Abstract
The brain tumour (BT) is an aggressive disease among others, which leads to a very short life expectancy. Therefore, early and prompt treatment is the main stage in enhancing patients’ quality of life. Biomedical imaging permits the non-invasive evaluation of diseases, depending upon visual assessments that lead to better medical outcome expectations and therapeutic planning. Numerous image techniques like computed tomography (CT), magnetic resonance imaging (MRI), etc., are employed for evaluating cancer in the brain. The detection, segmentation and extraction of diseased tumour regions from biomedical images are a primary concern, but are tiresome and time-consuming tasks done by clinical specialists, and their outcome depends on their experience only. Therefore, the use of computer-aided technologies is essential to overcoming these limitations. Recently, artificial intelligence (AI) models have been very effective in enhancing performance and improving the method of medical image diagnosis. This paper proposes an Enhanced Brain Tumour Segmentation through Biomedical Imaging and Feature Model Fusion with Bonobo Optimiser (EBTS-BIFMFBO) model. The main intention of the EBTS-BIFMFBO model relies on enhancing the segmentation and classification model of BTs utilizing advanced models. Initially, the EBTS-BIFMFBO technique follows bilateral filter (BF)-based noise elimination and CLAHE-based contrast enhancement. Furthermore, the proposed EBTS-BIFMFBO model involves a segmentation process by the DeepLabV3 + model to identify tumour regions for accurate diagnosis. Moreover, the fusion models such as InceptionResNetV2, MobileNet, and DenseNet201 are employed for the feature extraction. Additionally, the convolutional sparse autoencoder (CSAE) method is implemented for the classification process of BT. Finally, the hyper-parameter selection of CSAE is performed by the bonobo optimizer (BO) method. A vast experiment is conducted to highlight the performance of the EBTS-BIFMFBO approach under the Figshare BT dataset. The comparison results of the EBTS-BIFMFBO approach portrayed a superior accuracy value of 99.16% over existing models.
Similar content being viewed by others
Introduction
Despite significant progress being made to understand the path physiology of a tumour, its self-sufficient and highly dynamic nature poses some challenges relating to the recognition of the tumour and follow-up of treatment1. Even though technological progress holds promise to address these dilemmas, because of their challenges, complexities endure in multiple levels of cancer treatment, comprising segmentation of tumours, tracing of cancer evolution, detection of pre-cancerous lesions, and more2. The brain is the crucial organ in the human body, containing billions of cells. BT is segmented into two kinds of cancers, lower and higher grade. Lower-grade BT is named benign. Likewise, the higher-grade cancer is known as malignant3. Benign cancer is not a cancerous tumour. Consequently, it does not spread to other portions of the brain. Nevertheless, cancer is a cancerous tumour4. Subsequently, it induces immediate death. BT gets more complicated in shape and differs significantly in size, location, and texture; therefore, health records connected to cancers show higher structural and spatial variation from patient to patient. To recognize a tumour, a patient will experience various tests like MRI, CT, etc., which are presently employed to image cancers in the central nervous system5. The capability to image cancers with a ligand, which binds particularly to cancer cells and immediately crosses the intact blood-brain barrier, can substantially enhance cancer resolution from the brain and presents enhanced recognition of the exterior margin of tumour cells6.
During BT surveys, the health records embedded in MRI data are essential for investigating structural brain irregularities, tumour segmentation, recognizing pre-cancerous lesions, and identification, while contrasted with another model7.Conventional approaches to cancer assessment are based on qualitative aspects like intra-tumoral cellular composition, density of tumour, and anatomic relation with neighbouring tissue, also named semantic features. Technological development has permitted the quantitative analysis of neuropathology, assisting in quantifying cancer shape, textual patterns, and size8. Whereas earlier models of cancer identification depended on probabilistic cancer approaches, essential inventions in computer vision (CV) have resulted in faster growth in AI investigation in clinical imaging9. Deep learning (DL) and machine learning (ML) are subdivisions of AI and have a vast potential to assist clinicians in attaining insight from complicated, higher-dimensional, and diverse biomedical information. Compared with conventional statistics, DL and ML differ in their methods of BT quantification, with distinct weaknesses and strengths10. Segmentation of biomedical images with statistical modelling and DL and ML-based approaches is entrenched as an individual investigation region.
