Abstract
Gastric cancer (GC) is the fifth most common cancer and the third most common cause of cancer death globally, with high invasiveness, high recurrence rate, and poor prognosis. Multiple studies have shown that Metallothionein-1G (MT1G) is closely associated with oxidative stress, ferroptosis, and autophagy. However, the role and potential mechanisms of MT1G in GC have not been fully elucidated. This study aims to explore the biological functions and regulatory mechanisms of MT1G in GC. Perform bioinformatics analysis using the TCGA database to investigate the expression of MT1G in GC. RT-qPCR and Western blot were used to detect the expression of MT1G, ferroptosis related proteins, autophagy related proteins and ARNTL clock autophagy related proteins in Hgc27, MKN45 and AGS cell lines. Exploring the biological functions of MT1G overexpressing GC cell lines through wound healing and transwell experiments. Use specific fluorescence probes to examine mitochondrial membrane potential and Fe2+ fluorescence intensity. Using immunoprecipitation analysis (CO-IP) to elucidate the association between GC cell lines GPX4, SQSTM and ARNTL. Use flow cytometry to detect ROS expression. Observation of autophagy related morphological changes in cells using transmission electron microscopy. Compared with gastric mucosal cell lines, the expression of MT1G is decreased in three gastric cancer cell lines (Hgc27, MKN45 and AGS). Overexpression of MT1G inhibits the proliferation, migration, and invasion functions of GC cells, reduces SOD and GSH content, increases MDA content, cause the mitochondrial membrane potential to weaken and promote the transformation of JC-1 aggregates to JC-1 monomer, increases Fe2+, affects ROS, and reduces GPX4 and SLC7A11 protein expression, promoting ferroptosis. Overexpression of MT1G promotes the transformation of LC3B I to LC3B II, reduces SQSTM1 protein expression, and leads to the appearance of more autophagosomes and autolysosomes at low magnification. At high magnification, mitochondrial autophagy, endoplasmic reticulum autophagy, lipid droplet autophagy, and wrinkled mitochondrial cristae are observed, promoting autophagy. Overexpression of MT1G inhibits GPX4, thereby affecting SQSTM1 as a vector to promote ARNTL autophagy and EGLN2, promoting ARNTL clock autophagy through the GPX4/SQSTM1 axis. Our research findings elucidate that overexpression of MT1G promotes iron autophagy centered around ARNTL in GC cells via the GPX4/SQSTM1 axis, thereby inhibiting GC cell function and providing a new molecular mechanism and therapeutic target for the development of GC.
Similar content being viewed by others
Gastric cancer (GC) is the fifth most common cancer and the third most common cause of cancer death globally1. Over 95% of GC are adenocarcinomas, which are typically classified based on anatomic location and histologic type2. Epidemiological findings on GC, Eastern Asia presents the highest incidence of GC, followed by eastern and central Europe3,4. More than 1 million people are newly diagnosed with GC worldwide each year5. The main pathogenic factors of GC are Helicobacter pylori infection, excessive sodium intake and family history6,7. Surgical resection as the main treatment for GC, can only bring temporary therapeutic effects, but various factors lead to a decrease in the quality of life of patients after GC resection. Therefore, obtaining new disease targets from GC pathology is of great significance for GC treatment.
Metallothionein-1G (MT1G) have a high content of cysteine residues that bind various heavy metals, these proteins are transcriptionally regulated by both heavy metals and glucocorticoids, such as iron, zinc and copper8. The Metallothionein family proteins have typical functions in regulating cellular metal metabolism, and changes in metal metabolism can lead to various physiological changes in cells9,10. There is correlation of Metallothionein family in tumor growth, differentiation, angiogenesis, metastasis, microenvironment remodeling, immune escape and drug resistance11. Moreover, MT1G as a typical subtype of the Metallothionein family, exhibits outstanding characteristics in oxidative stress12,13. Therefore, studying new targets of GC from the perspective of MT1G is of great value.
Ferroptosis is a type of iron-dependent regulated cell death that plays a context-dependent role in health and disease14,15. Ferroptosis was recognized as a form of autophagy-dependent cell death16. Autophagy is a highly integrated process that maintains cell homeostasis by promoting cell survival or causing cell death17. A studies have shown that two important subtypes of autophagy (macroautophagy and chaperone-mediated autophagy) promote GPX4 degradation and ferritinophagy18.
Here, we found that MT1G exhibits low expression in GC patients, while overexpression of MT1G inhibits the proliferation, migration and invasion functions of GC cell lines. We further revealed that overexpression of MT1G promotes iron autophagy in GC cell lines, which is regulated by the GPX4/SQSTM1 axis. MT1G mainly initiates iron autophagy by promoting low expression of GPX4. Our research suggests that MT1G promoting iron autophagy in GC cell lines can be a valuable new approach as a GC replacement therapy strategy.
Results
High expression of MT1G inhibits the function of GC cell lines
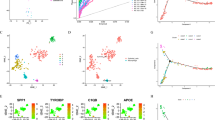
The results of TCGA pan cancer analysis showed that the expression level of MT1G in non cancerous tissues was higher than that in GC tissues (Fig. 1a). Comparing the MT1G mRNA content in different cell lines under different states using RT-qPCR, the MT1G mRNA content in normal gastric mucosal cell line GES1 and gastric mucosal inflammatory cell line MC was higher than that in GC cell lines Hgc27, MKN345 and AGS (Fig. 1b). The overexpression of GC cell line MT1G was forced by plasmid transfection, and the transfection effect was verified by RT-qPCR results (Fig. 1c). Observing the correlation between functional changes in GC cell lines and MT1G, we performed cell proliferation assays to study MT1G effect in GC cell function. We used transwell assay to detect the function of MT1G in cell migration and invasion. We found that overexpression of MT1G significantly suppressed the cell migration and invasion abilities in Hgc27, MKN345 and AGS cell lines (Fig. 1d–g). The results of scratch experiments at 12H and 24 H demonstrate that overexpression of MT1G significantly inhibits the proliferation ability of GC cell lines (Fig. 1h–j). All these results demonstrated that overexpression of MT1G inhibition the proliferation, migration and invasion of Hgc27, MKN345 and AGS cells.
High expression of MT1G inhibits the function of GC cell lines. (a) MT1G in TCGA pan cancer analysis results. (b) Comparison of MT1G mRNA expression in different cell lines using RT-qPCR. (c) RT-qPCR was used to test the transfection efficiency of MT1G plasmid. (d) Statistical analysis of 24-h cell migration in the Transwell experiment GC cell line. (e) Statistical analysis of 24-h cell invasion in the Transwell experiment GC cell line. (f) Microscopic observation of 24H cell migration in the Transwell experimental GC cell line. Scale bar = 50 μm. (g) Microscopic observation of 24-h cell migration in Transwell experiment GC cell line. Scale bar = 50 μm. (h) Statistical analysis of cell proliferation in the 12H GC cell line scratch test. (i) 24-h GC cell line scratch assay cell proliferation statistics. (j) GC cell line scratch experiment observed under a microscope. **p < 0.01; ***p < 0.001.
A correlation between the effect of MT1G on GC cell lines and the occurrence of ferroptosis
Further we explored the effect of MT1G in GC cell lines and detect changes in oxidative stress indicators in overexpressing cell lines. Compared with normal GC cell lines, the MT1G overexpressing cell line showed a significant increase in MDA (Fig. 2a), a significant decrease in SOD (Fig. 2b), and a significant decrease in GSH (Fig. 2c). The trend of MDA, SOD and GSH changes in MT1G overexpressing GC cell lines is consistent with the trend of MDA, SOD and GSH changes during ferroptosis. To clarify the association between ferroptosis and MT1G overexpressing GC cell lines, the expression of ferroptosis marker proteins in different cell lines was further detected (Fig. 2d). WB results showed that the expression of GPX4 protein in Hgc27, MKN345 and AGS cell lines overexpressing MT1G significantly decreased (Fig. 2e), while the expression of SLC7A11 protein in Hgc27 and AGS cell lines overexpressing MT1G decreased (Fig. 2g), and the change in MKN45 was not significant. The overexpression of MT1G in Hgc27, MKN45 and AGS cell lines showed no significant changes in FSP1 and DHODH (Fig. 2f,h). The fluorescence results of ferrous ion (Fe2+) showed that compared with the untreated cell line, the fluorescence intensity of the MT1G overexpressing cell line was significantly enhanced (Fig. 2i). By detecting changes in cellular ROS, the occurrence of ferroptosis in MT1G overexpressing GC cell lines was clarified. Compared with untreated cell lines, ROS expression in MT1G overexpressing cell lines was inhibited (Fig. 3a). Mitochondria are involved in maintaining the stability of ROS, and changes in JC-1 can intuitively reflect mitochondrial function. In the inner mitochondrial membrane, JC-1 forms aggregates according to the membrane potential. When the mitochondrial membrane potential weakens, JC-1 will transform from aggregates to monomers. The mitochondrial membrane potential fluorescence detection kit uses a specific fluorescent probe to make JC-1 aggregates emit red fluorescence and JC-1 monomers emit green fluorescence. Therefore, changes in mitochondrial membrane potential can be judged by fluorescence color and intensity. There were significant differences in the changes of different GC cell lines JC-1 (Fig. 3b). Compared with the untreated cell line, the fluorescence changes of MT1G overexpressing GC cell line tended to the positive control group(CCCP) fluorescence changes, JC-1 polymer fluorescence decreased, and JC-1 monomer fluorescence increased (Fig. 3c).
