Abstract
Fusing multimodal data play a crucial role in accurate brain tumor segmentation network and clinical diagnosis, especially in scenarios with incomplete multimodal data. Existing multimodal fusion models usually perform intra-modal fusion at both shallow and deep layers relying predominantly on traditional attention fusion. Rather, using the same fusion strategy at different layers leads to critical issues, feature redundancy in shallow layers due to repetitive weighting of semantically similar low-level features, and progressive texture detail degradation in deeper layers caused by the inherent feature of deep neural networks. Additionally, the absence of intra-modal fusion results in the loss of unique critical information. To better enhance the representation of latent correlation features from every unique critical features, this paper proposes a Hierarchical In-Out Fusion method, the Out-Fusion block performs inter-modal fusion at both shallow and deep layers respectively, in the shallow layers, the SAOut-Fusion block with self-attention extracts texture information; the deepest layer of the network, the DDOut-Fusion block which integrates spatial and frequency domain features, compensates for the loss of texture detail by enhancing the detail of the high frequency component. which utilizes a gating mechanism to effectively combine the tumor’s positional structural information and texture details. At the same time, the In-Fusion block is designed for intra-modal fusion, using multiple stacked Transformer-CNN blocks to hierarchical access modality-specific critical signatures. Experimental results on the BraTS2018 and the BraTS2020 datasets validate the superiority of this method, demonstrating improved network robustness and maintaining effectiveness even when certain modalities are missing. Our code is available https://github.com/liufangcoca-515/InOutFusion-main.
Similar content being viewed by others
Introduction
3D segmentation networks play a crucial role in accurate segmentation and clinical diagnosis of brain tumors. The extraction of features of brain tumors from magnetic resonance imaging (MRI) image scans plays an important role in further analysis and clinical diagnosis. MRI images typically encompass four modalities: T1-weighted (T1), T2-weighted (T2), T1 with contrast enhancement (T1ce), and Fluid-Attenuated Inversion Recovery (Flair)1. Four individual MRI modalities exhibit modality-specific critical signatures, for example, T1 have high soft-tissue contrast delineates anatomical structures but fails to distinguish edema from non-enhanced tumors; T2 have weighted imaging detects edema and necrosis yet suffers from cerebrospinal fluid interference; T1ce precisely localizes en hancing tumor cores but is ineffective for non-enhancing subtypes; Flair enhances edema boundaries by suppressing CSF signals but underperforms T1ce in core detection. each modality has inherent limitations due to distinct imaging principles. Thus, multi-modal fusion is essential for comprehensive characterization of tumor heterogeneity2, each modality provides unique and complementary information, the absence of any modality compromises information integrity, leading to loss of critical pathological features. Conventional data imputation or generation methods may introduce noise or artifacts, thereby degrading segmentation accuracy. Fusing multimodal data3 is essential especially in scenarios with incomplete multimodal data. Multimodal data fusion enhances the perceptual capabilities of machine learning models. The construction of a learning network that captures latent correlation features in multiple modalities to generate a shared representation4 is a key approach in segmentation networks, to fully utilize the unique key features within each modality and the complex interactions between modalities, while addressing the distinct feature requirements of the network’s shallow and deep layers, particularly at the bottleneck level. Consequently, a robust multimodal fusion method is highly desired to fuse latent correlated information to enhance clinical diagnosis and segmentation accuracy.
Currently, deep learning, particularly convolutional neural networks (CNNs)5, has shown outstanding performance in image processing, and U-Net6, along with its variants, has shown significant effectiveness in brain tumor segmentation. Attention mechanisms have been integrated into U-Net, combining CNNs with Transformers to leverage self-attention mechanisms for modeling long-range dependencies and more effectively capturing the relationships between modalities and tumor regions. In brain tumor MRI multimodal fusion research, while convolutional neural networks (CNNs) effectively extract local fine-grained texture features, their limited receptive fields struggle to preserve spatial consistency of anatomical structures. Although Transformer architectures model global dependencies, they exhibit insufficient sensitivity to local features in heterogeneous tumor regions. In fusion networks, existing models78use the same representation mode for both shallow and deep layers and predominantly rely on traditional attention fusion, the differences between layers are ignored, it’s necessary to focus on the latent correlated representations among all available modalities. Additionally, the emphasis on intermodal fusion often overlooks intra-modal fusion, leading to the loss of key information specific to each modality.
Recent research shows that Fourier-Based Mixers9 can replace multi-head attention layers, achieving comparable results. The convolution theorem10 indicates that dynamic adaptive filtering in the frequency domain can effectively function as a dynamic large-kernel convolution operation. GFNet11 has successfully employed frequency domain fusion for high-quality image deblurring. The theoretical and practical feasibility of these methods is evident. But in specific brain tumor segmentation networks, such as those with large variations in texture, shape, and size, and even with missing modalities, there are still significant challenges in developing dual-domain fusion methods that leverage the advantages of both the frequency and spatial domains.