This paper proposes an Enhanced Brain Tumour Segmentation through Biomedical Imaging and Feature Model Fusion with Bonobo Optimiser (EBTS-BIFMFBO) model. The main intention of the EBTS-BIFMFBO model relies on enhancing the segmentation and classification model of BTs utilizing advanced models. Initially, the EBTS-BIFMFBO technique follows bilateral filter (BF)-based noise elimination and CLAHE-based contrast enhancement. Furthermore, the proposed EBTS-BIFMFBO model involves a segmentation process by the DeepLabV3 + model to identify tumour regions for accurate diagnosis. Moreover, the fusion models such as InceptionResNetV2, MobileNet, and DenseNet201 are employed for the feature extraction. Additionally, the convolutional sparse autoencoder (CSAE) method is implemented for the classification process of BT. Finally, the hyperparameter selection of CSAE is performed by the bonobo optimizer (BO) method. A vast experiment is conducted to highlight the performance of the EBTS-BIFMFBO approach under the FigshareBT dataset. The significant contribution of the EBTS-BIFMFBO approach is listed below.
-
The EBTS-BIFMFBO model utilizes the BF method to eliminate noises from MRI images, significantly enhancing the quality of images by preserving edge details. This approach highlights the relevant features, allowing for more accurate tumour detection. Improving the clarity of the MRI scans contributes to the overall efficiency of the BT classification system.
-
The EBTS-BIFMFBO approach employs CLAHE to improve the MRI image contrast, enhancing the visibility of critical tumour features. This technique assists in highlighting subtle details, making it easier to detect and classify BT accurately. Improving image quality plays a crucial role in the overall performance of the tumour detection system.
-
The EBTS-BIFMFBO method employs DeepLabV3 + for accurate pixel-wise segmentation of BTs, one of the advanced architecture which precisely delineates tumour boundaries. This ensures accurate segmentation even in intrinsic MRI scans with varying tumour shapes. The precise segmentation contributes to the overall efficiency of the tumour detection and classification process.
-
The integration of pre-trained models such as InceptionResNetV2, MobileNet, and DenseNet201 improves the feature extraction process by utilizing their own merits for capturing a broader range of tumour characteristics. This multi-model approach improves the robustness of feature extraction, ensuring more detailed and accurate tumour representation. The system can handle diverse tumour discrepancies in MRI scans by incorporating these models.
-
The EBTS-BIFMFBO methodology implements the CSAE approach for robust and efficient classification, ensuring high accuracy in tumour identification. Integrating the BO model improves the model by fine-tuning hyper parameters, enhancing non-functional performance. This integration results in a more precise and reliable classification system for BT detection.
-
Integrating DeepLabV3 + for precise segmentation, with multiple pre-trained models such as InceptionResNetV2, MobileNet, and DenseNet201 for feature extraction. This approach utilizes CSAE for robust classification, while the BO-based optimization additionally fine-tunes hyper parameters for superior performance. The novelty of the proposed model integrates advanced segmentation, diverse feature extraction models and optimization techniques to improve BT detection and classification accuracy and efficiency.
Literature review
Rohilla and Jain11, precise segmentation and detection models for BT are projected in this work. A modified ResNet-50 technique is employed to recognize cancer, and a ResUNet model-based convolutional neural network (CNN) approach for segmentation is projected. The segmentation and detection were implemented on a similar database comprising FLAIR, pre- and post-contrast MRI images. Owing to the utilization of residual systems, the novelists monitored development in performance metrics, like precision for cancer recognition and Dice similarity coefficient for cancer segmentation. Ashimgaliyev et al.12 proposed a new incorporation of cutting-edge segmentation approaches with DL ensemble models to improve the classification precision of MRI-based BT identification. This work also performs a complete review of either conventional segmentation methods or contemporary developments in region-based and ML-driven segmentation models. This study examines the effectiveness of DL ensemble models, optimizing various method frameworks to enhance cancer classification stability and precision. Mansur et al.13 aimed at three segmentation techniques: region-based segmentation, threshold-based segmentation, and employing a CNN utilizing Unet to segment BT over MRI data. These models are intended to segment and differentiate the cancer region from normal tissue accurately. In14, a DL-based detection of BT in smart medical care methods with confidentiality preservation is proposed. In the initial stage, patients with BT are the main objective of an effective medical care method, which is being developed. During another stage, the novelists progress to a DL-based cancer identification platform that integrates the AES-128 models