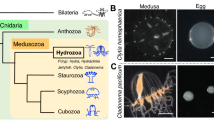
Abstract
Most host-parasite associations are explained by phylogenetically conservative capabilities for host utilization, and therefore parasite switches between distantly related hosts are rare. Here we report the first evidence of a parasitic spillover of the burrowing sea anemone Edwardsiella carnea from the invasive ctenophore Mnemiopsis leidyi to two scyphozoan hosts: the native Mediterranean barrel jellyfish Rhizostoma pulmo and the invasive Indo-Pacific nomad jellyfish Rhopilema nomadica, collected from the Eastern Mediterranean Sea. Edwardsiella carnea planulae found in these jellyfish were identified using molecular analyses of the mitochondrial 16S and nuclear 18S rRNA genes. Overall, 93 planulae were found on tentacles, oral arms, and inside of the gastrovascular canals of the scyphomedusae, whereas no infection was observed in co-occurring ctenophores. DNA metabarcoding approach indicated seasonal presence of Edwardsiella sp. in the Eastern Mediterranean mesozooplankton, coinciding with jellyfish blooms in the region. Our findings suggest a non-specific parasitic relationship between Edwardsiella carnea and various gelatinous hosts based on shared functionality rather than evolutionary history, potentially driven by shifts in host availability due to jellyfish blooms. This spillover raises questions about the ecological impacts of parasitism on native and invasive scyphozoan hosts and the potential role of Edwardsiella in controlling their populations.
Similar content being viewed by others
Introduction
Many marine sedentary organisms face significant challenges when dispersing in the ocean, especially if their pelagic stage is too short or lacks sufficient mobility1,2. Among these creatures are sea anemones (Actiniaria), globally distributed members of the class Anthozoa in the phylum Cnidaria. Unlike other classes in the phylum, anthozoans form polyps but lack the pelagic medusa stage. The only freely motile stage in their life cycle is a ciliated planula larva, which plays a crucial role in the dispersal of anthozoan populations3. The anemone planula is ephemerous—it exists for a short time, from a few days in the starlet sea anemone (Nematostella vectensis)4 to two months in table corals (Acropora digitifera)5. Together with a restricted gamete dispersion due to fertilization dynamics6, the short life span of the planula limits the anthozoan dispersal potential and settlement in areas distant from the parental polyps.
Several species of burrowing anemones from the family Edwardsiidae (genus Edwardsiella) cope with restricted dispersal by parasitizing planktonic ctenophores7,8,9,10. This interaction was previously described as facultative3,8. As in other Anthozoa, polyps of edwardsiids release gametes into the water, where fertilization occurs, and the zygotes develop into planulae. The free-swimming planula of Edwardsiella is pre-parasitic8 and can infect ctenophores either by burrowing through the epidermis7 or through the gastrodermal wall, after being ingested by the host11. Inside the host mesoglea, the parasitic planula grows and becomes elongated (vermiform). Its body is not differentiated into distinct regions, but a planula possesses a mouth and a pharynx8. The vermiform parasitic stage undergoes a body remodeling and generates a post-parasitic planula. This stage leaves the host and can re-infect other hosts, in which case it returns to the parasitic body plan, or can develop into a polyp and settle on the seafloor12. One host can harbor one or numerous parasitic edwardsiids7,8,9,10.
Ctenophore infection of Edwardsiella was first recorded in the warty comb jelly Mnemiopsis leidyi in the Northwest Atlantic in 1898 by Verrill13. However, he described the parasitic planula and adult polyp as different species: the polyp stage was assigned to Edwardsia lineata14 and the parasitic stage – to Edwardsiella lineata13. Later, Crowell showed that these two stages belong to one anemone species, Edwardsiella lineata, by rearing the larvae to the adult polyps7. Reitzel et al.12 sampled three ctenophore species in the Northwest Atlantic and found Edwardsiella lineata in two—M. leidyi and Beroe ovata. Edwardsiella lineata parasites were excised from both ctenophore species and exposed to uninfected ctenophores. Reitzel et al.12 found a strong preference of Edwardsiella to M. leidyi and concluded that M. leidyi is likely the only natural host. The presence of Edwardsiella inside B. ovata was explained by the predation of M. leidyi by B. ovata12.
The comb jelly M. leidyi is native to the western Atlantic and has spread over the past four decades to the Black, Azov, Caspian, Baltic and North seas15,16,17,18 . In the Mediterranean Sea, M. leidyi was first recorded in the western Aegean Sea in 199019, and by 2009, it had spread throughout the entire Mediterranean Sea20. Thus, it was not before 2007, one year after the first documentation of M. leidyi in the North Sea21, when Edwardsiella sp. was recorded in M. leidyi in the North Sea9.
