Abstract
The Mycobacterium avium complex (MAC) is a group of closely related nontuberculous mycobacteria that can cause various diseases in humans. In this study, genome sequencing, comprehensive genomic analysis, and antimicrobial susceptibility testing of 66 MAC clinical isolates from King Chulalongkorn Memorial Hospital, Bangkok, Thailand were carried out. Whole-genome average nucleotide identity (ANI) revealed the MAC species distribution, comprising 54 (81.8%) M. intracellulare, 6 (9.1%) M. avium, 5 (7.6%) M. colombiense, and 1 (1.5%) M. timonense. Phylogenetic analysis revealed a high diversity of M. intracellulare isolates and their evolutionary relationships which could be divided into 2 subspecies: M. intracellulare subsp. intracellulare and M. intracellulare subsp. chimaera. In addition, M. intracellulare subsp. chimaera mostly clustered in the distinct clades separated from M. intracellulare strains originating from other countries. Most MAC isolates were resistant to linezolid and moxifloxacin based on phenotypic antimicrobial susceptibility testing. Mutations within rrl gene associated with clarithromycin resistance were detected in M. intracellulare and M. colombiense. The pan-genome analysis presented clade-specific proteins for M. intracellulare, such as PE and PPE protein families. This study provides valuable insights into the genomic diversity and antimicrobial resistance profiles of MAC isolates circulating in Thailand, which are useful for clinical management, guiding the development of targeted diagnostic, and treatment strategies for MAC infections.
Similar content being viewed by others
Introduction
Mycobacterium avium complex (MAC) is a group of closely related nontuberculous mycobacteria (NTM) that can cause a range of clinical manifestations, including pulmonary disease, lymphadenitis, and disseminated infections, particularly in immunocompromised individuals1. The prevalence and clinical significance of MAC infections have been increasing worldwide, posing a significant public health challenge2. Recently, MAC has been classified into several species including M. avium, M. intracellulare, M. paraintracellulare, M. chimaera, M. colombiense, M. arosiense, M. vulneris, M. marseillense, M. timonense, M. bouchedurhonense, M. mantenii, and M. yongonense3. Lately, M. chimaera and M. yongonense have been reclassified as M. intracellulare subsp. chimaera and M. intracellulare subsp. yongonense, respectively, based on genome analyses4,5. Specific MAC species can exhibit varying degrees of virulence, antimicrobial resistance, and clinical outcomes6,7; among MAC members, M. avium and M. intracellulare are the most clinically significant for causing infections in humans8. In addition, the diversity and epidemiology of MAC species can vary significantly across different geographic regions, highlighting the need for region-specific studies9. Currently, individuals with MAC pulmonary disease typically receive treatment with a combination of macrolide, rifampicin, and ethambutol. In addition, an injectable aminoglycoside may be added for severe infections. Second-line drugs such as clofazimine, moxifloxacin, linezolid, and bedaquiline can be used as alternatives for patients who are intolerant of or whose isolate is resistant to first-line drugs10,11. However, the link between phenotypic antimicrobial susceptibility results for these antimicrobial agents and clinical outcomes in MAC patients remains limited10,12. To address this gap, this study aims to conduct a detailed genomic analysis of MAC from Thailand, which comprised 66 isolates (CU cohort) obtained in this study (King Chulalongkorn Memorial Hospital (KCMH), Bangkok, Thailand) and 23 isolates (KKU cohort) sequenced in a prior study (Srinagarind Hospital, Khon Kaen, Thailand)13. The findings of this study would provide a more comprehensive understanding of MAC species distribution, genomic diversity, phylogenetic relationships, and antimicrobial resistance profiles of MAC isolates from Thailand.
Results
Patient characteristics and clinical outcomes
During the study period from January 2016 to December 2022, 41 patients with MAC infections or colonization were included. The median patient age was 64 years, ranging from 12 to 85 years (interquartile range, IQR 19), with 24.4% being male. Predisposing factors included acquired immunodeficiency (22.0%), structural lung diseases (51.2%), and a combination of both (14.6%). Of the 41 patients, 29.3% (12/41) had NTM pulmonary colonization, 53.7% (22/41) had nontuberculous mycobacterial-pulmonary disease (NTM-PD) according to the 2020 ATS/ERS/ESCMID/IDSA diagnostic criteria10, and 17.1% (7/41) had disseminated infection diagnosed from clinical and confirmed by culture (Supplementary table S1). The treatment regimen for patients with NTM-PD and disseminated disease consisted of a susceptibility-based combination of at least three drugs in most patients (25 of 29, 86.2%). A macrolide-based regimen was used in 93.1% of patients. Ethambutol was included in the treatment regimen for 79.3% of patients, followed by rifampicin (69.0%), fluoroquinolone (34.5%), and amikacin (6.9%).
Treatment outcomes for 29 MAC-infected patients, according to the consensus statement of the Nontuberculous mycobacteria-Network European Trials Group (NTM-NET) for NTM-PD with modifications for disseminated infection14, were cure (10.3%), clinical cure (44.8%), culture conversion (3.4%), recurrence (13.8%), died from other conditions (10.3%), and loss to follow-up (17.2%). The median treatment duration for patients achieving a favorable outcome (cure, clinical cure, or culture conversion) was 14 months with an IQR of 4 months.
M. avium complex identification from clinical isolates
In this study, the 66 clinical isolates (CU cohort) were previously identified by line probe assay (LPA) as 53 (80.3%) M. intracellulare, 6 (9.1%) M. avium, 3 (4.5%) M. chimaera (currently M. intracellulare subsp. chimaera), and 4 (6.1%) MAC without species identification. The genomes of these isolates were classified based on average nucleotide identity (ANI) comparisons against reference genomes of M. avium complex species, using the FastANI algorithm . The FastANI results allow the species identification based on genomic data, which identified four species of MAC including 54 (81.8%) M. intracellulare, 6 (9.1%) M. avium, 5 (7.6%) M. colombiense, and 1 (1.5%) M. timonense.
