Abstract
This study evaluates long-read and short-read sequencing for mitochondrial DNA (mtDNA) heteroplasmy detection. 592,315 bootstrapped datasets generated from two single-nucleotide mismatched ultra-deep sequenced mtDNA samples were used to assess basecalling error and accuracy, limit of heteroplasmy detection, and heteroplasmy detection across various coverage depths. Results showed high Phred scores of data with GC-rich sequence bias for long reads. Limit of detection of 12% heteroplasmy was identified, showing strong correlation (R2 ≥ 0.955) with expected heteroplasmy but underreporting tendency of high-level variants. Nanopore sequencing shows potential for direct applicability in mitochondrial diseases diagnostics, but stringent validation processes to ensure diagnostic result quality are required.
Similar content being viewed by others
Introduction
Mitochondria are essential organelles in most eukaryotic cells, crucial for energy production1. They rely on approximately 1500 proteins from nuclear and mitochondrial genomes. The circular 16,569 bp mitochondrial genome encode 13 mRNAs, 22 tRNAs, and 2 rRNAs and exists in multiple copies in the mitochondrial matrix2. Mitochondrial DNA (mtDNA) replication is cell cycle-independent and maternally inherited in most animal species. Heteroplasmy, the presence of multiple mtDNA types within an individual, is common, with over 90% of the general population exhibiting low-level variants3,4,5.
Mitochondrial diseases are genetic disorders characterized by a shortage of cellular energy due to mitochondrial dysfunction. The organs that rely heavily on aerobic metabolism are often severely impacted6,7. Diagnosing these diseases is complex due to genetic heterogeneity, heteroplasmy and the presence of nuclear-mitochondrial segments (numts). Clinical symptoms of mitochondrial disease typically manifest when the heteroplasmy exceeds 60–90%, with the exact percentage depending on the mtDNA pathogenic variant and the affected tissue8,9. However, detecting low-level variants is essential for accurate diagnosis and genetic counselling of female carriers10.
Next-generation sequencing (NGS) technologies have revolutionized diagnostics by offering deeper coverage and simultaneous sequencing of multiple samples and concurrent analysis of mtDNA and nuclear genes related to mitochondrial disorders9,11. Ultra-deep mtDNA sequencing has significantly enhanced the heteroplasmy detection at low levels (0.5%)12 compared to the Sanger sequencing13. Long-read sequencing, such as Oxford Nanopore Technologies (ONT), allow for the detection of structural alterations in the mitochondrial and nuclear genomes. Studies have shown the power of long reads in mapping sporadic mtDNA deletions without mtDNA amplification and fragmentation14,15. Moreover, ONT facilitates the sequencing of native DNA, enabling the identification of epigenetic modifications of mtDNA16.
We evaluated heteroplasmy detection using PromethION data, focusing on data quality and k-mers impact. Long-read sequencing of two single-nucleotide mismatched mtDNA were generated by ONT, followed by bootstrapping and heteroplasmy simulation, generating more than half a million genomic datasets to evaluate the sensitivity of heteroplasmy detection and compared to short-read data. Additionally, we assessed low-level heteroplasmy detection across coverage depths. This study demonstrates the stepwise validation process of a molecular diagnostic test aimed at detecting low-level variants in mtDNA.
Methods
DNA samples and ultra-deep sequencing of mtDNA
DNA was extracted from whole blood samples of two individuals who shared the same mtDNA haplogroup but differed by a single-nucleotide variant in their mtDNA. One individual, diagnosed with Leigh syndrome, harbored a homoplasmic genetic variant at m.8993T > G, while the other, a healthy control, carried the homoplasmic wild-type T at the same position. DNA extraction was performed using the FlexiGene DNA Kit (QIAGEN, Hilden, Germany).
Following extraction, DNA concentration was quantified using the Qubit® dsDNA HS Assay Kit (Thermo Fisher Scientific, Waltham, Massachusetts, United States). The DS-11 Series Spectrophotometer Fluorometer (DeNovix Inc., Wilmington, DE, United States) was then used to assess the concentration, 260/280 and 260/230 absorbance ratios, confirming that the samples were free from protein, phenol, or other contaminants.
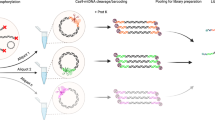
For library preparation, targets for Cas9 enrichment were designed using CHOPCHOP (Table S1)17. The sgRNA was synthesized using the EnGen sgRNA Synthesis Kit (S. pyogenes) (New England Biolabs, Massachusetts, USA) and purified with the Monarch® RNA Cleanup Kit (New England Biolabs, Massachusetts, USA). DNA libraries were prepared according to the SQK-CS9109 protocol, using 2.5 µg of DNA input for each sample. During library preparation, genomic DNA was dephosphorylated, followed by Cas9-mediated cleavage of the target DNA (mtDNA). This process leaves a 5’-phosphate group on the cleaved DNA, enabling the ligation of adapters specifically to the targeted regions and therefore enrichment. Sequencing was then performed on the PromethION platform, using an R9 flow cell (Oxford Nanopore Technology, Oxford, UK). The resulting data were basecalled and converted to FASTQ format using Guppy (version 6.3.9) in high-accuracy mode.
NGS libraries enriched with mitochondrial probes (xGen™ Human mtDNA Hyb Panel) (IDT, Iowa, USA) were prepared following the FC-141-1006 protocol, using 50 ng of DNA input for each sample. Briefly, genomic DNA was tagmented, followed by amplification and hybridization with the probes. The hybridized probes were then captured, and the enriched library was further amplified. The prepared library was subsequently sequenced on the MiSeq platform (Illumina, California, USA). For this study, informed consent was obtained from both subjects and/or their legal guardians. The study was ethically approved by the National Medical Ethics Committee of the Republic of Slovenia (Approval No. 132/03/15). All methods were performed in accordance with the relevant guidelines and regulations.
