Abstract
Early and precise diagnosis of craniosynostosis (CSO), which involves premature fusion of cranial sutures in infants, is crucial for effective treatment. Although computed topography offers detailed imaging, its high radiation poses risks, especially to children. Therefore, we propose a deep-learning model for CSO and suture-line classification using 2D cranial X-rays that minimises radiation-exposure risks and offers reliable diagnoses. We used data comprising 1,047 normal and 277 CSO cases from 2006 to 2023. Our approach integrates X-ray-marker removal, head-pose standardisation, skull-cropping, and fine-tuning modules for CSO and suture-line classification using convolution neural networks (CNNs). It enhances the diagnostic accuracy and efficiency of identifying CSO from X-ray images, offering a promising alternative to traditional methods. Four CNN backbones exhibited robust performance, with F1-scores exceeding 0.96 and sensitivity and specificity exceeding 0.9, proving the potential for clinical applications. Additionally, preprocessing strategies further enhanced the accuracy, demonstrating the highest F1-scores, precision, and specificity. A qualitative analysis using gradient-weighted class activation mapping illustrated the focal points of the models. Furthermore, the suture-line classification model distinguishes five suture lines with an accuracy of > 0.9. Thus, the proposed approach can significantly reduce the time and labour required for CSO diagnosis, streamlining its management in clinical settings.
Similar content being viewed by others
Introduction
Craniosynostosis (CSO) is a complex medical condition characterised by premature fusion of one or more cranial sutures in infants. Sutures are fibrous bands that connect the bones in the skull, allowing them to grow and expand to accommodate the developing brain. Early closure of these sutures can restrict skull and brain growth, leading to potential developmental issues and abnormal head shapes. Additionally, it is challenging to keep children stationary for prolonged periods to conduct detailed measurements or 3D computed photography (CT) scans. Moreover, there are concerns regarding exposing young children to high levels of radiation associated with such procedures, which could potentially lead to adverse outcomes further on in life. In contrast, standard skull X-rays, which require considerably less time and involve lower levels of radiation, offer a more practical and safer alternative for initial evaluations.
Various automatic CSO-classification methods have been developed for use with medical-imaging modalities. CT is the gold standard for diagnostic imaging and surgical planning and is routinely performed in many craniofacial centres worldwide. You et al.1. proposed an automated sagittal CSO-classification algorithm using features extracted from projected 3D skull images, which included a 3D multi-view learning algorithm, 3D rendering, patch-based erosion for data augmentation, transfer-learning strategy for training, and multi-view-based prediction strategy for classification. Liu et al.2 presented a novel end-to-end neural network architecture based on DenseNet3 with context regularisation to jointly label cranial bone plates and detect cranial base landmarks from CT images.
However, despite the accurate and precise diagnosis offered by CT scans4, prioritising 2D X-ray or radiation-free methods over CT is recommended because of the associated risks. The larger doses of radiation involved in CT scans significantly increase radiation-exposure risks in children compared with adults5,6. Furthermore, studies have shown that exposing infants to even low doses of ionising radiation can adversely influence their cognitive abilities in adulthood, underscoring the need for cautious diagnostic imaging for young patients7,8. To address the issue of radiation exposure, a statistical shape model (SSM) for radiation-free assessment and classification of CSO has been proposed9; this model successfully distinguished between three CSO classes (coronal, metopic, or sagittal) and a control group using photogrammetric surface scans. The authors of9 further explored the use of photogrammetric 3D surface scans by converting 3D scans into 2D distance maps, enabling the application of convolutional neural networks (CNNs) for CSO detection for the first time10. This method emphasised the reduction of radiation exposure, while preserving patient anonymity and enabling effective data augmentation, thereby achieving robust CSO detection using minimal 3D data. In the same context, de Jong et al.11 proposed a deep-learning algorithm for identifying CSO from 3D stereographs as a non-invasive technique that exhibited accurate performance for classifying the head shapes of children as either normal or CSO of one of the three subtypes (scaphocephaly, trigonocephaly, or anterior plagiocephaly). Subsequently, Schaufelberger et al.12 demonstrated the diagnostic value of 3D photogrammetry for assessing and diagnosing head deformities such as CSO, and Mashouri et al.13 developed machine learning image classifiers that enhance the triaging process for CSO referrals using 3D images for early identification of synostosis in children and qualifying them for less invasive surgical intervention.
