Abstract
To develop and validate a reliable and noninvasive prediction model for early identification of lobar pneumonia in children with mycoplasma pneumoniae (MP) infection. This retrospective study included 209 pediatric patients diagnosed with MP infection between January and December 2023 at the Children’s Hospital of Kunming Medical University. Clinical symptoms, laboratory findings, and imaging data were collected. Univariate analysis and receiver-operating-characteristic (ROC) curve analysis were performed to assess the predictive value of variables. least absolute shrinkage and selection operator (LASSO) regression was used to identify key predictive variables, which were then integrated into a logistic regression model. A nomogram was developed based on the model, and its performance was validated using ROC curves, calibration plots, and decision curve analysis (DCA). The predictive nomogram identified albumin (ALB), lactate dehydrogenase (LDH), presence of rales, and co-bacterial infection as significant predictors of lobar pneumonia in children with MP infection. The model demonstrated strong predictive accuracy, with area under the curve (AUC) values of 0.8463 (95% CI: 0.7793–0.9134) in the training set and 0.810 (95% CI: 0.742–0.8773) in the validation set. Calibration plots showed good agreement between predicted and actual outcomes, and DCA confirmed the model’s clinical utility, demonstrating greater net benefit compared to all-intervention or no-intervention strategies. This study presents a novel, accurate, and clinically useful nomogram for predicting lobar pneumonia in children with MP infection based on routine clinical and laboratory data. The model provides a non-invasive approach to early diagnosis, potentially reducing the need for repeated imaging and associated radiation exposure.
Similar content being viewed by others
Introduction
Mycoplasma pneumoniae pneumonia (MPP) is a common respiratory disease caused by Mycoplasma pneumoniae (MP) infection1,2,3. While MPP is generally considered to have a self-limiting course with a favorable prognosis, a subset of cases can develop into severe MPP (SMPP), potentially leading to systemic involvement and extrapulmonary organ damage4,5,6. Factors such as the emergence of macrolide-resistant MP strains, co-infections, and abnormal immune responses have contributed to an increasing number of cases of refractory mycoplasma pneumoniae pneumonia (RMPP)6,7,8,9. Lobar pneumonia is a frequent pulmonary complication following MP infection. Some children with MPP presenting with lobar pneumonia on imaging may progress to RMPP, resulting in complications such as atelectasis and necrotizing pneumonia10,11. Studies have identified lobar pneumonia as a risk factor for RMPP, often associated with macrolide-resistant MP infections12,13. Currently, macrolide-resistant MP infections account for up to 90%, rendering antibiotics like azithromycin less effective, necessitating the use of tetracycline or quinolone antibiotics2,14.
Once lobar pneumonia forms, mucous plugs obstructing the bronchi may require bronchoscopic intervention and lavage15,16, which carry risks of bronchial mucosal injury, bleeding, and anesthesia-related complications. Early identification of lobar pneumonia allows for the timely use of sensitive antibiotics and corticosteroids, which can shorten the disease course and reduce complications. The identification of lobar pneumonia typically relies on chest imaging, but in the early stages of MP infection, radiological abnormalities may be minimal. As the disease progresses, findings such as lobar consolidation or multi-segment atelectasis may become evident, necessitating repeated imaging. However, repeated imaging poses a risk of radiation exposure, which can increase the likelihood of secondary hematological malignancies17,18.
While many models for predicting SMPP and RMPP exist globally19,20, few address the prediction of lobar pneumonia resulting from MP infection. Therefore, we aim to establish a predictive model based on readily accessible clinical symptoms, signs, and laboratory findings to provide evidence-based guidance for the prevention and treatment of lobar pneumonia in children with MP infection.
Materials and methods
Study population
This study collected clinical data from 209 children diagnosed with MMP were collected at the Children’s Hospital of Kunming Medical University from January 2023 to December 2023. The study protocol received approval from the Ethics Review Committee of the Children’s Hospital of Kunming Medical University.
Inclusion criteria: Meeting the diagnostic standards for MP infection as outlined in the “Guidelines for the Diagnosis and Treatment of MPP in Children (2023 Edition)”; (1) Presenting with respiratory symptoms such as cough and fever; (2) A single serum MP antibody titer exceeding 1:160, or a positive MP DNA-PCR result from nasopharyngeal aspirate, bronchoalveolar lavage fluid, or pleural effusion. (3) Chest CT showed lobar pneumonia.