The comparison between LPA and FastANI results showed that all M. avium isolates had concordant results between the two methods as shown in Table 1. However, some M. intracellulare isolates identified by LPA were reclassified into M. colombiense and M. timonense based on the FastANI method. In addition, four MAC (previously unidentified MAC species) from LPA were finally classified as three M. intracellulare and one M. colombiense.
Phenotypic antimicrobial susceptibility patterns
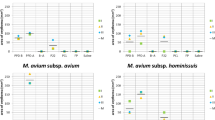
MAC clinical isolates were recovered from various types of clinical specimens, including sputa (n = 31), bronchoalveolar lavage (BAL) fluid (n = 30), endotracheal aspirate (n = 1), synovial fluid (n = 1), and tissue biopsy (n = 3). Most clinical isolates exhibited high resistance rates to linezolid and moxifloxacin with ranges of 60.0–100.0% and 55.3–100.0%, respectively, among three MAC species and their subspecies. In contrast, amikacin and clarithromycin were found to be effective against most MAC clinical isolates. None of the CU MAC clinical isolates was phenotypically resistant to amikacin. Clarithromycin resistance was present in 1 (20%) M. colombiense isolate and 3 (6.4%) M. intracellulare subsp. chimaera isolates (Table 2). For drugs without interpretative breakpoints, rifampicin, TMP/SMX, ciprofloxacin, streptomycin, isoniazid, ethambutol, doxycycline, and ethionamide exhibited high values of the minimal inhibitory concentration (MIC), MIC50, and MIC90 while rifabutin had the lowest values (1 and 2 µg/mL). The MIC50 and MIC90 were comparable among MAC species and their subspecies (Table 2).
Genomic diversity and core-gene based phylogenetic tree
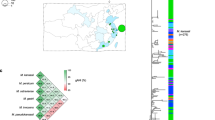
To investigate genomic diversity and phylogenetic relationships among all MAC isolates, comparative core genome analysis was performed using the Panaroo pipeline. The phylogenetic tree based on core genes (Fig. 1) revealed that most of the CU MAC clinical isolates belong to M. intracellulare/M. paraintracellulare (n = 54) and M. avium (n = 6) clades, while only five isolates and one isolate clustered with M. colombiense and M. timonense, respectively. In addition, we also included 23 isolates from KKU cohort13 from the Northeast region of Thailand in the analysis. Of these, 11 KKU isolates clustered with M. intracellulare/M. paraintracellulare, while eight isolates grouped with the M. colombiense cluster. In addition, one isolate was closely related to M. senriense TY59 and three isolates were closely related to M. arosiense DSM 45069 reference. The core-gene tree indicated high genomic diversity of MAC in Thailand. Nevertheless, this analysis may not accurately represent the subspecies of Thai isolates. Therefore, the SNP analysis was carried out to explore more details of their complex diversity.
Phylogenetic tree of Mycobacterium avium complex isolates from Thailand based on core-gene alignment. The core-gene tree, analyzed using the Panaroo pipeline, illustrates the phylogenetic relationships among Thai MAC isolates, including 66 isolates from CU cohort of KCMH, Bangkok, Thailand (pink; Current study) and 23 isolates from KKU cohort of Khon Kaen University, Khon Kaen, northeast of Thailand (orange; previous study13). Reference strains of MAC species (black) are also included, with T indicating type strains. The bootstrap values from 100 replicates are shown as purple circles on the branches.
SNP-based phylogenetic analysis and antimicrobial susceptibility of Thai M. avium complex isolates
According to the controversy of nomenclature and discrepancies of classification results, the genomic identification of subspecies for MAC cannot be directly inferred from the FastANI method and the core-gene tree. For isolates identified as M. avium, all six CU isolates were located within divergent clades within M. avium subsp. hominissuis and these isolates were resistant to linezolid and moxifloxacin (Fig. 2). The SNP-based phylogenetic tree of M. colombiense revealed that five CU isolates and eight KKU isolates monophyletically clustered in the same clade (Fig. 3). This clade included three reference strains: M. colombiense CECT 3035, M. colombiense TKK-01-0051, and M. colombiense 230. All M. colombiense isolates were resistant to moxifloxacin, although only two of KKU isolates were resistant to amikacin. Interestingly, based on the current analysis, M. colombiense can be clustered into three different clades which might be further explored for their origins and phenotypic characteristics. For drug resistance analysis, mutations on 23S rRNA (rrl) gene (including A215G, T288C, C1687T, A2270G, A2270C) associated with clarithromycin resistance were found in four M. colombiense isolates as shown in Table 3, three of which were KKU isolates derived from the same patient.
SNP-based phylogenetic tree of Mycobacterium avium isolates in Thailand. Phylogenetic tree based on SNPs from Snippy pipeline showed the diversity of M. avium isolates in Thailand and their phylogenetic relationship compared to global strains. All M. avium isolates in this study clustered within the M. avium subsp. hominissuis. Drug susceptibility, specimen types, and origins are presented as a color strip labelled beside the tips of the tree.
SNP-based phylogenetic tree of Mycobacterium colombiense isolates in Thailand. Phylogenetic tree based on SNPs from Snippy pipeline showed the diversity of M. colombiense isolates in Thailand and their phylogenetic relationships compared with global reference strains. All M. colombiense isolates in Thailand monophyletically clustered with M. colombiense CECT 3035, M. colombiense TKK-01-0051, and M. colombiense 230 (reference strains). Drug susceptibility, specimen types, and origins are presented as a color strip labelled beside the tips of the tree.
The SNPs-tree estimated for M. intracellulare/M. paraintracellulare presented three major clades, including M. intracellulare subsp. intracellulare group (MI), M. intracellulare subsp. chimaera group (MC), and an unknown clade (Fig. 4). Based on the current NCBI nomenclature (July 2024), the MI clade included the groups of M. intracellulare and M. paraintracellulare species, while the MC clade comprised M. intracellulare subsp. yongonense and M. intracellulare subsp. chimaera. Six CU isolates and three KKU isolates clustered within the unknown clade. These isolates were clustered with M. intracellulare E2190 and E3191 isolated from Cambodia. In the MI group, we defined three subclades: MI1 comprised two KKU isolates and other strains from USA, MI2 included all strains of previously known M. paraintracellulare, and MI3 included other strains of M. intracellulare and one CU isolate (CU MAC05).