However, a simpler and more accessible approach than statistical models involves the use of 2D images. To evaluate cranial deformities in infants, photographs of their heads have been used to calculate a craniometric index score14,15,16. Although 3D CT provides accurate scans, expecting young children to remain still for extended durations required for CT imaging is challenging. Moreover, exposing them to significant amounts of radiation can increase their risk of developing cancer during adolescence7. Therefore, methods that employ simple photographs that do not involve radiation exposure have been proposed as low-risk alternatives17,18,19. However, these methods struggle to obtain accurate and detailed skull information and determine the exact shape of the CSO, often necessitating subsequent X-ray imaging, rendering them inadequate for effective CSO diagnosis.
Despite the detailed insights offered by 3D imaging techniques11, their practical applications are limited owing to high costs and the specialised training required for their operation. Moreover, although SSMs provide statistically significant results, they are unsuitable for direct use in actual medical settings. Additionally, 2D photographs are insufficient for CSO interpretation and therefore, additional X-rays are required. To address these limitations, this study focused on 2D X-ray imaging for CSO classification. X-rays involve significantly lower radiation exposure compared with CT scans and are more accessible, which is crucial for frequent diagnostic assessments. Digital X-rays exhibit higher diagnostic accuracy without excessive radiation associated with traditional methods. Early detection of CSO is crucial for effective treatment planning, which may include surgical interventions to correct the skull shape and prevent further neurological damage. Moreover, accurate diagnosis and rapid categorisation of CSO are essential for optimising patient outcomes. Therefore, novel detection methods and classification criteria are vital areas of research for paediatric neurosurgeries and craniofacial surgeries. This study proposes a novel deep-learning model that aims to balance diagnostic efficacy with patient safety, setting a new benchmark in paediatric diagnostic imaging and advancing craniofacial medical practices. The proposed model can detect CSO and suture lines, and is specifically designed for application to children under the age of 2 y. Central to our approach is a preprocessing procedure for raw medical images that significantly enhances the model performance. Extensive ablation studies were conducted to ensure the robustness and reliability of the proposed model. The results showed that it can precisely identify and categorise CSO and can potentially serve as a new benchmark in paediatric diagnostic imaging. Thus, by facilitating earlier and more accurate diagnoses, our study makes a substantial contribution to the advancement of craniofacial surgery.
Materials and methods
Ethical approval
The study protocol adhered to the ethical guidelines set by the 1975 Declaration of Helsinki and revised in 1983, and was approved by the Institutional Review Board (IRB) of Ajou University Hospital (IRB number: AJOUIRB-DB2024-204). As this study used data previously collected at the Ajou University Medical Center, the requirement for informed consent was waived by the IRB of Ajou University Hospital (IRB number: AJOUIRB-DB2024-204). Following data acquisition, an anonymisation process was applied to remove any personal identifiable information and ensure patient confidentiality.
Data preprocessing
We developed fully automatic CSO and suture-line classification methods that employ anteroposterior (AP) and lateral-view images. This section outlines the preprocessing methods employed for effectively training our models on a relatively small amount of clinical data. As shown in Fig. 1A, the scale, orientation, and intensity of the input images were standardised, significantly reducing the difficulty for generalisation tasks.
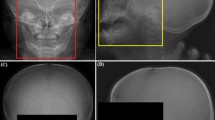
Illustration of the proposed preprocessing steps. (A) The first row contains AP-view X-rays whereas the second contains lateral-view X-rays. AP-view X-ray preprocessing: (a) input, (b) marker-removed, (c) cropped (scale standardisation), and (d) histogram-equalised (intensity standardisation ) images. Lateral-view X-ray preprocessing: (e) input, (f) marker-removed (the green line shows the detected connecting landmarks for visualisation), (g) cropped (scale and orientation standardisation), and (h) histogram-equalised (intensity standardisation) images. (B) Examples of (i) X-ray markers and (j) their removal process. (C) Head-pose standardisation and cropping.
