Abstract
Omics data provide a plethora of quantifiable information that can potentially be used to identify biomarkers targeting the physiological processes and ecological phenomena of organisms. However, omics data have not been fully utilized because current prediction methods in biomarker construction are susceptible to data multidimensionality and noise. We developed OmicSense, a quantitative prediction method that uses a mixture of Gaussian distributions as the probability distribution, yielding the most likely objective variable predicted for each biomarker. Our benchmark test using a transcriptome dataset revealed that OmicSense achieves accurate and robust prediction against background noise without overfitting. Weighted gene co-expression network analysis revealed that OmicSense preferentially utilized hub nodes of the network, indicating the interpretability of the method. Application of OmicSense to single-cell transcriptome, metabolome, and microbiome datasets confirmed high prediction performance (r > 0.8), suggesting applicability to diverse scientific fields. Given the recent rapidly expanding availability of omics data, the developed prediction tool OmicSense, can accelerate the use of omics data as a “biosensor” based on an assemblage of potential biomarkers.
Similar content being viewed by others
Introduction
One of the ways by which organisms adapt to their environment is by regulating their physiology1. Rapid progress in high-throughput technologies has enabled easy collection of various scales of omics data from a wide taxonomic range of organisms, representing a breakthrough in modern biology2. Omics data linked with environmental or physiological metadata can thoroughly depict the evolutionary strategies of organisms and, therefore, be potentially used to predict their metadata status3. Because omics data collection was expensive and time-consuming until the early 2000s, only a few “biomarker” genes and metabolites were identified to economically diagnose or monitor specific disease symptoms4,5. However, remarkable cost reduction achieved in recent times has enabled the use of omics data itself as a “biosensor”—representing an assemblage of potential biomarkers3.
Regression analysis is the most commonly used prediction method applied when multiple explanatory variables are available, such as in omics data; however, this method is often problematic owing to its multidimensionality or the “large p, small n” issue6. To overcome this problem, regression methods have been developed with solutions such as regularization. However, these methods have not been optimized for practical use because omics data often contain considerable noise7,8, including missing and erroneous values arising from stochasticity, genetic and environmental heterogeneity, and technical shortcomings in measurements, which cloud the true relationship between biological phenomena and omics data.
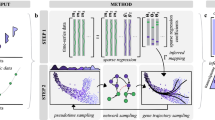
Herein, we introduce OmicSense—a quantitative prediction method that processes multidimensional omics data to produce highly accurate and robust predictions based on an ensemble learning-like framework (Fig. 1). OmicSense constructs a library of simple regression models between the target and each predictor variable and generates a conditional probability distribution of the target from the corresponding predictor variable using new input data from each of the model elements (Fig. 1a). Furthermore, OmicSense generates a mixture of Gaussian distributions by combining the distributions that yield the most likely target value (Fig. 1b). Because predictor variables are naturally weighted by the degree of association with the target value in the mixture distribution, OmicSense can construct a prediction model without any specific parameter tunings. The distribution combining process (Fig. 1c), considered ensemble learning, allows us to overcome the “large p, small n” issue in multidimensional omics data-based prediction. While general ensemble process such as bagging9 combines multiple predicted values (continuous values in regression), OmicSense combines probability distributions for prediction. We have made this prediction method freely available within CRAN and GitHub to promote its use in diverse fields of science. The method can be implemented using the downloadable R package “OmicSense”.
Flowchart of the prediction process using OmicSense. (a) The OmicSense prediction model is based on the model elements reflecting the relationship between each predictor variable (x) and the target values (y). (b) The values of x in new sample data are transformed into probability distributions of the y values using the model element library. (c) The most likely y value (y*) is obtained by combining the distributions.
