Abstract
Metabolomics is the measurement of metabolites in biological samples to reveal information on metabolic pathways and phenotypes. Cluster analysis is a popular multivariate technique employed in metabolomics to characterise observations with similar features. Previous work in the field has applied hard clustering approaches to group observations into distinct clusters. This approach can be overly restrictive in some practical applications. Therefore, there is a growing need for soft clustering methods that allow for the clustering of observations into more than one cluster. Simplex-structured matrix factorisation (SSMF) is proposed and applied in a simulation study and to a metabolomic dataset to demonstrate its utility for soft clustering. In the simulation study, the cluster prototypes and cluster memberships were well estimated. In the real data application to metabolomic data, the presence of four soft clusters was suggested by the gap statistic. Furthermore, the Shannon diversity index indicated that several observations have memberships in three clusters. Additionally, the introduction of the covariates sex, age and BMI revealed that sex and age mainly associated with the cluster memberships. The results indicate that a majority of men and young people were in the cluster predominantly characterised by high levels of amino acids and low levels of phosphatidylcholines and sphingomyelins. However, a high proportion of older people were characterised by low levels of amino acids, biogenic amines, acylcarnitines and lysophosphatidylcholines. The SSMF presented successfully estimates a soft clustering of the metabolomic data. It provides an interpretable representation of the data structure using the cluster prototypes combined with cluster memberships. A software package called MetabolSSMF has been developed, which is freely available as an R package, to facilitate the implementation of soft clustering in the field of metabolomics.
Similar content being viewed by others
Introduction
Metabolomics involves measurement of a range of small molecules called metabolites in biological samples, including body fluids and tissues. The metabolomic profile enables a comprehensive assessment of the metabolic status of the organism, which is linked to genetics, the microbiome, and environmental factors (such as exercise, pollutants, or diet) and provides detailed insights into metabolic pathways and biological processes1,2. Nuclear magnetic resonance spectroscopy (NMR) and mass spectrometry (MS) are the two primary platforms used for metabolomic analysis3. Since the data obtained through these methods often exhibit high dimensionality and complexity, the use of multivariate analysis techniques is recommended3,4.
Cluster analysis, a popular multivariate analysis technique, is often employed in metabolomics to group data into clusters based on specific criteria. It aims to ensure that the observations in the same cluster are more similar than observations in other clusters5,6. Cluster analysis methods can be divided into hard clustering and soft clustering7. Hard clustering methods, such as K-means and K-medoids clustering, assign observations to precisely one cluster with minimal overlapping of clusters. In contrast, soft clustering methods classify the data points into more than one cluster with different degrees of cluster membership, allowing clusters to overlap and indicating the strength of association between observation and cluster8. Hard clustering has several merits, such as fast convergence and easy implementation9,10. However, forcing data points to be in one cluster could lead to unrealistic classifications in practical applications7. For example, in an application to thyroid disease11, data measurements for 215 subjects from three known disease classes were analysed using clustering methods. Even though both hard clustering and soft clustering worked well, the authors conclude that soft clustering algorithms provide better separation and meaningful clustering with a high degree of compactness. In a study of type 2 diabetes (T2D)12, Bayoumi et al. employed unsupervised soft clustering to identify five clusters among 348 patients, providing valuable insights into T2D subtypes. An advantage of soft clustering methods is that they provide the flexibility to express data points belonging to multiple clusters simultaneously with different membership levels. Consequently, the application of soft clustering approaches has recently garnered attention for solving the problems on increasingly complex data7.
Currently, the clustering of metabolomic data is still dominated by hard clustering approaches. For example, in a study investigating the phenotypic and genotypic relationships in young women with hypercholesterolemia13, hierarchical clustering was carried out using plasma metabolomic data, leading to the identification of four subtypes of hypercholesterolemia. Galal et al.14 have proposed that machine learning techniques, such as k-nearest neighbour and neural networks, are capable of handling clustering of intricate metabolomic data.
Fuzzy C-means (FCM) is a popular soft clustering method. It utilises fuzzy partitioning to allocate the data points to the clusters in terms of a membership degree ranging from 0 to 1. Membership values close to 1 indicate a high degree of similarity between the observation and a cluster, and vice versa15. Matrix factorisation is a common method of data dimension reduction and recently has been used in a number of clustering approaches. Most popular clustering methods, such as K-means and FCM, can be reformulated as matrix factorisation problems16,17,18.
