Abstract
The fundamental question of how forces are generated in a motile cell, a lamellipodium, and a comet tail is the subject of this note. It is now well established that cellular motility results from the polymerization of actin, the most abundant protein in eukaryotic cells, into an interconnected set of filaments. We portray this process in a continuum mechanics framework, claiming that polymerization promotes a mechanical swelling in a narrow zone around the nucleation loci, which ultimately results in cellular or bacterial motility. To this aim, a new paradigm in continuum multi-physics has been designed, departing from the well-known theory of Larché–Cahn chemo-transport-mechanics. In this note, we set up the theory of network growth and compare the outcomes of numerical simulations with experimental evidence.
Similar content being viewed by others
Introduction
Actin-based motility (ABM) has an essential role in several biological processes in animal cells1,2,3: to name a few, embryogenesis4, angiogenesis5, tumor metastasis6, and the immune response of cells7. Cell motility relies on the assembly of actin filaments, which organize into a variety of architectures to promote protrusion (branched or crosslinked networks in the lamellipodium, parallel bundles in filopodia) and retraction (the antiparallel structures in contractile fibers). A detailed review of actin’s biochemical properties and interactions with other proteins can be found in8.
ABM is pivotal in cellular motility through the spatial and temporal coordination of three processes. Actin filaments undergo nucleation and primarily grow at the leading edge of a cell, immediately adjacent to the plasma membrane. Subsequently, actin filaments are interconnected into a network that shifts rearward toward the cell body, with different speeds in fast-moving cells or in stationary/slow-moving cells. Away from the leading edge, the network undergoes rapid depolymerization9.
ABM is also a key process in the life cycle of several bacteria, which propel into the cytoplasm and infect the human body10. The engine of their motion11 lies within a strip of actin filaments at the pathogen’s surface pole12, called the actin comet tail (ACT). It self-assembles using the host cell’s cytoskeletal machinery. Hijacking the cell’s functioning allows hosted pathogens to increase infectivity and virulence, spreading the infection to neighboring cells. Listeria monocytogenes13 has been selected for pioneering studies due to its experimental repeatability, from both biophysical and biochemical standpoints. The comet tail of Listeria bears a resemblance to a simplified lamellipodium, with the bacterial surface mimicking the plasma membrane at its forefront. Within living cells, actin monomers exhibit a preference for incorporating into the actin tail near the bacterial surface, mirroring the behavior observed in lamellipodia where new incorporation predominantly occurs at the leading edge. Fluorescence photoactivation experiments reveal that filaments within the tail of a moving bacterium remain stationary while only the bacterium itself advances, akin to rapidly moving lamellipodia. Furthermore, the rate of filament depolymerization within the comet tail appears to be unaffected by either its position within the tail or the speed of the bacterium. These filaments exhibit notably short half-lives, typically around 30 seconds.
Most of the abundant published research focuses on molecular questions about the nucleation, elongation, and depolymerization of single filaments (to name a few, see14,15,16,17). Countless publications have been devoted to ABM experimental biology. The interactions of actin filaments with the surface of Listeria are the cornerstone of the Brownian ratchet model18. Despite some limitations, the model successfully establishes a connection between the macroscopic movement of Listeria and microscopic polymerization events, establishing a force-dependent rate of filament growth. Key ingredients of the symmetry breaking of an actin gel around a bead have been studied in several publications, such as19,20 and references therein. The mechanism of polymerization in the elastic ratchet model of Mogilner and Oster is based on the fluctuations of filaments against the bacterium21. A two-compartment model, consisting of attached and detached filaments, was formulated. The detached, compressed fibers (the “working filaments”) generate the protrusive force, and the attached, stretched fibers resist the advancement of the bacterium. The Listeria ACT was modeled in22 as a continuum linear elastic gel. The addition of filaments at the bacterium surface builds up a new polymerized layer, which compresses the previously formed ones and deforms the gel. The shape of the ACT, the motion, and the external forces on the bacterium surface all result from the lowest energy conformation of the gel.
Examining the mechanics of biological systems at the tiniest level may not always offer a comprehensive understanding23. Theoretical and computational investigations into the physical response of biopolymer/cytoskeletal networks, with particular emphasis on constitutive theories, have been reviewed in24 at different scales. This note falls into the class of macroscopic, continuum models. It aims to deliver a chemo-transport-mechanical formulation of the biochemical processes (depicted in Sect. 2) that enable ABM, rooted in the modern theory of multiphysics of continua at finite strains25. Governing equations are derived in Sect. 3 from conservation and constitutive laws obtained from thermodynamic prescriptions, departing from the classical Larché–Cahn approach. We attribute the ABM at the continuum scale to a volumetric expansion of the branched and cross-linked actin network after polymerization.
A Larché–Cahn model25 depicts the flow of one or more generic species within a hosting material, which serves as the source of the mechanical strength of the system. This approach has been successfully applied to the flow of hydrogen in metals and lithium in batteries. Larché–Cahn models account for the influence of flowing species on the mechanical response of the system, for instance, because of alloying or phase transformations, and can be viewed as a limiting case of the theory of mixtures. Unfortunately, though, Larché–Cahn’s simple approach does not apply well to mechanobiologically complex systems, which would rather benefit from a compressible mixture theory. Literature on this topic is not abundant26,27, and its profound analysis goes beyond the scope of the present note.
The papers28,29 from Kimpton and coauthors, although limited to one-dimensional formulations, share common ideas with our manuscript. Specifically: (i) the idea of two phases, the network and the solution in which actin monomers flow; (ii) the mass transfer between phases, which corresponds to polymerization/depolymerization; (iii) the notion of swelling and contractility. However, all these concepts are depicted in a different way. Kimpton and coauthors use a two-phase mixture made of incompressible Newtonian (in28) or viscoelastic upper-convected Maxwell (in29) fluids, where the notion of swelling and contractility leads to a definition of pressure (swelling = compressive, contractile = tensile) related to the volume fractions. The notion of volumetric expansion is completely absent.