The data from the two sequenced samples formed the foundation for three datasets bundle, resulting in a total of 592,315 FASTQ files. These were used to evaluate basecalling accuracy, heteroplasmy detection, variant caller performance, and heteroplasmy detection across various coverage depths.
First dataset bundle generation and assessment of basecalling accuracy of long-read data
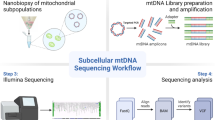
The first dataset bundle used for assessing basecalling accuracy was prepared only from long-read sequencing data of a healthy control with a homoplasmic wild-type T at m.8993 position (Fig. 1). Filtering was performed to retain only chrM sequences using Samtools view18. To obeserve variation and ensure dataset diversity, we employed the seqtk sample algorithm19. This algorithm was used to generate 100 FASTQ files, each retaining only 10% of the original data, selected randomly with repetitions. This approach was chosen to reduce bias in assessment of basecalling performance across different subsets of data and ensuring the stability of the results.
Sequencing and bootstrap simulations for evaluating heteroplasmy detection using long and short reads. Two individuals sharing the same mtDNA haplogroup yet distinguished by a single-nucleotide variant (m.8993T > G) in their mtDNA were sequenced using PromethION for long reads and MiSeq for short reads. The resulting datasets from these two sequenced samples served as the basis for conducting three dataset bundle, yielding a total of 592,315 datasets. The initial dataset was utilized for assessing basecalling accuracy, utilizing long-read sequencing data from a healthy individual sample. The second dataset bundle facilitated the evaluation of the limit of heteroplasmy detection. To analyze both long and short reads, 999 datasets were prepared for each heteroplasmy ratio by bootstrapping 10% of the original data separately. Subsequently, the third dataset bundle was employed for detecting heteroplasmy across varying coverage depths. Bootstrap simulations were generated to explore diverse levels of heteroplasmy from both samples across a range of coverage depths spanning from 30x to 1100x for short reads and 20x-1820x for long-reads.
For basecalling accuracy assessment, each of the 100 generated FASTQ files was aligned to the Homo sapiens (human) genome assembly GRCh38 (hg38)20, which includes the mitochondrial revised Cambridge Reference Sequence (rCRS) (GenBank number: NC_012920.1) using Minimap221. Alignment metrics, including coverage, base matches, insertions, deletions, and quality scores, were obtained using Samtools mpileup18. ASCII quality scores were then converted to Phred scores. We analyzed basecalling error rates and accuracy for each position in the mtDNA, excluding position 3107 due to an ambiguous reference nucleotide. Variant calling was conducted using Mutserve2 (v2.0.0-rc13)18 to identify both homoplasmic and heteroplasmic variants, with a minimum heteroplasmy threshold set at 1%. Variants were subsequently filtered out from the error rate and accuracy calculations. Accuracy was determined as the proportion of correctly matched bases to the reference genome, while the error rate was assessed by the proportion of insertions, deletions, and mismatches at each position.
Basecalling accuracy was further examined through a sliding window analysis of specific k-mers in the mtDNA, with a window size of 5 nucleotides (k = 5). This approach aligns with ONT’s use of 5-base segments to correlate sequence with changes in ion current22. Additionally, 4-base and 10-base segments were analyzed to understand how neighboring nucleotide sequences affect basecalling accuracy. Sets of low-quality k-mers were generated and statistically compared to the median Phred score of the total mtDNA.
Coverage calculation for the first dataset bundle
To estimate the average sequencing coverage across the mitochondrial genome. Sequence lengths were extracted directly from the FASTQ files by identifying and summing the lengths of all filtered reads. The cumulative sequence length, representing the total number of bases sequenced for chrM region, was divided by the full length of the mitochondrial genome (16,569 bp). This calculation yielded the average coverage. The resulting coverage metric reflects the average number of reads aligned to each nucleotide position within the mtDNA.
Second dataset bundle generation and analysis of heteroplasmy detection
The second dataset bundle included bootstrap simulations of heteroplasmy to evaluate the heteroplasmy detection limit. This dataset involved gradually increasing the proportion of reads from an individual with the m.8993T > G variant, compared to a sample without the variant. Datasets were generated separately for long and short reads (Fig. 1).
For long reads, filtering was performed to retain only chrM sequences using Samtools view, followed by trimming the first 50 bases with NanoFilt23. Reads shorter than 5,000 bp were excluded using Filtlong24 to ensure consistent mtDNA coverage and improve the likelihood of detecting the m.8993T > G variant. The combination of filtering, sgRNA design for Cas9 and mtDNA enrichment minimized the impact of numts on heteroplasmy detection. Multiple datasets for long reads were prepared by combining FASTQ files from the individual with m.8993T > G variant and healthy control. To assess variations in results and reduce bias in our estimates, we retained only 10% of the original data from the Leigh syndrome sample. The healthy sample was bootstrapped to match the number of reads in the Leigh syndrome sample. These filtering steps and the bootstrapping process reduced mtDNA coverage. Out of each bootstrapped dataset, reads from an individual with m.8993T > G variant and healthy control in various ratios ranging from 0 to 100 in increments of 1 were selected using the seqtk sample algorithm tool. This process generated 999 FASTQ files per heteroplasmy level, totaling 201,798 FASTQ files for long reads, which were used to analyze heteroplasmy detection.
These FASTQ files were aligned to the hg38 reference genome using Minimap2 (100,899 FASTQ files) and Ngmlr (100,899 FASTQ files)25, producing BAM files for variant analysis with Mutserve2 (v2.0.0-rc13).
