Abstract
Oral Squamous Cell Carcinoma (OSCC) causes a severe challenge in oncology due to the lack of diagnostic devices, leading to delays in detecting the disorder. The OSCC diagnosis through histopathology demands a pathologist expert because the cellular presentation is variable and highly complex. Existing diagnostic approaches for OSCC have specific efficiency and accuracy restrictions, highlighting the necessity for more reliable techniques. The increase of deep neural networks (DNN) model and their applications in medical imaging have been instrumental in disease diagnosis and detection. Automatic detection systems using deep learning (DL) approaches show tremendous promise in investigating medical imagery with speed, efficiency, and accuracy. In terms of OSCC, this system allows the diagnostic method to be streamlined, facilitating earlier diagnosis and enhancing survival rates. Automatic analysis of histopathological image (HI) can assist in accurately detecting and identifying tumorous tissue, reducing diagnostic turnaround times and increasing the efficacy of pathologists. This study presents a Squeeze-Excitation with Hybrid Deep Learning for Oral Squamous Cell Carcinoma Recognition (SEHDL-OSCCR) on HIs. The presented SEHDL-OSCCR technique mainly focuses on detecting oral cancer (OC) using hybrid DL models. The bilateral filtering (BF) technique is initially used to remove the noise. Next, the SEHDL-OSCCR technique employs the SE-CapsNet model to recognize the feature extractors. An improved crayfish optimization algorithm (ICOA) technique is utilized to improve the performance of the SE-CapsNet model. At last, the classification of the OSCC technique is performed by employing a convolutional neural network with a bidirectional long short-term memory (CNN-BiLSTM) model. The simulation results obtained using the SEHDL-OSCCR technique are investigated using a benchmark medical image dataset. The experimental validation of the SEHDL-OSCCR technique illustrated a greater accuracy outcome of 98.75% compared to recent approaches.
Similar content being viewed by others
Introduction
OC is the most prevalent cancer worldwide and is considered by late diagnosis, disease, and high death rates1. Using tobacco in any form and excessive alcohol consumption are critical risk factors that lead to OC. The cause ratio is very peak in South and Southeast Asia, which is the consumption of betel quid, which generally contains slaked lime, areca nut, and betel leaves containing tobacco2. These are easy commercials in packets and are common in public owing to robust advertising strategies. OC is usually connected with delay in presenting, especially in LMICs, but more than two-thirds present at the final stages and as outcome survival rates are worst3. Treatment of cancer, specifically at the final stages, is costly. The lack of public awareness and awareness of the medical profession regarding OC is a significant cause of late recognition. Several imaging processes can take images of oral lesions using various methods4. A binary-method image processor that incorporates autofluorescence and white light image is suggested to identify OC. Oral lesions presented in the final stage have an injurious effect on survival rates, with more than two-thirds of oral lesions being identified at the final stages5. The lesion care cost is expensive, especially in the final stage. Late analysis of oral lesions is a significant concern for medicinal staff. To enhance the early diagnosis of OC and decrease the control of late analysis, it is vital to improve an automatic method for recognizing OC with less human interface6.
Machine learning (ML) was established to help improve classifier accuracy in automatic methods. Particularly, DL is demonstrated to decrease the requirement for human contribution in the study of massive databases7. AI might address recent challenges in diagnosing and identifying the estimation of OC by decreasing the workloads, difficulty, and exhaustion for doctors over complex procedures. It is a technical improvement that has collected significant attention from experts globally as it emulates human cognitive proficiencies. In dentistry, its application is comparatively new growth, but the outcomes are encouraging8. OSCC remains a leading global health concern due to its often late diagnosis and high mortality rates. Addressing OSCC efficiently is challenging, specifically in areas with high incidence associated with specific lifestyle factors9. Enhancements in early detection and accurate diagnosis are significant for enhancing patient results. Implementing cutting-edge DL approaches can substantially improve diagnostic accuracy and assist in timely intervention, ultimately mitigating the impact of this disease. Enhancing diagnostic tools for OSCC is significant, as early and precise detection can significantly improve survival rates and mitigate the burden on healthcare systems, emphasizing the requirement for innovative techniques in cancer detection and assessment10.
This study presents a Squeeze-Excitation with Hybrid Deep Learning for Oral Squamous Cell Carcinoma Recognition (SEHDL-OSCCR) technique on HIs. The presented SEHDL-OSCCR technique mainly concentrates on detecting OC using hybrid DL methods. The bilateral filtering (BF) technique is employed to eliminate noise. Next, the SEHDL-OSCCR technique utilizes the SE-CapsNet model to recognize the feature extractors. An enhanced crayfish optimization algorithm (ICOA) approach is used to improve the performance of the SE-CapsNet model. Finally, the OSCC classification is accomplished by designing a convolutional neural network using a bidirectional long short-term memory (CNN-BiLSTM) technique. The obtained simulation outcomes of the SEHDL-OSCCR technique are examined by employing a benchmark medical image dataset. The key contribution of the SEHDL-OSCCR technique is as follows.
-
The BF approach was employed to efficiently mitigate noise in the images, thus improving the overall quality of the image. This preprocessing step is significant for enhancing subsequent feature extraction and classification processes. The technique also confirms more precise and more dependable data for evaluation.
-
The SE-CapsNet method was implemented for advanced feature extraction and recognition, allowing more accurate detection of relevant image characteristics. This methodology substantially improves the approach’s capability to distinguish between diverse features. By employing the SE-CapsNet model, the evaluation attains greater accuracy in recognizing intrinsic patterns.
