Abstract
Colorectal cancer (CRC) is the third most common cancer worldwide and the second leading cause of cancer-related deaths. Early detection through non-invasive methods is critical for improving patient outcomes. This study investigates the association between the presence and relative abundance of Fusobacterium nucleatum (Fn) in saliva and CRC, evaluating its potential as a non-invasive biomarker. A systematic search was performed in MEDLINE, Embase, CENTRAL, and Scopus databases on November 25, 2023. Studies analyzing Fn in salivary samples from adults with CRC, colorectal polyps (CRP), or healthy individuals were included. Statistical analyses were performed using random-effects models to calculate pooled odds ratios (OR) and mean differences (MD) with 95% confidence intervals (CI). The risk of bias was assessed using the Quality in Prognosis Studies (QUIPS) tool. Of the 14,200 studies identified, twelve were included in our systematic review. Of these, eight were analyzed by meta-analysis. The results indicated no significant difference in the presence (OR 1.40; 95% CI [0.77; 2.54]; I2 = 0% [0; 71%], p = 0.215) or relative abundance (MD -0.01; 95% CI [-0.13; 0.11]; I2 = 25% [0; 69%], p = 0.851) of salivary Fn among CRC patients, compared to a combined group of CRP and healthy controls. Our findings suggest that the presence and relative abundance of salivary Fn are not associated with CRC.
Similar content being viewed by others
Introduction
Colorectal cancer (CRC) ranks as the third most prevalent cancer globally, accounting for roughly 10% of all cancer cases. It is also the second leading cause of cancer-related deaths worldwide1. Early detection and appropriate treatment are essential for enhancing patient survival and lowering mortality rates2. Colonoscopy prevents early CRC by detecting and removing colorectal polyps3. However, this procedure is invasive, necessitating bowel preparation and intravenous analgesia, and carries risks such as bowel perforation, hemorrhage, and cardiovascular events4. Pain and fear associated with colonoscopy can discourage patients from participating in screenings5. Furthermore, geographic barriers, such as the lack of nearby healthcare facilities and insufficient availability of specialists in remote communities, limit colonoscopy access, leading to late diagnoses6.
Fusobacterium nucleatum (Fn) is an oral pathogenic bacterium associated with periodontitis7. It has been found to accelerate carcinogenesis in various types of cancer, including pancreatic, breast, and CRC8. Fn is involved in both the occurrence and metastasis of CRC through mechanisms such as regulating immune response, virulence factors, oncogenic microRNAs, and DNA damage9. An increased abundance of Fn has been observed in the colorectal tissues of patients with CRC and polyposis, with higher bacterial abundance correlating with lower survival rates10. Whole genome sequencing of paired oral and CRC Fn isolates demonstrated that oral fusobacteria could translocate to CRC by descending the digestive tract or through the bloodstream during transient bacteremia caused by activities such as chewing, daily hygiene, or dental procedures11. Interestingly, Zepeda-Rivera et al. found that the CRC tumor-isolated strains predominantly belong to Fn subspecies animalis (Fna). A specific clade of Fna, such as Fna C2, has been shown to alter the metabolic profile of the tumor microenvironment, promoting adenoma formation and cancer progression12.
Recent studies have suggested that Fn may serve as a potential biomarker for CRC13,14. Moreover, salivary Fn levels have shown promise as a non-invasive diagnostic tool15. One study has demonstrated that salivary Fn detection can differentiate CRC patients from healthy individuals with significant accuracy16. Saliva, an easily accessible biofluid, presents a promising medium for detecting CRC. Non-invasive diagnostic methods, such as detecting Fn in saliva, can significantly improve screening participation rates. Furthermore, understanding the diagnostic potential of salivary Fn may lead to the development of accessible and cost-effective screening tools, particularly in regions with limited access to colonoscopy. Therefore, this systematic review and meta-analysis aimed to explore the association between salivary Fn and CRC.
Results
Search and selection
Our search identified 14,200 studies. After duplicate removal and title and abstract selection (Cohen’s Kappa 0.94), 35 eligible articles remained for full-text analysis (Cohen’s Kappa 0.94). Of these, 23 articles were excluded due to overlapping populations (n = 5), no comparator (n = 2), different outcomes (n = 7), lack of salivary samples (n = 7), or abstract only (n = 2). Our systematic review and meta-analysis included 12 studies, 8 of which were suitable for a quantitative analysis. The selection process is summarized in Fig. 1.
Baseline characteristics of included studies
The 12 studies included 901 CRC patients, 203 CRP patients, and 1,004 healthy individuals from Canada17,18Japan19,20Turkey21Italy22,23Vietnam24China2,25,26and the USA27. ‘Healthy individuals’ refers to individuals without CRC or CRP. Two studies were bicentric21,23and the others were single-center. The studies used non-quantitative polymerase chain reaction (PCR), quantitative PCR (qPCR), next-generation sequencing (NGS), 16S rRNA techniques or a combination of them to detect the presence and relative abundance of salivary Fn. Eight studies excluded individuals who consumed antibiotics either one/three months17,19,22,25,26, or one week21,27,28 prior to sampling, while four studies2,18,23,24 did not report data regarding the use of antibiotics. The baseline characteristics of the included studies are detailed in Table 1.