Methods
Algorithms used in OmicSense
OmicSense is an approach that predicts quantitative target values using multidimensional omics data associated with it, which includes several predictor variables. The method computes a probability distribution, which follows a normal distribution to simplify calculations, to determine the most probable target value, as determined using the following equations:
where xiis the ith predictor value of the m predictors used for model construction; k is the order of the regression model and is expressed as f(xi);β0i, βki(k = 1, 2, 3), and eiis the intercept, coefficient, and error terms of the regression model of y on xi, respectively; and σiis the standard deviation of ei. The parameters β0i, βki(k = 1, 2, 3), and σiwere calculated from xiobserved in the data to generate the model element for xi. Using these equations, a conditional probability distribution of y, given xi, was obtained for each i. Homoscedasticity of σiwas assumed for each conditional probability distribution. By compiling the probability distributions obtained from m (or optionally selected) predictor variables, a mixture of Gaussian distributions that yields the most likely y* value was obtained. This works like ensemble process of the result y based on the framework of probability. By choosing the order k, regression models using a polynomial function (e.g., quadratic and cubic with k = 2 and 3, respectively), in addition to linear regression (k = 1), can be flexibly assumed for target–predictor relationships.
Transcriptome dataset preparation
Transcriptome data were obtained by conducting RNA sequencing of the roots of Pinus thunbergii seedlings. The plant materials were commercially available, and no specific permission was required for the data collection. All methods using the plant material were performed in accordance with the relevant guidelines and regulations. Seedlings were incubated at five temperatures (8, 13, 18, 23, and 28 °C; 12 seedlings/temperature) for 24 h, followed by RNA extraction. Six of the 12 seedlings from each treatment were used for constructing the prediction model (training data), and the remaining were used for the validation of the constructed model (validation data). Thus, the training and validation data were independent of each other.
Simulation test
The prediction model was constructed by randomly selecting 20 transcriptome samples from the training data containing either 30 or 300 genes. The prediction error was measured using the mean absolute error (MAE) by applying the prediction model to the validation data. To evaluate the overfitting of the constructed model to the training data, the increase in the MAE value when the model was applied to the validation data (∆MAEoverfit) was calculated as follows:
where y* is the predicted value of the observed y, r is the number of samples in the validation data, and MAEtrain is the MAE value obtained when the prediction model was applied to the training data for validation. To evaluate the robustness of the prediction against noise in the predictor variables, 10% of the total predictors in the training data was replaced by random or missing values, and the increase in the MAE value when the artificial noise (∆MAEnoise) was added to the training data was calculated. The simulation test was replicated 100 times for each simulation setting. All statistical analyses were performed using R (version 4.0.5)10.
Prediction model comparison
We compared MAE, ∆MAEoverfit, and ∆MAEnoiseamong the models based on OmicSense and six commonly used prediction methods: regressions using L1 and L2 regularization (lasso and ridge regressions)11,12, principal component and partial least squares regression (PCR and PLSR)13, and support vector and random forest regression (SVR and RFR)14,15. For OmicSense, two types of regression models were used to establish the target–predictor relationship: a linear function (referred to as OmicSense1) and a cubic function (referred to as OmicSense3). The lasso and ridge regressions were tuned using the optimum hyperparameter lambda, which was selected from a 10-fold cross-validation and implemented via the “cv.glmnet” function of the glmnet package in R16. Using the “pcr” and “plsr” functions of the pls package in R, the number of principal components used in the PCR and PLSR models, respectively, was selected as the minimum value within the range of standard deviation from the minimum MAE value of the prediction values selected via the 10-fold cross-validation17. The hyperparameter gamma of the radial basis function kernel for the SVR and RFR models was also selected from the 10-fold cross-validation, using the “ksvm” function of the kernlab package18and the train function with the RF option implemented in the caret19 and random forest packages in R. Prediction models were constructed using the same 30 or 300 variables for all the methods and hyperparameters were optimally tuned for each model.