-
Incorporating the ICOA model elevated the accomplishment of the SE-CapsNet technique. This optimization method fine-tunes the model’s parameters, improving feature extraction and recognition accuracy. Using the ICOA model substantially enhances the comprehensive efficiency in processing and classifying data.
-
The SEHDL-OSCCR technique implements the CNN-BiLSTM model to accomplish precise OSCC classification. This incorporation employs the CNN model for feature extraction and the BiLSTM technique for capturing temporal dependencies, resulting in improved accuracy. The method efficiently enhances the model’s ability to classify OSCC.
-
more accurately.
-
The SEHDL-OSCCR technique presented a novel integrated model by integrating SE-CapsNet with the ICOA and CNN-BiLSTM models. This innovative methodology improves both feature extraction and classification accuracy for OSCC detection. By incorporating these advanced methods, the technique attains greater accomplishment in detecting and classifying oral cancer.
The article is structured as follows: section “Literature review” presents the literature review, section “Modeling of SEHDL-OSCCR technique” outlines the proposed method, section “Result analysis and discussion” details the results evaluation, and section “Conclusion” concludes the study.
Literature review
In11, an intelligent SmSL method is specially aimed to depend upon Self-supervised Pre-training (SP) and Adaptive Threshold (AT), called SPAT_SmSL. At first, the SP and AT models are employed to develop the unlabeled data, combined into the SPAT_SmSL model for identifying stroma, cancer, and tumour-infiltrating lymphocyte (TIL) areas. Next, pathological variables containing depth of invasion (DOI) and TIL-score were numerically measured and built on the outcomes of image detection. Fati et al.12 used hybrid models based on combined features. The 1st developed model depends on a hybrid technique of CNN methods (ResNet18 and AlexNet) and the support vector machines (SVMs) technique. The 2nd projected technique depends upon the hybrid feature, which is removed by CNN models and united with the texture, colour, and shape properties removed utilizing the local binary pattern (LBP), fuzzy colour histogram (FCH), discrete wavelet transforms (DWT), and grey-level co-occurrence matrix (GLCM) techniques. The principal component analysis (PCA) technique has decreased the dimension and led to the artificial neural networks (ANNs) technique. In13, the classification of OSCC histopathology imageries is executed utilizing dual-developed techniques. In the 1st method, transfer learning (TL) aided deep CNN (DCNN) techniques are measured. In the 2nd model, a base DCNN structure, trained from scratch with ten convolutional layers, is projected. In14, an innovative technique is presented for the early classification of OSCC using DL models. The method includes a Cyclic Learning Rate (CLR) approach for dynamic alteration of the learning rate through the training model. ResNet18 profits from skip connection to improve the gradient flow, whereas DenseNet and AlexNet structures considerably donate to exact image classification. Begum and Vidyullatha15 aim to automatically identify malignant and benign oral biopsy HIs by applying a DL-based CNN technique for the early analysis of OSCC model. The four currently proposed candidates pretrained DL-CNN techniques, namely InceptionNet, NASNetLarge, DenseNet201, and Xception, are chosen for the TL method in this study. These pre-trained techniques are then adapted with extra layers for effectual OSCC recognition.
In16, a CAD technique has been developed. Feature extraction was executed from this database utilizing 4 DL techniques: AlexNet, ResNet50, VGG16, and InceptionV3. Binary PSO (BPSO) has been employed to select the finest feature. Once the finest features were removed and nominated, they were categorized utilizing the XGBoost. In17, an advanced plan for OC classification is presented. This contains the Feature Fusion DCNN with SGD-based LR for the analysis of OC. The following layers, containing pooling, fusion, and transformation, are intended to grip ranked features across dissimilar branches. Lastly, the developed method experiences training through LR on the removed data utilizing the Cross-Entropy Loss, and the optimizer (SGD with weight decay) was used to upgrade the model parameter. Ahmad et al.18 introduced a hybrid mechanism based on combined features. The primary tactic is TL utilizing the techniques of Inceptionv3, Xception, NASNetLarge, InceptionResNetV2, and DenseNet201. Next, it includes a pre-trained CNN model for the feature extractor and an SVM for identification. Notably, features were removed utilizing numerous pre-trained techniques. The last tactic uses an innovative hybrid feature fusion model, employing a CNN extraction technique. Kadhim and Mohammed19 analyze the current AI methods in kidney cancer diagnosis, compute their efficiency, explore future research areas, and address threats to enhance patient outcomes and treatment effectiveness. In20, the authors integrate various omics data by implementing the Quantum Cat Swarm Optimization (QCSO) technique for feature selection (FS), incorporated with K-means clustering and SVM technique, attaining improved performance and interpretability of the model.
Das et al.21 present a framework utilizing a two-phase methodology: TL with CNN methods in the initial phase and constructing an ensemble method with the top-performing CNNs in the subsequent phase. The presented classifier is compared with advanced approaches such as AlexNet, ResNet, InceptionNet, and XceptionNet. Das, Dash, and Mishra22 introduce a convolutional neural network (CNN) technique for the automatic and early detection of OSCC, utilizing histopathological oral cancer images for experimentation. Meer et al.23 propose a fully automated architecture incorporating Self-Attention CNN and Residual Network techniques with fusion and optimization. It comprises augmenting training and testing samples, then training two deep techniques: a Self-Attention MobileNet-V2 and a Self-Attention DarkNet-19, with hyperparameters tuned utilizing the Whale Optimization Algorithm (WOA) technique. Features from both methods are integrated using the Canonical Correlation Analysis (CCA) model, additionally refined with Quantum WOA technique in order to choose relevant features, which are later classified by implementing wide neural network models. Raj and Muneeswari24 introduce the Optimal Archimedes Shooty Tern Deep Network (OASTDN) approach. This method also utilizes a Deep Belief Network (DBN) model with weights optimized by a novel Archimedes Shooty Tern Optimization Algorithm (ASTOA) technique, integrating Archimedes Optimization Algorithm (AOA) and Shooty Tern Optimization (STO) approaches. Shukla, Ajwani, Sharma, and Das25 propose a novel machine vision methodology by implementing conventional supervised DL approaches. The proposed methodology also detects the nucleus in cancerous biopsy images, extracts it employing K-means clustering with thresholding, and implements a new classification technique for final cancer detection.