Overview
For each child, AP- and lateral-view images, with their gaze directed towards the left, were acquired in a 12-bit DICOM format. These images were converted into 8-bit files to fully exploit the pretrained models, yielding a dark background and light cranial structures20. Subsequently, two preprocessing methods were employed, as illustrated in Fig. 1A.
To automate the preprocessing, we fine-tuned the YOLOv3 detector21 on the skull regions and markers. Additionally, we used a landmark detector trained from scratch for pose standardisation. These automated preprocessing steps were robust, and all data were processed without critical failures.
Marker removal
Artefacts in images (e.g., “R” or “L” indicating the direction, as shown in Fig. 1B), especially those located near the regions of interest and/or having sharp contrasts, introduce undesirable effects in classification pipelines22. These effects can be mitigated through training on large datasets; however, it is desirable to remove these artefacts during preprocessing owing to the scarcity of medical images. To this end, we detected X-ray markers using the fine-tuned YOLOv3 and removed them using an inpainting algorithm23. As shown in Fig. 1B, the algorithm fills in the missing parts by propagating information from neighbouring pixels. Moreover, Gaussian blur with a kernel size of 7 × 7 was applied to the regions to further reduce boundary artefacts24. The entire inpainting process is illustrated in Fig. 1B.
Standardisation
Although the AP-view images contained regular poses, the lateral-view images showed strong inter-variation. The head pose, measured as the angle formed by the lines connecting the supraorbital ridge and central incisor, varied from 45–115°. These fluctuations can undermine the consistency of the analysis and impede accurate feature extraction and localisation. To address this issue, we employed a landmark detector, whose architecture is illustrated in Fig. 1C. It can predict the positions of the supraorbital ridge and the central incisor from an input image. Based on these detected points, we compensated for the rotation from the standard position, as shown in Fig. 1C (the standard angle was set to 81°).
We also detected skull regions using the fine-tuned YOLOv3 detector. The images were cropped to minimise changes in input scales and remove backgrounds, which improved the efficiency of the training process and generalisation performance of the model. Finally, we applied histogram equalisation to normalise the distribution of the intensity ranges25.
Training
The YOLOv3 model was fine-tuned on a dataset comprising 250 normal and 250 CSO cases to detect “Skull” and “X-ray marker” from cranial X-ray images. This model employs an input resolution of 416 × 416, a batch size of 8, and a learning rate of 1e-3. The model was optimised through classification, localisation, and confidence losses21, and exhibited a mean intersection over union (mIOU) value of 0.866 and mean average precision (mAP) of 0.991.
Additionally, the landmark detector shown in Fig. 1C, which was designed to yield the coordinates of the supraorbital ridge and central incisor from lateral-view images, was trained on 250 normal and 250 CSO cases. Model training was optimised using mean squared error (MSE) loss, with an input resolution of 400 × 500, batch size of 4, and learning rate of 1e-3. It exhibited an average root mean square error (RMSE) of 6.13.
CSO and suture-line classification
The overall process of the proposed CSO-classification method is shown in Fig. 2A. First, the preprocessed images are fed into the CSO classifier and if CSO is detected, five suture-line classifiers are applied to determine the presence of CSO on each suture line.
CSO classification
Deep-learning models, such as CNNs, can perform effective image identification26,27,28; however, they require a considerable amount of training data, which are often lacking in most medical contexts. Thus, we adopted a transfer-learning strategy to fine-tune the pretrained backbones.
The architecture of the proposed network comprises three blocks: (a) deep feature extraction, (b) feature fusion, and (c) fully connected layers. As shown in Fig. 2B, deep features from are extracted from two images (AP- and lateral-view) and concatenated. Subsequently, L2 normalisation is applied to each feature map to ensure that the results are not heavily biased towards a single side. Similar to Chandola et al29., we adopted a decision-level feature fusion approach wherein features are fused through concatenation30; this approach offers several advantages over other fusion methods, such as averaging and pooling. Fusion by concatenation allows expanding the feature space to ensure that the integrated features can easily preserve the original information. Finally, these fused features are fed into fully connected layers comprising two hidden layers with 128 and 32 neurones.