SNP-based phylogenetic tree of Mycobacterium intracellulare/Mycobacterium paraintracellulare isolates in Thailand. The phylogenetic tree based on SNPs from the Snippy pipeline showed the diversity of M. intracellulare/M. paraintracellulare isolates in Thailand and their phylogenetic relationships compared to global reference strains. The tree presented three clades, including M. intracellulare subsp. intracellulare group (MI; blue), M. intracellulare subsp. chimaera group (MC; red), and unknown clade. Drug susceptibility, specimen types, and origins are presented as a color strip labelled beside the tips of the tree.
For the MC group, 47 CU isolates and six KKU isolates were clustered and divided into nine subclades. Interestingly, several isolates which were members of subclades MC2-4 and MC6 were clustered without having any related reference strains, suggesting the high diversity of MC isolates in Thailand. The members of MC8 and MC9 monophyletically clustered with reference strains of M. intracellulare subsp. chimaera sensu stricto. Additional analysis of M. intracellulare subsp. chimaera-specific markers (SR1-SR6) reported by Zozaya-Valdés et al., 201715 was performed using a BLAST search against draft genomes of isolates. The results showed that MC8 carried SR3, SR4, and SR5 markers, while MC9 presented SR1, SR5, and SR6 within their genomes. These markers were completely absent in other members of the MC group (MC1-7). The reference genome of M. intracellulare subsp. yongonense, which is now reclassified as M. intracellulare subsp. chimaera16,17, clustered with three CU isolates in MC5.
For antimicrobial resistance of M. intracellulare/M. paraintracellulare, most of the isolates were resistant to linezolid and moxifloxacin. For moxifloxacin, we identified a non-synonymous mutation (S95T) in quinolone resistance-determining regions (QRDR) of gyrA of all M. intracellulare isolates, although this mutation was not correlated with the phenotypic antimicrobial susceptibility results. The mutations in rplC and rplD were not associated with linezolid resistance, despite the previously reported R148K mutation in rplD18, which was found in several isolates. Six isolates were resistant to clarithromycin, while seven isolates were resistant to amikacin. Based on SNPs analysis, some isolates showed mutations in the drug target genes. The two isolates of MC9 (CU MAC88 and CU MAC89), which were resistant to clarithromycin, had a mutation in 23S rRNA (rrl) gene at position A2268T (position based on M. intracellulare ATCC 13950). One isolate from the MC3 (KKU MAC80459) also had the A2268G mutation in rrl gene, which may be associated with clarithromycin resistance. For amikacin resistance, several mutations in 16S rRNA (rrs) gene which may be associated with amikacin resistance were identified. Two isolates (KKU MAC81283 and KKU MAC81569) had T87C mutation, while KKU MAC82837 had both T192C and A197T in the rrs gene (Table 3).
Pan-genome analysis of Mycobacterium intracellulare/Mycobacterium paraintracellulare
The core- and accessory-gene cluster analysis by PanExplorer showed that approximately 2,448 (9.9%) were core-gene clusters, while the others were strain-specific and dispensable clusters, which account for 8,105 (32.8%) and 14,131 (57.2%) genes, respectively (Fig. 5a). The genes in 65 genomes of CU and KKU isolates were analyzed for the subsystem of protein families (Fig. 5b) using the NCBI BV-BRC system. Most genes were related to metabolism (49,350 genes) and stress response, defense, and virulence (14,790 genes). These genes were classified into protein families, with specific protein families categorized for the MI group and unknown clade (n = 308) (Fig. 5c) and MC group (n = 1,870) (Fig. 5d).
Pan-genome analysis of Mycobacterium intracellulare/Mycobacterium paraintracellulare isolates from Thailand. (A) The pie chart shows the number of core- and accessory-gene clusters in the genomes of isolates and references of M. intracellulare/M. paraintracellulare (n = 176) analyzed by PanExplorer. The 65 genomes of CU and KKU isolates were analyzed for (B) subsystem of protein families shown as a pie chart using NCBI BV-BRC system. In addition, protein families which were specific for (C) MI group and unknown clade and (D) MC group are shown as heatmaps.
The gene families (Pfam) analysis provided information about the clade-specific gene families. Briefly, aliphatic amidase (AmiE) and proline-proline-glutamic acid (PPE) protein family 3 (PPE3) were specific for the M. intracellulare unknown clade. Proline-glutamic acid (PE) protein family 19 (PE19) was present in the unknown clade, except for two isolates (CU MAC36 and CU MAC38). For MI isolates, they have the methoxy mycolic acid synthase (MmaA2), which was absent in all other isolates. MI1 isolates exclusively carried cell surface lipoprotein (MPT83) and protein secretion chaperonin (CsaA), while the MI3 isolate had NADH: quinone reductase, acyl-CoA dehydrogenase (FadE17) and lactate-responsive regulator (LldR). The Pfam analysis of MC isolates showed that all MC members contained the partial REP13E12 repeat protein in their genome. The MC8 isolates carried acyl-CoA dehydrogenase (FadE1), while MC9 genomes exclusively contained several proteins, including RelB/StbD replicon stabilization protein (antitoxin to RelE/StbE), PPE protein family 4 (PPE4), and catalase (KatE). Another specific protein family for MC9 was associated with gamma-hexachlorocyclohexane degradation, as well as toluene and xylene degradation.