A correlation between the effect of MT1G on GC cell lines and the occurrence of ferroptosis. (a) MDA expression in different GC cell lines. (b) The expression of SOD in different GC cell line. (c) The expression of GSH in different GC cell lines. (d) The expression of ferroptosis related proteins in different GC cell lines. (e) The expression of GPX4 protein in different GC cell lines. (f) The expression of FSP1 protein in different GC cell lines. (g) Expression of SLC7A11 protein in different GC cell lines. (h) The expression of DHODH protein in different GC cell lines. (i) Fluorescence intensity of Fe2+ content. Scale bar = 50 μm. *p < 0.05; **p < 0.01; ***p < 0.001.The original images for western blot analysis were shown in “Supplementary information”.
The effect of MT1G overexpression on ROS metabolism during ferroptosis in GC cell lines. (a) ROS expression in MT1G overexpressing GC cell line. (b) Statistical analysis of fluorescence of JC-1 cell lines from different GC strains. (c) Fluorescence expression of JC-1 in different GC cell lines. Scale bar = 50 μm. ***p < 0.001.
The correlation between MT1G overexpression induced ferroptosis and autophagy in GC cell lines
To further explore the effect of MT1G overexpression on autophagy in GC cell lines, autophagy related proteins were detected in Hgc27, MKN45 and AGS cell lines (Fig. 4a). Compared with the untreated cell line, the MT1G overexpressing cell line showed a decrease in the ratio of LC3B I to LC3B II (Fig. 4b), a decrease in GPX4 protein expression (Fig. 4c) and a decrease in SQSTM1 protein expression (Fig. 4d). The Co-IP results confirmed a significant interaction between GPX4 and SQSTM1, which was observed in both Hgc27 and AGS cells (Fig. 4e). Further verification of autophagy in MT1G overexpressing GC cell lines was conducted using transmission electron microscopy. MT1G overexpressing GC cell lines showed more autophagosomes and autolysosomes, while MT1G overexpressing Hgc27 cell lines were found to contain endoplasmic reticulum autophagy and cytoplasmic autophagy. MT1G overexpressing MKN45 cell lines were found to contain autolysosomes and mitochondria with significant shrinkage. MT1G overexpressing AGS cell lines showed more organelles undergoing autophagy (Fig. 4f).
The correlation between MT1G overexpression induced ferroptosis and autophagy in GC cell lines. (a) Expression of autophagy related proteins in different GC cell lines. (b) The protein expression ratio of LC3B I and LC3B II in different GC cell lines. (c) The expression of GPX4 protein in different GC cell lines. (d) The expression of SQSTM1 protein in different GC cell lines. (e) Co-IP detection of the correlation between GPX4 and SQSTM1 in GC cell lines. (f) Observation of different GC cell lines under transmission electron microscopy. Scale bar = 2 μm .Scale bar = 500 nm. *p < 0.05; **p < 0.01; ***p < 0.001.The original images for western blot analysis were shown in “Supplementary information”.
MT1G overexpression in GC cell line intervenes in GPX4/SQSTM1 axis to initiate iron autophagy
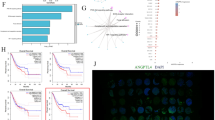
Overexpression of MT1G promotes varying degrees of ferroptosis and autophagy in Hgc27, MKN45 and AGS cell lines. To clarify the pathway through which MT1G overexpression initiates iron autophagy in GC cell lines, a comparison of protein associations between the common platform TCGA and non cancer and gastric cancer tissues was conducted (Fig. 5a). High expression of MT1G was observed in non cancer tissues, while low expression of GPX4, SLC7A11, SQSTM1, ARNTL and EGLN2 was observed, showing the opposite trend in gastric cancer tissues. By calculating the association between MT1G and GPX4/SQSTM1 axis related proteins through algorithms, MT1G showed a negative correlation with GPX4 and SQSTM1, but no direct correlation with SLC7A11, ARNTL and EGLN2. SQSTM1 showed a high correlation with ARNTL and EGLN2 (Fig. 5b). Comparing the changes in GPX4/SQSTM1 related proteins with high and low expression of MT1G in GC patients, the trends in GPX4, SLC7A11 and SQSTM1 were consistent with MT1G, while there was no significant correlation between ARNTL and EGLN2 changes and MT1G (Fig. 5c). The overexpression of MT1G in Hgc27 and AGS cell lines in MT1G has a certain impact on the downstream ARNTL protein of GPX4/SQSTM1 axis, while EGLN2 is weakly affected (Fig. 5d). Compared with the untreated group, the MT1G overexpression group showed a significant decrease in GPX4 protein expression (Fig. 5e), SQSTM1 protein expression (Fig. 5f), ARNTL protein expression (Fig. 5g), and partial decrease in EGLN2 protein expression (Fig. 5h).The Co-IP results confirmed a significant interaction between SQSTM1 and ARNTL, which was observed in both Hgc27 and AGS cells (Fig. 5i).
MT1G overexpression in GC cell line intervenes in GPX4/SQSTM1 axis to initiate iron autophagy. (a) Comparison of protein associations between TCGA non cancerous tissue and gastric cancer tissue. (b) Correlation heatmap between MT1G and GPX4/SQSTM1 axis related proteins. (c) The expression status of GPX4/SQSTM1 axis related proteins in the high and low expression queue of gastric adenocarcinoma. (d) The expression of GPX4/SQSTM1 axis related proteins in different GC cell lines. (e) The ratio of GPX4 protein expression in different GC cell lines. (f) The expression of SQSTM1 protein in different GC cell lines. (g) The expression of ARNTL protein in different GC cell lines. (h) The expression of EGLN2 protein in different GC cell lines. (i) Co-IP detection of the correlation between SQSTM1 and ARNTL in GC cell lines. *p < 0.05; **p < 0.01; ***p < 0.001.The original images for western blot analysis were shown in “Supplementary information”.
Intervention of GPX4 blocks iron autophagy induced by MT1G overexpression
MT1G overexpression promotes iron autophagy in GC cell lines by intervening in the GPX4/SQSTM1 axis. To further clarify the function of MT1G in inhibiting gastric cancer cell lines and promoting iron autophagy, the GPX4/SQSTM1 axis is involved. Fer-1 is an iron chelator that can protect membrane lipids from lipid peroxidation. The small molecule compound Fer-1 is used to inhibit ferroptosis to study the role of the GPX4/SQSTM1 axis.The CCK-8 method was used to detect the effects of DFOM, RSL3 and Fer-1 on the survival rate of gastric cancer cells. The results showed that the optimal intervention concentration of DFOM was 20 mM (Fig. 6a), the optimal intervention concentration of Fer-1 was 20 mM (Fig. 6b), the optimal intervention concentration of RSL3 was 10 mM (Fig. 6c).To further prove whether ferroptosis is involved, we conducted cell viability assays by using CCK-8. Compared with normal GC cells, cell viability decreased when MTG1 was overexpressed. This decrease should be inhibited by the application of Fer-1 and DFOM (Fig. 6d). The RT-qPCR results confirmed that the small molecule compounds RSL3 and Fer-1 could not alter the overexpression effect of MT1G (Fig. 6e). Small molecule compounds RSL3 and Fer-1 can significantly affect GPX4 expression (Fig. 6f,g). Overexpression of MT1G in GC cell lines will reduce the expression of GPX4, and then start the GPX4/SQSTM1 axis. Our research found that the use of the ferroptosis antagonist Fer-1 promotes high expression of GPX4 in MT1G-overexpressing GC cell lines (Fig. 6h). Compared with overexpressing cell lines, the group added with the small molecule compound Fer-1 showed an increase in GPX4 protein expression (Fig. 6i), SQSTM1 protein expression (Fig. 6j), ARNTL protein expression (Fig. 6k), and EGLN2 protein expression (Fig. 6l). The fluorescence results of JC-1 indicate that the small molecule compound Fer-1 can increase the fluorescence intensity of JC-1 polymer by blocking GPX4 (Fig. 6m) and decrease the fluorescence intensity of JC-1 (Fig. 6n). The small molecule compound Fer-1 blocks GPX4, reduces MDA content (Fig. 6o), and increases SOD and GSH content (Fig. 6p,q).