The proposed method was evaluated using six backbones: AlexNet31, VGG1932, ResNet18, ResNet5033, DenseNet-1213, and EfficientNet-B034.
Suture-line classification for CSO
We classified the suture lines into five types: sagittal, left-coronal, right-coronal, left-lambdoid, and right-lambdoid35,36. Because these labels are non-exclusive, it was natural to formulate this problem as multi-label classification (using sigmoid instead of softmax). However, multi-label classification using a single backbone yields poor performance probably due to the varying number of samples in each class and lack of data for some combinations. We addressed this problem by developing five independent detectors using the one-versus-all approach37. The overall methodology is illustrated in Fig. 2A. Additionally, each detector had the same architecture as the CSO detector, as shown in Fig. 2B.
Training details
This study employed six classification models: one for CSO classification and five for suture-line classification. All models adopted a batch size of 8 and learning rate of 1e-4. Model training was optimised through the cross-entropy loss function using the Adam optimiser38. The input images were resized to 224 × 224 pixels.
Experiments
This section presents the details of the datasets and training methods employed in this study. Additionally, the effectiveness of the proposed approach was evaluated using various metrics: accuracy (Acc), precision (Pre), sensitivity (Sen), specificity (Spec), and F1 Score (F1). The experiments were conducted using the PyTorch framework on a setup comprising an Intel(R) Core(TM) i5-13400 CPU running at 2.50 GHz, Nvidia GeForce RTX 3090 graphics card, and 32 GB of RAM.
Dataset
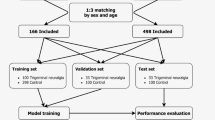
The CSO diagnoses in the data employed were validated by paediatric surgeons with more than 15 years of experience. The normal dataset comprised cases confirmed through medical examinations as having no skull-related symptoms or disorders. This study included 1,047 cases classified as normal and 277 diagnosed with CSO between 2006 and 2023. Each patient underwent AP- and lateral-view X-ray imaging. Analyses using the Mann–Whitney U test39for age distribution exhibited a p-value of 0.131, indicating no significant difference, and that using the chi-squared test40 for sex distribution yielded a p-value of 0.111. These results confirmed that the dataset was structured, thereby ensuring that these variables did not affect the classification results. A summary of the dataset is presented in Table 1.
Training, validation, and test split
As illustrated in Fig. 2A, we used six detectors with the same architecture. However, the training, validation, and test sets for each detector required adjustments according to their specific roles. For the CSO detector, a deep neural network was trained to differentiate C from N (see the notations in the bottom row of Table 1). For the sagittal detector, we used S and C ∩ Sc as we assumed that its input comprised least one CSO case. The sets for the other cases were defined in a similar manner.
Results and discussion
CSO classification
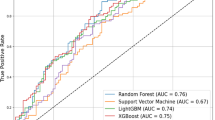
The CSO-classification performance was evaluated using six CNN backbones: EfficientNet-B0, DenseNet-121, ResNet50, ResNet18, VGG19, and AlexNet; the results are summarised in Table 2 and their confusion matrices are presented in Fig. 3A. The four CNN backbones with the best performances exhibited similar F-1 scores. These results can be attributed to two factors: (a) the number of parameters, which are important for transfer learning, especially if the amount of training data is limited (see the #Params column in Table 2) and (b) architectural optimisations, such as residual connections, are beneficial for X-ray images. As shown in Fig. 3B, all CNN backbones obtained area under the receiver operating characteristic curve (AUROC) values of > 0.93, suggesting potential for early monitoring and diagnosis of CSO.
Ablation study
Additionally, we conducted an ablation study to evaluate the effects of each preprocessing step. Only the results for ResNet18 are presented because the other backbones exhibited similar patterns. The results presented in Table 3indicate that each preprocessing step improved the performance. Similar to previous studies, we found that image standardisation is crucial. Additionally, they confirm that histogram equalisation is an essential component for X-ray image analysis41,42.