Discussion
In this study, comprehensive genomic analysis and detailed characterization provide an understanding of the species distribution and antimicrobial resistance profiles of MAC clinical isolates in Thailand, which could be beneficial for clinical management and as a guide for the development of targeted diagnostic and treatment strategies for MAC infections in the region. Based on the initial identification of the MAC clinical isolates using the LPA and the FastANI method, we found that the LPA has limitations for the identification of M. colombiense isolates. Moreover, LPA, specifically GenoType Mycobacterium CM VER 2.0 and INNO-LiPA MYCOBACTERIA v2, cannot differentiate M. chimaera (currently known as M. intracellulare subsp. chimaera) from M. intracellulare, whereas this can be identified by genome-based identification. This discrepancy highlights the limitations of LPA in accurately distinguishing between closely related MAC species19,20,21. Cross-reaction of M. intracellulare probe has been reported previously19, likely since the GenoType NTM-DR assay only targets the 23S rRNA (rrl) gene20. In addition, the target MAC species for LPA are limited to M. intracellulare, M. avium, and M chimaera (included only in GenoType NTM-DR). The identification of M. colombiense and M. timonense in this study expands the recognized species diversity of MAC in the region. Our study highlights that the presence of multiple isolates of M. colombiense leads to additional concerns regarding accurate identification and the effective treatment plans. Moreover, M. colombiense is a relatively recently described species within the MAC22. Although its clinical significance and epidemiology are not yet fully understood, recent findings indicate its potential involvement in opportunistic infections in immunosuppressed hosts such as HIV-infected patients23,24.
The core-genome phylogenetic analysis and SNP-based phylogenetic trees demonstrated the existence of distinct clades and subclades within MAC species, and the wide diversity of MAC isolates in Thailand across this spectrum. This diversity likely reflects the complex evolutionary history and distribution of MAC species causing diseases in humans. M. intracellulare was the most prevalent MAC species identified in this study. The high prevalence and diversity of M. intracellulare isolates observed is consistent with previous reports from Khon Kaen, Thailand13 and other countries such as China25, Japan26, and Korea27. Previously, the nomenclature of M. intracellulare and its closely related species has been controversial. The related species previously identified include M. intracellulare, M. paraintracellulare, M. chimaera, and M. yongonense3. The nomenclature of these MAC had been recently reviewed and updated16, but limited study has been conducted based on their comparative genomic analysis. To date, the NCBI taxonomy classifies them into two species, namely (1) M. paraintracellulare and (2) M. intracellulare which comprises three subspecies: M. intracellulare subsp. intracellulare, M. intracellulare subsp. chimaera, and M. intracellulare subsp. yongonense. The current analysis provides an in-depth analysis of their phylogenetic relationships. The clinical isolates from Thailand predominantly clustered into distinct clades that diverged from global reference stains, suggesting region-specific evolution. Interestingly, they also showed high diversity, which can be classified into three clades including M. intracellulare subsp. intracellulare (MI) group, M. intracellulare subsp. chimaera (MC) group, and a divergent unknown clade. Based on the genomic analysis, the MI group should include M. intracellulare subsp. intracellulare and former members of M. paraintracellulare, while the MC group comprises M. intracellulare subsp. chimaera and former members of M. intracellulare subsp. yongonense. This classification aligns with the previous analyses of M. intracellulare, which identified two divergent clades corresponding to M. intracellulare subsp. intracellulare and M. intracellulare subsp. chimaera28. In addition, the members of M. intracellulare subsp. yongonense should be classified as a group of MC members, supporting previous reports that M. intracellulare subsp. yongonense is a heterotypic synonym of M. intracellulare subsp. chimaera16.
The mutational analysis on drug target genes showed that M. intracellulare subsp. chimaera (MC3 and MC9) had A2268T/G, while M. colombiense carried A2270G/C mutations in rrl gene. These mutations correspond to A2058T/G and A2059G/C based on E. coli numbering, which are well-known as clarithromycin resistance-associated mutations reported previously in MAC and other NTM29,30,31. The current analysis also showed potential novel mutations (C1687T, A215G, T288C) associated with clarithromycin resistance in CU MAC45, but they were not statistically significant (Chi-Square test, P = 0.12, shown in supplementary table S2). These mutations should be further explored for larger cohorts of M. colombiense isolates. For amikacin resistance, the A1408G and C1409T substitutions reported previously32 were not identified in the current study. However, some novel mutations in M. intracellulare subsp. chimaera (MC4) from Khon Kaen including T87C, T192C, and A197T were potentially associated with amikacin resistance (Chi-Square test, P < 0.05). For the A197T mutation, it was also identified in other reference genomes (Supplementary table S3) from the USA. However, the association of the A197T mutation with amikacin resistance in those isolates remains unknown.
In this study, the results demonstrated that amikacin and clarithromycin remain effective against MAC clinical isolates in Thailand, while most of MAC members showed potential resistance to linezolid and moxifloxacin. However, the current analysis was limited by a small number of isolates and the use of breakpoints for MIC interpretation following Clinical and Laboratory Standards Institute (CLSI) guidelines, which are only applicable for four of the 13 drugs tested33,34. In this analysis, some isolates exhibited no potential mutations in the drug target genes for amikacin and clarithromycin, suggesting the involvement of other resistance mechanisms such as changes in membrane permeability, increased efflux of antibiotics, alterations in antibiotic-activating enzymes, or activation of resistant metabolic pathways35.
The comparative genome analysis from our study showed clade-specific gene families for M. intracellulare. Notably, some proteins in PE/PPE family were highly specific to distinct M. intracellulare clades. The role of PE/PPE family in M. tuberculosis complex (MTBC) may involve the ESX system, which plays a role in host cell interactions and immune response regulation36. Nevertheless, while the functions of PE/PPE in MAC are still unclear, they are potentially associated with virulence37,38. Previous studies have shown that PE and PPE are varied among MAC members39, consistent with the findings of the present study. Specifically, PPE3 and PE19 were related to M. intracellulare unknown clade, while PPE4 was specific to M. intracellulare subsp. chimaera (MC). However, they are not involved in the ESX operon as found in MTBC. For MmaA2 found in MI, this protein shared a redundant function with CmaA2 which is involved in cell envelop lipid and may be implicated in the pathogenesis of M. tuberculosis infection40. Cell surface lipoprotein MPT83, a cell wall-associated lipoglycoprotein that potentially stimulates protective immunity against M. tuberculosis, was specifically found in MI1 isolates41. In contrast, MI3 isolates carried the specific proteins related to electron transport chain (NADH: quinone reductase)42 and fatty acid metabolism (acyl-CoA dehydrogenase FadE17, which are possibly involved in mycobacterial adaptation25,43. For some MC members, MC8 also had the specific acyl-CoA dehydrogenase (FadE1), supporting the hypothesis of mycobacterial adaptation to survive in specific environments25. The RelB in MC9 overlapping with RelE genes in MC9 may associate with a regulation system of stress response and survival as proposed previously44. These results provide preliminary insights into the genomic functions for these MAC species, which will be valuable for further studies on their pathogenesis and therapeutic development.
