Abstract
Leukemia is a type of blood tumour that occurs because of abnormal enhancement in WBCs (white blood cells) in the bone marrow of the human body. Blood-forming tissue cancer influences the lymphatic and bone marrow system. The early diagnosis and detection of leukaemia, i.e., the accurate difference of malignant leukocytes with little expense at the beginning of the disease, is a primary challenge in the disease analysis field. Despite the higher occurrence of leukemia, there is a lack of flow cytometry tools, and the procedures accessible at medical diagnostics centres are time-consuming. Distinct researchers have implemented computer-aided diagnostic (CAD) and machine learning (ML) methods for laboratory image analysis, aiming to manage the restrictions of late leukemia analysis. This study proposes a new Falcon optimization algorithm with deep convolutional neural network for Leukemia detection and classification (FOADCNN-LDC) technique. The main objective of the FOADCNN-LDC technique is to classify and recognize leukemia. The FOADCNN-LDC technique utilizes a median filtering (MF) based noise removal process to eradicate the image noise. Besides, the FOADCNN-LDC technique employs the ShuffleNetv2 model for the feature extraction process. Moreover, the detection and classification of the leukemia process are performed by utilizing the convolutional denoising autoencoder (CDAE) model. The FOA is implemented to select the hyperparameter of the CDAE model. The simulation process of the FOADCNN-LDC approach is performed on a benchmark medical dataset. The investigational analysis of the FOADCNN-LDC approach highlighted a superior accuracy value of 99.62% over existing techniques.
Similar content being viewed by others
Introduction
Nowadays, Leukemia is one of the most popular kinds of blood cancer for all groups of people, especially for children1. This unusual problem is mainly affected by blood cell proliferation and young growth, which damages brain tissue, the immune system, and red blood cells. When to reproduce and die, a cell is directed by the genetic code. The modifications in gene appearance may result in defective instructions for cancer2. Protein construction is affected by cells due to the control of cells. A few types of genes change proteins to fix injured cells, which is associated with cancer3. Tumour sample classification and their subtypes assist in diagnosing and predicting dissimilar sorts of cancer. It aids in accurately predicting cancer types and recognizes sub-type-specific drug treatments. To handle such kind of challenges, the assessable study of diverse blood instances is implemented in the Computer-Aided-Diagnosis (CAD) technique, which is mainly proposed by utilizing both Deep Learning (DL) and ML models4. Many research studies exist in leukemic cancer detection is executed. In the past two years, most of the research studies employed ML and CAD techniques for laboratory image analysis to defeat the leukemia analysis and determine its subgroups5. Compared to the ML models, a different set of leukemic cell features can be removed by classification. Most researchers have recommended a segmentation procedure to remove exact features from the region of interest (RoI), which is segmented lymphocyte images6.
With the development of DL, numerous issues and challenges have been solved in image processing. These techniques used automated feature engineering. The most commonly utilized deep neural networks (DNNs) in computer vision (CV) are Convolutional Neural Networks (CNNs)7. The CNN has great generalization power, adaptability, and self-learning capability that are highly employed in IoT-based models and medical imaging issues. Conventional image identification approaches require handmade feature extraction and classification8. In several situations, an entire count of data instances is insufficient to test from the beginning for a CNN. In such conditions, transfer learning (TL) can be used to attain the probability of CNNs while reducing the cost of computing. Leukemia presents a crucial health challenge for children due to its impact on vital blood cell functions and overall immune system health7. Early and accurate detection is significant for effectual treatment and management. Advances in technology offer promising solutions for enhancing diagnostic accuracy and effectiveness. Employing modern computational techniques and optimization models can improve the precision of leukemia classification and detection precision, potentially leading to improved patient outcomes9.
This study proposes a new Falcon Optimization Algorithm with Deep Convolutional Neural Network for Leukemia Detection and Classification (FOADCNN-LDC) technique. The main objective of the FOADCNN-LDC technique is to classify and recognize leukemia. To accomplish this, the FOADCNN-LDC technique utilizes a median filtering (MF) based noise removal process to eradicate the noise in the images. Besides, the FOADCNN-LDC technique employs the ShuffleNetv2 model for the feature extraction process. Moreover, the detection and classification of the leukemia process are performed by utilizing the convolutional denoising autoencoder (CDAE) model. The FOA is implemented to select the hyperparameter of the CDAE model. The simulation process of the FOADCNN-LDC approach is performed on a benchmark medical dataset. The major contribution of the FOADCNN-LDC approach is listed below.
-
This FOADCNN-LDC model utilizes the MF method to improve the quality of the image by effectually reducing noise. This also enhances the clarity of images, which is significant for accurate analysis and subsequent processing. The improved image quality assists in more reliable outcomes in detecting and classifying leukemia.
