Abstract
The effect of long noncoding RNAs (lncRNAs) on the radiotherapy response has been gradually revealed. This systematic review and meta-analysis aimed to evaluate the association between the function and underlying mechanism of lncRNAs in regulating the radiosensitivity and radioresistance of different tumors. Hazard ratios (HRs) with corresponding 95% confidence intervals (CIs) were calculated to estimate the effect of lncRNAs on cancer patient prognosis, including overall survival (OS), recurrence-free survival (RFS), disease-free survival (DFS) and progression-free survival (PFS). Collectively, 23 lncRNAs in 11 cancer types were enrolled. Of them, 13 lncRNAs were downregulated and related to radiosensitivity, 11 lncRNAs were upregulated and related to radioresistance, and 3 lncRNAs were upregulated and related to radiosensitivity in cancers. Furthermore, 17 microRNAs and 20 pathways were targeted by different lncRNAs and contributed to the cancer radiotherapy response in this meta-analysis. The individual pooled HRs (95% CIs) of downregulated radiation-resistant and upregulated radiation-resistant lncRNAs for OS were 0.49 (0.40–0.60) and 1.88 (1.26–2.79), respectively. Our results showed that lncRNAs could modulate tumor radioresistance or sensitivity by affecting radiation-related signaling pathways and serve as potential biomarkers to predict radiotherapy response.
Similar content being viewed by others
Introduction
Multiple treatment options, including surgery, chemotherapy, radiotherapy, targeted therapy, and immunotherapy, have been applied to improve the survival of cancer patients. Of them, radiotherapy is carried out in the treatment of more than half of malignancies1. Radiation contributes to the formation of reactive oxygen species (ROS) and free radicals that lead to DNA single-strand breaks (SSBs) and double-strand breaks (DSBs), ultimately causing cancer cell death2. However, radiotherapy resistance limits its efficacy and has become a major clinical challenge. Although the mechanisms of radioresistance in cancer cells are not clear, they likely involve a series of biological and genetic alterations, such as aberrant DNA damage response (DDR) and enhanced DNA damage repair, disorder of cell proliferation and apoptosis, and related abnormal activation molecular mechanisms3,4.
Long noncoding RNAs (lncRNAs) are defined as transcripts of more than 200 nucleotides (nt), and the vast majority of them are not translated into proteins5,6. Recently, accumulating evidence has investigated the role of lncRNAs in regulating chromatin remodeling, transcription and cell biological behavior, such as proliferation, migration, immortality, and angiogenesis, in malignancies7,8. In particular, the relationship between lncRNAs and the cancer response to radiotherapy has received increased attention. Studies have shown that lncRNAs can modulate radioresistance through multiple processes, such as DNA damage repair, apoptosis, epithelial-mesenchymal transition (EMT) and cancer stem cell (CSC) activity3,9,10. For instance, the lncRNA DNM3OS has been confirmed to inhibit irradiation-induced DDR and confer significant radioresistance in esophageal squamous cell carcinoma (ESCC)11. LncRNA LINC02582 could promote breast cancer radioresistance by interacting with deubiquitinating enzyme ubiquitin specific peptidase 7 (USP7) to deubiquitinate and stabilize checkpoint kinase 1 (CHK1), a key molecule in DDR12. Inhibition of lncRNA HOTAIR could facilitate apoptosis, inhibit autophagy, and enhance the radiosensitivity of CRC after irradiation by regulating the miR-93/ATG12 axis13. LncRNA RBM5-AS1 was proven to be a novel inducer of medulloblastoma stemness, and silencing of RBM5-AS1 enhanced irradiation-induced apoptosis and DDR in medulloblastoma cells14.
These findings imply the potential use of lncRNAs as predictors or diagnostic biomarkers for identifying radioresistant patients in the clinic. Thus, in this study, we performed a systematic review and meta-analysis to investigate and classify the association of lncRNAs with cell response to radiotherapy and prognosis in different types of carcinomas.
Materials and methods
Search strategy
We prospectively registered the systematic review and meta-analysis with PROSPERO (CRD42022359669). This systematic review and meta-analysis was conducted in accordance with PRISMA (Preferred Reporting Items for Systematic Reviews and Meta-Analyses) guidelines15. The 2020 PRISMA flowchart and PRISMA checklist can be found in Fig. 1 and Table S1.
For the meta-analysis, a systematic literature search of the PubMed, Web of Science and Scopus databases (last search updated on September 2021) was conducted to identify studies that evaluated the association between the expression of lncRNAs and responses to radiotherapy and the prognosis of different tumors. The following search terms were used: “lnc RNA or long noncoding RNA” and “radioresistance or radiosensitivity or radiotherapy resistance or radiotherapy sensitivity or radioresistance or radiosensitivity” and “cancer or tumor or carcinoma”. In addition, a manual search for other relevant articles was performed using the reference lists of eligible studies.