The limitations of the existing studies include its dependence on self-supervised pre-training and adaptive thresholds, which may restrict generalizability. Another study implements complex hybrid techniques integrating CNNs with SVM techniques, potentially paving the way to overfitting issue. Techniques comparing pre-trained methods with those trained from scratch may only partially capture the advantages of the TL technique. Some methodologies concentrate on novel architectures or hybrid mechanisms, but their effectiveness might require to be validated more extensively, or difficulties with FS and computational complexity could be encountered. Models incorporating diverse omics data or implementing new optimization approaches may need assistance with model interpretability or additional validation. Furthermore, techniques based on conventional supervised DL might have limited adaptability to several datasets or scenarios. The present research on cancer detection faces gaps in generalizability across diverse datasets and validation of novel optimization models. Moreover, there is a requirement for enhanced model adaptability and interpretability in intrinsic hybrid techniques and integration models.
Modeling of SEHDL-OSCCR technique
The solution framework
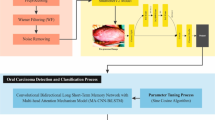
This study presented a new SEHDL-OSCCR approach to HIs. The technique mainly focuses on detecting OC using hybrid DL models. To accomplish that, the SEHDL-OSCCR approach contains distinct processes such as noise reduction, a SE-CapsNet-based feature extractor, ICOA-based parameter tuning, and a classifier selection process. Figure 1 demonstrates the entire flow of the SEHDL-OSCCR method.
Noise reduction
Initially, the BF technique is used to remove the noise. BF is an adaptable image processing method that intends to smooth images while maintaining edges26. BF was selected for the noise reduction process due to its capability to conserve edges while eliminating noise efficiently, which is significant in maintaining the integrity of crucial image features. Unlike other noise reduction methodologies that may blur or distort the image, BF selectively smooths regions depending on spatial distance and intensity differences, confirming that edges and fine details remain sharp. This makes BF specifically advantageous for processing medical images, where conserving substantial data is essential for precise evaluation. Its adaptability to varying noise levels and its capability to balance noise reduction with edge preservation offer crucial enhancements over other models, giving clearer and more reliable input for subsequent image processing tasks. Figure 2 shows the structure of the BF methodology.
BF considers the intensity variances amongst adjacent pixels in different classical smoothing filters that employ a weighted average solely based on spatial distance. This ensures that pixel with related intensity is smoothed together, maintaining edge sharpness. It is instrumental in applications where edge preservation and noise reduction are vital, like HDR tone mapping, computer vision, and image denoising. BF provides a flexible tool by adjusting parameters such as spatial distance and intensity similarity for manipulating and enhancing digital images with influence upon edge preservation and smoothing.
Architecture of the SE-CapsNet
Next, the SEHDL-OSCCR technique utilizes the SE-CapsNet model to recognize the feature extractors27. The SE-CapsNet method was chosen for recognizing feature extractors due to its advanced abilities in capturing and processing intrinsic features in data. Its architecture integrates the merits of capsule networks with squeeze-and-excitation blocks, improving the capability of the technique to learn and represent complex patterns more efficiently. Unlike conventional CNNs, SE-CapsNet can handle feature scale and context discrepancies better, paving the way to more accurate and robust feature extraction. This makes it suitable for applications needing detailed and precise analysis, namely recognizing subtle features in medical images. Its enhanced accomplishment in capturing spatial hierarchies and relationships gives a crucial advantage over other methods, which may need more depth in feature representation. The DL applications in the malware recognition area have attained outstanding research attainments; mainly, the CapsNet examines the relations among features, and this method has benefits that will be functional to lesser models. This research work unites a mechanism of channel attention, named SE block, by the CapsNet to construct the SE-CapsNet method, which mainly contains four layers. Figure 3 illustrates the architecture of the SE-CapsNet approach.
-
1)
CONVOLUTIONAL LAYER
The convolutional is the 1st layer, whose main aim is to remove local features utilizing \(\:3\text{x}3\) convolutional kernels with a \(\:step\:size\) of one in a mixture with \(\:ReLU\).