-
The FOADCNN-LDC approach employs the ShuffleNetv2 method for feature extraction, presenting an advanced methodology to capture relevant features from cleaned images. This methodology improves the capability to detect key patterns and details required for precise leukemia detection. By employing ShuffleNetv2, the study attains a more accurate and effective feature extraction process.
-
The CDAE technique is implemented for the precise detection and classification of leukemia, substantially enhancing the accuracy of the diagnosis. This method improves the ability to distinguish between leukemia and non-leukemia cases, paving the way to more reliable outcomes. The robust classification capabilities provided by CDAE confirm more precise and efficient diagnosis of leukemia.
-
The FOA model is for fine-tuning the hyperparameters of the CDAE method, improving its performance and robustness. By optimizing these hyperparameters, FOA enhances the accuracy and efficiency of the technique in classifying leukemia. This results in a more reliable and efficient diagnostic tool.
-
The FOADCNN-LDC methodology incorporates the FOA model for hyperparameter selection in the CDAE method, presenting a novel technique that improves its accuracy and effectiveness. This technique enhances the performance of leukemia classification by optimizing hyperparameters more efficiently. The implementation of FOA presents a new dimension of precision in model tuning.
The article is structured as follows: Section “Related works presents the literature review, Section “The proposed model” outlines the proposed method, Section “Image processing” details the results evaluation, and Section “Feature extraction” concludes the study.
Related works
Veeraiah et al.10 proposed a Mayfly enhanced with the Generative Adversarial Network (MayGAN) model. Moreover, the Generative Adversarial System (GAS) is combined with PCA in the feature extraction method for sorting out blood cancer types in the data. The morphological process and semantic method employing geometric features are utilized to part the cells that form leukaemia. In11, the authors present an external CNN-based model. The developed solution consists of a CNN-based technique, which employs a segment called Efficient Channel Attention (ECA) with VGG16 to remove enhanced features from the image datasets. Numerous augmentation models are used to boost the amount and excellence of testing data. The authors12 present an IoMT-based architecture. In this system, the clinical plans are associated with the network websites with the support of the CC platform. These models are applied to recognize leukemia subtypes in the recommended frameworks, namely ResNet34 and DenseNet121. Jawahar et al.13 designed a DNN-based (ALNett) technique that uses depth-wise convolutional of various dilation amounts for classifying microscopic white blood cell images. In particular, the group of layers involves max-pooling and convolution. Then, the normalization procedure produces enriched contextual and structural details to remove global and robust local features from the microscopic images for a precise estimate of ALL. Atteia14 proposed a new hybrid DL technique. From the features extraction, GoogleNet and Inception-v3 CNNs, in a combination of datasets of atomic blood smear images, usually design the deep image features. A sparse AE model is specially developed to create an abstract set of important hidden features from the distended image feature dataset. The latent feature executes image classification using the Support Vector Machine (SVM) model.
In15, two stages of ANN combined with the PSO model are presented. The performance is compared with multiclass NN-PSO and Backpropagation Neural Networks classification (multiclass NN-BP). The authors16 designed a new DL technique based on CNN to analyze Acute Lymphoblastic Leukemia (ALL). This technique does not need FE or any pre-training on another database. Therefore, it can be employed to detect leukaemia in real-time applications. In17, a deep FS-based technique, ResRandSVM, is projected. The presented method employs seven DL methods for deep feature extraction. Then, the three FS techniques, namely Random Forest (RF), Principal Component Analysis (PCA), and Analysis of Variance (ANOVA), are utilized to extract essential and valuable features. Next, the selected feature map is fed to four classification methods to categorize the images into standard images and leukaemia. Ali and Mohammed18 aim to review and analyze cutting-edge research on AI techniques in omics data analysis, emphasize the potential merits and critical difficulties, and present unique insights into optimizing AI applications in this field. In19, a model is introduced by incorporating diverse omics data utilizing the Quantum Cat Swarm Optimization (QCSO) method for feature selection (FS), combined with K-means clustering and SVM classification, ultimately improving the performance and interpretability of the technique. Benameur et al.20 compare the heating performance of two microwave antennas with various geometrical apertures in hyperthermia treatment, employing thermal simulations and Bland-Altman evaluation to determine which antenna gives improved temperature accuracy and treatment effectiveness.
Hosseinzadeh et al.21 implement a ResNet-based feature extractor for detecting ALL, utilizing several TL methods, namely ResNet, VGG, EfficientNet, and DenseNet for extracting features. It then employs several feature selectors: Genetic Algorithm, PCA, ANOVA, RF, Univariate, Mutual Information, Lasso, XGB, Variance, and Binary Ant Colony. It finds that MLP classifiers attain the optimum performance. In22, an effective pipeline for classifying ALL subtypes is proposed. It starts with a novel neighbourhood pixel transformation using differential evolution to improve blood cell image clarity. A hybrid feature extraction methodology employs the InceptionV3 and DenseNet201 models for comprehensive feature sets. Custom binary Grey Wolf Algorithm (GWA) optimization reduces feature size while retaining critical information, improving classification accuracy across diverse classifiers. Noshad and Fallahi23 present a hybrid two-layer framework for FS. The first layer comprises image segmentation utilizing an improved First-Spike-based model with a Gaussian function, followed by feature extraction employing a deep residual architecture. The second layer implements the effectual JAYA optimization algorithm for FS, and a support vector machine (SVM) is utilized to classify ALL images. Shree and Logeswari24 introduce an optimized deep recurrent neural network (DRNN) methodology for leukemia detection using blood sample microscopy images. The method utilizes DRNNs for diagnosis, optimized by the red deer optimization algorithm (RDOA), which fine-tunes DRNN weights based on deer roaring rate behaviour.