and PBKDF2 to protect clinical image storage on the data transmission and server. The projected method incorporates the VGG-16, Inception-V3, and ResNet-50 frameworks to develop a CNN-based BT identification method. Schwehr and Achanta15 developed a region of interest (ROI) recognition model that performs during data pre-processing to find salient aspects and extract irrelevant MRI data. It reduces the method’s input size, permitting more combative data augmentations and deeper NNs. Succeeding the pre-processing of the MRI conditions, a fully convolutional AE partitioned the diverse brain MRI utilizing attention gate and channel-wise attentions. Natha et al.16 introduced an innovative Stack Ensemble TL model named “SETL-BMRI” to identify BT in MRI images. The SETL-BMRI method integrates dual pre-trained methods, VGG19 and AlexNet, to enhance its capability for generalization. Stacking associated outputs from these techniques substantially enhanced the BT recognition precision compared to specialized techniques. Tiwari et al.17, inspected the projected approach called the adaptive neuro-fuzzy inference system-fusion-DBN (ANFIS-F-DBN) technique introduced in this investigation. Initially, the brain image attained from the identified database is pre-processed utilizing a Gaussian filter. Furthermore, deep fuzzy clustering (DFC) approach is applied to segment the region of BT, and multiple levels were developed for the image augmentation method. Finally, the recognition of BT is implemented and depends on hybrid DL techniques such as ANFIS-F-DBN.
Awudong and Li18 introduced an innovative deep semi-supervised learning method, which fully employs unlabeled data samples. A novel sliding window feature extractor approach was used to evade resizing images that can cause disregard and loss of smaller aspects, to precisely analyze brain tissue lesions of smaller BT, which is complex to identify. The feature extractor backbone system is developed with the convolutional block attention module (CBAM) that permits the system to fully recognize image data in either spatial or channel features, improving the system’s perception capability of essential characteristics. Sharif et al.19 developed an optimized DL technique for accurate and efficient multimodal BT classification using improved pre-processing, feature optimization, and multi-class support vector machine (MC-SVM) classification. Hekmat et al.20 developed an accurate BT classification model using contrast-limited adaptive histogram equalization (CLAHE), discrete wavelet transform (DWT), DenseNet121 and DenseNet169 architectures, and an attention mechanism (AM), attaining high accuracy on Kaggle and Figshare datasets. Zahid et al.21 proposed a fully automated BT classification system using optimized DL features and hybrid optimization techniques to mitigate redundancy, improve prediction speed, and maintain high accuracy. Rastogi et al.22 improved BT classification using a fine-tuned deep transfer learning (DTL) technique, with the Xception model achieving the highest accuracy for improved diagnosis. Nawaz et al.23 developed an improved UNET methodology using DenseNet77 for accurate and robust melanoma segmentation across varying skin lesion shapes, sizes, and colours. Karthik et al.24 developed a unified approach for BT multi-classification and segmentation by incorporating attention-augmented CNN, random forest (RF), and U-Net, achieving high accuracy and precision in MRI BT analysis. Mathivanan et al.25 explored DTL models for accurate BT diagnosis using MRI, with MobileNetv3 achieving the highest performance. Gao et al.26 proposed a novel BT classification method, SwinBTC, based on the healthcare Internet of Things (HIoT) and Swin transformer (ST) model, improving classification accuracy and reliability in MRI tumour detection. Asiri et al.27 developed a fast and accurate two-module system for BT detection using image enhancement and SVM-based classification, achieving high accuracy and efficiency. Semwal et al.28 developed a hybrid CNN-SVM methodology optimized by particle swarm optimization (PSO) for accurate, non-invasive BT classification, achieving superior performance in early tumour detection and treatment planning.
Several existing methods still encounter challenges related to the complexity of tumour heterogeneity, high computational costs, and limited generalization across diverse datasets. Some models depend on intrinsic methods and techniques, resulting in longer training times and potential overfitting. Although various methods illustrate high accuracy, a gap exists in developing more effective methods with mitigated computational overhead while maintaining or improving diagnostic performance. Furthermore, integrating hybrid and ensemble learning models exhibits real-time applicability and robustness limitations across diverse MRI datasets. Additionally, fusion of AM and transfer learning shows efficiency, but optimizing these techniques for handling several tumour types and imaging conditions remains underexplored. Moreover, some models fail to address the challenge of balancing model complexity with practical deployment in clinical settings, highlighting the requirement for more streamlined and adaptable solutions.
