Abstract
Metabolism, known to be temporally regulated to meet evolving energy demands, plays a crucial role in shaping developmental pace. Recent studies have demonstrated that two key proteins PARP1 and PARG play a regulatory role in the transcription of both morphogenic and metabolic genes. Intriguingly, in Drosophila, the depletion of PARP1 or PARG proteins causes a developmental arrest before pupation, resulting in individuals unable to complete their development. This phenotype highlights the critical involvement of poly(ADP-ribosyl)ating enzymes in regulating the metamorphic process. In this study, we provide compelling evidence that these enzymes intricately coordinate transcriptional changes in both developmental and metabolic pathways during metamorphosis. Specifically, they promote the expression of genes crucial for pupation, while simultaneously negatively regulating the expression of metabolic genes before the transition to the pupal stage. Additionally, these enzymes suppress the expression of genes that are no longer required during this transformative period. Our findings shed light on the intricate interplay between poly(ADP-ribosyl)ating enzymes, developmental processes, and metabolic regulation before metamorphosis and highlight a new role of poly(ADP-ribosyl)ating enzymes in the global regulation of transcription.
Similar content being viewed by others
Introduction
The orchestration of multicellular organism development relies on the sequential progression of proliferation and differentiation, each phase being accompanied by the activation of specific gene expression programs and diverse energy requirements1,2. Previous studies have demonstrated that metabolism is not a constant process throughout an organism's life but rather exhibits temporal regulation to match the dynamic energy demands3,4. Additionally, it has been observed that metabolism influences the pace of development5, further emphasizing the need for coordinated regulation between developmental progression and metabolic activities.
Poly(ADP-ribose) polymerase 1 (PARP1), a chromatin-associated protein, has emerged as a critical player in development6,7. PARP1 synthesizes poly(ADP-ribose) chains that modify chromatin proteins, resulting in their dissociation from DNA and subsequent chromatin opening8,9,10. Conversely, poly(ADP-ribose) glycohydrolase (PARG) reverses PARP1 activity by removing poly(ADP-ribose) moieties from acceptor proteins11,12,13. Recent studies have revealed a complementary, rather than antagonistic, relationship between PARG and PARP112,14,15,16. Upon activation, PARP1 undergoes automodification and dissociates from chromatin, leading to its inactivation8,17,18. In contrast, by removing poly(ADP-ribose) moieties from PARP1, PARG allows for its re-association with chromatin.
In fruit flies, both parp1C03256 hypomorph and parg27.1 null mutant animals suffer developmental arrest just prior to pupation stage, while their embryonic and larval development is temporally comparable to wildtype animals11,18. Consequently, the animals are viable but remain third instar larvae, unable to complete their development7. Previous findings suggest that both PARG and PARP1 regulate the transcription of developmental genes during the larva-to-pupa transition14. However, PARG lacks a DNA-binding domain (DBD) and seems to interact with chromatin only to a limited extent19, the specific interaction between PARG and chromatin in the context of the protein’s recruitment remains to be elucidated. We hypothesized that PARG is recruited to regulate transcription through its interaction with PARP1. To investigate this possibility, we conducted chromatin immunoprecipitation (ChIP) of PARG and PARP1 during the late third instar larval stage. Our results demonstrate that both PARP1 and PARG directly control global changes in the transcriptional profile that occurs prior to metamorphosis for both metabolic and developmental processes but through distinct mechanisms. Firstly, together with PARP1, PARG binds to the promoter region of genes crucial for pupation, promoting their expression. Secondly, in the absence of PARP1, PARG binds to the gene body of metabolic genes, repressing their expression, and binds to genes that are no longer required at the end of larval stages, repressing their expression as well. Together, these results suggest that PARP1 and PARG coordinate the regulation of metabolism and developmental genes during developmental progression.
Results
PARG binds to chromatin.
To assess the chromatin-binding capacity of PARG, we conducted chromatin immunoprecipitation followed by sequencing (ChIP-seq) assays using wandering third instar larvae (puffstage 7–9), expressing PARG-YFP in an endogenous parg mutant background, which was compared to a yellow white control line lacking GFP expression. We specifically chose this stage of wandering third instar larvae because it is the final stage before parg and parp1 mutants experience developmental arrest. Furthermore, the timing of this developmental stage is sharp, limiting potential developmental disparities between animals20. We have previously shown that PARG-YFP effectively rescues the developmental arrest observed in parg27.1 null mutants13 and exhibits, on average, a 60% higher expression level compared to endogenous Parg (Supplemental Fig. S1). However, this difference is not statistically significant when considering the inherent variability within the populations. Furthermore, PARG-YFP does not display any discernible phenotypic changes21, suggesting that PARG-YFP functions similarly to endogenous PARG.
We identified that PARG binds to 4639 genes (Supplemental Table S1). PARG is mainly enriched in the gene body and is almost depleted to the promoter region, except at 30 bp, upstream of the transcription start site (TSS) (Fig. 1A) (Supplemental Fig. S2). To confirm PARG binding, we performed ChIP-qPCR on kek1 locus, one of the top peaks (Supplemental Fig. S3). Since PARG protein does not have a DNA-binding domain (DBD), our central hypothesis was that PARG is recruited to chromatin via PARP1. Accordingly, we expected PARG and PARP1 chromatin binding profiles to be similar. Interestingly, however, we found that only 57.1% (2649) of PARG binding loci are shared with PARP1 (Supplemental Fig. S4) (Supplemental Table S1). Moreover, we observed that PARG is more enriched to the promoter region of PARG/PARP1 common targets compared to PARG-alone targets (Fig. 1B). In contrast, PARP1 distribution is similar between common and PARP1- alone targets (Fig. 1C). In parallel, we analyzed the distance between PARG peaks and the promoter. Results revealed that PARG binds significantly closer to the promoter of common targets, while PARP1 peaks show no significant differences (Fig. 1D). Collectively, these results show that PARG binds to chromatin and that its binding profile is affected by the presence of PARP1.