Simplex-structured matrix factorisation (SSMF) is a generalisation of non-negative matrix factorisation (NMF)19,20. SSMF decomposes a given data matrix into a prototype matrix and a soft membership matrix. The prototype matrix characterises the measurements from each of the clusters. The soft membership matrix contains non-negative values that sum to one, which record the soft clustering of each observation. SSMF represents each observation as a weighted combination of the cluster prototypes using the soft cluster membership values.
In this paper, we propose SSMF as a suitable method for soft clustering of metabolomic data. An overview of SSMF as a soft clustering method is introduced. Methods for selecting the number of clusters, uncertainties in the cluster prototypes and the soft membership values are provided. The soft adjusted Rand index and Shannon diversity index are introduced to study the soft clustering structure. Furthermore, SSMF will be illustrated using a simulation study and a specific metabolomic dataset. A discussion and conclusions are also provided.
Methods
Simplex-structured matrix factorisation
Let \(X \in \mathbb {R}^{n \times p}\) be a data matrix with n observations and p variables; each row of X corresponds to an observation in the p-dimensional space. Simplex-structured matrix factorisation (SSMF) finds a matrix \(H \in \mathbb {R}^{n \times k}\) and a matrix \(W \in \mathbb {R}^{k \times p}\) such that
where \(k \ll \text {min}(n,p)\) is the rank of the factorisation (k is also the number of clusters). H is the soft membership matrix where the \(i^{th}\) row, \(h_i \in \mathbb {R}^{k}\), gives the cluster membership of the \(i^{th}\) observation; the entries of \(h_i\) are positive and sum to one. W is the prototype matrix where the \(r^{th}\) row, \(w_r \in \mathbb {R}^{p}\), records the cluster prototype that characterises cluster r.
The problem of finding the matrix factorisation (1) is solved by minimising the the residual sum of squares (RSS),
such that \(\sum _{r=1}^k h_{ir}=1\) and \(h_{ir}\ge 0\) (unit simplex).
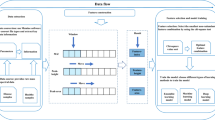
Illustration of the SSMF. (A) The decomposition of \(x_{i}\). Each value in the observation \(x_{i}\) is represented as a linear combination of the columns of W given the weights by \(h_{i}\) with unit simplex constraint. (B) An optimised convex hull of cluster prototypes \(w_{kj}\) for variable j was found to represent \(x_{ij}\) given the unit simplex weights \(h_{ik}\). The RSS is minimised during optimisation across all variables.
The SSMF approximates each row of X using a linear combination of the columns of W, where the coefficients are given by corresponding row of H (Fig. 1A); that is, each point is represented in the convex hull of the cluster prototypes (Fig. 1B):
An algorithm for minimising the RSS (2) is implemented using an iterative process that alternates between determining the optimal W for a given H (least squares) and determining the optimal H for a given W (convex least squares).
Simplex-structured matrix factorisation algorithm
- Step 0::
-
For a given a number of clusters (k); k is also the rank of the factorisation. Initialise the membership matrix \(\widehat{H}\), where each row is a random vector with a unit simplex constraint.
- Step 1::
-
Find the best \(\widehat{W}\) for the given \(\widehat{H}\). Solve p linear least squares problems (\(j=1, 2, \ldots ,p\)):
$$\begin{aligned} \hat{w}_j=\mathop {\textrm{argmin}}_{w_j}\Vert x_{j} - H w_{j} \Vert ^2. \end{aligned}$$This optimisation is completed using linear regression methods.
- Step 2::
-
Find the best \(\widehat{H}\) for the given value of \(\widehat{W}\). Solve n convex least squares problems (\(i=1, 2, \ldots ,n\)):
$$\begin{aligned} \hat{h}_{i}=\mathop {\textrm{argmin}}_{h_{i}}\Vert x_{i} - h_{i} \widehat{W} \Vert ^2, \end{aligned}$$such that \(h_{ir} \ge 0\) and \(\sum _{r=1}^{k} h_{ir} = 1\). This optimisation is achieved using quadratic programming methods21.