The existing studies for leukemia detection face various limitations. Methodologies incorporating MayGAN with PCA may face difficulty with overfitting and fail to address several cancer discrepancies. CNN methods with ECA and VGG16 techniques might need to handle large or varied datasets effectively, while IoMT-based systems may face data privacy and connectivity issues. Hybrid DL techniques such as GoogleNet and Inception-v3 may encounter difficulty with scalability. Two-stage ANN and DL models optimized by evolutionary algorithms might suffer from convergence and tuning difficulties. These approaches often need to address real-world discrepancies or integration complexities fully. Research gaps encompass the requirement for more robust methods to generalize across diverse datasets, enhanced handling of privacy and scalability issues, and improved optimization models to manage complexity better and ensure consistency.
The proposed model
This manuscript develops a model for the automated classification and detection of leukemia on medical images. The main objective of the FOADCNN-LDC approach is to classify and recognize the occurrence of leukemia. To accomplish this, the FOADCNN-LDC approach comprises a series of processes: MF-based preprocessing, ShuffleNetv2 feature extraction, CDAE-based classification, and FOA-based hyperparameter tuning. Figure 1 exhibits the entire flow of the FOADCNN-LDC method.
Image processing
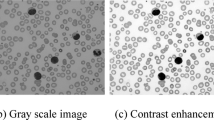
The MF technique is applied to preprocess the input images25. The MF approach was selected for image processing due to its efficiency in conserving edges while removing noise, which is significant for maintaining the integrity of medical images. Unlike other noise reduction techniques that may blur or distort important features, MF effectively removes salt-and-pepper noise and other random discrepancies without substantially affecting the image details. In the context of the FOADCNN-LDC technique, MF improves image clarity by mitigating noise, which directly affects the accuracy of leukemia detection. Cleaner images lead to more precise feature extraction and classification, enhancing the overall diagnostic precision compared to approaches that may introduce artefacts or reduce image quality. This makes MF a reliable choice for safeguarding high-quality input for subsequent evaluation. Figure 2 portrays the structure of the MF technique.
MF is a significantly utilized image processing method that supports decreased noise in images while maintaining edges and vital image features. It is primarily effective in eliminating salt-and-pepper noise, which is caused by arbitrarily removing bright and dark pixels from the image. The workings of MF are provided below.
Sliding Window: A lesser square or rectangular window (kernel/mask) can move with whole images. The window dimension is generally 3 × 3, 5 × 5, 7 × 7, or some other odd-sized square to ensure the centre of the pixel.
Pixel Sorting: The pixel values from the window can be arranged for every place in the window. Once the pixel values can be set in ascending order, the median value, which is the mid value, is chosen as the novel value for the centre pixel of the window.
Exchanging the Pixel: The median value can be allocated to the window’s centre pixel. This method can be repeated for all the pixels from the image.
Feature extraction
For the derivation of the feature vectors, the ShuffleNetv2 model is applied. Ma et al. presented four proposal guidelines for an effective network and enhanced ShuffleNet-Vl to present ShuffleNet-V226. The ShuffleNetv2 model was selected for feature extraction due to its effectual architecture, which balances performance and computational cost while maintaining high accuracy. Its lightweight design allows for rapid processing and mitigates the computational burden related to more convolutional techniques. This efficiency is beneficial when handling massive datasets or deploying models in resource-constrained environments. In the FOADCNN-LDC methodology, the capability of the ShuffleNetv2 model for extracting relevant features from cleaned images improves the technique’s overall performance. Its feature extraction process enhances the clarity and quality of input data, leading to improved subsequent classification results compared to other models, which may be less effective or introduce additional computational overhead. Figure 3 represents the structure of the ShuffleNetv2 technique.
The respective four guidelines are given in the following: (1) excessive utilization of group convolutional enhances MAC; (2) keeping the input and output channels equivalent to the convolutional process; (3) element operation on feature map could not be ignored; (4) branching and fragmented network structure leads to reduction from the parallelism of the model that decelerates the inference, and even though the FLOP for element-level operation is smaller, the MAC is larger.
A special case of grouped convolutional is Depthwise convolution (DW-Conv), followed by accessing a size of 1×1 convolution to form DW‐Conv. The DW‐Conv is computed in Eq. (1).