Contribution of genes in constructing prediction models using OmicSense and other models
We compared the gene profiles selected for constructing the prediction models using OmicSense and other methods. To this end, we performed gene co-expression network analysis using the WGCNA package in R20. To construct the network, we selected the minimum soft-thresholding power (β) that achieved scale-free fit R2 > 0.8 (β = 6). Next, we tested the correlation between the sixth root of degree centrality, which is the number of connections with other genes in the network, and the contribution to each prediction model for all genes. The contribution of genes in constructing the prediction models was represented by R2 values and the absolute value of regression coefficients for each method.
Gene Ontology (GO) enrichment analysis
GO terms were assigned to each gene using InterProScan21, which was implemented in the Blast2GO software22. We conducted GO enrichment analysis for each module in the gene co-expression network using the “goseq” function of the GOseq package in R.
Application of OmicSense for analyzing various types of omics data
We used OmicSense to analyze single-cell transcriptome, metabolome, and microbiome datasets linked with quantitative environmental or physiological parameters. The single-cell transcriptome dataset for neurons was obtained from the UBC Research Data Library (https://doi.org/10.14288/1.0347326), and the transcriptome data were extracted for action potential amplitude records (n = 31). The metabolome dataset for Drosophila melanogaster, which included records for hours after egg laying (n= 42), was retrieved from the online repository of a previous study23. For the microbiome dataset, we extracted total bacterial DNA from 49 soil samples collected from seven regions in Fukushima Prefecture, Japan, using the Extrap Soil DNA Kit (BioDynamics Laboratory Inc., Tokyo, Japan), following the manufacturer’s protocol. We amplified the V4 region of 16 S rRNA using the 515f–806r primer pair and performed 251-bp paired-end sequencing using the MiSeq system (Illumina, San Diego, CA, USA). The obtained 13,434,033 paired-end reads were cleaned and merged using DADA224, and a read-count matrix was generated using QIIME225. The merged sequences were labeled with bacterial taxonomic classifications using the q2-feature-classifier with Silva 138 and 99% operational taxonomic units from the 515 F/806R region of the sequences in QIIME2. The soil pH was measured for each sample using a pH meter (Orion Star A211; Thermo Fisher Scientific, Waltham, MA, USA).
Results and discussion
Prediction performance of OmicSense
We benchmarked the prediction performance of OmicSense using a temperature-associated transcriptome dataset (n = 30 each for model training and validation) against that of commonly used prediction methods, namely lasso and ridge, principal component, partial least squares, support vector, and random forest regression. We prepared two prediction models for OmicSense using linear and cubic regression functions to describe target–predictor relationships: OmicSense1 and OmicSense3, respectively. MAE, overfitting to the training dataset, and robustness against noise were compared among the eight models in simulation tests. Overfitting and robustness were measured using the increase in the MAE when the validation dataset was subjected to the prediction model (∆MAEoverfit) and added artificial noise (∆MAEnoise), respectively. These tests were conducted using 30 and 300 predictor variables.
Among the models with datasets of both sizes, simulation tests revealed that OmicSense3 yielded predictions with the smallest MAE values, followed by OmicSense1 (Fig. 2). These OmicSense predictions stably yielded a small ∆MAEoverfit value irrespective of the dataset size, suggesting that OmicSense is free from overfitting issues in its prediction. Compared with the other models, both OmicSense models produced the smallest ∆MAEnoise values for datasets of both sizes. Therefore, among the models evaluated, OmicSense was the most accurate quantitative prediction method using multidimensional omics data. In addition, OmicSense3 outperformed OmicSense1 throughout the tests, implying that a cubic function could flexibly capture target–predictor relationships. By contrast, one drawback of OmicSense is that it currently requires more processing time for prediction than the other methods (Supplementary Fig. S1) because the processing time of OmicSense depends on the number of predictor variables (Supplementary Fig. S2a). However, unlike other methods, OmicSense can achieve accurate prediction with fewer predictor variables (Supplementary Fig. S2b). This should be advantageous in situations where accuracy and robustness are prioritized more strongly than the speed of prediction, such as in medical diagnoses.