No difference in the presence of salivary Fn
We found no statistically significant difference in the presence of salivary Fn among CRC patients compared to healthy controls (OR 1.44; 95% CI [0.68; 3.06]; I2 = 0% [0; 79%], p = 0.215). Similarly, we found no significant association in the presence of salivary Fn in CRC compared to the combined group of healthy and CRP controls (OR 1.40; 95% CI [0.77; 2.54]; I2 = 0% [0; 71%], p = 0.215) (Fig. 2).
No difference in the relative abundance of salivary Fn
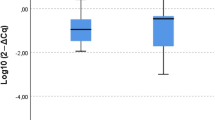
We evaluated five prospective cohort and case-control studies to assess the relative abundance of Fn in the saliva of CRC patients compared to control groups18,19,20,23,24. The analysis indicated no statistically significant difference between CRC patients and healthy controls (MD -0.02; 95% CI [-0.31; 0.27]; I2 = 47% [0; 84%]). Similarly, there was no statistically significant difference between CRC and combined healthy/CRP controls (MD -0.01; 95% CI [-0.13; 0.11]; I2 = 25% [0; 69%], p = 0.851) (Fig. 3).
Subgroup analysis based on the duration of antibiotic therapy prior to sample collection
We conducted a subgroup meta-analysis based on the exclusion criteria according to antibiotic use within one week, one/three months prior to the sample collection. We found significant difference neither in the presence (OR 1.40; 95% CI [0.77; 2.54]; I2 = 0% [0; 71%], p = 0.215) nor in the relative abundance (MD -0.01; 95% CI [-0.13; 0.11]; I2 = 25% [0; 69%], p = 0.851) of salivary Fn between CRC and controls (CRP and healthy individuals) (Figs. 4 and 5).
Studies contained individual patient data (IPD)
We assessed the relative abundance of salivary Fn using density plots to examine its relationship with tumor location (left vs. right side), sex, age, and stage of cancer (I-IV). In all plots, the peak in density was in the lower range of salivary Fn relative abundance; however, we could not evaluate the higher relative abundance range of salivary Fn, as salivary Fn is rarely found in this range (Appendices 2–6).
Studies assessed qualitatively showed varied results
Four studies showed a higher relative abundance of salivary Fn in CRC, CRP, or a combined CRC/CRP than in healthy patients2,17,21,26; three found no significant association22,27,29. However, the quantitative analysis could not evaluate these studies due to the following limitations: (1) different reference genes2,17(2) lack of data22,26,27,29or (3) incomparable units21.
Zhang X et al. detected a higher salivary Fn median relative abundance in CRC than in colorectal adenoma (CRA, a subgroup of CRP), hyperplastic polyp (HP, a subtype of serrated CRP, which is a broader subgroup of CRP), and healthy controls (CRC 1.324 [IQR 0.338–5.134] vs. CRA 0.209 [IQR 0.147–0.688] vs. HP 0.076 [IQR 0.046–0.389] vs. healthy 0.105 [IQR 0.018–0.203]; p < 0.001, in their training cohort, and CRC 2.352 [IQR 1.290–6.743] vs. CRA 0.230 [IQR 0.125–1.310] vs. HP 0.190 [IQR 0.145–0.605] vs. healthy 0.250 [IQR 0.165–0.560]; p < 0.001) in their test cohort. These values were calculated as the ratio of Fn DNA (NusG gene) levels to the geometric mean of GAPDH and TERT gene levels2.
Janati et al. showed a higher salivary Fn median relative quantification in a combined group of CRC/CRP (0.345 [IQR 0.15–0.82]) than in healthy controls (0.12 [IQR 0.05–0.65]), measured as 2−ΔCq using Fn 16 S rRNA as the target gene and MEFE as a reference gene17.
Zhang L et al. found an increased abundance of salivary Fn in CRP patients than in healthy controls (P < 0.05) using the total bacterial 16 S rRNA full-length26.
Wang et al. detected no significant difference in the relative abundance of salivary Fn in CRC (9.91 [IQR 8.799–11.216]) than in healthy controls (10.125 (IQR 9.1584–11.4306); p = 0.527) measured as the difference between total bacteria 16 S rRNA Cq and Fn 16 S rRNA Cq values. Lower Cq values mean a higher relative abundance of the gene25.
Similarly, neither Yang nor Russo found an association between salivary Fn and CRC using the 16 S rRNA target gene23,27. Finally, Guven et al. evaluated the absolute quantity of salivary Fn and found higher mean levels in CRC patients (6.89 ± 1.07 log10 copies/ml) than in healthy controls (6.35 ± 0.78 log10 copies/ml).(p = 0.001)21.