Exclusion criteria: (1) Children with severe underlying pulmonary diseases such as bronchopulmonary dysplasia, bronchiolitis obliterans, or pulmonary tuberculosis; (2) Children with other serious systemic conditions, including severe congenital heart disease or severe immune disorders; (3) Children with incomplete medical records.
Based on chest imaging results obtained after admission, lobar pneumonia was identified if chest radiographs or chest computed tomography (CT) revealed uniform consolidation in one or more lung segments/lobes. Patients were divided into the lobar pneumonia group (n = 96) and the general group (n = 113) (see Fig. 1).
Data extraction
Basic patient information, clinical manifestations, laboratory test results, and chest imaging data (from X-rays or CT scans) were collected from the hospital’s electronic medical record system. Laboratory tests included the initial white blood cell (WBC) count, hemoglobin (Hb), platelet count (PLT), C-reactive protein (CRP), procalcitonin (PCT), and percentage of neutrophils (N) on admission. Liver function including aspartate aminotransferase (AST), lactate dehydrogenase (LDH), and alanine aminotransferase (ALT), were also recorded.
Statistical analysis
Statistical analyses were performed using R software (version 4.2.2) and SPSS software (version 25). Continuous variables with normal distributions were compared using the t-test and presented as mean ± standard deviation. The t-test was selected because it is suitable for comparing the means of two independent groups when data are normally distributed. Non-normally distributed continuous variables were analyzed using the Mann-Whitney test and presented as the median with interquartile range [M (P25, P75)]. This test was chosen because it is more appropriate for skewed data and provides a robust comparison between groups. Categorical variables were evaluated using the χ2 test or Fisher’s exact test and expressed as counts (n) and percentages (%). The χ2 test was applied when all expected frequencies were ≥ 5, while Fisher’s exact test was used for small sample sizes to ensure statistical validity. Univariate analysis was initially conducted to assess intergroup differences for each variable. Subsequently, the predictive value of each continuous variable for lobar pneumonia caused by MP infection was analyzed using ROC curves, and heatmap was generated to evaluate correlations among continuous variables.
All samples were randomly divided into training and validation sets in a 7:3 ratio using the ‘caret’ package in R software. To construct a predictive model while addressing multicollinearity, a least absolute shrinkage and selection operator (LASSO) regression model was applied in the training set. LASSO regression performs variable selection by applying L1 regularization to shrink the regression coefficients. First, we plotted the LASSO coefficient profile for 30 candidate variables and compared it with the log of the penalty coefficient (log(λ)) to assess the importance of each variable. To select the optimal penalty coefficient λ, we used a ten-fold cross-validation method, determining the optimal λ value based on the criterion of minimizing binomial deviance. Specifically, the binomial deviance curve was plotted as a function of log(λ), with a vertical dashed line indicating the 1 standard error rule. Using this criterion, we selected the variables with non-zero regression coefficients, which were considered key predictors for the model. The training set was used to construct the logistic risk model, while the validation set served as the internal validation cohort to assess the model’s performance.
To improve the interpretability of the model, a nomogram was developed, and the model’s performance was evaluated through the following analyses: ROC curve analysis (to assess discriminative ability), calibration curve analysis (to examine the agreement between the model and observed outcomes), and decision curve analysis (DCA) (to evaluate the model’s net clinical benefit). Statistical significance was defined as P < 0.05.
Results
Clinical characteristics
A total of 209 children with lobar pneumonia caused by MP infection were included, comprising 119 boys (47%) and 90 girls (53%). The median age was 6 years in the general group and 5.54 years in the lobar pneumonia group (see Table 1). First, in order to demonstrate the differences in clinical characteristics (such as symptoms, signs, and laboratory indicators) between the general group and the lobar pneumonia group, we conducted a univariate analysis. This analysis revealed that seven variables—wheeze, shortness of breath, rales, co-bacterial infection, fever, LDH, and ALB—demonstrated significant statistical differences between the two groups (P < 0.05) (see Table 1). ROC curve analysis of these continuous variables revealed that LDH and ALB had high predictive values for lobar pneumonia, with areas under the ROC curve of 0.655 and 0.632, respectively (see Fig. 2a, b). Additionally, correlation calculations and heatmaps indicated significant correlations among many continuous variables (P < 0.05) (see Fig. 2c).