-
2)
Generally, the SE layer is effortless to utilize and can recover the feature extraction capability. So, it is beneficial to identify and contain dual processes such as squeeze and excitation. The squeeze process’s main aim was to attain an assumed network’s overall feature. \(\:{u}_{C}\) denotes the \(\:Cth\) feature mapping that yields by the convolutional layer. The channel-wise statistics \(\:{z}_{C}\) can be obtained in the global average pooling. The excitation process is the procedure of channel weight, whereas \(\:\sigma\:\) signifies sigmoid; \(\:\delta\:\) represents ReLU; \(\:{W}_{1}\) and \(\:{W}_{2}\) correspondingly denote the decreasing and increasing dimensionality activities. Over the excitation process, a non-linear interaction can be attained among networks, and the channel weights can be achieved. Lastly, over a scaling process, the weight of the channel is increased by the novel feature mappings to get the attention of the feature map as the yield of the SE layer. The Eqs. (1)-(3) shows the mathematical formulations for this layer.
$$z_{C} = F_{{sq}} \left( {uc} \right) = \frac{1}{{H \times W}}\sum\limits_{{i = 1}}^{H} {\sum\limits_{{j = 1}}^{W} {u_{C} \left( {i,j} \right)} }$$(1)$$s = F_{{ex}} \left( {z,W} \right) = \sigma \left( {g\left( {z,W} \right)} \right) = \sigma \left( {W_{2} \delta \left( {W_{z} } \right)} \right)$$(2)$$\widetilde{{x_{C} }} = F_{{scale}} \left( {u_{C} ,s_{C} } \right) = s_{C} \cdot u_{C}$$(3) -
3)
PRIMARYCAPS LAYER
Next, as a PrimaryCaps input, every feature map is taken using its equivalent attention weight. This layer is dissimilar from the standard convolutional layer. As per the explanation, once this layer is completed, capsules, also known as vectors, can store more data.
-
4)
DIGITCAPS LAYER
This layer is mainly employed to keep non-Ponzi and Ponzi capsules. The vector primarily signifies the last outcome. The CapsNet employed a squash function. When preserving the vector route, the output vector length was used as the potential of the present entity. The mathematical equations among capsules \(\:i\) and \(\:j\) were exposed in Eqs. (4)- (6):
$$\hat{u}_{{j1}} = W_{{ij}} u_{i}$$(4)$$s_{j} = \mathop \sum \limits_{i} c_{{ij}} \hat{u}_{{j|i}} ~$$(5)$$v_{j} = \frac{{\left\| {s_{j} } \right\|^{2} }}{{1 + \left\| {s_{j} } \right\|^{2} }}\frac{{s_{j} }}{{\left\| {s_{j} } \right\|}}$$(6)
Whereas, \(\:{W}_{ij}\) signifies the matrix of weight, indicating the association among capsules \(\:i\) capsule \(\:j\); \(\:{\widehat{u}}_{j|i}\) indicates the forecast that the \(\:ith\) lower-level capsule creates the \(\:jth\) higher-level capsule. \(\:{c}_{ij}\) refers to the concatenation factor attained over a dynamic route. The outcome of \(\:{v}_{j}\) is ranked through the result of the last squash function.
DL fine-tuning process
The ICOA technique is utilized to improve the performance of the SE-CapsNet model. The COA model is a newly presented SI approach28. The ICOA technique was selected for hyperparameter tuning due to its improved efficiency and effectiveness in navigating complex parameter spaces. ICOA integrates enhancements that refine the conventional crayfish optimization model, resulting in more precise and faster convergence on optimal hyperparameters. Its capability to handle non-linear and multi-modal optimization issues makes it appropriate for tuning parameters in DL methods. Compared to other techniques, the ICOA technique performs better in balancing exploration and exploitation, mitigating the likelihood of overfitting, and attaining improved model accuracy. This makes the ICOA approach compelling for optimizing hyperparameters in complex models where conventional techniques might face difficulties. Figure 4 specifies the overall structure of the ICOA model.
This method proposes to define the optimum performance of the problem by mimicking the competitive, heat avoidance, and foraging strategies. It contains two major phases, namely exploration and exploitation. This method employs \(\:X\) to signify the primary population position, whereas \(\:{X}_{ij}\) denotes the crayfish (CF) position \(\:i\) from dimensional \(\:j\). The value of \(\:{X}_{ij}\) has been computed utilizing the subsequent formula:
Whereas \(\:l{b}_{j}\) signifies the lower boundaries of the \(\:{j}^{th}\) dimensional, \(\:u{b}_{j}\) signifies the upper boundaries of the \(\:{j}^{th}\) dimensional, and \(\:rand\) signifies the random number.
CF thrives in environments with temperatures from 15-\(\:3{0}^{o}C\). If the temperature variable “temp” goes beyond \(\:3{0}^{o}C\), CF searches for refuge in caves. Under the condition that the count of accessible caves is restricted, cave scrambling events could occur. \(\:rand<0.5\) has been deployed to define the lack of cave scrambling event. The subsequent equation can be employed to indicate the CF’s entrance into the cave.
In which \(\:t\) represents the present iteration number, \(\:{C}_{2}\) implies the lessening curve, and \(\:{X}_{shade}\) stands for the cave position.
CF is employed in competition for burrows once the temperature exceeds 30 and the random variable is superior to or equivalent to 0.5.
Whereas \(\:z\) denotes the random individual of CF; \(\:z=round\left(rand\times\:\left(N-1\right)\right)+1.\)
If “temp” ≤ 30, then CF starts feeding. Because of its restricted body size, CF displays 2 different feeding behaviours. If the food was substantial, CF deployed its claws to shred it into controllable pieces previously feeding, employing its 2nd and 3rd walking feet in changing patterns. In contrast, if the size of the food can be appropriate, CF involves indirect feeding. The behaviours of foraging for normal and huge-size feed are defined as:
In this case, \(\:{x}_{food}\) signifies the food place, and \(\:p\) defines the mathematical equation of CF intake. Sin and cos functions can be deployed to demonstrate the changing feeding CF behaviour.
During distinct phases of the algorithm, the CF symbolizes the optimum performance once it enters the cave and eats the food. By constantly upgrading the CF position, it remains near the target variable, accomplishing the method’s optimizer function. The pseudocode for COA is depicted in Algorithm 1.
