Abstract
One of the critical issues in medical data analysis is accurately predicting a patient’s risk of heart disease, which is vital for early intervention and reducing mortality rates. Early detection allows for timely treatment and continuous monitoring by healthcare providers, which is essential but often limited by the inability of medical professionals to provide constant patient supervision. Early detection of cardiac problems and continuous patient monitoring by physicians can help reduce death rates. Doctors cannot constantly have contact with patients, and heart disease detection is not always accurate. By offering a more solid foundation for prediction and decision-making based on data provided by healthcare sectors worldwide, machine learning (ML) could help physicians with the prediction and detection of HD. This study aims to use different feature selection strategies to produce an accurate ML algorithm for early heart disease prediction. We have chosen features using chi-square, ANOVA, and mutual information methods. The three feature groups chosen were SF-1, SF-2, and SF-3. The study employed ten machine learning algorithms to determine the most accurate technique and feature subset fit. The classification algorithms used include support vector machines (SVM), XGBoost, bagging, decision trees (DT), and random forests (RF). We evaluated the proposed heart disease prediction technique using a private dataset, a public dataset, and different cross-validation methods. We used the Synthetic Minority Oversampling Technique (SMOTE) to eliminate inconsistent data and discover the machine learning algorithm that achieves the most accurate heart disease predictions. Healthcare providers might identify early-stage heart disease quickly and cheaply with the proposed method. We have used the most effective ML algorithm to create a mobile app that instantly predicts heart disease based on the input symptoms. The experimental results demonstrated that the XGBoost algorithm performed optimally when applied to the combined datasets and the SF-2 feature subset. It had 97.57% accuracy, 96.61% sensitivity, 90.48% specificity, 95.00% precision, a 92.68% F1 score, and a 98% AUC. We have developed an explainable AI method based on SHAP approaches to understand how the system makes its final predictions.
Similar content being viewed by others
Introduction
Globally, heart diseases consistently rank as the leading cause of death1. Heart disease and stroke account for 17.5 million annual deaths worldwide, according to the World Health Organization’s report. More than 75% of deaths caused by heart diseases occur mostly in countries with middle- and low-income populations. In addition, heart attacks and strokes are responsible for 80% of all fatalities caused by CVDs2. Observing the patient’s symptoms and conducting a physical examination often leads to the diagnosis of heart disease. Smoking, age, a family history of heart disease, high cholesterol levels, inactivity, high blood pressure, obesity, diabetes, and stress are some risk factors for cardiovascular disease3. Lifestyle modifications, such as quitting smoking, losing weight, exercising, and managing stress, may help to reduce some of these risk factors. We diagnose heart disease using medical history, physical examination, and imaging tests such as electrocardiograms, echocardiograms, cardiac MRIs, and blood tests. Lifestyle adjustments, drugs, medical treatments like angioplasty, coronary artery bypass surgery, or implanted devices like pacemakers or defibrillators can treat heart disease4. It is now possible to construct prediction models for heart disease with the assistance of the vast amounts of patient data that are easily accessible as a result of the growing number of recent healthcare systems. Machine learning is considered a data-sorting approach that analyzes large datasets from various viewpoints and then transforms the results into tangible knowledge5.
Several studies have utilized ML algorithms like SVM, artificial neural networks (ANN), DT, LR, and RF to analyze medical data and predict heart diseases. A recent study by6used ML models to predict the risk of cardiac disease in a multi-ethnic population. The authors utilized a large dataset of electronic health record data and linked it with socio-demographic information to stratify CVD risks. The models achieved high accuracy in predicting CVD risk in the multi-ethnic population. Similarly, another study by7applied a deep learning (DL) algorithm to predict coronary artery disease (CAD). The researchers utilized clinical data and coronary computed tomography angiography (CCTA) images to train the DL model. A study by8used different ML models to predict CVD based on clinical data. The models used by the researchers included DTs, K-nearest neighbor (KNN), and RFs. Using these models, the authors reported high accuracy in predicting CVD. Similarly, in a study by9, ML techniques were used to determine what factors contribute to heart disease risk. The authors utilized the National Health and Nutrition Examination Survey (NHANES) data to determine risk factors related to coronary heart disease. Another research study10 examined the effectiveness of various machine learning algorithms in predicting heart diseases. The authors reported that the models achieved high accuracy in predicting heart diseases.
Many researchers use ML classification techniques to predict heart disease. The ML classifiers used in this work have shown promising results in detecting the risk of CVD11,12,13,14,15,16,17,18,19,20,21,22,23. The LR, RF, and KNN algorithms have shown high accuracy in classifying the risk of CVD. Ensemble learning techniques, such as bagging, AdaBoost, and voting, have improved the classification accuracy compared to single classifiers. Employing several ML classifiers can enhance the accuracy of CVD risk prediction. We can conduct further research in this area to improve CVD forecasting and diagnosis. ML is a powerful tool for HD prediction. It has the potential to improve patient outcomes by allowing for early detection and personalized treatment. This part starts with a comparison of ten machine learning classifiers for predicting heart disease. The classifiers are Naive Bayes, SVM, voting, XGBoost, AdaBoost, bagging, KNN, DT, RF, and LR (see Table 1). The results indicated that ML classifiers could improve heart disease prediction accuracy, with the highest achieved being 97% by28using AdaBoost, DT, RF, KNN, and LR on the UCI dataset. Several studies utilized the Cleveland heart disease dataset (CHDD), with accuracies ranging from 77%32to 92%30 using various ML algorithms such as AdaBoost, DT, RF, KNN, LR, SVM, and Naive Bayes. Hence, ML classifiers could improve the certainty of heart disease forecasting, enabling early detection and personalized treatment. Nonetheless, more investigation is essential to validate these classifiers’ accuracy using larger datasets and increase the generalizability and reproducibility of the results.