We base our theory on the assumption that the cytosol occupies a large volume that encloses the network and fills all spaces within it at a much faster pace than any other motions. In this way, we write balance equations in the same domain for monomers and the network. We assume that they coexist in an ideal material, which has the transport properties of the cytosol and the mechanical features of the biopolymer network. The latter can be modeled with a thermodynamically consistent constitutive law. We selected the mechanical free energy density from an elastic material in the realm of finite strains, assuming that viscous effects do not have sufficient time to develop. When convenient, more sophisticated constitutive laws can be chosen.
Bacterial ABM does not rely on the host cell plasma membrane, allowing for reconstitution in a cell-free cytoplasmic extract. This has enabled biochemical studies to investigate the regulation of actin dynamics, demonstrating that polystyrene beads coated with purified ActA protein can form comet tails and move within cytoplasmic extracts. Measurements using a differential Atomic Force Microscope assay were performed in30. The model has been validated in "Balance equations" against this experimental evidence.
Biochemical Processes and Model Assumptions
Upon nucleation, actin filaments grow by the insertion of new monomers of globular (G) actin, which land on the ends of the filaments by diffusion in the cytoplasm. The addition of new monomers elongates the polymerized network and promotes the protrusion of cells or bacteria. Cells are capable of polymerizing or depolymerizing micrometer-thick sheets of densely packed filaments in seconds31. Diverse proteins regulate ABM9. Actin, Arp2/3, and capping protein are essential for the polymerization, while ADF (actin depolymerizing factor) and cofilin disassemble old filaments, VASP (vasodilator-stimulated phosphoprotein) and profilin increase the rate of movement, and \(\alpha\)-actinin stabilizes the network.
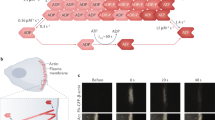
After invading the host cell cytoplasm, bacteria cover one surface pole with ActA, a protein capable of binding several available factors for the nucleation and branching of actin filaments—see Fig. 1. Interactions between host-cell proteins and ActA on the bacterial surface are crucial. Two ActA domains are vital for promoting normal motility. The amino-terminal domain stimulates actin filament nucleation through Arp2/3, while the central proline-rich domain binds VASP. Profilin interacts with VASP, enhancing filament elongation. Host protein functions are distributed throughout the comet tail. In addition to the factors at the bacterial surface, capping protein attaches to the barbed end of actin filaments to inhibit elongation of older filaments. \(\alpha\)-actinin links filaments together, reinforcing the tail structure, and ADF/cofilin breaks down aged filaments.
Recent experiments have convincingly shown that the actin network is attached to the surface of the pathogens, with no gap between filaments and the bacterial surface32,33. This evidence indicates that nascent filaments are nucleated by the protein complexes on the bacteria surface, then they dissociate and grow freely until they are eventually capped21.
Either in pathogens or in cells, ABM is based on three main processes: (i) assembly of the network, via the nucleation, polymerization, branching, and cross-linking of new actin filaments from the G-actin monomer pool; (ii) protrusion, related to actin filament growth at the nucleation surface; (iii) disassembly, which entails the depolymerization of the old filaments and the fragmentation or severing of the tail3,12.
Moving from the molecular to the macroscopic scale, where the laws of classical mechanics25 hold, chemo-transport-mechanics equations34,35 can describe these three coupled processes in a continuum framework. Two mass balance equations govern the transport and polymerization of the actin species, while the balances of momentum depict the mechanical response of the network. The assembly of the ACT is portrayed by a single chemical reaction
which models nucleation, branching, and cross-linking and is active in a volume surrounding the nucleation loci, defined by an intrinsic length scale denoted by \(\ell\) in what follows—see Fig. 2. Equation (1) identifies how many moles of G-actin in the cytosol are converted into the cross-linked network (F), defining a mole of F as the result of the polymerization of a mole of G. The structural arrangement of dispersed monomers into an organized network has been claimed36 to be the engine of the ABM, both for cellular lamellipodia and bacteria comet tails. The spatial reorganization entails an increase in the partial molar volume, depicted in Fig. 2, which is responsible for two major mechanical events: (i) a volumetric swelling in the material, induced by the different partial molar volumes of G and F-actin during assembly and disassembly—see Fig. 2; (ii) a mechanical flow of the F-actin network.
The volumetric swelling of the material is induced by the rearrangement of actin in space, and the F-actin mechanical flow is due to a compression state of the F-actin network. We emphasize that a phase transformation of actin from G to F not accompanied by a change in molar volume cannot explain the actin-based protrusion. Furthermore, it has been experimentally shown that a spot photobleached in the actin meshwork of the lamellipodium translocates backward slowly and moves rearward relative to the leading edge 9. These evidences align perfectly with the concept of volumetric swelling induced by actin polymerization that is depicted in Fig. 2 and detailed in the present work for both cellular lamellipodia and bacterial comet tails.
Schematic representation of volume increment and protrusion upon the structural arrangement of monomers after polymerization of the dense actin filament network in lamellipodia (brown). The sparser actin network in the lamella is represented in purple. Scale bar, 1 \(\upmu\)m. Inspired by12.
Modeling ABM at a continuum scale
The transport and reorganization of unconnected monomeric subunits into a network occur in the cytosol, see Fig. 2, a dense viscous medium that we do not equip with any mechanical strength. It surrounds the F-actin network and is capable of providing a viscous drag on bacteria, which opposes the retrograde motion of the F-actin network. The mechanical strength and stiffness are provided solely by the F-network upon its polymerization.
Balance equations
We assume that the cytosol occupies a large volume that encloses F and fills all spaces within it at a much faster pace than any other motions. In this way, the cytosol remains undisturbed by the spatial evolution of F and can be considered a medium that merely allows G-monomers to flow and reach the polymerization sites. In principle balance equations should be written in different domains for G and F. However, we assume that they coexist in an ideal material, which has the transport properties of the cytosol and the mechanical properties of F. The referential balance equations for monomers and network will be thus written at the same point \({\vec {X}}\)
\(c_{a_R}\) is the referential molarity (moles per unit undeformed volume) and \({\vec {H}}_{a}( {\vec {X}}, t)\) denotes the referential flux vector of species \(a=F,G\). In a simplistic scenario, the G monomers can be though of being replenished at infinite velocity, so that their concentration in the current configuration is constant in time and equal to the initial value. The equation
would then replace Eq. (2a).