Selection criteria
Papers were defined as eligible studies if they met the following criteria: (1) studies of human clinical trials; (2) lncRNA involvement in radioresistance or radiosensitivity data in cancers; (3) quantitative measurement of lncRNA expression in tumor tissues; (4) the association between lncRNA expression and survival was estimated; and (5) sufficient data for the assessment of hazard ratios (HRs) and 95% confidence intervals (CIs).
The exclusion criteria were as follows: (1) only cell or animal experiments, case reports, editorials, letters, meta-analyses, or reviews; (2) articles where the required data could not be extracted from the original papers; (3) nonrelated studies to radiotherapy; (4) studies without a definite patient sample size; and (5) lack of sufficient data to calculate HR and 95% CI.
Data extraction and quality assessment
The following data were extracted from all included studies: the first author’s name, year of study or publication, country, tumor type, involved lncRNAs, number and type of samples, patient gender, detection methods for lncRNA expression, cut-off level, outcomes, lncRNA expression and function, type of radiotherapy response, associated microRNAs and pathways. The prognostic endpoints included overall survival (OS), recurrence-free survival (RFS), disease-free survival (DFS) and progression-free survival (PFS). Additionally, the HR estimates were calculated from Kaplan–Meier survival curves using Engauge Digitizer V4.1 software according to the method of Tierney16 if these were not available (https://sourceforge.net/projects/digitizer/). Quality assessment of the primary studies was executed using the Newcastle–Ottawa Quality Assessment Scale17.
Statistical analysis
A Cochrane’s Q test was implemented to test heterogeneity among studies. The p value of the Q test was < 0.1, which suggested the presence of heterogeneity, and the random effects model (DerSimonian–Laird method)18 was used to calculate pooled HRs. Otherwise, heterogeneity was absent, and the fixed effects model (Mantel–Haenszel method)19 was more appropriate and applied. In addition, the degree of heterogeneity was assessed by the I2 test. The value of I2 ranged from 0 to 100% and was generally considered no heterogeneity for I2 = 0, moderate heterogeneity for 25%, large heterogeneity to 50%, and extreme heterogeneity for 75%20. Furthermore, a funnel plot and Begg and Egger’s tests21,22,23 were utilized to investigate any possible publication bias. The funnel plot was visually symmetrical, and the P value of Begger’s or Egger’s test was greater than 0.05, which indicated that there was no statistically significant publication bias. Sensitivity analysis was performed by consecutive omission of individual studies to verify the consistency of outcomes. Statistical analyses were performed with STATA software version 14.0 (https://www.stata.com/) and Review Manager 5.3 software (http://www.cochrane.org/cochrane/download.htm).
Results
Study selection and characteristics of eligible studies
The primary search strategy yielded a total of 2613 publications from PubMed (n = 1250), Web of Science (n = 597) and Scopus (n = 766), 1395 of which were excluded due to duplication. According to the inclusion and exclusion criteria, 29 studies about the association between lncRNAs and radiosensitivity or radioresistance in various cancers were ultimately used for this meta-analysis, as shown in Fig. 1. The main characteristics of the included studies are summarized in Table 1. A total of 2602 patients between 2015 and 2021 were included. They all came from China. The malignant tumors in the study included head and neck squamous cell carcinoma (HNSCC)24,25, nasopharyngeal carcinoma (NPC)26,27,28,29,30,31,32, glioma33,34, laryngeal carcinoma (LC)35, esophageal cancer (EC)36,37,38, non-small-cell lung cancer (NSCLC)39,40,41,42, lung adenocarcinoma (LUAD)43, cardiac cancer (CaC)44, colorectal cancer (CRC)13,45, cervical cancer (CeC)46,47, breast cancer (BrC)12,48,49, and bladder cancer (BlC)50. The level of lncRNA expression was measured with quantitative reverse transcription polymerase chain reaction (qRT–PCR) and/or in situ hybridization (ISH) in the included studies. The cut-off score was the median value when using qRT–PCR. In ISH, a score of simultaneous immunohistochemistry (SI) < 6 was considered low expression (Table 1).