Intervention of GPX4 blocks ferroptosis induced by MT1G overexpression. (a) The CCK-8 method was used to detect the survival of GC cells intervened with DFOM for 24 h. (b) The CCK-8 method was used to detect the survival of GC cells intervened with RSL3 for 24 h. (c) The CCK-8 method was used to detect the survival of GC cells intervened with Fer-1 for 24 h. (d) The effects of different ferroptosis inhibitors on the activity of GC cells. (e) RT-qPCR was used to examine the effects of small molecule compounds RSL3 and Fer-1 on MT1G transfection. (f) The effects of small molecule compounds RSL3 and Fer-1 on GPX4 protein expression in different GC cell lines. (g) Statistical analysis of GPX4 protein expression in different cell lines. (h) The effect of small molecule compound Fer-1 on the expression of GPX4/SQSTM1 axis protein. (i) The expression of GPX4 protein in different GC cell lines. (j) The expression of SQSTM1 protein in different GC cell lines. (k) The expression of ARNTL protein in different GC cell lines. (l) The expression of EGLN2 protein in different GC cell lines. (m) Fluorescence expression of different GC cell lines JC-1. Scale bar = 50 μm. (n) Fluorescence statistics of different GC cell lines JC-1. (o) MDA expression in different GC cell lines. (p) SOD expression in different GC cell lines. (q) The expression of GSH in different GC cell lines. *p < 0.05; **p < 0.01; ***p < 0.001.The original images for western blot analysis were shown in “Supplementary information”.
The fluorescence results of ferrous ion (Fe2+) showed that the small molecule compound Fer-1 weakened the fluorescence of Fe2+ in the MT1G overexpressing gastric cancer cell line (Fig. 7a). The results of transmission electron microscopy showed that the small molecule compound Fer-1 caused fewer autophagosomes and autolysosomes to be produced in the MT1G overexpressing gastric cancer cell line under low magnification, while clear mitochondrial cristae were observed under high magnification (Fig. 7b). The small molecule compound Fer-1 inhibits the occurrence of ferroptosis, inhibits the overexpression of MT1G, and restores the migration and invasion functions of gastric cancer cells through iron autophagy induced by the GPX4/SQSTM1 axis (Fig. 7c–f).
Effects of GPX4 intervention blocking MT1G overexpression on GC iron autophagy and function. (a) Fluorescence expression of Fe2+ in different GC cell lines Scale bar = 50 μm. (b) Observation of different GC cell lines under transmission electron microscopy. Scale bar = 2 μm .Scale bar = 500 nm. (c) Statistical analysis of 24-h cell migration in Transwell experimental GC cell line. (d) Transwell experiment GC cell line 24-h cell invasion statistics. (e) Microscopic observation of 24-h cell migration in Transwell experimental GC cell line. Scale bar = 50 μm. (f) Microscopic observation of 24-h cell migration in Transwell experiment GC cell line. Scale bar = 50 μm. *p < 0.05; **p < 0.01; ***p < 0.001.
Discussion
GC is a cancer with high mortality and incidence rate. More evidence supports that ROS has a better effect on GC than other cancers. The change of cellular ROS increases the incidence of gastrointestinal cancer in many species, including humans and mice19,20. The highly pathogenic factor of gastric cancer, Helicobacter pylori CagA, induces spermine oxidase to produce cells with oxidative DNA damage21. The tendency of Helicobacter pylori to produce DNA strand breaks can lead to genomic instability and promote the occurrence of gastric cancer22. The evidence related to Helicobacter pylori carcinogenesis still needs further improvement, but existing evidence points to oxidative stress as a bridge between Helicobacter pylori and GC23,24,25. In a study on long-term compliance with antioxidants to prevent GC, it was confirmed that the compliance of antioxidants with GC is negatively correlate26.
MT1G is involved in various oxidative stress processes and metal metabolism, and has good research value in oxidative stress27,28. As the main subtype of the metallothionein family, MT1G participates in the metal regulation process11,29. The commonality of the metallothionein family is that it has a strong affinity for heavy metals and can bind well with heavy metals to prevent metal toxicity (including cadmium poisoning)30. At the same time, the metallothionein family can bind to zinc and copper in cells and participate in regulating cell growth and proliferation31. MT1G shows obvious differences in different cancers, and our pan-cancer results also confirm this. In some cancers, researchers have found that MT1G gene silencing is related to high methylation of the promoter, further promoting the progression of thyroid carcinoma, prostate cancer, esophageal squamous cell carcinoma, oral cancer, hepatocellular carcinoma and hepatoblastoma32,33,34,35,36,37,38. Promoter methylation in thyroid cancer will lead to changes in MT1G39,40. Fu et al. revealed the molecular mechanism of MT1G inhibiting thyroid carcinogenesis. By regulating the phosphatidylinositol 3-kinase (PI3K)/AKT and Rb/E2F signaling pathways, it inhibits cell growth and invasion and induces cell cycle arrest and apoptosis41. Fu et al. demonstrated a significant positive correlation between MT1G hypermethylation and lymph node metastasis in 178 patients with thyroid carcinoma41. Chan et al. pointed out that the down-regulation of MT1G in hepatocellular carcinoma is due to allele deletion on chromosome 16q12.1-q23.142. Hirako et al. found that overexpression of MT1G can inhibit the differentiation of all-trans retinoic acid-induced acute promyelocytic leukemia cell NB443. Arriaga et al. transfected the MT1G-myc expression plasmid into colon cancer H29 cells and found that MT1G mainly participates in the tumor cell differentiation process through the Notch signaling pathway and unstable zinc chelation and redistribution44. Arriaga et al. showed that overexpression of MT1G makes colorectal cancer cells sensitive to the chemotherapy drugs oxaliplatin and 5-fluorouracil, which may be mediated by the activation of p53 and the inhibition of NF-κB activity45. Sun et al. demonstrated a new drug resistance molecular mechanism in hepatocellular carcinoma. Enhanced expression of MT1G leads to sorafenib resistance in hepatocellular carcinoma by inhibiting lipid peroxidation-mediated ferroptosis46.
Based on the TCGA database and our findings, MT1G expression is lower in GC than in non cancerous tissues. Further research was conducted to investigate the effect of overexpression of MT1G on the function of GC cells. Functional experiments confirmed that MT1G significantly affects the proliferation, migration and invasion of GC cell lines. This result is consistent with our prediction, confirming the anticancer effect of MT1G in GC cell lines, further promoting our exploration of the molecular mechanism of MT1G action.
In addition to Helicobacter pylori, smoking is also closely associated with oxidative stress47. Smoking is the main risk factor for male gastric cancer48, and tobacco use leads to approximately 80,000 cases of gastric cancer annually (11% of all cases)49. The release of ROS and subsequent oxidative stress in smoking have a significant impact on inflammation and carcinogenesis50,51. Oxidative stress is closely related to the occurrence of GC, and antioxidants have inhibitory effects on GC52,53. Our study found that overexpression of MT1G can inhibit the SOD and GSH content of GC cell lines, and increase the MDA content, indicating that overexpression of MT1G is involved in the changes of oxidative stress in GC cells. Cancer cells are under sustained oxidative stress, and the basic level of ROS increases. Oxidative stress-induced ferroptosis helps to inhibit tumor growth and increase chemical sensitivity54,55. Our subsequent research noted ferroptosis, where overexpression of MT1G promotes low expression of GPX4 and SLC7A11, but has no significant effect on DHODH and FSP1. Fe2+ that important indicator of ferroptosis is also affected by MT1G56, and overexpression of MT1G can significantly enhance the fluorescence intensity of Fe2+. The sources of ROS are divided into exogenous and endogenous. Exogenous ROS is mainly caused by radiation and other conditions, while endogenous ROS is mainly produced by mitochondria and endoplasmic reticulum57,58. In the acidic microenvironment of tumors, Fe3+ ions are first released and reduced to Fe2+ by tannic acid for ROS generation59. The increase of Fe2+ will cause changes in ROS. Our study also confirms that overexpression of MT1G affects ROS expression, and MT1G affects mitochondrial function and converts JC-1 polymer into JC-1 monomer.