The objective of the study is to provide an ML approach for heart disease prediction. We evaluated ML algorithms on large, open-access heart disease prediction datasets. This study aims to construct an innovative machine learning technique that is capable of properly classifying several high-definition datasets and then evaluate its performance in comparison to that of other first-rate models. One of the key contributions to this research is the use of a private HD dataset. Egyptian specialized hospitals voluntarily provided 200 data samples between 2022 and 2024. We were able to collect approximately 13 features from these participants. This work deals with the immediate requirement for early HD prediction in Egypt and Saudi Arabia, where the HD rate is rapidly increasing. The authors evaluated the proposed model’s performance by applying ML classification algorithms to a combined dataset consisting of both CHDD and private datasets. Using a combined dataset, this approach accurately predicts HD. It is a new method compared to earlier studies. The research’s stated goal was to predict HD using the combined datasets and the SF-2 feature subset. The following rates were achieved: 97.57% for accuracy, 96.61% for sensitivity, 90.48% for specificity, 95.00% for precision, 92.68% for F1 score, and 98% for AUC. To understand how the system predicts its outcomes, an explainable artificial intelligence approach utilizing SHAP methodologies has been developed. The use of SMOTE to increase the overall number of balanced cases in the dataset is of additional importance to this study. The proposed technique is trained on a balanced dataset using SMOTE to increase the performance of heart disease prediction. The ML techniques applied in this article were additionally optimized with hyperparameters. We have tuned the hyperparameters for all the ML classifiers. The proposed method got 97.57% accuracy rates with hyperparameters that were optimized when the combined datasets and the SF-2 feature subset were used. Additionally, to identify the classifier that achieves the most accurate HD prediction rate, the study assessed 10 distinct ML classification algorithms. The XGBoost technique was identified as a highly accurate classifier to predict HD after assessing the performance of ten algorithms. The proposed app’s capacity for adaptability is shown by applying a domain adaptation method. This shows the ability of the proposed approach to be implemented in various environments and communities, in addition to the initial datasets used in this article.
The proposed study offers several unique contributions that significantly enhance its novelty and relevance in the heart disease prediction field, including:
-
1.
Comprehensive Feature Selection Methodology: Our research introduces a comprehensive feature selection process using three distinct methods: chi-square, analysis of variance (ANOVA), and mutual information (MI). Unlike prior studies that may rely on a single or less systematic approach, we rigorously evaluated the importance of each feature through these methods, resulting in three specialized feature subsets (SF-1, SF-2, SF-3). This methodological approach guarantees the inclusion of only the most relevant and impactful features in the predictive model, thereby enhancing its accuracy and efficiency.
-
2.
Evaluation Across Multiple ML Classifiers: We conducted an extensive comparative analysis of ten different ML classifiers, including state-of-the-art algorithms like XGBoost, AdaBoost, and ensemble methods. We identified the optimal classifier-feature combination, a topic not commonly addressed in existing literature, through a broad evaluation across various algorithms and the use of selected feature subsets. We demonstrated the superior performance of the XGBoost classifier with the SF-2 feature subset, highlighting the significance of our feature selection strategy.
-
3.
Utilization of a Private Health Dataset: In addition to using a publicly available dataset, we employed a private health dataset that has not been explored in previous research. This inclusion of a novel dataset adds a layer of originality to our study, as it allows us to validate the model’s robustness and generalizability across different data sources. This dataset’s results offer fresh perspectives on heart disease prediction, especially in areas where comparable data has not undergone extensive analysis.
-
4.
We implemented the Synthetic Minority Oversampling Technique (SMOTE) to address the issue of unbalanced data, which is often a challenge in medical datasets. By ensuring balanced training data, our study improves the reliability and accuracy of the predictive models, particularly in detecting early-stage heart disease. This step is crucial for enhancing the practical applicability of the model in real-world scenarios, where data imbalance is common.
-
5.
Development of an Explainable AI Approach: To our knowledge, the integration of SHAP (Shapley Additive Explanations) methodologies to provide an explainable AI framework in the context of heart disease prediction is a novel contribution. This approach not only enhances the model’s trustworthiness by providing transparency into the prediction process, but also assists healthcare professionals in comprehending the underlying factors that influence the diagnosis.
-
6.
Practical Application Through a Mobile App: Finally, the development of a mobile application based on the best-performing ML model marks a significant step towards practical, real-world implementation. This app enables users to input symptoms and quickly receive a heart disease prediction, offering a user-friendly, cost-effective tool for early detection. The translation of our research findings into a tangible product underscores the novelty of our study by bridging the gap between theoretical research and practical healthcare solutions.
Methods
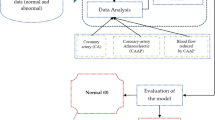
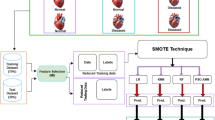
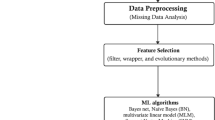
Figure 1 shows the proposed system’s sequences for predicting heart diseases. We first gathered and preprocessed the dataset to remove any necessary inconsistencies, such as replacing null occurrences with average values. We divided the dataset into two distinct groups, named the test dataset and the training dataset, respectively. Next, we implemented several distinct classification algorithms to determine which one achieved the highest accuracy for these datasets.
The proposed methodology
This study investigates ML techniques such as Naive Bayes, SVM, voting, XGBoost, AdaBoost, bagging, DT, KNN, RF, and LR classifiers. These algorithms can aid doctors and data analysts in making correct diagnoses of cardiac disease. This article incorporates recent data on cardiovascular illness, as well as relevant journals, research, and publications. The methodology, as in1, provides a framework for the suggested model. The methodology is a set of steps that transform raw data into consumable and identifiable data patterns. The proposed approach consists of three stages: the first stage is data collection; the second stage extracts specific feature values; and the third stage is data exploration, as shown in Fig. 1. Depending on the procedures employed, data preprocessing deals with the missing values, cleansing of the data, and normalization2. We then classified the pre-processed data using the ten classifiers (A1, A2,., A10). Finally, after putting the suggested model into practice, we evaluated its performance and accuracy using a range of performance measures. This model developed a Reliable Prediction System for Heart Disease (RPSHD) using a variety of classifiers. This model uses 13 medical factors for prediction, among which are age, sex, cholesterol, blood pressure, and electrocardiography3.
Datasets and dataset features
This research employs both the CHDD and a private dataset for heart disease prediction. The CHDD dataset has 303 samples, while the private dataset has 200, and they have the same features. The combined dataset contains 503 records, and 13 features are associated with each one (including demographic, clinical, and laboratory parameters). The datasets have many features that can be used for heart disease prediction including age, gender, blood pressure, cholesterol levels, electrocardiogram readings-ECG, chest pain, exercise-induced angina, blood sugar with fasting condition, max heart rate achieved, oldpeak, coronary artery, thalassemia, and other clinical and laboratory measurements, as shown in Table 2. The outcome variable known as “Target” takes a binary value and refers to the heart disease predicting feature (i.e., it indicates whether or not cardiac disease is present).
Figure 2 shows the percentage distribution of individuals with heart disease in the combined datasets. A total of 503 samples have been gathered, and 45.9% of those have been diagnosed with HD, while the remaining 54.1% of individuals have not been infected with the disease.
