 2 h and 4 h at either 35 °C, 37 °C or 40 °C, respectively.
RNA extraction and cDNA synthesis
Cells were lysed in homemade Trizol solution and RNA was extracted as published before77. Purified RNA was quantified using NanoDrop 2000c spectrophotometer (#ND-2000c, Thermo Fisher Scientific) and 800 ng RNA were subsequently utilized for cDNA synthesis with the LunaScript RT SuperMix Kit (#E3010L, New England BioLabs).
qRT-PCR
qRT-PCR reactions were carried out with EvaGreen Fluorescent DNA stain (# PCR-379, Jena Bioscience), ROX as an internal reference dye (#PCR-351, Jena Bioscience) and GoTaq Polymerase (#M3001, Promega) according to manufacturers’ instructions. The reaction setup was as follows: 95 °C for 2 min; (95 °C for 30 s, 62 °C for 30 s, 72 °C for 1 min) for 40 cycles; 72 °C for 10 min and infinite hold at 4 °C. Unless stated otherwise, each qPCR reaction was performed with 100 ng cDNA. To minimize pipetting errors, the template was diluted and 5 µl were used for each qRT-PCR reaction. Each primer pair was added separately into different wells. Measurements were performed with the QuantStudio 6 Flex real-time PCR system (#4485691, Applied Biosystems). Melting curves were performed within the temperature range from 60.16 °C to 94.885 °C in steps of 0.193 °C, respectively.
Establishment of standard curves and examination of amplification efficiency via qRT-PCR
Standard curves of all reference genes in the qRT-PCR reaction were generated with copy numbers from 300,000 to 3 copies in ten-fold dilution steps. To achieve accurate copy numbers, amplicon sizes of each reference gene were used to calculate the specific weight of each amplicon78, resulting in 2.12 × 10–19 g for ACTB, 1.78 × 10–19 g for EEF1A1, 1.32 × 10–19 g for GAPDH and 1.62 × 10–19 g for PGK1. The amplification efficiency of all primer pairs was subsequently determined with the slope of the standard curve according to the equation 10–1/slope-1. The amplification efficiency of favourable primers ranges between 90–110%. Linear dynamic range (LDR) is described as the highest to the lowest quantifiable copy numbers from standard curves. LDR should cover at least 3 orders of magnitude, ideally 5–6 orders. Precision refers to intra-assay variation and is defined as standard deviation (SD) of technical replicates30.
Amplicon purification and sequencing
Amplicons of all candidate reference genes were visualized in 1.5% agarose gels and bands were cut and purified using the QIAquick gel extraction kit (#28506, Qiagen). Purified amplicons were subsequently sequenced using the Eurofins tubeseq platform.
Stability analyses of reference gene candidates
To investigate the expression stability of the four reference gene candidates, two statistical algorithms were used: BestKeeper33 and NormFinder35. The highest stability was defined as the lowest variation of expression levels under all the selected experimental conditions33,35. In brief, for BestKeeper, raw Ct values without any normalization are subjected to the calculation and the parameters of interest are SD (std dev [± CP]) and the Pearson coefficient of correlation (r). NormFinder utilizes normalized Ct values (2−ΔCt) to provide a direct readout for the estimated expression variation and can separately calculate the stability under each condition (IFN-I treatment, temperature) or total stability.
Statistical analysis
Statistical analysis was performed with GraphPad Prism 8 (GraphPad Software Inc., USA). To determine statistical significance among investigated groups, one-way analysis of variance (ANOVA) with Holm-Šidák’s post-hoc test was performed. A P value of < 0.05 was considered to be significant.
References
Irving, A. T., Ahn, M., Goh, G., Anderson, D. E. & Wang, L.-F. Lessons from the host defences of bats, a unique viral reservoir. Nature 589, 363–370. https://doi.org/10.1038/s41586-020-03128-0 (2021).
Amman, B. R. et al. Oral shedding of Marburg virus in experimentally infected Egyptian fruit bats (Rousettus aegyptiacus). J. Wildl. Dis. 51, 113–124. https://doi.org/10.7589/2014-08-198 (2015).