The mechanism of oxidative stress-induced autophagy has been elucidated in various diseases, with mitochondrial and endoplasmic reticulum changes being most significant in oxidative stress-induced autophagy60,61. We further explored the localization of autophagy in promoting ferroptosis in GC cell lines by MT1G. Through experiments, we demonstrated that overexpression of MT1G induces a transition from LC3B I to LC3B II and reduces SQSTM1 protein expression. We first validated the association between GPX4 and SQSTM1 in GC cell lines using CO-IP. Under transmission electron microscopy, more autophagosomes and autolysosomes were observed in MT1G overexpressing cells. At high magnification, mitochondrial autophagy and endoplasmic reticulum autophagy were observed, with obvious mitochondrial cristae shrinkage and lipid droplet autophagy. The shrinkage of mitochondrial cristae confirms that overexpression of MT1G promotes the transition of JC-1 polymer to JC-1 monomer. Mitochondrial autophagy and endoplasmic reticulum autophagy confirmed that overexpression of MT1G promotes changes in ROS. We verified through experiments that overexpression of MT1G promotes ferroptosis and autophagy in GC cell lines, and there is a close correlation between ferroptosis and autophagy in GC cell lines. Therefore, we speculate that overexpression of MT1G promotes iron autophagy in GC cells, which significantly alters GPX4 and SQSTM1.
The transcription factor ARNTL (aryl hydrocarbon receptor nuclear translocation factor like protein 1) is a core component of the mammalian circadian clock17,62, and the autophagic degradation of the circadian clock protein ARNTL depends on the cargo receptor SQSTM1. The Co-IP results confirm the correlation between SQSTM1 and ARNTL. ARNTL exerts a previously unknown pro survival function by inhibiting the expression of EGLN2 (proline hydroxylase 2) during ferroptosis63. Under normal conditions, EGLN2 is an oxygen sensor that hydroxylates the proline residue of HIF1A (hypoxia inducible factor 1 subunit), thereby promoting HIF1A degradation64. Under hypoxic conditions, EGLN2 activity decreases, thereby increasing the stability of HIF1A protein65. ARNTL increases HIF1A levels and subsequent ferroptosis resistance by directly downregulating EGLN1 expression. HIF1A inhibition restored the sensitivity of EGLN1 knockdown or ARNTL overexpression cells to ferroptosis. Lipid droplets are dynamic storage organelles found in most eukaryotic cells. HIF1A promotes lipid storage and reduces fatty acid beta oxidation, which helps tumor cell survival66,67. Therefore, we cannot ignore the changes in ARNTL and EGLN2 proteins downstream of the GPX4/SQSTM1 axis. Through experiments, we confirmed that MT1G caused changes in GPX4 and SQSTM1, resulting in changes in ARNTL, but EGLN2 was weakly affected. To further explore the effect of GPX4 on the overexpression of MT1G in GC cell lines SQSTM1 and ARNTL, we inhibited the decrease of GPX4 with the small molecule compound Fer-1 and found that the protein expression of SQSTM1, ARNTL, and EGLN2 increased to varying degrees. Not only that, after the small molecule compound Fer-1 inhibited the decrease of glutathione peroxidase 4 (GPX4), the contents of superoxide dismutase (SOD) and glutathione (GSH) increased, malondialdehyde (MDA) decreased, the fluorescence expression of JC-1 polymer increased, the fluorescence of Fe2+ weakened, and the number of autophagosomes and autolysosomes decreased under transmission electron microscopy.Clear mitochondria and endoplasmic reticulum were observed under high magnification, and the migration and proliferation functions of GC cell lines were also improved.In GC cell lines, ARNTL can activate downstream EGLN2. EGLN2 further activates HIF1A to perform fat uptake and store. After overexpressed MT1G enters GC cells through plasmid transfection, it causes a decrease in GPX4. The decrease in GPX4 activates SQSTM1, forming a double-layer membrane structure of autophagy. Autophagy of ARNTL occurs, and fat uptake and store are interrupted. At the same time, the decrease in GPX4 causes a decrease in SLC7A11. Mitochondria in cells shrink, and ferroptosis occurs (Fig. 8). In summary, targeting MT1G on GC cell lines is a novel molecular mechanism and therapeutic target.
Of course, we must admit that our research has certain limitations. We focused on the effect of MT1G overexpression on the function of GC cell lines in the study of the GPX4/SQSTM1 axis, and proved that overexpressed MT1G in GC cell lines will induce ferritinophagy, affect the GPX4/SQSTM1 axis, and inhibit the function of gastric cancer cells. In the study of hepatocellular carcinoma by Wang et al., it was found that MT1G can interact with P53 to inhibit some functions of cancer. Iron loss induced by iron chelators can also induce mitochondrial autophagy independently of SQSTM1 and is related to HIF-1α signaling8. SQSTM1 is a common ubiquitin-dependent selective autophagy receptor. In addition, NBR1, OPTN and NDP52 are also common autophagy receptors68,69. They can bind cargo and ubiquitin at the same time to initiate pathways leading to autophagy initiation and membrane recruitment. There is a classic correlation between ferritinophagy and the HIF-1α signaling pathway. That is, when GPX4 decreases, autophagy receptors will be activated, prompting the circadian rhythm regulatory protein ARNTL to undergo autophagy. The physiological function of ARNTL is to affect downstream EGLN2. EGLN2 activates downstream HIF-1α. Activation of HIF-1α can play a role in fat uptake and storage17,70. When ARNTL undergoes autophagy, the downstream will be inhibited, and EGLN2 and HIF-1α will also change accordingly71. Our experiment also confirmed this. In the study of hepatocellular carcinoma, it is also indicated that when ARNTL undergoes autophagy and changes downstream HIF-1α, other autophagy receptors appear8. This new receptor may be different from the autophagy receptors found in other existing studies. However, in existing research reports, there is a lack of research reports on new autophagy receptors when ARNTL undergoes autophagy. We believe that the new autophagy receptor of ARNTL will be a very interesting research and worthy of in-depth exploration. Similarly, we also noticed the importance of NCOA4 in the study of ferritinophagy72,73. As a star protein in the study of ferritinophagy, NCOA4 has clear evidence to prove its important mechanism in the occurrence of ferritinophagy74. NCOA4 can directly interact with ferritin and degrade it in a ferritinophagy-dependent manner, and then release a large amount of iron. Cytoplasmic Fe2+ further activates the expression of iron-containing proteases on the mitochondrial membrane, and then transports cytoplasmic Fe2+ into mitochondria, producing mitochondrial ROS and ferroptosis. The promotion of ferritinophagy by NCOA4 has been confirmed in diseases such as diabetic complications, sepsis, Parkinson’s disease, acute kidney injury, cerebral ischemia–reperfusion injury and osteoarthritis75,76,77,78,79,80,81. NCOA4 promoting ferritinophagy has also been found in cancers such as glioma, pancreatic cancer, hepatocellular carcinoma and head and neck cancer82,83,84,85. We believe that NCOA4 is involved in ferritinophagy in GC cells. Our research did not conduct in-depth research on the role of NCOA4 in the process of ferritinophagy in GC cells caused by MT1G overexpression. Our research is limited to MT1G overexpression promoting the GPX4/SQSTM1 axis, occurrence of ferritinophagy, and inhibiting the function of GC cell lines. We also have to admit that there are certain limitations in our experiment. In the experiment to prove ferroptosis, we detected ROS but not lipid ROS. Lipid ROS is an important observation index in ferroptosis experiments as it can directly reflect the level of oxidative stress in cells. We used the detection of GSH, MDA, SOD and mitochondrial function to make up for this problem. Although a large amount of evidence has proven the occurrence of ferroptosis at the current level, there are some limitations for scientific issues. We look forward to continuing to demonstrate our conclusion with supplementary evidence in subsequent experiments.