In conclusion, this study demonstrates the high genomic diversity of MAC clinical isolates in Thailand, with M. intracellulare being the predominant species, followed by M. avium, M. colombiense, and M. timonense. The identification of M. colombiense as an emerging cause of MAC infections in Thailand is a notable finding. The present study suggests that M. intracellulare members could be reclassified into 2 species: M. intracellulare subsp. intracellulare (MI) and M. intracellulare subsp. chimaera (MC). The mutations associated with resistance to clarithromycin among the MAC isolates in this study are concerning and highlight the need for improved antimicrobial strategies and the development of novel therapeutics. The comprehensive genomic characterization of MAC isolates in this study provides valuable insights that can inform clinical management and guide the development of targeted diagnostic and treatment strategies for MAC infections in Thailand and other regions. Lastly, the genomic information will be helpful for further studies of MAC genomic features that influence pathogenesis and host-pathogen interactions.
Methods
Study population
MAC isolates (CU cohort) were recovered from clinical specimens from patients at KCMH, Bangkok, Thailand, between January 2016 and December 2022. Data related to microbiology, epidemiology, clinical presentations, and clinical outcomes of patients were examined from the medical records. A total of 66 MAC clinical isolates, including single and sequential isolates from 41 patients, were included in the study. All patients met the diagnostic criteria of NTM-PD according to clinical, radiological, and microbiological criteria (≥ 2 positive sputa or endotracheal aspirates with the same species isolated, or ≥ 1 positive BAL or sterile sites)8,10 or extrapulmonary NTM infection. All experiments were performed in accordance with relevant guidelines and regulations. This study has been approved by the Institutional Review Board (IRB) of the Faculty of Medicine, Chulalongkorn University (IRB No. 0840/65, COA No. 1723/2022) with the exemption of patient informed consent.
Mycobacterial isolation and culture
MAC isolates were recovered from clinical specimens using the sodium hydroxide-N-acetyl-L-cysteine-sodium citrate method45. These isolates were identified to species or complex level based on LPA kits using the GenoType Mycobacterium CM VER 2.0 (Hain Lifescience) (M. avium and M. intracellulare), GenoType NTM-DR assay (Hain Lifescience) (M. avium, M. intracellulare, and M. chimaera), or INNO-LiPA MYCOBACTERIA v2 (Fujirebio) (M. avium and M. intracellulare) following the manufacturer’s protocol. The isolates with mixed bands of M. avium and M. intracellulare were reported as M. avium complex. The frozen stocks of MAC clinical isolates were sub-cultured on Lowenstein-Jensen slants at 35 °C for 7–10 days for antimicrobial susceptibility testing and genomic DNA extraction.
Antimicrobial susceptibility testing
The MIC of 13 antimicrobial agents was examined using Sensititre™ Myco SLOMYCOI (Thermo Fisher Scientific) and results were interpreted according to the CLSI guidelines M24, Ed3 and M24S33,34 (Supplementary table S4). For drugs without addressed breakpoints, the MIC values were reported and calculated for the MIC50 and MIC90. The MIC plates were incubated at 35 ºC for 7 days until a positive control had sufficient growth33. The MIC50 and MIC90 are determined as the MIC values that inhibit the growth of 50% and 90% of the tested MAC clinical isolates, respectively. The colonies of M. avium ATCC 700898 was included as quality controls.
Genomic DNA extraction and next-generation sequencing (NGS)
For next-generation sequencing (NGS), genomic DNA was extracted from bacterial colonies using the ZymoBIOMICS DNA Miniprep Kit (Zymo Research) following the manufacturer’s protocol. Genomic DNA libraries were prepared using the NEBNext® Ultra™DNA Library Prep Kit for Illumina® (New England Biolabs) following the manufacturer’s protocol. NGS was carried out using a 2 × 150 paired-end (PE) configuration on an Illumina NovaSeq 6000 instrument. The sequence data were deposited and are publicly available in the NCBI Sequence Read Archive (SRA) with BioProject accession: PRJNA 1142684.
Bioinformatics analysis
Sequence preprocessing and genome assembly
The raw paired-end FASTQ sequences of 66 MAC clinical isolates from KCMH, Bangkok, Thailand (CU cohort; current study; collected during 2016–2022; under BioProject accession No. PRJNA1142684) and 23 MAC isolates previously sequenced from Khon Kaen University, Khon Kaen, Thailand located approximately 450 km northeast of Bangkok (KKU cohort; collected during 2013–2016; under BioProject accession No. PRJNA972846) were included for analysis. The FASTQ sequences were quality-checked by FASTQC v0.11.8. Then, the sequences were subjected to adaptor trimming and quality trimming (< Q30) using Trimmomatic v0.3946. The sequences longer than 100 bp were retained for further analysis. The draft genome assembly was carried out by Unicycler v0.5.047. The quality and completeness of the assembled genomes was evaluated by QUAST 5.1.048. The genomes which had completeness > 99% and no contamination were retained for further analysis. The filtered genomes were annotated using Prokka v1.1149.
Genome classification and core genome analysis
The assembled genomes were compared to the reference genomes of MAC species available in https://www.ncbi.nlm.nih.gov/genome using FastANI50. The classification of MAC species was determined based on the best hits to the references. The list of reference genomes used for FastANI-based comparison are in supplementary table S5. The annotated genomes from Prokka and reference genomes were used for core genome analysis by Panaroo 1.3.4 with option --clean-mode strict --remove-invalid-genes51. The core gene alignment was then used for the phylogenetic analysis conducted by IQ-TREE v2.3.0 with a GTR substitution model52 and 100 bootstrap replicates. The phylogenetic tree was used for the determination of MAC species based on clade clustering with reference strains. The classification results from FastANI and the core-gene phylogeny were compared with the results obtained using the LPA method.