Kalunda, M. et al. Kasokero virus: A new human pathogen from bats (Rousettus aegyptiacus) in Uganda. Am. J. Trop. Med. Hyg. 35, 387–392. https://doi.org/10.4269/ajtmh.1986.35.387 (1986).
Amman, B. R. et al. A recently discovered pathogenic paramyxovirus, Sosuga virus, is present in Rousettus aegyptiacus fruit bats at multiple locations in Uganda. J. Wildl. Dis. 51, 774–779. https://doi.org/10.7589/2015-02-044 (2015).
Schlottau, K. et al. SARS-CoV-2 in fruit bats, ferrets, pigs, and chickens: an experimental transmission study. Lancet. Microbe 1, e218–e225. https://doi.org/10.1016/S2666-5247(20)30089-6 (2020).
Balkema-Buschmann, A. et al. Productive propagation of rift valley fever phlebovirus vaccine strain MP-12 in Rousettus aegyptiacus fruit bats. Viruses https://doi.org/10.3390/v10120681 (2018).
Paweska, J. T. et al. Experimental Inoculation of Egyptian Fruit Bats (Rousettus aegyptiacus) with Ebola Virus. Viruses https://doi.org/10.3390/v8020029 (2016).
Seifert, S. N. et al. Rousettus aegyptiacus bats do not support productive Nipah virus replication. J. Infect. Dis. 221, S407–S413. https://doi.org/10.1093/infdis/jiz429 (2020).
Kulzer, E. Temperaturregulation bei Flughunden der Gattung Rousettus Gray. Z. Vergl. Physiol. 46, 595–618. https://doi.org/10.1007/BF00298161 (1963).
Zhou, P. et al. Contraction of the type I IFN locus and unusual constitutive expression of IFN-α in bats. Proc. Natl. Acad. Sci. U.S.A. 113, 2696–2701. https://doi.org/10.1073/pnas.1518240113 (2016).
Pavlovich, S. S. et al. The Egyptian Rousette genome reveals unexpected features of bat antiviral immunity. Cell 173, 1098-1110.e18. https://doi.org/10.1016/j.cell.2018.03.070 (2018).
Subudhi, S., Rapin, N. & Misra, V. Immune system modulation and viral persistence in bats: Understanding viral spillover. Viruses https://doi.org/10.3390/v11020192 (2019).
Banerjee, A., Rapin, N., Bollinger, T. & Misra, V. Lack of inflammatory gene expression in bats: a unique role for a transcription repressor. Sci. Rep. 7, 2232. https://doi.org/10.1038/s41598-017-01513-w (2017).
Ahn, M. et al. Dampened NLRP3-mediated inflammation in bats and implications for a special viral reservoir host. Nat. Microbiol. 4, 789–799. https://doi.org/10.1038/s41564-019-0371-3 (2019).
Guito, J. C. et al. Asymptomatic infection of Marburg virus reservoir bats is explained by a strategy of immunoprotective disease tolerance. Curr. Biol. 31, 257-270.e5. https://doi.org/10.1016/j.cub.2020.10.015 (2021).
Wang, L.-F., Gamage, A. M., Chan, W. O. Y., Hiller, M. & Teeling, E. C. Decoding bat immunity: the need for a coordinated research approach. Nat. Rev. Immunol. https://doi.org/10.1038/s41577-021-00523-0 (2021).
Gamage, A. M. et al. Immunophenotyping monocytes, macrophages and granulocytes in the Pteropodid bat Eonycteris spelaea. Sci. Rep. 10, 309. https://doi.org/10.1038/s41598-019-57212-1 (2020).
Martínez Gómez, J. M. et al. Phenotypic and functional characterization of the major lymphocyte populations in the fruit-eating bat Pteropus alecto. Sci. Rep. 6, 37796. https://doi.org/10.1038/srep37796 (2016).
Periasamy, P. et al. Studies on B cells in the fruit-eating black flying fox (Pteropus alecto). Front. Immunol. 10, 489. https://doi.org/10.3389/fimmu.2019.00489 (2019).