Boxplots are an effective visualization technique for understanding the distribution of data and identifying potential outliers. By applying boxplots to a dataset related to HD, one can get insights into the distribution of a variety of HD-related features or variables. The HD dataset’s boxplots are illustrated in Fig. 3. Boxplots are used to illustrate the distribution of scores for HD detection in this figure. Every graph we obtained had an anomaly. Removing them will cause the median of the data to drop, which might make it harder to detect HD accurately. On the other hand, this method offers more benefits than the others; by identifying heart disease infection at an early stage, when medical care is most beneficial, this diagnostic could preserve lives.
Dataset preparation
In this research, preprocessing was performed on collected data. The CHDD has four inaccurate CMV records and two erroneous TS entries. Incorrect data is updated to reflect the best possible values for all fields. Then, StandardScaler is employed to normalize all the features to the relevant coefficient, ensuring each feature has a zero mean and one variance. By considering the patient’s history of cardiac problems and following other medical concerns, an organized and composed augmented dataset was chosen.
The dataset studied in this research is a combination of accessible public WBCD and chosen private datasets. Partitioning the two datasets in this way allows us to use the holdout validation method. In this study, 25% of the data is in the test dataset, compared to 75% in the training dataset. The mutual information method is used in this research to measure the interdependence of variables. Larger numbers indicate greater dependency and information gathering.
The importance of features provides valuable insights into the relevance and predictive power of each feature in a dataset. Using this reciprocal information technique, the thalach feature is given the highest value of 13.65%, while the fbs feature is given the lowest importance of 1.91%, as illustrated in Fig. 4.
Feature selection
In this research, we perform feature selection and classification using the Scikit-learnmodule of Python20. Initially, the processed dataset was analyzed using several different ML classifiers, including RF, LR, KNN, bagging, DT, AdaBoost, XGBoost, SVM, voting, and Naive Bayes, which were evaluated for their overall accuracy. In the second step, we used the Seaborn libraries from Python to create heat maps of correlation matrices and other visualizations of correlations between different sets of data. Thirdly, a wide variety of feature selection methods (FSM) such as analysis of variance (ANOVA), chi-square, and mutual information (MI) were applied. These strategies are explained in Table 3 and are indicated by the acronyms FSM1, FSM2, and FSM3, respectively. Finally, the performance of several algorithms was compared for the identified features. The validity of the analysis was demonstrated using accuracy, specificity, precision, sensitivity, and F1 score. The StandardScaler method was used to standardize every feature before it passed into the algorithms.
The outcome of different feature selection methods
The F value for each pair of features is determined by using the ANOVA F value technique and the feature weights. Table 4(a) presents the findings of the ANOVA F test. The EIA, CPT, and OP features provide the most importance to the score, while the RES, CM, and FBS features contribute the least. Chi-square is another approach that determines the degree to which every feature relates to the target. Table 4(b) shows the chi-square outcomes. In this method, the first three features that are the most significant are MHR, OP, and CMV, whereas TS, REC, and FBS, respectively, are the least important ones. The MI technique is utilized in FSM3. To evaluate the degree of mutual dependency between features, this approach calculates the mutual information between them. A score of 0 indicates complete independence between the two features under consideration; a larger number indicates a greater dependence. The MI score results are shown in Table 4(c). CPT, TS, and CMV are the three features that are most dependent on each other in this case, whereas FBS and REC are the features that are independent of each other. Table 4 illustrates important factors that can be utilized for predicting the probability of having heart disease. Furthermore, REC, FBS, RBP, and CM all have lower total scores across all three FSMs. Because of all these features, three distinct groups are chosen to be included depending on their score. SF-1, SF-2, and SF-3 were the abbreviations that were given to each of the three different sets of features, respectively. Table 5 shows these feature sets that were selected for additional investigation.
Based on the research’s assessment of performance criteria (see Table 6), we chose the XGBoost classifier with SMOTE using the combined datasets and SF-2 feature subset. We will embed the most accurate technique in a mobile app and deploy the model using a variety of integrated development environments (IDEs), including Android Studio 14.0, Python 3.10, Spyder, Java 11, and Pickle 526.
The use of SMOTE and SHAP methods
To overcome the problem of imbalanced datasets, ML prediction applications employ the strong Synthetic Minority Oversampling Technique (SMOTE). This technique plays an important role in various applications.
-
1.
Balancing class distribution: In many prediction tasks, such as medical diagnosis and prediction, the dataset is often imbalanced. This implies that a particular class, typically the one of interest, has a lower representation than the other class. SMOTE interpolates minority class examples to create synthetic minority class samples. This balanced class distribution ensures the prediction model gets enough minority class examples to learn from.
-
2.
Improving predictive accuracy: In predictive modeling, an imbalanced dataset can cause the model to be biased towards the majority class, leading to poor performance in predicting the minority class. Accurate prediction of the minority class poses a significant challenge. Applying SMOTE trains the model on a more balanced dataset, improving accuracy and predictive performance, particularly for the minority class. This is critical in applications where missing the minority class (e.g., disease cases) can have significant consequences.
-
3.
Enhancing recall and precision: Predictive models trained on imbalanced datasets often exhibit high precision for the majority class but low recall for the minority class. This means they miss a large portion of the minority class instances, even if the ones they do identify are accurate. SMOTE helps improve recall without sacrificing precision, leading to a more balanced and effective model. In practical terms, this means the model is better at identifying all relevant cases, not just a select few.
-
4.
Reducing model bias: In prediction applications, a biased model can result in unfair outcomes, especially when the minority class is underrepresented. By exposing the model to a sufficient number of minority class examples during training, SMOTE mitigates this bias. This helps create a more equitable model that makes fairer predictions across all classes.
-
5.
Improving generalization: Models trained on imbalanced data may perform well on the majority class during training, but they fail to generalize well to new, unseen data, particularly for the minority class. By using SMOTE to create a balanced training set, the model is better equipped to generalize its predictions to new data, leading to more reliable and consistent performance in real-world applications.
-
6.
Enhancing robustness in deployment: In deployed machine learning applications, robustness is key. Predictive models often face real-world data that is skewed or imbalanced. SMOTE helps create a more robust model that can handle such data more effectively, reducing the risk of failure in production environments. This is crucial for applications like predictive maintenance, where identifying rare but critical failures can prevent costly downtime.

On the other hand, SHAP (Shapley Additive Explanations) is a powerful tool in ML that helps to interpret and explain the predictions made by complex models. The following are the benefits that SHAP offers in ML applications:
-
1.