The managers of the connected matrix are \({\text{I}},\)\(j,\)\(w,\) and \(h,\) and the outcome feature matrix and convolution kernel weighted matrix is G and K, correspondingly. DW-Conv is capable of replacing the regular convolutional with lesser computation and is nearly standard for lightweight processes:
Here, the computation amount of DW-Conv and standard convolution is indicated as\(~{Q_1}\) and \({Q_2}\); respectively, the side dimensional of the feature matrix and the convolutional kernel are specified as \({D_f}\) and \({D_k}\). The count of input and output mapping feature channels are depicted as N and M. During feature extraction, the convolutional function of 3×3 dimensional is used. Consequently, the computation amount of DW‐Conv is only about 1/9 of that of standard convolution. ShuffleNet introduced the concept of a vital innovation point: recognizing channels’ data change during the extraction feature process with low calculation costs.
Image classification
This work uses the CDAE model for leukemia detection and classification27. The CDAE method was chosen for image classification due to its robust capability to learn robust features from noisy data while reconstructing clean images, which is crucial for accurate classification. Its architecture outperforms in denoising and conserving significant image details, improving the quality of input data for subsequent evaluation. For leukaemia detection, feature extraction capabilities of the CDAE technique enhance the technique’s sensitivity to subtle discrepancies in medical images, paving the way to more accurate classification. When incorporated with the FOA technique for hyperparameter tuning, CDAE benefits from FOA’s effectual exploration of parameter space, optimizing the performance and robustness of the method. This integration confirms precise and reliable leukemia detection by fine-tuning the CDAE to adapt efficiently to the dataset. Figure 4 represents the infrastructure of CDAE.
The DAE incorporates the denoising code into the classical AE. It is used to accept the corrupted input dataset and reconstruct the uncorrupted original data as output via training to increase the anti-noise capability of the network, as well as the structure of DAE. Convolutional autoencoder (CAE) is used to combine the AE and CNN. The CDAE model, which goes to the unsupervised learning model, is proposed by DAE and CAE. It fuses CNN’s pooling and convolution layers and enhances an upsampling layer that allows decoding and encoding processes.
While accomplishing feature extraction, CDAE is used to retain the 2D data structure of the images. Local perception feature could reflect the configuration data of the image, and weighted sharing can diminish the parameter count that needs to be trained in the network:
where\(~\hat {\alpha }\) is the transformed image after adding noise to the original signal, the original image transmuted by the signal is represented as \(\alpha\), the original vibration signal is x, and the noise signal conforming to the specific distribution is\(~n\left( t \right)\).
Then, for any \({j^{th}}\) feature maps, Eq. (7) is described as:
where \(\sigma =\left\{ {{W^j},{b^j}} \right\},\)\({\text{*~}}\)is the convolution function, \(and~\phi\) is the activation function. The weighted matrix and deviation vector are \({W^j}\) and \({b^j}\) correspondingly, and the parameter is shared on all maps. Using the decoder, the encoded images are recreated as follows:
where N shows the set of possible feature maps. \(\sigma ^{\prime}=\left\{ {{W^{{{\prime }}j}},{b^{{{\prime }}j}}} \right\}.\)\({,^*}~\)is the convolution function, \(\phi ^{\prime}\left( \cdot \right)\) is the activation function, and the weighted matrix and deviation vector are\(~{W^{{{\prime }}j}}\) and \({b^{{{\prime }}j}}\), correspondingly. The ReLU and sigmoid functions are the commonly used activation functions as follows:
The backpropagation model performs the σ and σ’ parameter optimization. The CDAE should be enhanced to minimalize the objective function for suppressing the noise as follows:
In Eq. (12), the sample size is M, and the loss function is\(~L\). In general, the MSE loss function can be employed, and then the primary function is formulated by Eq. (14):
After decoding, the noisy images are restored to the original image using unsupervised training of the CDAE network, which enhances the model’s capability to express denoising features.
Hyperparameter tuning using FOA
Finally, the FOA adjusts the hyperparameter values of the CDAE model28. The FOA method is well-suited explicitly for hyperparameter selection of the CDAE approach due to its effectiveness in exploring complex search spaces and averting local optima. FOA replicates falcons’ hunting behaviour, efficiently balancing exploration and exploitation during optimization. This results in an enhanced convergence speed and accuracy compared to conventional methodologies. Furthermore, the adaptability of the FOA model to high-dimensional problems and the capability to handle various hyperparameter configurations make it a powerful tool for fine-tuning CDAE techniques, leading to improved performance and model robustness. Figure 5 illustrates the steps involved in the FOA method.
FOA is robust and reliable for stochastic population-based problems that require arrangement from a set of parameters to its three-phase action settlement. The motivation behind that was the chasing strategy of falcons while looking for prey during the flight. The foraging process of falcons can be discussed as follows:
Stage 1: Parameter initialization.