(a) ROC curves of continuous variables for diagnosing lobar pneumonia. The sensitivity and specificity of each parameter are shown, with different colors representing different variables. (b) Bar chart of AUCs for continuous variables in diagnosing lobar pneumonia. Higher AUC values indicate better predictive performance. (c) Heatmap of correlations among continuous variables. This figure presents a correlation matrix depicting the relationships among various clinical parameters. The size and color of each circle represent the strength and direction of the correlation, with red indicating a strong positive correlation and blue representing a strong negative correlation. The color gradient scale on the right ranges from − 1 (strong negative correlation) to + 1 (strong positive correlation).
Variable selection
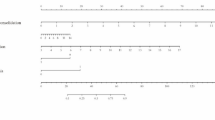
Multicollinearity in regression models occurs when two or more independent variables are strongly correlated, which can adversely affect model performance. Therefore, it is crucial to preprocess the data to detect and address multicollinearity issues before constructing the model to ensure optimal predictive performance. Given the observed correlations among continuous variables, LASSO regression was employed for variable selection to mitigate the impact of multicollinearity. Ultimately, LASSO regression identified ALB, LDH, rales, and co-bacterial infection as significant risk factors for lobar pneumonia caused by MP infection (see Fig. 3a and b).
LASSO regression for predictor selection: (a) LASSO coefficient profiles for the 30 variables, plotted against log(λ). (b) Selection of the optimal penalty coefficient lambda using tenfold cross-validation and minimization criteria, with the binomial deviance curve plotted versus log(λ) and dotted vertical lines representing 1 standard error criteria. Four variables with nonzero coefficients were selected by the 1 standard error criteria.
Nomogram construction
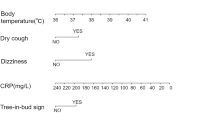
LASSO regression addresses multicollinearity by selecting variables with minimal collinearity. However, in some cases, selected variables may still exhibit some collinearity.To assess multicollinearity among the variables, we utilized the Variance Inflation Factor (VIF). The results revealed that the VIF values for ALB, LDH, rales, and co-bacterial infection were all close to 1, indicating that there was minimal linear relationship among the independent variables. To enhance the model’s interpretability and improve its clinical applicability, we constructed a visual nomogram. A nomogram was developed based on logistic regression analysis, with lobar pneumonia caused by MP infection as the dependent variable, and ALB, LDH, rales, and co-bacterial infection as independent variables. For example, if a patient has an ALB of 32 g/L (25 points), LDH of 390 U/L (20 points), co-bacterial infection (46 points), and no rales in the lungs (22 points), the total score would be 113 points. At the score of 113 on the nomogram, the corresponding predicted probability is approximately 0.692, indicating a 69.2% chance of the child having lobar pneumonia (see Fig. 4a).
(a) Nomogram based on the four optimal predictors for lobar pneumonia caused by mycoplasma pneumoniae infection. For example, if a patient has an ALB of 32 g/L (25 points), LDH of 390 U/L (20 points), co-bacterial infection (46 points), and no rales in the lungs (22 points), the total score would be 113 points. At the score of 113 on the nomogram, the corresponding predicted probability is approximately 0.692, indicating a 69.2% chance of the child having lobar pneumonia (b) ROC curve of the predictive nomogram in the training set. The area under the curve (AUC) is 0.846, indicating good discriminatory ability. (c) Calibration plot of the nomogram in the training set. The calibration plot for the apparent model shows the relationship between predicted and observed probabilities. The dashed line represents the ideal (perfect) calibration, while the solid red line shows the apparent calibration of the model, which is close to the ideal. The bias-corrected calibration curve (orange line) adjusts for overfitting. (d) Calibration plot of the nomogram in the validation set. (e) ROC curve of the predictive nomogram in the validation set, suggesting a slight decrease in performance but still strong model accuracy.
Nomogram accuracy evaluation and validation
The Hosmer-Lemeshow test results indicated a good model fit, with χ2 = 8.852, P = 0.354 in the training set, and χ2 = 7.247, P = 0.510 in the validation set. The area under the ROC curve for the training set was 0.8463 (95% CI: 0.7793–0.9134) (see Fig. 4b), and for the validation set, it was 0.810 (95% CI: 0.742–0.8773) (see Fig. 4e), suggesting good discriminative ability of the model. Calibration lines for both the training and validation sets were close to the ideal 45° line, indicating accurate predictive performance (see Fig. 4c, d).