For comparative purposes, 100,899 FASTQ files were generated with bootstrap simulations using short reads. Short reads were filtered to the chrM region using Samtools view. Although short reads were not subjected to the same filtering processes as the long reads, their coverage depth was matched to that of the long reads to ensure balanced representation. This approach maintained comparable coverage across both sequencing types, which is crucial for comparing heteroplasmy detection between the two methods. The same bootstrapping protocol, as applied for long-read sequencing, was used to generate different heteroplasmy ratios, and these FASTQ files were mapped to reference hg38 using BWA26, followed by variant calling with Mutserve2.
The thresholds for reliable detection of heteroplasmy in both short-read and long-read sequencing were established by identifying the inflection points derived from the model fitted to the data, whereas model considered the number of successfully identified variants across different levels of heteroplasmy.
During heteroplasmy detection evaluation, information loss was noted and further analyzed using Clair327 for long reads and MtoolBox28 for short reads. Additional FASTQ files with bootstrap simulations representing heteroplasmy ratios from 85% to 100% were prepared. Clair3 was used to call variants in the long-read dataset (n = 15,984), while MtoolBox was used for the short-read dataset (n = 15,984).
Third dataset bundle generation and analysis of heteroplasmy detection across various coverages
The third dataset bundle involved simulation of heteroplasmy to assess heteroplasmy detection across different coverage depths (Fig. 1). The long-read sequencing data were first filtered as described in the section »Second dataset generation and analysis of heteroplasmy detection«, which included retaining only the chrM region, trimming the first 50 bases, and excluding reads shorter than 5,000 bp. Multiple datasets for long reads were then prepared by combining FASTQ files from the individual with m.8993T > G variant and healthy control. To achieve different coverage levels, the number of reads was incrementally decreased in each dataset. Using the seqtk sample algorithm, reads were selected in various ratios, ranging from 0% to 100% in 1% increments, representing different levels of heteroplasmy. Our initial pool of reads consisted of 6500 reads per sample, and we simulated heteroplasmy from 0% to 100%. At each step, we reduced the total pool of reads by 10 random reads and again generated set of heteroplasmy from 0% to 100%. In total, we performed 650 steps and produced 65,650 FASTQ files with coverage depths ranging from 30x to 1820x.
For long reads, FASTQ files were aligned to the hg38 reference genome using Minimap2 (65,650 FASTQ files) and Ngmlr (65,650 FASTQ files), producing BAM files for variant analysis with Mutserve2 (v2.0.0-rc13).
Similarly, the short-read sequencing data were first filtered to chrM region. Multiple datasets for short reads were then prepared by combining FASTQ files from the individual with m.8993T > G variant and healthy control. Our initial pool of reads consisted of 125,000 reads per sample, and we simulated heteroplasmy from 0% to 100%. At each step, we reduced the total pool by 100 random reads and again generated set of heteroplasmy from 0% to 100%. In total, we performed 1250 steps and produced 126,250 FASTQ files with coverage depths ranging from 30x to 1100x. FASTQ files from short-read data (126,250 FASTQ files) were aligned to hg38 using BWA, followed by variant calling with Mutserve2 (v2.0.0-rc13). The aim was to determine the minimum detectable heteroplasmy across varying depths of coverage.
Coverage calculation for the second and third dataset bundle
Coverage at the mt.8993T > G position was assessed by calculating the number of reads specifically aligned to this nucleotide. Reads overlapping the mt.8993 position were identified, and the total count of these reads was used to determine the coverage at this site. This position-specific coverage was crucial for quantifying the number of reads supporting the heteroplasmic variant at mt.8993T > G, enabling precise analysis of heteroplasmy at this location within the mitochondrial genome. All datasets used for heteroplasmy analysis were considered, and the median coverage with the interquartile range (IQR) across all files was calculated.
Assessment of variant caller efficacy
To assess the performance of the variant caller, we aligned 100,899 FASTQ files from long-read sequencing, originally generated for heteroplasmy detection during the second bootstrap simulations. These files were processed using Minimap2 and Mutserve2, based on findings from long-read heteroplasmy detection analysis (from second dataset bundle). The resulting VCF files were then used to compute precision (Eq. 1), recall (Eq. 2), and F1 score (Eq. 3). True Positive (TP) variants were those present in the true variant sets and correctly identified by Mutserve2. True Negative (TN) variants did not exist in the true variant sets and were not detected by Mutserve2. False Positives (FP) represented variants not present in the true variant sets but detected by Mutserve2, while False Negatives (FN) were variants present in the true variant sets but missed by Mutserve2. Notably, the heteroplasmic variant at m.8993 was excluded from the performance analysis of Mutserve2 if the heteroplasmy was below a threshold of 12%, as determined in heteroplasmy detection analysis for long reads (from second dataset bundle).
Equation 1: Precision calculation for variant caller.
Equation 2: Recall calculation for variant caller.
Equation 3: F1 score calculation for variant caller.
Results
Assessment of basecalling accuracy of long-read sequencing
The PromethION provided a sequencing mtDNA depth of 12,466x for the healthy control and 6,264x for the individual with Leigh syndrome. Due to the higher depth in the healthy control, it was selected for basecalling accuracy assessment. Utilizing only 10% of read per iteration reduced the coverage depth to 1247x. The analysis was restricted to non-variant positions, resulting in the exclusion of 137 positions and the inclusion of 16,432 positions for analysis. Phred scores per-base across the mtDNA predominantly (97.8%) fell within the 20 to 35 range. Phred quality scores provide an estimate of the accuracy of base calls in DNA sequencing. A higher Phred score reflects a higher probability that the base is correctly called. Phred score 20 to 35 corresponds to a probability of an incorrect base call, equating to a base call accuracy of approximately 99–99.9% (Fig. S1). NanoPlot analysis showed a mean read quality of 12.1, with a 33.3% of the reads above 15 (Fig. S1).