PARG binds to chromatin. (A) Graph representing the average distribution of PARG along the genes. On the X-axis, the region of all target genes is scaled to a 2 kb region (from TSS to TES) and includes the 1 kb region upstream from TSS and downstream from TES. On the Y-axis, the PARG signal is normalized to the lower (0%) and the higher value (100%) to better visualize the difference in distribution. This graph was generated using all the genes bound by PARG (PARG and PARG/PARP1 categories of Supplemental Table S1). (B) Graph representing the average distribution of PARG along the PARG/PARP1 common target genes (blue) or PARG-alone target genes (orange) (respectively PARG/PARP1 and PARG categories of Supplemental Table S1). (C) Graph representing the average distribution of PARP1 along the PARG/PARP1 common target genes (blue) or PARP1- alone target genes (orange) (respectively PARG/PARP1 and PARP1 categories of Supplemental Table S1). (D) Distribution of the distance of PARG (left) and PARP1 (right) peaks to promoter in log2 scales for the promoter of PARG/PARP1 common targets (blue) and PARG- or PARP1-alone targets (orange) (respectively PARG/PARP1 and PARG or PARP1 categories of Supplemental Table S1). The significance of the difference between the two populations was addressed with a Mann–Whitney U test. ***: p-value < 0.01. N.S: Non-significant.
PARG presents two distinct binding profiles
Next, we investigated the transcription profile of PARG and PARP1 targets during late third instar larvae. We found that PARG/PARP1 common and PARP1-alone targets are mainly highly expressed genes, while PARG-alone targets are mainly silent genes (Fig. 2A). Furthermore, the expression of PARG-alone targets is, on average, significantly lower, while PARG/PARP1 common and PARP1-alone target expression is higher compared to other genes (Fig. 2B). Then, we analyzed the chromatin accessibility of PARP1 and PARG targets using the publicly available third instar larvae Assay for Transposase-Accessible Chromatin (ATAC) data22. This analysis revealed that PARG/PARP1 common and PARP1-alone target loci are more accessible than PARG- alone target loci (Fig. 2C). Moreover, the occupancy of active chromatin marks (H3K4me3 and H3K27Ac), but also PolII occupancy, is higher in PARG/PARP1 common and PARP1-alone targets while the distribution pattern of these markers is not affected (Fig. 2D). Specifically, at the promoters of common and PARP1-alone targets, the occupancy of PolII and H3K27Ac was found to be five times higher than that observed at PARG-alone target promoters. Similarly, at the promoters of PARP1-alone targets and common targets, H3K4me3 occupancy was five and three times higher, respectively, than at PARG-alone target promoters. In contrast, the occupancy of repressive chromatin marks (H3K27me3, H3K9me3 and H3K9me2) is higher for PARG-alone targets while the distribution pattern of these markers is not affected (Fig. 2E). Specifically, H3K9me3 was observed at the promoters of PARG-alone targets but absent at the promoters of common and PARP1-alone targets. Moreover, the gene bodies of PARG-alone targets exhibited a 43% increase in H3K9me2 occupancy compared to common and PARP1-alone targets. Furthermore, H3K27me3 occupancy was found to be five times higher at the gene bodies of PARG-alone targets and four times higher at the gene bodies of common targets compared to PARP1-alone targets. Overall, PARG binding profile correlates more with active marks for common targets (Fig. 2F), while it correlates more with repressive marks for PARG-alone targets (Fig. 2F). Taken together, these results suggest that PARG binds to highly active genes when PARP1 is present, and to silent genes when alone.
The presence of PARP1 affects PARG-binding profile. (A) Expression level of PARG-alone, PARP1-alone, and PARG/PARP1 common target genes. Genes were sorted based on their expression level during late third instar larvae (larval puffstage 7-9)23 and split into four quartiles, each including 25% of the genes. From the lowest to the highest expression, the quartiles were named as follows: Silent, Low, Moderate and High. The Y-axis represents the number of genes in percentage that belong to each group for PARG/PARP1 common targets (left), PARG-alone (middle), and PARP1-alone targets (right) (respectively PARG/PARP1, PARG and PARP1 categories of Supplemental Table S1). Error bars are standard error of the proportion and statistical test was Fisher exact test, PARG alone and PARP1 alone were compared to Common. ***: p-value < 0.01, N.S.: Non-significant. (B) Box plot representing the distribution of the log2 expression of PARG/PARP1 common targets (orange), PARG-alone (gray), PARP1- alone (yellow), and all other gene targets (gray) during late third instar larvae23. The difference among populations was analyzed with a Mann–Whitney U test. ***: p-value < 0.01. (C) Third instar larvae ATAC-seq signal22 to quantify chromatin accessibility for PARG/PARP1 common targets (blue), PARP1-alone (orange) and PARG-alone targets (gray). (D) Occupancy of polymerase II (Pol II) (left) and occupancy of the active marks H3K27Ac (middle) and H3K4me3 (right) to PARG/PARP1 common targets (blue), PARP1-alone (orange), or PARG-alone targets (gray). (E) Occupancy of the repressive marks H3K9me3 (left), H3K9me2 (middle), and H3K27me3 (right) to PARG/PARP1 common targets (blue) or PARP1-alone (orange), or PARG-alone targets (gray). (F) Spearman correlation of genome-wide signal for PARG/PARP1 common targets (left) or PARG-alone (right) targets (respectively PARG/PARP1 and PARG categories of Supplemental Table S1). (D, E) See the “Methods” section for information about the datasets that were used.
