Abstract
Selective Serotonin Reuptake Inhibitor (SSRI) therapy is common among perinatal populations for the treatment of mood disorders. Medications can affect diversity and composition of the gut microbiome, which plays a key role in modulating health. While previous studies have examined the effects of antidepressant exposure on the maternal gut microbiome, whether SSRI exposure affects the offspring gut microbiome is unknown. We investigated the effects of maternal fluoxetine exposure on the gut microbiome of maternal and offspring mice during pregnancy and lactation (embryonic day 10–lactation day 21; E10–L21). Stool samples collected on E17, L11, L15, and L21 were examined using 16S rRNA sequencing. Our results suggest that maternal fluoxetine exposure may result in decreased alpha diversity of the offspring gut microbiome in early life. Furthermore, we observed several genera-specific differences in the gut microbiome based on treatment, specifically of Turicibacter, Parasutterella, and Romboutsia. These findings support our understanding of gut health, as dysbiotic development of the gut microbiome has been associated with local and systemic health problems including gastrointestinal morbidities and interrupted growth patterns in infants. Future research should pursue study in human populations and those at high risk for gut microbial dysbiosis and intestinal injury.
Similar content being viewed by others
Introduction
Perinatal mental health disorders occurring during pregnancy and up to 12 months following delivery affect up to 20% of the population and represent one of the most common complications of pregnancy and postpartum1. Those with mental health disorders pre-gestation are at particularly high risk for experiencing a disorder in the pregnancy or postpartum period2. The burden and cost to the U.S. heathcare system is immense, with recent estimates suggesting that mental health disorders increase severe maternal morbidity by up to 50%, with delivery-related hospitalization costs in excess of $102 million dollars annually3,4. Recently, the Joint Commission declared that the United States is facing a maternal health crisis and listed maternal mental health conditions as a leading cause of pregnancy-related death5.
Approximately 5–13% of mothers receive treatment with a common class of antidepressants, selective serotonin reuptake inhibitors (SSRIs). Estimates suggest that the highest rates of SSRI exposure worldwide occur in the United States6,7,8. Although many medications exist within the SSRI class, sertraline (Zoloft®), citalopram (Celexa®) and fluoxetine (Prozac®) are among the most frequently prescribed SSRIs in perinatal populations9. Like many other medications, SSRIs can cross the blood-placental and blood-milk barrier to reach both fetal circulation and human milk10. The decision to initiate or continue treatment with SSRIs during pregnancy requires consideration of risk to benefit and can be complicated by the sparsity of research on how perinatal SSRI exposure might affect short and long-term health outcomes.
Antidepressants of the SSRI class are thought to act by manipulating serotonin signaling in the central nervous system, although oral administration may result in greater systemic effects11. In fact, SSRIs have antimicrobial properties which have sparked growing interest in the relationship between antidepressant treatment, gut serotonin bioavailability and the gut microbiome12,13. Disruptions to serotonin homeostasis through antidepressant exposure have been shown to affect the fitness of the resident gut microorganisms14,15. In turn, members of the gut microbiome can modulate peripheral serotonin production, likely through the secretion of metabolites14. The competing influences between antidepressant exposure, gut serotonin dynamics and microbial activity illustrate the critical yet complex role of serotonin in the gut microbial ecosystem and raises further questions about the the gut microbiome’s role in mental health.
While accumulating evidence links depression and SSRI antidepressant exposure to gut microbial composition, the specific nature of influence across populations may vary16,17. Research conducted with adult nulliparous participants affected by major depressive disorder suggests that antidepressant exposure can decrease the Firmicutes/Bacteriodetes ratio and negatively impact alpha diversity18. At higher resolution, reductions in the Turicibacter genre have also been described19. Despite the burden of maternal mental health conditions during perinatality, less is known about the effects of maternal antidepressant exposure on the gut microbiome in mothers and offspring. Study of perinatal rodent models suggest SSRI exposure could potentially decrease maternal populations of Bacteroides while increasing Lachnospiraceae, Lachnoclostridium and Blautia20,21. Consideration of antidepressant effects on the gut microbiome of perinatal populations is crucial, as it’s possible that effects may be unique or potentially compounded by gut microbiome plasticity or transitions that occur during pregnancy and lactation20,21.