- Step 3::
-
Repeat Step 1 and Step 2 until the RSS reduction is sufficiently small (less than 0.01) or the number of maximum iterations (50 iterations) is reached.
Model selection
In many cases of cluster analysis, there is not a single “best” definition of a cluster and no exact rule for selecting the correct number of clusters22,23. A popular method to determining the number of cluster (k) involves running the algorithm with various values of k and identifying an “elbow” point where the residual sum of squares (RSS) begins to decrease at a much slower rate. However, this method is difficult to apply in practice when the data are noisy or lack a clear cluster structure.
The gap statistic offers a statistical method to formalise the heuristic elbow approach24. In this paper, the gap method selects the value of k with the biggest difference between the original RSS and the RSS under the appropriate null reference distribution of the data, which is defined to be
where B is the number of samples from the reference distribution, \(\textrm{RSS}^*_{kb}\) is the residual sum of squares for the \(b^{th}\) sample from the reference distribution fitted the SSMF model using k clusters and \(\textrm{RSS}_{k}\) is the residual sum of squares for the original data X fitted the model using the same k. The estimated gap suggests the number of cluster (\(\hat{k}\)) using
where \(s_{k+1}=sd_k \sqrt{1+\frac{1}{B}}\) is standard error, \(sd_k=\sqrt{ \frac{1}{B} \sum _{b=1}^{B} [\log (\textrm{RSS}^*_{kb})-\bar{l}]^2}\) is standard deviation using \(\bar{l} = \frac{1}{B} \sum _{b=1}^{B} \log (\textrm{RSS}^*_{kb})\).
Bootstrap resampling
Confidence intervals (CIs) are important for indicating the uncertainty of estimating a population parameter using a given sample25 and can be used to evaluate significant differences among the cluster prototypes. Estimating the CIs for the cluster prototypes is achieved through the bootstrap algorithm:
- Step 1::
-
Create bootstrap samples of size n by sampling from the dataset with replacement and repeat this step M times. The \(m^{th}\) bootstrap sample is denoted as
$$\begin{aligned} X^{*(m)}=(x_1^{*(m)}, x_2^{*(m)},\ldots ,x_n^{*(m)}), \end{aligned}$$where each \(x_i^{*(m)}\) is a random sample (with replacement) from the dataset.
- Step 2::
-
Apply the SSMF algorithm to each bootstrap sample and calculate the \(m^{th}\) bootstrap replicate of the prototype matrix, which is denoted as \(W^{*(m)}\)
- Step 3::
-
The estimated standard deviation of M bootstrap replicates is calculated by
$$\begin{aligned} sd(W^*) = \sqrt{\frac{1}{M-1} \sum _{m=1}^{M} [W^{*(m)}-\overline{W}^{*}]^2}, \end{aligned}$$where \(\overline{W}^{*}=\frac{1}{M} \sum _{m=1}^{M} W^{*(m)}\).
Therefore, the 95% CIs for the cluster prototypes are given as,
where \(t_{(0.025, M-1)}\) and \(t_{(0.975, M-1)}\) are the quantiles of the student t distribution with \((M-1)\) degrees of freedom.
Soft adjusted Rand index
The soft adjusted Rand index (sARI) is proposed to compare the performance of both classification and clustering methods26. Given a known and an estimated soft membership matrix H and \(\widehat{H}\), \(p_{rci} = h_{ri} \cdot \hat{h}_{ci}\) is the product of the probabilities of assigning observation i to the \(r^{th}\) cluster in H and to the \(c^{th}\) cluster in \(\widehat{H}\). The soft pairwise agreement is then calculated by \(p_{rc.} = \sum _{i=1}^n p_{rci}\). Therefore, the sARI for H and \(\widehat{H}\) is defined as
where \(\Lambda _{rc} = \sum _{r} \frac{\Gamma (p_{r..}+1)}{\Gamma (p_{r..}-1)} + \sum _{c} \frac{\Gamma (p_{.c.}+1)}{\Gamma (p_{.c.}-1)}\), \(p_{r..} = \sum _{c=1}^C \sum _{i=1}^n p_{rci}\) and \(p_{.c.} = \sum _{r=1}^R \sum _{i=1}^n p_{rci}\). A larger sARI illustrates the more similarity between the two soft partitions.