SNP-based phylogenetic analysis
The cleaned reads were mapped to the reference genome of M. avium subsp. hominissuis (CP018019.1), M. intracellulare ATCC 13950 (CP003322.1) or M. colombiense CECT 3035 (GCF_000222105.3) depending on the classified species and subjected to haploid variant calling using Snippy v4.6.0 with default parameters. The reference genomes of the described species derived from the NCBI genome (Supplementary table S5) were also included in the analysis. The alignment of single nucleotide polymorphisms (SNPs) was checked for recombination using Gubbins v3.3.4. After that, the phylogenetic trees of M. avium, M. intracellulare, and M. colombiense were constructed using IQ-TREE with GTR model and 100 bootstrap replicates. The phylogenetic trees were visualized using the iTol web tool v6.9 (https://itol.embl.de/).
Genomic analysis for antimicrobial resistance
In addition, the variant calling files (VCFs) generated by Snippy were also utilized to analyze genetic variants in specific genes, including 16S rRNA (rrs), 23S rRNA (rrl), gyrA, gyrB, large ribosomal subunit protein L3 (rplC), and large ribosomal subunit protein L4 (rplD) to investigate the mutations associated with drug resistance. The rrs gene is a target for amikacin, while the rrl gene is a target for clarithromycin and linezolid. The gyrA and gyrB genes are targets for moxifloxacin, whereas rplC and rplD are linezolid targets. The putative variants were determined based on the criterion that these mutations should be exclusively present in the resistant strains compared to the closest susceptible strains based on SNPs trees.
Pan-genome analysis for M. intracellulare/M. paraintracellulare
The annotated genomes from Prokka (Genbank format) and reference genomes of M. intracellulare/M. paraintracellulare were submitted to PanExplorer53 web-based tool (https://panexplorer.southgreen.fr/cgi-bin/home.cgi) for investigating core- and accessory-gene clusters. The 65 genomes of M. intracellulare from CU (n = 54) and KKU (n = 11) cohorts were also annotated and used for comparative systems analysis using the NCBI BV-BRC system (https://www.bv-brc.org/). This analysis provides information on gene families, pathways, and subsystems.
Data availability
The raw sequence data are publicly available in the NCBI Sequence Read Archive (SRA), under the BioProject accession number PRJNA 1142684.
References
Nishiuchi, Y., Iwamoto, T. & Maruyama, F. Infection sources of a common non-tuberculous mycobacterial pathogen, Mycobacterium avium complex. Front. Med. (Lausanne). 4, 27. https://doi.org/10.3389/fmed.2017.00027 (2017).
Stout, J. E., Koh, W. J. & Yew, W. W. Update on pulmonary disease due to non-tuberculous mycobacteria. Int. J. Infect. Dis. 45, 123–134. https://doi.org/10.1016/j.ijid.2016.03.006 (2016).
van Ingen, J., Turenne, C. Y., Tortoli, E., Wallace, R. J. Jr. & Brown-Elliott, B. A. A definition of the Mycobacterium avium complex for taxonomical and clinical purposes, a review. Int. J. Syst. Evol. Microbiol. 68, 3666–3677. https://doi.org/10.1099/ijsem.0.003026 (2018).
Castejon, M., Menendez, M. C., Comas, I., Vicente, A. & Garcia, M. J. Whole-genome sequence analysis of the Mycobacterium avium complex and proposal of the transfer of Mycobacterium yongonense to Mycobacterium intracellulare subsp. yongonense subsp. Nov. Int. J. Syst. Evol. Microbiol. 68, 1998–2005. https://doi.org/10.1099/ijsem.0.002767 (2018).
Nouioui, I. et al. Genome-based taxonomic classification of the phylum actinobacteria. Front. Microbiol. 9, 2007. https://doi.org/10.3389/fmicb.2018.02007 (2018).
Koh, W. J. et al. Clinical significance of the differentiation between Mycobacterium avium and Mycobacterium intracellulare in M. Avium complex lung disease. Chest 142, 1482–1488. https://doi.org/10.1378/chest.12-0494 (2012).
Boyle, D. P., Zembower, T. R., Reddy, S. & Qi, C. Comparison of clinical features, virulence, and relapse among Mycobacterium avium complex species. Am. J. Respir Crit. Care Med. 191, 1310–1317. https://doi.org/10.1164/rccm.201501-0067OC (2015).
Griffith, D. E. et al. An official ATS/IDSA statement: diagnosis, treatment, and prevention of nontuberculous mycobacterial diseases. Am. J. Respir Crit. Care Med. 175, 367–416. https://doi.org/10.1164/rccm.200604-571ST (2007).
Hoefsloot, W. et al. The geographic diversity of nontuberculous mycobacteria isolated from pulmonary samples: an NTM-NET collaborative study. Eur. Respir J. 42, 1604–1613. https://doi.org/10.1183/09031936.00149212 (2013).
Daley, C. L. et al. Treatment of nontuberculous mycobacterial pulmonary disease: an official ATS/ERS/ESCMID/IDSA clinical practice guideline. Clin. Infect. Dis. 71, 905–913. https://doi.org/10.1093/cid/ciaa1125 (2020).
Haworth, C. S. et al. British Thoracic Society guidelines for the management of non-tuberculous mycobacterial pulmonary disease (NTM-PD). Thorax 72, ii1–ii64. https://doi.org/10.1136/thoraxjnl-2017-210927 (2017).
Griffith, D. E. Treatment of Mycobacterium avium complex (MAC). Semin Respir Crit. Care Med. 39, 351–361. https://doi.org/10.1055/s-0038-1660472 (2018).