Petriccione, M., Mastrobuoni, F., Zampella, L. & Scortichini, M. Reference gene selection for normalization of RT-qPCR gene expression data from Actinidia deliciosa leaves infected with Pseudomonas syringae pv. actinidiae. Sci. Rep. 5, 16961. https://doi.org/10.1038/srep16961 (2015).
McMillan, M. & Pereg, L. Evaluation of reference genes for gene expression analysis using quantitative RT-PCR in Azospirillum brasilense. PLoS ONE 9, e98162. https://doi.org/10.1371/journal.pone.0098162 (2014).
Radonić, A. et al. Guideline to reference gene selection for quantitative real-time PCR. Biochem. Biophys. Res. Commun. 313, 856–862. https://doi.org/10.1016/j.bbrc.2003.11.177 (2004).
Cruz-Neto, A. P., Garland, T. & Abe, A. S. Diet, phylogeny, and basal metabolic rate in phyllostomid bats. Zoology (Jena) 104, 49–58. https://doi.org/10.1078/0944-2006-00006 (2001).
Hock, R. J. The metabolic rates and body temperatures of bats. Biol. Bull. 101, 289–299. https://doi.org/10.2307/1538547 (1951).
O’Mara, M. T. et al. Cyclic bouts of extreme bradycardia counteract the high metabolism of frugivorous bats. Elife https://doi.org/10.7554/eLife.26686 (2017).
Noll, U. G. Body temperature, oxygen consumption, noradrenaline response and cardiovascular adaptations in the flying fox, Rousettus aegyptiacus. Comp. Biochem. Physiol. A Physiol. 63, 79–88. https://doi.org/10.1016/0300-9629(79)90631-5 (1979).
Brook, C. E. et al. Accelerated viral dynamics in bat cell lines, with implications for zoonotic emergence. Elife https://doi.org/10.7554/eLife.48401 (2020).
Koh, J. et al. ABCB1 protects bat cells from DNA damage induced by genotoxic compounds. Nat. Commun. 10, 2820. https://doi.org/10.1038/s41467-019-10495-4 (2019).
Xie, J. et al. Dampened STING-dependent interferon activation in bats. Cell Host Microbe 23, 297-301.e4. https://doi.org/10.1016/j.chom.2018.01.006 (2018).
Bustin, S. A. et al. The MIQE guidelines: Minimum information for publication of quantitative real-time PCR experiments. Clin. Chem. 55, 611–622. https://doi.org/10.1373/clinchem.2008.112797 (2009).
La Cruz-Rivera, P. C. et al. The IFN response in bats displays distinctive IFN-stimulated gene expression kinetics with atypical RNASEL induction. J. Invest. Med. 200, 209–217. https://doi.org/10.4049/jimmunol.1701214 (2018).
Klie, M. & Debener, T. Identification of superior reference genes for data normalisation of expression studies via quantitative PCR in hybrid roses (Rosa hybrida). BMC. Res. Notes 4, 518. https://doi.org/10.1186/1756-0500-4-518 (2011).
Pfaffl, M. W., Tichopad, A., Prgomet, C. & Neuvians, T. P. Determination of stable housekeeping genes, differentially regulated target genes and sample integrity: BestKeeper–Excel-based tool using pair-wise correlations. Biotechnol. Lett. 26, 509–515. https://doi.org/10.1023/b:bile.0000019559.84305.47 (2004).
Pombo, M. A., Zheng, Y., Fei, Z., Martin, G. B. & Rosli, H. G. Use of RNA-seq data to identify and validate RT-qPCR reference genes for studying the tomato-Pseudomonas pathosystem. Sci. Rep. 7, 44905. https://doi.org/10.1038/srep44905 (2017).
Andersen, C. L., Jensen, J. L. & Ørntoft, T. F. Normalization of real-time quantitative reverse transcription-PCR data: a model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Can. Res. 64, 5245–5250. https://doi.org/10.1158/0008-5472.CAN-04-0496 (2004).
Dheda, K. et al. Validation of housekeeping genes for normalizing RNA expression in real-time PCR. BioTechniques 37, 112–4, 116, 118–9. https://doi.org/10.2144/04371RR03 (2004).