PARG regulates the expression of its target genes
We previously reported that PARG controls the transcription of developmental genes during third instar larvae, mainly as a repressor of transcription14. We then investigated if the expression of PARG/PARP1 common and PARG-alone targets is affected differently in the absence of PARG. First, we found that PARG binds to the loci of 68.3% of the genes significantly misregulated in parg mutant. Conversely, only 62.8% of the genes bound by PARG are misregulated in parg mutant. However, PARG presents a higher occupancy to the loci of genes that are significantly misregulated in parg mutant compared to other genes (Fig. 3A). Interestingly, PARP1 presents a higher occupancy to genes significantly downregulated in the absence of PARG (Fig. 3B). Furthermore, PARG/PARP1 common targets are significantly downregulated in parg mutant, while PARG-alone targets are significantly upregulated (Fig. 3C). Surprisingly, 72.1% of PARG-alone targets are also upregulated in parp1 mutant while PARP1 does not bind to them. This suggests that PARG promotes gene expression when PARP1 is present and participates in repression of expression when alone.
PARG regulates the expression of its target genes. (A, B) Average distribution of PARG (A) or PARP1 (B) along the genes upregulated (red), downregulated (blue) or those not misregulated (gray) in parg mutant. On the X-axis, the region of all target genes is scaled to a 2 kb region (from TSS to TES) and includes the 1 kb region upstream from TSS and downstream from TES. On the Y-axis, the distribution of PARG is normalized to the lower (0%) and the higher value (100%) to better visualize the difference in distribution. (C) Box plot representing the distribution of the log2 fold change of PARG/PARP1 common targets (blue), PARG-alone (orange), and all other gene targets (gray) during late third instar larvae in parg mutant compared to wild-type larvae. Difference among populations was analyzed with a with a Mann–Whitney U test. *: p-value < 0.05. ***: p-value < 0.01. (D) Average time course expression of PARG/PARP1 common (blue) and PARG-alone (orange) target genes from embryo to adult. Average expression is normalized from 0% (lowest expression) to 100% (highest expression). Error bar is standard error of the mean (SEM). Black arrow indicates the studied stage. L1, First instar larvae; L2, Second instar larvae; L3, Third instar larvae; h, Hours; PS, puff stage; PP, Prepupa; d, Days. Time course expression data were obtained from23. (E, F) Heatmap representing the time course expression of genes coding for a transcription factor bound by both PARG and PARP1 (E) and all other genes coding for a transcription factor (F) from first instar larvae to 30-day-old adult flies. Normalized expressions are shown as row z-scores. L1, First instar larvae; L2, Second instar larvae; L3, Third instar larvae; hr, Hours; PS, puff stage; PP, Prepupa; d, Days; Am, Adult males; Af, Adult females. Time course expression data were obtained from23.
PARG/PARP1 common targets are developmental genes upregulated during pupal stage
Next, we analyzed the time course expression of PARG/PARP1 common target genes in wild-type animals23. Overall, PARG/PARP1 common targets are genes that present a high expression during late instar larval stages, with a peak of expression during pupal stage (Fig. 3D, Supplemental Fig. S5A) with around 60% of PARG/PARP1 common targets upregulated during the transition from early to late third instar larvae (Supplemental Fig. 5B). In contrast, PARP1-alone target genes present high expression along all larval stages (Supplemental Fig. 5C). Gene ontology analysis reveals that PARG/PARP1 common target genes are involved in development, morphogenesis, cell differentiation and metamorphosis (Supplemental Fig. S6). A specific subset of 134 genes among the PARG/PARP1 common targets are specifically implicated in the process of metamorphosis (Supplemental Table S2). These genes exhibit a notable increase in expression prior to the pupal transition while their expression is reduced in both parg and parp1 mutants (Supplemental Table S2). Furthermore, among PARG/PARP1 common targets, we identified 196 genes coding for transcription factors (Supplemental Table S3). These transcription factors are mainly expressed during the end of third instar larvae and pupal stage (Fig. 3E, F, Supplemental Fig. 5D). Among them, we identified several transcription factors involved in ecdysone-response, including br, eip74ef, eip75b, eip78c, hr3, hr38, and ftz-f1, but also the gene coding for ecdysone receptor (ecr) that plays a critical role during metamorphosis24,25. Collectively, our findings underscore the collaborative role of PARG and PARP1 in promoting the expression of critical developmental genes that play a pivotal role in metamorphosis.
PARG binds in the absence of PARP1 to the loci of metabolic genes and of cuticle components that are downregulated before pupal stage
Finally, we analyzed the time course expression of PARG-alone target genes in wild-type animals23. Overall, PARG-alone targets are genes silenced during late third instar larvae and start their expression again during late pupa (Fig. 3D, Supplemental Fig. S5B). We found that only 20% of PARG-alone targets are upregulated during the transition from early to late third instar larvae (Fig. 3D). PARG-alone targets are mainly involved in metabolism and cuticle formation (Supplemental Fig. S7). We identified 39 genes coding for cuticle constituents that are bound by either PARG or PARP1, with 27 being exclusively bound by PARG. Strikingly, all these PARG-bound genes exhibit a significant downregulation during the transition from larva to pupa and display a marked increase in expression in parg mutant individuals (Supplemental Table S4). Interestingly, we identified 12 genes involved in cuticle formation that are co-bound by both PARP1 and PARG. These genes collectively exhibit an increase of their expression levels before the larva-to-pupa transition, while they experience a significant downregulation in parg and parp1 mutant animals (Supplemental Table S4). Given the well-established change in cuticle composition during the larva-to-pupa transition26, our findings suggest a regulatory role of PARG and PARP1 in orchestrating this transformation.