To address these challenges, we propose a Hierarchical In-Out Fusion strategy. First, the In-Fusion block performs intra-modal fusion by combining global and local features, extracting modality-specific critical signature of each modality layer by layer. Subsequently, the Out-Fusion block performs inter-modal fusion at both shallow and deep layers, employing different strategies to meet the specific needs of each layer. In the shallow layers, the SAOut-Fusion block with self-attention extracts local information to facilitate information flow via skip connections. In the deepest layer layer, the DDOut-Fusion block integrates spatial and frequency domain feature information through a dual-domain fusion module, which uses a gating mechanism to effectively combine the tumor’s positional structural information and texture details while reducing noise. The frequency domain fusion employs dynamic adaptive filters to determine which frequency features should be retained, while the spatial domain fusion enhances global structural positional information through a location-based attention mechanism. The network follows a hierarchical encoder-decoder architecture12 that dynamically leverages different levels of encoders and fusion techniques. This flexibility allows the network to effectively handle various input data types, making it suitable for scenarios involving incomplete multimodal brain tumor segmentation. The contributions of our work are outlined as below:
-
(1)
We designed a Hierarchical In-Out Fusion strategy, where intra-modal fusion via the In-Fusion block is performed before inter-modal fusion at each layer, allowing the preservation of unique critical features when modalities are missing, and this hierarchical fusion process also enhances information flow and improves the network’s robustness.
-
(2)
At the deepest representation stage of the convolutional neural network, we propose a Dual-Domain Fusion (DDOut-Fusion), which integrates spatial and frequency domain features, compensates for the loss of texture detail by enhancing the detail from the high frequency component and preserving the global structural information from the spatial domain, complementing each other to enhance the representation of latent correlation features across different domains.
-
(3)
We evaluated the network on the publicly available BraTS20187 and BraTS202013 datasets. Experimental results demonstrate that our method achieves excellent performance in segmentation tasks and is well-suited for scenarios of incomplete multimodal brain tumor segmentation.
Related work
Image segmentation networks
Deep learning is the foundational technology for implementing segmentation networks. Common segmentation network architectures include Fully Convolutional Networks (FCNs)5, Segmentation Network (SegNet)14, U-net6, encoder-decoder-based fusion15, and Transformers16. In recent years, Convolutional Neural Networks (CNNs) have effectively captured local and detailed features in images. U-Net17adopts the concept of Fully Convolutional Networks (FCNs)5and has been successful, but CNNs lack the ability to model long-range dependencies. Pure Transformer architectures, such as Swin U-Net18 and misformer19, are effective in extracting global features but struggle to learn local features effectively. Attention mechanisms computational paradigms that dynamically allocate feature weights through query-key-value interactions to focus on semantically relevant regions have been integrated into U-Net20, a typical segmentation network, including Attention U-Net21, and U-Net++22, as well as RFNet22, ViTAE, PVT23, and Swin Transformer24, which introduce a mechanism to adaptively model the relationship between modalities and tumor regions, achieving good segmentation results2526. Hybrid-Fusion27 and ParaTransCNN28 involve a parallelized encoder consisting of both CNNs and Transformers, allowing for better fusion of local and global information. In addition, MLPs2930 also mix tokens with deterministic network parameters, such as Halo-Net and swin Transformer24, which forms networks by repeating attention layers followed by point-wise convolution MLPs. The models like TransUNet31 utilizing skip connections.When segmentation networks are employed for multimodal fusion, existing research often fails to address the distinct requirements of both shallow and deep layers due to the use of a uniform fusion strategy. This leads to redundant information in the shallow layers, a loss of detailed texture in the deepest layers, and an inability to effectively capture potential multimodal correlations.
Fusion for incomplete multimodal
Fusion of incomplete multimodes aligns cross-modal features by generating missing data or building shared representations, and adaptively fuses modalities to maintain performance under partial observations. In many scenarios, medical images may be missing due to artifacts and various conditions. Many methods require the addition of extra networks and lead to increased computational costs, Hyper-Networks32 propose an adaptive multimodal MR image synthesis method among the available modalities. HeMIS33 was designed to work with hierarchical feature learning and multi-scale processing to handle complex image data effectively. Average fusion strategies34 strategies learns embeddings of multimodal information by calculating the mean and variance of features, selection rules3536 based on probability, and HVED treat the input of each modality as a Gaussian distribution. These mathematically-based fusion methods integrate features in inter-modal but overlook the contribution weight of each modality. Recently, approaches for generating shared representations have also been proposed to learn the latent correlation relationships across multiple data. Gated fusion28 applied a gate fusion strategy to extract more distinctive features for segmentation tasks, enhancing feature fusion representation, while a flexible fusion network37 can flexibly fuse varying numbers of modalities. SFusion4 utilizes self-attention modules to fuse and generate potential multimodal correlations. Saving intra-modal fusion information layer by layer is conducive to obtaining rich correlation information during inter-modal encoding. However, current methods often neglect intra-modal fusion, potentially leading to the loss of critical information. the mmFormer7 considers intra-modal fusion, the basic Transformer-based fusion mechanism struggles to effectively extract key features and global information within the modalities. Although the CNN-Transformer Network for brain tumor segmentation with the multimodal feature distillation38 build a cross modal fusion module to explicitly align the global correlations among different modalities, but the issue of the loss of the lowest-level details still remains unresolved. Simultaneously, adopting different fusion methods to meet the varying needs of shallow and deep layers during inter-modal fusion within the network is also an urgent issue that needs to be addressed.