The rate of the reaction depicted in Eq. (1), \(w^(1)_R\), is defined via the law of mass action and, considering that species concentrations remain far from saturation in their own binding sites, we can assume
In most forms of ABM, there is an external energy input that effectively reverses the polymerization reaction and maintains the actin filaments far from chemical equilibrium. The best-controlled form of this external energy operates in continuous cycles at the expense of ATP hydrolysis. In this way, the polymer can use the free energy of ATP hydrolysis to translate the kinetic differences between subunit addition rates at the plus and minus ends into energetic differences that can potentially be exploited by the polymerization motor37. In addition to hydrolysis, accessory proteins such as Arp2/3, VASP and capping protein, are summoned to support the F-actin network formation during actin polymerization.
As done in38, we embody all these biochemical events in a single function \({{\mathcal {C}}}({\vec {x}},t)\) within the forward reaction rate parameter \(k_{f}\)
where \({k}^*_{f}\) is a constant. Function \({{\mathcal {C}}}({\vec {x}},t)\) will be named the activation signal function. Its role is to localize the polymerization reaction (1) where it takes place, i.e., by a pathogen surface pole or at the leading edge of a cell, immediately adjacent to the plasma membrane. Since reaction (1) transforms monomers into fully formed network (and not into single actin filaments) the activation signal function is non-vanishing in a small volume rather than along a manifold, to incorporate the role of ATP and accessory proteins described above.
In a similar way, we will not indulge in modeling every single complex detail of the network disassembly3. Rather than scrupulously accounting for every fine-scale mechanism of network severance and depolymerization, the average lifetime \(\tau\) of filaments in cells9 is used.
The backward reaction rate parameter is defined as:
The assumption that both the forward and backward reaction parameters are force-independent is a simplification of the real process, since the presence of the load force can affect both \(k_{f}\) and \(k_{b}\)37.
The difference \(\Omega _F=\omega _F-\omega _G\) between the partial molar volumes (\(\omega\)) of F and G in the reaction (1) is the mechanical essence of the polymerization motor. The rearrangement of monomers results in a volume deformation, which is captured by the multiplicative decomposition
of the deformation gradient. The polymerization tensor \({\textbf {F}}^c\) is the local distortion of the neighborhood at point \(\vec {X}\) in the comet tail or in the lamella after polymerization of monomers into a network of filaments and vice versa39. In this note an isotropic chemical swelling
was considered, since it appears well designed for the complex orientation of filaments in the ACT and in the lamella (see Fig. 2). The elastic tensor \({\textbf {F}}^{e}\) recovers the (visco)-elasticity of the network. As stated in25 for the Kröner decomposition, the elastic and the polymerization tensors are not the gradients of point-field maps. In fact, an intermediate and stress-free configuration in which referential vectors are mapped via \({\textbf {F}}^c\) can only be idealized since it would lack geometrical compatibility.
The balance of linear momentum in the referential local form reads
where \({ {\textbf {P}}}\) denotes the first Piola–Kirchhoff stress tensor, \({\vec {B}}\) represents the body forces (per unit referential volume), and the right-hand side is the time derivative of the linear momentum. Note that the time derivative also involves the network concentration, which is not constant over time; in other words, the network density evolves in time according to the polymerization reaction (1) and such time evolution enters the balance of linear momentum. We assume henceforth that \(M_F\), the molar mass of the F-actin network, is small, so that the inertial effects in (8) are second-order and can be neglected.
At time \(t=0\) only the cytosol is assumed to exist, together with a signaling locus at which polymerization develops at all times \(t>0\). Since the F-network is initially absent, the notion of hosting material is meaningless and one concludes that differently from several well established theories40, either accounting for trapping or not34,35, the present framework for ABM does not fall into the class of Larché–Cahn models. In other words, the material density of the hosting material is not associated to \(J^c\), but is also related to the density of the polymer network in a stress free configuration.
Thermodynamics
Thermodynamic prescriptions have been derived from a rigorous, now classical25, approach that entails the energy balance, the entropy imbalance, the Clausius–Duhem inequality, and the Coleman-Noll procedure. By selecting the referential Helmholtz free energy density \(\psi _R\) as the thermodynamic potential, the second Piola–Kirchhoff stress \({\textbf {S}}\) and the chemical potentials \(\mu _G\), \(\mu _F\) are obtained
with the Green-Lagrange tensor \({\textbf {E}}\) and concentrations \(c_{G_R}, \, c_{F_R}\) as state variables. The first and second Piola–Kirchhoff stress tensors are related by the identity \({ {\textbf {P}}} = {\textbf {F}} \, {\textbf {S}}\), with \({\textbf {F}}\) as in (6). To satisfy the Clausius–Duhem inequality, isotropic fluxes can be expressed through the definition of positive mobilities \({M}_{G_R}\), \({M}_{F_R}\) as
\(a=F,G\). The Helmholtz free energy density is defined via an additive decomposition
which accounts for the entropic and the mechanical contributions. The entropic free energy does not involve the F-actin network concentration. This choice has a fundamental consequence: the chemical potential \({\mu }_{F}\) in (9a) is not entropic in nature; it is solely due to the mechanics of protrusion via the swelling tensor defined in (7). The same interpretation applies to the mass flux \(\vec {H}_F\) in (9b).
The entropic free energy density
where R is the gas constant, T is temperature, \(c^{sat}_{G_R}\) is the saturation limit for the actin monomers, is inferred from statistical mechanics.