Association between lncRNAs and radiotherapy response in vitro
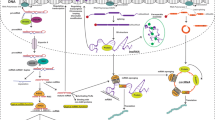
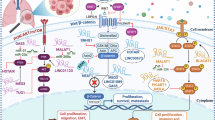
There were 23 lncRNAs in 11 cancer types included in our systematic review and meta-analysis. Of them, 13 lncRNAs were downregulated by small-interfering RNA (siRNA) transfection, and 14 were upregulated by lentiviral vector infection. Furthermore, 16 lncRNAs were observed to be related to radiosensitivity, while 11 were found to be associated with radioresistance. Reduced expression of BLACAT1 in HNSCC; MALAT1, NEAT1 and PVT1 in NPC; FAM201A, PVT1 and LINC00857 in NSCLC; LINC00473, CCAT2and Rpph1 in EC; HOTAIR and LINC00958 in CRC; AFAP1-AS1 and LINC00511 in breast cancer were correlated with radiosensitivity. Increased expression of LINC00473 in HNSCC; LINC00114, MINCR and PVT1 in NPC; Linc-RA1 in glioma; LINC00473and CYTOR in NSCLC; NEAT1 and LINC00958 in cervical cancer; H19 in cardiac cancer; LINC02582 in breast cancer; and TUG1 in bladder cancer were associated with radioresistance. However, some elevated expression of LINC00312 in NPC, TPTEP1 in glioma, and NKILA in laryngeal carcinoma was found to be related to radiosensitivity. Four lncRNAs appeared in more than one type of cancer and were differentially regulated, including downregulation of LINC00473 in ESCC radiosensitivity and upregulation in HNSCC and NSCLC radioresistance, downregulation of NEAT1 in NPC radiosensitivity and upregulation in cervical cancer radioresistance, downregulation of LINC00958 in colorectal cancer radiosensitivity and upregulation in cervical cancer radioresistance, downregulation of PVT1 in NPC and NSCLC radiosensitivity and upregulation in NPC radioresistance. Additionally, 17 miRNAs and 20 pathways or genes were targeted by different lncRNAs, which contributed to DNA damage repair, apoptosis, CSC regulation, EMT, etc., causing cancer cell radioresistance or sensitivity (Table 2 and Fig. 2).
Association between OS and lncRNA expression with regard to radiotherapy response
A total of 27 studies were included for OS analysis. Under the random-effects model, the individual overall HR (95% CI) of downregulated radiotherapyresistant lncRNAs was 0.49 (0.40–0.60), with extreme heterogeneity (I2 = 0.0%; Ph = 0.486) (Fig. 3A). The individual overall HR (95% CI) of upregulated radiotherapy-resistant lncRNAs was 1.88 (1.26–2.79) (Fig. 3B). The individual overall HR (95% CI) of upregulated radiotherapy-sensitive lncRNAs was 0.37 (0.23–0.58) from few studies (Fig. 3C). Our results showed that downregulated radiotherapy-resistant lncRNAs were associated with better OS in cancer patients treated with radiotherapy. Similar patterns were found for DFS (random-effects model: pooled HR 0.28; 95% CI 0.10–0.75), although there were few qualified articles included in the analysis. However, no significant difference was found in the RFS and PFS analyses (Fig. S1).
Subgroup analysis of the association between OS and lncRNA expression with regard to radiotherapy response
Subgroup analyses were performed based on the type of cancer. Head and neck cancers included HNSCC, NPC, glioma and LC; respiratory system tumors included NSCLC and LUAD; and digestive tract system tumors included cardiac cancer, CRC and EC. Under the random-effects model, the HR and 95% CI of upregulated radiotherapy-resistant lncRNAs were 2.33 (1.21–4.48) for cervical cancer studies, and 3.83 (2.03–7.22) for bladder cancer studies (Fig. 4). Our results showed that upregulated radiotherapy-resistant lncRNAs were associated with worse OS in these cancer patients treated with radiotherapy.
However, for head and neck cancer studies, the individual HR (95% CI) of downregulated radiotherapy-resistant lncRNAs was 0.47 (0.35–0.64), the individual HR (95% CI) of upregulated radiotherapy-resistant lncRNAs was 1.66 (1.11–2.48) and the individual HR (95% CI) of upregulated radiotherapy-sensitive lncRNAs was 0.34 (0.22–0.52), respectively (Fig.S2). For breast cancer studies, the individual HR (95% CI) of downregulated radiotherapy-resistant lncRNAs was 1.02 (0.44–2.36). The results implied that there was no statistically significant in the association of radiotherapy-related lncRNAs and survival in breast cancer (Fig. S2).
Furthermore, in the respiratory system and digestive tract system subgroups, the individual HRs (95% CIs) of downregulated radiotherapy-resistant lncRNAs were 0.45 (0.30–0.68) and 0.43 (0.25–0.76), respectively. Correspondingly, the HRs of upregulated radiotherapy-resistant lncRNAs were 2.82 (1.03–7.72) and 1.62 (1.07–2.46), respectively. The results also indicated that downregulated radiotherapy-resistant lncRNA expression was associated with radiosensitivity and could be a better prognostic marker for those patients (Fig. S2).
Publication bias
Begg’s funnel plot and Egger’s test were performed to detect publication bias. There was no obvious asymmetry in Begg’s funnel plots of OS (Fig. 5, p = 0.967). The p values of Egger’s tests were all greater than 0.05, indicating no potential publication bias in our study.
Sensitivity analysis
To assess the stability of the pooled results in the current meta-analysis, we conducted a sensitivity analysis by gradually removing each single study from the merged analysis. The results indicated that the overall pooled estimates could not be affected by a single study (Fig. S3).
Discussion
Radiotherapy is considered a highly effective cancer treatment. Radiotherapy resistance is still the main cause of treatment failure, leading to tumor recurrence and metastasis. Tumor and microenvironment heterogeneity is considered to be responsible for the differing sensitivities of tumor cells to cancer treatment, including radiotherapy, which explains why tumor subpopulations are not equally affected by this treatment51,52. Therefore, it is crucial to explore the targets and mechanisms of radiotherapy resistance, providing some potential therapeutic points for improving cancer radiotherapy.