The method of spatial and frequency domain fusion
The Dual-Domain fusion method to fuses spatial textures with frequency spectra through dual-stream processing, to effectively combine the tumor’s positional structural information and texture detail. The convolution theorem has been generalized10 to illustrate that a dynamic adaptive filter is analogous to a dynamic large-kernel convolution operation. Fourier-Based Mixers9and Fourier Transform methods39have been proposed. such as FNet9, GFNet11, FNO40, and AFNO41 leverage the Fourier transform to achieve spatial token mixing, resulting in fewer parameters compared to traditional convolutional methods. Similarly, Global Filter Networks (GFNs)11 and the Adaptive Fourier Transform (AFT)10has been proposed for global feature fusion using adaptive filters. The research42demonstrates that transformers operating in the frequency domain exhibit superior capabilities for high-quality image deblurring. Additionally, it has been established that integrating spectral and multi-headed attention layers enhances transformer architectures, particularly by incorporating spectral layers in the initial stages and multi-headed attention layers in subsequent stages. In spatial domains, the local Vision43 approach converts the space into a set of small windows and performs attention operations, characterized by sparse connectivity, weight sharing, and dynamic weights44. This method can significantly improve memory and computational efficiency45 when processing local content in long texts. There is still no two-domain fusion strategy that is suitable for multimodal fusion segmentation networks with significant changes in texture, shape and tumor image size.
Methods
Methodology
This study proposes Hierarchical In-Out Fusion for Segmentation network based on the TransUNet architecture. When performs inter-modal, at the shallow layer of the network, the self-attention mechanism is adopted. At the deepest layer of the network, a dual-domain fusion mechanism of spatial and frequency domains is adopted to extract richer high-level semantic information. Meanwhile, globally and locally collaborative encoding is fused in intra-modal to enhance the specific key features of each modality.
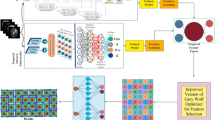
The overview of network structure diagram is shown as Figure 1. The input tokens set P first undergoes intra-modal fusion, followed by inter-modal fusion. Finally, a softmax function is applied to generate the shared representation. This network structure adopts a method of calling encoders and fusion modules to flexibly handle incomplete multimodal fusion, which n denoting the number of available modalities.
The structure diagram of the Hierarchical In-Out Fusion Network assumes three available modalities\(f_{1}\), \(f_{2}\), \(f_{3}\) with \(f_{n}\) representing additional modalities that can be added. The In-Fusion block is used for intra-modal fusion, while the Out-Fusion block is used for inter-modal fusion; and finally, an multiplication is performed between the input feature map and the corresponding weight map(\(a_{1}\), \(a_{2}\) and \(a_{3}\)) to obtain the shared representation \(f_{s}\), which is used for segmentation.
Firstly, given the input tokens set P, which represents the feature tokens for all available 3D image modalities. At each layer of the encoder, the In-Fusion block performs intra-modal fusion to obtain the token set from \((l-1)_{th}\) layer to \(l_{th}\) layer. This process can be formulated as follows:
where, \(\overline{P} _{N}^{IN_{l-1}}\) represents tokens set from any complete modality at the the \((l-1)_{th}\) layer, \(\overline{P} _{N}^{IN_{l}}\) represents tokens set of all available modalities at the the \(l_{th}\) layer of encode, \(\phi _{1}\) represents the operation function of Infusion block, \(\theta _{n}^{attn}\) is corresponding network parameters.
Next, the Out-Fusion block performs inter-modal fusion at both shallow and deep layers. The tokens are processed through the SAOut-Fusion block, which merges them into the corresponding decoder layer. And at the bottleneck layer of the network, the DDOut-Fusion block performs Dual-Domain fusion to obtain the output token set. This process can be formulated as follows:
where, \(\widehat{P} _{N}^{Out}\) represents tokens set through inter-modal fusion, \(\overline{P} _{N}^{IN}\) represents tokens set through intra-modal fusion, \(\phi _{2}\) represents the operation function of Out-Fusion block, \(\theta _ {n}^{'attn}\) is Corresponding network parameters.
Finally, \(\widehat{P} _{N}^{Out}\) are restored 3D feature representation map through split. Using a softmax function after segmentation network to generate the weight maps. We can obtain the shared representation by element-wise multiplying input feature map with the corresponding weight map and summing all the modalities, the process can be summarized as follows:
where, the \(f_{n}\) is 3D feature representation of the \(n_{th}\) image modality, \(n\in \left\{ 1,2...N \right\}\), N denotes the maximum number of available modes. Meanwhile, let \(m \in M \subseteq \left\{ 1,2,..., N\right\}\) index a particular subset of modalities, where M represents the set of available modalities, the feature tokens set P for all 3D image available modalities, expressed as \(P= \left\{ f_{m} \mid m \in M \right\}\), and the fused feature set reverted through a decoder, which is represented as \(f'_m\). An multiplication is performed between the input feature map and the corresponding weight map(\(a_{1}\), \(a_{2}\) and \(a_{3}\)) to obtain the shared representation \(f_{s}\), which is used for segmentation.
In-fusion block in intra-modal via transformer-CNN
The In-Fusion block is designed to facilitate intra-modal fusion within the encoder network architecture. Our synergistic hybrid architecture addresses these limitations: CNN branches capture tumor sub-region details, such as tumor cores through multi-scale kernels, while transformer branches establish global semantic correlations across modalities. A bidirectional gated feature interaction module enables hierarchical dynamic fusion. This architecture progressively combines global and local features, extracts high-level features from the input data through a series of stacked stage and downsampling operations, with downsampling achieved via max pooling3D. The In-Fusion block consists of 4 stages, which serves as the fundamental component of the network, comprising three main parts: a 3D convolution layer, a parallel branch feature extractor, and a feedforward neural network (FFN). The framework considers the interplay between longrange and shortrange information, thereby facilitating the extraction of highlevel, rich features. Furthermore, the use of residual connections alleviates common training challenges associated with deep networks and ensures a smooth flow of information throughout the architecture. The corresponding network structure is shown in Figure 2.