The COA population-initialized approach restricts the speed and direction with the optimum result found, affecting the process’s entire efficiency. Chaotic mapping has been presented to enhance the population initialized process and improve its global searching ability. Chaotic mapping was employed for orders that reveal features of ergodicity, arbitrariness, and orbital instability. Generally utilized chaotic mapping comprises tent mapping, sin mapping, logistic mapping, circle mapping, and singer mapping. The circle map has been recognized for its constancy and wide range of chaotic rates. During this case, the circle map has been deployed for initializing the population CF as:
On the other hand, \(\:n\) refers to the size of the solutions.
The ICOA is used to derive an FF to enhance the efficiency of a classifier. It defines a positive number to signify the superior performance of the candidate solution. In this research, the minimizer of the classifier rate of error is measured as the FF, as set in Eq. (13).
Classifier selection
At last, the classification of OSCC is performed by utilizing the CNN-BiLSTM model29. LSTM is a different RNN structure, as presented by S. Hochreiter and J. Schmidhuber in 1997. Its internal configuration involves the memory cell, forget, input, and output gates. The CNN-BiLSTM technique was chosen for OSCC classification due to its unique capability to integrate CNNs with BiLSTM networks. The CNN component outperforms in extracting spatial features from histopathological images, capturing complex patterns and structures. The BiLSTM approach improves this by capturing temporal dependencies and contextual data from sequential data, enhancing the technique’s understanding of intrinsic patterns. This incorporation allows for more precise classification by employing spatial and sequential features, making it specifically efficient for the complex evaluation needed in OSCC detection. The incorporation of CNN and BiLSTM presents a robust approach that can outperform other techniques that may only partially capture the temporal and spatial aspects of the data. Figure 5 illustrates the structure of the CNN-BiLSTM model.
Eventually, integrating these two layers and the forget gate upgrades the LSTM storage cell based on Eq. (17).
In which, \(\:{i}_{t},{\:f}_{t}\), and \(\:{o}_{t}\) refer to the input, forget, and output gates at a \(\:t\) time step of the LSTM cell; correspondingly, \(\:{C}_{t}\) implies the candidate cell layer describing the long-term memory at time step \(\:t\), \(\:{h}_{t-1}\) defines the outcome of the preceding time step \(\:t-1\), \(\:{x}_{t}\) illustrates the input of present time step \(\:t\),\(\:\:\sigma\:\) demonstrates the sigmoid activation function, \(\:{b}_{i}\) stands for the bias term of input gate, \(\:{w}_{ix}\) illustrates the weight factor of the input gate \(\:{x}_{t}\), \(\:{w}_{hich}\) shows the weighted factor of the preceding hidden layer \(\:{h}_{t-1}\), \(\:{b}_{f}\) indicates the biased term of forget gate, \(\:{w}_{fx}\) represents the weighted factor of forget gate \(\:{x}_{t}\),\(\:\:{b}_{c}\) signifies the bias term, \(\:{w}_{cx}\) denotes the weighted coefficient of cell \(\:{x}_{t}\), \(\:{w}_{fh}\) defines the preceding hidden layer \(\:{h}_{t-1}\), \(\:{w}_{ch}\) indicates the weighted coefficient of cell \(\:{h}_{t-1}\), \(\:{b}_{o}\) demonstrates the bias term of output gate, \(\:{w}_{ox}\) refers to the weighted coefficient of the output gate \(\:{x}_{t}\), \(\:{w}_{oh}\) defines the weighted coefficient of output gate \(\:{h}_{t-1}.\)
Write Eq. (14) in vector method as follows
Eqs. (15) and (16) are expressed as:
whereas \({y}_{1}= [i_{t} f_{t} o_{t} ]^{T}\), \(w_{1} = \left[ {\begin{array}{*{20}l} {w_{{ix}} w_{{fx}} w_{{ox}} } \hfill \\ {\:w_{{ih}} w_{{fh}} w_{{oh}} } \hfill \\ \end{array} } \right]\), \(x_{1} = [x_{t} h_{{t - 1}} ]^{T}\), \(b_{1} = [b_{i} b_{f} b_{o} ]^{T}\), \(y_{2} = [c_{t} ]\), \(w_{2} = [w_{{cx}} w_{{ch}} ]^{T}\), \(x_{2} = [x_{t} h_{{t - 1}} ]^{T}\) and \({b}_{2}=\left[{b}_{c}\right]\). The resultant of LSTM cell for every time step is
In this case, \(\:{c}_{t}\) represents the cell layer at time step \(\:t\), and \(\:{c}_{t-1}\) defines the cell layer from the preceding time step \(\:t-1.\)
The architecture of the BiLSTM network, whereas \(\:{x}_{t}\) refers to the input data at time step \(\:t\), \(\:{\overrightarrow{h}}_{t}=\left({\overrightarrow{h}}_{1},{\overrightarrow{h}}_{2},\:\cdots\:,\:{\overrightarrow{h}}_{n}\right)\) denotes the resultant of the forward LSTM hidden layer at time step \(\:t\), \(\:{\overleftarrow{h}}_{t}=\)\(\:\left({\overleftarrow{h}}_{1}{,\overleftarrow{h}}_{2},\:\cdots\:,\:{\overleftarrow{h}}_{n}\right)\) refers to the resultant of reverse LSTM hidden layer at time step \(\:t,\)\(\:{y}_{t}=({y}_{1},\:{y}_{2},\:\cdots\:{y}_{n})\) denotes the resultant of the Bi-LSTM network at time step \(\:t\). The last resultant vector is an integrated effect of forward or reverse data flow \(\:{y}_{t}=f\left({\overrightarrow{h}}_{t},{\overleftarrow{h}}_{t}\right)\). The mathematical formulae of the Bi-LSTM are given below.