Performance of prediction models based on OmicSense and six commonly used methods evaluated using a transcriptome dataset. The upper two graphs show the prediction errors (mean absolute error; MAE) of the models. The middle and bottom figures show the increase in prediction errors upon application of OmicSense to validation data (∆MAE overfit ) and upon the addition of noise (∆MAE noise ), respectively. The left and right figures show the results using 30 and 300 genes in the transcriptomes, respectively. Different colors indicate different prediction methods, and different letters indicate a significant difference between group means tested using the Tukey’s honest significant difference test (P < 0.05).
OmicSense prediction based on biological background
Gene co-expression network analysis helped us identify the co-expressed gene modules in the transcriptomes26. We examined the module structures of predictor variables in the network. OmicSense showed a strong positive correlation between degree centrality in the network and contribution of predictor variables (r > 0.5, P < 0.001), whereas the other models did not (r < 0.4, Fig. 3). This finding indicates that OmicSense can successfully capture omics data structures and preferentially utilize “hub” nodes of the network, which fulfill core functions in gene regulation26. Moreover, OmicSense used hub genes related to heat stress (GO: 0031072, P < 0.05). This is of great benefit in terms of model interpretability, ensuring OmicSense’s high reliability for practical use.
Gene co-expression network of temperature-associated transcriptomes. The colored nodes represent the genes related to response to heat stress (red) and those forming three different modules (purple, green, and sky blue). Gene Ontology (GO) terms commonly enriched in each module are listed, and the terms related to heat stress are indicated in red. The contribution of predictor variables in the models is mapped in the gene co-expression network using colors. The scatter plots show the relationship between the sixth root of degree centrality and the contributions.
Application of OmicSense prediction to other types of omics data
Currently, omics data are being increasingly utilized at various scales in many fields, including food27, healthcare28, and agricultural sciences29. These data are essential in modern biology because of their high quantitative and high-throughput advantages. We further applied OmicSense to single-cell transcriptome30, metabolome24, and microbiome datasets associated with quantitative measurements and found that OmicSense worked well (r > 0.8, P < 0.001) with the indicated types of omics data (Fig. 4). For example, OmicSense achieved better prediction (R2 = 0.88, Fig. 4) using the single-cell transcriptome dataset than that originally reported using a regularized regression method (R2= 0.63)30. We found that Lasso outperformed for metabolome dataset, in which the targeted morphotype could be governed by a few metabolites. Because Lasso model is constructed using a few features, Lasso might result in the best fit.
Application of OmicSense to various types of omics data. (A) Prediction errors (mean absolute error; MAE) in OmicSense compared with those in other commonly used methods for action potential amplitude, age, and pH associated with single-cell transcriptome (n = 31), metabolome (n = 42), and microbiome (n = 49) data structures, respectively. Different colors indicate different prediction methods, and different letters indicate significant differences between group means tested using the Tukey’s honest significant difference test (P < 0.05). (B) Correlation between the observed and predicted values in the prediction using OmicSense. For all cases, models were constructed using two-thirds of the available data, and prediction performance was validated using the remaining data.
Given that biomarker techniques are currently being applied in various fields10,11,12and are expected to be implemented further for practical uses, including preventive and personalized healthcare31, smart farming in agriculture32, and ecosystem engineering33, OmicSense represents a solution to various global issues by maximizing the potential prediction power of omics data with excellent accuracy, robustness, and interpretability.
General workflow of the R package OmicSense
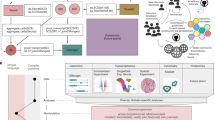
The OmicSense algorithm can be implemented on the R platform using the OmicSense package. This package provides basic functions that help identify strong target–predictor relationships from omics datasets, resulting in accurate, robust, and stable prediction using newly obtained data (Fig. 5).