Experimental evidence in ABM shows a large spectrum of F-network velocities. The observed velocities of the free-expansion F-network reported in the literature vary widely. For example, the study30 measures \(85 \pm 68 \mathrm{nm/min}\), while41 reports \(74 \pm 50 \mathrm{nm/s} = 4.4 \pm 0.3 {\upmu }\mathrm{m/min}\), and42 finds \(2 {\mu }\mathrm{m/min}\). These three measurements show significant discrepancies, suggesting that the experimental conditions may differ considerably and that the velocity at zero force is highly sensitive to these conditions. The biological motivations of these observations warrant further studies. Bacterial ABM is rapid in the host cytoplasm, reaching speeds of up to 1 \(\upmu \mathrm{m/s}\)9. Given this speed, viscous effects do not have enough time to develop, although some rearrangement of cross-links may still occur. Accordingly, we derive the mechanical free energy density from an elastic material, i.e.
with the right Cauchy–Green tensors \({\textbf {C}} = {{\textbf {F}}}^\textsf{T} {\textbf {F}}\), \({\textbf {C}}^e = {{\textbf {F}}^{e}}^\textsf{T} {\textbf {F}}^{e}\), the Green-Lagrange tensors \({\textbf {E}} = \frac{1}{ 2} ( {\textbf {C}} - \mathbbm {1})\), \({\textbf {E}}^e = \frac{1}{ 2} ( {\textbf {C}}^e - \mathbbm {1})\), and
—see Appendix A—and \(\psi _R^{el}= 0\) at vanishing \({{\textbf {E}}^e}\). Easy algebra shows that, in view of Eqs. (9), (10), and (12),
The model permits any specification of the Helmholtz free energy \(\psi _R^{el}({{\textbf {E}}^e} )\). Since the focus of this note is the ABM engine rather than the mechanical response of the F-network, we arbitrarily and simplistically consider in the numerical simulations an elastic strain energy of Saint-Venant type
where \(\lambda\), G are the Lamé parameters43. Viscosity is purposely neglected since it is argued that the cross-linking rearrangement in the F-actin network has insufficient time to develop3. The mechanics of the actin network is determined by its mesh size, which can be defined either as the spacing between actin filaments within the network or the distance between fixed cross-link points. There is evidence that the mesh size affects the Lamé parameters, and thus the Helmholtz free energy \(\psi _R^{el}({{\textbf {E}}^e} )\) would depend on the concentration \(c_{F_R}\) and possibly on further internal variables to capture the history dependence of the constitutive law. Since we limit our investigation to the ABM engine in this note, we postpone the mechanical response of the F-network to future studies and assume constant Lamé parameters.
Signaling
As stated, all biological and biochemical events that trigger actin nucleation, branching, and cross-linking have been encapsulated in the activation signal function \({{\mathcal {C}}}(\vec {x},t)\), which localizes the signaling in Eq. (4) to the locations in the current configuration where the protein ActA is present, such as the bacterium surface. We take \({{\mathcal {C}}}(\vec {x}, t)\) to vanish everywhere except in the specific domain
with characteristic length \(\ell\) about the nucleation surface \(\partial _b\), where it holds
Mechanobiological properties
The forward reaction constant \({k}^*_{f} =10.00\, \textrm{s}^{-1}\) in (4) has been estimated for bacteria in44. The backward reaction rate parameter has been set to \(k^*_{b}=0.25\, \textrm{s}^{-1}\), consistent with the severing parameter r in44. The rate of filament depolymerization is independent of the position in the tail and the bacterial speed. According to9, filaments have a half-life in the order of \(\tau =30\) s. The G-actin diffusion coefficient in the cytoplasm \(\textrm{D} \hspace{-4.44443pt}| \,_{G_R}=3.00 \,{\upmu \mathrm{m}^{2} \mathrm{s}^{-1}}\) has been reported in45. According to46, the approximate length of the ACT is \(l \approx 10\, \mathrm {\mu m}\). The G-actin has a molar volume of \(32.2\, \mathrm{l \, mol^{-1}}\), according to47. The equilibrium temperature is assumed to be \(\textrm{T}=310 \, \textrm{K}\)48. Following22, the Young’s modulus of the F-network ranges from 1 to 10 kPa. Elasticity measurements on fibroblast actin cortex showed that the Poisson ratio is about 0.422. Following46, the viscosity of the cytoplasm is taken as \(\nu =0.03\, \textrm{poise}\).
Discussion
Free expansion without disassembly
Measurements of F growth in a differential AFM assay were performed in30. The Listeria nucleation promotion factor, ActA, was non-specifically adsorbed onto one cantilever, initiating the formation of a branched F-network between the cantilever and a nearby surface. Network growth was monitored through epifluorescence imaging of labeled actin. Allowing the growing actin network to deflect the cantilever, a compressive state was induced in the F-network. The relationship between the F growth velocity and the compressive forces was measured and reported in Fig. 2 in30.
Before entering a stall phase, the network length increases in a load-independent manner. Although velocities (\(85 \pm 68 \; \text{nm \, min}^{-1}\)) varied significantly among trials, it is reasonable to expect that their trend will remain unchanged at small loads, even at no loads whatsoever. Such a condition indeed occurs when the F-network is not yet in contact with the glass surface and is developing without mechanical opposition. In view of Eq. (14),
it follows that: (i) the flux \(\vec {H}_F\) vanishes, (ii) the reference configuration remains unchanged over time time, (iii) the current configuration is stress-free and materializes the idealized intermediate configuration, and (iv) the mass balance (2b) reduces to an ordinary differential equation in time. If G monomers are replenished at infinite velocity in the current configuration, i.e., \(c_G=c_G^0\), and disassembly is neglected, Eq. (2b) simplifies as
Equation (19) can be solved for the concentration \(c_F( \vec {x},t)\) in the current configuration, see Appendix B. At all places \(\vec {x}\) in which \({{\mathcal {C}}}( \vec {x},t) =1\) at all times, and assuming that no network exists at \(t=0\), i.e., \(c_F({\vec {x}}, 0) =0\), \(c_F\) yields
According to Eq. (20): (i) \(c_F\) is limited and its asymptotic value is
(ii) the higher \({k}^*_{f} \, c_G^0 \, \Omega _F > 0\) the faster the convergence to a steady-state condition, in the sense that \({ \textrm{d} {c_{F}} }/{ \textrm{d} t}\) = 0—see Fig. 3.