The network structure diagram of In-Fusion block in intra-modal. The left three circles (colored blue, green, and purple) represent the token sets corresponding to the three different modalities, \(\overline{P} _{N}^{In_{l-1}}\) denotes the tokens set from the \((l-1)_{th}\) layer input at stage i; the right three circles with gradient colors represent the fused token sets, and \(\overline{P} _{N}^{In_{l}}\) denotes the tokens set from the \(l_{th}\) layer output at stage i.
The parallel branch feature extractor consists of two paths: one for local feature extraction based on CNNs and another for global feature extraction via transformers. Each branch produces one-dimensional features that are subsequently integrated and encoded by the FFN, which integrates both global and local features. The local feature extraction path focuses on capturing local texture details through multiple residual blocks, with each block containing two 3D convolution layers with a stride of 1 and a kernel size of \(3\times 3\times 3\). Following each convolution operation, instance normalization is applied in conjunction with the Leaky ReLU activation function. Another branch for global feature extraction based on Vision Transformer (VIT) architecture, capturing overall information from each 3D tumor image to enhance the representation of global features. Each block within the ViT consists of layer normalization, multi-head self-attention, and a feedforward neural network. The outputs from preceding layers are connected through a multi-layer perceptron (MLP) using residual connections. the process can be summarized as follows:
where, \(\overline{P} _{n}^{local_{l} }\) represents the local fused features of the \(n_{th}\) available modalities at the \(l_{th}\) layer, \(\overline{P}_{n}^{global_{l} }\) represents the global fused features of the \(n_{th}\) available modalities at the \(l_{th}\) layer, and \(\overline{P} _{N}^{IN_{l}}\) represents the fused features through In-Fusion block at the \(l_{th}\) layer.
Out-fusion block in inter modal
DDOut-fusion via dual-domain fusion at the deepest layer
In the deepest (bottleneck)layers, the Dual-Domain OutFusion (DDOut-Fusion) block integrates spatial and frequency domain features through a dual-domain fusion module, which utilizes a gating mechanism to effectively combine the tumor’s positional structural information and texture details while reducing noise. The spatial fusion is achieved through location-based local attention, and the frequency domain fusion is performed using adaptive frequency filters, The network structure diagram of DDOut-Fusion in intra-modal is illustrated as Figure 3. The process can be summarized as follows:
where, \(\overline{P} _{S}^{Out}\) represents tokens set through spatial fusion, \(\overline{P} _{F}^{Out}\)represents tokens set through frequency fusion, \(\overline{P} _{N}^{Out}\) represents tokens set through DDOut-Fusion block.
In spatial fusion branch, the location-based local attention weights are computed by focusing solely on the positional information within the window using relative position encoding43. The input token sequences Pn as a window of n consecutive inputs \(X_{t-n} ,...X_{t}\) in parallel, the attention output for the query \(h_{t}\) at the center position is the aggregation of the corresponding values in the local window. Let \(h_{t}\) be the feature value corresponding to the center position at time t in the given window, defined as \(h_{t}\in R^{1\times d }\), while \(h_{\le t}\in R^{t\times d }\) represents the feature values at the preceding positions in the same window, it reasons over time using multi-head attention. When the attention head is the \(i_{th}\) head, \(q_{i}=h_{t} Q_{i}\) is defined as query, \(k_{i}=\bar{h} _{\le t} K_{i}\) is defined as key, and \(v_{i}=\bar{h} _{\le t} V_{i}\) is defined as value. The attention weight is computed as the softmax normalization of the dot product between the query q and key k to generate a similarity matrix, which are a set of learnable weight matrices, the query q and key k share weights to reduce computational costs, the corresponding attention value is defined as follows:
where, \(\sigma\) (\(\cdot\)) is defined as the softmax operator. Attention is the linear combination of each attention head, let u is the number of attention heads, then last attention value is calculated as follows:
where, \(W_i\) is the weight of the \(i_{th}\) head, and \(\ attn_i\) is the attention value of the \(i_{th}\) head. \(\overline{P} _{S}^{Out1}\) is output tokens set in spatial domains after attention. Each transformer within each modality incorporates both multi-head attention (MSA) and a feedforward network (FFN), the process can be summarized as follows:
where, \(\overline{P} _{N}^{IN}\) denotes input tokens set, \(\overline{P} _{S}^{Out}\) denotes output tokens set in spatial fusion branch and Z denotes output tokens set through MSA.
In frequency fusion branch, the Dynamic Adaptive Frequency Filters (DAFT) was designed, we devise the \(G\left( \cdot \right)\) function as the \(G(\mathcal {F}(\overline{P}_{N}^{IN} ))\) mask tensor matrix, enables the learning of an instance-adaptive mask from the frequency representation \(\mathcal {F}(\overline{P}_{N}^{IN} )\), which is the tokens set \(\overline{P}_{N}^{IN}\) through fast Fourier transform. The \(G\left( \cdot \right)\) function employs a \(1\times 1\) convolution as a linear layer with a non-linear activation function (ReLU), followed by another linear layer to represent the tokens set. This process can be summarized as follows:
The matrix \(G\left( \cdot \right)\) has the same shape as \(\mathcal {F}(\overline{P}_{N}^{IN} )\) function, and the dot product operation of \(G\left( \cdot \right)\)and \(\mathcal {F}(\overline{P}_{N}^{IN} )\) yields the fused output in frequency domain. This process can be formulated as follows:
where, \(\odot\) denotes the Hadamard product, \(\overline{P}_{F}^{Out1}\) denotes output tokens after the \(G\left( \cdot \right)\) function.