Glare, E. M., Divjak, M., Bailey, M. J. & Walters, E. H. beta-Actin and GAPDH housekeeping gene expression in asthmatic airways is variable and not suitable for normalising mRNA levels. Thorax 57, 765–770. https://doi.org/10.1136/thorax.57.9.765 (2002).
Suzuki, T., Higgins, P. J. & Crawford, D. R. Control selection for RNA quantitation. Biotechniques 29, 332–337. https://doi.org/10.2144/00292rv02 (2000).
Falkenberg, V. R., Whistler, T., Murray, J. R., Unger, E. R. & Rajeevan, M. S. Identification of Phosphoglycerate Kinase 1 (PGK1) as a reference gene for quantitative gene expression measurements in human blood RNA. BMC. Res. Notes 4, 324. https://doi.org/10.1186/1756-0500-4-324 (2011).
Molina, C. E. et al. Identification of optimal reference genes for transcriptomic analyses in normal and diseased human heart. Cardiovasc. Res. 114, 247–258. https://doi.org/10.1093/cvr/cvx182 (2018).
Sarwar, M. B. et al. Identification and validation of superior housekeeping gene(s) for qRT-PCR data normalization in Agave sisalana (a CAM-plant) under abiotic stresses. Physiol. Mol. Biol. Plants 26, 567–584. https://doi.org/10.1007/s12298-020-00760-y (2020).
Aminfar, Z., Rabiei, B., Tohidfar, M. & Mirjalili, M. H. Selection and validation of reference genes for quantitative real-time PCR in Rosmarinus officinalis L. in various tissues and under elicitation. Biocatal. Agric. Biotechnol. 20, 101246. https://doi.org/10.1016/j.bcab.2019.101246 (2019).
Bai, B., Ren, J., Bai, F. & Hao, L. Selection and validation of reference genes for gene expression studies in Pseudomonas brassicacearum GS20 using real-time quantitative reverse transcription PCR. PLoS ONE 15, e0227927. https://doi.org/10.1371/journal.pone.0227927 (2020).
Ham, S., Harrison, C., Southwick, G. & Temple-Smith, P. Selection of internal control genes for analysis of gene expression in normal and diseased human dermal fibroblasts using quantitative real-time PCR. Exp. Dermatol. 25, 911–914. https://doi.org/10.1111/exd.13091 (2016).
Huggett, J., Dheda, K., Bustin, S. & Zumla, A. Real-time RT-PCR normalization. Strategies and considerations. Genes Immun. 6, 279–284. https://doi.org/10.1038/sj.gene.6364190 (2005).
Panina, Y., Germond, A., Masui, S. & Watanabe, T. M. Validation of common housekeeping genes as reference for qPCR gene expression analysis during iPS reprogramming process. Sci. Rep. 8, 8716. https://doi.org/10.1038/s41598-018-26707-8 (2018).
Sullivan-Gunn, M., Hinch, E., Vaughan, V. & Lewandowski, P. Choosing a stable housekeeping gene and protein is essential in generating valid gene and protein expression results. Br. J. Cancer 104, 1055; author reply 1056. https://doi.org/10.1038/bjc.2011.35 (2011).
Fujii, H. et al. Functional analysis of Rousettus aegyptiacus “signal transducer and activator of transcription 1” (STAT1). Dev. Comp. Immunol. 34, 598–602. https://doi.org/10.1016/j.dci.2010.01.004 (2010).
Sabath, D. E., Broome, H. & Prystowsky, M. B. Glyceraldehyde-3-phosphate dehydrogenase mRNA is a major interleukin 2-induced transcript in a cloned T-helper lymphocyte. Gene 91, 185–191. https://doi.org/10.1016/0378-1119(90)90087-8 (1990).
Graven, K. K., McDonald, R. J. & Farber, H. W. Hypoxic regulation of endothelial glyceraldehyde-3-phosphate dehydrogenase. Am. J. Physiol. 274, C347–C355. https://doi.org/10.1152/ajpcell.1998.274.2.C347 (1998).