Various studies have confirmed that deregulated lncRNAs contribute to radiotherapy resistance by regulating abnormal DDR, apoptosis, CSCs and EMT in many cancers, such as NPC, breast cancer, NSCLC, ESCC, cervical cancer and colorectal cancer12,29,42,45,46,53,54. In the current study, we performed the first comprehensive systematic review and meta-analysis aimed to evaluate the effect of lncRNA expression on radiotherapy response and patient prognosis in various human cancers.
Our meta-analysis showed that lncRNAs with lower expression were significantly associated with a better radiotherapy response and OS, while lncRNAs with higher expression were related to radiotherapy resistance. In the subgroup analysis, based on cancer type, lncRNAs were determined to be a potential predictor for prognosis in various cancers, includingseveral neoplasms of the genitourinary system (e.g., cervical and bladder cancers)46,47,55. Previous studies have shown that urothelial cells in bladder cancer have high radioresistance. LncRNAs may act as potential regulators of radioresistance in these cancers, and they may represent promising therapeutic targets by interfering with their expression55. Most head and neck cancers are radiation-sensitive squamous carcinomas. However, the impact of radiotherapy-related lncRNAs on the prognosis of patients with head and neck cancers is diverse and inconsistent. In addition, we could not find a significant role of lncRNAs in the radiotherapy response for breast cancer due to insufficient studies reporting clinical outcomes12,48,49. More research needs to be done to clarify the effect of lncRNAs on radiotherapy in these cancers.
Furthermore, lncRNAs mediate the tumor response to radiotherapy by acting on a wide range of target genes and pathways. Numerous studies have confirmed that lncRNAs may serve as endogenous sponges to regulate the expression and function of miRNAs and subsequently radiotherapy regulatory pathways10,37,42. We found that more than one lncRNA/miRNA interaction has been investigated in distinctive cancer cell lines and tissues (e.g., NPC, NSCLC, CRC, cervical cancer and breast cancer). NPC, where radiotherapy is the main and essential treatment, has been the most studied to explore the effect of lncRNAs on radiotherapy26,27,28,29,30,31,32. In NPC, PVT1 regulates DNA repair and cell apoptosis to induce radioresistance via the miR-515-5p/PIK3CA axis and modulates the AKT pathway by interacting with miR-515-5p26. MALAT1 modulates CSC activity and radioresistance by inactivating miR-131. NEAT1 modulates EMT and radioresistance through the miR-204/ZEB1 axis30. Thus, lncRNAs could act as promising theragnostic biomarkers for predicting radioresistance and radiosensitivity. In addition, a previous meta-analysis indicated that miRNAs contribute to the cellular response to irradiation by affecting radiation-related signaling pathways, such as the PI3K/AKT/mTOR pathway56. It seems that miRNAs accompanied by lncRNAs act as theragnostic biomarkers to predict and monitor the clinical response to radiotherapy. In addition, specific lncRNAs can also appear in different types of tumors. For example, LINC00473 facilitated radioresistance in ESCC, HNSCC and NSCLC by modulating the miR‐374a‐5p/SPIN1 axis, activating the Wnt/β-catenin pathway and sponging miR-513a-3p, respectively25,36,40. Numerous studies have proven that LINC00473, a novel lncRNA, is upregulated and acts as an oncogene in most human cancers, leading to tumor progression57,58. Furthermore, the overexpression of LINC00473 is strongly associated with poor survival59,60. Our study showed that LINC00473 regulated the sensitivity of radiotherapy for several different neoplasms25,36,40. In addition, the MAPK pathway and p53 pathway were found to be the main mechanisms involved in the regulation of the radiotherapy response mediated by lncRNAs.
There were several limitations in our meta-analysis. Heterogeneity was observed to be high within the forest plots of the total lncRNAs and some subgroups, suggesting that HRs vary across studies. The indications of radiation therapy in different cancers and sometimes in different stages of the same cancer type are different. It can be with curative intent, palliative, adjuvant or neoadjuvant, etc. which should contribute to the heterogeneity among the studies. Thus, the random-effects model was applied in these analyses. This could be explained by the number of patients, various types of tumors, variable cut-off points and, most importantly, the upregulation and downregulation of lncRNA expression integrated for calculation. Fortunately, according to the sensitivity analyses, we could suggest that, despite heterogeneity, the pooled HR can be considered quite reliable and representative. Although we performed a systematic and extensive literature search, the studies included only Asian patients and caused singleness of the population. Thus, there were not enough eligible studies for all types of cancers involving radiotherapy (e.g., lack of soft tissue tumors, lymphoma, etc.). Additionally, only nearly 14% of the included papers had direct hazard values that could be applied for the current meta-analysis, leading to bias and decreased accuracy of the results. The HR for OS in our study was retrieved by KM curve; thus, this is a univariable analysis. However, HR should be extracted by multivariable analysis to avoid selection bias and remove confounders as much as possible. These results suggest that further associated studies are needed to precisely determine the effect of lncRNA expression on radiotherapy response and prognosis.