Diagnostic value of salivary Fn in the detection of CRC
Only two studies assessed the diagnostic accuracy of salivary Fn in detecting CRC; thus, we could not conduct a quantitive analysis. Zhang X et al. reported that salivary Fn can detect CRC with high diagnostic accuracy (area under the curve (AUC) = 0.841; 95% CI 0.797–0.879), achieving a sensitivity of 71.5% and a specificity of 82.1% in the training set2. Similarly, high diagnostic accuracy was achieved in the test cohort (AUC = 0.860; 95% CI 0.774–0.922), reaching a sensitivity of 86.7% and a specificity of 67.2% 2. Guven et al. achieved no significant results when assessing the diagnostic accuracy of salivary Fn for CRC diagnosis21.
Risk of bias assessment
The risk of bias was evaluated using the QUIPS tool30. One study demonstrated a potential bias in the study participation domain due to a small sample size22. Three studies showed a low overall risk of bias2,25,27. The other nine studies presented a moderate overall risk of bias, as they did not mention whether blinding was used, demonstrating a potential bias in the outcome measurement domain (Fig. 6). Additionally, they did not adjust the confounding factors, showing a possible bias in the study confounding domain17,18,19,20,21,22,23,24.
Discussion
The major findings of this research revealed no significant difference in the relative abundance of salivary Fn in patients with CRC, compared to CRP, and healthy controls. Not surprisingly, we found no significant difference in the presence of salivary Fn in CRC patients compared to CRP and healthy controls, as Fn is part of the oral microbiome. The diagnostic accuracy of salivary Fn in detecting CRC could not be assessed by the meta-analysis due to the low number of eligible studies.
Previous meta-analyses have investigated the association between CRC and Fn using fecal samples or tissue biopsies. Villar-Ortega et al. showed significantly higher odds for the positivity of Fn in CRC than in CRA tissue biopsies (OR 3.244; 95% CI: 2.359 − 4.462), as well as in healthy controls (OR 4.558; 95% CI: 3.312–6.272)31. Kim et al. indicated that high Fn levels in tissue biopsies of CRC patients are associated with poor overall survival (OS) (Hazard ratio (HR) 1.58; 95% CI 1.28–1.94), disease-free survival (HR 1.76; 95% CI 1.06–2.93), and cancer-specific survival (HR 1.72; 95% CI 1.05–2.83)32. Huangfu et al. found a similar association with OS (HR 1.40; 95% CI: 1.40–1.63)14. Zhang X et al. assessed the diagnostic accuracy of fecal Fn. They found a promising diagnostic accuracy for CRC with a sensitivity of 71% (95% CI 61%-79%) and a specificity of 76% (95% CI 66%‐84%), but a much lower value for the diagnosis of CRA, with a sensitivity of 36% (95% CI 27%‐46%) and a specificity of 73% (95% CI 65%‐79%)33. In light of these meta-analyses and the fact that Fn may originate from the oral cavity and colonize the CRC tissues11we expected to detect similar outcomes using saliva samples.
The results of our study may be attributed to the fact that various factors, including age, gender, diet, and oral hygiene, influence saliva composition, and its microbiome. Consequently, it exhibits significant individual variations34. This presents substantial challenges in acquiring standardized saliva samples, diminishing the comparability and reliability of the outcomes35.
In contrast to the results of this research, two of the included studies showed an association between salivary Fn relative abundance and CRC2,21. One of those studies did not assess essential cofounders, such as periodontal disease, which is associated with Fn. Furthermore, the control group was not evaluated for precancerous lesions using colonoscopy, potentially confounding the bacterial analysis21.
This meta-analysis could not assess the diagnostic accuracy of salivary Fn in detecting CRC. Nonetheless, it is essential to acknowledge the inherent advantages of saliva as a diagnostic medium. Notably, detecting alterations in saliva composition may still serve as a valuable diagnostic tool in other contexts. A significant advantage of using saliva as a diagnostic tool is its non-invasive nature. Unlike blood or tissue biopsies, saliva collection is easy and painless, which improves patient compliance and facilitates repeated sampling over time36. In addition, saliva collection poses less risk to patients and healthcare providers than other sample types. It has a lower risk of exposure to blood-borne pathogens such as HIV or hepatitis viruses, and unlike blood, it does not clot and is, therefore, easier to manipulate37. Saliva testing is generally more cost-effective compared to fecal tests. It requires less specialized equipment and training for collection and processing, making it a more economical option for large-scale screenings and routine diagnostics38.