Enhanced Transparency: SHAP makes black-box models more transparent, fostering trust among users and stakeholders. This is especially crucial in industries like finance, healthcare, and legal, where understanding model decisions is essential.
-
2.
Regulatory Compliance: Many industries are subject to regulations that require model decisions to be explainable. SHAP ensures compliance by providing clear, understandable explanations for each decision, facilitating documentation, and sharing with regulators.
-
3.
Improved user trust and adoption: When end-users understand why a model is making certain predictions, they are more likely to trust and adopt the technology. User interfaces can incorporate SHAP explanations to improve the user friendliness of AI-powered applications.
-
4.
Actionable Insights: SHAP doesn’t just explain predictions; it also provides actionable insights. For example, in prediction models, SHAP can identify key factors for effective features, allowing doctors to take proactive steps to detect disease.
-
5.
Facilitates Collaboration: SHAP explanations can bridge the gap between data scientists and non-technical stakeholders, facilitating better communication and collaboration. By providing a common understanding of model behavior, teams can work more effectively together.
Experimental results and analysis
We use Jupyter Notebook 7 to predict heart diseases from a dataset. It simplifies the visualization of different data relationship graphs in the dataset and facilitates document creation, including live coding. The first step of this research involves cleaning the CHDD using Python’s Pandas and NumPy libraries (version 24.2.0). Next, the StandardScaler method from Python’s Scikit-learnmodule preprocesses the dataset34. The second step of the process calculates the importance of each feature using a feature selection approach, generating three sets of features (SF). Thirdly, we separated the dataset into training and testing sets. We use 75% of the data for training and the remaining 25% for testing. Finally, we trained ten distinct ML algorithms using this 75% of test data. We selected the method with the best performance to predict heart disease35.
Performance evaluation
In this subsection, the authors evaluate and explain the proposed system’s performance. The authors presented various algorithms and their comparative performances using evaluation metrics such as accuracy, sensitivity, specificity, and F1-score. We evaluated these performance measures using true positive (TP), true negative (TN), false positive (FP), and false negative (FN) data. The next subsection focuses on these measurements. After this evaluation, we provided the algorithm that produced the best results. Figure 5 illustrates the use of the confusion matrix in assessing the performance of a classification model.
Figure 5 illustrates the predicted values of TP, FP, TN, and FN for the XGBoost classifier using SMOTE. Each element in this confusion matrix represents the number of cases for both the actual classes and the predicted classes that have a particular set of labels. As an illustration, the matrix has a total of 63 cases (TP) of heart disease classifications, 3 cases (FP) of diagnosis classified as “heart disease”, 4 cases (FN) of diagnosis classed as “no heart disease”, and 66 cases (TN) of distinct “heart disease” classifications.
Figure 6 presents the correlation between the important features of SF-2 using SMOTE. The y-axis values include thalach, chol, sex, age, slope, exang, oldpeak, ca., cp., and thal. Positive or negative correlation coefficients show a significant relationship between the two variables, whereas − 1 and 1 indicate no association. It is essential to keep in mind that the only thing that can be detected via the use of correlation is the linear link that exists between the variables. The prediction for the patient is correlated with each of those variables at a level of at least 70% correlation.
Figures 7 and 8 show the scatter and density plots among the four selected features in the SF-2 dataset. These scatter and density graphs are beneficial for exploring the relationships and distributions of variables in the HD dataset. They can provide insights into correlation, concentration, outliers, and patterns that may exist among the four variables (exang, cp., ca., and thal).
-
Accuracy: The proposed model’s accuracy was developed to determine what percentage of samples has been accurately classified. Accuracy is computed using the formula given in (Eq. 1), which is based on the confusion matrices:
-
Sensitivity (or recall): Sensitivity measures the rate of truly positive results and implies that all values should be evaluated positively. Additionally, sensitivity is calculated as “the proportion of correctly detected positive samples”. Sensitivity is determined by the following formula:
-
Specificity: It predicts that all values will be negative and is determined by calculating the fraction of real negative situations. Specificity is determined mathematically by.
-
Precision: It determines classifier accuracy and may be calculated from the information given. This is presented by comparing real TP versus predicted TP. The formula in (Eq. 4) shows how the accuracy measure verifies the proposed method’s behavior:
-
F-measure: It is a statistical measure that is employed in the process of evaluating the efficacy of a classification model. It does this by determining the harmonic mean of the accuracy and recall measurements, giving each of these metrics an equal amount of weight. It enables the performance of a model to be described and compared using a single score that takes into consideration both the recall and precision of the model’s predictions and is calculated using the following formula:
The performance of a classifier has been represented and evaluated with the use of a confusion matrix, as shown in Fig. 5. TP measures how many individuals are accurately classified into the sick positive class. The percentage of healthy people who are appropriately labeled as being in the negative class is known as TN. The number of times that healthy persons were incorrectly diagnosed as being sick is referred to as the FP. When the number of healthy persons is mistakenly predicted, this is known as FN. A comparison of the various performance indicators across 10 ML algorithms is presented in Table 6. These AI classifiers were applied to the combined dataset that contained SF-1, SF-2, and SF-3 feature subsets. Based on its accuracy of 97.75%, sensitivity of 96.61%, specificity of 90.48%, precision of 95.00%, and F1 score of 92.68% for the SF-2 feature group (see Table 6), the XGBoost classifier had the best overall performance.
Experimental evaluation of system performance
Table 6 displays the accuracy of each technique and the processed dataset that underwent analysis using those algorithms. In terms of accuracy for each technique, A4’s accuracy calculation for SF-2 was the most accurate (97.57%), followed by its accuracy calculations for SF-1 and SF-3 (93.17% and 94.19%), respectively. A9 computed an accuracy of 93.07% over all three SFs, putting it in second place. On the other hand, A5 determined that SF-1 and SF-3 had a low accuracy of 85.15% among all classifiers. A3 and A10 likewise provided a low level of accuracy for SF-2 and SF-3, coming in at 86.14% and 86.12%, respectively. The other methods have an accuracy between 87.13% and 90.00%. Furthermore, this finding shows that the XGBoost algorithm method using the SF-2 is the most effective for processing the dataset. Figure 9 illustrates the range of accuracy rates possible for the ten machine learning techniques, all utilizing SF-2.