Although our understanding surrounding maternal antidepressant exposure and the gut microbiome is rapidly expanding, whether maternal antidepressant exposure affects early infant gut microbiome diversity and composition has not been explored. The infant gut microbiome is shaped by a variety of social, cultural and environmental determiants and ultimately, may be related to both short and long-term health22,23,24. Here, we sought to address this important gap in our knowledge by examining the effect of fluoxetine (SSRI) exposure on the maternal and offspring gut microbiome in a rodent model.
Results
In cohort one, 20 female mice were ordered and 11/20 completed the model of pregnancy. Six were randomized to the fluoxetine (treatment) group and five were randomized to the saline (control) group. During the experimental period, a total of 53 stool samples were collected, sequenced, and used in the analysis (E10 = 9, E17 = 11, L11 (maternal) = 11, L11 (offspring) = 22). In cohort two, 12/40 mice completed the model of pregnancy. Five were randomized to the fluoxetine group and 7 were randomized to the saline group. A total of 79 samples were collected, sequenced and used in the analysis (E10 = 12, E17 = 12, L15 (maternal) = 12, L15 (offspring) = 10, L21 (maternal) = 12, L21 (offspring) = 21). Six negative control samples were sequenced during DNA extraction and sequencing procedures, for a grand total of 138 sequenced samples.
A total of 3,535,190 reads were sequenced with an average of 25,617+/− 64,612 reads per sample. After processing through QIIME II, our dataset included 1,742 taxa, with 13 taxa that were removed during the decontam procedure25. After removal of negative controls and samples with read counts below 5,000, a total of 114 samples remained. Of the 17 samples removed for low read counts, 4 were maternal (2 = E10, 1 = E17, 1 = L11) and 14 were offspring (11 = L11, 2 = L15, 1 = L21). Following rarefaction of the data, 331 ASVs were removed resulting in a final dataset of 1,389 taxa and 114 samples. One sample (offspring L15) was removed from the dataset as an outlier following alpha diversity analyses.
Alpha diversity
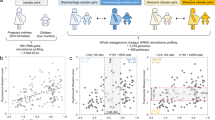
Maternal and offspring populations displayed distinct patterns in the progression of gut microbial alpha diversity over time. In maternal rodents that received either treatment (fluoxetine) or control (saline) exposures, alpha diversity appeared to trend upwards as gestation and lactation progressed. Upon comparison, there were no statistically significant differences in alpha diversity of the maternal gut microbiome in treatment and control mice. In contrast, the offspring of mothers treated with fluoxetine had lower alpha diversity values than the offspring of mothers treated with saline at L11 (p = 0.03), with increasing alpha diversity and similarity observed with time (Table 1, Fig. 1).
Maternal and offspring alpha diversity between treatment groups. Alpha Diversity was calculated with Shannon’s Alpha Diversity Index and visualized across time. Associated comparisons were calculated with Wilcoxon Signed Rank Test (Table 1). The offspring Alpha Diversity comparison at L11 is statistically significant with p = 0.03. E10 = embryonic day 10, E17 = embryonic day 17, L11 = lactation day 11, L15 = lactation day 15, L21 = lactation day 21.
Beta diversity
Measures of beta diversity were calculated for maternal and offspring populations and compared by treatment group using Weighted Unifrac distances, Unweighted Unifrac distances and the Bray–Curtis index. Beta diversity measures of maternal samples were similar across treatment groups. In contrast, Weighted Unifrac distance and the Bray–Curtis index revealed statistically significant differences between samples from offspring with maternal exposure to fluoxetine or the control group at L11 (p = 0.01; p = 0.02; Table 2, Figs. 2, S1). Differences in the Bray–Curtis and Weighted Unifrac measures between treatment groups of offspring at L11 suggest that the relative abundances of microorganisms between treatment groups may be responsible for the observed differences, rather than presence or absence of unique taxa.
Offspring beta diversity at lactation day 11 between treatment groups. Beta Diversity was calculated with Weighted Unifrac distance and visualized through non-metric multidimensional scaling (NMDS) at lactation day 11. Associated values were calculated with PERMANOVA are statistically significant, p = 0.01.
Community composition
Composition of the maternal and offspring gut microbiome was determined using the analysis of compositions of microbiome with bias correction (ANCOM-BC). In maternal samples collected at E17 and L11, there were no significant ASVs found to be different between mothers in the treatment or control groups. However, mothers treated with fluoxetine had higher relative abundances of Parasutterella at L15 (p = 0.008), and lower relative abundances of Turicibacter at L21 (p = 0.002; Table 3; Fig. 3). The offspring of mothers treated with fluoxetine had lower relative abundances of Turicibacter and Romboutsia at L11 (p = 0.08; p = 0.02; Fig. 4). Similarly, the offspring of mothers treated with fluoxetine had lower relative abundances of Romboutsia at L21 (p = 0.05).