Clinical utility assessment
Clinical decision analysis of the prediction model (sky-blue curve) demonstrated that the net clinical benefit of the model consistently exceeded that of all intervention or no intervention strategies, underscoring its high clinical value (see Fig. 5a, b). Clinical impact curves for both the training and validation sets showed that when the high-risk threshold was greater than 0.4, the predicted number of patients closely matched the actual number of patients (see Fig. 5c, d).
(a) Decision curve analysis (DCA) for the predictive model in the training set. The net benefit curve for the training set shows the relationship between the high-risk threshold and the net benefit. The curve for the training set (blue) is compared with the “All” and “None” strategies, with the former showing higher net benefits at various threshold levels. (b) Decision curve analysis (DCA) for the predictive model in the validation set. The net benefit curve for the testing set (orange) is similar to the training set, but with slight variations. This demonstrates the model’s performance and its predictive ability in an independent testing set. (c) Clinical impact curve for the predictive model in the training set. The plot shows the number of individuals classified as high risk (blue) at each threshold and the number of those who had the event (orange dashed line). As the high-risk threshold increases, fewer individuals are categorized as high risk, while the number of true events (dashed line) gradually decreases. (d) Clinical impact curve for the predictive model in the validation set.
Discussion
In this study, we developed a predictive model for lobar pneumonia in children with MP infection using clinical and laboratory data, providing a non-invasive approach for early identification. The nomogram (Fig. 3a), which incorporated ALB, LDH, rales, and co-bacterial infection, demonstrated good predictive accuracy and clinical utility. This is a significant step toward reducing reliance on imaging for diagnosis, thereby minimizing radiation exposure in pediatric patients.
CT imaging, while helpful in diagnosing pneumonia, cannot specifically identify MPP, as many other types of pneumonia, such as pneumococcal pneumonia, may also exhibit similar imaging features to lobar pneumonia caused by MP. This creates limitations in early pathogen identification in clinical practice. Typically, outpatient patients with fever and cough can have MP infection detected through throat swabs or blood tests. However, whether the infection has caused severe lobar pneumonia due to MP can only be definitively confirmed by chest CT. However, chest CT carries radiation risks, and not all patients undergo such examinations. Our model, by combining clinical and laboratory data, aims to predict lobar pneumonia caused by MP. The goal of our study is to replace chest CT to some extent with simple clinical indicators for predicting lobar pneumonia. Our findings are consistent with prior research5,7,21,22,23, which has identified key clinical factors associated with SMPP progression, including elevated LDH and ALB levels. For example, similar studies5,7,21 have reported that increased LDH levels correlate with severe pneumonia, highlighting its role as a biomarker for inflammatory and tissue injury responses in pulmonary diseases. Our study extends this understanding by demonstrating that LDH can be effectively used in a predictive model for lobar pneumonia caused by MP. LDH is an enzyme present in the cytoplasm of all tissue cells throughout the body. When cells are damaged or undergo lysis, LDH is released into the bloodstream. Due to this, LDH serves as a valuable biomarker for detecting tissue injury. Elevated LDH reflects cell injury and inflammation, which are commonly seen in more severe pulmonary conditions like lobar pneumonia. Additionally, hypoalbuminemia, frequently found in patients with severe infections[23], has been previously linked to poor clinical outcomes, reinforcing our inclusion of ALB in the nomogram.
The inclusion of clinical signs, such as rales and co-bacterial infection, further strengthened our model. Rales, indicative of fluid or mucus in the airways, are frequently observed in lobar pneumonia due to the obstruction and consolidation of lung segments. The presence of a co-bacterial infection is another critical predictor, as dual infections can complicate the course of MPP and lead to more extensive lung involvement, which is consistent with the results of previous studies21,24.
A unique aspect of our study is the use of LASSO regression to address multicollinearity among continuous variables, ensuring the robustness of the final model (Fig. 2a and b)[25]. LASSO regression effectively reduced the dimensionality of our data, selecting only the most significant variables. This methodological approach enhances the clinical applicability of the nomogram by focusing on a concise set of predictors that are easily measured during routine clinical assessments.
Validation of the model showed strong concordance between predicted and actual outcomes, with AUC values of 0.8463 in the training set and 0.810 in the validation set (Fig. 4b and e). The calibration (Fig. 4c and d) plots indicated good agreement, and decision curve analysis (DCA) (Fig. 5a and b) demonstrated the model’s potential to guide clinical decision-making. By integrating readily available clinical and laboratory data, this model could assist clinicians in identifying high-risk patients early, allowing for timely intervention and potentially reducing the need for invasive diagnostic procedures such as bronchoscopy.