Parasitic spillover happens when a parasite endemic to one host species infects a new host22,23. Such host switches can occur from invasive to native species in the invaded ecosystem24. As most host-parasite associations are explained by phylogenetically conservative capabilities for host utilization25, parasite switches between distantly related hosts were found to be infrequent26,27. Here, we report for the first time a parasitic spillover of Edwardsiella carnea from the ctenophore M. leidyi to two scyphozoan hosts—the native Mediterranean barrel jellyfish Rhizostoma pulmo, and the invasive Indo-Pacific nomad jellyfish Rhopilema nomadica. We applied molecular identification of Edwardsiella carnea planulae and described their morphology at successive developmental stages. In addition, we used DNA metabarcoding approach of bulk mesozooplankton samples to indicate the seasonality of the parasitic infection and discussed the potential implications of this spillover.
Results
In total, 86 planulae were found in four R. nomadica and 7 planulae in one R. pulmo (Fig. 1a, b). In both species of medusa, the parasitic planulae were located on the tentacles, oral arms, and in the subumbrella (Fig. 1c), and two individuals of R. nomadica contained planulae inside gastro-vascular canals. The first observations of planulae in R. nomadica were at the end of March 2024 (temperature 19.4 ºC, salinity 38.7 ppt). Later, in the middle of April 2024 (temperature 22.2 ºC, salinity 39.1 ppt), planulae were found in R. pulmo, when these medusae began to appear off the coast of Israel. None of the ctenophore specimens examined was infected with Edwardsiella carnea parasites.
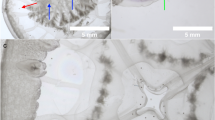
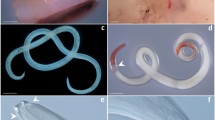
General view of scyphomedusa hosts Rhizostoma pulmo (a) and Rhopilema nomadica (b) and early stages of the Edwardsiella carnea parasitic planulae (c–g) that were found in them. (c). planula (black arrowhead) on the oral arm of R. nomadica. (d). planulae (pl) with developing septa (se) and vacuolated spheres that resembled lipid droplets (v). (e). spherical planula with mouth (mo). Box contents are detailed in panel (g). (f). vermiform planula with distinguishable mouth (mo) and aboral end (ab). Box contents are detailed in panel (h). (g). extensively ciliated (ci) epidermis of spherical planulae. (h). extensively ciliated (ci) epidermis of vermiform planula. um – umbrella (bell), oa – oral arms, tn – tentacles. Scale bar: 1 mm (c), 300 µm (d,e,f), 25 µm (g, h).
Morphology of planulae
Most planulae had slightly flattened spherical bodies, were semi-transparent, and were greyish/yellow (Fig. 1d, e). Diameter of the planulae ranged between 400 and 600 µm. All individuals had a mouth opening in the center of the body (Fig. 1e). In the planulae, two epithelial layers (epidermal and gastrodermal) and a thin layer of mesoglea were distinguishable. Epidermis and gastrodermis were extensively ciliated (Fig. 1g, h; Supplementary video 1). Two larger specimens (750 µm and 600 µm) had eight developing mesenteries (Fig. 1d). The gastrovascular cavity was filled with vacuolated spheres that resembled lipid droplets. Several planulae were found to have vermiform (worm-like) bodies ≥ 1.5 mm length and ≥ 360 µm diameter (Fig. 1f). Vermiform planulae had distinguishable oral end, and aboral end. In the lab, once excised from the medusae, the planulae swam freely in the seawater (Supplementary Material Video 2).
Molecular analyses
We compared the 16S rRNA and 18S rRNA gene sequences amplified from the DNA of the planulae collected in this study with available sequences of other Anthozoa in the NCBI. BLASTn search results showed that the highest identity percentage of our 16S rRNA and 18S rRNA gene sequences with the family Edwardsiidae (Nematostella, Edwardsia, Edwardsiella, Scolanthus, and Edwardsianthus). Among the family representatives, the highest identity of Edwardsiella sp. planulae sequences from R. nomadica and R. pulmo, with Edwardsiella carnea (99.25% and 99.06% respectively). The ML (Maximum Likelihood) trees based on the 16S (Fig. 2) and 18S (Fig. 3) rRNA gene sequences confirmed that the planulae from both medusae form a cluster within Edwardsiidae and belong to the genus Edwardsiella. In the 18S based ML tree, our sequences form a cluster with E. lineata and E. carnea (Fig. 3).
Maximum-likelihood phylogenetic tree of Edwardsiidae family based on the 16S rRNA gene, using the T92 + G substitution model. The outgroup Zoanthus kuroshiro was used as a root. The numbers near branches indicate the percentage of ML bootstrap support (1000 replicates) for nodes that received at least 60% support. The scale bar denotes the estimated number of nucleotide substitutions per site.
Maximum-likelihood phylogenetic tree of Edwardsiidae family based on the 18S rRNA gene, using the T92 + G substitution model. The outgroup Actinia equina was used as a root. The numbers near the branches indicate the percentage of ML bootstrap support (1000 replicates) for nodes that received at least 60% support. The scale bar denotes the estimated number of nucleotide substitutions per site.