Sirichoat, A., Kaewprasert, O., Hinwan, Y. & Faksri, K. Phenotypic drug-susceptibility profiles and genetic analysis based on whole-genome sequencing of Mycobacterium avium complex isolates in Thailand. PLoS One. 18, e0294677. https://doi.org/10.1371/journal.pone.0294677 (2023).
van Ingen, J. et al. Treatment outcome definitions in nontuberculous mycobacterial pulmonary disease: an NTM-NET consensus statement. Eur. Respir J. 51 https://doi.org/10.1183/13993003.00170-2018 (2018).
Zozaya-Valdes, E. et al. Target-specific assay for rapid and quantitative detection of Mycobacterium chimaera DNA. J. Clin. Microbiol. 55, 1847–1856. https://doi.org/10.1128/JCM.00197-17 (2017).
Armstrong, D. T. & Parrish, N. Current updates on mycobacterial taxonomy, 2018 to 2019. J. Clin. Microbiol. 59, e0152820. https://doi.org/10.1128/JCM.01528-20 (2021).
Wengenack, N. L. et al. This is giving me a complex: a practical attempt to streamline nontuberculous mycobacteria nomenclature for clinical purposes. J. Clin. Microbiol. 62, e0153123. https://doi.org/10.1128/jcm.01531-23 (2024).
Kim, S. Y. et al. Genetic mutations in linezolid-resistant Mycobacterium avium complex and Mycobacterium abscessus clinical isolates. Diagn. Microbiol. Infect. Dis. 94, 38–40. https://doi.org/10.1016/j.diagmicrobio.2018.10.022 (2019).
Huh, H. J. et al. GenoType NTM-DR performance evaluation for identification of Mycobacterium avium complex and Mycobacterium abscessus and determination of clarithromycin and amikacin resistance. J. Clin. Microbiol. 57. https://doi.org/10.1128/JCM.00516-19 (2019).
Mok, S., Rogers, T. R. & Fitzgibbon, M. Evaluation of GenoType NTM-DR assay for identification of Mycobacterium chimaera. J. Clin. Microbiol. 55, 1821–1826. https://doi.org/10.1128/JCM.00009-17 (2017).
Richter, E., Rusch-Gerdes, S. & Hillemann, D. Evaluation of the GenoType Mycobacterium assay for identification of mycobacterial species from cultures. J. Clin. Microbiol. 44, 1769–1775. https://doi.org/10.1128/JCM.44.5.1769-1775.2006 (2006).
Murcia, M. I., Tortoli, E., Menendez, M. C., Palenque, E. & Garcia, M. J. Mycobacterium colombiense sp. nov., a novel member of the Mycobacterium avium complex and description of MAC-X as a new ITS genetic variant. Int. J. Syst. Evol. Microbiol. 56, 2049–2054. https://doi.org/10.1099/ijs.0.64190-0 (2006).
Andreani, J., Barrassi, L. & Davoust, B. La Scola, B. evidence of an environmental reservoir for emergent Mycobacterium colombiense. New. Microbes New. Infect. 35, 100666. https://doi.org/10.1016/j.nmni.2020.100666 (2020).
Yu, X. & Jiang, W. Mycobacterium colombiense and Mycobacterium avium complex causing severe pneumonia in a patient with HIV identified by a novel molecular-based method. Infect. Drug Resist. 14, 11–16. https://doi.org/10.2147/IDR.S282190 (2021).
Song, Z. et al. High diversity of clinical Mycobacterium intracellulare in China revealed by whole genome sequencing. Front. Public. Health. 10, 989587. https://doi.org/10.3389/fpubh.2022.989587 (2022).
Ichikawa, K. et al. Genetic diversity of clinical Mycobacterium avium subsp. hominissuis and Mycobacterium intracellulare isolates causing pulmonary diseases recovered from different geographical regions. Infect. Genet. Evol. 36, 250–255. https://doi.org/10.1016/j.meegid.2015.09.029 (2015).
Kim, S. Y. et al. Genotyping of Mycobacterium intracellulare isolates and clinical characteristics of lung disease. Int. J. Tuberc. Lung Dis. 17, 669–675. https://doi.org/10.5588/ijtld.12.0575 (2013).
Hasan, N. A. et al. Population genomics and inference of Mycobacterium avium complex clusters in cystic fibrosis care centers, United States. Emerg. Infect. Dis. 27, 2836–2846. https://doi.org/10.3201/eid2711.210124 (2021).
Meier, A. et al. Identification of mutations in 23S rRNA gene of clarithromycin-resistant Mycobacterium intracellulare. Antimicrob. Agents Chemother. 38, 381–384. https://doi.org/10.1128/AAC.38.2.381 (1994).
Nash, K. A. & Inderlied, C. B. Genetic basis of macrolide resistance in Mycobacterium avium isolated from patients with disseminated disease. Antimicrob. Agents Chemother. 39, 2625–2630. https://doi.org/10.1128/AAC.39.12.2625 (1995).
Tait-Kamradt, A. et al. Mutations in 23S rRNA and ribosomal protein L4 account for resistance in pneumococcal strains selected in vitro by macrolide passage. Antimicrob. Agents Chemother. 44, 2118–2125. https://doi.org/10.1128/AAC.44.8.2118-2125.2000 (2000).
Zhang, Z. et al. Inducible resistance to amikacin in Mycobacterium abscessus isolated in Beijing, China. Infect. Drug Resist. 15, 2287–2291. https://doi.org/10.2147/IDR.S357887 (2022).
CLSI. Susceptibility testing of mycobacteria, Nocardia. spp., and other aerobic Actinomycetes, 3rd ed. CLSI document M24 (Clinical and Laboratory Standards Institute, 2018).
CLSI. Performance standards for susceptibility testing of mycobacteria, Nocardia. spp., and other aerobic actinomycetes. 2nd edition. CLSI supplement M24S (Clinical and Laboratory Standards Institute, 2023).
Parker, H., Lorenc, R., Ruelas Castillo, J. & Karakousis, P. C. Mechanisms of antibiotic tolerance in Mycobacterium avium complex: lessons from related mycobacteria. Front. Microbiol. 11, 573983. https://doi.org/10.3389/fmicb.2020.573983 (2020).