The limit concentration \(c_{\infty }\) is the uniform concentration of F in ideal conditions, when the full supply of G is granted and no mechanics affects the polymerization (i.e., at stress free conditions). It might be considered the ideal result of the polymerization of a mole of G and the definition of molar volume \(\Omega _F\) for the network shall refer to the \(c_{\infty }\) concentration. In view of Eq. (17), at all places \(\vec {x}\) in which \({{\mathcal {C}}}( \vec {x},t) <1\) the ideal concentration \(c_{\infty }\) is an upper bound for \(c_F\).
The referential concentration can easily be inferred from \(c_{F_R}({\vec {X}}, t) = J^c \, {c_F}({\vec {x}}({\vec {X}},t), t)\), i.e.,
showing that \(c_{F_R}\) is unbounded as \(c_F \rightarrow c_{\infty }\). Furthermore, in view of Eq. (7), the hypothesis of free expansion implies
and
upon simple boundary conditions on displacements. Accordingly: (i) the F-network growth velocity is achieved at the length-scale \(\ell >0\), where it holds \(v_F := \ell \, {k}^*_{f} \, c_G^0 \, \Omega _F \, / 3\); (ii) the pullback of the length scale \(\ell\) in the reference configuration becomes smaller and smaller with time, as described by
This eventually generates a boundary-layer problem, which may justify interface-based approaches such as the ones in49,50; (iii) in these approaches, one may assume that the velocity jump at the cell membrane or bacteria surface amounts to \(v_F\), although the evidence that the actin network is attached to the nucleation loci may induce lateral constraints and localized mechanical stresses.
The influence of mechanics on the F growth
Free expansion is an extremely rare event. In cellular motility, the cytoskeletal structure mechanically deforms to transmit forces to the extracellular matrix through focal adhesions. In bacterial motion, the viscous drag of the cytosol is a necessary constraint for motility, as well as a source of stress in the network36, acting against external forces that resist the forward progress of the pathogen.
The departure from ideal conditions is attributed to two competing effects: the shortage of G-actin, captured through the mass balance (2a), and the influence of mechanics on the F growth. This influence may manifest in a stress dependency of the reaction rate constants34,35, resulting in the densification of the polymer network. Mechanical stresses may promote a reorganization of actin filaments and their cross-linkers in the network, such as filamin, allowing \(c_F({\vec {x}}, t)\) to exceed the ideal limit \(c_{\infty }\) even when the signaling function is below unity, as indicated in (17). This speculation is confirmed by numerical simulations obtained through the implementation of the weak form of the governing equations in a finite element code, via the high-performance computing open library deal.ii51. Details on the implementation and numerical analysis will be provided in a forthcoming publication.
Figure 4 presents the numerical simulation of the Listeria monocytogenes comet tail growth. According to the findings in32,33 (see also21,52 and references therein), the tail is firmly attached to the bacterium. Therefore, imposing continuity of displacements for the F-network and the bacterium at its surface pole as a boundary condition for Eq. (8) is an appropriate choice. The bacterium’s surface is represented as the circular boundary of the gray area in Fig. 4a. A full restraint applied at the bacterium surface to the F-network promotes the evolution of a stress field, as depicted in Fig. 4a, which in turn affects the concentration \(c_F\). Figure 4a also shows that the formation of the comet tail aligns well with the images of actin polymerization in Listeria monocytogenes provided in22,53. Figuure 4b depicts the evolution of actin concentration over time at point A on the bacterium’s pole surface, comparing it with the free expansion curve in Fig. 3a, which corresponds to \(\Omega _F=0.25\) \(\hbox {m}^{3}\) \(\hbox {moles}^{-1}\) , \({k}^*_{f} = 0.05\) \(\hbox {s}^{-1}\).
The growth of the F-network on an AFM cantilever, as described in the previous section, has been simulated by restricting all lateral expansions, resembling the “soap effect” conditions described in22. Figure 5 depicts the outcomes of such simulation. A proliferation of \(c_F\) is seen at the tip of the network, see Fig. 5a. Figure 5b shows the time evolution of the velocity at the tip of the network, comparing it with the free expansion curve in Fig. 3a for \(\Omega _F=0.25\) \(\hbox {m}^{3}\) \(\hbox {moles}^{-1}\) , \({k}^*_{f} = 0.05\) \(\hbox {s}^{-1}\). It can be observed that the network velocity, following an initial rapid increase, aligns well with the data reported in30.
Overcoming the ideal limit, as reported in Figs. 4b and 5a, might either be a purely elastic event or promote a reorganization of the cross-linkers. This effect may alter the mesh size of the F-network. According to3,54, “the mechanics of the actin network is defined by its mesh size, which is variously taken as the spacing between actin filaments within the network or the distance between fixed cross-link points”. There is evidence that the mesh size strongly affects the elastic modulus32. The compressive stress, the relative concentration \(c_F - c_\infty\), the cell phenotype, and the nature of the polymer cross-linkers may affect mesh densification. To include this feature in the model, an internal variable related to the Helmholtz free energy is suggested, and studies in this regard are ongoing.
(a) Transient evolution of \(c_F\) in time, for \(\Omega _F=0.25\) \(\hbox {m}^{3}\) \(\hbox {moles}^{-1}\) , \({k}^*_{f} = 0.05\) \(\hbox {s}^{-1}\), and \(\, c_G^0 = 2.42\) moles \(\hbox {m}^{-3}\); (b) Evolution of the network velocity in time, for \(\Omega _F=0.25\) \(\hbox {m}^{3}\) \(\hbox {moles}^{-1}\) , \({k}^*_{f} = 0.05\) \(\hbox {s}^{-1}\), and \(\, c_G^0 = 2.42\) moles \(\hbox {m}^{-3}\). After an initial rapid development, the velocity is in agreement with the data, highlighted by the shaded strip. Data reported in30 correspond to the violet band (\(85 \pm 68 \mathrm{nm/min}\)).