Shannon diversity index
Shannon diversity index27 is proposed to evaluate the average uncertainty in the soft clustering partition. Choosing base 2 logarithm, the entropy of \(h_i\) is defined as (4),
where the value of \(h_{ir} \textrm{log}_2 (h_{ir})\) is taken to be 0 when \(h_{ir}=0\)27.
We use the Shannon Diversity index, which is given as \(2^{\textrm{E}(h_{i})}\), to indicate the effective number of fuzzy clusters that observation i is assigned to and it takes values from 1 to k. The maximum of \(2^{\textrm{E}(h_{i})}\) is achieved when \(h_{i1} = \cdots = h_{ik} = 1/k\). In other words, if an observation is spread equally across k clusters, \(2^{\textrm{E}(h_{i})} = k\) which indicates membership across k clusters. Otherwise, the observation has membership in r cluster(s) when the value of \(2^{\textrm{E}(h_{i})}\) is between r and \(r+1\), for \(r=1, 2, \dots , k-1\).
Simulated metabolomic dataset
A simulated dataset was created by the product of a soft membership matrix H and a prototype matrix W, both of them were generated using designed information. A Dirichlet distribution was used to generate values for the soft membership matrix H; the Dirichlet distribution is a widely studied distribution on the unit simplex28, providing a flexible approach for modelling contributions from various clusters. Specifically, four independent vectors were simulated from a gamma distribution with a scale parameter of 1 and shape parameters are 0.5, 0.8, 0.3 and 1.2. To better align the simulated data with real-world scenarios, each gamma value was randomly replaced by zero with probability 0.1. Then, each vector was then normalised so that the values summed to one.29.
Runtimes of the algorithm including the SSMF, the gap statistic and the bootstrap (Table S1 in supplementary) revealed the runtimes increased with increasing the number of observations. To save time and align the dimensions of the simulated data with the structure of the metabolomic dataset, the prototype matrix W was designed with four prototypes for 138 variables (Fig. 2), while the membership matrix H was simulated for 175 observations.
Simulated prototypes. The simulated dataset is created based on 4 prototypes. The first prototype has all values as 1; values of the second prototype increase from 0.1 to 0.8; values of the third prototype start at 0.25, then they increase to 0.9 and drop to 0.4; values of the last prototype decrease from 0.7 to 0.2.
Metabolomic dataset
The metabolomic dataset has 177 observations with 138 metabolites and 3 covariates: age, sex, and body mass index (BMI)2. Ethical approval was granted by University College Dublin Sciences Human Research Ethics Committee (LS-16-91-Gibbons-Brennan). Written informed consent was obtained. Among the 177 observations, 52 were men and 125 were women. The average age and BMI of the observations were 35 years (±13 years) and 24 \(\mathrm {kg/m}^{2}\) (±3.0 \(\mathrm {kg/m}^{2}\)), respectively (Table 1). A total of 138 variables from 7 metabolite categories (Table 2), including 20 amino acids, 11 biogenic amines, 10 acylcarnitines, 10 lysophosphatidylcholines, 72 phosphatidylcholines, 14 sphingomyelins and 1 hexose were presented in the dataset. For analysis of the dataset, Min-Max scaling was applied to transform all values into the interval [0, 1]. Dirichlet regression https://doi.org/10.57938/ad3142d3-2fcd-4c37-aec6-8e0bd7d077e1 was used to examine the association between covariates and soft memberships.
Results
Application of SSMF to simulated metabolomic data
To evaluate the model’s performance, the SSMF model was fitted to the simulated metabolomic dataset. The gap statistic (Fig. 3A) suggested using a model with 4 prototypes and the four soft clusters are shown in Fig. 3B, where the area of colour in the pie charts depicts the soft membership values for each observation. For the given SSMF, the 4 estimated cluster prototypes for the 138 simulated metabolites are shown in Fig. 3C. These estimates also show the 95% confidence intervals and the results reproduced the known prototypes in Fig. 2.