Maternal genera of interest between treatment groups. Relative abundance of Parasutterella, Romboutsia, and Turicibacter at E17, L11, L15 and L21. See Table 3 for inferential tests of statistical significance.
Offspring genera of interest between treatment groups. Relative abundance of Romboutsia and Turicibacter at L11, L15 and L21. See Table 3 for inferential tests of statistical significance.
Discussion
Here we show that maternal fluoxetine exposure may result in decreased offspring alpha diversity in early life, a finding that may be particularly relevant for high-risk infants. Medically complex populations including low birthweight infants often already display lower gut microbial diversity compared to healthy, term-born infants26. A low diversity gut microbiome may be less stable than those that are higher and could be more easily perturbed by influences known to affect the infant gut microbiome such as antibiotic exposure27. It is also possible that a low diversity gut microbiome may be deficient in certain metabolites produced by missing microorganisms or suffer from a lack of functional redundancy. As such, infants with gut microbiomes deficient in certain metabolites may suffer from increased infection susceptibility and poor postnatal growth28. We note that early life growth patterns are particularly important, considering that positive growth patterns are associated with a variety of infant health outcomes, including sufficient neurocognitive development and markers of metabolic health29.
Our study also identified associations between maternal fluoxetine exposure and altered abundance of certain bacteria such as those belonging to the genus Turicibacter. Species of Turicibacter are known to possess a serotonin sensor (CUW_0748) with high structural homology to SERT, a serotonin transporter in mammalian cells that serves as the binding target of SSRI antidepressants30. Intestinal conditions high in serotonin have been shown to support the growth of Turicibacter sanguinis, but their abundance is significantly depleted by exposure to fluoxetine15. Our study supports this finding, as we observed that mothers treated with fluoxetine had lower relative abundance of Turicibacter at L21. However, we acknowledge that the decreased abundance we observed may result from other factors such as competitive colonization by other gut microbiota. We also observed similar changes in offspring Turicibacter abundance at L11, and our data suggests that Turicibacter are higher in abundance in the gut microbiomes of both mothers and offspring without fluoxetine exposure (Figs. 3 and 4).
It is important to note that there is accumulating evidence that supports the role of Turicibacter in systemic health. Members of the Turicibacter genus have been associated with depression severity and antidepressant exposure in both animal and human populations, and higher abundances of Turicibacter have been linked to lower reports of depression symptom severity31. This observation may be related to changes in serotonin signaling, as Turicibacter have been shown to respond positively to serotonin availability30. Gut microbial Turicibacter abundance may also serve as a determinant of metabolic health. Studies of Turicibacter sanguinis have shown altered intestinal expression of genes related to lipid metabolism and reduced host serum triglyceride levels in monocolonized mice. Moreover, T. sanguinis appears to regulate fat mass in sex-specific patterns where only female T. sanguinis-colonized mice exhibited reduced mass of inguinal white adipose tissue, relative to males15. In human populations, low abundance of Turicibacter were observed in children of mothers that were overweight32. Turicibacter are producers of butyrate and low abundances of this important short-chain fatty acid may represent a health risk, particularly in offspring with medical complications including those born with intestinal underdevelopment or in premature infants with low birth weight27.
In addition to Turicibacter our study also identifies other gut microbes associated with fluoxetine exposure that are likely related to systemic health. For example, we observed lower relative abundances of bacteria in the genus Romboutsia in offspring of mothers treated with fluoxetine. Human studies of SSRI exposure have found reduced abundances of Romboutsia in adults treated with SSRIs and that Romboutsia is positively correlated with depression symptom severity31. Furthermore, high Romboutsia abundance has been repeatedly associated with weight gain in children33,34. We also found that perinatal fluoxetine exposure is associated with a decrease in the relative abundance of maternal Parasutterella. While Parasutterella is commonly observed in the mouse and human gut microbiome, studies have suggested that patterns of increased Parasuterella abundance may be related to the development and progression of irritable bowel syndrome35,36. Parasutterella colonization has also been observed in the context of elevated intestinal inflammatory activity and an abundance of Parasutterella may exert effects on local and systemic health through production of succinate, an intermediate metabolite that functions in cross-feeding metabolic pathways35,36.