Two 18S rRNA ASVs classified as Edwardsiidae (100% identity, BLASTn, deposited in GenBank as accession numbers PP955977 and PP955978) were found in the High-Throughput Sequences of the mesozooplankton samples collected from Hadera (0–26 m) in July 2020, and from Haifa Section stations H01 (0–50 m), H02 (150–100 m), and H04 (800–500 m) in August 2022. All duplicates (subsamples) fully corresponded, thus providing the same indication of the presence/absence of edwardsiid anemones. These sampling events co-occurred with blooms of R. nomadica.
Discussion
We assert that the parasitic planulae that we found in scyphomedusae belong to the burrowing anemone species Edwardsiella carnea. Dnyansagar et al.3 showed that both E. lineata and E. carnea are very close, phylogenetically-related species that are geographically isolated, and suggested that populations of these species could interbreed. Thus, we suggest that the planulae parasitizing scyphomedusae species can represent a Mediterranean population E. carnea. Our results show, for the first time, a spillover of the parasitic anemone Edwardsiella carnea from the ctenophore Mnemiopsis leidyi to two scyphozoan medusae, the Mediterranean-native barrel jellyfish Rhizostoma pulmo, and the invasive, Indo-Pacific nomad jellyfish Rhopilema nomadica (Fig. 4).
Life cycle and spillover of the burrowing sea anemone Edwardsiella carnea. (a). adult female and male polyps release gametes to the water column. (b). a free-swimming pre-parasitic planula is formed following fertilization. (c). the planula infects the ctenophore host Mnemiopsis leidyi and develops into vermiform parasitic stage. (d). a post-parasitic planula leaves the ctenophore host into the water column (e) where it can either settle in the seabed (f) and develop into a polyp (a) or reinfect another ctenophore or infect schyphozoan host Rhopilema nomadica or Rhizostoma pulmo (hypothetical spillover) (g-h). a post-parasitic planula may leave the schyphozoan host, traveling in the water column (i) where it can settle on the seabed (f) and develop into a polyp. The dashed line represents an alternate pathway.
The Edwardsiella carnea planulae that we found in these medusae were in early successive stages, including pre-parasitic and parasitic planulae, described here for the first time. The pre-parasitic stage included spherical planulae and planulae with developing septa, and the parasitic stage consisted of vermiform planulae. The developmental stage determination was based on the following morphological characters: extensive ciliation, as described by Reitzel et al.8; lack of body region differentiation and no differentiated siphonoglyph; and eight planula mesenteries. Crowell7 determined that the penetration of the planulae into the host is oral and forward through the epidermis and mesoglea, while Reitzel et al.11 found that the planulae infect the host via ingestion, through the gastrodermal wall. Here, we found Edwardsiella carnea planulae on the epithelium covering tentacles, on the oral arms, in the subumbrella and in the gastrovascular canals. Since the mesoglea in scyphozoans is thicker and denser than in ctenophores28,29,30,31, we propose that Edwardsiella planulae could not penetrate the medusa epithelium and mesoglea as burrowing through these structures is harder and requires more time. Hence, planulae were found on the epithelial surface and inside the gastrovascular canals of the medusae.
Three species of Edwardsiella are known in the Mediterranean Sea: E. carnea, E. janthina and E. loveni. Of them, only E. carnea was described as parasitic in Bolinopsis32, a high latitude distributed ctenophore, and later in M. leidyi9 in the Northeast Atlantic. To date, anemones from genus Edwardsiella as parasites were not reported in the Mediterranean Sea. Here we suggest that Edwardsiella carnea was introduced with its host M. leidyi from the East Atlantic waters to the Mediterranean Sea33, reaching the Southeastern Mediterranean. In our study, we found Edwardsiella carnea planulae exclusively in scyphomedusae but not in the ctenophores examined. This finding can be explained by parasite host-switch, which is not a rare phenomenon, yet it is often conservative within phylogenetically related taxa26,27. Both ctenophores and scyphozoans provide an ample pelagic substrate due to their relatively large size and frequency of appearance in blooms. Thus, it can be inferred that the choice of host by Edwardsiella carnea is non-specific and based on function rather than evolutionary history.