The nomogram developed in this study (Fig. 3a), by integrating easily accessible clinical and laboratory data, can assist healthcare providers in making timely decisions, thereby facilitating early intervention and improving patient prognosis. The non-invasive nature of the model enhances its potential value, particularly in pediatric populations, where minimizing radiation exposure is a critical concern. Moreover, incorporating this predictive model into routine clinical practice could yield substantial benefits, particularly in emergency departments and outpatient settings, where rapid diagnosis and treatment decisions are essential. The ability to accurately predict lobar pneumonia based on simple clinical and laboratory markers can assist clinicians in prioritizing further diagnostic steps, such as imaging or microbiological testing, thereby reducing unnecessary procedures.
One limitation of our study is its retrospective design, which may introduce selection bias. Although our model performed well in internal validation, the limited sample size and single-source data make external validation crucial. To further confirm the model’s generalizability, future research should conduct external validation in larger, more representative multicenter cohorts. Particularly, validating the model’s performance across different regions and diverse population backgrounds is essential to ensure its applicability and robustness in varied patient groups.
In conclusion, the proposed nomogram offers a reliable and clinically useful tool for predicting lobar pneumonia in children with MP infection. By facilitating early identification of at-risk patients, this model has the potential to optimize treatment strategies and reduce the reliance on imaging, ultimately improving patient outcomes. Further research is warranted to validate and refine the model across different clinical settings.
Data availability
The datasets are available from the corresponding author on reasonable request.
References
Jain, S., Williams, D. J., Arnold, S. R., Ampofo, K., Bramley, A. M., Reed, C., Stockmann, C., Anderson, E. J., Grijalva, C. G. & Self, W. H. et al. Community-acquired pneumonia requiring hospitalization among U.S. children. N. Engl. J. Med.372(9), 835–845 (2015).
Gao, L. W. et al. The epidemiology of paediatric Mycoplasma pneumoniae pneumonia in North China: 2006 to 2016. Epidemiol. Infect 147, e192 (2019).
Roh, E. J. et al. Analysis of national surveillance of respiratory pathogens for community-acquired pneumonia in children and adolescents. BMC Infect Dis. 22(1), 330 (2022).
Wang, X. et al. Necrotizing pneumonia caused by refractory Mycoplasma pneumonia pneumonia in children. World J. Pediatr. 14(4), 344–349 (2018).
Shen, W. & Sun, X. Construction of a nomogram for early diagnosis of refractory Mycoplasma pneumoniae pneumonia in children. Transl. Pediatr. 13(7), 1119–1129 (2024).
Shen, F. et al. Development of a nomogram for predicting refractory mycoplasma pneumoniae pneumonia in children. Front Pediatr. 10, 813614 (2022).
Wei, D., Zhao, Y., Zhang, T., Xu, Y. & Guo, W. The role of LDH and ferritin levels as biomarkers for corticosteroid dosage in children with refractory Mycoplasma pneumoniae pneumonia. Respir Res. 25(1), 266 (2024).
Li, M. et al. Recognition of refractory Mycoplasma pneumoniae pneumonia among Myocoplasma pneumoniae pneumonia in hospitalized children: Development and validation of a predictive nomogram model. BMC Pulm Med. 23(1), 383 (2023).
Zhan, X. W. et al. Correlation between Mycoplasma pneumoniae drug resistance and clinical characteristics in bronchoalveolar lavage fluid of children with refractory Mycoplasma pneumoniae pneumonia. Ital. J. Pediatr. 48(1), 190 (2022).
Luo, Y. & Wang, Y. Risk Prediction Model for Necrotizing Pneumonia in Children with Mycoplasma pneumoniae Pneumonia. J Inflamm Res 16, 2079–2087 (2023).
Luo, Y., Dai, J., Tang, G., He, S. & Fu, W. Development and validation of a simple-to-use nomogram for predicting the delayed radiographic recovery in children with mycoplasma pneumoniae pneumonia complicated with atelectasis. J. Investig. Med. 71(7), 722–729 (2023).
Sung, M. et al. Assessment of variables associated with prolonged admission duration in children with Mycoplasma pneumoniae pneumonia. Clin. Respir. J. 16(11), 756–767 (2022).