The FOA initially defines the decision variables, limits, and optimization difficulties. Next, the parameters \({d_d},\)\({g_d}\) and \({t_d}{\text{~}}\)represent cognitive, following, and social constants. \(OQ\) indicates the falcon quantities. The awareness and diving probabilities are correspondingly specified as\(~BQ\) and \(EQ\), and the maximum acceptable speed is represented as\(~{u_{{\text{max}}}}\).
Stage 2: Initialize the values of velocity and position of the falcon.
In this stage, the boundary condition is considered for locating the falcons randomly in D-dimension.
The falcon position is now specified as\(~y\), based on the number of falcon \(OQ\) in D-dimensional space. Using Eqs. (13) & (14), maximum and minimum velocity values can be defined.
Here, maximum and minimum acceptable velocities are represented as \({u_{{\text{max}}}}\) and \({u_{{\text{min}}}}\), correspondingly, and the upper bound limit range is termed as \(ub\).
Stage 3: Finding the individual position, value of best global, and determination of fitness value.
The \(pg\) represents the fitness value, which can be defined by generating vectors.
Here, the best position of the individual falcon is characterized as \({y_{best}}\). Lastly, the new location was created using \({y_{best}}\), based on awareness and diving probabilities.
Stage 4: Establishment and upgrade of the falcon’s new location.
To begin with, the awareness and diving probabilities are contrasted by producing the two random integers, which are indicated as \({q_{BQ}}\) and \({q_{EQ}}\) correspondingly. Next, the value of \({q_{BQ}}\) is contrasted with awareness probability \(BQ\). If \({q_{BQ}}<BQ\), then the experience of the falcon is taken into account for detecting its prey, and it is shown as follows:
Now, the velocity and existing location of the falcon are specified as \({y_{ltr - 1}},{u_{ltr - 1}}\), correspondingly.
If \({q_{BQ}}<BQ,\) the fitness values of falcon and prey are contrasted. Eventually, the falcon follows the prey by using a diving approach, and it is represented as follows:
In Eq. (16), \({y_{cho}}\) is the chosen prey. If the abovementioned criteria are not met, the falcon’s displacement is based on an individual’s fittest location and given as.
Lastly, velocity and boundary conditions are examined for the new location, and the fitness values are calculated for the new location. The new position of \({y_{best}}\) is determined.
Stage 5: Repeat the steps until the maximum number of iterations is attained.
The fitness optimum is a key feature in the FOA methodology. Encoded performance has been deployed to improve the efficiency of candidate results. Presently, the accuracy value is a primary condition for designing an FF.
whereas \(FP\) and \(TP\) denote the false and true positive values.
Results and discussion
In this section, the leukemia detection outcomes of the FOADCNN-LDC methodology can be tested utilizing the leukemia database from the Kaggle repository29. The database comprises 518 samples with five classes, as represented in Table 1. The suggested technique is simulated using the Python 3.6.5 tool on PC i5-8600k, 250GB SSD, GeForce 1050Ti 4GB, 16GB RAM, and 1 TB HDD. The parameter settings are provided: learning rate: 0.01, activation: ReLU, epoch count: 50, dropout: 0.5, and batch size: 5.
Figure 6 shows the confusion matrices achieved by the FOADCNN-LDC technique under 80:20 and 70:30 of the TR/TS phase. The simulated values reported the effective recognition of all 5 classes. In the TR phase (80%), the confusion matrix exhibits high accuracy for distinguishing between ALL, AML, CLL, CML, and Healthy samples, with no misclassifications among certain classes. The confusion matrix in the TR phase (70%) portrays misclassifications, specifically with CML and CLL. During the TS phase (20%), the model accomplishes well with most samples correctly classified, though there are some misclassifications of ALL and CLL. The TS phase (30%) illustrates an enhanced performance with the majority of samples precisely classified, though some misclassifications remain, specifically with AML and CML.
In Table 2; Fig. 7, the leukaemia classification outcomes of the FOADCNN-LDC methodology are illustrated under 80:20 of the TR/TS phase. The results highlighted that the FOADCNN-LDC model classifies the samples proficiently. With 80% of the TR Phase, the FOADCNN-LDC model gains an average \(acc{u_y}\) of 99.32%, \(sen{s_y}\) of 98.53%, \(spe{c_y}\) of 99.54%, \({F_{score}}~\)of 98.22%, and MCC of 97.75%. Also, based on 20% of the TS Phase, the FOADCNN-LDC methodology gains an average \(acc{u_y}\) of 99.62%, \(sen{s_y}\) of 97.14%, \(spe{c_y}\) of 99.78%, \({F_{score}}~\)of 97.59%, and MCC of 97.47% correspondingly.