In accuracy assessment, 99.6% of positions displayed rates from 0.8 to 1, with exceptions (0.15% of positions) dropping to as low as 0.6. Despite excluding the “N” at position 3107, reduced accuracy was noted at position 3106 (Fig. S2). Error rates analysis revealed that 99.2% of the positions had error rates under 0.2, consistent with the calculated accuracy values (Fig. S2).
K-mer profiling for identifying sequence patterns and their impact on basecalling accuracy
Specific sequence patterns that might influence the accuracy of basecalling and sequencing were identified by calculating the median Phred scores for k-mers. The 4-mer “CGGG” had the lowest Phred score of 22.04 (1.81) [median (IQR)]. Similarly, “CGGGG” among 5-mers had a Phred score of 20.15 (2.02) [median (IQR)] (Table 1). The median Phred scores of the top five lowest-scoring k-mers were significantly lower compared to the entire mtDNA sequence, confirmed by the Kruskal-Wallis test (Table 1).
For each k-mer length, sequencing quality was assessed by examining the GC content based on Phred scores. This included 5% and 10% of the k-mers with the highest and lowest Phred scores, respectively. Neither narrow (5%) nor broader (10%) selection altered GC content percentages. However, k-mers with lowest Phred scores consistently showed higher GC content (0.60 to 0.80 [median (IQR)]), while high-quality k-mers had lower GC content (0.00 to 0.3 [median (IQR)]), with significant difference (P = 0.0036, Kruskal-Wallis). This pattern suggests that a higher GC content may be associated with lower sequencing quality (Table 2).
Quantifying heteroplasmy limits of detection
To evaluate the limit of heteroplasmy detection on long versus short reads, multistep data processing was utilized, significantly reducing the coverage of the original data set by 82-fold for the healthy control (from 12,466x to 152x) and 41-fold for the patient with Leigh syndrome (from 6,264x to 152x). The depth of coverage of short reads was adjusted to that of long reads, resulting in a coverage of 166 × (816x) [median (IQR)]. This large range in coverage is attributed to the nature of short-read sequencing based on hybridization with probes.
Mutserve2 detected heteroplasmy as low as 3% in long reads regardless of the aligner used (Minimap2 and Ngmlr). However, consistently identifying these extremely low-level variants proved challenging, with only 2 out of 999 tests successfully identified heteroplasmy at such levels. To assess the reliability of heteroplasmy detection, we established a detection threshold with a 95% confidence interval. The threshold was determined at 12%, for coverage of 152 × (23x) [median (IQR)], indicating the point at which heteroplasmy detection became significantly more reliable for long-read sequencing (Fig. 2A).
Heteroplasmy above 21%, provided there is a minimum mtDNA coverage depth of 150x, achieves 90% reliability. The reliability reflects the proportion of successful variant detections, indicating that 899 out of 999 tests successfully identified the variant at position m.8993. However, for heteroplasmy between 91% and 98%, there is a decrease in the proportion of called variants. This suggests that the variant predominant in the majority of the reads was not identified in 3–8% of tests with combination of Minimap2 aligner and Mutserve2 variant caller and in 5–13% of tests with combination of Ngmlr aligner and Mutserve2 due to being flagged as strand bias and subsequently filtered from the final VCF file (Fig. 2A and B). Notably, there appears to be a tendency for Mutserve2 to underestimate heteroplasmy in long-read sequencing data. However, there is a strong correlation (R2 = 0.960 for Minimap2, R2 = 0.955 for Ngmlr) between the detected and expected heteroplasmy (Fig. 2A and B), and it is not expected that underestimated calls would negatively impact the diagnostic result.
Detecting heteroplasmy using Mutserve2 on bioinformatically simulated heteroplasmy ratios. The detection of heteroplasmy concerning a variant at position m.8993 utilizing both long (A, B) and short (C) reads via various aligners: Minimap2 (A), Ngmlr (B), and BWA (C). Heteroplasmy was identified within the BAM files utilizing Mutserve2. The plots depict the detection of heteroplasmy at the individual level (I.), with the mean of called heteroplasmy at each level represented by an orange line for long reads aligned with Minimap2 (A), a blue line for long reads aligned with Ngmlr (B), and a red line for short reads aligned with BWA (C). In each plot, the expected heteroplasmy based on the FASTQ mixture is displayed as a black line, while the proportion of successfully called variants (II.) is shown with a green dashed line along with the corresponding model (dotted line). For each model, the inflection point was determined, along with a 95% confidence interval, beyond which heteroplasmy detection became more consistent and reliable.
Additionally, we simulated varying heteroplasmy using short reads to compare the detection across both sequencing platforms. Mutserve2 detected 1% heteroplasmy on short reads; however, calls at such low levels were not consistent. The threshold at which heteroplasmy calls became significantly more reliable was determined above 3% heteroplasmy at a coverage above 150x. A 90% reliability of variant calling (899 of 999 performed tests) was achieved at 9% heteroplasmy with coverage above 150x. Strand bias flags persisted with short reads, affecting 2–5% of tests of heteroplasmy detection for heteroplasmy between 91% and 98%. Compared to long reads, Mutserve2 showed greater sensitivity with short reads, with a strong correlation between detected and expected heteroplasmy (R2 = 0.998) (Fig. 2C).