This network structure operates in two main stages: transformation and concatenation. In the first stage, the input tokens set \(\overline{P}_{N}^{IN}\) are normalized using layer normalization (LN), and then passed through the DAFT block add to obtain \(\widehat{P}_{F}\), the subsequent stage, the tokens set \(\widehat{P}_{F}\) undergo further normalization and activation using the ReLU function, then concatenated with the original normalized tokens and passed through layer normalization once more to produce the final output tokens set in frequency domains fusion branch \(\overline{P}_{F}^{Out}\). This process can be formulated as follows:
SAOut-fusion via skip connection at shallow layers
In the shallow layers, SAOut-Fusion achieves inter-modal fusion through skip connections, which is an improved transformer module that combines attention and feedforward networks. This module enhances the model’s expressiveness through layer normalization, residual connections, and DropPath, improving the network’s generalization ability. The input tokens set at the \(l_{th}\) layer \(\overline{P} _{N}^{IN_{l}}\) is first standardized through a normalization layer; next, the data are processed through the self-attention mechanism, and the output is added to the input via a residual connection; then, DropPath regularization is applied by using the input drop probability dropprob, randomly generating a keep mask, and adding the attention output with the original input to get the \(P_{N}^{Out_{l_{1} }}\), This process can be formulated as follows:
where, the Q, K and V represent the query, key, and value corresponding to the feature tokens, respectively. Afterward, the data \(P_{N}^{Out_{l_{1} }}\) are passed through the feedforward network, the output tokens set \(P_{N}^{Out_{l_{ffn} }}\) is then passed through DropPath and added to the original input tokens set \(P_{N}^{IN_{l} }\) to get the out tokens set \(P_{N}^{Out_{l} }\) at the \(l_{th}\) layer. Finally, the output of the entire SAOut-Fusion layer is the features processed through both the attention mechanism and the feedforward network.This process can be formulated as follows:
Experiments and results
Dataset and implementation
The experiments were conducted on the BraTS202013 and the BraTS20187 dataset. the dataset are public dataset that can be used for scientific research. the BraTS202013 dataset comprises 369 multicontrast MRI scans including Flair(F), T1, T1 contrast enhanced (T1c), T2 and experienced imaging experts annotated ground truth. To better reflect clinical application tasks, the dataset includes three tumor regions: whole tumor(WT), enhance tumor(ET), and tumor core(TC), which need to be segmented into three parts: ET, TC, and WT. The BraTS20187 contains 3D multimodal brain MRIs, including Flair, T1, T1c, T2 and ground truth. It includes 285 cases for training (210 gliomas with high grade and 75 gliomas with low grade). The dataset was randomly divided into 70% for training, 10% for validation, and 20% for testing, ensuring that all methods were evaluated on the same dataset and data split. To mitigate the risk of overfitting, we employed two data enhancement techniques: random flipping along the axes and rotation within a random angle range of [−10\(^{\circ }\), 10\(^{\circ }\)]. Each volume was individually normalized with the z score [15], followed by random cropping of 128 \(\times\) 128 \(\times\) 128 patches to serve as input to the networks. The framework was implemented using PyTorch 1.9.1 on a NVIDIA RTX 4090 (24GB) GPU. The input size consisted of 128 \({\times }\) 128 \({\times }\) 128 patches, with a batch size of 1. The network was trained using the Adam optimizer with an initial learning rate of 0.0002 for 250 epochs. The total training time was approximately 26 hours, utilizing 17GB memory on each GPU.
Baseline methods
The baseline methods aim to leverage Transformers to build a unified model for incomplete multimodal learning in brain tumor segmentation. The SFusion method4 proposes a self-attention fusion block based on the self-attention mechanism, where features are extracted from the upstream processing model, projected as tokens, and fed into the self-attention module to capture latent multimodal correlations. The Multimodal Medical Transformer (mmFormer)7employs hybrid modality-specific encoders and a modality-correlated encoder to capture long-range dependencies both within and across different modalities. It extracts modality-invariant representations by explicitly building and aligning global correlations between modalities.
Results
Incomplete multimodal activity recognition
To evaluate the performance of our model, we conducted a comparative analysis with four recent methods: UHeMIS33, mmFormer7, SFusion4 and CNNTra38. Our method achieves the highest performance among the evaluated models.
We conducted experiments on the BraTS202013 dataset, the Dice score was employed as the primary evaluation metric, as demonstrated in in Table 1, adhering to the data partitioning strategy outlined in [24], and we reference the results from that study. Our method outperforms the current state-of-the-art (SOTA) method, when compared with SFusion, which achieved average Dice scores of 57.33, 71.81 and 82.29 for the WT, TC, ET, respectively, our method achieving average Dice scores of 62.43, 73.54, and 83.14 for ET, TC, and WT respectively. This results in enhancements of 5.10%, 1.73%, and 0.85%, respectively across these tumor regions. when compared with CNNTra38, which achieved average Dice scores of 59.61, 73.21, and 83.07 for the ET, TC, and WT, respectively, our method demonstrates improvements of 2.81%, 0.33%, and 0.07% across these tumor regions.
We conducted experiments on the BraTS20187 dataset, compared to mmFormer, the Dice score was employed as the primary evaluation metric, as demonstrated in Table 2, which achieved average Dice scores of 59.85, 73.01 and 82.94 for the WT, TC, ET, respectively, our method achieving average Dice scores of 63.83, 74.37, and 84.97 for ET, TC, and WT respectively (epoch 1000). This results in enhancements of 3.98%, 1.41%, and 2.03%, respectively, across these tumor regions. when compared with CNNTra38, which achieved average Dice scores of 57.44, 73.01 and 84.91 for the ET, TC, and WT, respectively, our method demonstrates improvements of 6.39%, 1.36%, and 0.06% across these tumor regions.