The CNN-BiLSTM method integrates the benefits of CNNs and Bi-LSTM. A CNN has been deployed to capture the local features of input text, gradually decreasing the size and count of feature data with the sequence of convolution and pooling layers. Afterwards, the Bi-LSTM network removes global feature data in the CNN output by assuming the entire infrastructure and long‐term dependencies. The forward and backward networks of Bi-LSTM individual method the resultant features of CNN, recollecting the primary data with the memory unit and gating mechanism to acquire the last feature data. Eventually, the Bi-LSTM output is distributed to a fully connected (FC) layer. Combining the local feature extractor proficiency of the CNN with the global data processing proficiency of BiLSTM, the CNN‐BiLSTM technique effectually improves the accuracy.
Result analysis and discussion
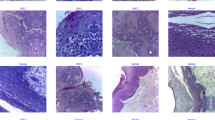
The SEHDL-OSCCR technique’s performance evaluation is examined using the OSCC dataset from the Kaggle repository30,31. The images were captured using a Leica ICC50 HD microscope from H&E-stained tissue slides prepared by medical experts from 230 patients. The dataset comprises 528 samples with two classes, as demonstrated in Table 1. Figure 6 represents the sample images. The suggested technique is simulated using the Python 3.6.5 tool on PC i5-8600k, 250GB SSD, GeForce 1050Ti 4GB, 16GB RAM, and 1 TB HDD. The parameter settings are provided: learning rate: 0.01, activation: ReLU, epoch count: 50, dropout: 0.5, and batch size: 5.
Figure 7 demonstrates the confusion matrices produced by the SEHDL-OSCCR method on different epochs. The outcomes indicate that the SEHDL-OSCCR approach effectively recognizes samples under all classes.
In Table 2; Fig. 8, the detection results of the SEHDL-OSCCR technique are depicted under several epochs. The results signified that the SEHDL-OSCCR technique correctly identified the normal and OSCC samples. With 500 epochs, the SEHDL-OSCCR technique gains an average \(\:acc{u}_{y}\) of 94.89%, \(\:pre{c}_{n}\) of 91.65%, \(\:rec{a}_{l}\) of 89.76%, \(\:F{1}_{score}\) of 90.67%, and MCC of 81.39%. Also, with 1000 epochs, the SEHDL-OSCCR method gets an average \(\:acc{u}_{y}\) of 96.97%, \(\:pre{c}_{n}\) of 93.93%, \(\:rec{a}_{l}\) of 95.49%, \(\:F{1}_{score}\) of 94.69%, and MCC of 89.41%. Concurrently, with 1500 epochs, the SEHDL-OSCCR method attains an average \(\:acc{u}_{y}\) of 97.16%, \(\:pre{c}_{n}\) of 94.42%, \(\:rec{a}_{l}\) of 95.60%, \(\:F{1}_{score}\) of 95.00%, and MCC of 90.02%. Simultaneously, with 2000 epochs, the SEHDL-OSCCR approach achieves an average \(\:acc{u}_{y}\) of 98.11%, \(\:pre{c}_{n}\) of 96.25%, \(\:rec{a}_{l}\) of 97.07%, \(\:F{1}_{score}\) of 96.65%, and MCC of 93.31%. At last, with 2500 epochs, the SEHDL-OSCCR approach gains an average \(\:acc{u}_{y}\) of 97.73%, \(\:pre{c}_{n}\) of 95.95%, \(\:rec{a}_{l}\) of 95.95%, \(\:F{1}_{score}\) of 95.95%, and MCC of 91.89%.
In Fig. 9, the training and validation accuracy results of the SEHDL-OSCCR technique are established. The accuracy values are computed in the interval of 0-3000 epochs. The result emphasizes that the training and validation accuracy values show a rising tendency, which alerts the ability of the SEHDL-OSCCR approach to improve performance over numerous iterations. Moreover, the training and validation accuracy remain closer over the epochs, which states low least overfitting and displays amended performance of the SEHDL-OSCCR approach, guaranteeing consistent prediction on hidden samples.
In Fig. 10, the training and validation loss graph of the SEHDL-OSCCR method is demonstrated. The loss values are computed for 0-3000 epochs. It is indicated that the training and validation accuracy values demonstrate a decreasing tendency, notifying the capability ability of the SEHDL-OSCCR method in balancing a tradeoff between data fitting and generalization. The continual reduction in loss values also ensures the upgraded performance of the SEHDL-OSCCR approach and tunes the prediction outcomes over time.
In Fig. 11, the precision-recall (PR) curve examination of the SEHDL-OSCCR technique under dissimilar epochs delivers an interpretation of its performance by plotting Precision against Recall for all the classes. The figure denotes that the SEHDL-OSCCR method continuously accomplishes improved PR values across diverse class labels, representing its capability to maintain a significant part of true positive predictions between every positive prediction (precision) while capturing a substantial proportion of actual positives (recall). The stable growth in PR outcomes among every class signifies the efficacy of the SEHDL-OSCCR methodology in the classification process.
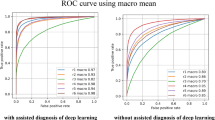
In Fig. 12, the ROC curve of the SEHDL-OSCCR model under discrete epochs is studied. The outcomes suggest that the SEHDL-OSCCR method improves ROC results over every class, representing the critical skill of discriminating the classes. This reliable trend of improved ROC values over many classes indicates the proficient performance of the SEHDL-OSCCR approach in forecasting classes, highlighting the robust nature of the identification process.