We could assess the distribution of salivary Fn among different subgroups using density plots (appendices 2–6). Four studies indicated that CRC and CRP patients are more likely to have a low range of salivary Fn relative abundance than healthy controls18,23,24,28. From two studies, it was observed that patients with right-sided tumors are more likely to have a low range of salivary Fn relative abundance than those with left-sided tumors23,24. Interestingly, when analyzing the salivary Fn relative abundance based on sex, we found that males with CRC are more likely to have salivary Fn relative abundance at the low range than women; however, females and males with CRP have the same likelihood of having salivary Fn relative abundance in the low range18,23,24. Assessing the salivary Fn relative abundance among CRC patients at different stages, it was noted that stage I CRC patients are more likely to have Fn relative abundance at the low range than the other stages23,24. Tran et al. analyzed the salivary Fn relative abundance among CRP and CRC at different age groups. However, no correlation between the salivary Fn relative abundance and the patient’s age could be noticed24. Looking at the higher ranges of salivary Fn, it was impossible to assess the relative abundance of the bacteria at these ranges. One possible explanation is that after a certain threshold, Fn can not be detected in saliva.
Abed et al. observed that Fn may be transmitted from the oral cavity to the bloodstream11. This hematogenous spread allows Fn to reach distant organs, including the pancreas, promoting cancer cell proliferation and migration8. Similar to pancreatic cancer, Fn can translocate from the oral cavity to breast tissue through multiple routes, including the mammary-intestinal axis, direct nipple contact, and hematogenous transmission39. In addition, Parhi et al. have indicated that Fn colonizes breast cancer and is secondary to tumor initiation40.
Even though Fn is primarily known for its association with various diseases, particularly CRC and periodontal disease, it is essential to note some of its positive effects. The bridging role of Fn is crucial for developing a stable and diverse oral microbiome, which is vital for maintaining oral health. A balanced oral microbiome can help prevent the overgrowth of pathogenic bacteria and maintain oral homeostasis7. Hsieh et al. have shown the ability of Fn to induce an interferon response in gastric cancer cells, which might affect immune system activation and regulation. Although this immune modulation can contribute to disease progression in some contexts, it may also have potential therapeutic implications in modulating immune responses41.
While Fn plays a significant role in CRC development, other members of the oral microbiota may also be associated with CRC. Guven et al. found increased levels of Streptococcus gallolyticus in the saliva of CRC patients21. Kageyama et al. indicated a higher relative abundance of other bacteria, such as Porphyromonas gingivalis, Streptococcus parasanguinis, Neisseria species, and Actinomyces odontolyticus, in CRC patients. To exclude the influence of periodontal infection on the oral microbiota, they conducted a dental examination and found no significant difference in the periodontal health of CRC patients and healthy controls; however, the small sample size may limit this finding19. Furthermore, Kudra et al. indicated that P. gingivalis increases the risk for CRC and perhaps even stimulates its growth. Several studies imply that P. gingivalis would be beneficial as a non-invasive biomarker in the future. However, since these bacteria induces periodontal disease, which occurs much more frequently than CRC, it may be of little value. Moreover, establishing microbial biomarkers is a complex and challenging task. Various factors must be taken into consideration, including the type of samples (dental plaque, stimulated or unstimulated saliva), analytical methods, CRC stage, age, dental health, and oral hygiene levels42.
The limitations of individual studies should be considered; Zhang. X et al. could not follow up with patients long enough2. Kageyama et al. indicated that the sample size was insufficient to confirm the association independently of the gingival condition19. Guven et al. used the qPCR method rather than NGS. In addition, potential confounders were not assessed21. Furthermore, the observational design of the studies could not explain the causal relationship between salivary Fn and CRC. Chen et al. explained how the structure of a study should be performed to prove causality43. Thus, it is important to note that studies done so far included small sample sizes making difficult to control for the various confounding factors when assessing the association between salivary Fn and CRC.
The limitations of this research were: (1) the use of different DNA extraction kits among studies, which may affect microbiota profiling44. Variations in DNA extraction methodologies and sequence curation steps employed among laboratories - the processes of filtering and processing raw genetic or microbial sequence data - significantly reduce consistency and comparability between studies, potentially leading to confounding results45; (2) the expression of salivary Fn levels was inconsistent among studies, which prevented us from conducting a quantitative assessment in some cases; (3) there was a low number of eligible studies, and about half of the included studies could not be analyzed by the meta-analysis; (4) the small number of studies did not allow us to interpret the diagnostic accuracy of salivary Fn quantitively.
On the other hand, the strengths of this study should also be mentioned. We adhered to the pre-registered protocol and applied a strict methodology. Where possible, data on possible influencing factors, such as patient sex, age, tumor stage, and location, were extracted from each patient’s data. Lastly, studies from different regions with low to moderate risk of bias were included.
Implementing scientific findings in practice is crucial46,47. Therefore, on the basis of the major findings of this research, salivary Fn alone should not be recommended as a biomarker for CRC. Adopting unified methods for detecting salivary Fn is essential to enhance the reliability of the results. Moreover, we encourage researchers to investigate other non-invasive biomarkers that can replace colonoscopy.