Data cleaning
The construction of a prediction model using an omics data matrix is obstructed by missing or erroneous predictor values. To address this issue, the “os.clean” function removes such variables from the matrix and facilitates the identification of candidate predictor variables for the construction of OmicSense. This results in a quality-controlled omics data matrix that is essential for better prediction performance using OmicSense.
Data visualization
The performance of a prediction model is dependent on the significance and number of predictor variables used in the model. The “os.rank” and “os.pca” functions are provided for data visualization using rank-order and principal component analysis plots, respectively, to capture the entire picture of omics data linked with such information. These functions use a certain regression method for determining target–predictor relationships and calculate the R2 value in the regression for each predictor variable; the output can be used as a reference to determine the number (or lower the threshold for R2) of predictor variables to be used for model construction.
Data preparation
The OmicSense algorithm constructs a prediction model using (a specified number of) predictor variables in descending order of the R2 value. The function “os.sort” provides matrix-sorted predictor variables for OmicSense modeling, resulting in an output matrix that can be used in subsequent analyses.
Modeling and prediction
The function “os.pred” constructs a prediction model based on the OmicSense algorithm using an omics dataset prepared by users, applies it to newly provided omics data, and predicts a quantitative status of interest. This function allows users to specify the regression method assumed in the target–predictor relationships as well as the number of predictor variables to be used for model construction.
Data Availability
The R package OmicSense is freely available at https://CRAN.R-project.org/package=OmicSense and https://github.com/takakoizumi/OmicSense. RNA sequencing data were deposited in the DDBJ Sequence Read Archive under the accession number DRA015920 (https://ddbj.nig.ac.jp/resource/bioproject/PRJDB15408).
References
Yablonovitch, A. L. et al. Regulation of gene expression and RNA editing in Drosophila adapting to divergent microclimates. Nat. Commun. 8, 1570 (2017).
Nathan, R. et al. Big-data approaches lead to an increased understanding of the ecology of animal movement. Science 375, eabg1780 (2022).
Kurokawa, M. et al. Metagenomic thermometer. bioRxiv (2022).
Vargas, A. J. & Harris, C. C. Biomarker development in the precision medicine era: lung cancer as a case study. Nat. Rev. Cancer. 16, 525–537 (2016).
Varma, V. R. et al. Brain and blood metabolite signatures of pathology and progression in Alzheimer disease: a targeted metabolomics study. PLOS Med. 15, e1002482 (2018).
Bellman, R. Dynamic programming treatment of the travelling salesman problem. J. ACM. 9, 61–63 (1962).
Van Bakel, H., Nislow, C., Blencowe, B. J. & Hughes, T. R. Most dark matter transcripts are associated with known genes. PLOS Biol. 8, e1000371 (2010).
Djebali, S. et al. Landscape of transcription in human cells. Nature 489, 101–108 (2012).
Breiman, L. Bagging predictors. Mach. Learn. 24, 123–140 (1996).
R Core Team. R: A language and environment for statistical computing. (2020). http://www.r-project.org/index.html (R Foundation for Statistical Computing, Vienna, Austria.
Tibshirani, R. Regression shrinkage and selection via the lasso. J. R Stat. Soc. B (Methodol). 58, 267–288 (1996).
Hoerl, A. E. & Kennard, R. W. Ridge regression: biased estimation for nonorthogonal problems. Technometrics 12, 55–67 (1970).
Wold, S., Esbensen, K. & Geladi, P. Principal component analysis. Chemom Intell. Lab. Syst. 2, 37–52 (1987).
Drucker, H., Burges, C. J., Kaufman, L., Smola, A. & Vapnik, V. Support vector regression machines. Adv. Neural Inf Proceeding Syst. 28, 779–784 (1997).
Liaw, A. & Wiener, M. C. Classification and regression by random forest. R News. 2, 18–22 (2002).
Friedman, J., Hastie, T. & Tibshirani, R. Regularization paths for generalized linear models via coordinate descent. J. Stat. Softw. 33, 1–22 (2010).