Conclusion
In conclusion, this comprehensive systematic review and meta-analysis showed the role of lncRNA expression on radiotherapy response and survival in various malignancies. Furthermore, we provided lists of potential radiation-related signaling pathways controlled by lncRNA/miRNA interactions that might help to identify new therapeutic targets to counter radioresistance.
Data availability
All data generated or analyzed during this study are included in this published article and the Supplementary Information files.
Abbreviations
- BrC:
-
Breast cancer
- BlC:
-
Bladder cancer
- CSC:
-
Cancer stem cell
- CHK1:
-
Checkpoint kinase 1
- CaC:
-
Cardiac cancer
- CRC:
-
Colorectal cancer
- CeC:
-
Cervical cancer
- DFS:
-
Disease-free survival
- DSB:
-
Double-strand break
- DDR:
-
DNA damage response
- EMT:
-
Epithelial-mesenchymal transition
- ESCC:
-
Esophageal squamous cell carcinoma
- EC:
-
Esophageal cancer
- HR:
-
Hazard ratio
- HNSCC:
-
Head and neck squamous cell carcinoma
- ISH:
-
In situ hybridization
- lncRNA:
-
Long noncoding RNA
- LC:
-
Laryngeal carcinoma
- LUAD:
-
Lung adenocarcinoma
- miRNA:
-
MicroRNA
- NSCLC:
-
Non-small-cell lung cancer
- NPC:
-
Nasopharyngeal carcinoma
- OS:
-
Overall survival
- PFS:
-
Progression-free survival
- PRISMA:
-
Preferred Reporting Items for Systematic Reviews and Meta-Analyses
- qRT–PCR:
-
Quantitative reverse transcription polymerase chain reaction
- RFS:
-
Recurrence-free survival
- ROS:
-
Reactive oxygen species
- SSB:
-
Single-strand break
- SI:
-
Simultaneous immunohistochemistry
- siRNA:
-
Small-interfering RNA
- USP7:
-
Ubiquitin specific peptidase 7
References
Citrin, D. E. Recent developments in radiotherapy. N. Engl. J. Med. 377, 1065–1075. https://doi.org/10.1056/NEJMra1608986 (2017).
Rey, S., Schito, L., Koritzinsky, M. & Wouters, B. G. Molecular targeting of hypoxia in radiotherapy. Adv. Drug Deliv. Rev. 109, 45–62. https://doi.org/10.1016/j.addr.2016.10.002 (2017).
Zhang, X. et al. Role of non-coding RNAs and RNA modifiers in cancer therapy resistance. Mol. Cancer 19, 47. https://doi.org/10.1186/s12943-020-01171-z (2020).
Ebahimzadeh, K. et al. Emerging role of non-coding RNAs in response of cancer cells to radiotherapy. Pathol. Res. Pract. 218, 153327. https://doi.org/10.1016/j.prp.2020.153327 (2021).
Kopp, F. & Mendell, J. T. Functional classification and experimental dissection of long noncoding RNAs. Cell 172, 393–407. https://doi.org/10.1016/j.cell.2018.01.011 (2018).
Palazzo, A. F. & Koonin, E. V. Functional long non-coding RNAs evolve from junk transcripts. Cell 183, 1151–1161. https://doi.org/10.1016/j.cell.2020.09.047 (2020).
Fang, Y. & Fullwood, M. J. Roles, functions, and mechanisms of long non-coding RNAs in cancer. Genomics Proteomics Bioinform. 14, 42–54. https://doi.org/10.1016/j.gpb.2015.09.006 (2016).
Li, J., Tian, H., Yang, J. & Gong, Z. Long noncoding RNAs regulate cell growth, proliferation, and apoptosis. DNA Cell Biol. 35, 459–470. https://doi.org/10.1089/dna.2015.3187 (2016).
Zhu, J. et al. Molecular mechanisms of lncRNAs in regulating cancer cell radiosensitivity. LID—BSR20190590. 10.1042/BSR20190590.
Podralska, M., Ciesielska, S., Kluiver, J. & van den Berg, A. Non-coding RNAs in cancer radiosensitivity: MicroRNAs and lncRNAs as regulators of radiation-induced signaling pathways. Cancers 12, 1662. https://doi.org/10.3390/cancers12061662 (2020).
Zhang, H. et al. Cancer-associated fibroblast-promoted LncRNA DNM3OS confers radioresistance by regulating DNA damage response in esophageal squamous cell carcinoma. Clin. Cancer Res. 25, 1989–2000. https://doi.org/10.1158/1078-0432.ccr-18-0773 (2019).
Wang, B. Y. et al. Long noncoding RNA LINC02582 acts downstream of miR-200c to promote radioresistance through CHK1 in breast cancer cells. Cell Death Dis. 10, 10 (2019).