Conclusion
The findings of this research show that the presence and relative abundance of salivary Fn are not associated with CRC. Therefore, salivary Fn may not be a reliable biomarker for detecting CRC.
Methods
This systematic review and meta-analysis was conducted using the PRISMA 2020 guidelines48 and followed the Cochrane Handbook49. The study protocol was registered on PROSPERO (registration number CRD42023474939), and we adhered to it.
Eligibility criteria
The PECO and PIRD frameworks were used to address the clinical questions. We included studies if they reported on salivary samples of adults over 18 years old (P) with colorectal cancer (E), polyps (C), or healthy individuals (C). Moreover, articles had to contain data on the presence, quantity, or relative abundance of salivary Fn (O). In addition, we collected data on the diagnostic performance of salivary Fn if they reported on adults over 18 years old (P), salivary Fn (I), histology (R), and CRC (D). Interventional and analytical observational studies were eligible. Reports were excluded if the investigators obtained samples from crevicular fluid or subgingival plaque. Also excluded were studies that contained other gastrointestinal diseases.
Information source
The systematic search was conducted on the MEDLINE (via Pubmed), Embase, Cochrane Central Register of Controlled Trials (CENTRAL), and Scopus databases on November 25, 2023.
Search strategy
The systematic search was performed using a search key consisting of four domains: (1) saliva AND (2) fusobacterium AND (3) colorectal AND (4) cancer. See Appendix 1 for a detailed search key. No restrictions or filtering options were applied.
Selection process
Duplicate articles were automatically and manually removed using the citation manager Endnote 20. Two independent review authors (EG and BZ-S) used the Rayyan Intelligent Systematic Review program50 to select articles by title, abstract, and full texts based on eligibility criteria. A third author (PM) then resolved the conflicts.
Data collection process and data items
Two independent authors (EG and BZ-S) extracted data from included studies based on eligibility criteria. The following data were extracted into a Microsoft Excel spreadsheet (Microsoft 2016, Redmond, WA, USA): first author, publication year, study period, sample size, and population characteristics such as age and sex distribution. We also collected information about the presence/abcsence, quantity, and relative abundance of salivary Fn, the bacterial analysis methods, and the DNA extraction kit type. The relative abundance of salivary Fn was manually extracted from the SRA run selector archive by National Library of Medicine in four studies18,20,23,24whereas, in one study, it was provided by the author19 .
Study risk of bias assessment
Two independent authors (EG and BZ-S) assessed the risk of bias using the Quality In Prognosis Studies (QUIPS) tool30. This tool evaluates six domains: study participation, study attrition, prognostic factor measurement, outcome measurement, study confounding, and statistical analysis and reporting.
Statistical methods
As considerable between-study heterogeneity was assumed in all cases, a random-effects model was used to pool effect sizes. Statistical analysis was performed with R 4.3.251. and package meta 6.5.052.
CRC patients were compared to healthy, or healthy and CRP patients by calculating the mean differences with 95% confidence intervals of Fn relative abundances. Sample sizes, mean and standard deviation values were extracted from the studies to calculate mean differences. Mean differences were calculated by extracting the mean values of the control group from the mean values of the CRC group.
For the presence of salivary Fn, odds ratios with 95% confidence intervals were used as outcome measures. Results are presented as the odds of the CRC group compared to the odds of the same event in the control group. Results were considered statistically significant if the pooled CI did not contain the null effect value. Egger’s test and funnel plots were planned to visualize publication bias if at least ten studies were involved in the analysis. Pooled OR was calculated using the Mantel-Haenszel method53,54. Confidence intervals were created with the Paule-Mandel method55as recommended by Veroniki et al.56.
Pooled mean differences were computed with the inverse variance method. The restricted maximum-likelihood estimator was used with the Q profile method for confidence intervals by Harrer et al.57and Veroniki et al.56. Hartung-Knapp adjustments were also applied58,59 We summarized the results on forest plots. Between-study heterogeneity was assessed with the Higgins and Thompson’s I² statistic60and the Cochrane Q test Harrer et al.57.
Data availability
The datasets used and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Colorectal cancer. https://www.who.int/news-room/fact-sheets/detail/colorectal-cancer (2023).
Zhang, X. et al. Salivary Fusobacterium nucleatum serves as a potential biomarker for colorectal cancer. iScience 25. https://doi.org/10.1016/j.isci.2022.104203 (2022).
Song, M. & Bretthauer, M. Interpreting epidemiologic studies of colonoscopy screening for colorectal cancer prevention: Understanding the mechanisms of action is key. Eur. J. Epidemiol. 38, 929–931. https://doi.org/10.1007/s10654-023-01043-y (2023).
Koffas, A., Papaefthymiou, A., Laskaratos, F. M., Kapsoritakis, A. & Epstein, O. Colon capsule endoscopy in the diagnosis of Colon polyps: who needs a colonoscopy?? Diagnostics (Basel). 12. https://doi.org/10.3390/diagnostics12092093 (2022).