This study evaluated the sensitivity of all the algorithms. Table 6 displays the sensitivity scores obtained from the ten ML techniques using SF-1, SF-2, and SF-3, respectively. A5’s sensitivity to SF-3 was the lowest (88.14%). A8 rated both SF-1 and SF-3 (89.83% and 89.83%, respectively). A4 (XGBoost) reported the highest sensitivity for SF-2 as well, at 96.61%; A2, A3, A4, A6, and A9 reported the second-highest sensitivity, at 94.92%.
We performed the analysis of specificity on each of these techniques, and Table 6 summarizes the results. A3 scored the lowest (73.81%) for SF-2 and FS-3. According to the analysis, A4 and A9 scored the highest (90.48%) for all SFs, based on the results of the analysis. When compared to the results of the other techniques, A7 for SF-3 (92.86%) provided the best score with SF-3 only.
We have adopted several strategies to mitigate the risk of overfitting and ensure the real-world applicability of the proposed heart disease prediction model, including (1) Cross-validation: We have used the k-fold cross-validation technique in Python (using scikit-learn) to assess the generalization performance of the models on multiple subsets of the data. This helps identify models that are less prone to overfitting and provides a more reliable estimate of their true performance. (2) Regularization: incorporate regularization techniques, such as L1 or L2 regularization, dropout, or early stopping, to prevent the models from becoming overly complex and reduce the risk of overfitting. (3) Feature engineering: In addition to using feature selection methods such as ANOVA, chi-square, and MI, we expanded our work to include feature engineering techniques that capture meaningful relationships and domain knowledge relevant to heart disease prediction. (4) External validation: Validate the models on independent datasets or real-world patient data to assess their performance in a variety of settings and ensure their reliability in clinical practice.
Discussion
This study used a combined dataset (CHDD and private datasets) for both testing and training, implementing a variety of ML techniques for the early recognition of CVD. We then tested and trained the ML model on the source and target datasets using a domain adaptation approach. We first trained the proposed HD prediction technique in this study using a private dataset containing 200 cases. After that, we evaluated the system using the combined dataset of 503 cases. To be more specific, we employed a total of ten well-known ML algorithms, including Naive Bayes, SVM, voting, XGBoost, AdaBoost, bagging, DT, KNN, RF, and LR, denoted by (A1, A2, …, A10) as shown in Table 6, each with a unique set of selected features. The ANOVA F statistic, the chi-square test, and the MI statistic were the statistical methods used to group the important factors that were better at predicting CVD. We used five different evaluation standards: accuracy, sensitivity, precision, specificity, and F1 score to compare and rate the performance of the different ML techniques that used SMOTE. The experiment showed that algorithm A4 had the highest rate of accuracy (97.57%) for SF-2. Algorithm A9 had the second highest rate of accuracy (93.17%) for all three SFs shown in Table 6. A4 likewise obtained the greatest score possible for sensitivity (96.61%), as well as the best score possible for specificity (90.48%), while testing for SF-2, as shown in Table 6. The result of the F1 score demonstrated that A4 had the highest score of 92.68% for SF-2 (see Table 6), while A9 obtained the highest score of 91.57% for SF-1, SF-2, and SF-3, and A6 obtained the highest score of 90.24% for SF-2. Because A4 performs best when used with SF-2, this method is the most reliable technique in terms of accuracy, specificity, and sensitivity. In terms of F1 score, A9 is the more accurate predictive model for all SFs, making it the second-best predictive algorithm overall. As a result of this research, we have concluded that it provides the highest performance rate. Therefore, we can conclude that XGBoost is an effective method for predicting heart diseases. Most cases achieved an accuracy range of 85.15 to 97.57% when combining the results of multiple different ML algorithms.
The XGBoost classifier’s receiver operating characteristic (ROC) curve with SF-2 is shown in Fig. 10. This curve shows how well the model works across all classification thresholds, with an AUC of 0.98 (see Table 7).
An explainable AI method applying SHAP libraries is used to comprehend the model’s decision-making.
Figure 11 shows the importance of SHAP library XGBoost with SMOTE features.
Using ML classifiers for HD prediction is the goal of this work. The experiment findings proved that the XGBoost algorithm was the most accurate percentage for predicting the occurrence of HD. The following features are classified as important for HD prediction according to the mutual information-based feature selection approach: thalach, chol, oldpeak, age, trestbps, ca., thal, cp., exang, slope, restecg, sex, and fbs. We have used the SMOTE method to optimize hyperparameters and oversample using the data that was collected. The XGBoost technique with SMOTE produced the best results. The study reached its goal of predicting HD, with the combined datasets, and the experimental results were 97.57% for accuracy, 96.61% for sensitivity, 90.48% for specificity, 95.00% for precision, 92.68% for F1 score, and 98% for AUC.
In the context of heart disease prediction, the high accuracy of 97.57% suggests that the XGBoost model is very reliable in distinguishing between patients who do and do not have heart disease. However, it is critical to interpret what this high accuracy means in clinical practice:
-
1.
Early Detection: The model’s high sensitivity (96.61%) indicates that it can effectively identify patients in the early stages of heart disease, which is crucial for timely intervention and treatment.
-
2.
Minimizing False Positives: The specificity (90.48%) indicates a relatively low rate of false positives, implying fewer patients would receive an incorrect heart disease diagnosis, thereby reducing unnecessary anxiety and treatments.
-
3.
Balanced Prediction: The high F1 score (92.68%) reflects a balance between precision (correctly predicting heart disease when present) and recall (identifying as many true cases as possible), which is critical for practical applications where both false positives and false negatives have serious consequences.
To understand the breakthrough, the performance of XGBoost must be compared to existing gold standards in heart disease prediction, typically involving established clinical scoring systems or other ML models that have been widely accepted in healthcare.
-
1.
If XGBoost surpasses existing models, this improvement to 97.57% accuracy would signify a significant advancement, potentially offering a more reliable tool for clinicians.
-
2.
If Comparable to Current Standards: If this accuracy is only slightly better or comparable to current methods, the significance of the improvement must be critically evaluated. Factors such as the model’s generalizability, ease of integration into clinical workflows, and interpretability for healthcare professionals become critical in deciding whether it truly represents a breakthrough.
To consider this performance a breakthrough, future work must incorporate the following justifications:
-
Comparison to Baseline Methods: Show that XGBoost significantly outperforms existing prediction methods in terms of accuracy and other key metrics.
-
Clinical Impact: Discuss how this improvement could translate into better patient outcomes, such as reduced mortality or morbidity, due to more accurate early diagnosis.
-
Scalability and Implementation: Describe the potential integration of this model with current medical systems and its applicability to various patient populations in real-world settings.