Moreover, our study identifies PARG as a key regulator of genes implicated in various metabolic functions (Supplemental Fig. S7). More precisely, we found that PARG binds to the loci of genes participating in the digestive processes (Supplemental Table S5) (and based on27). Strikingly, these genes exhibit a reduction in expression prior to the transition to pupa, while their expression is substantially upregulated in parg mutant, has we previously reported14. Simultaneously, our investigation unveiled PARG's independent binding to the loci of metabolic genes governing fatty acid biosynthesis, glycogen metabolism, and amino acid metabolism (Supplemental Table S6). Notably, all these metabolic genes display a downregulation before the transition to pupa, and a notable upregulation in parg mutant individuals. Intriguingly, we also identified four genes involved in amino acid metabolism, which are bound by both PARP1 and PARG. These genes are collectively upregulated before transition to pupa while they are downregulated in parg and parp1 mutants. In summary, our results collectively demonstrate PARG’s role in repressing the expression of metabolic genes prior to the transition to the pupal stage, shedding light on its regulatory impact in these critical developmental processes.
Discussion
As expected, we found PARP1 to be enriched at the promoter of highly active genes. Interestingly, we identified that PARG/PARP1 common targets are genes that increase their expression rapidly at the end of third instar larvae, while PARP1-alone target genes present a high expression level along all larval stages (Supplemental Fig. S5B, C). This result suggests that the presence of both PARG and PARP1 is required to increase gene expression rapidly, while only the presence of PARP1 is required to maintain a constant expression level.
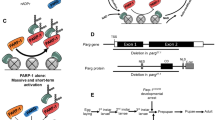
PARP1 has been shown to directly regulate gene expression as an activator of transcription28,29,30,31,32, but sometimes as a repressor18,33,34. PARG activity is essential for PARP1 function since PARP1 mainly targets itself18. Accordingly, PARG was expected to participate in PARP1-mediated transcriptional regulation, at least by allowing PARP1 to return to chromatin. Despite the absence of a DBD, we showed in this study that PARG binds to chromatin, confirming the in vitro results of a previous study on human cells35. Remarkably, we identified two distinct PARG-binding profiles. First, PARG, together with PARP1, binds to the promoter region of genes involved in metamorphosis to promote their expression (Fig. 4A). Second, PARG binds to the gene body of genes involved in metabolism and cuticle formation to repress their expression before the transition from larval to pupal stage (Fig. 4B). Notably, it is crucial to highlight that the observed downregulation of these metabolic genes before the transition to the pupal stage does not entail complete silencing, signifying a nuanced regulatory mechanism at play. Indeed, metamorphosis is well-documented to orchestrate a U-shaped pattern in global metabolism dynamics, as corroborated by previous studies36,37,38,39. Specifically, metabolism rate is downregulated preceding the transition to the pupal stage and is upregulated during the middle of pupal stage.
PARG exhibits two distinct mechanisms of action at chromatin. (A) PARG binds, together with PARP1, to the promoter region of target genes to promote their expression. PARG/PARP1 common targets are mainly genes upregulated during larval/pupal transition, including transcription factors mainly expressed at the end of the larval stage and during pupal stage. (B) PARG binds, without PARP1, to the gene body of its target genes to repress their expression. PARG-alone targets are mainly genes involved in metabolism and larval cuticle formation; two processes downregulated before larval/pupal transition.
One possibility is that parg mutant animals experience subtle developmental delays during the wandering stage, contributing to the observed difference between mutant and control larvae. However, this possibility is challenged by the binding of PARG to these metabolic loci, suggesting a more direct involvement of PARG in the regulation of the expression of these metabolic genes. Our findings shed light on the distinctive role of PARG, particularly in the absence of PARP1, as a key contributor to the downregulation of metabolic genes prior to pupation, simultaneously, PARG’s involvement in orchestrating changes in cuticle formation underscores its multifaceted role in shaping developmental processes. Collectively, our results offer compelling insights into the potential global transcriptional regulatory roles of both PARP1 and PARG in the context of metamorphosis, further expanding our understanding of their multifunctional contributions to this intricate biological phenomenon.
Our prior research revealed that the absence of either PARP1 or PARG leads to the upregulation of genes that are expected to be downregulated before pupation14. In this study, our analysis reveals that 72.1% of PARG-alone target genes exhibit upregulation in the absence of PARP1, despite the absence of direct binding by PARP1 to these genes. This outcome underscores the synergistic interplay between PARG and PARP1; the absence of one affects the function of the other12,35. Therefore, these genes are not directly impacted by the absence of PARP1 but by a defect in PARG function caused by the absence of PARP1. Interestingly, we found that PARG binds the loci of 4639 genes in total. This number is coherent with the range of binding sites recorded for other transcription factors that can range from 1 binding site to more than 9000 in Drosophila40. However, it is worth noting that around 30% of loci bound by PARG are not misregulated in parg mutant at the analyzed time-point. Collectively, the integration of our previous RNA-seq data with the current ChIP-seq results provides a more comprehensive understanding of the roles played by PARP1 and PARG in gene expression regulation before the transition from larva to pupa.
In the absence of a DNA-binding domain, it is plausible that PARG operates as part of more extensive protein complexes. When considering the shared targets of PARG and PARP1, it is reasonable to postulate that PARP1 takes on the role of recruiting PARG to the promoters of these loci. In this context, PARG effectively serves as a cofactor for PARP1. Additionally, our data concerning genes regulated by PARG indicate that PARG predominantly associates with the gene bodies rather than their promoters. This pattern of binding aligns more closely with the anticipated behavior of chromatin-associated factors.
In this study we showed that PARP1 and PARG play critical roles in the transition from larva to pupa, roles consistent with the death of both parp1 hypomorph and parg null mutants before metamorphosis11,18. However, the expression of PARG-alone and PARG/PARP1 common targets is dynamic throughout development (Fig. 3D), not only before metamorphosis. Furthermore, we previously reported that affecting PARP1 or PARG maternal contribution leads to lethality during embryonic stage7,19, suggesting that PARG and PARP1 might control transcriptional changes before metamorphosis and earlier in development. We found that PARG also exhibits a chromatin-binding pattern that is PARP1-independent. PARG binds to the gene body of genes involved in cuticle formation and metabolism to repress their expression before metamorphosis. However, since PARG lacks a DBD, it is not clear how PARG is recruited to chromatin in the absence of PARP1.