We present this study as the first to investigate the effects of maternal antidepressant exposure on the gut microbiome of offspring and acknowledge that our study is not without a number of limitations. While mouse models provide a level of experimental control not feasible in human studies, mice have the capacity to model depression with varying validity37. This study did not include a model of depression, although strong evidence supports the effects of depression on the gut microbiome in human populations17. Rather, we present this study as proof of concept that maternal fluoxetine exposure has the potential to affect both the maternal and offspring gut microbiome. Mice in this experiment were also treated with fluoxetine via subcutaneous injection which contrasts the traditional oral route of SSRI administration in humans. The use of subcutaneous injection was necessary to standardize antidepressant exposure in an intervention tolerable to maternal mice but to prevent unintentional offspring exposure. It is possible that the antimicrobial activity of SSRI antidepressants was muted by preventing direct administration of the SSRI into the gastrointestinal tract, as in humans. Furthermore, the period of maternal SSRI exposure in this study was 29 days. Because human populations are counseled to initiate antidepressant therapy for a minimum of several weeks before therapeutic effect is expected, the period of exposure in this experiment could be considered relatively short for the assessment of change in bacterial diversity or composition. It is possible that SSRI therapy may affect the human gut microbiome in ways not observed here, due to the inability to model the length of expected SSRI exposure, perinatal stage, and bacterial lifecycle at a level commensurate to what is experienced by humans.
In this study, we found that maternal antidepressant exposure has the capacity to affect both the maternal and offspring gut microbiome during pregnancy and early life. Fluoxetine exposure affected the relative abundance of several gut microbial genera and was associated with reduced alpha diversity of offspring in the early postnatal period. Changes in the relative abundance of specific bacteria within the gut microbiome may be relevant to the mental, gut, and metabolic health of perinatal populations. Future studies should consider examining the effects of other antidepressant medications commonly prescribed to pregnant and lactating people. Our findings support moving towards inquiry in human populations where impacts of depression on the gut microbiome can be considered alongside perinatal antidepressant exposure. Because concurrent antibiotic exposure or other determinants may act on the gut microbiome alongside SSRIs, researchers may consider inclusion of particularly vulnerable populations at high risk for gut microbial dysbiosis and intestinal morbidities such as preterm infants.
Methods
Animals
All experiments were approved by the University of Wisconsin-Madison’s Research Animal Care and Use Committee (RARC) under protocol #A005789-R03-A02 and all study procedures were performed in accordance with institutional and RARC guidelines. Reporting is consistent with The Animal Research: Reporting of In Vivo Experiments (ARRIVE) recommendations38. Mice were housed in a controlled environmental vivarium for biological research in the Department of Animal and Dairy Science at the University of Wisconsin-Madison. Wild-type C57BL/6J mice were obtained from Jackson Laboratories (Bar Harbor, ME, USA). Animal facilities were maintained at a temperature of 25 °C and a humidity of 50% to 60% with a 12:12 h light–dark cycle and mice had ad libitum access to both food (LabDiet 5015, TestDiet, Richmond, IN, USA) and water. All animals in our study were fed from the same food lot which was kept separate from food used for other experiments and were euthanized via carbon dioxide followed by cervical dislocation.
Experimental design
Mice were ordered at four weeks of age and housed according to RARC guidelines (based on weight and sex, up to six mice per cage unit) for two weeks. During this time, we followed a procedure to standardize the starting gut microbiome between animals, which included mixing cage bedding between study animals weekly39. We created a perinatal mouse model by breeding female mice with male mice beginning at six weeks of age. After detection of a vaginal plug, dams were individually housed. Mice were weighed on embryonic day 7 (E7) and embryonic day 10 (E10) with a 1.5g increase in weight indicating pregnancy40,41. At E10 pregnant mice were randomly assigned through block randomization to treatment groups and began receiving subcutaneous injections of 10 mg/kg/day fluoxetine in sterile saline or an equivalent volume of sterile saline as a control. Treatments continued through pregnancy and birth (embryonic day 18, E18) and until weaning at lactation on day 21 (L21). Animals were excluded for loss of pregnancy or litter. Mice were ordered, bred, and enrolled in two cohorts to support study feasibility in coordination with vivarium resources (Fig. 5).