Because almost homoplasmic variants (more than 90% heteroplasmy) not being detected in 2–13% of tests of heteroplasmy detection using Mutserve2, we reanalyzed the data with two independent variant callers for heteroplasmy ranging from 85% to 100%. Clair3 was used for long reads, while MToolBox was used for short reads. Both variant callers consistently detected the variants in all tests indicating potential bias in the Mutserve2 algorithm.
Sensitivity of heteroplasmy detection at different coverage depths
To explore low-level heteroplasmy detection relative to sequence coverage depth, the minimum detected heteroplasmy across sequencing depths was determined. This systematic coverage variation was applied to both sequencing platforms.
Our findings indicate that increased coverage substantially enhances the detection of heteroplasmy. However, there appears to be a saturation point beyond which additional coverage does not markedly improve the identification of low-level heteroplasmy. Specifically, we detected 5% heteroplasmy at a depth of coverage of 378.5 × (202.4x) [median (IQR)] (Fig. 3C) for short reads and 810.6 × (368.8x) [median (IQR)] (Fig. 3A) and 707.6 × (389.3x) [median (IQR)] (Fig. 3B) for long reads aligned with Minimap2 and Ngmlr, respectively. Notably, the fitted curve asymptotic value stands at 3.0% heteroplasmy for short reads and 3.6% heteroplasmy for both long reads aligned with Minimap2 and Ngmlr (Fig. 3).
Detecting heteroplasmy using Mutserve2 on bootstrapped simulated heteroplasmy ratios across a spectrum of depth of coverage. The detection of low-level heteroplasmy at different depths of coverage using Mutserve2 at bootstrapped simulated heteroplasmy. Plots illustrating the detection of heteroplasmy using BAM files, employing both long (A, B) and short reads (C) through different aligners: Minimap2 (A), Ngmlr (B), and BWA (C). The plotted data represent the lowest detected heteroplasmy concerning the depth of coverage. The orange line corresponds to a fitted curve with a 95% confidence interval for long reads aligned with Minimap2 (A), the blue line corresponds to long reads aligned with Ngmlr (B), and the red line corresponds to short reads aligned with BWA (C). Data is plotted only for coverage below 1100x.
Performance of variant caller
The performance of Mutserve2 was evaluated based on the 100,899 VCF files from long reads aligned with Minimap2. At a coverage depth of 152 × (23x) [median (IQR)], the precision was 0.70 (0.070) [mean (SD)], the recall was 0.91 (0.033) [mean (SD)], the F1 score was 0.79 (0.048) [mean (SD)] (Fig. 4), and the accuracy was 1.0 (0.0001) [mean (SD)].
Discussion
Precise genetic variant identification is crucial for disease risk assessment, treatment response monitoring and genetic counselling. Transitioning to long-read sequencing addresses challenges in the complex genome regions but introduces novel difficulties. Simulating heteroplasmy using mtDNA, we demonstrated that the accuracy of long-read sequencing and basecalling is adequate for the accurate detection of genetic variants, including being effective for identifying low-level heteroplasmic variants. Furthermore, we ascertained a 12% threshold for reliable detection of low-level variants at 150x coverage, critical for the diagnosis of mitochondrial diseases and genetic counselling10.
Long-read sequencing has advanced significantly in recent years, offering improved accuracy and extended capabilities, particularly in identifying structural variants that may be missed by short-read sequencing29. One of the advantages of long-read sequencing lies in its ability to facilitate real-time analysis, making it a powerful tool for genomics research. Notably, long-read sequencing has the ability to help deconvolute numts, which is especially important when detecting low-level heteroplasmic variants. Previous studies have demonstrated that long-read sequencing can successfully map pseudogenes, revealing missense variants that short-read datasets failed to identify due to preferential mapping of variant-containing reads to downstream pseudogenes30. This capability can significantly improve clinical diagnostics by reducing misalignment errors associated with numts. Furthermore, long-read sequencing technology can sequence native DNA, allowing for the detection of epigenetic modifications that are likely to play a crucial role in the future diagnosis of mitochondrial diseases, thus enhancing its clinical utility. Progress in flow cells and library preparation kits has further improved sequencing accuracy, offering better detection of low-level heteroplasmy, although identifying extremely low levels remains a challenge. With these advancements and the ongoing development of reliable bioinformatics tools, long-read sequencing is poised to become a critical element in mitochondrial diagnostics. These improvements will likely enable more accurate and comprehensive analyses of genetic mechanisms. While many of these technical capabilities have been demonstrated, their application in clinical settings remains a promising area for future research.
One objective of this study is to evaluate error rates in ONT sequencing. Our focus on ONT arises from the need to better understand its error profiles, especially as ONT technology becomes more prevalent in research and clinical applications. While error rate evaluations for short-read sequencing have been extensively documented in the literature31,32,33,34, including studies that identified sequence-specific errors associated with inverted repeats, GGC sequences35, and motifs such as “CGG” and “GGG”33, there is a gap in knowledge regarding ONT error profiles. Additionally, short-read sequencing has been shown to have higher error rates and increased miscalls in GC-rich regions36. Given that these issues with short-read sequencing are well-documented, our study specifically targets the quality assessment of ONT to highlight the unique error characteristics and challenges of this technology.