In Table 3; Fig. 8, the leukaemia classification analysis of the FOADCNN-LDC technique is exhibited at 70:30 in the TR/TS phase. The simulated values show that the FOADCNN-LDC method classifies the samples proficiently. Additionally, based on 70% of the TR Phase, the FOADCNN-LDC methodology attains an average \(acc{u_y}\) of 99.23%, \(sen{s_y}\) of 97.41%, \(spe{c_y}\) of 99.48%, \({F_{score}}~\)of 97.45%, and MCC of 96.94%. Besides, based on 30% of the TS Phase, the FOADCNN-LDC approach gets an average \(acc{u_y}\) of 99.23%, \(sen{s_y}\) of 97.66%, \(spe{c_y}\) of 99.46%, \({F_{score}}~\)of 97.47%, MCC of 97.01% respectively.
To determine the performance of the FOADCNN-LDC method with 80:20 of TR/TS phase, TR and TS \(acc{u_y}\) curves are defined, as represented in Fig. 9. The TR and TS \(acc{u_y}\) curves exhibit the performance of the FOADCNN-LDC method over numerous epochs. The figure provides essential details about the learning tasks and generalization capabilities of the FOADCNN-LDC technique. With an improvement in epoch count, it is observed that the TR and TS \(acc{u_y}\) curves acquire enhanced. The FOADCNN-LDC technique attains improved testing accuracy, which can potentially recognize patterns in the TR and TS data.
Figure 10 represents the overall TR and TS loss values of the FOADCNN-LDC method with 80:20 of TR/TS phase over epochs. The TR loss represented the model loss acquired reduced over epochs. The loss values acquired mainly decreased as the model adapted the weight to diminish the predicted error on the TR and TS data. The loss curves illustrate how much the model fits the training data. It is remarked that the TR and TS loss is gradually lessening and described that the FOADCNN-LDC methodology successfully learns the patterns exhibited in the TR and TS data. It has also been noticed that the FOADCNN-LDC methodology changes the parameters to decrease the difference between the predicted and actual training labels.
The PR performance of the FOADCNN-LDC technique with 80:20 of TR/TS phase is illustrated by plotting precision against recall as reported in Fig. 11. The simulated values confirm that the FOADCNN-LDC technique gets increased PR values with each 5 class. The figure represents that the model learns to recognize different 5 class labels. The FOADCNN-LDC model improves outcomes in recognizing positive samples with decreased false positives.
Figure 12 shows the ROC analysis offered by the FOADCNN-LDC methodology with 80:20 of the TR/TS phase, which can differentiate the 5 class labels. The figure points out valuable perceptions of the trade-off between the TPR and FPR rates over diverse classification thresholds and changing numbers of epochs. It accurately provides the predictive performance of the FOADCNN-LDC approach on the classification of distinct 5 classes.
Table 4 represents an extensive comparison study of the FOADCNN-LDC technique with recent models12. Figure 13 exhibits a comparative \(acc{u_y}\) analysis of the FOADCNN-LDC method. The results indicate the FOADCNN-LDC method performed better. Based on \(acc{u_y}\), the FOADCNN-LDC technique offers an increased \(acc{u_y}\) of 99.62% while the AlexNet, GA-ANN, SCA-DCNN, CNN-SVM, ResNet-34, and DenseNet-121 models attain decreased \(acc{u_y}~\)values of 96.76%, 97.05%, 98.60%, 98.90%, 99.11%, and 99.22% respectively.
Figure 14 represents a comparison \(sen{s_y}\) and \(spe{c_y}~\)analysis of the FOADCNN-LDC technique. The simulated values represent the excellent performance of the FOADCNN-LDC technique. According to the value of \(sen{s_y}\), the FOADCNN-LDC methodology offers an improved \(sen{s_y}\) of 97.14%. In contrast, the AlexNet, GA-ANN, SCA-DCNN, CNN-SVM, ResNet-34, and DenseNet-121 models achieve reduced \(sen{s_y}~\)values of 96.98%, 96.44%, 97.03%, 97.09%, 96.91%, and 96.59% individually. Also, with \(spe{c_y}\), the FOADCNN-LDC technique gives an improved \(spe{c_y}\) of 99.78%, but the AlexNet, GA-ANN, SCA-DCNN, CNN-SVM, ResNet-34, and DenseNet-121 methodologies get diminished \(spe{c_y}~\)values of 97.29%, 97.07%, 97.72%, 98.88%, 98.76%, and 98.54% correspondingly. These simulated values confirmed the enhanced performance of the FOADCNN-LDC technique.
The computational time (CT) of the FOADCNN-LDC model is shown in Table 5; Fig. 15. The computational time for diverse techniques exhibits notable differences in efficiency. The FOADCNN-LDC method stands out with the shortest processing time of 3.88 s, illustrating its superior effectiveness compared to other methodologies. AlexNet and DenseNet-121 portray longer processing times of 7.60 and 8.69 s, reflecting their more complex architectures. GA-ANN and DenseNet-121 are the slower techniques, taking 8.76 and 8.69 s, respectively. On the contrary, CNN-SVM and SCA-DCNN models present moderate processing times of 5.66 and 6.31 s, suggesting a balance between performance and effectualness. Overall, the FOADCNN-LDC method provides a faster alternative while maintaining efficient performance.