i Cohort one
Cohort one mice were ordered, bred and enrolled into the experiment in September-December of 2021. Stool samples were collected from dams using sterile tweezers on embryonic day 10 (E10), embryonic day 17 (E17), and lactation day 11 (L11). At lactation day 11, a male and female pup from each litter were sacrificed and stool was collected from the gut using sterile utensils. Stool collection from the gut of L11 pups proved challenging due to small size of the animals and therefore, we piloted two collection procedures including (1) collection of an intact gut segment (containing stool) for extraction and (2) sectioning and scraping the gut interior for stool. Prior to collection, equipment was cleaned with a laboratory-grade detergent and sterilized with 70% ethanol. Bedding used in the bedding transfer procedure for Cohort one was collected and stored at − 80 °C and later introduced to the environment of Cohort two upon arrival. All stool samples collected in this cohort were immediately placed into microcentrifuge tubes and stored at − 80 °C.
ii Cohort two
Cohort two mice were ordered, bred and enrolled into the experiment in March–June of 2022. Stool samples were collected from dams using sterile tweezers on embryonic day 10 (E10), embryonic day 17 (E17), lactation day 15 (L15) and lactation day 21 (L21). From the offspring, stool samples were collected on lactation day 15 (L15) and lactation day (21). Samples were not collected from offspring at L11 due to the infrequency and unpredictability of stool sample availability at L11 in Cohort one. Instead, we collected a pooled stool sample from all offspring in each litter at L15. Samples were collected by placing pups on sterile gauze and collecting stool with a sterile tweezers. On L21, a sample was collected from a male and female pup from each litter. All stool samples collected were immediately placed into microcentrifuge tubes and stored at − 80 °C.
DNA extraction and sequencing
All samples were thawed to room temperature and processed individually. A total of 0.03–0.05g of stool was used for each extraction. Total genomic DNA was obtained by using a bead-beating protocol combined with a phenol:chloroform extraction with the following modification: all aqueous phase washes used 25:24:1 phenol:chloroform:isoamyl alcohol instead of phenol:chloroform for a total of three washes42. Four extraction controls were processed with the stool samples and one control was processed during the final PCR reactions.
DNA was quantified using a Qubit fluorometer reagents (Invitrogen, Waltham, MA) and a Synergy 2 microplate reader (BioTek, Winooski, VT, USA). The V4 region of the 16S rRNA gene was amplified via polymerase chain reaction (PCR) using a universal bacterial primer (F-GTGCCAGCMGCCGCGGTAA; R-GGACTACHVGGGTWTCTAAT). Each of these primers were barcoded with individual custom indices to facilitate demultiplexing, as previously described43. Each PCR reaction consisted of 12.5 μl KAPA 2 × HiFi Master Mix (KAPA Biosystems, Wilmington, MA, USA), 0.5 μl of 10 μM forward primer, 0.5 μl of 10 μM reverse primer and up to 11.5 μl of 10ng/μl DNA to a total volume of 25 μl with nuclease-free water (IDT, Coralville, Iowa, USA). PCR was conducted on a C1000 Touch™ thermal cycler (Bio-Rad Laboratories, Hercules, CA, USA) with the following amplification protocol: 95 °C for 3 min, 35 cycles of 95° for 30 s, 55 °C for 30 s, and 72 °C for 30 s, followed by a final extension at 72 °C for 5 min. PCR products were quantified on a 1% (w/v) low-melt agarose gel using AquaPor low-melt agarose (National Diagnositcs, Atlanta, GA) using SYBRSafe DNA gel stain (Invitrogen, Waltham, CA), with bands at ~ 380 bp indicating successful amplification. These bands were excised, extracted, and purified using a Zymoclean Gel DNA Recovery Kit (Zymo Research, Irving, CA). A no-template negative control was included with each set of PCRs and if a band was present in the negative control, all samples in that set were redone starting with PCR set-up and amplification. Negative controls for which no band was present had the approximate location of the amplicon (~ 380 bp) excised and sequenced as further confirmation that no contamination was present. The gel-extracted DNA was then quantified using a Qubit fluorometer and a 96-well plate spectrophotometer, and a library was created using a 4 nmol/L equimolar pool of all PCR products. This library was then sequenced on an Illumina MiSeq (Illumina Inc., San Diego, CA) following standard sequencing protocols using a MiSeq v2 2 × 250 bp sequencing kit. Raw sequences from this study were deposited into the National Center for Biotechnological Information’s (NCBI) Short Read Archive (SRA) and are publicly accessible under accession #PRJNA1084828.