In discussing the quality of DNA sequencing, it is crucial to differentiate between Phred scores, which are commonly used in short-read sequencing37, and mean read quality scores, which are reported by quality control tools for long-read sequencing (ONT)23,38. Phred scores are assigned to each DNA base in a sequence39,40, and are calculated from the ASCII code in the FASTQ format. Phred scores aims to assess the confidence of base calls41,42. Analysis of mtDNA Phred scores from FASTQ data revealed that 97.8% of base positions had scores above 20. The mean read quality scores used in long-read sequencing provide an overall assessment of the quality of an entire DNA sequence read rather than focusing on individual bases. Nanoplot yielded a mean read quality score of 12.1. The mean read quality score is computed by averaging the per-base Phred score across the entire read. Transitioning from per-base Phred to mean read quality involves logarithmic scaling, linearization, averaging and back conversion to a logarithmic scale, potentially causing discrepancies due to data dispersion alteration43. Although position 3107 was excluded, low accuracy issues were observed at position 3106, attributed to aligning data to the rCRS, containing an “N” at the position 3107, an error present in the original Cambridge sequence44. Direct alignment with the rCRS can facilitate the identification of false heteroplasmies and small indels near position 310745. Therefore, removing the “N” at position 3107 in the reference mitochondrial genome, correcting base positions post-alignment, and aligning to a shifted genome to account for the circular nature of the mitochondrial genome are considered best practices. These steps ensure meticulous interpretation of findings near this critical position46.
In bioinformatic applications for long-read sequencing data, analyzing k-mers is crucial. Our k-mer analysis uncovered a bias in sequencing quality between GC-rich and AT-rich regions. Lower quality k-mers have a GC content ranging from 0.6 (0.20) [median (IQR)] to 0.8 (0.20) [median (IQR)]. Conversely, higher quality k-mers have a significantly lower GC content, ranging from 0.2 (0.25) [median (IQR)] to 0.3 (0.28) [median (IQR)]. Despite mtDNA GC/AT ratio of 44% GC to 56% AT, lower quality k-mers exhibit higher GC content compared to AT. Although previous studies47,48,49 suggested that long-read sequencing is not affected by GC content bias, our analysis offers a contrasting perspective. We found that the GC content markedly reduced the Phred score, which is consistent with the findings of50. GC-rich regions showed notably lower Phred scores, decreasing from an average of 24.98 for mtDNA to a minimum of 19.08 for 10-mers (p < 0.001). This reduction translates to a 3.86-fold increase in the probability of error rate from 0.0032 to 0.0124. Such higher error rates in GC-rich regions may lead to increased false-positive low-level genetic variants and, more critically, diminish the detection of heteroplasmy if errors coincide with the variant position.
Discrepancies in mtDNA variant detection accuracy among software tools have been noted51. Mutserve2, Freebayes and Nanopanel2 were compared in detecting low-level variants in long-read sequencing data, with Mutserve2 exhibiting the highest F1 score52. Our study focused on identifying heteroplasmy and assessing detection accuracy between long reads aligned with Minimap2 and Ngmlr, and short reads aligned with BWA. Short reads detected heteroplasmy as low as 1%, while long reads detected heteroplasmy as low as 3% at a coverage depth above 150×. However, at these low levels of heteroplasmy, variant calls were not reliable. Detection stability increased above 3% heteroplasmy for short reads and 12% heteroplasmy for long reads. These thresholds were determined based on the inflection points calculated from the fitted model that assessed the number of correctly identified variants at each level of heteroplasmy. Long-read sequencing reached a 90% reliability threshold for heteroplasmy detection when the levels exceeded 21% with over 150x coverage. Notably, decrease in identified variants was observed with Mutserve2 for heteroplasmy between 91% and 98%, a phenomenon not previously documented to our knowledge. Reanalysis with Clair3 for long reads and MToolBox for short reads detected all variants across datasets with heteroplasmy ranging from 85% to 100%, contrasting with Mutserve2 performance. The diagnostic significance of this discrepancy is crucial, as failure to detect a highly heteroplasmic variant in a patient could lead to false negative results and the misidentification of a potential disease-causing variant. Hence, caution is advised when considering the use of Mutserve2 for diagnostic purposes, and stringent validation protocols should be implemented before its clinical application.
Our analysis focused on the effectiveness of low-level heteroplasmy detection concerning coverage depth, revealing that higher coverage enhances heteroplasmy detection capabilities but plateaus beyond a certain threshold. For detecting 5% heteroplasmy events, the minimal required coverage was 378.5 × (202.4x) [median (IQR)] for short reads, 810.6 × (368.8x) [median (IQR)] for long reads aligned using Minimap2, and 707.6 × (389.3x) [median (IQR)] for long reads aligned with Ngmlr. Short-read sequencing requires lower coverage depth for stable heteroplasmy calling compared to long-read sequencing due to more accurate detection and lower error rates. Previous studied utilizing short-read data show false positives and negatives persist even at depths exceeding 3000x. To accurately identify 3% heteroplasmy, 1000x depth is necessary, reducing false negatives53. The ability of ONT to detect low-level variants was reported in52, revealing the ability to detect variants down to 1% heteroplasmy. Nevertheless, the accuracy of determination relies significantly on the depth of coverage. We assert that the reliable detection of 5% heteroplasmy typically requires coverage of approximately 850x for long reads.
In assessing Mutserve2 efficacy for long-read sequencing, we observed a high recall of 0.91 (0.033) [mean (SD)] but a slightly lower precision of 0.70 (0.070) [mean (SD)]. This discrepancy arises from two main factors. First, distinguishing low-level heteroplasmy from sequencing errors challenges variant callers, often resulting in elevated false positives. Second, mtDNA has a lower variant ratio normalized to the length compared to the nuclear genome.
The performance of Mutserve2 was previously evaluated in52, benchmarked against Freebayes and Nanopanel2 using laboratory-prepared heteroplasmy. Mutserve2 outperformed the others, achieving an F1 score of 0.82 for data aligned with Minimap2, albeit with coverage exceeding 1000x due to PCR amplification. In our bootstrapped stimulated dataset, we achieved an F1 score of 0.79 (0.048 [mean (SD)], with a notably lower coverage depth of 152 × (23x) [median (IQR)]. Comparing Mutserve2 performance between short and long reads, an F1 score of 0.74 was reported in 51 for short reads, whereas we determined an F1 score of 0.79 for long reads.