Over the past decades, the Eastern Mediterranean waters experienced an intensification of medusan blooms, mostly caused by the scyphozoan R. nomadica34,35,36. R. nomadica was first recorded in Israel in 1977 and considered a Lessepsian invader, introduced via the Suez Canal37. Since then, it has expanded its distribution westwards and was recorded in Lebanon, Syria, Turkey, Greece, Malta, Tunisia, Egypt, Sardinia, and Sicily34,38,39,40,41,42. This venomous species affects human health, fisheries, nutrient dynamics, coastal recreation and tourism, and pose a threat to coastal installations by clogging intake pipes43,44,45,46,47.The barrel-jellyfish R. pulmo is the most widespread scyphomedusa in the Mediterranean Sea, with prominent blooms from the western to the eastern basins48. In the Eastern Mediterranean, individuals of R. pulmo co-occurr with early summer swarms of R. nomadica34. During 2024, the Southeastern Mediterranean experienced consecutive blooms of the scyphomedusae R. nomadica and R. pulmo, while M. leidyi were less frequent and abundant. Since Edwardsiella carnea planulae were found in two scyphomedusa species but none in Mnemiopsis leidyi, it is possible that due to the shift in host availability, Edwardsiella carnea “spilled over” from ctenophores to scyphomedusae. As scyphomedusae are not feeding on ctenophores, this parasitic presence cannot be interpreted as an opportunistic predation, but rather as a parasitic host-switch.
In laboratory experiments, Edwardsiella lineata planulae excised from its ctenophore host quickly developed into a free-swimming planula larva, that could re-infect another host and assume the parasitic body plan11,12, or undergo settlement and develop into a free-living polyp11,12,49,50. The experimental manipulation was hypothesized to mimic the process that would occur in nature when the host dies, or when the parasite leaves the living host. Following these findings, it was concluded that the interaction between Edwardsiella lineata and M. leidyi is facultative8. Nonetheless, there is no evidence of direct development from planulae to polyps of Edwardsiella lineata or E. carnea in nature, without infecting a planktonic host. Our finding of a host-switch from ctenophores to scyphomedusae rather supports an obligate, yet non-specific interaction between Edwardsiella carnea and its diverse gelatinous hosts.
Parasites can have adverse impacts on their hosts, affecting their growth, reproduction, behavior, survival, and population dynamics51,52. Bumann & Puls53 studied the impacts of the burrowing parasitic anemone E. lineata on the host by comparing infested and non-infested M. leidyi. Non-infested ctenophores had higher growth rates than infested individuals, which had zero or negative growth rates, however no impact on egg production was found. They concluded that E. lineata could be partly responsible for the sharp decline of M. leidyi populations in autumn in US coastal waters, and recommended considering E. lineata as a biological control agent of the introduced M. leidyi populations. Chiaverano et al.54 reported edwardsiid-like organisms inside Aurelia sp. in Croatia. They concluded that parasitized medusae were significantly smaller and produced fewer but larger oocytes. Nonetheless, neither morphological nor molecular identification of the parasites was provided, thus this finding should be treated with caution.
Climate change can impact parasitic stages either directly or indirectly, by affecting the host55. The effects of temperature and salinity on the survival and development of vermiform parasitic E. lineata were experimentally tested56. Survival was impaired at temperatures above 30 ºC and salinity below 6 ppt. These thresholds do not appear to impede the colonization of the Edwardsiella carnea parasites in the Eastern Mediterranean Sea. Our metabarcoding data from mesozooplankton samples indicated the presence of Edwardsiella carnea in the water column of the Eastern Mediterranean Sea during summer months, when sea surface temperatures exceeded 31 ºC, and salinity was > 40 ppt. R. nomadica medusae are able to endure summer maxima beyond 31 ºC34, while R. pulmo medusae prevail in regions with winter minima below 15 ºC57, ensuring a widespread distribution of Edwardsiella carnea via these native and invasive scyphomedusan hosts.
Further research is required to understand the geographical and taxonomic extent of the parasitic infection of scyphomedusae by Edwardsiella carnea anemone and underpin its host-switch dynamics. Experiments are needed to assess the potential impact of this interaction on scyphozoan growth, reproduction, behavior, and survival. Future climate change may contribute to additional spillover of Edwardsiella carnea among scyphozoans, while potentially controlling their populations.
Methods
Sample collection and examination
The scyphomedusae Rhizostoma pulmo and Rhopilema nomadica and the ctenophores Mnemiopsis leidyi and Beroe ovata were sampled in the Israeli Southeastern Mediterranean Sea at the following sites: Hadera (32.46527°N, 34.861567°E), Mikhmoret (32.424963°N, 34.848131°E and 32.411922°N, 34.835943°E), and Shikmona (32.826474°N, 34.956926°E). Animals were sampled intact at 0-1m depths using scoop nets and buckets. Specimens were collected between December 2023 and May 2024. After collection, the medusae and ctenophores were brought to the lab at the National Institute of Oceanography, Israel Oceanographic and Limnological Research (IOLR), for morphometric analysis. For each medusa/ctenophore individual, weight and umbrella diameter/oral-aboral length were measured.
A total of 53 medusae (36 individuals of R. nomadica and 17 of R. pulmo) and 59 ctenophores (16 M. leidyi and 43 B. ovata) were collected and analyzed alive for the presence of parasites. Whole animals (medusae and ctenophores) were inspected visually, and later individuals were dissected by scalpel into parts the size of a petri dish for investigation under a stereomicroscope (SZX16, Olympus, Japan). The presence of planulae in each part was evaluated under a stereomicroscope. Morphology of the detected planulae was subsequently studied using a light microscopy and phase-contrast imaging (BX43, Olympus, Japan).