The In-Fusion block is used to capture modality-specific key information, effectively preserving unique critical features in incomplete multimodal scenarios. Different fusion strategies applied at different layers improve the representation of latent correlation features, resulting in significantly improved scores for ET and TC. Furthermore, the rich texture details from the frequency domain and the global structural information from the spatial domain complement each other, leading to clearer tumor boundary segmentation in 3D image comparisons. The average performance of Hierarchical In-Out Fusion is superior, and the average value of 11 Out of all 15 possible pattern combinations has been improved. Moreover, our method demonstrates a clear advantage in the segmentation of enhancing tumors (ET) and tumor cores (TC).
Visual comparison results of 3D brain tumor segmentation
In addition to the quantitative analysis, we also performed a visual comparison to further validate the performance of our method. Randomly selected three input images which have significant differences in the size, shape, and location of the tumors, comparing our fusion method with traditional networks such as UHeMIS33, as well as certain fusion methods like SFusion4, and mmFormer7. To evaluate the robustness of our method for incomplete multimodal segmentation, we used the same data partitioning as SFusion to ensure a fair comparison and directly referenced their results. The segmentation images produced by our fusion method exhibit greater detail fidelity and clearer tumor boundary delineation, closely aligning with the ground truth images. The quantitative segmentation results are shown in Figure 4.
Visual comparison results between ours In-Out Fusion and other four compared methods on the BraTS202013 benchmark (red, geen, and yellow regions represent the ET, TC, and WT, respectively), the red box represents regions that may be tumor areas.
In any scenario where the T1c modality is present (whether with one, two, or three modalities), the performance scores achieved by In-Out Fusion show a significant improvement compared to cases where the T1c modality is absent. Furthermore, by incorporating the T1c modality, we performed brain tumor segmentation across various combinations of missing modalities, demonstrating favorable segmentation results for WT, TC,and ET in comparison to SFusion4. The Visual comparison results are shown in Figure 5.
Brain tumor segmentation maps corresponding to different combinations of modalities, comparing Ours Fusion and SFusion on the BraTS202013 images. Red, green, and yellow regions represent ET, TC, and WT. The red box highlights potential tumor areas.
Based on the BraTS202013 public dataset, a patient’s brain tumors multimodal MRI scans (T1, T2, T1c, Flair; stored in the NIfTI standard compressed format *.nii.gz) were processed using the medical imaging software ITK-SNAP (version 4.0.0)46. The original *.nii.gz file was imported into ITK-SNAP for multi-planar reformation (MPR), generating orthogonal cross-sectional views in the axial (top-left), coronal (top-right), and sagittal (bottom-right) planes, along with a 3D volume-rendered spatial distribution of the tumor (bottom-left) after annotation. To validate the segmentation results, the predicted mask (*.nii.gz) generated by our method and the BraTS ground truth mask (*.nii.gz) were co-registered in ITK-SNAP after spatial alignment. We formatted the graph displayed in ITK-SNAP (version 4.0.0) to obtainvisual comparison resultst as shown in Figure6. The 3D segmentation produced by the Hierarchical In-Out Fusion Network achieved an average similarity of 83.9% compared to the ground truth 3D segmentation on the lable WT, effectively illustrating that the three-dimensional structure of brain tumor has good 3D segmentation effect.
The visualization images generated by ITK-SNAP (version 4.0.0)46 were integrated and formatted as follows. The picture on the left: three cross-sectional views (axial, coronal, sagittal) and a 3D-rendered tumor image of a selected BraTS202013 case, derived from four MRI modalities (T1, T2, T1c, Flair). The picture on the right: The fused segmentation mask (.nii.gz) generated by our method and the ground truth mask (.nii.gz), both visualized in ITK-SNAP for comparative analysis. Yellow, green, and red represent WT, TC, and ET.
Ablation experiments
We evaluated the average Dice scores under two configurations: retaining the Out-Fusion block while using CNN alone, Transformer alone(Tra), or their combined architecture (CNN-Tra) in the In-Fusion block; and employing only the In-Fusion block without Out-Fusion block with the same architectural variations. As shown in Table 3, our method achieves competitive performance, validating that the integration of Transformer and CNN architectures effectively enables local-global feature complementarity. We investigate the effectiveness of the In-Fusion block in intra-modal fusion (IF), the SAOut-Fusion block in inter-modal fusion (SAF), and the DDOut-Fusion block in inter-modal fusion (DDF) as three critical components of our method. We evaluate In-Out Fusion without SAF and DDF. Specifically, In-Out Fusion without SAF and DDF indicates that feature representations are directly fed into the IF module, meaning that we directly add the calculated feature representations up to get the fusion result. Our findings reveal that InOut Fusion without SAF and DDF performs worse than the other methods. Similarly, configurations lacking either IF and DDF or IF and SAF also yield suboptimal results. We compare the performance of our method using IF alone, SAF alone, or DDF alone, as shown in Table4, which presents the average performance over the 15 possible combinations on the BraTS202013 dataset. The results demonstrate that the IF block, SAF block, and DDF block contribute to performance improvements across all tumor regions. This underscores the importance of IF, SAF, and DDF in the overall effectiveness of InOut-Fusion.