Final remarks
Research conducted by various laboratories has advanced our understanding to the extent that we now possess knowledge of the majority of crucial molecules implicated in regulating actin at the forefront of cells. Nevertheless, despite having a comprehensive list of molecules, we have yet to formulate the precise set of rules necessary for generating a mobile cell, a lamellipodium, or even a comet tail at a continuum scale. This paper suggests that the force in ABM is generated by mechanical swelling in a narrow zone around the nucleation loci, which, in turn, affects the chemical potential and the kinetics of actin polymerization according to Eqs. (14) and (19).
To prove this claim, a chemo-transport-mechanics model for ABM has been formulated using a rigorous thermodynamic approach. The fundamental idea is to associate a volumetric increment with the organization of G monomers into the F-network in the cytosol. While we recognize that further improvements are necessary, we believe that our conceptual framework paves the way for quantitative investigations that can help interpret biochemical experimental outcomes and address key challenges in cellular mechanobiology, such as tumor metastasis, which, given their timescale, can hardly be followed in vivo.
The model is characterized by several assumptions and limitations. We defer the multiscale and multiphysics description of the energetics of nucleation, polymerization, and branching of actin filaments in the F-network to further studies. All biological events have been condensed into a single activation signal function \({{\mathcal {C}}}({\vec {x}},t)\) , which encompasses the activity of several proteins at the nucleation loci. We do not account for ADF and cofilin; instead, we consider the average lifetime of filaments in the cellular domain to model network disassembly.
Since the distribution of filaments in the lamellum and in the comet tail does not show a preferential direction12,55, we have chosen the volume as the mechanical descriptor of swelling, without accounting for anisotropy in the multiplicative decomposition of the deformation gradient. However, it could be objected that the isotropy of the network may lead to an isotropic mechanical response (i.e. an isotropic choice for \(\psi _R^{el}({{\textbf {E}}^e} )\)) but not necessarily to an isotropic swelling. This concept would indeed emerge if the polymerization resulted in surface accretion, with its own growth direction \(\vec {n}\). To account for specific orientations \(\vec {n}\) of the swelling, a polymerization tensor of the form
would replace the isotropic form (7). The consequences of such a choice will discussed in further publications. By the way, note that definition (26) would be mandatory for filopodia.
We believe that our model could shed light on one of the several fundamental mechanistic inquiries that remain unresolved at the continuum scale. The model permits any specification of the Helmholtz free energy. Since the focus of this note is the ABM engine, we arbitrarily and simplistically consider an elastic strain energy of the Saint-Venant kind in the numerical simulations. Viscosity is purposely neglected, as it is argued that the cross-linking rearrangement in the F-actin network has insufficient time to develop3. We also neglect the dependence of material parameters on F-actin concentration, although we are well aware that the mechanical behavior of the network changes during its evolution30; focused studies are required to capture the complex response of the network to external loads.
Translating the engine of actin-based motility depicted in this note into a mechanism for complete cell motility demands a notable increase in complexity, primarily due to the inclusion of a plasma membrane and the contractile machinery at the rear of the cell. Additionally, thoroughly understanding cell motility will necessitate comprehending how the regulation of cell adhesion and nuclear translocation is coordinated with lamellipodial protrusion to facilitate the movement of the entire cell. Motility relies not just on the overall mechanical behavior of the cell but also on various proteins that regulate the extension and detachment of the cell’s cytoskeletal structure and its interaction with the surrounding environment56,57. Several studies, either based on stochastic network models58 or on deterministic force balance at the leading edge—without neglecting forces acting on different parts of the cell, including the lamella59—have been proposed recently.
It is a reality that the free-expansion F-network velocities measured in the literature are extremely scattered. To cite a few examples, paper30 reports \(85 \pm 68 \mathrm{nm/min}\), paper41 reports \(74 \pm 50 \mathrm{nm/s} = 4.4 \pm 0.3 {\upmu }\mathrm{m/min}\), while paper42 reports \(2 {\upmu }\mathrm{m/min}\). The three measurements differ significantly from one another. It is likely that the experimental conditions vary considerably, and that the velocity at zero force is very sensitive to these conditions.
This evidence does not diminish the significance of the proposed model, but rather highlights that experiments and models are far from reaching conclusive results on this fascinating topic. Further developments, including a thorough investigation of the mechanobiology of membrane resistance, are essential for simulating cellular motility60, the cell-to-cell spread of bacteria through protrusion- and vesicle-mediated transfer, and the life cycles of intracellular bacteria that utilize actin-based motility61.
Data availibility
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Borisy, G. G. & Svitkina, T. M. Actin machinery: Pushing the envelope. Curr. Opin. Cell Biol. 12(1), 104–112 (2000).
Pantaloni, D., Le Clainche, C. & Carlier, M.-F. Mechanism of actin-based motility. Science 292(5521), 1502–1506 (2001).
Blanchoin, L., Boujemaa-Paterski, R., Sykes, C. & Plastino, J. Actin dynamics, architecture, and mechanics in cell motility. Physiol. Rev. 94(1), 235–263 (2014).
Marston, D. J. & Goldstein, B. Actin-based forces driving embryonic morphogenesis in caenorhabditis elegans. Curr. Opin. Genet. Dev. 16(4), 392–398 (2006).
Barrasa-Ramos, S., Dessalles, C., Hautefeuille, M. & Barakat A. Mechanical regulation of the early stages of angiogenesis. J. R. Soc. Interface19(20220360) (2022).
Stuelten, C. H., Parent, C. A. & Montell, D. J. Cell motility in cancer invasion and metastasis: Insights from simple model organisms. Nat. Rev. Cancer 18(5), 296–312 (2018).
Acharya, D. et al. Actin cytoskeleton remodeling primes RIG-I-like receptor activation. Cell 185(19), 3588–360221 (2022).