Hazell, A. S., Desjardins, P. & Butterworth, R. F. Increased expression of glyceraldehyde-3-phosphate dehydrogenase in cultured astrocytes following exposure to manganese. Neurochem. Int. 35, 11–17. https://doi.org/10.1016/s0197-0186(99)00024-8 (1999).
Nakayama, T. et al. Assessment of suitable reference genes for RT-qPCR studies in chronic rhinosinusitis. Sci. Rep. 8, 1568. https://doi.org/10.1038/s41598-018-19834-9 (2018).
O’Neill, L. A. J., Kishton, R. J. & Rathmell, J. A guide to immunometabolism for immunologists. Nat. Rev. Immunol. 16, 553–565. https://doi.org/10.1038/nri.2016.70 (2016).
O’Neill, L. A. J. & Pearce, E. J. Immunometabolism governs dendritic cell and macrophage function. J. Exp. Med. 213, 15–23. https://doi.org/10.1084/jem.20151570 (2016).
Tannahill, G. M. et al. Succinate is an inflammatory signal that induces IL-1β through HIF-1α. Nature 496, 238–242. https://doi.org/10.1038/nature11986 (2013).
Cham, C. M., Driessens, G., O’Keefe, J. P. & Gajewski, T. F. Glucose deprivation inhibits multiple key gene expression events and effector functions in CD8+ T cells. Eur. J. Immunol. 38, 2438–2450. https://doi.org/10.1002/eji.200838289 (2008).
Cham, C. M. & Gajewski, T. F. Glucose availability regulates IFN-gamma production and p70S6 kinase activation in CD8+ effector T cells. J. Invest. Med. 174, 4670–4677. https://doi.org/10.4049/jimmunol.174.8.4670 (2005).
Chang, C.-H. et al. Posttranscriptional control of T cell effector function by aerobic glycolysis. Cell 153, 1239–1251. https://doi.org/10.1016/j.cell.2013.05.016 (2013).
Macintyre, A. N. et al. The glucose transporter Glut1 is selectively essential for CD4 T cell activation and effector function. Cell Metab. 20, 61–72. https://doi.org/10.1016/j.cmet.2014.05.004 (2014).
Gerriets, V. A. et al. Metabolic programming and PDHK1 control CD4+ T cell subsets and inflammation. J. Clin. Investig. 125, 194–207. https://doi.org/10.1172/JCI76012 (2015).
Keating, S. E. et al. Metabolic reprogramming supports IFN-γ production by CD56bright NK cells. J. Immunol. (Baltimore, Md., 1950 ) 196, 2552–2560. https://doi.org/10.4049/jimmunol.1501783 (2016).
Mah, A. Y. et al. Glycolytic requirement for NK cell cytotoxicity and cytomegalovirus control. JCI Insight https://doi.org/10.1172/jci.insight.95128 (2017).
Fekete, T. et al. Human plasmacytoid and monocyte-derived dendritic cells display distinct metabolic profile upon RIG-I activation. Front. Immunol. 9, 3070. https://doi.org/10.3389/fimmu.2018.03070 (2018).
Burke, J. D., Platanias, L. C. & Fish, E. N. Beta interferon regulation of glucose metabolism is PI3K/Akt dependent and important for antiviral activity against coxsackievirus B3. J. Virol. 88, 3485–3495. https://doi.org/10.1128/JVI.02649-13 (2014).
Pantel, A. et al. Direct type I IFN but not MDA5/TLR3 activation of dendritic cells is required for maturation and metabolic shift to glycolysis after poly IC stimulation. PLoS Biol. 12, e1001759. https://doi.org/10.1371/journal.pbio.1001759 (2014).
Zhang, W. et al. Lactate is a natural suppressor of RLR signaling by targeting MAVS. Cell 178, 176-189.e15. https://doi.org/10.1016/j.cell.2019.05.003 (2019).
Kelm, D. H., Simon, R., Kuhlow, D., Voigt, C. C. & Ristow, M. High activity enables life on a high-sugar diet: Blood glucose regulation in nectar-feeding bats. Proc. Biol. Sci. 278, 3490–3496. https://doi.org/10.1098/rspb.2011.0465 (2011).