Choi, Y. J., Chung, E. H., Lee, E., Kim, C. H., Lee, Y. J., Kim, H. B., Kim, B. S., Kim, H. Y., Cho, Y. & Seo, J. H., et al Clinical characteristics of macrolide-refractory mycoplasma pneumoniae pneumonia in Korean Children: A multicenter retrospective study. J. Clin. Med., 11(2) (2022).
Kim, K. et al. Global trends in the proportion of macrolide-resistant mycoplasma pneumoniae infections: A systematic review and meta-analysis. JAMA Netw. Open 5(7), e2220949 (2022).
Zhang, J. et al. Prediction of risk factors of bronchial mucus plugs in children with Mycoplasma pneumoniae pneumonia. BMC Infect Dis. 21(1), 67 (2021).
Xu, X. et al. Nomogram for prediction of bronchial mucus plugs in children with mycoplasma pneumoniae pneumonia. Sci. Rep. 10(1), 4579 (2020).
Mathews, J. D. et al. Cancer risk in 680,000 people exposed to computed tomography scans in childhood or adolescence: Data linkage study of 11 million Australians. BMJ 346, f2360 (2013).
Pearce, M. S. et al. Radiation exposure from CT scans in childhood and subsequent risk of leukaemia and brain tumours: A retrospective cohort study. Lancet 380(9840), 499–505 (2012).
Xie, Q., Zhang, X., Cui, W. & Pang, Y. Construction of a nomogram for identifying refractory mycoplasma pneumoniae pneumonia among macrolide-unresponsive mycoplasma pneumoniae pneumonia in children. J. Inflamm. Res. 15, 6495–6504 (2022).
Liu, G. et al. A lung ultrasound-based nomogram for the prediction of refractory mycoplasma pneumoniae pneumonia in hospitalized children. Infect. Drug Resist. 15, 6343–6355 (2022).
Li, L. et al. Construction and validation of a nomogram model to predict the severity of mycoplasma pneumoniae pneumonia in children. J. Inflamm. Res. 17, 1183–1191 (2024).
Chen, Q., Hu, T., Wu, L. & Chen, L. Clinical features and biomarkers for early prediction of refractory mycoplasma pneumoniae pneumonia in children. Emerg. Med. Int. 2024, 9328177 (2024).
Chang, Q., Chen, H. L., Wu, N. S., Gao, Y. M., Yu, R. & Zhu, W. M. Prediction model for severe mycoplasma pneumoniae pneumonia in pediatric patients by admission laboratory indicators. J. Trop. Pediatr. 68(4) (2022).
Yan, M., Tao, R., Li, S., Xiong, J. & Xiang, J. Clinical characteristics and logistic regression analysis of macrolide-resistant Mycoplasma pneumoniae pneumonia in children. Eur. J. Clin. Microbiol. Infect. Dis. 43(9), 1825–1835 (2024).
Wang, H. et al. Construction of a nomogram with IrAE and clinic character to predict the survival of advanced G/GEJ adenocarcinoma patients undergoing anti-PD-1 treatment. Front Immunol. 15, 1432281 (2024).
Acknowledgements
None.
Funding
This work was supported by the Joint project of Yunnan Science and Technology Department Kunming Medical University (202101AY070001—217); Kunming Health Science and technology personnel training project (2020 (Province)-21); Yunnan key specialty of pediatric infection (Training and education program/Kunming key specialty of pediatric infection; Key Science and Technology Program of Yunnan Province (No.202102AA100019).
Author information
Authors and Affiliations
Contributions
YL and YW conducted data screening and quality evaluation. YW and YW helped design study. YL wrote the main manuscript text. YL and YW made contributions to the revision of the paper. HB, FJ, YG and XY did a lot of data collection work. All authors revised, read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
This study was approved by the ethics committee of Ethics Review Committee of the Children’s Hospital of Kunming Medical University(2023-03-049-K01) and was carried out in accordance with the ethical standards of the Declaration of Helsinki. Informed consent has been waived off by the Ethics Review Committee of the Children’s Hospital of Kunming Medical University as this is a retrospective study.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Luo, Y., Bai, H., jiao, F. et al. Establishment and validation of a predictive model for Lobar pneumonia caused by Mycoplasma pneumoniae infection in children. Sci Rep 15, 22811 (2025). https://doi.org/10.1038/s41598-025-05548-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-05548-2