Materials and methods
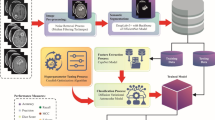
In this study, an EBTS-BIFMFBO model is proposed. The main intention of the EBTS-BIFMFBO model is to enhance the segmentation and classification model of BTs by utilizing advanced models. It holds image pre-processing, segmentation, fusion of feature models, BT detection using CASE, and parameter selection. Figure 1 demonstrates the entire flow of the EBTS-BIFMFBO approach.
Image pre-processing
Initially, the EBTS-BIFMFBO technique utilizes BF-based noise removal and CLAHE-based contrast enhancement. This fusion model was chosen for its capability to effectively address common threats in medical imaging, specifically MRI scans. The BF model effectually eliminates noise while preserving edge information, an essential factor for precise detection. CLAHE improves the local contrast of the image, allowing for improved visibility of subtle tumour features that other models may miss. These methodologies work synergistically to enhance image quality without over-enhancing artefacts, which is often risky with alternative methods. BF and CLAHE give a balanced model, ensuring noise reduction and contrast improvement, which is significant for accurate classification and segmentation tasks in BT detection. This integration assists in maintaining crucial image details while enhancing the visibility of tumours, thereby giving a reliable foundation for additional analysis with DL techniques.
BF-based noise elimination
BF is the denoising model applied for smoothing images by reflecting the intensity and distance amongst pixels29. BF can eliminate noise in the image and successfully preserve the image edge details. During BF, all pixels are substituted by the total amount of pixel intensities to eliminate noise. The weightage is chosen according to the Gaussian distribution. The BF procedure is mathematically seen in Eq. (1).
Whereas \(\:B(x,\:y)\) represents BF outcome at pixel point \(\:(x,\:y)\), \(\:({x}^{{\prime\:}},\:{y}^{{\prime\:}})\) denotes neighboring pixel of \(\:(x,\:y)\),\(\:\:(x,\:y)\) signifies co-ordinates of image pixels from \(\:xth\) row and \(\:yth\) column, \(\:{\sigma\:}_{s}\) signifies spatial kernel region, and \(\:{\sigma\:}_{\gamma\:}\) denotes minimal amplitude value.
CLAHE-based contrast enhancement
The CLAHE model is executed to show hidden characteristics by matching the histogram for all values of image pixels. This mechanism gives maximal boundary values for the histogram height (clip limit). A clip limit is applied to split the image into smaller portions with equal dimensions according to the parameters. The Eq. (2) is used by the clip limit measurement procedure.
Whereas \(\:{l}_{\text{m}\text{a}\text{x}}\) refers to the maximal pixel value of gray-scale, \(\:{c}_{i,j}\) stands for the value of clip limit in \(\:jth\) column and\(\:ith\) row, \(\:k\) signifies area dimensions, \(\:h\) refers to the gray-scale value in the interval of 1-256, and \(\:\alpha\:\) represents clip feature in the interval of 0-100.
Segmentation using DeepLabV3+
In the proposed EBTS-BIFMFBO model, segmentation process is carried out by DeepLabV3 + architecture to identify tumour regions for accurate diagnosis30. This model is chosen for its superiority in segmenting complex images like MRI scans. This model efficiently captures multi-scale contextual data, enabling the model to detect both small and large tumour features with high accuracy. Furthermore, the encoder-decoder structure of DeepLabV3 + assists in preserving spatial resolution, which is significant for precisely delineating tumour boundaries. Compared to conventional segmentation models, DeepLabV3 + better handles tumour size, shape, and location discrepancies. It’s efficient segmenting complex structures makes it an ideal choice for BT segmentation, where fine details are crucial for precise diagnosis and treatment planning. This model illustrates superior performance in medical imaging tasks and making it an ideal selection for BT detection.
The DeepLab-v3 + method is tailored for segmentation, which includes classifying all pixels in the image into pre-defined classes. The structure incorporates numerous essential modules, namely the Atrous Spatial Pyramid Pooling (ASPP) component, Atrous convolution, and an encoder-decoder architecture.