The major actor involved in metamorphosis is the ecdysone pathway24,25. We previously reported that 20-hydroxyecdysone, an effector in the ecdysone pathway41, can activate PARP1 activity12. Furthermore, we previously showed that the expression of genes coding for enzymes involved in the synthesis of 20-hydroxyecdysone are not affected during L3 larval puff stage 7-9 in parg and parp1 mutant14, suggesting that PARP1 acts downstream to ecdysone rather than upstream. Additionally, the expression of glue genes, which are located within puffs of polytene chromosomes and belong to the early response to ecdysone pathway that starts during L3 larval puff stage 7-942, are still initiated in both parg and parp1 mutants, suggesting that PARP1 does not affect ecdysone early-response genes. Interestingly, we found that PARG and PARP1 bind together to the promoter region of the transcription factors involved in ecdysone mid-response (Supplemental Table S1). Their expression is not significantly affected in parg and parp1 mutants, but most are downregulated14. Together, these results suggest that PARP1 and PARG might play a role in the ecdysone mid-response, and that a defect in the ecdysone pathway could be the cause of the developmental arrest that exhibits both parg and parp1 mutant animals. However, the broad binding of PARP1 and PARG to loci involved in the metamorphosis, infers that the function of these dual proteins is not limited to the transition from ecdysone early to mid-response but is more global in the promotion of transcription.
In conclusion, our study reveals that poly(ADP-ribosylating) enzymes directly orchestrate transcriptional changes that start before metamorphosis by promoting the expression of genes required for pupation while in parallel regulating negatively the expression of metabolic genes before the transition to pupal stage.
Methods
Drosophila strains and genetics
Flies were raised at 20 °C. The PARG-YFP line used in ChIP-seq corresponds to parg27.1/parg27.1; pP{w1, UAST::PARG-EYFP}, 69B-GAL4/TM2, as described in19. The parg27.1 mutant was described in11, P{w1, UAST::PARG-EYFP} was described in43, and 69B-GAL4 driver was described in7. The expression of PARG-EYFP rescues the lethality of parg mutant. The PARP1-YFP line used in ChIP-seq was P{w1, UAST::PARP1-EYFP}, P{GAL4}Mz1087.hx ; parp1C03256/parp1C03256. UAST-PARP1-EYFP was described in32 and Mz1087.hx GAL4 driver was described in44. The expression of PARP1-EYFP rescues the lethality of parp1 mutant. A yellow white strain carrying the mutations y1, w1118 was used as a YFP-negative control line.
Quantitative RT-PCR assay
This assay was performed in triplicate. Twelve wandering third instar larvae were collected for control yellow white and PARG-YFP flies. Total RNA was extracted from cells using the QIAshredder column and RNeasy kit (Qiagen). Contaminating genomic DNA was removed by the g-column provided in the kit. cDNA was obtained by reverse transcription using M-NLV reverse transcriptase (Invitrogen). Real-time PCR assays were run using SYBR Green master mix (Bio-Rad) and an Applied Biosystems StepOnePlusTM instrument. The amount of DNA was normalized using the difference in threshold cycle (CT) values (ΔCT) between rpL32 and parg genes.
The quantitative real-time PCR (qPCR) primer sequences for Drosophila melanogaster ribosomal protein L32 gene (rpL32) were 5′-GCTAAGCTGTCGCAACAAAT-3′ (forward) and 5′-GAACTTCTTGAATCCGGTGGG-3′ (reverse).
Sequences for parg were 5′-AGAAACACCCTCAAGAGGAAG-3′ (forward) and 5′-CGCTCTGTGGGACACAC-3′ (reverse).
Chromatin immunoprecipitation assay
ChIP was performed with PARG-YFP and PARP1-YFP lines and with a yellow white line (control line) that does not express YFP as a negative control line. We performed ChIPseq experiment on YFP-tagged PARG due to the unavailability of antibodies capable of effectively binding to Drosophila PARG. This experiment was performed in duplicates for PARG-YFP (two biological replicates for PARP-YFP and two biological replicates for control lines) and in triplicates for PARP1-YFP. 75 wandering third instar larvae (L3 puff stage 7-9) were collected in a 2 ml DNA LoBind Eppendorf tube and washed twice with 1 ml 1× PBS. The larvae were homogenized in ice-cold lysis buffer (200 µl of 1× protease inhibitor cocktail, 250 µl of 100 mM PMSF, 800 µl of 1× PBS, and 1 µl of Tween 20) using a pellet pestle. The homogenized lysate was crosslinked using 244.5 µl of 11% formaldehyde to a final concentration of 1.8% for 15 min at room temperature on a rotator. 500 mM Glycine was added to quench the fixative on ice for 5 min. The larval debris was pelleted at 1,000 g for 3 min, and the supernatant was removed. The pellet was resuspended in 1 ml sonication buffer (0.5% SDS, 20 mM Tris pH 8.0, 2 mM EDTA, 0.5 mM EGTA, 0.5 mM PMSF, and 1× protease inhibitor cocktail), and chromatin was fragmented to 300–500 bp by Bioruptor sonicator (UCD-200) for 20 cycles (30 s high frequency sonication, 1.5 s pause) in a cold room. The sonicated material was pelleted at 10,000 g for 10 min at 4 °C, supernatant was collected, and then fragment size was checked prior to immunoprecipitation. The sonicated chromatin was pre-cleared and incubated with an anti-GFP polyclonal antibody (TP-401, Origene) at 4 °C overnight. TP401 recognizes YFP as well. The immunoprecipitated chromatin was then collected with prewashed Protein A agarose beads for 2 h. The beads were sequentially washed with the following buffers: 1 low-salt buffer wash (0.1% SDS, 1% Triton X-100, 2 mM EDTA, 20 mM Tris442 HCL pH 8.0, and 150 mM NaCl), 3 high-salt buffer washes (0.1% SDS, 1% Triton X-100, 2 mM EDTA, 20 mM Tris–HCL pH 8.0, and 500 mM NaCl), 1 LiCL wash (2 mM EDTA, 20 mM Tris–HCl pH 8.0, and 0.25 M LiCl, 1% NP-40), and 2 TE buffer washes before elution. Bound chromatin on beads was eluted twice at room temperature using 250 µl of freshly prepared ChIP elution buffer (1% SDS and 100 mM NaHCO3) for 15 min and reverse-crosslinked overnight at 65 °C. The eluates were then treated with RNase A and proteinase K prior to DNA extraction via phenol–chloroform extraction and ethanol precipitation. The library was prepared using NEBnext multiplex oligos for Illumina kit (Index primer set 4), according to the manufacturer’s protocol. Paired-end sequencing was performed by Novogene Corporation, Sacramento CA. Raw data are available here: (GSE228898).