Mesozooplankton samples were collected monthly in Hadera and biannually offshore Haifa (Haifa Section) in the framework of the Israeli National Monitoring Program by IOLR. The samples were collected by vertical hauls of WP2 net (Ø = 57 cm, Hydro-Bios, Germany) in the coastal stations (depth 15–60 m), and by MultiNet Midi (50 × 50 cm, Hydro-Bios, Germany) in the offshore stations (100–1500 m depth). The samples were kept at –20 °C pending molecular analysis.
Sampling locations and dates are detailed in Supplementary Table S1.
DNA extraction, amplification and sequencing
After removing anemone planulae from the medusae, the planulae were individually preserved in absolute ethanol for molecular analysis. Total genomic DNA was extracted from five planulae using InviSorb Spin Tissue Mini Kit (Invitek Diagnostics, Germany) according to the manufacturer’s specifications. The amplifications of the following ribosomal RNA genes were performed: nuclear 18S rRNA gene with primers #3F (5’ GYGGTGCATGGCCGTTSKTRGTT 3’)58 and 9R (5’ GATCCTTCCGCAGGTTCACCTAC 3’)59, mitochondrial 16S rRNA gene with primers 16S-L (5' GACTGTTTACCAAAAACATA 3')60 and Aa_H16S_1541H (5' AGATTTTAATGGTCGAACAGAC 3')61. Reaction conditions for 18S rRNA gene amplification were as follows: 94 °C for 2 min, followed by 34 cycles of 94 °C for 15 s, 49°C for 30 s, and 72 °C for 1 min, and a final elongation step of 72 °C for 7 min. Reaction conditions for 16S rRNA gene amplification were as follows: 95°C for 5 min followed by 35 cycles of 95°C for 1 min, 45°C for 1 min, and 72°C for 1 min, and a final elongation step of 72°C for 10 min. Obtained PCR products were separated on 1.5% agarose gel and stained with GelRed (Biotium Inc., USA). Purification and Sanger sequencing of the PCR products were performed by Hy Laboratories Ltd. (Rehovot, Israel).
Frozen mesozooplankton samples were thawed and processed in duplicates following the protocol described in Guy-Haim et al.62. The 18S rRNA gene V9 region (ca. 192 bp) was amplified using 1391F-EukBr primer set63 amended with CS1/CS2 tags. Library preparation from the PCR products and Next Generation Sequencing (NGS) of 2 × 150 bp Illumina MiniSeq reads were performed by Hy Laboratories Ltd. (Israel).
Bioinformatic and phylogenetic analyses
A total of twelve 16S rRNA gene sequences of Hexacorallia were analysed, including two sequences of Edwardsiella carnea planulae obtained in this study, and nine sequences of Edwardsiidae from NCBI GenBank (https://www.ncbi.nlm.nih.gov/genbank/). A sequence of Zoanthus kuroshiro (AB219189) was used as an outgroup. A total of fifteen 18S rRNA gene sequences of Actinaria were analysed, including two sequences of Edwardsiella sp. planula obtained in this study, and thirteen sequences of Edwardsiidae from NCBI GenBank. A sequence of Actinia equina (AJ133552) was used as an outgroup. The sequences generated in this study were deposited in GenBank under the accession numbers PP874669 and PP958816 (18S), and PP955979-PP955980 (16S).
Sequence alignments were conducted using ClustalW embedded in MEGA v11.064. The best-fitting substitution model was selected according to the Bayesian information criterion using ML model selection in MEGA. ML analyses were performed using the T92 + G model with 1000 bootstrapping replicates.
The NGS demultiplexed paired-end reads were processed in the QIIME2 V2022.2 environment65. Reads were truncated based on quality plots, checked for chimeras, merged, and grouped into amplicon sequence variants (ASVs) with DADA266, as implemented in QIIME2. The 18S rRNA amplicons were classified with a scikit-learn classifier that was trained on the Silva 138 database or BLAST against the Silva 138 and PR2 databases (0.9 minimum identity cutoff).
Data availability
Sequence data that support the findings of this study have been deposited in the NCBI GenBank with the accession numbers: PP874669 and PP958816 (18S), PP955979-PP955980 (16S), PP955977- PP955978 (18S v9).
References
Shanks, A. L. Pelagic larval duration and dispersal distance revisited. Biol. Bull. 216, 373–385 (2009).
Guy-Haim, T., Hyams-Kaphzan, O., Yeruham, E., Almogi-Labin, A. & Carlton, J. T. A novel marine bioinvasion vector: Ichthyochory, live passage through fish. Limnol Oceanogr Lett 2, 81–90 (2017).