Backbone Network (Encoding)
The encoding stage of DeepLab-v3+, extracts deeper feature representations from the input image. Numerous backbone methods, like ResNet50, ResNet18, Xception, and MobileNet-V2, are tested to recognize the appropriate configurations for these tasks. ResNet50 was created to present the optimal balance between computational efficiency and segmentation accuracy.
Let \(\:x\) represent the input images. The encoding removes feature maps.\(\:{L}_{j}\) over a series of convolutional layers:
While \(\:{f}_{enc}\) characterizes the convolutional layers series in the backbone, \(\:{W}_{enc}\) refers to a collection of weights for the encoder.
The backbone removes higher-level feature mapping for input to the ASPP component and maintains lower level features from previous layers. Lower level features are essential for tasks such as segmentation, whereas precise pixel-wise predictions are required. During the framework, lower level features \(\:{L}_{j}\) from the encoding were distributed immediately to the decoding for fine grained particulars, whereas higher level features \(\:{\text{L}}_{ASPP}\) blocks handled LASPP to take global and contextual information.
ASPP
The ASPP component takes multiple scale context data by utilizing parallel atrous convolutions with dissimilar dilation rates, in addition to average global pooling. The output of every atrous convolution is specified by:
Whereas \(\:{y}_{ASPP}^{\left(i\right)}\) denotes output feature mapping for the rate of dilation \(\:{r}_{i}\) and \(\:{W}_{{r}_{i}}\) represents weight with rates \(\:{r}_{i}\), \(\:*\)signifies convolutional, and \(\:\varphi\:\) denotes activation function (for example,\(\:ReLU\)).
Meanwhile, \(\:N\) denotes total pixel counts \(\:(H\)x\(\:W\text{x}C)\) in an input, and \(\:{L}_{ASPP}\left[i\right]\) represents the vector of a feature.
The last output of ASPP is gained by concatenation of the outputs of the global average pooling and the parallel atrous convolutions, accompanied by a \(\:1\)x\(\:1\) activation and convolution:
Here, \(\:{W}_{ASPP}\) characterizes the weighting for the last convolution after concatenation.
Feature fusion models
The fusion models such as InceptionResNetV2, MobileNet, and DenseNet201 are chosen for its capability of capturing intrinsic features from MRI images. InceptionResNetV2 integrates the Inception modules and residual connections, enabling them to learn multi-level features and capture fine-grained details from varied image regions, making it highly effective for BT detection. MobileNet is a lightweight, computationally efficient model appropriate for real-time applications. DenseNet201 model utilizes dense connections that promote feature reuse and enhance the data flow throughout the network, improving its capability to detect complex tumour patterns. By using these three models, the system benefits from both accuracy and efficiency, utilizing the merits of all architecture to handle the complexity and diversity of BT images better than conventional methods.
InceptionResNetV2 technique
InceptionResNetV2 is an advanced neural network (NN) structure, which integrates Inception modules with residual links to improve the accuracy and efficacy of image recognition31. During this presented study, the primary extraction of the features takes place in a block that contains pooling and convolutional layers for downsampling the inputs. These blocks start with three convolutional layers: 32 filters (3 × 3, no padding), 32 filters (3 × 3, stride 2, no padding), and 64 filters (3 × 3, padding), accompanied by a max-pooling layer. It proceeds with a convolution layer of 80 filters (1 × 1), another with 192 filters (3 × 3), and finishes with a max-pooling layer (3 × 3 pool dimensions, stride 2). Inception-ResNet‐A contains parallel convolution routes with different residual connections and kernel dimensions (1 × 1, 3 × 3, 5 × 5). Inception ResNet contains complex residual connections and convolutional routes. This network ends with a globally average pooling layer to decrease sizes to 1 × 1, and dense layers for the last classification. It successfully combines residual and inception models, presenting a robust method for great performance image recognition with effective feature extraction.
MobileNet approach
MobileNet was presented to present low computational usage and improved implementation on mobile devices with lower hardware requirements32. This method decreases the learnable parameter counts by utilizing various convolution layers, like depth-wise separable 1 × 1 convolutions, to manage all input channels. It utilizes this convolution to decrease the feature size map when extracting the features. Its special feature is its usage of depth-wise separable convolutions, regular convolutions separated into depth-wise and one-by-one pointwise convolutions. All input channels are filtered out in depth-wise convolutions autonomously. The one-by-one pointwise convolutions unite each of the depth-wise convolution outputs linearly. These factorizations result in an essential decrease in the calculation cost or the model dimensions. By succeeding in all convolutional layers in the method, the nonlinear ReLU activation function and batch normalization were utilized. Stride procedures are applied for downsampling in the depth-wise convolutional layers or the initial convolutions. Formerly, the softmax classifier, fully connected (FC) layer, and average pooling layer were added. The elementary MobileNet comprises 28 layers, comprising pointwise and depthwise convolution layers.