Pontone, S. et al. Do difficulties in emotional processing predict procedure pain and shape the patient’s colonoscopy experience? BMJ Open. 12, e050544. https://doi.org/10.1136/bmjopen-2021-050544 (2022).
Smith, H., Brunet, N., Tessier, A., Boushey, R. & Kuziemsky, C. Barriers to colonoscopy in remote Northern canada: an analysis of cancellations. Int. J. Circumpolar Health. 79, 1816678. https://doi.org/10.1080/22423982.2020.1816678 (2020).
Chen, Y. et al. More than just a periodontal pathogen -the research progress on Fusobacterium nucleatum. Front. Cell. Infect. Microbiol. 12, 815318. https://doi.org/10.3389/fcimb.2022.815318 (2022).
Wang, B. et al. The roles and interactions of Porphyromonas gingivalis and Fusobacterium nucleatum in oral and Gastrointestinal carcinogenesis: A narrative review. Pathogens 13 https://doi.org/10.3390/pathogens13010093 (2024).
Li, R., Shen, J. & Xu, Y. Fusobacterium nucleatum and colorectal Cancer. Infect. Drug Resist. 15, 1115–1120. https://doi.org/10.2147/idr.S357922 (2022).
Gethings-Behncke, C. et al. Fusobacterium nucleatum in the colorectum and its association with Cancer risk and survival: A systematic review and Meta-analysis. Cancer Epidemiol. Biomarkers Prev. 29, 539–548. https://doi.org/10.1158/1055-9965.Epi-18-1295 (2020).
Abed, J. et al. Colon Cancer-Associated Fusobacterium nucleatum May originate from the oral cavity and reach Colon tumors via the circulatory system. Front. Cell. Infect. Microbiol. 10, 400. https://doi.org/10.3389/fcimb.2020.00400 (2020).
Zepeda-Rivera, M. et al. A distinct Fusobacterium nucleatum clade dominates the colorectal cancer niche. Nature 628, 424–432. https://doi.org/10.1038/s41586-024-07182-w (2024).
Wang, Y. et al. Clinicopathological differences of high Fusobacterium nucleatum levels in colorectal cancer: A review and meta-analysis. Front. Microbiol. 13, 945463. https://doi.org/10.3389/fmicb.2022.945463 (2022).
Huangfu, S. C. et al. Clinicopathological and prognostic significance of Fusobacterium nucleatum infection in colorectal cancer: a meta-analysis. J. Cancer. 12, 1583–1591. https://doi.org/10.7150/jca.50111 (2021).
Zhao, R. et al. Improved diagnosis of colorectal cancer using combined biomarkers including Fusobacterium nucleatum, fecal occult blood, transferrin, CEA, CA19-9, gender, and age. Cancer Med. 12, 14636–14645. https://doi.org/10.1002/cam4.6067 (2023).
Flemer, B. et al. The oral microbiota in colorectal cancer is distinctive and predictive. Gut 67, 1454–1463. https://doi.org/10.1136/gutjnl-2017-314814 (2018).
Idrissi Janati, A. et al. Investigation of Fusobacterium Nucleatum in saliva and colorectal mucosa: a pilot study. Sci. Rep. 12, 5622. https://doi.org/10.1038/s41598-022-09587-x (2022).
Nearing, J. T., DeClercq, V. & Langille, M. G. I. Investigating the oral Microbiome in retrospective and prospective cases of prostate, colon, and breast cancer. NPJ Biofilms Microbiomes. 9, 23. https://doi.org/10.1038/s41522-023-00391-7 (2023).
Kageyama, S. et al. Characteristics of the salivary microbiota in patients with various digestive tract cancers. Front. Microbiol. 10 https://doi.org/10.3389/fmicb.2019.01780 (2019).
Uchino, Y. et al. Colorectal Cancer patients have four specific bacterial species in oral and gut microbiota in Common—A metagenomic comparison with healthy subjects. Cancers 13, 3332 (2021).
Guven, D. C. et al. Analysis of Fusobacterium nucleatum and Streptococcus gallolyticus in saliva of colorectal cancer patients. Biomark. Med. 13, 725–735. https://doi.org/10.2217/bmm-2019-0020 (2019).
Russo, E. et al. Preliminary comparison of oral and intestinal human microbiota in patients with colorectal cancer: A pilot study. Front. Microbiol. 8 https://doi.org/10.3389/fmicb.2017.02699 (2018).
Russo, E. et al. From adenoma to CRC stages: the oral-gut Microbiome axis as a source of potential microbial and metabolic biomarkers of malignancy. Neoplasia 40, 100901. https://doi.org/10.1016/j.neo.2023.100901 (2023).
Tran, H. N. H. et al. Tumour microbiomes and Fusobacterium genomics in Vietnamese colorectal cancer patients. NPJ Biofilms Microbiomes. 8, 87. https://doi.org/10.1038/s41522-022-00351-7 (2022).