Qian, J., Chen, R., Wang, H. & Zhang, X. Role of the PE/PPE family in host-pathogen interactions and prospects for anti-tuberculosis vaccine and diagnostic tool design. Front. Cell. Infect. Microbiol. 10, 594288. https://doi.org/10.3389/fcimb.2020.594288 (2020).
Li, Y., Miltner, E., Wu, M., Petrofsky, M. & Bermudez, L. E. A Mycobacterium avium PPE gene is associated with the ability of the bacterium to grow in macrophages and virulence in mice. Cell. Microbiol. 7, 539–548. https://doi.org/10.1111/j.1462-5822.2004.00484.x (2005).
Ramakrishnan, L., Federspiel, N. A. & Falkow, S. Granuloma-specific expression of Mycobacterium virulence proteins from the glycine-rich PE-PGRS family. Science 288, 1436–1439. https://doi.org/10.1126/science.288.5470.1436 (2000).
Mackenzie, N., Alexander, D. C., Turenne, C. Y., Behr, M. A. & De Buck, J. M. Genomic comparison of PE and PPE genes in the Mycobacterium avium complex. J. Clin. Microbiol. 47, 1002–1011. https://doi.org/10.1128/JCM.01313-08 (2009).
Barkan, D., Rao, V., Sukenick, G. D. & Glickman, M. S. Redundant function of cmaA2 and mmaA2 in Mycobacterium tuberculosis cis cyclopropanation of oxygenated mycolates. J. Bacteriol. 192, 3661–3668. https://doi.org/10.1128/JB.00312-10 (2010).
Kao, F. F. et al. The secreted lipoprotein, MPT83, of Mycobacterium tuberculosis is recognized during human tuberculosis and stimulates protective immunity in mice. PLoS One 7, e34991. https://doi.org/10.1371/journal.pone.0034991 (2012).
Yano, T. et al. Mycobacterium tuberculosis type II NADH-menaquinone oxidoreductase catalyzes electron transfer through a two-site ping-pong mechanism and has two quinone-binding sites. Biochemistry 53, 1179–1190. https://doi.org/10.1021/bi4013897 (2014).
Mokrousov, I. et al. Highly lethal strain cluster of the Mycobacterium tuberculosis Beijing genotype endemically prevalent in Buryatia, Russia. Eur. Respir. J. 58 (PA2475). https://doi.org/10.1183/13993003.congress-2021.PA2475 (2021).
Singh, R., Barry, C. E., Boshoff, H. I. & rd & The three RelE homologs of Mycobacterium tuberculosis have individual, drug-specific effects on bacterial antibiotic tolerance. J. Bacteriol. 192, 1279–1291. https://doi.org/10.1128/JB.01285-09 (2010).
Global Laboratory Initiative a Working Group of the Stop TB Partnership. (2014). Mycobacteriology laboratory manual.
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120. https://doi.org/10.1093/bioinformatics/btu170 (2014).
Wick, R. R., Judd, L. M., Gorrie, C. L. & Holt, K. E. Unicycler: resolving bacterial genome assemblies from short and long sequencing reads. PLoS Comput. Biol. 13, e1005595. https://doi.org/10.1371/journal.pcbi.1005595 (2017).
Gurevich, A., Saveliev, V., Vyahhi, N. & Tesler, G. QUAST: quality assessment tool for genome assemblies. Bioinformatics 29, 1072–1075. https://doi.org/10.1093/bioinformatics/btt086 (2013).
Seemann, T. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069. https://doi.org/10.1093/bioinformatics/btu153 (2014).
Jain, C., Rodriguez, R. L., Phillippy, A. M., Konstantinidis, K. T. & Aluru, S. High throughput ANI analysis of 90K prokaryotic genomes reveals clear species boundaries. Nat. Commun. 9, 5114. https://doi.org/10.1038/s41467-018-07641-9 (2018).
Tonkin-Hill, G. et al. Producing polished prokaryotic pangenomes with the Panaroo pipeline. Genome Biol. 21, 180. https://doi.org/10.1186/s13059-020-02090-4 (2020).
Nguyen, L. T., Schmidt, H. A., von Haeseler, A. & Minh, B. Q. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 32, 268–274. https://doi.org/10.1093/molbev/msu300 (2015).
Dereeper, A., Summo, M. & Meyer, D. F. PanExplorer: a web-based tool for exploratory analysis and visualization of bacterial pan-genomes. Bioinformatics 38, 4412–4414. https://doi.org/10.1093/bioinformatics/btac504 (2022).
Acknowledgements
This research is funded by Thailand Science research and Innovation Fund Chulalongkorn University (HEA663000042) and Ratchadapiseksompotch Fund, Faculty of Medicine, Chulalongkorn University, grant number RA66/026. We also would like to thank the Mahidol Artificial Intelligent Center (MAI) and Center for Artificial Intelligence in Medicine – Chulalongkorn university (CU-AIM) for high performance computing (HPC) clusters used for bioinformatics analysis.
Author information
Authors and Affiliations
Contributions
V.S. conceived the study design, performed the bioinformatics analysis, and wrote the manuscript. K.W. wrote the manuscript and collected clinical data. S.Pe., U.S., and Y.Y. performed the experiments. K.F. collected data and supervised the project for KKU cohort. T.F. and S.Pa. designed and supervised the research. S.R. designed and supervised the research, collected and analyzed data, and wrote the manuscript. All authors reviewed and critiqued the draft manuscript and approved the final manuscript prior to submission for publication.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
This study related with human was approved by the Institutional Review Board (IRB) of the Faculty of Medicine, Chulalongkorn University with IRB No. 0840/65 (COA No. 1723/2022).
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Sawaswong, V., Wongjarit, K., Petsong, S. et al. Diversity and antimicrobial resistance profiles of Mycobacterium avium complex clinical isolates in Thailand based on whole genome comparative analysis. Sci Rep 15, 772 (2025). https://doi.org/10.1038/s41598-024-84511-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-84511-z