Table 3; Fig. 13 show a widespread comparison study of the SEHDL-OSCCR technique clearly illustrated32. The results indicate that the Resnet50-VGG16 model has shown ineffective performance. Additionally, three kinds of CNN and MobileNetv3-GTO models have exhibited slightly boosted results. Meanwhile, Resnet50-Feature fusion, VGG16, Mobile Inception, and Swin-Transformer models have demonstrated moderately closer results. However, the SEHDL-OSCCR technique outperforms the other models with an increased \(\:acc{u}_{y}\) of 98.75%, \(\:pre{c}_{n}\) of 96.69%, \(\:rec{a}_{l}\) of 98.75%, and \(\:F{1}_{score}\) of 97.69%.
The computation time (CT) results of the SEHDL-OSCCR technique are compared with other DL models in Table 4; Fig. 14. The results show that the SEHDL-OSCCR technique attains a minimal CT of 2.70s. On the other hand, the ResNet50-feature fusion, ResNet50-DCNN, ResNet50-VGG16, three kinds of CNN, VGG16 and Mobile Inception, MobilenetV3-GTO, and Swin Transformer models obtain increased CT values of 7.49s, 8.59s, 5.71s, 4.62s, 6.88s, 3.84s, and 4.66s, correspondingly. Thus, the SEHDL-OSCCR technique can be used to identify OC.
Conclusion
This study developed a novel SEHDL-OSCCR approach to HIs. The presented SEHDL-OSCCR technique mainly focuses on detecting OC using hybrid DL models. To accomplish that, the SEHDL-OSCCR approach contains distinct processes such as noise reduction, SE-CapsNet-based feature extractor, ICOA-based parameter tuning, and classifier selection process. Initially, the BF technique is used to remove the noise. Next, the SEHDL-OSCCR technique applies the SE-CapsNet model to recognize the feature extractors. The ICOA is utilized to boost the performance of the SE-CapsNet model. Finally, the classification of OSCC is performed using the CNN-BiLSTM model. The obtained simulation results of the SEHDL-OSCCR technique can be tested under a benchmark medical image dataset. The experimental validation of the SEHDL-OSCCR technique illustrated a greater accuracy outcome of 98.75% compared to recent approaches. The limitations of the SEHDL-OSCCR approach encompass challenges in managing diverse data discrepancies and complexities that can affect its overall performance. Moreover, the technique may need help with scalability problems as the dataset size increases, potentially impacting processing effectiveness. There may also be limitations in handling subtle variations between classes, which could affect the precision of the technique in specific scenarios. Future studies should address these limitations by expanding the dataset to include a broader range of conditions and focusing on improving the scalability and capability of the system to differentiate subtle features. Incorporating advanced techniques and innovative strategies will significantly enhance robustness, accuracy, and effectiveness. Future research should also include incorporating multi-modal data, namely genomic and clinical information, with the present histopathological images to improve the capability of the model to make more comprehensive and precise diagnoses.
Data availability
The datasets used and analyzed during the current study available from the corresponding author on reasonable request.
References
Fu, Q. et al. A deep learning algorithm for detection of oral cavity squamous cell carcinoma from photographic images: A retrospective study. EClinicalMedicine 27, 100558 (2020).
Rahman, T. Y., Mahanta, L. B., Das, A. K. & Sarma, J. D. Study of morphological and textural features for classification of OSCC by traditional machine learning techniques. Cancer Rep. 3, e1293 (2020).
Welikala, S., Rajapakse, J., Karunathilaka, S. & Abeywickrama, P. Automated detection and classification of oral lesions using deep learning for early detection of oral cancer. Sci. Rep. 10, 1–10 (2020).
Rahman, M. S., Hossain, M. A., Khan, M. S. R. & Kaiser, M. S. Histopathologic oral cancer prediction using oral squamous cell carcinoma biopsy empowered with transfer learning. Comput. Methods Prog Biomed. 22, 107143 (2022).
Ibrar, M. W., Khan, M. S. R. & Kaiser, M. S. Early diagnosis of oral squamous cell carcinoma is based on histopathological images using deep and hybrid learning approaches. Comput. Methods Prog Biomed. 252, 107372 (2023).
Haq, U., Ahmed, M., Assam, M., Ghadi, Y. Y. & Algarni, A. Unveiling the future of oral squamous cell carcinoma diagnosis: An innovative hybrid AI approach for accurate histopathological image analysis. IEEE Access. 11, 118281–118290 (2023).
Deif, M. A. et al. Diagnosis of oral squamous cell carcinoma using deep neural networks and binary particle swarm optimization on histopathological images: An AIoMT approach. Comput Intell Neurosci. 1–13 (2022).
Alanazi, A. A., Khayyat, M. M., Khayyat, M. M., Elamin Elnaim, B. M. & Abdel-Khalek, S. Intelligent deep learning enabled oral squamous cell carcinoma detection and classification using biomedical images. Comput. Intell. Neurosci. 2022, 1–11 (2022).
Rahman, T., Mahanta, L., Das, A. & Sarma, J. Histopathological imaging database for oral cancer analysis. Data Brief. 29, 105114 (2020).
Fatapour, Y., Abiri, A., Kuan, E. C. & Brody, J. P. Development of a machine learning model to predict recurrence of oral tongue squamous cell carcinoma. Cancers 15, 2769 (2023).