Wang, Y. et al. A clinical nomogram incorporating salivary Desulfovibrio desulfuricans level and oral hygiene index for predicting colorectal cancer. Ann. Transl. Med. 9(9), 15 (2021).
Zhang, L. et al. Salivary and fecal microbiota: potential new biomarkers for early screening of colorectal polyps. Front. Microbiol. 14 https://doi.org/10.3389/fmicb.2023.1182346 (2023).
Yang, Y. et al. Prospective study of oral Microbiome and colorectal cancer risk in low-income and African American populations. Int. J. Cancer. 144, 2381–2389. https://doi.org/10.1002/ijc.31941 (2019).
Uchino, Y. et al. Colorectal Cancer patients have four specific bacterial species in oral and gut microbiota in Common-A metagenomic comparison with healthy subjects. Cancers (Basel). 13. https://doi.org/10.3390/cancers13133332 (2021).
Wang, Y. et al. Alterations in the oral and gut Microbiome of colorectal cancer patients and association with host clinical factors. Int. J. Cancer. https://doi.org/10.1002/ijc.33596 (2021).
Hayden, J., van der Windt, D., Cartwright, J., Côté, P. & Bombardier, C. Assessing Bias in studies of prognostic factors. Ann. Intern. Med. 158, 280–286. https://doi.org/10.7326/0003-4819-158-4-201302190-00009 (2013).
Villar-Ortega, P. et al. The association between Fusobacterium nucleatum and cancer colorectal: A systematic review and meta-analysis. Enferm Infecc Microbiol. Clin. (Engl Ed). 40, 224–234. https://doi.org/10.1016/j.eimce.2022.02.007 (2022).
Kim, Y., Cho, N. Y. & Kang, G. H. Prognostic and clinicopathological significance of Fusobacterium nucleatum in colorectal cancer: a systemic review and meta-analysis. J. Pathol. Transl Med. 56, 144–151. https://doi.org/10.4132/jptm.2022.03.13 (2022).
Zhang, X. et al. Fecal Fusobacterium nucleatum for the diagnosis of colorectal tumor: A systematic review and meta-analysis. Cancer Med. 8, 480–491. https://doi.org/10.1002/cam4.1850 (2019).
Bhattarai, K. R., Kim, H. R. & Chae, H. J. Compliance with saliva collection protocol in healthy volunteers: strategies for managing risk and errors. Int. J. Med. Sci. 15, 823–831. https://doi.org/10.7150/ijms.25146 (2018).
Li, Y., Ou, Y., Fan, K. & Liu, G. Salivary diagnostics: opportunities and challenges. Theranostics 14, 6969–6990. https://doi.org/10.7150/thno.100600 (2024).
Kuwabara, H. et al. Salivary metabolomics with machine learning for colorectal cancer detection. Cancer Sci. 113, 3234–3243. https://doi.org/10.1111/cas.15472 (2022).
Segal, A. & Wong, D. T. Salivary diagnostics: enhancing disease detection and making medicine better. Eur. J. Dent. Educ. 12 (Suppl 1), 22–29. https://doi.org/10.1111/j.1600-0579.2007.00477.x (2008).
Warsi, I. et al. Saliva exhibits high sensitivity and specificity for the detection of SARS-COV-2. Diseases 9, 38 (2021).
Guo, X., Yu, K. & Huang, R. The ways Fusobacterium nucleatum translocate to breast tissue and contribute to breast cancer development. Mol. Oral Microbiol. 39, 1–11. https://doi.org/10.1111/omi.12446 (2024).
Parhi, L. et al. Breast cancer colonization by Fusobacterium nucleatum accelerates tumor growth and metastatic progression. Nat. Commun. 11, 3259. https://doi.org/10.1038/s41467-020-16967-2 (2020).
Hsieh, Y. Y. et al. Fusobacterium nucleatum colonization is associated with decreased survival of helicobacter pylori-positive gastric cancer patients. World J. Gastroenterol. 27, 7311–7323. https://doi.org/10.3748/wjg.v27.i42.7311 (2021).
Kudra, A. et al. Insights into oral Microbiome and colorectal cancer – on the way of searching new perspectives. Front. Cell. Infect. Microbiol. 13–2023. https://doi.org/10.3389/fcimb.2023.1159822 (2023).
Chen, J., Domingue, J. C. & Sears, C. L. Microbiota dysbiosis in select human cancers: evidence of association and causality. Semin Immunol. 32, 25–34. https://doi.org/10.1016/j.smim.2017.08.001 (2017).
Lazarevic, V., Gaïa, N., Girard, M., François, P. & Schrenzel, J. Comparison of DNA extraction methods in analysis of salivary bacterial communities. PLoS One. 8, e67699. https://doi.org/10.1371/journal.pone.0067699 (2013).