Dnyansagar, R. et al. Dispersal and speciation: The cross Atlantic relationship of two parasitic cnidarians. Mol Phylogenet Evol 126, 346–355 (2018).
Technau, U. & Steele, R. E. Evolutionary crossroads in developmental biology: Cnidaria. Development 138, 1447–1458 (2011).
Nishikawa, A. & Sakai, K. Settlement-competency period of planulae and genetic differentiation of the scleractinian coral Acropora digitifera. Zoolog Sci 22, 391–399 (2005).
Marshall, D. J., Semmens, D. & Cook, C. Consequences of spawning at low tide: Limited gamete dispersal for a rockpool anemone. Mar Ecol Prog Ser 266, 135–142 (2004).
Crowell, S. An edwardsiid larva parasitic in Mnemiopsis. In Coelenterate ecology and behavior (ed. Mackie, G. O.) 247–250 (Plenum Press, New York, 1976).
Reitzel, A. M., Daly, M., Sullivan, J. C. & Finnerty, J. R. Comparative anatomy and histology of developmental and parasitic stages in the life cycle of the lined sea anemone edwardsiella lineata. J. Parasitol. 95, 100–112 (2009).
Selander, E., Møller, L. F., Sundberg, P. & Tiselius, P. Parasitic anemone infects the invasive ctenophore Mnemiopsis leidyi in the North East Atlantic. Biol. Invasions 12, 1003–1009 (2009).
Östman, S. C. & Friis Møller, L. Gross morphology and cnidae of edwardsiella anemones and larvae (Anthozoa, Edwardsiidae) from the Swedish West Coast. Acta Zoologica 102, 467–482 (2021).
Reitzel, A. M., Sullivan, J. C. & Finnerty, J. R. Qualitative shift to indirect development in the parasitic sea anemone Edwardsiella lineata. Integr. Comp. Biol. 46, 827–837 (2006).
Reitzel, A. M. et al. Ecological and developmental dynamics of a host-parasite system involving a sea anemone and two ctenophores. J. Parasitol. 93, 1392–1402 (2007).
Verrill, A. E. Descriptions of new American actinians, with critical notes on other species. Am. J. Sci. 6, 493–498 (1898).
Verrill, A. E. Report upon the Invertebrate Animals of Vineyard Sound and Adjacent Waters, with an Account of the Physical Features of the Region. Report upon the invertebrate animals of Vineyard sound and adjacent waters, with an account of the physical features of the region. (1873).
Harbison, G. R. & Volovik, S. P. The ctenophore, Mnemiopsis leidyi, in the Black Sea: a holoplanktonic organism transported in the ballast water of ships. Nonindigenous Estuarine and Marine Organisms (NEMO) 25–36 (1994).
Shiganova, T. A. et al. The invader into the Caspian Sea ctenophore Mnemiopsis and its initial effect on the pelagic ecosystem. Oceanology (Wash D C) 41, 517–524 (2001).
Javidpour, J., Sommer, U. & Shiganova, T. First record of Mnemiopsis leidyi A. Agassiz 1865 in the Baltic sea. Aquat. Invasions 1, 299–302 (2006).
Faasse, M. A. & Bayha, K. M. The ctenophore Mnemiopsis leidyi A. Agassiz 1865 in coastal waters of the Netherlands: An unrecognized invasion?. Aquat. Invasions 1, 270–277 (2006).
Shiganova, T. A. et al. Population development of the invader ctenophore Mnemiopsis leidyi, in the Black Sea and in other seas of the Mediterranean basin. Mar. Biol. 139, 431–445 (2001).
Ghabooli, S. et al. Invasion pathway of the ctenophore Mnemiopsis leidyi in the Mediterranean Sea. PLoS One 8, 1–9 (2013).
Boersma, M., Malzahn, A. M., Greve, W. & Javidpour, J. The first occurrence of the ctenophore Mnemiopsis leidyi in the North Sea. Helgol. Mar. Res. 61, 153–155 (2007).
Daszak, P., Cunningham, A. A. & Hyatt, A. D. Emerging infectious diseases of wildlife - Threats to biodiversity and human health. Science 1979(287), 443–449 (2000).
Goedknegt, M. A. et al. Parasites and marine invasions: Ecological and evolutionary perspectives. J. Sea Res. 113, 11–27. https://doi.org/10.1016/j.seares.2015.12.003 (2016).
Chalkowski, K., Lepczyk, C. A. & Zohdy, S. Parasite ecology of invasive species: Conceptual framework and new hypotheses. Trends Parasitol. 34, 655–663 (2018).
Hoberg, E. P. & Brooks, D. R. A macroevolutionary mosaic: Episodic host-switching, geographical colonization and diversification in complex host-parasite systems. J. Biogeogr. 35, 1533–1550 (2008).
Gómez, J. M., Verdú, M. & Perfectti, F. Ecological interactions are evolutionarily conserved across the entire tree of life. Nature 465, 918–921 (2010).