Zhou, J. et al. A pathology-based diagnosis and prognosis intelligent system for oral squamous cell carcinoma using semi-supervised learning. Expert Syst. Appl. 124242 (2024).
Fati, S. M., Senan, E. M. & Javed, Y. Early diagnosis of oral squamous cell carcinoma based on histopathological images using deep and hybrid learning approaches. Diagnostics 12(8), 1899 (2022).
Panigrahi, S. et al. Classifying histopathological images of oral squamous cell carcinoma using deep transfer learning. Heliyon 9(3) (2023).
Yadav, A. & Yadav, S. Enhancing oral squamous cell carcinoma detection: A transfer learning perspective on histopathological analysis using ResNet-18, AlexNet, DenseNet-169, and DenseNet-201 with cyclic learning rate. Int. J. Intell. Syst. Appl. Eng. 12 (17s), 689–699 (2024).
Begum, S. H. & Vidyullatha, P. Deep Learning Model for Automatic detection of oral squamous cell carcinoma (OSCC) using histopathological images. Int. J. Comput. Digit. Syst. (2023).
Deif, M. A. et al. Diagnosis of oral squamous cell carcinoma using deep neural networks and binary Particle Swarm optimization on histopathological images: An AIoMT approach. Comput. Intell. Neurosci. 2022(1), 6364102 (2022).
Sampath, P., Sasikaladevi, N., Vimal, S. & Kaliappan, M. OralNet: Deep learning fusion for oral cancer identification from lips and tongue images using stochastic gradient based logistic regression. Netw. Model. Anal. Health Inform. Bioinform., 13(1), p.24. (2024).
Ahmad, M. et al. Multi-method analysis of histopathological image for early diagnosis of oral squamous cell carcinoma using deep learning and hybrid techniques. Cancers 15(21), 5247 (2023).
Kadhim, D. A. & Mohammed, M. A. A comprehensive review of artificial intelligence approaches in kidney cancer medical images diagnosis, datasets, challenges and issues and future directions. Int. J. Math. Stat. Comput. Sci. 2, 199–243 (2024).
Mohammed, M. Enhanced Cancer Subclassification using Multi-omics Clustering and Quantum Cat Swarm optimization. Iraqi J. Comput. Sci. Math. 5 (3), 552–582 (2024).
Das, M., Dash, R., Mishra, S. K. & Dalai, A. K. An Ensemble deep learning model for oral squamous cell carcinoma detection using histopathological image analysis. IEEE Access (2024).
Das, M., Dash, R. & Mishra, S. K. Automatic detection of oral squamous cell carcinoma from histopathological images of oral mucosa using deep convolutional neural network. Int. J. Environ. Res. Public Health 20(3), 2131 (2023).
Meer, M. et al. Deep convolutional neural networks information fusion and improved whale optimization algorithm based smart oral squamous cell carcinoma classification framework using histopathological images. Expert Syst. pe13536. (2024).
Raj, M. P. & Muneeswari, G. Intelligent optimal archimedes shooty tern deep network (OASTDN) for oral squamous cell carcinoma detection and classification in oral cancer. Multimedia Tools Appl. 1–25 (2024).
Shukla, R., Ajwani, B., Sharma, S. & Das, D. Identifying oral carcinoma from histopathological image using unsupervised nuclear segmentation. In 2024 IEEE 9th International Conference for Convergence in Technology (I2CT) 1–6 (IEEE, 2024).
DUMAN, E. A. An Edge Preserving Image Denoising Framework based on statistical edge detection and bilateral Filter. Mehmet Akif Ersoy Üniversitesi Fen Bilimleri Enstitüsü Dergisi 12(Ek (Suppl.) 1), 519–531 (2021).
Bian, L., Zhang, L., Zhao, K., Wang, H. & Gong, S. Image-based scam detection method using an attention capsule network. IEEE Access 9, 33654–33665 (2021).
Wang, J. et al. Soil salinity inversion in yellow river delta by regularized extreme learning machine based on ICOA. Remote Sens. 16(9), 1565 (2024).
Chang, Y. & Bao, G. Enhancing rolling bearing fault diagnosis in motors using the OCSSA-VMD-CNN-BiLSTM model: A novel approach for fast and accurate identification. IEEE Access. (2024).
Rahman, T., Yesmin, Mahanta, L. B., Das, A. K. & Sarma, J. D. Histopathological imaging database for oral cancer analysis. Mendeley Data 29, 105114 (2023).
Albalawi, E. et al. Oral squamous cell carcinoma detection using EfficientNet on histopathological images. Front. Med., 10, 1349336 (2024).
Funding
This Project was funded by the Deanship of Scientific Research (DSR) at King Abdulaziz University (KAU), Jeddah, Saudi Arabia, under grant no. (GPIP: 7-611-2024). The authors, therefore, acknowledge with thanks DSR at KAU for technical and financial support.
Author information
Authors and Affiliations
Contributions
Conceptualization, Investigation and Methodology, Writing—original draft: Turky Omar Asar. Data curation, Formal analysis, Project administration, Resources, Supervision, Validation and Visualization, Writing—review and editing: Mahmoud Ragab. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval
This article contains no studies with human participants performed by any authors.
Consent to participate
Not applicable.
Institutional Review Board Statement
Not applicable.
Informed consent
Not applicable.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ragab, M., Asar, T.O. Deep transfer learning with improved crayfish optimization algorithm for oral squamous cell carcinoma cancer recognition using histopathological images. Sci Rep 14, 25348 (2024). https://doi.org/10.1038/s41598-024-75330-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-75330-3