Vesty, A., Biswas, K., Taylor, M. W., Gear, K. & Douglas, R. G. Evaluating the impact of DNA extraction method on the representation of human oral bacterial and fungal communities. PLoS One. 12, e0169877. https://doi.org/10.1371/journal.pone.0169877 (2017).
Hegyi, P. et al. Academia Europaea position paper on translational medicine: the cycle model for translating scientific results into community benefits. J. Clin. Med. 9 https://doi.org/10.3390/jcm9051532 (2020).
Hegyi, P., Erőss, B., Izbéki, F., Párniczky, A. & Szentesi, A. Accelerating the translational medicine cycle: the academia Europaea pilot. Nat. Med. 27, 1317–1319. https://doi.org/10.1038/s41591-021-01458-8 (2021).
Page, M. J. et al. The PRISMA 2020 statement: an updated guideline for reporting systematic reviews. BMJ 372, 71. https://doi.org/10.1136/bmj.n71 (2021).
Higgins, J. P. T. et al. (eds) VA. Cochrane Handbook for Systematic Reviews of Interventions version 6.4 (updated August Cochrane, 2023. (2023). Available from www.training.cochrane.org/handbook
Ouzzani, M., Hammady, H., Fedorowicz, Z. & Elmagarmid, A. Rayyan-a web and mobile app for systematic reviews. Syst. Rev. 5, 210. https://doi.org/10.1186/s13643-016-0384-4 (2016).
R: A Language and Environment for Statistical Computing. https://www.R-project.org/ (2023).
Schwarzer, G. Meta: General Package for Meta-Analysis. (2023). https://doi.org/10.1007/978-3-319-21416-0.
Mantel and Haenszel. (1959). https://doi.org/10.1093/jnci/22.4.719
Robins, J., Greenland, S. & Breslow, N. E. A general estimator for the variance of the Mantel-Haenszel odds ratio. Am. J. Epidemiol. 124, 719–723. https://doi.org/10.1093/oxfordjournals.aje.a114447 (1986).
Paule, R. C. & Mandel, J. Consensus Values and Weighting Factors. J. Res. Natl. Bur. Stand . 87, 377–385. https://doi.org/10.6028/jres.087.022 (1982).
Veroniki, A. A. et al. Methods to estimate the between-study variance and its uncertainty in meta-analysis. Res. Synthesis Methods. 7, 55–79. https://doi.org/10.1002/jrsm.1164 (2016).
Mathias Harrer, P. C., Furukawa, T. & Ebert, D. Doing Meta-Analysis with R. 1st Edition edn, 500 (London: Chapman & Hall/CRC Press., (2021).
Knapp, G. & Hartung, J. Improved tests for a random effects meta-regression with a single covariate. Stat. Med. 22, 2693–2710. https://doi.org/10.1002/sim.1482 (2003).
IntHout, J., Ioannidis, J. P. & Borm, G. F. The Hartung-Knapp-Sidik-Jonkman method for random effects meta-analysis is straightforward and considerably outperforms the standard DerSimonian-Laird method. BMC Med. Res. Methodol. 14, 25. https://doi.org/10.1186/1471-2288-14-25 (2014).
Higgins, J. P. & Thompson, S. G. Quantifying heterogeneity in a meta-analysis. Stat. Med. 21, 1539–1558. https://doi.org/10.1002/sim.1186 (2002).
Funding
Open access funding provided by Semmelweis University. The open-access funding was provided by Semmelweis University. This work was supported by the Ministry of Innovation and Technology of Hungary from the National Research, Development and Innovation Fund, financed under the TKP2021-EGA-23 funding scheme. Sponsors had no role in the design, data collection, analysis, interpretation, and manuscript preparation.
Author information
Authors and Affiliations
Contributions
EG: conceptualization, investigation, project administration, methodology, formal analysis, accessing and verifying data, writing—original draft; BZ-S: conceptualization, investigation, writing—review & editing; PM: conceptualization, methodology, project administration, visualization, validation, writing—review & editing; SK-D: conceptualization, formal analysis, software, writing—review & editing; GA: conceptualization, software, writing—review & editing; PH: conceptualization, writing—review & editing; AB: conceptualization, supervision, validation, accessing and verifying data, writing—original draft; AZ: conceptualization, supervision, validation, accessing and verifying data, writing—original draft.All authors certify that they have participated sufficiently in the work to take public responsibility for the content, including participation in the concept, design, analysis, writing, or revision of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval and consent for publication
No ethical approval was required for this systematic review with meta-analysis, as all data were already published in peer-reviewed journals. No patients were involved in the design, conduct, or interpretation of our study. The datasets used in this study can be found in the full-text articles included in the systematic review and meta-analysis.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gutmacher, E., Sárai, B.Z., Martineková, P. et al. The presence and relative abundance of salivary Fusobacterium nucleatum are not associated with colorectal cancer: a systematic review and meta-analysis. Sci Rep 15, 24815 (2025). https://doi.org/10.1038/s41598-025-07465-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-07465-w