Horká, I., De Grave, S., Fransen, C. H. J. M., Petrusek, A. & Duriš, Z. Multiple host switching events shape the evolution of symbiotic palaemonid shrimps (Crustacea: Decapoda). Sci. Rep. 6, 1–13 (2016).
Franc, S. Collagen of Coelenterates. In Biology of Invertebrate and Lower Vertebrate Collagens (eds Bairati, A. & Garrone, R.) 197–210 (Springer, US, Boston, MA, 1985).
Hernandez-Nicaise, M. L. Ctenophora. In Microscopic Anatomy of Invertebrates (eds Harrison, F. W. & Westfall, J. A.) 359–418 (Springer, US, 1991).
Lesh-Laurie, G. E. & Suchy, P. E. Cnidaria: Scyphozoa and Cubozoa. in Microscopic Anatomy of Invertebrates (eds. Harrison, F. W. & Westfall, J. A.) vol. 2 185–266 (1991).
Fidler, A. L. et al. Collagen IV and basement membrane at the evolutionary dawn of metazoan tissues. Elife https://doi.org/10.7554/eLife.24176 (2017).
Stephenson, T. A. British Sea Anemones (The Ray Society, 1935).
Bolte, S. et al. Population genetics of the invasive ctenophore Mnemiopsis leidyi in Europe reveal source-sink dynamics and secondary dispersal to the Mediterranean Sea. Mar Ecol Prog Ser 485, 25–36 (2013).
Edelist, D. et al. Phenological shift in swarming patterns of Rhopilema nomadica in the Eastern Mediterranean Sea. J. Plankton Res. 42, 211–219 (2020).
Edelist, D. et al. Tracking jellyfish swarm origins using a combined oceanographic-genetic-citizen science approach. Front. Mar. Sci. 9, 1–15 (2022).
Katsanevakis, S. et al. Impacts of invasive alien marine species on ecosystem services and biodiversity: A pan-European review. Aquat. Invasions 9, 391–423 (2014).
Galil, B., Spanier, E. & Ferguson, W. The Scyphomedusae of the Mediterranean coast of Israel, including two Lessepsian migrants new to the Mediterranean. Zoologische Mededelingen 64, 95–105 (1990).
Lotan, A., Fine, M. & Benhillel, R. Synchronization of the life cycle and dispersal pattern of the tropical invader Scyphomedusan Rhopilema nomadica is temperature dependent. Mar. Ecol. Prog. Ser. 109, 59–66 (1994).
Brotz, L. & Pauly, D. Jellyfish populations in the Mediterranean Sea. Acta Adriat. 53, 213–231 (2012).
Yahia, M. N. D. et al. The invasive tropical scyphozoan Rhopilema nomadica Galil, 1990 reaches the Tunisian coast of the Mediterranean Sea. Bioinvasions Rec. 2, 319–323 (2013).
El-regal, M. A. & Temraz, T. Blooming of nomad jelly fish Rhopilema nomadica along the Egyptian Mediterranean coasts. Rapport Comm. Int. Mer. Mediterrane 41, 490 (2016).
Madkour, F. F., Safwat, W. & Hanafy, M. H. Record of Aggregation of Alien Tropical Schyphozoan Rhopilema nomadica Galil, 1990 in the Mediterranean Coast of Egypt. Int. Marine Sci. J. 1, 1–7 (2019).
Angel, D. L., Edelist, D. & Freeman, S. Local perspectives on regional challenges: jellyfish proliferation and fish stock management along the Israeli Mediterranean coast. Reg. Environ. Change 16, 315–323 (2016).
Galil, B. Poisonous and Venomous: Marine Alien Species in the Mediterranean Sea and Human Health. in CAB International 2018. Invasive species and human health (eds. Mazza, G. & Tricarico, E.) (2018). https://doi.org/10.1079/9781786390981.0000.
Ghermandi, A., Galil, B., Gowdy, J. & Nunes, P. A. L. D. Jellyfish outbreak impacts on recreation in the Mediterranean Sea: Welfare estimates from a socioeconomic pilot survey in Israel. Ecosyst. Serv. 11, 140–147 (2015).
Guy-Haim, T. et al. The effects of decomposing invasive jellyfish on biogeochemical fluxes and microbial dynamics in an ultra-oligotrophic sea. Biogeosciences 17, 5489–5511 (2020).
Rahav, E. et al. Jellyfish swarm impair the pretreatment efficiency and membrane performance of seawater reverse osmosis desalination. Water Res. 215, 118231 (2022).
Fuentes, V. et al. Life cycle of the jellyfish Rhizostoma pulmo (Scyphozoa: Rhizostomeae) and its distribution, seasonality and inter-annual variability along the Catalan coast and the Mar Menor (Spain, NW Mediterranean). Mar. Biol. 158, 2247–2266 (2011).