The soft adjusted Rand index (sARI), calculated by equation (3), was 0.344. In this simulation study, the upper bound of the sARI was calculated using \(\textrm{sARI}(H, H)=0.382\); so the estimated soft clustering was close to the true soft clustering. In a study of sARI26, model-based clustering solutions for hard and soft clustering were compared, revealing that sARI tends to be smaller than ARI. This is because the overconfidence in hard correct classifications generally outweighs the overconfidence in hard misclassifications. In addition, the Shannon diversity index was calculated and the average values of \(2^{\textrm{E}(h_{i \cdot })}\) were 2.4 for the known soft memberships and 2.7 for the estimated soft memberships. This means most observations clustered by SSMF had fuzzy clustering across two clusters, which was consistent with the known information. The application of K-means to the simulated data was also performed using the same value of k. K-means gave the \(\textrm{RSS}_\textrm{kmeans}=513.8\) while \(\textrm{RSS}_\textrm{SSMF}=286.7\). This indicates that the SSMF represented the data more accurately when using the same number of clusters.
Application of SSMF to plasma metabolomic data
Results of fitting SSMF model to the metabolomic dataset. (A) Results of the gap statistic for choosing a suitable number of prototypes. (B) A two-dimensional PCA projected scatter pie chart representation of the clusters. (C) The estimated prototypes with the confidence intervals. The vertical grey dashed lines separate 7 categories of variables corresponding to Table 2.
For the metabolomic data, the gap statistic suggested six soft clusters (Fig. 4). However, it is worthwhile noting that at \(k=4\), \(\textrm{Gap}(k) \approx \textrm{Gap}(k+1) - s_{k+1}\), which suggested that this could also be an appropriate number of prototypes. Therefore, the smaller number \(k=4\) was selected for the clustering. A K-means clustering using the same value of k achieved an \(\textrm{RSS}_\textrm{kmeans}\) of 577.1 which is greater than the results for SSMF, where \(\textrm{RSS}_\textrm{SSMF}=439.7\).
The resulting four soft clusters are shown in Fig. 4B, with the majority of observations having the highest memberships in clusters 1, 2 or 4. A small number of observations had the highest memberships in cluster 3. The Shannon diversity index indicated that most of the observations had fuzzy membership to three clusters. The four cluster prototypes are displayed in the Fig. 4C. The grey dashed lines separate the variables into 7 categories shown in the Table 2. The prototypes of clusters 1, 2 and 4 are separated well (Fig. 4C). The first and second prototypes had similarly high values in amino acids, biogenic amines and lysophosphatidylcholines metabolites but they separated into different levels in categories phosphatidylcholines, sphingomyelins and hexose metabolites. Cluster prototype 4 had the lowest level in amino acids, biogenic amines, biogenic amines and lysophosphatidylcholines metabolites but its levels of phosphatidylcholines and sphingomyelins metabolites were located between prototype 1 and 2. Cluster prototype 3 had high level of acylcarnitines metabolites. Specifically, there are 9 observations having the largest memberships in cluster 3 (Fig. 5) and showing their metabolite levels peak for acylcarnitines metabolites. The 95% CIs of these prototypes are shown in supplementary Table S2. Non-overlapping confidence intervals (CIs) indicate that the estimated prototypes are significantly different. For example, seven acylcarnitine metabolites (C4, C6C41DC, C8, C10, C101, C141, and C181) show significant differences between prototype 3 and prototypes 1, 2, and 4.
An examination of the covariates sex, age and BMI using Dirichlet regression revealed that membership varied mainly across sex and age (Fig. 6A–C). A significantly higher proportion of men exhibited greater membership in cluster 2 (\(p < 0.001\)). Older observations (age \(> 50\) years) tended to have higher membership in cluster 4 (\(p < 0.001\)), while younger observations (age \(< 30\) years) showed higher membership in cluster 2 (\(p < 0.001\)). The observations generally had low membership values in cluster 3. However, the observations with lower BMI (<25 \(\mathsf {kg/m}^\textsf{2}\)) showed significantly greater membership (\(p < 0.001\)) in this cluster compared to those who with higher BMI. In contrast, BMI had no significant influence on membership values in the other clusters.
Compared with other techniques (Table S3), the SSMF had ability to find more meaningful and interpretable clusters in the dataset. K-means structured the data using two clusters (Fig. S1) with the main differences in the phosphatidylcholines, sphingomyelins and hexose metabolite. It did not find a cluster with higher levels of acylcarnitines, for example, as was found with the SSMF approach (Fig. 5). The NMF approach (Fig. S2) captured more clusters than K-means, but it missed the first prototype that is presented in Fig. 4C.