Suture-line classification
The suture-line classification performances of the six CNN backbones are summarised in Suppl. Table S1 and Suppl. Figure S3, which support our previous discussion on CNN backbones. The F1-scores in Suppl. Table S1 indicate that the classification performances for the sagittal and right-coronal suture lines were higher than those for the other lines. This can be attributed to the presence of more samples for these lines, as listed in Table 1. Despite architectural advances in deep learning, the quality and quantity of data remain crucial for model performance. Finally, the classification performances of the ResNet18 backbone for each suture-line detector are listed in Suppl. Table S2.
The results of gradient-weighted class activation mapping (Grad-CAM)43 are shown in Suppl. Figure S1. All Grad-CAM results are averages of the test set results. The CAM was overlaid on each suture line. The two coronal detectors tended to focus on the frontal parts, whereas the two lambdoid detectors focused on the head (Suppl. Figure S1). However, each detector did not simply focus on a single suture line but rather on the overall shape of the head.
Experiments on the number of training samples
Figure 4A illustrates a typical pattern of training curves: as the number of training samples increases, training error rise while validation error decrease. It is crucial to have enough data to reduce the gap between these errors to an acceptable range, ensuring good generalization. In our task, we determined that a total of 440 positive samples and 50 negative samples are required, as shown in Fig. 4B. Specifically, more than 50 samples per class are necessary. We attribute the relatively small number of required samples to the simple network structure, preprocessing, and pretraining.
Experiments incorporating Towne view images
In practical clinical applications, Towne-view images, such as those shown in Suppl. Figure S2, aid in diagnosing CSO because they provide a clear view of the posterior and basal structures of the skull44. We added an additional CNN backbone to the architecture shown in Fig. 2B (three inputs in parallel) to exploit Towne-view images. They were preprocessed in the same manner as that employed for the AP-view images shown in Fig. 1A, and the results are shown in Suppl. Figure S2. The models were trained using the same protocol described above. Since the classification accuracy for CSO is nearly saturated with two views, further improvements are marginal—the accuracy increases by only 0.01. However, considering that the number of normal samples decreases from 628 to 428, and the CSO samples from 165 to 93 in the three-view experiments, we can conclude that incorporating Towne view images effectively enhances CSO classification accuracy.
We also incorporated Towne view images into the suture line classification, with the results presented in Table 4. In this experiment, we used the same number of training samples for both the three-view and two-view methods. This approach allows us to quantify the contribution of Towne view images while maintaining a constant number of training samples. Specifically, we used 44, 17, 23, 13, and 10 positive samples for each respective suture line classifier.
When compared to Suppl. Table S2, performance seems to decrease with the addition of Towne view images (three views). However, when comparing results using the same number of samples (within Table 4), the addition of Towne views results in a slight performance improvement. Based on these experiments, we cautiously anticipate that performance will likely improve when the number of samples for each suture line exceeds 50, with the inclusion of Towne views further enhancing the results.
However, our conclusions should be interpreted with caution. Our findings indicate that a minimum of 50 images per class is needed to adequately capture the variability in the data. Performance tends to deteriorate below this threshold. To confirm the generalizability of the proposed method, a significant number of additional clinical tests will be required.
Conclusions
This study introduced a deep-learning model designed for CSO and suture-line classification using 2D cranial X-ray imaging. Although the experiments involving Towne-view images showed suboptimal results, we believe that the poor performance was due to the limited number of samples. By incorporating various views, including sagittal, anteroposterior, and Towne, the proposed model is expected to achieve more robust performance. This highlights the potential of artificial intelligence-assisted methodologies for enhancing the diagnostic accuracy and efficiency for CSO. Owing to the demonstrated efficacy of our CNN model for processing 2D skull X-ray images for CSO classification, it is reasonable to anticipate that it can potentially enhance access to CSO treatments. Deployment of this model as a screening tool in clinical settings is expected to increase early treatment rates by enabling faster and more accurate diagnoses. Moreover, this approach offers the potential to incidentally detect CSO during X-ray assessments of unrelated cranial anomalies, thereby broadening the scope of classification and ensuring that early interventions can be performed for a higher number of affected infants. Such advancements in diagnostic technology signify a pivotal step in the comprehensive management of CSO, promising not only to improve patient outcomes, but also to facilitate more efficient allocation of healthcare resources. Our analysis of the CNN backbones suggests that skull X-ray-based diagnostic models can be potentially extended to various diseases.