Statistical analysis
Bioinformatic processing and statistical analysis of sequencing reads was performed using QIIME II (2023.5) and R (v 4.1.2)44,45. The standard operating procedure described in the QIIME II “Moving Pictures” tutorial, accessed on June 2, 202346, was used to process all sequences. Briefly, paired-end demultiplexed sequences with quality scores in the Casava 1.8 format were imported into QIIME II and the q2-dada2 (v.1.18.0) plugin was used for quality control, trimming, and denoising47. A feature table of amplicon sequence variants (ASVs), which are unique sequences not grouped by percentage threshold, was generated. ASVs were aligned with MAFFT (v.7.475), and a phylogenetic tree was constructed using FastTree (v.2.1.1). Taxonomy was assigned using the Silva database (v.138.1, released August 27, 2020) and the classify-sklearn naïve Bayes taxonomy classifier (v0.24.1)48,49. The resulting feature table and phylogenetic tree were imported into R for downstream statistical analyses.
The Qiime II feature table and experimental metadata (animal number, treatment, etc.; Microsoft Excel v.2310) were imported into R (1.38.0)50. Possible contaminants (Mitochondria, Chloroplasts, Eukaryotes, unclassified) were pruned and ASVs prevalent in negative controls and low read samples (< 5000) were removed with decontam (v.1.14.0)25,50. Sequences were rarefied to an even depth of 8000 reads per sample50. Alpha diversity was calculated using Shannon’s alpha diversity index and using the Shapiro Wilk test, values were found to have a non-normal distribution. Subsequently, inferential comparisons were made using non-parametric Wilcoxon Signed Rank tests. Beta diversity was visualized using non-metric multidimensional scaling (NMDS) with Bray–Curtis and Unifrac (weighted/unweighted) distance matrixes and inferential comparisons were made using PERMANOVA in the phyloseq package50. Testing for differentially abundant taxa was performed using ANCOM-BC (v.1.4.0), a linear regression based method designed to correct biases introduced through differences in sampling fraction51. The reported p values were adjusted for false discovery rate using parameters internal to ANCOM-BC51. For the purposes of this study, results were considered differentially abundant at an (adjusted) alpha level of 0.1.
Data availability
The datasets generated during and/or analysed during the current study are available in the National Center for Biotechnological Information’s (NCBI) Short Read Archive (SRA) and are publicly accessaible under accession #PRJNA1084828 at https://www.ncbi.nlm.nih.gov/sra.
References
The American College of Obstetrics and Gynecology. ACOG Committee opinion no. 757 summary: Screening for perinatal depression. Obstet. Gynecol. 132, e208–e212 (2018).
Kiewa, J. et al. Lifetime prevalence and correlates of perinatal depression in a case-cohort study of depression. BMJ Open 12, e059300 (2022).
World Health Organization. Launch of the WHO guide for integration of perinatal mental health in maternal and child health services. https://www.who.int/news/item/19-09-2022-launch-of-the-who-guide-for-integration-of-perinatal-mental-health (2023).
Brown, C. C., Adams, C. E., George, K. E. & Moore, J. E. Mental health conditions increase severe maternal morbidity by 50 percent and cost $102 million yearly in the United States. Health Aff. 40, 1575–1584 (2021).
The Joint Commission. Eliminating racial and ethnic disparities causing mortality and morbidity in pregnant and postpartum patients. Sentinel Event Alert. https://www.jointcommission.org/-/media/tjc/newsletters/sea-66-maternal-mm-and-he-1-13-23-final.pdf (2023).
Cooper, W. O., Willy, M. E., Pont, S. J. & Ray, W. A. Increasing use of antidepressants in pregnancy. Am. J. Obstet. Gynecol. 196(544), e1-544.e5 (2007).
Petersen, J. M., Esposito, D. B. & Werler, M. M. Selective serotonin reuptake inhibitor use patterns among commercially insured US pregnancies (2005–2014). Arch. Womens Ment. Health 24, 155–164 (2021).
Molenaar, N. M., Kamperman, A. M., Boyce, P. & Bergink, V. Guidelines on treatment of perinatal depression with antidepressants: An international review. Aust. N. Z. J. Psychiatry 52, 320–327. https://doi.org/10.1177/0004867418762057 (2018).
Molenaar, N. M. et al. The international prevalence of antidepressant use before, during, and after pregnancy: A systematic review and meta-analysis of timing, type of prescriptions and geographical variability. J. Affect. Disord. 264, 82–89. https://doi.org/10.1016/j.jad.2019.12.014 (2020).
Komorowski, J. Antidepressants in pregnancy. In Clinical Pharmacology During Pregnancy 2nd edn (eds Mattison, D. & Halbert, L.-A.) 311–321 (Academic Press, 2022).