ChIP-qPCR assay was repeated with three biological replicates per condition for ChIP-qPCR confirmation. To confirm that PARG binds to the promoter region of kek1 locus, we performed another set of ChIP assay on PARG-YFP and a yellow white line. This experiment was performed in triplicates. We designed two sets of primers (Supplemental Fig. S3), one set to amplify kek1 promoter region (positive region) and one set to amplify the beginning of kek1 gene body where PARG is not detected (negative region).
Primers used for kek1 positive region: 5′-GTTGCGTCGTTCCCGCTGTAGC-3′ (forward) and 5′-CGGTGTGTCCTGGCTAGCGGTAC-3′ (reverse).
Primers used for kek1 negative region: 5′-GGATCTGCATGTGGATGAGTTTGCC-3’ (forward) and 5′-GCTCGACATGTAATCGAGGCATTCTC-3′ (reverse).
Real-time PCR assays were run using SYBR Green master mix (Bio-Rad) volume and an Applied Biosystems StepOnePlus™ instrument. The amount of DNA in the immunoprecipitated fraction (IP) was compared to the Input fraction for both PARG-YFP and yellow white control lines.
Genome-wide datasets
In this study, we compared different publicly available ChIP-seq and ATAC-seq datasets. All were performed on whole Drosophila third instar larvae and included PARG-YFP ChIP-seq (this study), (GSE228898). parg mutant RNA-seq from our previous study14, GSE200499. PARP1-YFP ChIP-seq, (GSE217729). parp1 mutant microarray from our previous study45, ATAC-seq22, GSE96922. Pol II ChIP-seq46 (GSE15292). H3K4me3 ChIP-seq46, GSE49491. H3K27Ac ChIP-seq 46, GSE49488. H3K27me3 ChIP-seq46, GSE49490. H3K9me2 ChIP-seq46, GSE47260. H3K9me3 ChIP-seq46, and GSE47258. All these datasets were analyzed from raw data. In addition, we used the Developmental time-course RNA-seq dataset23, SRP001065.
ChIP-seq analysis
ChIP-seq data were analyzed with Galaxy47. The quality of raw reads was checked using FastQC (version. 0.11.9), and adapters were removed with fastp48. Trimmed raw reads were aligned to the Drosophila genome (dm6) using Bowtie249. Unmapped and low-quality reads were discarded (< = 20 mapQuality) using BamTools50. Duplicate reads were identified and removed from mapped reads using Picard MarkDuplicates (http://broadinstitute.github.io/picard). MACS251 was used to call peaks against control (Input or negative control, depending from the dataset) using default settings. Peaks were annotated to genomic features with ChIPseeker 52. Pairwise correlation of peaks was determined using Intervene53. MACS2 bedGraph pileups were used to generate normalized coverage of ChIP-seq signals using Deeptools bigWigCompare by computing the ratio of the signals (IP vs. Control/Input) using a 50 bp bin size. Deeptools multiBigwigSummary and plotCorrelation54 were used to determine genome-wide signal correlation using a 10 kb bin size. Deeptools computematrix Deeptools plotHeatmap54 was used with a 50 bp bin size to create enrichment profiles around peak centers (± 1 kb) in reference mode, or in a 2 kb scale region mode from transcription start site (TSS) to transcription end site (TES) to create enrichment profiles along the genes. Scale region mode also included the 1 kb flanking regions before TSS and after TES.
Gene ontology (GO) terms were determined using g:profiler55. The list of Drosophila transcription factors was obtained from Flybase56 (Gene Ontology “DNA-binding transcription factor activity”). Time-course heatmap was generated with heamap2 after computing z-scores and performing an Euclidian clustering. Gene set of a specific GO-term were retrieved from AmiGO database57,58.
Data availability
Mutant strains and transgenic stocks are available upon request. The authors state that all data necessary to confirm the conclusions presented in the article are represented fully within the article. PARG ChIP-seq raw and processed data are accessible upon demand or on GEO platform: GSE228898. PARP1 ChIP-seq raw and processed data are accessible upon demand or on GEO platform: GSE217729. The PARG ChIP-seq data can be accessed with the accession number GSE228898. Similarly, the PARP1 ChIP-seq data can be accessed with the accession number GSE217729.
References
Dillman, A. A. & Cookson, M. R. Transcriptomic changes in brain development. Int. Rev. Neurobiol. 116, 233–250 (2014).
Ohler, U. & Wassarman, D. A. Promoting developmental transcription. Development 137(1), 15–26 (2010).