Crowell, S. & Oates, S. Metamorphosis and reproduction by transverse fission in an edwardsiid anemone. in Developmental and cellular biology of coelenterates Proceedings of the 4th International Coelenterate Conference (eds. Tardent, P. & Tardent, R.) 139–142 (Elsevier, Amsterdam, 1980).
Daly, M. Taxonomy, anatomy, and histology of the lined sea anemone, Edwardsiella lineata (Verrill, 1873) (Cnidaria: Anthozoa: Edwardsiidae). Proc. Biol. Soc. Washington 115, 868–877 (2002).
Dobson, A. P. The population biology of parasite-induced changes in host behavior. Q. Rev. Biol. 63, 139–165 (1988).
Lafferty, K. D. & Kuris, A. M. How environmental stress affects the impacts of parasites. Limnol. Oceanogr. 44, 925–931 (1999).
Bumann, D. & Puls, G. Infestation with larvae of the sea anemone Edwardsia lineata affects nutrition and growth of the ctenophore Mnemiopsis leidyi. Parasitology 113, 123–128 (1996).
Chiaverano, L. M., Graham, W. M. & Costello, J. H. Parasites alter behavior, reproductive output, and growth patterns of Aurelia medusae in a marine lake. Mar. Ecol. Prog. Ser. 540, 87–98 (2015).
Byers, J. E. Marine parasites and disease in the era of global climate change. Ann. Rev. Mar. Sci. 13, 397–420 (2021).
Lee, J. et al. Temperature and salinity affect development of the parasitic sea anemone Edwardsiella lineata potentially limiting its impact as a biological control on the ctenophore Mnemiopsis leidyi. J. Parasitol. 109, 574–579 (2023).
Leoni, V., Bonnet, D., Ramírez-Romero, E. & Molinero, J. C. Biogeography and phenology of the jellyfish Rhizostoma pulmo (Cnidaria: Scyphozoa) in southern European seas. Global Ecol. Biogeogr. 30, 622–639 (2021).
Machida, R. J. & Knowlton, N. PCR primers for metazoan nuclear 18S and 28S ribosomal DNA sequences. PLoS One 7, e46180 (2012).
Giribet, G., Carranza, S., Baguñà, J., Riutort, M. & Ribera, C. First molecular evidence for the existence of a Tardigrada + Arthropoda clade. Mol. Biol. Evol. 13, 76–84 (1996).
Ender, A. & Schierwater, B. Placozoa are not derived cnidarians: Evidence from molecular morphology. Mol. Biol. Evol. 20, 130–134 (2003).
Bayha, K. M., Dawson, M. N., Collins, A. G., Barbeitos, M. S. & Haddock, S. H. D. Evolutionary relationships among scyphozoan jellyfish families based on complete taxon sampling and phylogenetic analyses of 18S and 28S ribosomal DNA. Integr. Comp. Biol. 50, 436–455 (2010).
Guy-Haim, T. et al. A new record of the rapidly spreading calanoid copepod Pseudodiaptomus marinus (Sato, 1913) in the Levantine Sea using multi-marker metabarcoding. Bioinvasions Rec. 11, 964–976 (2022).
Amaral-Zettler, L. A., McCliment, E. A., Ducklow, H. W. & Huse, S. M. A method for studying protistan diversity using massively parallel sequencing of V9 hypervariable regions of small-subunit ribosomal RNA Genes. PLoS One 4, 1–9 (2009).
Tamura, K., Stecher, G. & Kumar, S. MEGA11: Molecular evolutionary genetics analysis version 11. Mol. Biol. Evol. 38, 3022–3027 (2021).
Bolyen, E. et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 37, 852–857 (2019).
Callahan, B. J. et al. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
Acknowledgements
This study was funded by ISF grant no. 1655/21 to T.G.-H and the IOLR National Monitoring Program of the Israeli Mediterranean Sea. A.I. thanks the Bloom Scholarship for Doctoral Studies from the Bloom Graduate School, University of Haifa, Israel. We express our gratitude to Shai Mienis for providing underwater jellyfish photos. We thank Meduzot Ba’am jellyfish observation initiative (https://www.meduzot.co.il) and its Facebook group participants for providing up-to-date information on the location of jellyfish. We thank Dr. Jacob Douek for providing 16S primers. Finally, we thank the IOLR sea-going team and Dr. Zafrir Kuplik for their help with the sampling.
Author information
Authors and Affiliations
Contributions
A.I. and T.G.-H conceived the study, A.I. and A.R.M. collected and analysed the data, A.I. and T.G.-H interpret the findings and wrote original draft. T.G.-H. and D.A. supervised the research. All coauthors reviewed and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Iakovleva, A., Morov, A.R., Angel, D. et al. From ctenophores to scyphozoans: parasitic spillover of a burrowing sea anemone. Sci Rep 14, 20785 (2024). https://doi.org/10.1038/s41598-024-72168-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-72168-7