Amitai, O. et al. Fruit bats (Pteropodidae) fuel their metabolism rapidly and directly with exogenous sugars. J. Exp. Biol. 213, 2693–2699. https://doi.org/10.1242/jeb.043505 (2010).
Tristan, C., Shahani, N., Sedlak, T. W. & Sawa, A. The diverse functions of GAPDH: views from different subcellular compartments. Cell. Signal. 23, 317–323. https://doi.org/10.1016/j.cellsig.2010.08.003 (2011).
Tarze, A. et al. GAPDH, a novel regulator of the pro-apoptotic mitochondrial membrane permeabilization. Oncogene 26, 2606–2620. https://doi.org/10.1038/sj.onc.1210074 (2007).
Millet, P., Vachharajani, V., McPhail, L., Yoza, B. & McCall, C. E. GAPDH binding to TNF-α mRNA contributes to posttranscriptional repression in monocytes: A novel mechanism of communication between inflammation and metabolism. J. Invest. Med. 196, 2541–2551. https://doi.org/10.4049/jimmunol.1501345 (2016).
Frick, W. F., Kingston, T. & Flanders, J. A review of the major threats and challenges to global bat conservation. Ann. N. Y. Acad. Sci. 1469, 5–25. https://doi.org/10.1111/nyas.14045 (2020).
Gorbunova, V., Seluanov, A. & Kennedy, B. K. The world goes bats: Living longer and tolerating viruses. Cell Metab. 32, 31–43. https://doi.org/10.1016/j.cmet.2020.06.013 (2020).
Luis, A. D. et al. A comparison of bats and rodents as reservoirs of zoonotic viruses: are bats special?. Proc. Biol. Sci. 280, 20122753. https://doi.org/10.1098/rspb.2012.2753 (2013).
Turmelle, A. S. & Olival, K. J. Correlates of viral richness in bats (order Chiroptera). EcoHealth 6, 522–539. https://doi.org/10.1007/s10393-009-0263-8 (2009).
Halwe, N. J. et al. Egyptian fruit bats (Rousettus aegyptiacus) were resistant to experimental inoculation with avian-origin Influenza A virus of subtype H9N2, but are susceptible to experimental infection with bat-borne H9N2 virus. Viruses https://doi.org/10.3390/v13040672 (2021).
Chomczynski, P. & Sacchi, N. Single-step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction. Anal. Biochem. 162, 156–159. https://doi.org/10.1006/abio.1987.9999 (1987).
Lee, C., Kim, J., Shin, S. G. & Hwang, S. Absolute and relative QPCR quantification of plasmid copy number in Escherichia coli. J. Biotechnol. 123, 273–280. https://doi.org/10.1016/j.jbiotec.2005.11.014 (2006).
Acknowledgements
We thank Dr. Donata Hoffmann and Nico Joel Halwe (Institute of Diagnostic Virology, Friedrich-Loeffler-Institut, Germany) for providing homogenized R. aegyptiacus tissue samples as well as Lisa Loerzer and Silke Rehbein for skilled technical assistance.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
Data curation and data analysis: V.F. and G.P.; sample preparation: A.B.B., V.F. and G.P.; funding, study design and supervision: G.P. and A.D.; manuscript writing and editing: V.F., A.B.B., A.D., G.P.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Friedrichs, V., Balkema-Buschmann, A., Dorhoi, A. et al. Selection and stability validation of reference gene candidates for transcriptional analysis in Rousettus aegyptiacus. Sci Rep 11, 21662 (2021). https://doi.org/10.1038/s41598-021-01260-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-021-01260-z
This article is cited by
-
Bat organoids reveal antiviral responses at epithelial surfaces
Nature Immunology (2025)
-
Expression patterns of housekeeping genes and tissue-specific genes in black goats across multiple tissues
Scientific Reports (2024)
-
Evaluation and validation of reference genes for RT-qPCR gene expression in Naegleria gruberi
Scientific Reports (2023)