The proposed ML-based heart disease prediction technique has the potential to improve healthcare. By allowing early detection and treatment, accurate heart disease prediction can greatly reduce mortality. In resource-constrained situations with limited expert access, doctors can use this tool to diagnose patients. Integrating the technology with electronic health record (EHR) systems would enable real-time risk assessments to improve patient outcomes and decision-making.
This study’s mobile app may help patients, especially those in remote places or without access to healthcare, self-assess. It lets users enter their symptoms and get a quick heart disease risk assessment, pushing them to seek medical care. This tool puts individuals in charge of their health, which may help diagnose and treat heart problems earlier.
The study contributes to the broader goal of digital health improvement by providing a scalable, cost-effective solution for heart disease prediction. By leveraging ML and explainable AI (through SHAP methodologies), the authors have created a system that is not only accurate but also interpretable, ensuring that healthcare professionals can trust and understand the predictions made by the model. This level of transparency is critical for the adoption of AI tools in clinical practice.
Moreover, the use of a mobile app extends the reach of this technology, making it accessible to a larger population. This democratization of healthcare tools aligns with global efforts to improve public health and reduce the burden of cardiovascular diseases, which are a leading cause of death worldwide.
Limitations
Despite the promising results and potential uses of the proposed ML-based technique for heart disease prediction, there are several limitations to consider:
-
1.
Dataset quality and availability: The performance and reliability of ML models depend on the quality and availability of testing and training datasets. We employed Cleveland heart disease, and private databases in our study. There may be limitations in availability, representativeness, and data quality. This limitation could make it hard to apply the proposed approach to a broader sample with a variety of additional sources.
-
2.
Imbalanced classes: SMOTE generates synthetic minority class samples to overcome class imbalance, but its effectiveness depends on the dataset and situation. Class imbalance becomes a major issue in heart disease prediction when disease frequency may be minimal. Class imbalance can cause models to perform well for the majority group but poorly for the minority class, which is often the class of interest. To address this issue, it’s essential to discuss and compare different approaches for handling class imbalances as alternatives to SMOTE such as:
-
Class Weights: Advantages include (a) being simple to implement, as it involves adjusting the misclassification penalties for different classes; (b) not requiring modifying the dataset or generating synthetic samples, making it computationally efficient; and (3) being effective in improving the model’s performance on the minority class without introducing additional complexity. The limitations include (a) assuming that the misclassification costs are known and properly specified, which may not always be the practice case; (b) assuming that the decision boundary between classes is highly non-linear or complex.
-
Ensemble Methods: Advantages include (a) naturally handling class imbalance by aggregating predictions from multiple base models trained on balanced subsets of the data; (b) tending to be robust to noise and overfitting, making them suitable for imbalanced datasets; and (c) capturing complex relationships between features and target variables, improving predictive performance. The limitations include (a) requiring more computational resources and longer training times compared to simpler algorithms; (b) not providing explicit control over the balance between classes in the final predictions.
-
Cost-sensitive Learning: Advantages include (a) explicitly considering the costs associated with misclassifying instances from different classes, allowing for fine-tuning of the model’s behavior; (b) being able to accommodate varying degrees of class imbalance and adjust the decision boundary accordingly. The limitations include (a) requiring knowledge of misclassification costs, which may be difficult to collect or subjective in real-world circumstances; and (b) making model selection more complicated by tuning more hyperparameters.
-
Anomaly Detection: Pros include (a) being able to be used in situations where the minority class represents rare or unusual events, like finding rare heart conditions; and (b) not needing explicit labeling of minority class instances, which means it can be used in either semi-supervised or unsupervised settings. The limitations include (a) assuming that the minority class instances are outliers or deviate significantly from the majority class, which may not always be the case; and (b) struggling with detecting subtle or nuanced patterns in the data, particularly when the boundary between normal and abnormal instances is ambiguous.
-
3.
Algorithm selection: To determine the optimal algorithm for predicting HD, the researchers used a variety of ML techniques. Nonetheless, the selection of algorithms is arbitrary and may affect the outcome. Other algorithms that were not considered in this study might be able to achieve different trade-offs or greater accuracy. As a result, future research should carefully consider and evaluate the ML algorithms selected.
-
4.
Domain adaptation: The use of domain adaptation techniques demonstrated the proposed system’s adaptability. The application of the proposed technique to a variety of different populations or environments may still face some limitations. More research is required to determine the technique’s efficacy in a range of populations with different lifestyles, demographics, and healthcare systems. It is also important to fully address any potential restrictions and difficulties related to domain adaptation.
-
5.
Missing data: The study does not specify if the ML model training and testing datasets contain missing data. In real-world healthcare, missing data is widespread and can dramatically impair predictive models. The discussion should focus on how missing data can lead to erroneous predictions, misdiagnosis, or delayed treatment. The authors could discuss imputation or robust algorithms for partial datasets. We must address missing data to ensure the reliability and generalizability of clinical prediction models.
Conclusions and future work
In this research study, we employed diverse methods to select features, and then applied ten distinct machine learning techniques with SMOTE to these selected features. This process allowed us to identify the most significant features that are highly effective in predicting heart disease. Every algorithm generates a unique score based on a different combination of features. We used three methods to choose features: ANOVA, chi-square, and MI. We applied these methods to three selected feature groups, namely SF-1, SF-2, and SF-3, respectively. Ten ML classifiers determined the best model and feature subset. The classifiers used were Naive Bayes, SVM, voting, XGBoost, AdaBoost, bagging, DT, KNN, RF, and LR. We employed a well-known open-access dataset and numerous cross-validation processes to evaluate the selected algorithms and measure the performance accuracy of the heart disease detection system. When compared to all other algorithms, the performance of XGBoost was more significant. The XGBoost classifier performed best with the SF-2 feature subset, with 97.64% accuracy, 96.61% sensitivity, 90.48% specificity, 95.00% precision, a 92.68% F1 score, and a 98% AUC. We developed an explainable AI method using SHAP techniques to understand how the system predicts its outcomes. Furthermore, the study demonstrated that the proposed system is adaptable using a domain adaptation approach. This work has made a significant contribution to the field of ML-based HD prediction applications by introducing unique insights and techniques. These findings have the possibility of aiding in the diagnosis and prediction of HD in Egypt and Saudi Arabia.