Dos Remedios, C. G. & Thomas, D. D. (eds) Molecular Interactions of Actin: Actin-Myosin Interaction and Actin-Based Regulation (Springer, Berlin, 2001).
Cameron, L. A., Giardini, P. A., Soo, F. S. & Theriot, J. A. Secrets of actin-based motility revealed by a bacterial pathogen. Nat. Rev. Mol. Cell Bio. 1(2), 110–119 (2000).
Choe, J. E. & Welch, M. D. Actin-based motility of bacterial pathogens: Mechanistic diversity and its impact on virulence. Pathog. Dis. 74(8), ftw0919 (2016).
Kühn, S. & Enninga, J. The actin comet guides the way: How listeria actin subversion has impacted cell biology, infection biology and structural biology. Cell Microbiol. 22(4), e13190 (2020).
Svitkina, T. The actin cytoskeleton and actin-based motility. Csh. Perspect. Biol. 10(1), a018267 (2018).
Marchand, J. B. et al. Actin-based movement of Listeria monocytogenes: Actin assembly results from the local maintenance of uncapped filament barbed ends at the bacterium surface. J. Cell Biol. 130(2), 331–343 (1995).
Hu, L. & Papoian, G. A. Mechano-chemical feedbacks regulate actin mesh growth in lamellipodial protrusions. Biophys. J. 98(8), 1375–1384 (2010).
Ni, H. & Papoian, G. A. Membrane-medyan: Simulating deformable vesicles containing complex cytoskeletal networks. J. Phys. Chem. B 125(38), 10710–10719 (2021).
Carlsson, A. E. Growth of branched actin networks against obstacles. Biophys. J . 81(4), 1907–1923 (2001).
Carlsson, A. E. Growth velocities of branched actin networks. Biophys. J . 84(5), 2907–2918 (2003).
Lin, Y., Shenoy, V. B., Hu, B. & Bai, L. A microscopic formulation for the actin-driven motion of listeria in curved paths. Biophys. J. 99(4), 1043–1052 (2010).
John, K., Caillerie, D. & Misbah, C. Spontaneous polarization in an interfacial growth model for actin filament networks with a rigorous mechanochemical coupling. Phys. Rev. E 90, 052706 (2014).
John, K., Peyla, P., Kassner, K., Prost, J. & Misbah, C. Nonlinear study of symmetry breaking in actin gels: Implications for cellular motility. Phys. Rev. Lett. 100, 068101 (2008).
Mogilner, A. & Oster, G. Force generation by actin polymerization II: The elastic ratchet and tethered filaments. Biophys. J . 84(3), 1591–1605 (2003).
Gerbal, F., Chaikin, P., Rabin, Y. & Prost, J. An elastic analysis of listeria monocytogenes propulsion. Biophys. J . 79(5), 2259–2275 (2000).
Egan, P., Sinko, R., LeDuc, P. R. & Keten, S. The role of mechanics in biological and bio-inspired systems. Nat. Commun. 6(1), 7418 (2015).
Gong, B., Wei, X., Qian, J. & Lin, Y. Modeling and simulations of the dynamic behaviors of actin-based cytoskeletal networks. ACS Biomater. Sci. Eng. 5(8), 3720–3734 (2019).
Gurtin, M. E., Fried, E. & Anand, L. The Mechanics and Thermodynamics of Continua (Cambridge University Press, Cambridge, 2010).
Humphrey, J. D. Constrained mixture models of soft tissue growth and remodeling—twenty years after. J. Elasticity 145(1–2), 49–75 (2021).
Humphrey, J. D. & Rajagopal, K. R. A constrained mixture model for growth and remodeling of soft tissues. Math. Mod. Meth. Appl. S 12, 407–430 (2002).
Kimpton, L. S., Whiteley, J. P., Waters, S. L., King, J. R. & Oliver, J. M. Multiple travelling-wave solutions in a minimal model for cell motility. Math. Med. Bio. 30(3), 241–272 (2012).
Kimpton, L. S., Whiteley, J. P., Waters, S. L. & Oliver, J. M. On a poroviscoelastic model for cell crawling. J. Math. Biol. 70(1), 133–171 (2015).
Parekh, S. H., Chaudhuri, O., Theriot, J. A. & Fletcher, D. A. Loading history determines the velocity of actin-network growth. Nat. Cell Biol. 7(12), 1219–1223 (2005).
Higgs, H. Getting down to basics with actin. Nat. Cell Biol. 3(8), E189–E189 (2001).
Noireaux, V. et al. Growing an actin gel on spherical surfaces. Biophys. J . 278, 1643–1654 (2000).
Kuo, S. C. & McGrath, J. L. Steps and fluctuations of listeria monocytogenes during actin-based motility. Nature 407, 1026–1029 (2000).
Salvadori, A., McMeeking, R. M., Grazioli, D. & Magri, M. A coupled model of transport-reaction-mechanics with trapping. Part I-small strain analysis. J. Mech. Phys. Solids 114, 1–30 (2018).
Arricca, M. et al. A coupled model of transport-reaction-mechanics with trapping, Part II: large strain analysis. J. Mech. Phys. Solids 181, 105425 (2023).
Bonanno, C., Serpelloni, M., Arricca, M., McMeeking, R. M. & Salvadori, A. Actin based motility unveiled: How chemical energy is converted into motion. J. Mech. Phys. Solids 175, 105273 (2023).
Theriot, J. A. The polymerization motor. Traffic 1(1), 19–28 (2000).
Deshpande, V. S., McMeeking, R. M. & Evans, A. G. A bio-chemo-mechanical model for cell contractility. PNAS 103(45), 17064–17065 (2006).
Serpelloni, M., Arricca, M., Bonanno, C. & Salvadori, A. Modeling cells spreading, motility, and receptors dynamics: A general framework. Acta Mech. Sin. 37(6), 1013–1030 (2021).
Anand, L. A Cahn–Hilliard-type theory for species diffusion coupled with large elastic-plastic deformations. J. Mech. Phys. Solids 60(12), 1983–2002 (2012).