Liu, Y. et al. Long non-coding RNA HOTAIR knockdown enhances radiosensitivity through regulating microRNA-93/ATG12 axis in colorectal cancer. Cell Death Dis. 11, 175. https://doi.org/10.7150/ijms.4570610.1038/s41419-020-2268-8 (2020).
Zhu, C., Li, K., Jiang, M. & Chen, S. RBM5-AS1 promotes radioresistance in medulloblastoma through stabilization of SIRT6 protein. Acta Neuropathol. Commun. 9, 123. https://doi.org/10.1186/s40478-021-01218-2 (2021).
Page, M. J. et al. The PRISMA statement: An updated guideline for reporting systematic reviews. BMJ (Clin. Res. Ed.) 372(71), 2021. https://doi.org/10.1136/bmj.n71 (2020).
Tierney, J. F., Stewart, L. A., Ghersi, D., Burdett, S. & Sydes, M. R. Practical methods for incorporating summary time-to-event data into meta-analysis. Trials 8, 16. https://doi.org/10.1186/1745-6215-8-16 (2007).
Deeks, J. J. et al. Evaluating non-randomised intervention studies. Health Technol. Assess. (Winch. Eng.) 7, 1–173. https://doi.org/10.3310/hta7270 (2003).
DerSimonian, R. & Laird, N. Meta-analysis in clinical trials. Control. Clin. Trials 7, 177–188. https://doi.org/10.1016/0197-2456(86)90046-2 (1986).
Mantel, N. & Haenszel, W. Statistical aspects of the analysis of data from retrospective studies of disease. J. Natl. Cancer Inst. 22, 719–748 (1959).
Higgins, J. P., Thompson, S. G., Deeks, J. J. & Altman, D. G. Measuring inconsistency in meta-analyses. BMJ (Clin. Res. Ed.) 327, 557–560. https://doi.org/10.1136/bmj.327.7414.557 (2003).
Begg, C. B. & Mazumdar, M. Operating characteristics of a rank correlation test for publication bias. Biometrics 50, 1088–1101 (1994).
Egger, M., Davey Smith, G., Schneider, M. & Minder, C. Bias in meta-analysis detected by a simple, graphical test. BMJ (Clin. Res. Ed.) 315, 629–634. https://doi.org/10.1136/bmj.315.7109.629 (1997).
Sterne, J. A. & Egger, M. Funnel plots for detecting bias in meta-analysis: Guidelines on choice of axis. J. Clin. Epidemiol. 54, 1046–1055. https://doi.org/10.1016/s0895-4356(01)00377-8 (2001).
Gou, C. et al. Knockdown of lncRNA BLACAT1 enhances radiosensitivity of head and neck squamous cell carcinoma cells by regulating PSEN1. Br. J. Radiol. 93, 20190154. https://doi.org/10.1259/bjr.20190154 (2020).
Han, P. B., Ji, X. J., Zhang, M. & Gao, L. Y. Upregulation of lncRNA LINC00473 promotes radioresistance of HNSCC cells through activating Wnt/γ-catenin signaling pathway. Eur. Rev. Med. Pharmacol. Sci. 22, 7305–7313. https://doi.org/10.26355/eurrev-201811-16267 (2018).
Han, Y., Li, F., Xie, J., Wang, Y. & Zhang, H. PVT1 mediates cell proliferation, apoptosis and radioresistance in nasopharyngeal carcinoma through regulating miR-515-5p/PIK3CA axis. Cancer Manag. Res. 12, 10077–10090. https://doi.org/10.2147/cmar.s257583 (2020).
He, Y. et al. Long non-coding RNA PVT1 predicts poor prognosis and induces radioresistance by regulating DNA repair and cell apoptosis in nasopharyngeal carcinoma. Cell Death Dis. 9, 235. https://doi.org/10.1038/s41419-020-02977-x10.1038/s41419-018-0265-y (2018).
Guo, Z. et al. LncRNA linc00312 suppresses radiotherapy resistance by targeting DNA-PKcs and impairing DNA damage repair in nasopharyngeal carcinoma. Cell Death Dis. 12, 2 (2021).
Han, Y. Y. et al. LINC00114 promoted nasopharyngeal carcinoma progression and radioresistance in vitro and in vivo through regulating ERK/JNK signaling pathway via targeting miR-203. Eur. Rev. Med. Pharmacol. Sci. 24, 2491–2504. https://doi.org/10.26355/eurrev_202003_20517 (2020).
Lu, Y. et al. The long non-coding RNA NEAT1 regulates epithelial to mesenchymal transition and radioresistance in through miR-204/ZEB1 axis in nasopharyngeal carcinoma. Tumour Biol. 37, 11733–11741. https://doi.org/10.1007/s13277-015-4773-4 (2016).
Jin, C., Yan, B., Lu, Q., Lin, Y. & Ma, L. The role of MALAT1/miR-1/slug axis on radioresistance in nasopharyngeal carcinoma. Tumour Biol. 37, 4025–4033. https://doi.org/10.1007/s13277-015-4227-z (2016).