Finally, the authors are working on developing a smartphone app that allows users to enter symptoms and predict heart disease quickly and accurately. We will embed the best XGBoost technique in the mobile app to predict heart disease and display the detection result instantly. Because the mobile app is a symptom-based heart disease prediction, we will consider and address the impact of “dark data” during its implementation, which refers to information that exists but remains ungathered or underutilized due to data collection limitations, poor reporting, or ignorance. Unreported heart disease instances are considered “black data” when predicting heart disease. Therefore, in our future work, we will examine how dark data impacts the real-world implementation of the proposed mobile apps, specifically focusing on (1) asymptomatic cases in which patients with early-stage cardiac disease may not exhibit any symptoms. (2) Limited scope for symptom-based prediction: The models consider only a limited set of symptoms. Due to diagnostic testing and imaging examinations, heart disease can manifest in ways other than typical symptoms, so focusing solely on symptoms may miss critical signs. (3) Data collection: The absence of asymptomatic cases could potentially impact the quality of the app’s prediction model dataset. If most of the training data is symptomatic, the model’s predictions may prioritize symptomatic presentations, thereby intensifying the dark data effect.
To address the limitations imposed by the dark data effect and enhance the real-world applicability of the mobile app for heart disease prediction, we will consider several strategies, such as (1) comprehensive risk assessment: Expand the scope of the predictive model to incorporate additional risk factors beyond symptoms, such as demographic information, medical history, lifestyle factors, and biomarkers. (2) Integration with diagnostic tools: Connect the mobile app to diagnostic tools or wearable devices capable of measuring physiological parameters associated with heart health, such as blood pressure, heart rate variability, or electrocardiogram (ECG) signals. (3) Population screening programs: Partner with healthcare providers or public health agencies to promote population screening programs aimed at identifying individuals with undiagnosed heart disease. (4) Education and awareness campaigns: Launch educational initiatives to raise awareness about the importance of regular cardiovascular screenings, even in the absence of symptoms. The authors will also consider a cost-effectiveness argument for a heart disease prediction app, provide evidence, and consider a variety of factors, including development, implementation, and maintenance costs. Claiming that certain features are cheaper necessitates a thorough evaluation and comparison with alternative approaches to ensure the claim’s validity.
Due to their inability to explain the decision-making process, ML-based system developers tend to treat AI-based apps as a mystery. In this study, we used SHAP and feature importance techniques to explain and interpret the prominent features that were most influential in the decision. In our future work, we plan to expand our research by incorporating other explainable artificial intelligence (XAI) techniques that can improve transparency and interpretability, such as: (1) Partial Dependence Plots (PDPs): PDPs show the link between a feature and the expected outcome while ignoring other features. By showing each feature separately, we explicitly understand its effect on heart disease prediction. (2) Individual Conditional Expectation (ICE) Plots: ICE plots show how a feature affects each data point, not just the average. The effects of feature changes on various people can be better understood. (3) Local Interpretable Model-Agnostic Explanations (LIME): To explain predictions, LIME develops local surrogate models around specific examples. By changing the input data and watching how the predictions change, LIME gets close to the model’s local behavior and gives simple explanations for specific predictions. (4) Rule-based models, like decision trees or rule lists, connect input features directly to predictions. These models explain the criteria for heart disease, providing transparency. By employing these explainable AI methods, machine learning-based systems for heart disease prediction can provide healthcare professionals and patients with transparent, interpretable, and actionable insights, facilitating informed decision-making and improving trust in AI-driven healthcare applications.
Data availability
The corresponding author will share study datasets upon reasonable request.
References
World Health Organization. Cardiovascular Diseases (CVDs). Available online: (2023). https://www.afro.who.int/health-topics/cardiovascular-diseases, (accessed on 5 May).
Alom, Z. et al. Early Stage Detection of Heart Failure Using Machine Learning Techniques. In Proceedings of the International Conference on Big Data, IoT, and Machine Learning, Cox’s Bazar, Bangladesh, 23–25 September (2021).
Gour, S., Panwar, P., Dwivedi, D. & Mali, C. A machine learning approach for heart attack prediction. In Intelligent Sustainable Systems (eds Nagar, A. K., Jat, D. S., Marín-Raventós, G. & Mishra, D. K.) 741–747 (Springer, Singapore, 2022). https://doi.org/10.1007/978-981-16-6309-3_70.
Gupta, C., Saha, A., Reddy, N. S. & Acharya, U. D. Cardiac Disease Prediction using Supervised Machine Learning Techniques. In Journal of Physics: Conference Series; IOP Publishing: Bristol, UK, Volume 2161, p. 012013 (2022).
Shameer, K. et al. Machine learning predictions of cardiovascular disease risk in a multi-ethnic population using electronic health record data. Int. J. Med. Informatics. 146, 104335 (2021).
Liu, M. et al. Deep learning-based prediction of coronary artery disease with CT angiography. Japanese J. Radiol. 38 (4), 366–374 (2020).
Zakria, N., Raza, A., Liaquat, F. & Khawaja, S. G. Machine learning based analysis of cardiovascular disease prediction. J. Med. Syst. 41 (12), 207 (2017).
Yang, M., Wang, X., Li, F. & Wu, J. A machine learning approach to identify risk factors for coronary heart disease: a big data analysis. Comput. Methods Programs Biomed. 127, 262–270 (2016).
Ngufor, C., Hossain, A., Ali, S. & Alqudah, A. Machine learning algorithms for heart disease prediction: a survey. Int. J. Comput. Sci. Inform. Secur. 14 (2), 7–29 (2016).
Shoukat, A., Arshad, S., Ali, N. & Murtaza, G. Prediction of Cardiovascular diseases using machine learning: a systematic review. J. Med. Syst. 44 (8), 162. https://doi.org/10.1007/s10916-020-01563-1 (2020).
Shankar, G. R., Chandrasekaran, K. & Babu, K. S. An analysis of the potential use of Machine Learning in Cardiovascular Disease Prediction. J. Med. Syst. 43 (12), 345. https://doi.org/10.1007/s10916-019-1524-8 (2019).
Khandadash, N., Ababneh, E. & Al-Qudah, M. Predicting the risk of coronary artery disease in women using machine learning techniques. J. Med. Syst. 45, 62. https://doi.org/10.1007/s10916-021-01722-6 (2021).
Moon, S., Lee, W. & Hwang, J. Applying machine learning to Predict Cardiovascular diseases. Healthc. Inf. Res. 25 (2), 79–86. https://doi.org/10.4258/hir.2019.25.2.79 (2019).
Lakshmi, M. & Ayeshamariyam, A. Machine learning techniques for Prediction of Cardiovascular Risk. Int. J. Adv. Sci. Technol. 30 (3), 11913–11921. https://doi.org/10.4399/97888255827001 (2021).