Discussion and conclusion
In this paper, we propose Hierarchical In-Out Fusion for incomplete multimodal brain tumor segmentation. Compared to existing multimodal fusion networks, our approach achieves more effective results without adding extra encoders that would increase computational cost. Hierarchical fusion, which employs different fusion strategies at different layers, enhances the representation of latent correlation features, resulting in significantly improved scores for ET and TC. Due to dual-domain fusion, leading to clearer tumor boundary segmentation in 3D image comparisons. As a result, the average Dice score is improved compared to similar fusion networks. We demonstrate that our approach can be seamlessly integrated into existing backbone networks by replacing their fusion operations or blocks, performing individual segmentation for each image modality and learning latent correlation representation through a shared encoder can improve segmentation performance, but it increases the network’s computational overhead, to designing a lightweight network is an important problem to address in the future.
Data availability
The experiments were conducted on the BraTS202013 and the BraTS20187 dataset. The datasets are public datasets that can be used for scientific research. You can download the BraTS202013 at https://www.kaggle.com/datasets/awsaf49/brats20-dataset-training-validation. You can download the BraTS20187 at https://www.med.upenn.edu/sbia/brats2018/data.html.
References
Menze, B. H. et al. The multimodal brain tumor image segmentation benchmark (brats). IEEE transactions on medical imaging 34, 1993–2024 (2014).
Xing, D. et al. Optimised diffusion-weighting for measurement of apparent diffusion coefficient (adc) in human brain. Magn. Reson. Imaging15, 771–784 (1997).
Song, H. et al. Multimodal separation and cross fusion network based on raman spectroscopy and FTIR spectroscopy for diagnosis of thyroid malignant tumor metastasis. Sci. Rep.14, 29125 (2024).
Liu, Z., Wei, J., Li, R. & Zhou, J. Sfusion: Self-attention based n-to-one multimodal fusion block. In International Conference on Medical Image Computing and Computer-Assisted Intervention, 159–169 (Springer, 2023).
Long, J., Shelhamer, E. & Darrell, T. Fully convolutional networks for semantic segmentation. In Proceedings of the IEEE conference on computer vision and pattern recognition, 3431–3440 (2015).
Huang, H. et al. Unet 3+: A full-scale connected unet for medical image segmentation. In ICASSP 2020-2020 IEEE international conference on acoustics, speech and signal processing (ICASSP), 1055–1059 (IEEE, 2020).
Zhang, Y. et al. mmformer: Multimodal medical transformer for incomplete multimodal learning of brain tumor segmentation. In International Conference on Medical Image Computing and Computer-Assisted Intervention, 107–117 (Springer, 2022).
Ding, Y., Yu, X. & Yang, Y. Rfnet: Region-aware fusion network for incomplete multi-modal brain tumor segmentation. In Proceedings of the IEEE/CVF international conference on computer vision, 3975–3984 (2021).
Lee-Thorp, J., Ainslie, J., Eckstein, I. & Ontanon, S. Fnet: Mixing tokens with fourier transforms. arXiv preprint arXiv:2105.03824 (2021).
Huang, Z. et al. Adaptive frequency filters as efficient global token mixers. In Proceedings of the IEEE/CVF International Conference on Computer Vision, 6049–6059 (2023).
Rao, Y., Zhao, W., Zhu, Z., Lu, J. & Zhou, J. Global filter networks for image classification. Adv. in neural information processing systems 34, 980–993 (2021).
Chen, H., Wang, Z., Qin, H. & Mu, X. Dhfnet: Decoupled hierarchical fusion network for rgb-t dense prediction tasks. Neurocomputing583, 127594 (2024).
Bakas, S. et al. Advancing the cancer genome atlas glioma mri collections with expert segmentation labels and radiomic features. Sci. data 4, 1–13 (2017).
Badrinarayanan, V., Kendall, A. & Cipolla, R. Segnet: A deep convolutional encoder-decoder architecture for image segmentation. IEEE transactions on pattern analysis and machine intelligence 39, 2481–2495 (2017).
Dorent, R., Joutard, S., Modat, M., Ourselin, S. & Vercauteren, T. Hetero-modal variational encoder-decoder for joint modality completion and segmentation. In Medical Image Computing and Computer Assisted Intervention–MICCAI 2019: 22nd International Conference, Shenzhen, China, October 13–17, 2019, Proceedings, Part II 22, 74–82 (Springer, 2019).
Wang, P., Yang, Q., He, Z. & Yuan, Y. Vision transformers in multi-modal brain tumor mri segmentation: A review. Meta-Radiology 100004 (2023).
Ronneberger, O., Fischer, P. & Brox, T. U-net: Convolutional networks for biomedical image segmentation. In Medical image computing and computer-assisted intervention–MICCAI 2015: 18th international conference, Munich, Germany, October 5-9, 2015, proceedings, part III 18, 234–241 (Springer, 2015).
Cao, H. et al. Swin-unet: Unet-like pure transformer for medical image segmentation. In European conference on computer vision, 205–218 (Springer, 2022).
Huang, X., Deng, Z., Li, D., Yuan, X. & Fu, Y. Missformer: An effective transformer for 2d medical image segmentation. IEEE Transactions on Med. Imaging 42, 1484–1494 (2022).
Yang, H., Aydi, W., Innab, N., Ghoneim, M. E. & Ferrara, M. Classification of cervical cancer using dense capsnet with seg-unet and denoising autoencoders. Sci. Rep.14, 31764 (2024).
Oktay, O. et al. Attention u-net: Learning where to look for the pancreas. arXiv preprint arXiv:1804.03999 (2018).
Zhou, Z., Siddiquee, M. M. R., Tajbakhsh, N. & Liang, J. Unet++: Redesigning skip connections to exploit multiscale features in image segmentation. IEEE transactions on medical imaging 39, 1856–1867 (2019).