Zhong, Q., Chen, Y. & Chen, Z. LncRNA MINCR regulates irradiation resistance in nasopharyngeal carcinoma cells via the microRNA-223/ZEB1 axis. Int. J. Mol. Sci. 19, 53–66. https://doi.org/10.3390/ijms2118678710.1080/15384101.2019.1692176 (2020).
Zheng, J. et al. Linc-RA1 inhibits autophagy and promotes radioresistance by preventing H2Bub1/USP44 combination in glioma cells. Gastric Cancer 11, 758. https://doi.org/10.1007/s10120-020-01096-y10.1038/s41419-020-02977-x (2020).
Tang, T., Wang, L. X., Yang, M. L. & Zhang, R. M. lncRNA TPTEP1 inhibits stemness and radioresistance of glioma through miR-106a-5p-mediated P38 MAPK signaling. Mol. Med. Rep. 22, 4857–4867. https://doi.org/10.1038/s41419-018-0265-y10.3892/mmr.2020.11542 (2020).
Yang, T. et al. lncRNA-NKILA/NF-κB feedback loop modulates laryngeal cancer cell proliferation, invasion, and radioresistance. Cancer Med. 7, 2048–2063. https://doi.org/10.1016/j.canlet.2021.03.02510.1002/cam4.1405 (2018).
Lamperska, K., Chen, W. & Zhang, Y. LINC00473/miR-374a-5p regulates esophageal squamous cell carcinoma via targeting SPIN1 to weaken the effect of radiotherapy. Non-coding RNA 120, 14562–14572. https://doi.org/10.3390/ncrna404003310.1002/jcb.28717 (2019).
Wang, M. et al. lncRNA CCAT2 promotes radiotherapy resistance for human esophageal carcinoma cells via the miR-145/p70S6K1 and p53 pathway. Int. J. Oncol. 56, 327–336. https://doi.org/10.2174/138920022166620010610520110.3892/ijo.2019.4929 (2020).
Li, Z. Y. et al. Value of long non-coding RNA Rpph1 in esophageal cancer and its effect on cancer cell sensitivity to radiotherapy. J. Cell. Physiol. 26, 1775–1791. https://doi.org/10.1002/jcp.2890210.3748/wjg.v26.i15.1775 (2020).
Ren, D. et al. Long noncoding RNA FAM201A involves in radioresistance of non-small-cell lung cancer by enhancing EGFR expression via miR-370. Mol. Cancer 23, 5802–5814. https://doi.org/10.1186/s12943-020-01217-210.26355/eurrev_201907_18319 (2019).
Qin, P., Li, Y., Liu, J. & Wang, N. Knockdown of LINC00473 promotes radiosensitivity of non-small cell lung cancer cells via sponging miR-513a-3p. Free Radic. Res. 54, 756–764. https://doi.org/10.1080/10715762.2020.1841900 (2020).
Wu, D., Li, Y., Zhang, H. & Hu, X. Knockdown of lncrna PVT1 enhances radiosensitivity in non-small cell lung cancer by sponging Mir-195. Cell. Physiol. Biochem. 42, 2453–2466. https://doi.org/10.1159/000480209 (2017).
Zhang, J. & Li, W. Q. Long noncoding RNA CYTOR sponges miR-195 to modulate proliferation, migration, invasion and radiosensitivity in nonsmall cell lung cancer cells. Biosci. Rep. 38, 20181599. https://doi.org/10.1042/bsr20181599 (2018).
Han, F. et al. Silencing of lncRNA LINC00857 enhances BIRC5-dependent radio-sensitivity of lung adenocarcinoma cells by recruiting NF-κB1. Mol. Ther. Nucleic Acids 22, 981–993. https://doi.org/10.1016/j.omtn.2020.09.020 (2020).
Jia, J. et al. LncRNA H19 interacted with miR-130a-3p and miR-17-5p to modify radio-resistance and chemo-sensitivity of cardiac carcinoma cells. Cancer Med. 8, 1604–1618. https://doi.org/10.1002/cam4.1860 (2019).
Liang, H., Zhao, Q., Zhu, Z., Zhang, C. & Zhang, H. Long noncoding RNA LINC00958 suppresses apoptosis and radiosensitivity of colorectal cancer through targeting miR-422a. Cancer Cell Int. 21, 477. https://doi.org/10.1186/s12935-021-02188-0 (2021).
Zhao, H. et al. Long noncoding RNA LINC00958 regulates cell sensitivity to radiotherapy through RRM2 by binding to microRNA-5095 in cervical cancer. J. Cell Physiol. 234, 23349–23359. https://doi.org/10.1111/jgh.1360610.1002/jcp.28902 (2019).
Han, D. M., Wang, J. F. & Cheng, G. H. LncRNA NEAT1 enhances the radio-resistance of cervical cancer via miR-193b-3p/CCND1 axis. Oncotarget 9, 2395–2409. https://doi.org/10.18632/oncotarget.23416 (2018).