Miyazawa, H. & Aulehla, A. Revisiting the role of metabolism during development. Development 145, 19 (2018).
Sharpley, M. S., Chi, F., Hoeve, J. T. & Banerjee, U. Metabolic plasticity drives development during mammalian embryogenesis. Dev. Cell. 56(16), 2329–2347 (2021).
Diaz-Cuadros, M. et al. Metabolic regulation of species-specific developmental rates. Nature 613(7944), 550–557 (2023).
Menissier-de-Murcia, J. et al. Functional interaction between PARP-1 and PARP-2 in chromosome stability and embryonic development in mouse. EMBO J. 22(9), 2255–2263 (2003).
Tulin, A., Stewart, D. & Spradling, A. C. The Drosophila heterochromatic gene encoding poly(ADP-ribose) polymerase (PARP) is required to modulate chromatin structure during development. Genes Dev. 16(16), 2108–2119 (2002).
D’Amours, D., Desnoyers, S., D’Silva, I. & Poirier, G. G. Poly(ADP-ribosyl)ation reactions in the regulation of nuclear functions. Biochem. J. 342(Pt 2), 249–268 (1999).
Liu, C. & Yu, X. ADP-ribosyltransferases and poly ADP-ribosylation. Curr. Protein Pept. Sci. 16(6), 491–501 (2015).
Tulin, A. & Spradling, A. Chromatin loosening by poly(ADP)-ribose polymerase (PARP) at Drosophila puff loci. Science 299(5606), 560–562 (2003).
Hanai, S. et al. Loss of poly(ADP-ribose) glycohydrolase causes progressive neurodegeneration in Drosophila melanogaster. Proc. Natl. Acad. Sci. U. S. A. 101(1), 82–86 (2004).
Kotova, E., Jarnik, M. & Tulin, A. V. Poly (ADP-ribose) polymerase 1 is required for protein localization to Cajal body. PLoS Genet. 5(2), e1000387 (2009).
Tulin, A., Naumova, N. M., Menon, A. K. & Spradling, A. C. Drosophila poly(ADP-ribose) glycohydrolase mediates chromatin structure and SIR2-dependent silencing. Genetics 172(1), 363–371 (2006).
Bordet, G., Karpova, I. & Tulin, A. V. Poly(ADP-ribosyl)ating enzymes cooperate to coordinate development. Sci. Rep. 12(1), 22120 (2022).
Gibson, B. A. & Kraus, W. L. New insights into the molecular and cellular functions of poly(ADP-ribose) and PARPs. Nat. Rev. Mol. Cell Biol. 13(7), 411–424 (2012).
Harrision, D., Gravells, P., Thompson, R. & Bryant, H. E. Poly(ADP-Ribose) Glycohydrolase (PARG) vs Poly(ADP-Ribose) Polymerase (PARP)—function in genome maintenance and relevance of inhibitors for anti-cancer therapy. Front. Mol. Biosci. 7, 191 (2020).
Kraus, W. L. & Lis, J. T. PARP goes transcription. Cell 113(6), 677–683 (2003).
Kotova, E., Jarnik, M. & Tulin, A. V. Uncoupling of the transactivation and transrepression functions of PARP1 protein. Proc. Natl. Acad. Sci. U. S. A. 107(14), 6406–6411 (2010).
Bordet, G., Kotova, E. & Tulin, A. V. Poly(ADP-ribosyl)ating pathway regulates development from stem cell niche to longevity control. Life Sci. Alliance 5, 3 (2022).
Wegman, L. J., Ainsley, J. A. & Johnson, W. A. Developmental timing of a sensory-mediated larval surfacing behavior correlates with cessation of feeding and determination of final adult size. Dev. Biol. 345(2), 170–179 (2010).
Pinnola, A., Naumova, N., Shah, M. & Tulin, A. V. Nucleosomal core histones mediate dynamic regulation of poly(ADP-ribose) polymerase 1 protein binding to chromatin and induction of its enzymatic activity. J. Biol. Chem. 282(44), 32511–32519 (2007).
Meers, M. P. et al. Transcription start site profiling uncovers divergent transcription and enhancer-associated RNAs in Drosophila melanogaster. BMC Genom. 19(1), 157 (2018).
Graveley, B. R. et al. The developmental transcriptome of Drosophila melanogaster. Nature 471(7339), 473–479 (2011).
Ou, Q. & King-Jones, K. What goes up must come down: Transcription factors have their say in making ecdysone pulses. Curr. Top. Dev. Biol. 103, 35–71 (2013).
Yamanaka, N., Rewitz, K. F. & O’Connor, M. B. Ecdysone control of developmental transitions: Lessons from Drosophila research. Annu. Rev. Entomol. 58, 497–516 (2013).
Ali, M. S. & Takaki, K. Transcriptional regulation of cuticular genes during insect metamorphosis. Front. Biosci. (Landm. Ed.) 25(1), 106–117 (2020).
Lemaitre, B. & Miguel-Aliaga, I. The digestive tract of Drosophila melanogaster. Annu. Rev. Genet. 47, 377–404 (2013).
Petesch, S. J. & Lis, J. T. Activator-induced spread of poly(ADP-ribose) polymerase promotes nucleosome loss at Hsp70. Mol. Cell. 45(1), 64–74 (2012).
Weaver, A. N. & Yang, E. S. Beyond DNA repair: Additional functions of PARP-1 in cancer. Front. Oncol. 3, 290 (2013).
Lodhi, N., Kossenkov, A. V. & Tulin, A. V. Bookmarking promoters in mitotic chromatin: Poly(ADP-ribose)polymerase-1 as an epigenetic mark. Nucleic Acids Res. 42(11), 7028–7038 (2014).
Thomas, C. J. et al. Kinase-mediated changes in nucleosome conformation trigger chromatin decondensation via poly(ADP-ribosyl)ation. Mol. Cell 53(5), 831–842 (2014).