Despite these advancements, it is imperative to acknowledge that the gold standard for CSO diagnosis is CT modelling because of its unparalleled precision for detailing cranial structures. Therefore, in a future work, we will explore the feasibility of 2D to 3D CT reconstruction using skull X-ray data to bridge the gap between the practical applications of X-ray imaging and the diagnostic superiority of CT models45,46. In conclusion, this study underscores the impact of integrating artificial intelligence with medical imaging, while further evidence is required to confirm that this is sufficient for clinical diagnosis. These results highlight the possibility but should not be taken as definitive proof, and additional methods may need to be explored to enhance early diagnosis. Our findings not only reinforce the potential of artificial intelligence in medical diagnostics but also open avenues for further exploration of 3D imaging and reconstruction techniques, promising considerable advancements in the screening procedures of cranial disorders.
Data availability
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
References
You, L. et al. A novel sagittal craniosynostosis classification system based on multi-view learning algorithm. Neural Comput. Appl. 34, 14427–14434 (2022).
Liu, J. et al. Joint cranial bone labeling and landmark detection in pediatric CT images using context encoding. IEEE Trans. Med. Imaging. 42, 3117–3126 (2023).
Huang, G., Liu, Z., Van Der Maaten, L. & Weinberger, K. Q. Densely connected convolutional networks. IEEE Conf. Comput. Vis. Pattern Recognit. CVPR. 2017, 2261–2269 (2017).
Fearon, J. A., Singh, D. J., Beals, S. P. & Yu, J. C. The diagnosis and treatment of single-sutural synostoses: are computed tomographic scans necessary? Plast. Reconstr. Surg. 120, 1327–1331 (2007).
Huang, W. Y. et al. Paediatric head CT scan and subsequent risk of malignancy and benign brain tumour: a nation-wide population-based cohort study. Br. J. Cancer. 110, 2354–2360 (2014).
Meulepas, J. M. et al. Radiation exposure from pediatric CT scans and subsequent cancer risk in the Netherlands. J. Natl. Cancer Inst. 111, 256–263 (2019).
Brenner, D. J., Elliston, C. D., Hall, E. J. & Berdon, W. E. Estimated risks of radiation-induced fatal cancer from pediatric CT. AJR Am. J. Roentgenol. 176, 289–296 (2001).
Hall, P. et al. Effect of low doses of ionising radiation in infancy on cognitive function in adulthood: Swedish population based cohort study. BMJ. 328, 19 (2004).
Schaufelberger, M. et al. A Statistical Shape Model for Radiation-Free Assessment and Classification of Craniosynostosis (2022).
Schaufelberger, M. et al. 3D-2D Distance maps conversion enhances classification of craniosynostosis. IEEE Trans. Biomed. Eng. 70, 3156–3165 (2023).
de Jong, G. et al. Combining deep learning with 3D stereophotogrammetry for craniosynostosis diagnosis. Sci. Rep. 10, 15346 (2020).
Schaufelberger, M. et al. CNN-based classification of craniosynostosis using 2D distance maps. Annu. Int. Conf. IEEE Eng. Med. Biol. Soc. 44th Annu. Int. Conf. IEEE Eng. Med. Biol. Soc. EMBC 2022, 446–449 (2022).
Mashouri, P. et al. 3D photography based neural network craniosynostosis triaging system in Proc. Mach. Learn. Health NeurIPS Workshop, PMLR 226–237. (2020).
Anderson, M. G. et al. Incorporation of a biparietal narrowing metric to improve the ability of machine learning models to detect sagittal craniosynostosis with 2D photographs. Neurosurg. Focus. 54, E9 (2023).
Sabeti, M., Boostani, R., Moradi, E. & Shakoor, M. H. Machine learning-based identification of craniosynostosis in newborns. Mach. Learn. Appl. 8, 100292 (2022).
Callejas Pastor, C. A. et al. Two-dimensional image-based screening tool for infants with positional cranial deformities: a machine learning approach. Diagnostics (Basel). 10, 495 (2020).