Discussion
Cluster analysis is widely employed in the field of metabolomics, with hard clustering being the dominant application. Hard clustering methods group the observations into distinct clusters, which are easy for metabolomic researchers to interpret. However, the scope of hard clustering approaches is limited in some metabolomic applications, as it can oversimplify the data structure and impose clusters where the boundaries are not clear. We applied the simplex-structured matrix factorisation (SSMF) to perform soft clustering of metabolomic data.
In this paper, the metabolite levels of observations were not specified by one cluster prototype but instead can be described by combinations of 4 cluster prototypes based on the defined characteristics of each prototype. The membership values in each observation indicate the possibilities that the observation relates to the clusters. An important advantage of this approach is that it better represents the data and enables a certain amount of flexibility. Interestingly for the metabolomics application in this paper use of the soft clustering approach uncovered a prototype with elevated levels of acylcarnitine. The ability to identify individuals with high membership of this prototype is important given the associations between acylcarnitines and a number of health outcomes30,31,32. As reported in12, soft unsupervised clustering avoids preconceived notions present in hard clustering, allowing for the potential discovery of the true underlying data structure and heterogeneity. In a study of type 2 diabetes33, soft clustering was applied to effectively identify four archetypes, representing continuous combinations of dysfunction across five underlying etiological processes in individuals with type 2 diabetes. However, users should be careful when applying soft clustering because overlapping clusters generated through this method might present a drawback. For instance, in healthcare domains, this ambiguity in patient group assignment could hinder effective clinical decision-making.34,35,36.
From a mathematical aspect, K-means clustering is a hard clustering variant of SSMF model when a binary constraint is imposed on membership matrix. By comparing the RSS for the same number of clusters (k), SSMF is able to give a smaller RSS and thus a better model in both the simulation study and the application to the metabolomic data. Consequently, from the perspective of mathematical modelling, soft clustering models are superior to hard clustering models. They allow more diverse information to be extracted from the data, thereby enriching the structure of the original data.
The selection of the number of soft clusters can be subjective and an objective approach the gap statistic has been proposed. While the approach performed well, it is important to acknowledge that there is no exact definition of a ‘cluster’. Previous work applied the method to well-separated data and reported that it easily identified the “correct” number of clusters in simulation studies. However, when the data contains overlaps, the more overlaps there are, the more likely gap statistic will select a single cluster24. Consequently, selection of the appropriate number of clusters is not straightforward. Moreover, it is proposed that the optimal clustering depends on background information and the research purpose or question at hand37. Furthermore, others report that the data alone cannot determine the “optimal” clustering but researcher input is needed38. Therefore, it is important that researchers are explicit about the purpose of the research and how the number of clusters was selected.
The SSMF aims to find a convex combination of prototype vectors that best represent the data and is moderately sensitive to noise, especially outliers. It is suggested to apply data cleaning methods before using the model, particularly for data points that significantly deviate from the rest of the dataset and those with missing or incomplete values.
Conclusion
This study outlines an innovative application of soft clustering in metabolomic data using simplex-structured matrix factorisation. This analysis characterises 177 observations using four soft cluster prototypes combined using the soft cluster membership values. Finally, we have shown that in the soft clustering patterns are impacted by sex and age but not by BMI in this population group.
The code is accessible through the R package named MetabolSSMF.
Data availability
The simulated metabolomic dataset generated and analysed during the current study are available in the MetabolSSMF repository, https://github.com/WenxuanLiu1996/MetabolSSMF/tree/main/data. The plasma metabolomic dataset used and analysed during the current study are available from the corresponding author on reasonable request.
References
Deidda, M., Piras, C., Bassareo, P. P., Cadeddu Dessalvi, C. & Mercuro, G. Metabolomics, a promising approach to translational research in cardiology. IJC Metab. Endocr. https://doi.org/10.1016/j.ijcme.2015.10.001 (2015).
Yin, X., Prendiville, O., McNamara, A. E. & Brennan, L. Targeted metabolomic approach to assess the reproducibility of plasma metabolites over a four month period in a free-living population. J. Proteome Res. https://doi.org/10.1021/acs.jproteome.1c00440 (2022).
Nyamundanda, G., Brennan, L. & Gormley, I. C. Probabilistic principal component analysis for metabolomic data. BMC Bioinform. https://doi.org/10.1186/1471-2105-11-571 (2010).