Liu, L., Zhu, Y., Liu, A. M., Feng, Y. & Chen, Y. Long noncoding RNA LINC00511 involves in breast cancer recurrence and radioresistance by regulating STXBP4 expression via miR-185. Eur. Rev. Med. Pharmacol. Sci. 23, 7457–7468. https://doi.org/10.1016/j.yexmp.2018.11.01010.26355/eurrev_201909_18855 (2019).
Bi, Z. et al. Nanoparticles (NPs)-meditated LncRNA AFAP1-AS1 silencing to block Wnt/β-catenin signaling pathway for synergistic reversal of radioresistance and effective cancer radiotherapy. Adv. Sci. 7, 2000915. https://doi.org/10.1002/advs.202000915 (2020).
Tan, J., Qiu, K., Li, M. & Liang, Y. Double-negative feedback loop between long non-coding RNA TUG1 and miR-145 promotes epithelial to mesenchymal transition and radioresistance in human bladder cancer cells. FEBS Lett. 589, 3175–3181. https://doi.org/10.1016/j.febslet.2015.08.020 (2015).
Barker, H. E., Paget, J. T., Khan, A. A. & Harrington, K. J. The tumour microenvironment after radiotherapy: Mechanisms of resistance and recurrence. Nat. Rev. Cancer 15, 409–425. https://doi.org/10.1038/nrc3958 (2015).
Schaue, D. & McBride, W. H. Opportunities and challenges of radiotherapy for treating cancer. Nat. Rev. Clin. Oncol. 12, 527–540. https://doi.org/10.1038/nrclinonc.2015.120 (2015).
Olivares-Urbano, M. A. & Griñán-Lisón, C. CSC radioresistance: A therapeutic challenge to improve radiotherapy effectiveness in cancer. Cells 9, 1651. https://doi.org/10.3390/cells9071651 (2020).
Sun, Y. et al. LINC00657 played oncogenic roles in esophageal squamous cell carcinoma by targeting miR-615-3p and JunB. Biomed. Pharmacother. 108, 316–324. https://doi.org/10.1016/j.biopha.2018.09.003 (2018).
Jiang, H., Hu, X., Zhang, H. & Li, W. Down-regulation of LncRNA TUG1 enhances radiosensitivity in bladder cancer via suppressing HMGB1 expression. Radiat. Oncol. (Lond.) 12, 65. https://doi.org/10.1186/s13014-017-0802-3 (2017).
Mohammadi, C., Gholamzadeh Khoei, S., Fayazi, N., Mohammadi, Y. & Najafi, R. miRNA as promising theragnostic biomarkers for predicting radioresistance in cancer: A systematic review and meta-analysis. Crit. Rev. Oncol. Hematol. 157, 103183. https://doi.org/10.1016/j.critrevonc.2020.103183 (2021).
Bai, J., Zhao, W. Y., Li, W. J., Ying, Z. W. & Jiang, D. Q. Long noncoding RNA LINC00473 indicates a poor prognosis of breast cancer and accelerates tumor carcinogenesis by competing endogenous sponging miR-497. Eur. Rev. Med. Pharmacol. Sci. 23, 3410–3420. https://doi.org/10.26355/eurrev_201904_17705 (2019).
Mo, J. et al. LINC00473 promotes hepatocellular carcinoma progression via acting as a ceRNA for microRNA-195 and increasing HMGA2 expression. Biomed. Pharmacother. 120, 109403. https://doi.org/10.1016/j.biopha.2019.109403 (2019).
Li, L., Zhang, X., Liu, N., Chen, X. & Peng, C. LINC00473: A novel oncogenic long noncoding RNA in human cancers. J. Cell Physiol. 236, 4174–4183. https://doi.org/10.1002/jcp.30176 (2021).
Chen, Z., Lin, S. & Li, J. L. CRTC1-MAML2 fusion-induced lncRNA LINC00473 expression maintains the growth and survival of human mucoepidermoid carcinoma cells. Oncogene 37, 1885–1895. https://doi.org/10.1038/s41388-017-0104-0 (2018).
Wang, M. et al. IncRNA CCAT2 promotes radiotherapy resistance for human esophageal carcinoma cells via the miR-145/p70S6K1 and p53 pathway. Int. J. Oncol. 56, 327–336. https://doi.org/10.3892/ijo.2019.4929 (2020).
Funding
This work was supported by the National Natural Science Foundation of China (Grant Number: 81902723).
Author information
Authors and Affiliations
Contributions
Y.X. and Q.G. conceived the study concept and design. Y.X. and J.H. collected the data. Y.X. performed statistical analysis. Y.X. and K.X. drafted the manuscript, and all authors significantly contributed to the critical revision of the manuscript and data interpretation, and approved the submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Xie, Y., Han, J., Xie, K. et al. LncRNAs as biomarkers for predicting radioresistance and survival in cancer: a meta-analysis. Sci Rep 12, 18494 (2022). https://doi.org/10.1038/s41598-022-21785-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-022-21785-1
This article is cited by
-
Non-coding RNAs as modulators of radioresponse in triple-negative breast cancer: a systematic review
Journal of Biomedical Science (2024)