Sabeti, M., Boostani, R., Taheri, B. & Moradi, E. Image processing and machine learning for diagnosis and screening of craniosynostosis in children. Interdiscip Neurosurg. 36, 101887 (2024).
Sabeti, M., Boostani, R., Shakoor, M. H., Moradi, E. & Mohammadi, H. R. An efficient image segmentation scheme for determination of cranial index in scaphocephalic patients. Intell. -Based Med. 6, 100074 (2022).
Bookland, M. J., Ahn, E. S., Stoltz, P. & Martin, J. E. Image processing and machine learning for telehealth craniosynostosis screening in newborns. J. Neurosurg. Pediatr. 27, 581–588 (2021).
Hržić, F. et al. Estimation of missing parameters for DICOM to 8-bit x-ray image export Int. Conf. Electr. Comput. Commun. Mechatron. Eng. ICECCME, 2022 1–6. (2022).
Redmon, J. & Farhadi, A. YOLOv3: an Incremental Improvement, (2018).
Kim, K. D. et al. Enhancing deep learning based classifiers with inpainting anatomical side markers (L/R markers) for multi-center trials. Comput. Methods Programs Biomed. 220, 106705 (2022).
Telea, A. An image inpainting technique based on the fast marching method. J. Graph Tool. 9, 23–34 (2004).
Gedraite, E. S. & Hadad, M. Investigation on the effect of a gaussian blur in image filtering and segmentation in Proc. ELMAR-2011 393–396. (2011).
Pizer, S. M. et al. Adaptive histogram equalization and its variations. Comput. Vis. Graph Image Process. 39, 355–368 (1987).
Hassantabar, S., Ahmadi, M. & Sharifi, A. Diagnosis and detection of infected tissue of COVID-19 patients based on lung x-ray image using convolutional neural network approaches. Chaos Solitons Fract. 140, 110170 (2020).
Akter, S., Shamrat, F. M. J. M., Chakraborty, S., Karim, A. & Azam COVID-19 detection using deep learning algorithm on chest x-ray images. Biology. 10, 1174 (2021).
Avola, D. et al. Study on transfer learning capabilities for pneumonia classification in chest-x-rays images. Comput. Methods Programs Biomed. 221, 106833 (2022).
Chandola, Y., Virmani, J., Bhadauria, H. S. & Kumar, P. Chapter 10. Comparative analysis of computer-aided classification systems designed for chest radiographs: conclusion and future scope in Deep Learn. Chest Radiogr (ed. Chandola, Y., Virmani, J., Bhadauria, H. S. & Kumar, P.) 205–209 (Academic, 2021).
Haghighat, M., Abdel-Mottaleb, M. & Alhalabi, W. Discriminant correlation analysis: real-time feature level fusion for multimodal biometric recognition. IEEE Trans. Inf. Forensics Secur. 11, 1984–1996 (2016).
Krizhevsky, A., Sutskever, I. & Hinton, G. E. ImageNet classification with deep convolutional neural networks. Adv. Neural Inf. Process. Syst. 25 (Curran Associates, Inc., 2012).
Simonyan, K. & Zisserman, A. (eds), San Diego CA USA. Very deep convolutional networks for large-scale image recognition in Conf. Track Proc. 3rd Int. Conf. Learn. Represent. ICLR, 2015 (ed. Bengio, Y. & LeCun, Y.). (2015).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition IEEE Conf. Comput. Vis. Pattern Recognit. CVPR, 2016 770–778. (2016).
Tan, M. & Le, Q. EfficientNet: rethinking model scaling for convolutional neural networks in Proc. 36th Int. Conf. Mach. Learn., PMLR 6105–6114. (2019).
Chumas, P. D., Cinalli, G., Arnaud, E., Marchac, D. & Renier, D. Classification of previously unclassified cases of craniosynostosis. J. Neurosurg. 86, 177–181 (1997).
Trench, L., Bracken, S., McInerney, N., Caird, J. & Murray, D. J. OC7 nonsyndromic craniosynostosis: referral patterns for metopic, coronal and lambdoid craniosynostosis in Ireland; 2009–2018, a 10-year review. Arch. Dis. Child. 104, (2019). A3.2–A3A3.