DenseNet201 model
DenseNets have gained more popularity. It is a short form of densely linked convolution systems that correspond to ResNets but additionally have different main changes33. Unlike ResNet, which utilizes a preservative model while the preceding output is input for upcoming layers, Densenet captures each preceding output as input for forthcoming layers. It was tailored to address the vanishing gradient problems in DNNs; however, information vanished before reaching its target owing to the longer distance between the output and input layers. Figure 2 depicts the framework of DenseNet201.
During this DenseNets architecture, the layer’s output \(\:{a}^{\left[l\right]}\)is linked to the outputs of each preceding layer over functions, as stated in Eq. (7).
Here, let’s focus on the system with \(\:L\) layers. In the convention system, \(\:L\) links are similar to connections amongst the layers. Nevertheless, in the DenseNets, there should be around\(\:\frac{L(L+1)}{2}\) connections. It has better connections than other smaller layers. This new architecture effectively permits DenseNets to use training methods with over 100 layers.
BT classification using CSAE
The EBTS-BIFMFBO technique utilizes CSAE to classify BT34, which effectively learns sparse and discriminative features from MRI images. It can effectively detect subtle patterns in BT. The model also mitigates noise and irrelevant details by learning sparse representations, improving the overall classification performance. Furthermore, the model shows efficiency in performing unsupervised feature learning and capturing both spatial and semantic relationships within the data, resulting in an enhanced accuracy in tumour classification. CSAE’s auto-encoding architecture allows it to concentrate on crucial features while discarding redundant data, ensuring more efficient and robust classification. This capability makes CSAE superior to other classification techniques that may face difficulty with intrinsic, high-dimensional medical imaging data.
An autoencoder (AE) is a specialized feed-forward NN created to duplicate the input signal as closely as possible at the output. The AE is made from two major sections: a decoder and an encoder. The encoding converts the input signal \(\:x\in\:{R}^{N}\)into the latent code, or representation, utilizing a convolutional layer count. Afterwards, the decoder rebuilds the novel input by mapping these latent representations to return to the input area. CSAE is a specialized form of NNk created for effective feature extraction from signals, utilizing the powers of either sparse coding or convolutional operations methods. The convolutional layers in CSAEs permit the technique to effectively seize temporal or spatial variation in data, making them mainly efficient for processing designed signals, namely audio, time-series, or image data. By presenting sparsity constraints, CSAEs guarantee that just a few smaller neurons are active at some provided time, encouraging the finding of more distinct and interpretable features. This sparsity is usually enforced utilizing methods, namely \(\:{L}_{1}\) regularisation, promising the system for learning contracts and substantial representation. Therefore, CSAEs can isolate the most appropriate designs inside the input data, resulting in stronger and generalizable characteristics for downstream tasks like regression or classification. Their capability to incorporate local receptive areas with sparsity creates CSAEs, an excellent device for removing higher quality characteristics from composite signals.
Let \(\:x\in\:{R}^{N}\) characterize the input 1D signal to the AE with dimension \(\:N\). During this encoding, the \(\:lth\) convolutional layer use filters \(\:{\left\{{W}_{l}^{\left(k\right)}\right\}}_{k=1}^{{K}_{l}} \in{\text{R}}^{{K}_{l}\times{S}_{l}}\) to the input signal, while \(\:{K}_{l}\) signifies filter counts and \(\:{S}_{l}\)represents filter dimensions. The \(\:lth\) convolutional layer’s output (feature mapping) is gained by:
Whereas, \(\:{b}_{l}^{\left(k\right)}\in\:{R}^{{K}_{l}}\) refers to bias, \(\:'*'\) signifies the convolutional process, and \(\:\sigma\:\) denotes the activation function. The final convolutional layer is accompanied by a flattening layer that connects each of the 261 feature mappings of \(\:the\:lth\) layer:
A dense layer emulates the flattening procedure, generating the encoder’s output. To apply sparsity in the dense layer, a regularization term is added to the loss function, like the \(\:{L}_{1}\) norm of the feature mapping:
Whereas \(\:\lambda\:\) denotes the regularization parameter. In addition, kernel regularization applied for the convolutional layers. This regularizer is a \(\:{L}_{2}\) norm (ridge regularization) that penalizes the squared amount of the kernel weights:
Here, \(\:\alpha\:\) refers to the regularisation parameter for the kernel regularisation.