Conclusion
In this study, a model for automated classification and detection of leukemia on medical images is developed. The main objective of the FOADCNN-LDC technique is to classify and recognize the occurrence of leukemia. To accomplish this, the FOADCNN-LDC technique comprises a series of processes: MF-based preprocessing, ShuffleNetv2 feature extraction, CDAE-based classification, and FOA-based hyperparameter tuning. Meanwhile, the FOADCNN-LDC technique utilizes the ShuffleNetv2 approach for the feature extraction process. Furthermore, the detection and classification of leukemia are performed using the CDAE model. The FOA is employed to select the hyperparameter of the CDAE model. The simulation process of the FOADCNN-LDC approach is performed on a benchmark medical dataset. The investigational analysis of the FOADCNN-LDC approach highlighted a superior accuracy value of 99.62% over existing techniques. The primary challenges in leukaemia classification utilizing the FOADCNN-LDC technique include managing noisy images, extracting relevant features, and fine-tuning the model’s parameters. The proposed technique addresses these problems by implementing the MF model to remove noise, which improves the clarity of the image and assists in more accurate feature extraction employing the ShuffleNetv2 model. The CDAE technique efficiently handles the complexities of leukemia detection and classification by reconstructing clean, informative features from noisy data. Moreover, the FOA model optimizes the hyperparameters of the CDAE model, enhancing overall model performance. However, limitations encompass potential computational overhead from the complex model and possible threats in generalizing across several datasets. Future work should focus on refining the model for improved scalability and exploring alternative optimization methods to improve performance. Future research may also explore incorporating advanced augmentation models to enhance model robustness against varied data conditions. Furthermore, developing lightweight versions of the model to mitigate computational needs while maintaining accuracy would be beneficial. Examining other optimization models beyond FOA could also lead to additional enhancements in hyperparameter tuning and overall performance.
Data availability
The datasets used and analyzed during the current study available from the corresponding author on reasonable request.
References
Mallick, P. K., Mohapatra, S. K., Chae, G. S. & Mohanty, M. N. Convergent learning–based model for leukemia classification from gene expression. Personal. Uniquit. Comput.27(3), 1103–1110 (2023).
Abhishek, A., Jha, R. K., Sinha, R. & Jha, K. Automated classification of acute leukemia on a heterogeneous dataset using machine learning and deep learning techniques. Biomed. Signal Process. Control72, 103341 (2022).
Gondal, C. H. A. et al. Automated leukemia screening and sub-types classification using deep learning. Comput. Syst. Sci. Eng., 46(3), 3541–3558 (2023).
Das, P. K. & Meher, S. An efficient deep convolutional neural network based detection and classification of acute lymphoblastic leukemia. Expert Syst. Appl.183(115311), (2021).
Bukhari, M., Yasmin, S., Sammad, S., El-Latif, A. & Ahmed, A. A deep learning framework for leukemia cancer detection in microscopic blood samples using squeeze and excitation learning. Math. Problems Eng. (2022).
Hagar, M., Elsheref, F. K. & Kamal, S. R. A new model for blood cancer classification based on deep learning techniques. Int. J. Adv. Comput. Sci. Appl.14(6), (2023).
Arivuselvam, B. & Sudha, S. Leukemia classification using the deep learning method of CNN. J. X-Ray Sci. Technol.30 (3), 567–585 (2022).
Ramagiri, A. et al. March. Image classification for optimized prediction of leukemia cancer cells using machine learning and deep learning techniques. In 2023 International Conference on Innovative Data Communication Technologies and Application (ICIDCA) (pp. 193–197). IEEE. (2023).
Abhishek, A., Jha, R. K., Sinha, R. & Jha, K. Automated detection and classification of leukemia on a subject-independent test dataset using deep transfer learning supported by Grad-CAM visualization. Biomed. Signal Process. Control83, 104722 (2023).
Veeraiah, N., Alotaibi, Y. & Subahi, A. F. MayGAN: Mayfly optimization with generative adversarial network-based deep learning method to classify leukemia form blood smear images. Comput. Syst. Sci. Eng.46(2), 2039–2058 (2023).
Zakir Ullah, M. et al. An attention-based convolutional neural network for acute lymphoblastic leukemia classification. Appl. Sci.11(22), 10622 (2021).
Bibi, N., Sikandar, M., Ud Din, I., Almogren, A. & Ali, S. IoMT-based automated detection and classification of leukemia using deep learning. J. Healthcare Eng. (2020).
Jawahar, M., Sharen, H. & Gandomi, A. H. ALNett: A cluster layer deep convolutional neural network for acute lymphoblastic leukemia classification. Comput. Biol. Med.148, 105894 (2022).