Our research highlights both the strengths and challenges of long-read sequencing in detecting mtDNA heteroplasmy, showcasing its precision in identifying low frequency variants. The objective was to gain insights into the significant improvement of ONT and its potential for direct application in the field of mitochondrial disease diagnostics. Through a detailed evaluation of mtDNA, focusing on quality metrics, we found that long-read sequencing provides accurate detection of genetic variants, including low-level heteroplasmic variants. Specifically, we identified a 12% heteroplasmy detection threshold at a coverage depth of 150x. Although not entirely comparable to short-read sequencing, the quality is expected to improve with advancements in the technology. More importantly, from a diagnostic perspective, long-read sequencing is particularly compelling due to its ability to provide real-time analysis; sequences can be analyzed as they are being sequenced, allowing for variant detection within half an hour of sequencing. Additionally, it enables the identification of structural variants and the extraction of epigenetic information from the same data. These findings can refine bioinformatics practices and enhance the use of long-read sequencing for detailed genetic variant analysis.
However, this study has limitations, including the small sample size potentially limiting the generalizability of the findings. This limitation is primarily due to the specificity and uniqueness of the samples included in our study. To address this, we employed bootstrapping, which involves selecting reads with replacement, to observe variations in the results and mitigate potential bias in our estimates. Another limitation is the study’s focus on only a single-nucleotide variant, which may not fully capture the complexity of heteroplasmy detection.
Data availability
The datasets supporting the current study have not been deposited in a public repository due to privacy and ethical restrictions but are available from the corresponding author on request.
References
Archibald, J. M. Endosymbiosis and eukaryotic cell evolution. Curr Biol. 25, R911–R921 (2015).
Andrews, R. M. et al. Reanalysis and revision of the Cambridge reference sequence for human mitochondrial DNA. Nat Genet. 23, 147 (1999).
Chinnery, P. F. & Hudson, G. Mitochondrial genetics. Br Med Bull. 106, 135–159 (2013).
Pakendorf, B. & Stoneking, M. Mitochondrial DNA and human evolution. Annu Rev Genomics Hum Genet. 6, 165–183 (2005).
Ye, K., Lu, J., Ma, F., Keinan, A. & Gu, Z. Extensive pathogenicity of mitochondrial heteroplasmy in healthy human individuals. Proc Natl Acad Sci USA 111, 10654–10659 (2014).
Gorman, G. S. et al. Mitochondrial diseases. Nat Rev Dis Primers 2 (2016).
Aldossary, A. M. et al. Recent advances in mitochondrial diseases: From molecular insights to therapeutic perspectives. Saudi Pharm J. 30, 1065–1078 (2022).
Rossignol, R. et al. Mitochondrial threshold effects. Biochem J. 370, 751–762 (2003).
Schon, K. R., Ratnaike, T., van den Ameele, J., Horvath, R. & Chinnery, P. F. Mitochondrial diseases: A diagnostic revolution. Trends Genet. 36, 702–717 (2020).
Fazzini, F. et al. Analyzing low-level mtDNA Heteroplasmy-Pitfalls and challenges from bench to benchmarking. Int J Mol Sci. 22, 1–13 (2021).
Mavraki, E. et al. Genetic testing for mitochondrial disease: the United Kingdom best practice guidelines. Eur J Hum Genet. 31, 148–163 (2023).
Duan, M. et al. Evaluating heteroplasmic variations of the mitochondrial genome from whole genome sequencing data. Gene. 699, 145–154 (2019).
Rohlin, A. et al. Parallel sequencing used in detection of mosaic mutations: comparison with four diagnostic DNA screening techniques. Hum Mutat. 30, 1012–1020 (2009).
Vandiver, A. R. et al. Long read mitochondrial genome sequencing using Cas9-guided adaptor ligation. Mitochondrion. 65, 176–183 (2022).
Vandiver, A. R. et al. Nanopore sequencing identifies a higher frequency and expanded spectrum of mitochondrial DNA deletion mutations in human aging. Aging Cell 22 (2023).
Goldsmith, C. et al. Low biological fluctuation of mitochondrial CpG and non-CpG methylation at the single-molecule level. Sci Rep 11 (2021).
Labun, K. et al. CHOPCHOP v3: expanding the CRISPR web toolbox beyond genome editing. Nucleic Acids Res. 47, W171–W174 (2019).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. Gigascience 10 (2021).
Toolkit for processing sequences in FASTA/Q formats. Github. https://github.com/lh3/seqtk. Accessed: 2024 Mar 20.
Schneider, V. A. et al. Evaluation of GRCh38 and de novo haploid genome assemblies demonstrates the enduring quality of the reference assembly. Genome Res. 27, 849–864 (2017).
Li, H. Minimap2: Pairwise alignment for nucleotide sequences. Bioinformatics. 34, 3094–3100 (2018).
Wick, R. R., Judd, L. M. & Holt, K. E. Performance of neural network basecalling tools for Oxford Nanopore sequencing. Genome Biol. 20 (2019).
De Coster, W., D’Hert, S., Schultz, D. T., Cruts, M. & Van Broeckhoven, C. NanoPack: Visualizing and processing long-read sequencing data. Bioinformatics. 34, 2666–2669 (2018).
Ryan Wick. Filtlong. Github. 2017. https://github.com/rrwick/Filtlong. Accessed: 2024 Mar 20.
Sedlazeck, F. J. et al. Accurate detection of complex structural variations using single-molecule sequencing. Nat Methods. 15, 461–468 (2018).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics. 26, 589–595 (2010).