In summary, we have demonstrated that MT1G is lowly expressed in GC and closely associated with GC cell lines. In terms of functionality, MT1G overexpression can effectively inhibit the proliferation, migration and invasion functions of GC cell lines. In terms of mechanism, overexpression of MT1G induces a decrease in GPX4 in GC cells, leading to ferroptosis of GC cell lines. The decrease in GPX4 further promotes SQSTM1 as an ARNTL autophagy vector, facilitating the occurrence of clock autophagy and inhibiting EGLN2 expression. In summary, our research findings elucidate that overexpression of MT1G promotes iron autophagy centered around ARNTL in GC cells via the GPX4/SQSTM1 axis, thereby inhibiting GC cell function and providing a new molecular mechanism and therapeutic target for the development of GC.
Methods
Cell lines
Gastric cancer cell lines Hgc27, MKN45, AGS and human gastric mucosal cells GES1 and MC (as the non cancer control group) were purchased from the National Collection of Authenticated Cell Cultures (ShaanXi, China). AGS was cultured in F-12 K medium (Boster) with 10% fetal bovine serum (Boster) , other cells were cultured in DMEM medium (Boster) with 10% fetal bovine serum (Boster) ,all cells under the condition of 5% CO and 37 °C.
Vectors and transfection
Cells were transfected 1 h before irradiation using lipofectamine 3000 (Thermo Fisher) according to manufacturer’s protocol. Lipofectamine 3000 reagent (Invitrogen) was used for MT1G overexpression plasmid transfection transient transfection. For MT1G overexpression, cells were infected by using the overexpression plasmid transfection homo-MT1G(NM_001301267.2)_pcDNA3.1( +)-EGFP(1) (HyCyte). The transfection efficiency was determined by quantitative real-time PCR.
RT-qPCR assay
TRIzol® Plus RNA Purification Kit (Invitrogen, Carlsbad, CA, USA) was used to perform total RNA extraction. First-strand cDNA was synthesized using SuperScript™ III First-strand Synthesis SuperMix for RT-qPCR (Invitrogen, Carlsbad, CA, USA). cDNA was then synthesized using Power SYBR® Green PCR Master Mix Kit (Applied Biosystems, USA) for real-time fluorescent quantitative PCR.The following primers were used:
Human GAPDH forward: 5ʹ-CAGGAGGCATTGCTGATGAT-3ʹ;
Human GAPDH reverse: 5ʹ-GAAGGCTGGGGCTCATTT-3ʹ;
Human MT1G forward: 5ʹ-GCAAGUGCAAAGAGUGCAATT-3ʹ;
Human MT1G reverse: 5ʹ-UUGCACUCUUUGCACUUGCTT-3ʹ.
Cell counting kit-8 (CCK-8) assay for cell proliferation
Cells were seeded into 96-well plates at 5,000 cells/well. Thereafter, add different concentrations of small molecule compounds (RSL3, Fer-1 and DFMO) to cells for incubation, and at 0, 24, 48 and 72 h, 10 μL CCK-8 solution (Dojindo, Kyushu Island, Japan) and 100 μL fresh medium were supplemented into each well. After 1 h of incubation at 37 °C, the optical density (OD) value at 450 nm was measured using a microplate reader (Bio-Rad 680, Bio-Rad). The average OD values from each group were used to reflect the cell proliferation.
Migration and invasion
Migration and invasion were assayed in 6-well transwell plates with 8.0 μm pore size. After adding matrix gel to the upper chamber of transwell for 12 h, cell suspension was added for cell invasion experiments. After 24 h of migration or invasion in the medium (Boster) with 10% fetal bovine serum, cells at the bottom of the insert were fixed with 4% paraformaldehyde (PFA) for 10 min, washed three with PBS, and stained with 0.5% crystal violet for 0.5 h. Remaining cells at the top of the insert were removed with a cotton swab. Pictures were taken at microscope and quantified using Image J software.
Scratch wound healing assay
The number of normal cultured cells in 6-well plates was ≥ 90% and reached confluence. Use a sterile pipette tip to produce a clean scratch in the center of the cell layer. Photographs were taken under a microscope after 0 h, 12 h and 24 h and cell migration distance was estimated by Image J software.
Protein blotting assay
The expression level of ferroptosis key protein of gastric cancer cell line Hgc27, MKN45 and AGS was confirmed by protein blotting. Cells were treated with RIPA lysis buffer (Fdbio: #FD009) containing phosphatase inhibitors and protease inhibitors. Protein concentration was detected using BCA Protein Assay Kit (Boster: AR0146). Equivalent proteins were then separated by 10% Tris-Tricine SDS-PAGE and transferred to polyvinylidene difluoride (PVDF) membranes. After 2 h of closure with skimmed milk in TBST, the membranes were analyzed using SQSTM1 (Immununoway: YT7058), LC3B (Immununoway: YN5524), EGLN2 (Boster: BM3977), GAPDH (Boster: BA2913), GPX4 (Boster: BM5231), FSP1 (Proteintech: #20,886–1-AP), DHODH (Boster: M04035-1) and antibodies to SLC7A11 (Boster: BM5318) were incubated overnight at 4 °C. The membranes were subsequently washed with Tris-buffered saline containing Tween and then incubated with the corresponding coupled anti-rabbit (anti-mouse) antibodies for 1 h at 37 °C. Finally, the bands on the membrane were visualized using a ChemiDoc™ imaging system.
Data collection and processing
RNA-seq data of 33 tumor items STAR process were downloaded and organized from TCGA database (Table S1) and extracted in TPM format. Data processing and visualization were performed using the (ggplot2 [3.3.6], stats [4.2.1], car [3.1–0]) R package.
Measurement of MDA, SOD and GSH levels
Malondialdehyde (MDA), superoxide dismutase (SOD) and Reduced glutathione (GSH) assay kits were purchased from Nanjing Jiancheng Biotechnology Institute (Nanjing, China). Cell samples were assayed for MDA, SOD and GSH levels using the kits according to the instructions. The samples were treated with RIPA lysis buffer containing phosphatase inhibitors and protease inhibitors for 30 min, and then centrifuged at 12,000 r/min for 10 min to produce supernatants for the detection of MDA, SOD and GSH levels.
Co-IP (Co-immunoprecipitation) assay
For Co-IP assays, cells were collected and washed three times with PBS. Total protein was extracted using RIPA lysis buffer. The lysates were incubated with primary antibodies anti-GPX4 (Boster, 1:100), anti-SQSTM1 (Immununoway, 1:200) and anti-ARNTL (Proteintech, 1:200) at room temperature for 3 h. IgG of the same species as the endogenous antibody was used as a negative control. Followed by overnight incubation with protein A/G PLUS-Agarose beads at 4 °C. Beads were washed three times with PBS, resuspended in SDS-PAGE loading buffer, and subjected to Western blot analysis. For sample loading, whole cell lysates were used as Input group, the positive control group.
Mitochondrial membrane potential assay
The mitochondrial membrane potential assay kit (Beyoime:C2006) was purchased from Shanghai Beyoime Biotechnology Co. Ltd (Shanghai, China), and the mitochondrial membrane potential was determined using JC-1 according to the instructions. Briefly, cells of different groups were stained with JC-1 and observed by fluorescence microscopy. The excitation wavelength of JC-1 monomer was 490 nm, and the emission light was set to 530 nm; the excitation wavelength of JC-1 aggregates was 525 nm, and the emission light was set to 590 nm.
Fe2+ fluorescent reference reagent manufacturer’s manual. FerroOrange (Dojindo, F374) is a new type of fluorescent probe that can perform fluorescence imaging of Fe2+ in living cells. GC cells are seeded in fluorescent culture dishes and incubated overnight in a 37 °C, 5% CO2 incubator. Discard the supernatant and wash the cells three times with HBSS. Transfect the cells in the experimental group and incubate in a 37 °C, 5% CO2 incubator. Add FerroOrange working solution at a concentration of 1 μmol/l and incubate in a 37 °C, 5% CO2 incubator. Observe the cells under a fluorescence microscope after adding the FerroOrange staining agent (Green laser (Excitation wavelength: 532 nm)).
Cellular reactive oxygen species (ROS) assay
Gastric cancer cells were seeded in 6-well culture plates at a density of 1 × 10⁶ cells per well and treated with different conditions for 24 h. Subsequently, the cells were collected and incubated with the fluorescent probe 2,7-dichlorodihydrofluorescein diacetate (DCFH-DA, Beyoime:S0033). To prevent interference from fluorescent markers in the experiment, no fluorescent label was added during plasmid construction. Therefore, there is no interference from other fluorescences in the experiment. After incubating for 30 min at 37 °C in the dark, the cells were washed three times with serum-free medium and detected using the Cytoflex flow cytometer (Beckman, USA). According to the excitation/emission wavelengths (488 nm/522 nm) of the DCFH-DA probes in the manufacturer’s instructions, appropriate channels were selected to measure the fluorescence values, and the data were analyzed using Flowjo V10 software (FlowJo LLC, USA). During flow detection, the acquisition voltage, rate, and gating settings were consistent for each group of cells.