McGrath, J. L. et al. The force-velocity relationship for the actin-based motility of listeria monocytogenes. Curr. Biol. 13(4), 329–332 (2003).
Marcy, Y., Prost, J., Carlier, M. F. & Sykes, C. Forces generated during actin-based propulsion: A direct measurement by micromanipulation. PNAS 101(16), 5992–5997 (2004).
Holzapfel, G. Nonlinear Solid Mechanics: A Continuum Approach for Engineering (Wiley, Hoboken, 2001).
Aroush, D.R.-B. et al. Actin turnover in lamellipodial fragments. Curr. Biol. 27(19), 2963-29731.e4 (2017).
Kiuchi, T., Nagai, T., Ohashi, K. & Mizuno, K. Measurements of spatiotemporal changes in G-actin concentration reveal its effect on stimulus-induced actin assembly and lamellipodium extension. J. Cell Biol. 193(2), 365–380 (2011).
Mogilner, A. & Oster, G. Cell motility driven by actin polymerization. Biophys. J . 71(6), 3030–3045 (1996).
Quirion, F. & Gicquaud, C. Changes in molar volume and heat capacity of actin upon polymerization. Biochem. J. 295(3), 671–672 (1993).
Damioli, V., Salvadori, A., Beretta, G. P., Ravelli, C. & Mitola, S. Multi-physics interactions drive VEGFR2 relocation on endothelial cells. Sci. Rep.-UK 7(1), 16700 (2017).
Kiana Naghibzadeh, S., Walkington, N. & Dayal, K. Surface growth in deformable solids using an Eulerian formulation. J. Mech. Phys. Solids 154, 104499 (2021).
Abeyaratne, R., Puntel, E. & Tomassetti, G. Treadmilling stability of a one-dimensional actin growth model. Int. J. Solids Struct. 198, 87–98 (2020).
Arndt, D. et al. The deal.II library, version 9.5. J. Numer. Math. 31(3), 231–246 (2023).
Gerbal, F. et al. Measurement of the elasticity of the actin tail of listeria monocytogenes. Eur. Biophys. J. 29(2), 134–140 (2000).
Kocks, C., Hellio, R., Gounon, P., Ohayon, H. & Cossart, P. Polarized distribution of Listeria monocytogenes surface protein Acta at the site of directional actin assembly. J. Cell Sci. 105(3), 699–710 (1993).
Kawska, A. et al. How actin network dynamics control the onset of actin-based motility. PNAS 109, 14440–14445 (2012).
Svitkina, T. M. & Borisy, G. G. Arp2/3 complex and actin depolymerizing factor/cofilin in dendritic organization and treadmilling of actin filament array in lamellipodia. J. Cell Biol. 145(5), 1009–1026 (1999).
Hodge, N. & Papadopoulos, P. Continuum modeling and numerical simulation of cell motility. J. Math. Biol. 64(7), 1253–1279 (2012).
Serpelloni, M., Arricca, M., Bonanno, C. & Salvadori, A. Chemo-transport-mechanics in advecting membranes. Int. J. Eng. Sci. 181, 103746. https://doi.org/10.1016/j.ijengsci.2022.103746 (2022}.
Weichsel, J. & Schwarz, U. S. Two competing orientation patterns explain experimentally observed anomalies in growing actin networks. PNAS 107(14), 6304–6309 (2010).
Schreiber, C., Amiri, B., Heyn, J. C. J., Rädler, J. O. & Falcke, M. On the adhesion-velocity relation and length adaptation of motile cells on stepped fibronectin lanes. PNAS 118(4), e2009959118 (2021).
Doyle, A. D., Sykora, D. J., Pacheco, G. G., Kutys, M. L. & Yamada, K. M. 3D mesenchymal cell migration is driven by anterior cellular contraction that generates an extracellular matrix Prestrain. Dev. Cell 56(6), 826–841 (2021).
Shenoy, V. B., Tambe, D. T., Prasad, A. & Theriot, J. A. A kinematic description of the trajectories of listeria monocytogenes propelled by actin comet tails. PNAS 104(20), 8229–8234 (2007).
Acknowledgements
Authors express their gratitude to the Ferriera Valsabbia foundation and to Comipont for the generous donation given in order to fund studies in the field of Mechanobiology. This paper has been funded by the Mechanobiology research center at UNIBS (https://www.mechanobiology-unibs.it/) through the support of companies COMSOL, Copan and Antares Vision. Insightful discussions with Prof. V. Shenoy are gratefully acknowledged. Figure 1 is an artwork by Caterina Salvadori.
Author information
Authors and Affiliations
Contributions
All authors contributed to the paper. A.S. provided the paper conceptualization, all authors worked at the problem formulation, A.S. and M.S. coded in deal.ii whereas C.B .coded in MATLAB, A.S., R.M., and C.B. wrote the paper. A.S. and R.M. supervised the work of M.S. and C.B.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Appendices
Proof of Eq. (13)
From the multiplicative decomposition (6) and the assumption of isotropic chemical swelling (7), the deformation gradient reads
By definition of the right Cauchy-Green tensors \({\textbf {C}} = {{\textbf {F}}}^\textsf{T} {\textbf {F}}\), \({\textbf {C}}^e = {{\textbf {F}}^{e}}^\textsf{T} {\textbf {F}}^{e}\), the Green Lagrange tensors yield
Proof of Eq. (20)
Consider (19) and a fixed location \(\vec {x}\) at which \({{\mathcal {C}}}( \vec {x},t) =1\) at all times. It holds:
Under free expansion conditions, \(J=J^c\) and
(29) thus becomes at a fixed location \(\vec {x}\)
(20) comes out by direct integration of the ordinary differential Eq. (30) with homogeneous initial conditions for \(c_F\) at given location \(\vec {x}\).
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Salvadori, A., Bonanno, C., Serpelloni, M. et al. On the generation of force required for actin-based motility. Sci Rep 14, 18384 (2024). https://doi.org/10.1038/s41598-024-69422-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-69422-3