Zheng, Z. et al. Symphonizing pileup and full-alignment for deep learning-based long-read variant calling. Nat Comput Sci. 2, 797–803 (2022).
Calabrese, C. et al. MToolBox: A highly automated pipeline for heteroplasmy annotation and prioritization analysis of human mitochondrial variants in high-throughput sequencing. Bioinformatics. 30, 3115 (2014).
Frascarelli, C. et al. Nanopore long-read next-generation sequencing for detection of mitochondrial DNA large-scale deletions. Front Genet. 14 (2023).
Watson, C. M. et al. Long-read nanopore sequencing resolves a TMEM231 gene conversion event causing Meckel-Gruber syndrome. Hum Mutat. 41, 525–531 (2020).
Benjamini, Y. & Speed, T. P. Summarizing and correcting the GC content bias in high-throughput sequencing. Nucleic Acids Res. 40, e72–e72 (2012).
Meacham, F. et al. Identification and correction of systematic error in high-throughput sequence data. BMC Bioinformatics. 12, 451 (2011).
Schirmer, M., D’Amore, R., Ijaz, U. Z., Hall, N. & Quince, C. Illumina error profiles: Resolving fine-scale variation in metagenomic sequencing data. BMC Bioinformatics. 17, 125 (2016).
Stoler, N. & Nekrutenko, A. Sequencing error profiles of Illumina sequencing instruments. NAR Genom Bioinform. 3 (2021).
Nakamura, K. et al. Sequence-specific error profile of Illumina sequencers. Nucleic Acids Res. 39, e90–e90 (2011).
Dohm, J. C., Lottaz, C., Borodina, T. & Himmelbauer, H. Substantial biases in ultra-short read data sets from high-throughput DNA sequencing. Nucleic Acids Res. 36 (2008).
Andrews S. FastQC: A quality control tool for high throughput sequence data. http://www.bioinformatics.babraham.ac.uk/projects/fastqc. Accessed: 2024 Mar 20.
Leger, A. & Leonardi, T. pycoQC, interactive quality control for Oxford Nanopore Sequencing. J Open Source Softw. 4, 1236 (2019).
Ewing, B., Hillier, L. D., Wendl, M. C. & Green, P. Base-calling of automated sequencer traces using phred I. Accuracy assessment. Genome Res. 8, 175–185 (1998).
Ewing, B. & Green, P. Base-calling of automated sequencer traces using phred. II. Error probabilities. Genome Res. 8, 186–194 (1998).
Kwon, S., Park, S., Lee, B. & Yoon, S. In-depth analysis of interrelation between quality scores and real errors in Illumina reads. Annu Int Conf IEEE Eng Med Biol Soc. 2013, 635–638 (2013).
Cock, P. J. A., Fields, C. J., Goto, N., Heuer, M. L. & Rice, P. M. The Sanger FASTQ file format for sequences with quality scores, and the Solexa/Illumina FASTQ variants. Nucleic Acids Res. 38, 1767–1771 (2010).
EPI2ME. https://labs.epi2me.io/quality-scores/. Accessed: 2024 Mar 20.
MITOMAP. https://www.mitomap.org/foswiki/bin/view/MITOMAP/WebHome. Accessed: 2024 Mar 20.
Guo, Y. et al. The use of next generation sequencing technology to study the effect of radiation therapy on mitochondrial DNA mutation. Mutat Res. 744, 154–160 (2012).
Maude, H. et al. NUMT confounding biases mitochondrial heteroplasmy calls in favor of the reference allele. Front Cell Dev Biol. 7 (2019).
Karlsson, E., Lärkeryd, A., Sjödin, A., Forsman, M. & Stenberg, P. Scaffolding of a bacterial genome using MinION nanopore sequencing. Sci Rep. 5 (2015).
Browne, P. D. et al. GC bias affects genomic and metagenomic reconstructions, underrepresenting GC-poor organisms. Gigascience. 9 (2020).
Goldstein, S., Beka, L., Graf, J. & Klassen, J. L. Evaluation of strategies for the assembly of diverse bacterial genomes using MinION long-read sequencing. BMC Genomics 20 (2019).
Delahaye, C. & Nicolas, J. Sequencing DNA with nanopores: Troubles and biases. PLoS ONE 16 (2021).
Ip, E. K. K. et al. Benchmarking the effectiveness and accuracy of multiple mitochondrial DNA variant callers: Practical implications for clinical application. Front Genet. 13, 692257 (2022).
Lüth, T. et al. Benchmarking low-frequency variant calling with long-read data on mitochondrial DNA. Front Genet. 13 (2022).
González, M del M., Ramos, A., Aluja, M. P., Santos, C. Sensitivity of mitochondrial DNA heteroplasmy detection using Next Generation Sequencing. Mitochondrion 50, 88–93 (2020).
Acknowledgements
Slovenian Research Agency is acknowledged for continuous support.
The National Medical Ethics Committee of the Republic of Slovenia ethical approval No. 132/03/15. Young Research Fellowship Slovenian Research Agency (SRA) #56916, Tertiary UMCLJ project 20210119, SRA research project J3-50122.
Author information
Authors and Affiliations
Contributions
B.S. and J.K. conceived the original idea. B.S. and R.Š. conducted the experiments and performed the computations and analysis. B.S., R.Š., and K.Č. contributed to the interpretation of the results. J.K. provided project supervision. B.S. drafted the manuscript with support from R.Š., K.Č., J.K., T.T., and B.J.B. All authors participated in discussion of the results and contributed to the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Slapnik, B., Šket, R., Črepinšek, K. et al. The quality and detection limits of mitochondrial heteroplasmy by long read nanopore sequencing. Sci Rep 14, 26778 (2024). https://doi.org/10.1038/s41598-024-78270-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-78270-0