Thomas, C. et al. Hit and run versus long-term activation of PARP-1 by its different domains fine-tunes nuclear processes. Proc. Natl. Acad. Sci. U. S. A. 116(20), 9941–9946 (2019).
Ding, L. et al. PARP1 suppresses the transcription of PD-L1 by Poly(ADP-Ribosyl)ating STAT3. Cancer Immunol. Res. 7(1), 136–149 (2019).
Tian, Y. N., Chen, H. D., Tian, C. Q., Wang, Y. Q. & Miao, Z. H. Polymerase independent repression of FoxO1 transcription by sequence-specific PARP1 binding to FoxO1 promoter. Cell Death Dis. 11(1), 71 (2020).
Frizzell, K. M. et al. Global analysis of transcriptional regulation by poly(ADP-ribose) polymerase-1 and poly(ADP-ribose) glycohydrolase in MCF-7 human breast cancer cells. J. Biol. Chem. 284(49), 33926–33938 (2009).
Arbeitman, M. N. et al. Gene expression during the life cycle of Drosophila melanogaster. Science 297(5590), 2270–2275 (2002).
Merkey, A. B., Wong, C. K., Hoshizaki, D. K. & Gibbs, A. G. Energetics of metamorphosis in Drosophila melanogaster. J. Insect. Physiol. 57(10), 1437–1445 (2011).
Nishimura, T. Feedforward regulation of glucose metabolism by steroid hormones drives a developmental transition in drosophila. Curr. Biol. 30(18), 3624–3632 (2020).
Farkas, R. & Slama, K. Respiratory metabolism of salivary glands during the late larval and prepupal development of Drosophila melanogaster. J. Insect. Physiol. 81, 109–117 (2015).
MacArthur, S. et al. Developmental roles of 21 Drosophila transcription factors are determined by quantitative differences in binding to an overlapping set of thousands of genomic regions. Genome Biol. 10(7), R80 (2009).
Nakagawa, Y. & Sonobe, H. Subchapter 98A—20-Hydroxyecdysone. In (eds. Takei, Y. et al.) Handbook of Hormones 560-e98A-2 (Academic Press, 2016).
Lehmann, M. Drosophila Sgs genes: Stage and tissue specificity of hormone responsiveness. Bioessays 18(1), 47–54 (1996).
Ji, Y. & Tulin, A. V. Poly(ADP-ribose) controls DE-cadherin-dependent stem cell maintenance and oocyte localization. Nat. Commun. 3, 760 (2012).
Fossgreen, A. et al. Transgenic Drosophila expressing human amyloid precursor protein show gamma-secretase activity and a blistered-wing phenotype. Proc. Natl. Acad. Sci. U. S. A. 95(23), 13703–13708 (1998).
Bordet, G., Lodhi, N., Guo, D., Kossenkov, A. & Tulin, A. V. Poly(ADP-ribose) polymerase 1 in genome-wide expression control in Drosophila. Sci. Rep. 10(1), 21151 (2020).
Consortium EP. An integrated encyclopedia of DNA elements in the human genome. Nature 489(7414), 57–74 (2012).
Afgan, E. et al. The Galaxy platform for accessible, reproducible and collaborative biomedical analyses: 2018 update. Nucleic Acids Res. 46(W1), W537–W544 (2018).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: An ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34(17), i884–i890 (2018).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9(4), 357–359 (2012).
Barnett, D. W., Garrison, E. K., Quinlan, A. R., Stromberg, M. P. & Marth, G. T. BamTools: A C++ API and toolkit for analyzing and managing BAM files. Bioinformatics 27(12), 1691–1692 (2011).
Feng, J., Liu, T., Qin, B., Zhang, Y. & Liu, X. S. Identifying ChIP-seq enrichment using MACS. Nat. Protoc. 7(9), 1728–1740 (2012).
Yu, G., Wang, L. G. & He, Q. Y. ChIPseeker: An R/Bioconductor package for ChIP peak annotation, comparison and visualization. Bioinformatics 31(14), 2382–2383 (2015).
Khan, A. & Mathelier, A. Intervene: A tool for intersection and visualization of multiple gene or genomic region sets. BMC Bioinform. 18(1), 287 (2017).
Ramirez, F. et al. deepTools2: A next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44(W1), W160–W165 (2016).
Raudvere, U. et al. g:Profiler: A web server for functional enrichment analysis and conversions of gene lists (2019 update). Nucleic Acids Res. 47(W1), W191–W198 (2019).
Thurmond, J. et al. FlyBase 2.0: The next generation. Nucleic Acids Res. 47(D1), D759–D765 (2019).
Ashburner, M., Golic, K. G. & Hawley, R. S. Drosophila: A Laboratory Handbook (Cold Spring Harbor Laboratory Press, 2004).
Gene Ontology, C. et al. The Gene Ontology knowledgebase in 2023. Genetics 224, 1 (2023).
Funding
Funding for this project was supported by the National Science Foundation MCB-2231403 and Department of Defense grant PC160049 to A.V.T. Funding for this project was supported by the Doctoral Research Grant to Gbolahan Bamgbose from the Doctoral Incentive Program Fund through the School of Graduate Studies of the University of North Dakota. Funding agencies had no role in study design, data collection, data analysis, interpretation, or writing of the report.
Author information
Authors and Affiliations
Contributions
The conceptualization and investigation of this work was performed by G.B., G.B. and A.V.T. This work was supervised byA.V.T. This manuscript was written by G.B. and A.V.T.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bordet, G., Bamgbose, G. & Tulin, A.V. Poly(ADP-ribosyl)ating enzymes coordinate changes in the expression of metabolic genes with developmental progression. Sci Rep 13, 20320 (2023). https://doi.org/10.1038/s41598-023-47691-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-023-47691-8