Transmission electron microscope (TEM)
Cells were fixed in 2% glutaraldehyde overnight at 4 °C and fixed with 1% osmium tetroxide (OsO4) for 1 h. Epon-812 pure embedding agent embedding, Use an ultra-thin slicer to make ultra-thin slices of 60–90 nm and transfer them onto a copper mesh. First, stain with uranium acetate for 10–15 min, then stain with lead citrate for 1–2 min, and stain at room temperature. The JEM-1400FLASH transmission electron microscope produced by Japan Electronics Corporation (JEOL) was used to capture images of copper mesh. Each copper mesh was observed at low magnification for all tissues, and images were collected from the selected areas to observe specific lesions.
Statistical analysis
Statistical analyses were performed using GraphPad Prism 8 and SPSS 26.0. Protein fluorescence intensity was quantified using ImageJ software. Data are presented as mean ± SD. One-way ANOVA was used for comparisons among multiple groups, while the Student’s t-test was used for two-group comparisons. Kaplan–Meier survival curves were generated, and multivariate survival analysis was conducted using the Cox proportional hazards model. A p-value of < 0.05 was considered statistically significant.
Data availability
The data sets used to support the findings of this study are available from the corresponding author on reasonable request.
Change history
24 January 2025
A Correction to this paper has been published: https://doi.org/10.1038/s41598-025-86444-7
Abbreviations
- GC:
-
Gastric cancer
- MT1G:
-
Metallothionein-1G
- GPX4:
-
Phospholipid hydroperoxide glutathione peroxidase
- SLC7A11:
-
Cystine/glutamate transporter
- FSP1:
-
Ferroptosis suppressor protein 1
- DHODH:
-
Dihydroorotate dehydrogenase (quinone)
- SQSTM1:
-
Sequestosome-1
- LC3B:
-
Recombinant Human Microtubule-Associated Protein 1 Light Chain 3 Beta
- ARNTL:
-
Basic helix-loop-helix ARNT-like protein 1
- EGLN2:
-
Egl nine homolog 2
- RSL3:
-
(1S,3R)-RSL3
- Fer-1:
-
Ferrostatin-1
- DFOM:
-
Deferoxamine mesylate
- MDA:
-
Malondialdehyde
- SOD:
-
Superoxide dismutase
- GSH:
-
Reduced glutathione
- Co-IP:
-
Co-immunoprecipitation
- ROS:
-
Reactive oxygen species
- RT‒qPCR:
-
Real-time quantitative polymerase chain reaction
- TEM:
-
Transmission electron microscope
References
Smyth, E. C., Nilsson, M., Grabsch, H. I., van Grieken, N. C. & Lordick, F. Gastric cancer. Lancet (London, England) 396, 635–648. https://doi.org/10.1016/s0140-6736(20)31288-5 (2020).
Ajani, J. A. et al. Gastric cancer, version 2.2022, NCCN clinical practice guidelines in oncology. J. Natl. Compr. Cancer Netw. JNCCN 20, 167–192. https://doi.org/10.6004/jnccn.2022.0008 (2022).
López, M. J. et al. Characteristics of gastric cancer around the world. Crit. Rev. Oncol. Hematol. 181, 103841. https://doi.org/10.1016/j.critrevonc.2022.103841 (2023).
Lordick, F. et al. Gastric cancer: ESMO clinical practice guideline for diagnosis, treatment and follow-up. Ann. Oncol. Off. J. Eur. Soc. Med. Oncol. 33, 1005–1020. https://doi.org/10.1016/j.annonc.2022.07.004 (2022).
Thrift, A. P. & El-Serag, H. B. Burden of Gastric Cancer. Clin. Gastroenterol. Hepatol. Off. Clin. Pract. J. Am. Gastroenterol. Assoc. 18, 534–542. https://doi.org/10.1016/j.cgh.2019.07.045 (2020).
Yan, L. et al. Effect of helicobacter pylori eradication on gastric cancer prevention: updated report from a randomized controlled trial with 26.5 years of follow-up. Gastroenterology 163, 154–162. https://doi.org/10.1053/j.gastro.2022.03.039 (2022).
Zeng, Y. & Jin, R. U. Molecular pathogenesis, targeted therapies, and future perspectives for gastric cancer. Semin. Cancer Biol. 86, 566–582. https://doi.org/10.1016/j.semcancer.2021.12.004 (2022).
Wang, Y. et al. MT1G serves as a tumor suppressor in hepatocellular carcinoma by interacting with p53. Oncogenesis 8, 67. https://doi.org/10.1038/s41389-019-0176-5 (2019).
Chen, B. et al. Cellular zinc metabolism and zinc signaling: from biological functions to diseases and therapeutic targets. Signal Transduct. Target. Ther. 9, 6. https://doi.org/10.1038/s41392-023-01679-y (2024).
Xue, Q. et al. Copper metabolism in cell death and autophagy. Autophagy 19, 2175–2195. https://doi.org/10.1080/15548627.2023.2200554 (2023).
Si, M. & Lang, J. The roles of metallothioneins in carcinogenesis. J. Hematol. Oncol. 11, 107. https://doi.org/10.1186/s13045-018-0645-x (2018).
Wei, T. et al. Epigenetic regulation of the DNMT1/MT1G/KLF4/CA9 axis synergises the anticancer effects of sorafenib in hepatocellular carcinoma. Pharmacol. Res. 180, 106244. https://doi.org/10.1016/j.phrs.2022.106244 (2022).
Wang, S. et al. MT1G induces lipid droplet accumulation through modulation of H3K14 trimethylation accelerating clear cell renal cell carcinoma progression. Br. J. Cancer https://doi.org/10.1038/s41416-024-02747-y (2024).
Tang, D., Chen, X., Kang, R. & Kroemer, G. Ferroptosis: molecular mechanisms and health implications. Cell Res. 31, 107–125. https://doi.org/10.1038/s41422-020-00441-1 (2021).
Stockwell, B. R. et al. Ferroptosis: a regulated cell death nexus linking metabolism, redox biology, and disease. Cell 171, 273–285. https://doi.org/10.1016/j.cell.2017.09.021 (2017).
Liu, J. et al. TMEM164 is a new determinant of autophagy-dependent ferroptosis. Autophagy 19, 945–956. https://doi.org/10.1080/15548627.2022.2111635 (2023).
Yang, M. et al. Clockophagy is a novel selective autophagy process favoring ferroptosis. Sci. Adv. 5, eaaw2238. https://doi.org/10.1126/sciadv.aaw2238 (2019).
Liu, Y., Wang, Y., Liu, J., Kang, R. & Tang, D. Interplay between MTOR and GPX4 signaling modulates autophagy-dependent ferroptotic cancer cell death. Cancer gene therapy 28, 55–63. https://doi.org/10.1038/s41417-020-0182-y (2021).
Liu, Y. et al. Signaling pathways of oxidative stress response: the potential therapeutic targets in gastric cancer. Front. Immunol. 14, 1139589. https://doi.org/10.3389/fimmu.2023.1139589 (2023).
Yang, Z. et al. ACTL6A protects gastric cancer cells against ferroptosis through induction of glutathione synthesis. Nat. Commun. 14, 4193. https://doi.org/10.1038/s41467-023-39901-8 (2023).
Chaturvedi, R. et al. Spermine oxidase mediates the gastric cancer risk associated with Helicobacter pylori CagA. Gastroenterology 141, 1696–1708. https://doi.org/10.1053/j.gastro.2011.07.045 (2011).
Della Bella, C. et al. The helicobacter pylori CagY protein drives gastric Th1 and Th17 inflammation and B cell proliferation in gastric MALT lymphoma. Int. J. Mol. Sci. 22, 9459. https://doi.org/10.3390/ijms22179459 (2021).
González, A. et al. Repurposing dihydropyridines for treatment of helicobacter pylori infection. Pharmaceutics 11, 681. https://doi.org/10.3390/pharmaceutics11120681 (2019).
Rao, S. V. et al. Gastrin activates autophagy and increases migration and survival of gastric adenocarcinoma cells. BMC cancer 17, 68. https://doi.org/10.1186/s12885-017-3055-5 (2017).