Md, R. et al. Early detection of cardiovascular autonomic neuropathy: A multi-class classification model based on feature selection and deep learning feature fusion. Information Fusion, vol. 77, P 70–80, January (2022).
Wongkoblap, A., Vadillo, M. A. & Curcin, V. Machine learning classifiers for early detection of Cardiovascular Disease. J. Biomed. Inform. 88, 44–51. https://doi.org/10.1016/j.jbi.2018.09.003 (2018).
Delavar, M. R., Motwani, M. & Sarrafzadeh, M. A. Comparative study on feature selection and classification methods for Cardiovascular Disease diagnosis. J. Med. Syst. 39 (9), 98. https://doi.org/10.1007/s10916-015-0333-5 (2015).
Yong, K., Kim, S., Park, S. J. & Kim, J. A. Clinical decision support system for Cardiovascular Disease Risk Prediction in type 2 diabetes Mellitus patients using decision Tree. Comput. Biol. Med. 89, 413–421. https://doi.org/10.1016/j.compbiomed.2017.08.024 (2017).
Mirza, Q. Z., Siddiqui, F. A. & Naqvi, S. R. The risk prediction of cardiac events using a decision Tree Algorithm. Pakistan J. Med. Sci. 36 (2), 85–89. https://doi.org/10.12669/pjms.36.2.1511 (2020).
Farag, A., Farag, A. & Sallam, A. Improving Heart Disease prediction using boosting and bagging techniques. Proc. Int. Conf. Innovative Trends Comput. Eng. (ITCE). 90-96https://doi.org/10.1109/ITCE.2016.7473338 (2016).
Jhajhria, S. & Kumar, R. Predicting the risk of Cardiovascular diseases using ensemble learning approaches. Soft. Comput. 24 (7), 4691–4705. https://doi.org/10.1007/s00500-019-04268-8 (2020).
Samadiani, N., Moghadam, E., Motamed, C. & A. M., & SVM-based classification of Cardiovascular diseases using feature selection: a high-dimensional dataset perspective. J. Med. Syst. 40 (11), 244. https://doi.org/10.1007/s10916-016-0573-7 (2016).
Zhang, X., Zhang, Y., Du, X. & Li, B. Application of XGBoost algorithm in clinical prediction of coronary heart disease. Chin. J. Med. Instrum. 43 (1), 12–15 (2019).
Liu, Y., Li, X. & Ren, J. A comparative analysis of machine learning algorithms for heart disease prediction. Comput. Methods Programs Biomed. 200, 105965 (2021).
Hussein, N. S., Mustapha, A. & Othman, Z. A. Comparative study of machine learning techniques for heart disease diagnosis. Comput. Sci. Inform. Syst. 17 (4), 773–785 (2020).
Akbar, S., Tariq, R. & Basharat, A. Heart disease prediction using different machine learning approaches: a critical review. J. Ambient Intell. Humaniz. Comput. 11 (5), 1973–1984 (2020).
Zarshenas, A., Ghanbarzadeh, M. & Khosravi, A. A comparative study of machine learning algorithms for predicting heart disease. Artif. Intell. Med. 98, 44–54 (2019).
Kaur, I. & Singh, G. Comparative analysis of machine learning algorithms for heart disease prediction. J. Biomed. Inform. 95, 103208 (2019).
Li, Y., Jia, W. & Li, J. Comparing different machine learning methods for predicting heart disease: a telemedicine case study. Health Inform. Sci. Syst. 6, 7 (2018).
Zhang, X., Zhou, Y. & Xie, D. Heart disease diagnosis using machine learning and expert system techniques: a survey paper. J. Med. Syst. 42 (7), 129 (2018).
Wu, J. & Roy, J. Stewart, & W. F. A comparative study of machine learning methods for the prediction of heart disease. Journal of Healthcare Engineering, 7947461 (2017). (2017).
Ahmed, Z., Mohamed, K. & Zeeshan, S. Comparison of machine learning algorithms for predicting the risk of heart disease: A systematic review. Journal of Healthcare Engineering, 7058278 (2016). (2016).
Chen, X., Hu, Z. & Cao, Y. Heart disease diagnosis using decision tree and naïve Bayes classifiers. World Congress Med. Phys. Biomedical Eng. 14, 1668–1671 (2007).
Pedregosa, F. et al. Scikit-learn: machine learning in python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Hosam El-Sofany, S. A., El-Seoud, O. H., Karam, Yasser, M., Abd El-Latif, Islam, A. T. F. & Taj-Eddin A Proposed Technique Using Machine Learning for the Prediction of Diabetes Disease Through a Mobile App. International Journal of Intelligent Systems, volume ID 6688934, (2024). https://doi.org/10.1155/2024/6688934, (2024).
Chintan, M. B., Parth, P., Tarang, G. & Pier, L. M. Effective Heart Disease Prediction Using Mach. Learn. Techniques Algorithms, 16, 88, https://doi.org/10.3390/a16020088, (2023).
Acknowledgements
The authors extend their appreciation to the Deanship of Research and Graduate Studies at King Khalid University for funding this work through small group research under grant number (RGP1/129/45).
Author information
Authors and Affiliations
Contributions
Hosam El-Sofany: Create the original concept for the research. Methodology, design, and implementation. Writing, reviewing, and editing. Proofreading and checking against plagiarism using the iThenticate program provided by King Khalid University. Belgacem Bouallegue: Methodology, design, writing, reviewing, and editing. Yasser M. Abd El-Latif: Methodology, design, writing, reviewing, and editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
This study did not involve human participants, human tissue, or any personal data. The dataset used for this research is publicly available and anonymized, and no ethical approval or informed consent was required.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
El-Sofany, H., Bouallegue, B. & El-Latif, Y.M.A. A proposed technique for predicting heart disease using machine learning algorithms and an explainable AI method. Sci Rep 14, 23277 (2024). https://doi.org/10.1038/s41598-024-74656-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-74656-2
Keywords
This article is cited by
-
Polynomial-SHAP as a SMOTE alternative in conglomerate neural networks for realistic data augmentation in cardiovascular and breast cancer diagnosis
Journal of Big Data (2025)
-
Dual level dengue diagnosis using lightweight multilayer perceptron with XAI in fog computing environment and rule based inference
Scientific Reports (2025)
-
Context matters in machine learning based disease prediction with insights from diverse clinical and symptom data
Scientific Reports (2025)
-
Harris Hawks optimization based deep learning models for heart disease diagnosis
Scientific Reports (2025)
-
Dual explainability framework for heart disease prediction using LIME and permutation feature importance
Discover Applied Sciences (2025)