Atteia, G. E. Latent space representational learning of deep features for acute lymphoblastic leukemia diagnosis. Comput. Syst. Sci. Eng.45(1), (2023).
Agustin, R. I., Arif, A. & Sukorini, U. Classification of immature white blood cells in acute lymphoblastic leukemia L1 using neural networks particle swarm optimization. Neural Comput. Appl.33(17), 10869–10880 (2021).
Chand, S. & Vishwakarma, V. P. A novel deep learning framework (DLF) for classification of acute lymphoblastic leukemia. Multimedia Tools Appl.81(26), 37243–37262 (2022).
Sulaiman, A. et al. ResRandSVM: Hybrid approach for acute lymphocytic leukemia classification in blood smear images. Diagnostics, 13(12), p.2121. (2023).
Ali, A. M. & Mohammed, M. A. A comprehensive review of artificial intelligence approaches in omics data processing: Evaluating progress and challenges. Int. J. Math. Stat. Comput. Sci.2, 114–167 (2024).
Mohammed, M. Enhanced cancer subclassification using multi-omics clustering and quantum cat swarm optimization. Iraqi J. Comput. Sci. Math.5(3), 552–582 (2024).
Benameur, N. et al. Numerical study of two microwave antennas dedicated to superficial cancer hyperthermia. Procedia Comput. Sci.239, 470–482 (2024).
Hosseinzadeh, M. et al. A Diagnostic model for acute lymphoblastic leukemia using metaheuristics and deep learning methods. arXiv preprint arXiv:2406.18568. (2024).
Awais, M. et al. An efficient decision support system for leukemia identification utilizing nature-inspired deep feature optimization. Front. Oncol., 14, p.1328200. (2024).
Noshad, A. & Fallahi, S. A new hybrid framework based on deep neural networks and JAYA optimization algorithm for feature selection using SVM applied to classification of acute lymphoblastic leukaemia. Comput. Methods Biomech. Biomedical Engineering: Imaging Visualization. 11(4), 1549–1566 (2023).
Shree, K. D. & Logeswari, S. ODRNN: Optimized deep recurrent neural networks for automatic detection of leukaemia. Signal. Image Video Process.18(5), 4157–4173 (2024).
Singh, P., Bhandari, A. K. & Kumar, R. Naturalness balance contrast enhancement using adaptive gamma with cumulative histogram and median filtering. Optik, 251, p.168251. (2022).
Chen, Z., Yang, J., Feng, Z. & Chen, L. RSCNet: An efficient remote sensing scene classification model based on lightweight convolution neural networks. Electronics, 11(22), p.3727. (2022).
Shi, H., Chen, J., Si, J. & Zheng, C. Fault diagnosis of rolling bearings based on a residual dilated pyramid network and full convolutional denoising autoencoder. Sensors, 20(20), p.5734. (2020).
Nadarajan, D. & Perumal, T. P. Logistic 2D map based elliptic curve cryptography encryption Scheme with Key optimization using pathfinder and Falcon algorithms for providing enhanced security in Open Social Networks. Rivista Italiana Di Filosofia Analitica Junior. 14(2), 797–821 (2023).
https://www.kaggle.com/datasets/nikhilsharma00/leukemia-dataset
Acknowledgements
This work was funded by the University of Jeddah, Jeddah, Saudi Arabia, under grant No. (UJ-23-DR-199). Therefore, the authors thank the University of Jeddah for its technical and financial support.
Author information
Authors and Affiliations
Contributions
Conceptualization, Data curation, Formal analysis, Investigation, Methodology, Writing—original draft: Turky Omar Asar Project administration and Resources, Supervision, Validation and Visualization, Writing—review and editing: Mahmoud RagabAll authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval
This article does not contain any studies with human participants performed by any of the authors.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Asar, T.O., Ragab, M. Leukemia detection and classification using computer-aided diagnosis system with falcon optimization algorithm and deep learning. Sci Rep 14, 21755 (2024). https://doi.org/10.1038/s41598-024-72900-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-72900-3
Keywords
This article is cited by
-
DSTL-NET: Multi-level Leukemia Stage Classification Via Dual-Stage Segmentation Based Transfer Learning Networks
International Journal of Computational Intelligence Systems (2026)
-
Advanced deep learning framework for automated hematological malignancy classification: integrating FCMAE V2-WPAT with ACDB-GAN for enhanced leukemia subtype detection
Signal, Image and Video Processing (2026)
-
A two stage blood cell detection and classification algorithm based on improved YOLOv7 and EfficientNetv2
Scientific Reports (2025)
-
A Novel Modified AlexNet with Spatial Pyramid Pooling for Robust Detection of Acute Myeloid Leukemia from Blood Smear Images
Iranian Journal of Science and Technology, Transactions of Electrical Engineering (2025)
-
Transformative Advances in AI for Precise Cancer Detection: A Comprehensive Review of Non-Invasive Techniques
Archives of Computational Methods in Engineering (2025)