Ji, X. K. et al. Genetic variant of cyclooxygenase-2 in gastric cancer: more inflammation and susceptibility. World J. Gastroenterol. 27, 4653–4666. https://doi.org/10.3748/wjg.v27.i28.4653 (2021).
Janssen, A. M. et al. Superoxide dismutases in gastric and esophageal cancer and the prognostic impact in gastric cancer. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 6, 3183–3192 (2000).
Houessinon, A. et al. Metallothionein-1 as a biomarker of altered redox metabolism in hepatocellular carcinoma cells exposed to sorafenib. Mol. Cancer 15, 38. https://doi.org/10.1186/s12943-016-0526-2 (2016).
Zhou, Z. et al. Extracellular vesicles activated cancer-associated fibroblasts promote lung cancer metastasis through mitophagy and mtDNA transfer. J. Exp. Clin. Cancer Res. : CR 43, 158. https://doi.org/10.1186/s13046-024-03077-w (2024).
Peng, B. et al. Ferroptosis-related gene MT1G as a novel biomarker correlated with prognosis and immune infiltration in colorectal cancer. Front. Cell Dev. Biol. 10, 881447. https://doi.org/10.3389/fcell.2022.881447 (2022).
Li, K. et al. Metallothionein-1G suppresses pancreatic cancer cell stemness by limiting activin A secretion via NF-κB inhibition. Theranostics 11, 3196–3212. https://doi.org/10.7150/thno.51976 (2021).
Hung, K. C. et al. The expression profile and prognostic significance of metallothionein genes in colorectal cancer. Int. J. Mol. Sci. 20, 3849. https://doi.org/10.3390/ijms20163849 (2019).
Roth, M. J. et al. p16, MGMT, RARbeta2, CLDN3, CRBP and MT1G gene methylation in esophageal squamous cell carcinoma and its precursor lesions. Oncol. Rep. 15, 1591–1597 (2006).
Kumar, A., Chatopadhyay, T., Raziuddin, M. & Ralhan, R. Discovery of deregulation of zinc homeostasis and its associated genes in esophageal squamous cell carcinoma using cDNA microarray. Int. J. Cancer 120, 230–242. https://doi.org/10.1002/ijc.22246 (2007).
Sakamoto, L. H., De Camargo, B., Cajaiba, M., Soares, F. A. & Vettore, A. L. MT1G hypermethylation: a potential prognostic marker for hepatoblastoma. Pediatr. Res. 67, 387–393. https://doi.org/10.1203/PDR.0b013e3181d01863 (2010).
Brazão-Silva, M. T. et al. Metallothionein gene expression is altered in oral cancer and may predict metastasis and patient outcomes. Histopathology 67, 358–367. https://doi.org/10.1111/his.12660 (2015).
Liu, H. et al. Identification and validation of a prognostic signature for prostate cancer based on ferroptosis-related genes. Front. Oncol. 11, 623313. https://doi.org/10.3389/fonc.2021.623313 (2021).
Wang, S. et al. Betulinic acid arrests cell cycle at G2/M phase by up-regulating metallothionein 1G inhibiting proliferation of colon cancer cells. Heliyon 10, e23833. https://doi.org/10.1016/j.heliyon.2023.e23833 (2024).
Cheng, B. et al. MT1G, an emerging ferroptosis-related gene: a novel prognostic biomarker and indicator of immunotherapy sensitivity in prostate cancer. Environ. Toxicol. 39, 927–941. https://doi.org/10.1002/tox.23997 (2024).
Ferrario, C. et al. Metallothionein 1G acts as an oncosupressor in papillary thyroid carcinoma. Lab. Investig. J. Tech. Methods Pathol. 88, 474–481. https://doi.org/10.1038/labinvest.2008.17 (2008).
Huang, Y., de la Chapelle, A. & Pellegata, N. S. Hypermethylation, but not LOH, is associated with the low expression of MT1G and CRABP1 in papillary thyroid carcinoma. Int. J. Cancer 104, 735–744. https://doi.org/10.1002/ijc.11006 (2003).
Fu, J. et al. Metallothionein 1G functions as a tumor suppressor in thyroid cancer through modulating the PI3K/Akt signaling pathway. BMC cancer 13, 462. https://doi.org/10.1186/1471-2407-13-462 (2013).
Chan, K. Y. et al. Positional expression profiling indicates candidate genes in deletion hotspots of hepatocellular carcinoma. Mod. Pathol. Off. J. United States Can. Acad. Pathol. 19, 1546–1554. https://doi.org/10.1038/modpathol.3800674 (2006).
Hirako, N., Nakano, H. & Takahashi, S. A PU.1 suppressive target gene, metallothionein 1G, inhibits retinoic acid-induced NB4 cell differentiation. PloS One 9, e103282. https://doi.org/10.1371/journal.pone.0103282 (2014).
Arriaga, J. M., Bravo, A. I., Mordoh, J. & Bianchini, M. Metallothionein 1G promotes the differentiation of HT-29 human colorectal cancer cells. Oncol. Rep. 37, 2633–2651. https://doi.org/10.3892/or.2017.5547 (2017).
Arriaga, J. M., Greco, A., Mordoh, J. & Bianchini, M. Metallothionein 1G and zinc sensitize human colorectal cancer cells to chemotherapy. Mol. Cancer Therap. 13, 1369–1381. https://doi.org/10.1158/1535-7163.Mct-13-0944 (2014).
Sun, X. et al. Metallothionein-1G facilitates sorafenib resistance through inhibition of ferroptosis. Hepatology (Baltimore, Md.) 64, 488–500. https://doi.org/10.1002/hep.28574 (2016).
Siegel, R. L., Giaquinto, A. N. & Jemal, A. Cancer statistics. CA: Cancer J. Clin. 74, 12–49. https://doi.org/10.3322/caac.21820 (2024).
Li, N. & Sohal, D. Current state of the art: immunotherapy in esophageal cancer and gastroesophageal junction cancer. Cancer Immunol. Immunother. CII 72, 3939–3952. https://doi.org/10.1007/s00262-023-03566-5 (2023).
Larsson, S. C. et al. Smoking, alcohol consumption, and cancer: a mendelian randomisation study in UK Biobank and international genetic consortia participants. PLoS Med. 17, e1003178. https://doi.org/10.1371/journal.pmed.1003178 (2020).
Badran, M. & Laher, I. Waterpipe (shisha, hookah) smoking, oxidative stress and hidden disease potential. Redox Biol. 34, 101455. https://doi.org/10.1016/j.redox.2020.101455 (2020).
Caliri, A. W., Tommasi, S. & Besaratinia, A. Relationships among smoking, oxidative stress, inflammation, macromolecular damage, and cancer. Mutat. Res. Rev. Mutat. Res. 787, 108365. https://doi.org/10.1016/j.mrrev.2021.108365 (2021).
Bhattacharyya, A., Chattopadhyay, R., Mitra, S. & Crowe, S. E. Oxidative stress: an essential factor in the pathogenesis of gastrointestinal mucosal diseases. Physiol. Rev. 94, 329–354. https://doi.org/10.1152/physrev.00040.2012 (2014).
Chen, X. et al. Celastrol induces ROS-mediated apoptosis via directly targeting peroxiredoxin-2 in gastric cancer cells. Theranostics 10, 10290–10308. https://doi.org/10.7150/thno.46728 (2020).
Li, K. et al. Role of oxidative stress-induced ferroptosis in cancer therapy. J. Cell. Mol. Med. 28, e18399. https://doi.org/10.1111/jcmm.18399 (2024).
Panda, S. K. et al. Repression of the aryl-hydrocarbon receptor prevents oxidative stress and ferroptosis of intestinal intraepithelial lymphocytes. Immunity 56, 797-812.e794. https://doi.org/10.1016/j.immuni.2023.01.023 (2023).
Jiang, M. et al. Exosome-mediated miR-144-3p promotes ferroptosis to inhibit osteosarcoma proliferation, migration, and invasion through regulating ZEB1. Mol. Cancer 22, 113. https://doi.org/10.1186/s12943-023-01804-z (2023).
Shao, F. et al. Synergistic ROS generation and directional overloading of endogenous calcium induce mitochondrial dysfunction in living cells. Biomaterials 301, 122284. https://doi.org/10.1016/j.biomaterials.2023.122284 (2023).
Qi, R. et al. Cancer-associated fibroblasts suppress ferroptosis and induce gemcitabine resistance in pancreatic cancer cells by secreting exosome-derived ACSL4-targeting miRNAs. Drug Resist. Updates Rev. Comment. Antimicrob. Anticancer Chemother. 68, 100960. https://doi.org/10.1016/j.drup.2023.100960 (2023).