Vahora, I. S., Tsouklidis, N., Kumar, R., Soni, R. & Khan, S. How serotonin level fluctuation affects the effectiveness of treatment in irritable bowel syndrome. Cureus https://doi.org/10.7759/cureus.c36 (2020).
Macedo, D. et al. Antidepressants, antimicrobials or both? Gut microbiota dysbiosis in depression and possible implications of the antimicrobial effects of antidepressant drugs for antidepressant effectiveness. J. Affect. Disord. 208, 22–32. https://doi.org/10.1016/j.jad.2016.09.012 (2017).
McGovern, A. S., Hamlin, A. S. & Winter, G. A review of the antimicrobial side of antidepressants and its putative implications on the gut microbiome. Aust. N. Z. J. Psychiatry 53, 1151–1166. https://doi.org/10.1177/0004867419877954 (2019).
Jones, L. A., Sun, E. W., Martin, A. M. & Keating, D. J. The ever-changing roles of serotonin. Int. J. Biochem. Cell Biol. 125, 105776. https://doi.org/10.1016/j.biocel.2020.105776 (2020).
Fung, T. C. et al. Intestinal serotonin and fluoxetine exposure modulate bacterial colonization in the gut. Nat. Microbiol. 4, 2064–2073. https://doi.org/10.1038/s41564-019-0540-4 (2019).
Duan, J. et al. Characterization of gut microbiome in mice model of depression with divergent response to escitalopram treatment. Transl. Psychiatry 11, 303 (2021).
Amirkhanzadeh Barandouzi, Z., Starkweather, A. R., Henderson, W. A., Gyamfi, A. & Cong, X. S. Altered composition of gut microbiota in depression: A systematic review. Front. Psychiatry 11, 1–10. https://doi.org/10.3389/fpsyt.2020.00541 (2020).
Shen, Y., Yang, X., Li, G., Gao, J. & Liang, Y. The change of gut microbiota in MDD patients under SSRIs treatment. Sci. Rep. 11, 14918 (2021).
Jackson, M. A. et al. Gut microbiota associations with common diseases and prescription medications in a population-based cohort. Nat. Commun. 9, 2655 (2018).
Vuong, H. E. et al. Interactions between maternal fluoxetine exposure, the maternal gut microbiome and fetal neurodevelopment in mice. Behav. Brain Res. 410, 113353 (2021).
Ramsteijn, A. S., Jašarević, E., Houwing, D. J., Bale, T. L. & Olivier, J. D. A. Antidepressant treatment with fluoxetine during pregnancy and lactation modulates the gut microbiome and metabolome in a rat model relevant to depression. Gut Microbes 11, 735–753 (2020).
Sarkar, A., Yoo, J. Y., Dutra, S. V. O., Morgan, K. H. & Groer, M. The association between early-life gut microbiota and long-term health and diseases. J. Clin. Med. 10, 1–24 (2021).
Levin, A. M. et al. Joint effects of pregnancy, sociocultural, and environmental factors on early life gut microbiome structure and diversity. Sci. Rep. 6, 31775 (2016).
Lewis, C. R. et al. Family SES is associated with the gut microbiome in infants and children. Microorganisms 9, 1608 (2021).
Davis, N. M., Proctor, D. M., Holmes, S. P., Relman, D. A. & Callahan, B. J. Simple statistical identification and removal of contaminant sequences in marker-gene and metagenomics data. Microbiome 6, 226 (2018).
Arboleya, S. et al. Establishment and development of intestinal microbiota in preterm neonates. FEMS Microbiol. Ecol. 79, 763–772 (2012).
Groer, M. W. et al. Contributors to dysbiosis in very-low-birth-weight infants. J. Obstet. Gynecol. Neonatal Nurs. 49, 232–242 (2020).
Groer, M. et al. Predicted metabolic pathway distributions in stool bacteria in very-low-birth-weight infants: Potential relationships with NICU faltered growth. Nutrients 12, 1345 (2020).
Castanys-Muñoz, E. et al. Systematic review indicates postnatal growth in term infants born small-for-gestational-age being associated with later neurocognitive and metabolic outcomes. Acta Paediatr. 106, 1230–1238. https://doi.org/10.1111/apa.13868 (2017).
Hoffman, J. M. & Margolis, K. G. Building community in the gut: A role for mucosal serotonin. Nat. Rev. Gastroenterol. Hepatol. 17, 6–8. https://doi.org/10.1038/s41575-019-0227-6 (2020).