Mevik, B. H. & Wehrens, R. The pls package: principal component and partial least squares regression in R. J. Stat. Soft. 18, 1–23 (2007).
Karatzoglou, A., Smola, A., Hornik, K. & Zeileis, A. Kernlab – an S4 package for Kernel methods in R. J. Stat. Soft. 11, 1–20 (2004).
Kuhn, M. Building predictive models in R using the caret package. J. Stat. Soft. 28, 1–26 (2008).
Langfelder, P. & Horvath, S. W. G. C. N. A. An R package for weighted correlation network analysis. BMC Bioinform. 9, 559 (2008).
Jones, P. et al. InterProScan 5: genome-scale protein function classification. Bioinformatics 30, 1236–1240 (2014).
Conesa, A. et al. Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21, 3674–3676 (2005).
Avanesov, A. S. et al. Age- and diet-associated metabolome remodeling characterizes the aging process driven by damage accumulation. eLife 3, e02077 (2014).
Callahan, B. J. et al. DADA2: high-resolution sample inference from Illumina amplicon data. Nat. Methods. 13, 581–583 (2016).
Bolyen, E. et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 37, 852–857 (2019).
Ichihashi, Y. et al. Transcriptomic and metabolomic reprogramming from roots to haustoria in the parasitic plant, Thesium chinense. Plant. Cell. Physiol. 59, 724–733 (2018).
Erban, A. et al. Discovery of food identity markers by metabolomics and machine learning technology. Sci. Rep. 9, 9697 (2019).
Shilo, S., Rossman, H. & Segal, E. Axes of a revolution: challenges and promises of big data in healthcare. Nat. Med. 26, 29–38 (2020).
Rohr, A. D. et al. Identification and validation of early genetic biomarkers for apple replant disease. PLOS ONE. 15, e0238876 (2020).
Tripathy, S. J. et al. Transcriptomic correlates of neuron electrophysiological diversity. PLOS Comput. Biol. 13, e1005814 (2017).
Sommer, F., Anderson, J. M., Bharti, R., Raes, J. & Rosenstiel, P. The resilience of the intestinal microbiota influences health and disease. Nat. Rev. Microbiol. 15, 630–638 (2017).
Singh, B. K., Trivedi, P., Egidi, E. & Macdonald, C. A. Delgado-Baquerizo, M. D. Crop microbiome and sustainable agriculture. Nat. Rev. Microbiol. 18, 601–602 (2020).
Wang, J. et al. Embracing mountain microbiome and ecosystem functions under global change. New. Phytol. 234, 1987–2002 (2022).
Acknowledgements
We thank Atsushi Fukushima for technical support in the publication of the R package and metabolome data manipulation and Naoto Nihei for environmental data collection for microbiome analyses. This work was supported by the Japan Society for Promotion of Science (JSPS) with a Research Fellowship for Young Scientists (grant number 21J00762 to T.K.); JSPS KAKENHI (grant number 20K15557 to T.K.); and the Cabinet Office, Government of Japan, Cross-ministerial Moonshot Agriculture, Forestry and Fisheries Research and Development Program, “Technologies for Smart Bio-industry and Agriculture” (funding agency: Bio-oriented Technology Research Advancement Institution to Y.I.), and RIKEN TRIP Initiative Field-omics.
Author information
Authors and Affiliations
Contributions
T.K. conceived the ideas, designed the methodology, analyzed the data, and led the writing of the manuscript; T.K., I.M., and K.K. collected the data; T.K., K.S., and Y.I. contributed to the R package. All authors contributed critically to the draft and gave final approval for publication.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Koizumi, T., Suzuki, K., Mizuki, I. et al. A quantitative prediction method utilizing whole omics data for biosensing. Sci Rep 15, 1928 (2025). https://doi.org/10.1038/s41598-024-84323-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-84323-1