Yu, Q. et al. EGCG attenuated acute myocardial infarction by inhibiting ferroptosis via miR-450b-5p/ACSL4 axis. Phytomed. Int. J. Phytother. Phytopharmacol. 119, 154999. https://doi.org/10.1016/j.phymed.2023.154999 (2023).
Baechler, B. L., Bloemberg, D. & Quadrilatero, J. Mitophagy regulates mitochondrial network signaling, oxidative stress, and apoptosis during myoblast differentiation. Autophagy 15, 1606–1619. https://doi.org/10.1080/15548627.2019.1591672 (2019).
O’Malley, J., Kumar, R., Inigo, J., Yadava, N. & Chandra, D. Mitochondrial Stress Response and Cancer. Trends in cancer 6, 688–701. https://doi.org/10.1016/j.trecan.2020.04.009 (2020).
Partch, C. L., Green, C. B. & Takahashi, J. S. Molecular architecture of the mammalian circadian clock. Trends Cell Biol. 24, 90–99. https://doi.org/10.1016/j.tcb.2013.07.002 (2014).
Liu, J., Yang, M., Kang, R., Klionsky, D. J. & Tang, D. Autophagic degradation of the circadian clock regulator promotes ferroptosis. Autophagy 15, 2033–2035. https://doi.org/10.1080/15548627.2019.1659623 (2019).
Appelhoff, R. J. et al. Differential function of the prolyl hydroxylases PHD1, PHD2, and PHD3 in the regulation of hypoxia-inducible factor. J. Biol. Chem. 279, 38458–38465. https://doi.org/10.1074/jbc.M406026200 (2004).
Majmundar, A. J., Wong, W. J. & Simon, M. C. Hypoxia-inducible factors and the response to hypoxic stress. Mol. Cell 40, 294–309. https://doi.org/10.1016/j.molcel.2010.09.022 (2010).
Bensaad, K. et al. Fatty acid uptake and lipid storage induced by HIF-1α contribute to cell growth and survival after hypoxia-reoxygenation. Cell Rep. 9, 349–365. https://doi.org/10.1016/j.celrep.2014.08.056 (2014).
Huang, D. et al. HIF-1-mediated suppression of acyl-CoA dehydrogenases and fatty acid oxidation is critical for cancer progression. Cell Rep. 8, 1930–1942. https://doi.org/10.1016/j.celrep.2014.08.028 (2014).
Vargas, J. N. S., Hamasaki, M., Kawabata, T., Youle, R. J. & Yoshimori, T. The mechanisms and roles of selective autophagy in mammals nature reviews. Mol. Cell Biol. 24, 167–185. https://doi.org/10.1038/s41580-022-00542-2 (2023).
Marsh, T. & Debnath, J. Autophagy suppresses breast cancer metastasis by degrading NBR1. Autophagy 16, 1164–1165. https://doi.org/10.1080/15548627.2020.1753001 (2020).
Rabinovich-Nikitin, I. et al. Mitochondrial autophagy and cell survival is regulated by the circadian Clock gene in cardiac myocytes during ischemic stress. Autophagy 17, 3794–3812. https://doi.org/10.1080/15548627.2021.1938913 (2021).
Li, C. et al. Mitochondrial DNA stress triggers autophagy-dependent ferroptotic death. Autophagy 17, 948–960. https://doi.org/10.1080/15548627.2020.1739447 (2021).
Wu, H., Liu, Q., Shan, X., Gao, W. & Chen, Q. ATM orchestrates ferritinophagy and ferroptosis by phosphorylating NCOA4. Autophagy 19, 2062–2077. https://doi.org/10.1080/15548627.2023.2170960 (2023).
Zhu, M. et al. STAT3 signaling promotes cardiac injury by upregulating NCOA4-mediated ferritinophagy and ferroptosis in high-fat-diet fed mice. Free Radic. Biol. Med. 201, 111–125. https://doi.org/10.1016/j.freeradbiomed.2023.03.003 (2023).
Mancias, J. D., Wang, X., Gygi, S. P., Harper, J. W. & Kimmelman, A. C. Quantitative proteomics identifies NCOA4 as the cargo receptor mediating ferritinophagy. Nature 509, 105–109. https://doi.org/10.1038/nature13148 (2014).
He, J. et al. Ferroptosis and ferritinophagy in diabetes complications. Mol. Metab. 60, 101470. https://doi.org/10.1016/j.molmet.2022.101470 (2022).
Li, N. et al. Ferritinophagy-mediated ferroptosis is involved in sepsis-induced cardiac injury. Free Radic. Biol. Med. 160, 303–318. https://doi.org/10.1016/j.freeradbiomed.2020.08.009 (2020).
Sun, K. et al. JNK-JUN-NCOA4 axis contributes to chondrocyte ferroptosis and aggravates osteoarthritis via ferritinophagy. Free Radic. Biol. Med. 200, 87–101. https://doi.org/10.1016/j.freeradbiomed.2023.03.008 (2023).
Li, C. et al. Nuclear receptor coactivator 4-mediated ferritinophagy contributes to cerebral ischemia-induced ferroptosis in ischemic stroke. Pharmacol. Res. 174, 105933. https://doi.org/10.1016/j.phrs.2021.105933 (2021).
Jin, L. et al. STING promotes ferroptosis through NCOA4-dependent ferritinophagy in acute kidney injury. Free Radic. Biol. Med. 208, 348–360. https://doi.org/10.1016/j.freeradbiomed.2023.08.025 (2023).
Liu, X. et al. Ferritinophagy and ferroptosis in cerebral ischemia reperfusion injury. Neurochem. Res. 49, 1965–1979. https://doi.org/10.1007/s11064-024-04161-5 (2024).
Tian, Y. et al. FTH1 Inhibits Ferroptosis Through Ferritinophagy in the 6-OHDA Model of Parkinson’s Disease. Neurotherap. J. Am. Soc. Exp. NeuroTherap. 17, 1796–1812. https://doi.org/10.1007/s13311-020-00929-z (2020).
Santana-Codina, N. et al. NCOA4-mediated ferritinophagy is a pancreatic cancer dependency via maintenance of iron bioavailability for iron-sulfur cluster proteins. Cancer Discov. 12, 2180–2197. https://doi.org/10.1158/2159-8290.Cd-22-0043 (2022).
Li, K. et al. TRIM7 modulates NCOA4-mediated ferritinophagy and ferroptosis in glioblastoma cells. Redox Biol. 56, 102451. https://doi.org/10.1016/j.redox.2022.102451 (2022).
Xiu, Z. et al. Caryophyllene oxide induces ferritinophagy by regulating the NCOA4/FTH1/LC3 pathway in hepatocellular carcinoma. Front. Pharmacol. 13, 930958. https://doi.org/10.3389/fphar.2022.930958 (2022).
Zhang, J. et al. YAP1 alleviates sepsis-induced acute lung injury via inhibiting ferritinophagy-mediated ferroptosis. Front. Immunol. 13, 884362. https://doi.org/10.3389/fimmu.2022.884362 (2022).
Acknowledgements
We would like to thank the Core Facility of Shaanxi University of Traditional Chinese Medicine Medical Research and Experiment Center for its help in the detection of experimental samples. Thanks to Dr. Liang Song and Ms. Dan Chen for their help in the experimental study.
Funding
This study was partially supported by the Qin Chuang Yuan Traditional Chinese Medicine Innovation Research and Development and Transformation Project (2023-QCYZH-038), and the special project of the Graduate Quality Improvement Project of Shaanxi University of Traditional Chinese Medicine (CXSJ202306).
Author information
Authors and Affiliations
Contributions
Kaiqiang Meng: performed experiments, data analysis and drafted the original manuscript. Zhichao Fang and Jiamin Li: collected data and performed western blot analysis. Fan Qi and Liang Song: designed the study. Jian Song: revised the manuscript. All authors approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this Article was revised: The Funding section in the original version of this Article was incorrect. Full information regarding the corrections made can be found in the correction for this Article.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Meng, K., Song, J., Qi, F. et al. MT1G promotes iron autophagy and inhibits the function of gastric cancer cell lines by intervening in GPX4/SQSTM1. Sci Rep 14, 28539 (2024). https://doi.org/10.1038/s41598-024-80160-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-80160-4
Keywords
This article is cited by
-
A bibliometric analysis of trends and hotspots in autophagy in gastric cancer
Discover Oncology (2025)