Chen, Y., Li, E.-M. & Xu, L.-Y. Guide to metabolomics analysis: A bioinformatics workflow. Metabolites https://doi.org/10.3390/metabo12040357 (2022).
Aggarwal, C. C. An introduction to cluster analysis. In Data Clustering (eds Aggarwal, C. C. et al.) 1–27 (CRC Press, 2014). https://doi.org/10.1201/9781315373515-1.
Scitovski, R., Sabo, K., Martínez-Álvarez, F. & Ungar, S. Introduction. In Cluster Analysis and Applications (eds Scitovski, R. et al.) 1–3 (Springer, 2021). https://doi.org/10.1007/978-3-030-74552-3_1.
Ferraro, M. B. & Giordani, P. Soft clustering. WIREs Comput. Stat. https://doi.org/10.1002/wics.1480 (2020).
Bora, D. J. & Gupta, D. A. K. A comparative study between fuzzy clustering algorithm and hard clustering algorithm. Int. J. Comput. Trends Technol. https://doi.org/10.14445/22312803/IJCTT-V10P119 (2014).
Yuan, C. & Yang, H. Research on K-Value Selection Method of K-Means Clustering Algorithm https://doi.org/10.3390/j2020016 (2019).
Arora, P., Deepali, & Varshney, S. Analysis of K-means and K-medoids algorithm for big data. Procedia Comput. Sci. https://doi.org/10.1016/j.procs.2016.02.095 (2016).
Azar, A. T., El-Said, S. A. & Hassanien, A. E. Fuzzy and hard clustering analysis for thyroid disease. Comput. Methods Programs Biomed. https://doi.org/10.1016/j.cmpb.2013.01.002 (2013).
Bayoumi, R. et al. Etiologies underlying subtypes of long-standing type 2 diabetes. PLoS ONE https://doi.org/10.1371/journal.pone.0304036 (2024).
Zhang, X. et al. Use of plasma metabolomics to analyze phenotype-genotype relationships in young hypercholesterolemic females. J. Lipid Res. https://doi.org/10.1194/jlr.M088930 (2018).
Galal, A., Talal, M. & Moustafa, A. Applications of machine learning in metabolomics: Disease modeling and classification. Front. Genet. https://doi.org/10.3389/fgene.2022.1017340 (2022).
Bezdek, J. C., Ehrlich, R. & Full, W. FCM: The fuzzy c-means clustering algorithm. Comput. Geosci. https://doi.org/10.1016/0098-3004(84)90020-7 (1984).
Carmona-Saez, P., Pascual-Marqui, R. D., Tirado, F., Carazo, J. M. & Pascual-Montano, A. Biclustering of gene expression data by non-smooth non-negative matrix factorization. BMC Bioinform. https://doi.org/10.1186/1471-2105-7-78 (2006).
Fogel, P. et al. Applications of a novel clustering approach using non-negative matrix factorization to environmental research in public health. Int. J. Environ. Res. Public Health https://doi.org/10.3390/ijerph13050509 (2016).
Binesh, N. & Rezghi, M. Fuzzy clustering in community detection based on nonnegative matrix factorization with two novel evaluation criteria. Appl. Soft Comput. https://doi.org/10.1016/j.asoc.2016.12.019 (2018).
Abdolali, M. & Gillis, N. Simplex-structured matrix factorization: Sparsity-based identifiability and provably correct algorithms. SIAM J. Math. Data Sci. https://doi.org/10.1137/20M1354982 (2021).
Gillis, N. & Kumar, A. Exact and heuristic algorithms for semi-nonnegative matrix factorization. SIAM J. Matrix Anal. Appl. https://doi.org/10.1137/140993272 (2015).
Lawson, C. L. & Hanson, R. J. 23. Linear Least Squares with Linear Inequality Constraints, 158–173 https://doi.org/10.1137/1.9781611971217.ch23 (1995).
Schubert, E. Stop using the elbow criterion for k-means and how to choose the number of clusters instead. ACM SIGKDD Explor. Newsl. https://doi.org/10.1145/3606274.3606278 (2023).
Eugster, M. J. A. & Leisch, F. From spider-man to hero—Archetypal analysis in R. J. Stat. Softw. (8) https://doi.org/10.18637/jss.v030.i08 (2009).