Read, J., Pfahringer, B., Holmes, G. & Frank, E. Classifier chains for multi-label classification. Mach. Learn. 85, 333–359 (2011).
Kingma, D. P. San Diego CA USA. Ba J. Adam: a method for stochastic optimization in Conf. Track Proc., San Diego, CA, USA. 3rd Int. Conf. Learn. Represent. ICLR, 2015 (ed. Bengio, Y. & LeCun, Y.). (2015).
Mann, H. B. & Whitney, D. R. On a test of whether one of two random variables is stochastically larger than the other. Ann. Math. Statist. 18, 50–60 (1947).
Pearson, K. X. On the criterion that a given system of deviations from the probable in the case of a correlated system of variables is such that it can be reasonably supposed to have arisen from random sampling. Lond. Edinb. Dublin Philos. Mag J. Sci. 50, 157–175 (1900).
Salem, N., Malik, H. & Shams, A. Medical image enhancement based on histogram algorithms. Procedia Comput. Sci. 163, 300–311 (2019).
Qiu, J., Li, H., Zhang, H., Ma, T., Yang, D. & F. & Automatic x-ray image contrast enhancement based on parameter auto-optimization. J. Appl. Clin. Med. Phys. 18, 218–223 (2017).
Selvaraju, R. R. et al. Grad-CAM: visual explanations from deep networks via gradient-based localization IEEE Int. Conf. Comput. Vis. ICCV, 2017 618–626. (2017).
Mathijssen, I. M. J. Guideline for care of patients with the diagnoses of craniosynostosis: working group on craniosynostosis. J. Craniofac. Surg. 26, 1735–1807 (2015).
Kyung, D., Jo, K., Choo, J., Lee, J. & Choi, E. Perspective projection-based 3d CT reconstruction from biplanar x-rays. ICASSP IEEE Int. Conf. Acoust. Speech Signal. Process. ICASSP, 2023–20231. (2023).
Ying, X. et al. X2CT-GAN: reconstructing CT from biplanar x-rays with generative adversarial networks IEEECVF Conf. Comput. Vis. Pattern Recognit. CVPR, 2019 10611–10620. (2019).
Acknowledgements
This research was supported by a grant from the Korea Health Technology R&D Project through the Korea Health Industry Development Institute (KHIDI) funded by the Ministry of Health & Welfare, Republic of Korea (Grant no. HR22C1734) and a National Research Foundation of Korea (NRF) grant funded by the Korean Government (MSIT) (Grant no. RS-2023-00253964). This research was supported in part by the Ministry of Science and ICT (MSIT), Korea, under the Information Technology Research Center (ITRC) support program (IITP-2024-2020-0-01461) supervised by the Institute for Information & Communications Technology Planning & Evaluation (IITP).
Author information
Authors and Affiliations
Contributions
S.M.K.: Conceptualisation, Software, Visualisation, Writing – Original draft, Writing – Reviewing and editing. J.S.Y.: Investigation, Conceptualisation, Data curation, Methodology, Software, Writing – Original draft, Writing - Reviewing and editing. J.W.H.: Formal analysis, Methodology, Software, Writing – Original draft. H.I.K.: Methodology, Writing – Reviewing and editing, Conceptualization, Supervision, Project administration, Funding acquisition. T.H.R.: Methodology, Validation, Supervision, Conceptualisation, Supervision, Project administration, Funding acquisition. S.H.Y.: Resources - Patients, Conceptualisation, Data curation.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Kim, S.M., Yang, J.S., Han, J.W. et al. Convolutional neural network-based classification of craniosynostosis and suture lines from multi-view cranial X-rays. Sci Rep 14, 26729 (2024). https://doi.org/10.1038/s41598-024-77550-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-77550-z
Keywords
This article is cited by
-
Towards Automated Craniosynostosis Diagnosis Using EfficientNet-Based Artificial Intelligence Models: A Two-Class and Multi-Class Approach
Journal of Imaging Informatics in Medicine (2025)