Wang, W. et al. Pyramid vision transformer: A versatile backbone for dense prediction without convolutions. In Proceedings of the IEEE/CVF international conference on computer vision, 568–578 (2021).
Liu, Z. et al. Swin transformer: Hierarchical vision transformer using shifted windows. In Proceedings of the IEEE/CVF international conference on computer vision, 10012–10022 (2021).
Chu, X. et al. Twins: Revisiting spatial attention design in vision transformers. arXiv preprint arXiv:2104.138402 (2021).
Guo, M.-H., Liu, Z.-N., Mu, T.-J. & Hu, S.-M. Beyond self-attention: External attention using two linear layers for visual tasks. IEEE Transactions on Pattern Analysis and Machine Intelligence 45, 5436–5447 (2022).
Cho, J. & Park, J. Hybrid-fusion transformer for multisequence mri. In International Conference on Medical Imaging and Computer-Aided Diagnosis, 477–487 (Springer, 2022).
Chen, C. et al. Robust multimodal brain tumor segmentation via feature disentanglement and gated fusion. In Medical Image Computing and Computer Assisted Intervention–MICCAI 2019: 22nd International Conference, Shenzhen, China, October 13–17, 2019, Proceedings, Part III 22, 447–456 (Springer, 2019).
Zhang, D. J. et al. Morphmlp: A self-attention free, mlp-like backbone for image and video. arXiv preprint arXiv:2111.125272 (2021).
Alexey, D. An image is worth 16x16 words: Transformers for image recognition at scale. arXiv preprint arXiv: 2010.11929 (2020).
Chen, J. et al. Transunet: Transformers make strong encoders for medical image segmentation. arXiv preprint arXiv:2102.04306 (2021).
Yang, H., Sun, J. & Xu, Z. Learning unified hyper-network for multi-modal MR image synthesis and tumor segmentation with missing modalities. IEEE Trans. Med. Imaginghttps://doi.org/10.1109/TMI.2023.3301934 (2023).
Havaei, M., Guizard, N., Chapados, N. & Bengio, Y. Hemis: Hetero-modal image segmentation in: International conference on medical image computing and computer-assisted intervention (2016).
Wang, Z. & Hong, Y. A2fseg: Adaptive multi-modal fusion network for medical image segmentation. In International Conference on Medical Image Computing and Computer-Assisted Intervention, 673–681 (Springer, 2023).
Ouyang, J., Adeli, E., Pohl, K. M., Zhao, Q. & Zaharchuk, G. Representation disentanglement for multi-modal brain mri analysis. In Information Processing in Medical Imaging: 27th International Conference, IPMI 2021, Virtual Event, June 28–June 30, 2021, Proceedings 27, 321–333 (Springer, 2021).
Alzahrani, A. A. Enhanced multimodal medical image fusion via modified dwt with arithmetic optimization algorithm. Sci. Reports 14, 19261 (2024).
Yang, H., Zhou, T., Zhou, Y., Zhang, Y. & Fu, H. Flexible fusion network for multi-modal brain tumor segmentation. IEEE J. Biomed. Heal. Informatics 27, 3349–3359 (2023).
Kang, M., Ting, F. F., Phan, R. C.-W., Ge, Z. & Ting, C.-M. A multimodal feature distillation with cnn-transformer network for brain tumor segmentation with incomplete modalities. arXiv preprint arXiv:2404.14019 (2024).
Guibas, J. et al. Efficient token mixing for transformers via adaptive fourier neural operators. In International Conference on Learning Representations (2021).
Li, Z. et al. Fourier neural operator for parametric partial differential equations. arXiv preprint arXiv:2010.08895 (2020).
Guibas, J. et al. Adaptive fourier neural operators: Efficient token mixers for transformers. arXiv preprint arXiv:2111.13587 (2021).
Kong, L., Dong, J., Ge, J., Li, M. & Pan, J. Efficient frequency domain-based transformers for high-quality image deblurring. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 5886–5895 (2023).
Han, Q. et al. On the connection between local attention and dynamic depth-wise convolution. arXiv preprint arXiv:2106.04263 (2021).
Vaswani, A. et al. Scaling local self-attention for parameter efficient visual backbones. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 12894–12904 (2021).
Touvron, H. et al. Feedforward networks for image classification with data-efficient training. Feedforward networks for image classification with data-efficient training” in (2021).
Yushkevich, P. A. et al. User-guided 3d active contour segmentation of anatomical structures: significantly improved efficiency and reliability. Neuroimage 31, 1116–1128 (2006).
Acknowledgements
This work was supported in part by the National Natural Science Foundation of China under Grant 62072350, Grant 62171328, Grant 62401410; in part by Central Government Guides Local Science and Technology Development Special Projects under Grant ZYYD2022000021; in part by the National Natural Science Foundation of Hubei under Grant 2023AFB158; in part by Enterprise Technology Innovation Project under Grant 2022012202015060; and in part supported by the Scientific Research Team Plan of Wuhan Technology and Business University.
Author information
Authors and Affiliations
Contributions
Fang.Liu. conceived the experiment(s), JiaMing.Wang. and LiWei.Wang. conducted the experiment(s), Tao.Lu. and YanDuo.Zhang. analysed the results. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liu, F., Zhang, Y., Lu, T. et al. Hierarchical in-out fusion for incomplete multimodal brain tumor segmentation. Sci Rep 15, 23017 (2025). https://doi.org/10.1038/s41598-025-07466-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-07466-9