Lin, S. K. K. et al. Exploring the human gut microbiota targets in relation to the use of contemporary antidepressants. J. Affect. Disord. 344, 473–484 (2024).
Gilley, S. P. et al. Associations between maternal obesity and offspring gut microbiome in the first year of life. Pediatr. Obes. 17, e12921 (2022).
Pan, L. Y., Zhou, Y. Y., Zhang, X. & Jiang, H. Y. Gut microbiota is associated with weight gain in children treated with atypical antipsychotic: A pilot longitudinal study. Psychiatry Res. 316, 114784 (2022).
Wei, Y. et al. The associations of the gut microbiome composition and short-chain fatty acid concentrations with body fat distribution in children. Clin. Nutr. 40, 3379–3390 (2021).
Ju, T., Kong, J. Y., Stothard, P. & Willing, B. P. Defining the role of Parasutterella, a previously uncharacterized member of the core gut microbiota. ISME J. 13, 1520–1534 (2019).
Chen, Y. J. et al. Parasutterella, in association with irritable bowel syndrome and intestinal chronic inflammation. J. Gastroenterol. Hepatol. 33, 1844–1852 (2018).
Acikgoz, B., Dalkiran, B. & Dayi, A. An overview of the currency and usefulness of behavioral tests used from past to present to assess anxiety, social behavior and depression in rats and mice. Behav. Process. 200, 104670. https://doi.org/10.1016/j.beproc.2022.104670 (2022).
du Sert, N. P. et al. The arrive guidelines 2.0: Updated guidelines for reporting animal research. PLoS Biol. 18, e3000410 (2020).
Miyoshi, J. et al. Minimizing confounders and increasing data quality in murine models for studies of the gut microbiome. PeerJ 6, e5166 (2018).
Heyne, G. W. et al. A simple and reliable method for early pregnancy detection in inbred mice. J. Am. Assoc. Lab. Anim. Sci. 54, 368–371 (2015).
Domingues, R. R. et al. Effect of low and high doses of two selective serotonin reuptake inhibitors on pregnancy outcomes and neonatal mortality. Toxics 10, 11 (2022).
Eggers, S. et al. Wisconsin microbiome study, a cross-sectional investigation of dietary fibre, microbiome composition and antibiotic-resistant organisms: Rationale and methods. BMJ Open 8, e019450 (2018).
Kozich, J. J., Westcott, S. L., Baxter, N. T., Highlander, S. K. & Schloss, P. D. Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the miseq illumina sequencing platform. Appl. Environ. Microbiol. 79, 5112–5120 (2013).
Bolyen, E. et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 37, 852–857 (2019).
R Core Team. R: A language and environment for statistical computing (2021).
QIIME 2 Development Team. QIIME 2: Moving Pictures Tutorial. https://docs.qiime2.org/2022.2/tutorials/moving-pictures/ (2021).
Callahan, B. J. et al. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
Katoh, K., Misawa, K., Kuma, K. & Miyata, T. MAFFT: A novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 30, 3059–3066 (2002).
Price, M. N., Dehal, P. S. & Arkin, A. P. FastTree 2—Approximately maximum-likelihood trees for large alignments. PLoS One 5, e9490 (2010).
McMurdie, P. J. & Holmes, S. phyloseq: An R package for reproducible interactive analysis and graphics of microbiome census data. PLoS One 8, e61217 (2013).
Lin, H. & Peddada, S. D. Analysis of compositions of microbiomes with bias correction. Nat. Commun. 11, 3514 (2020).
Acknowledgements
Research reported in this manuscript was supported by The National Institutes of Health under award number R01HD094759. KDS was supported by the University of Wisconsin-Madison Metabolism and Nutrition Training Program (5T32-DK007665) and IZ-C was supported by the University of Wisconsin-Madison Department of Bacteriology Roland and Nina Girolami Predoctoral Fellowship.
Author information
Authors and Affiliations
Contributions
K.M.D.S. is the corresponding author responsible for the conception, design, analysis and interpretation of the work. M.R. is credited with roles in study design and data acquisition. I.Z.-C. strongly and significantly contributed to analysis and interpretation. G.S. and L.L.H. provided mentorship through all phases including conception, design analysis and interpretation.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Desorcy-Scherer, K., Zuniga-Chaves, I., Reisner, M.A. et al. Investigating the influence of perinatal fluoxetine exposure on murine gut microbial communities during pregnancy and lactation. Sci Rep 14, 13762 (2024). https://doi.org/10.1038/s41598-024-62224-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-62224-7