Tibshirani, R., Walther, G. & Hastie, T. Estimating the number of clusters in a data set via the gap statistic. J. R. Stat. Soc. Ser. B Stat Methodol. https://doi.org/10.1111/1467-9868.00293 (2001).
Severiano, A., Carriço, J. A., Robinson, D. A., Ramirez, M. & Pinto, F. R. Evaluation of jackknife and bootstrap for defining confidence intervals for pairwise agreement measures. PLoS ONE https://doi.org/10.1371/journal.pone.0019539 (2011).
Flynt, A., Dean, N. & Nugent, R. sARI: A soft agreement measure for class partitions incorporating assignment probabilities. Adv. Data Anal. Classif. https://doi.org/10.1007/s11634-018-0346-x (2019).
Jiroušek, R. & Shenoy, P. P. A new definition of entropy of belief functions in the Dempster-Shafer theory. Int. J. Approx. Reason. https://doi.org/10.1016/j.ijar.2017.10.010 (2018).
Ongaro, A. & Migliorati, S. A generalization of the Dirichlet distribution. J. Multivar. Anal. https://doi.org/10.1016/j.jmva.2012.07.007 (2013).
Lange, K. Applications of the Dirichlet distribution to forensic match probabilities. Genetica https://doi.org/10.1007/BF01441156 (1995).
Gander, J. et al. Metabolic impairment in coronary artery disease: Elevated serum acylcarnitines under the spotlights. Front. Cardiovasc. Med. https://doi.org/10.3389/fcvm.2021.792350 (2021).
Luo, H. & Zhu, Z. Serum acylcarnitines levels as a potential predictor for gestational diabetes: A systematic review and meta-analysis. Front. Public Health https://doi.org/10.3389/fpubh.2023.1217237 (2023).
Dambrova, M. et al. Acylcarnitines: Nomenclature, biomarkers, therapeutic potential, drug targets, and clinical trials. Pharmacol. Rev. 74(3), 506–551. https://doi.org/10.1124/pharmrev.121.000408 (2022).
...Wesolowska-Andersen, A. et al. Four groups of type 2 diabetes contribute to the etiological and clinical heterogeneity in newly diagnosed individuals: An IMI DIRECT study. Cell Rep. Med. https://doi.org/10.1016/j.xcrm.2021.100477 (2022).
van Smeden, M., Harrell, F. E. & Dahly, D. L. Novel diabetes subgroups. Lancet Diabetes Endocrinol. https://doi.org/10.1016/S2213-8587(18)30124-4 (2018).
Dennis, J. M., Shields, B. M., Henley, W. E., Jones, A. G. & Hattersley, A. T. Disease progression and treatment response in data-driven subgroups of type 2 diabetes compared with models based on simple clinical features: an analysis using clinical trial data. Lancet Diabetes Endocrinol. https://doi.org/10.1016/S2213-8587(19)30087-7 (2019).
Dennis, J. M. Precision medicine in type 2 diabetes: Using individualized prediction models to optimize selection of treatment. Diabetes https://doi.org/10.2337/dbi20-0002 (2020).
Hennig, C. What are the true clusters?. Pattern Recogn. Lett. https://doi.org/10.1016/j.patrec.2015.04.009 (2015).
Akhanli, S. E. & Hennig, C. Comparing clusterings and numbers of clusters by aggregation of calibrated clustering validity indexes. Stat. Comput. https://doi.org/10.1007/s11222-020-09958-2 (2020).
Funding
This publication has emanated from research conducted with the financial support of Science Foundation Ireland under Grant number 18/CRT/6049. The opinions, findings and conclusions or recommendations expressed in this material are those of the author(s) and do not necessarily reflect the views of the Science Foundation Ireland.
Author information
Authors and Affiliations
Contributions
W.L. conducted modelling and research, analysed data and drafted the manuscript; T.B.M. and L.B. designed and conducted research, reviewed and edited the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Ethical approval was granted by University College Dublin Sciences Human Research Ethics Committee (LS-16-91-Gibbons-Brennan). Written informed consent was obtained.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liu, W., Murphy, T.B. & Brennan, L. Simplex-structured matrix factorisation: application of soft clustering to metabolomic data. Sci Rep 15, 17817 (2025). https://doi.org/10.1038/s41598-025-02361-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-02361-9