The decoding rebuilds the signal from the latent representation utilizing deconvolution (moved convolution). The output of the lth layer of the decoder is gained by
.
Now, \(\:{\stackrel{\sim}{W}}_{l}^{\left(k\right)}\) denotes a collection of filters applied to convolution with the encoding output signal, and \(\:{\stackrel{\sim}{b}}_{l}^{\left(k\right)}\) signifies a biased term, correspondingly, in the \(\:lth\) layer of a decoder. At last, the decoder output \(\:\stackrel{\sim}{x}={\stackrel{\sim}{h}}_{l}^{\left(k\right)}\) represents the reconstructed signal of the input signal\(\:\:x\). The CSAE training is established by reducing the following loss function:
The initial term of the statement (13) is the reconstruction loss, which calculates the change between the reconstructed signal and the input signal, usually utilizing the mean squared error (MSE) and is provided by
Parameter selection using BO approach
Finally, the hyper parameter selection of CSAE is performed by utilizing the BO method35.This model is chosen for its efficiency in optimizing hyper parameters in complex models. This method adapts to intrinsic, high-dimensional search spaces of DL methods, finding optimal solutions with fewer iterations. Its capability in exploring the parameter space ensures it can avert local minima and find the most effective hyper parameters for the task. The technique integrates a probabilistic model, which allows for better exploration-exploitation trade-offs, enhancing the model’s overall performance. BO presents better optimization with lower computational cost than grid or random search methods. The robustness and adaptability of the approach to diverse DL models make it appropriate for fine-tuning intrinsic techniques such as CSAE and DeepLabV3 + for BT detection.
BO is a robust optimizer model stimulated by bonobos’ natural social behaviour approaches. The arithmetical representation of BO is presented in the subsequent sections.
Fission fusion social approach
The initial stage is applying this approach, which selects the bonobo to mate with the present individual. This chosen bonobo (\(\:pth\) bonobo) is selected from a smaller short-term subgroup. Initially, the maximal dimensions \(\:tsg{s}_{\text{m}\text{a}\text{x}}\) of the sub-group are computed as shown:
Whereas \(\:tsg{s}_{factor}\) means the feature of the short-term sub group and \(\:N\) characterizes the population size. The \(\:tsg{s}_{factor}\) indicates dynamical parameters that should be presented, and its primary value is initiated as shown:
Here, \(\:tsg{s}_{factor\_max}\) signifies the maximal value of \(\:tsg{s}_{factor}\), which is set at 0.07.
Promiscuous and restrictive mating approach
This approach should generate the novel bonobo once the bonobo population is in a positive phase (\(\:PP\)). The mathematical representation is built as shown:
Whereas \(\:{\alpha\:}_{j}^{bonobo}\), bonobo \(\:{i}_{j}\), and bonobo \(\:{p}_{j}\) represent locations in the \(\:jth\:\)size of the optimal bonobo, present bonobo, and chosen bonobo, respectively. \(\:{r}_{1}\) refers to randomly generated numbers uniformly distributed among (0,1). \(\:scab\) and \(\:scsb\) represent distribution coefficients for the \(\:{\alpha\:}_{bonobo}\) and the \(\:pth\) bonobo, respectively. The flag is set to \(\:-1\) or 1.
Consortship and extra group mating approach
On the contrary, this approach should be accepted in the negative phase (NP) to make the original bonobo. Once, randomly generated values \(\:{r}_{2}\) amongst (0,1) is higher than the parameter \(\:{p}_{xgm}\), the consortship mating approach should be utilized, which is represented below:
Meanwhile, \(\:{r}_{3}\) and \(\:{r}_{4}\) represent a randomly generated numberin [0,1]. When \(\:{r}_{2}\) is discovered to be less than or equivalent to the \(\:{p}_{xgm}\), the extra group mating approach should be implemented as shown below:
On the other hand, \(\:{r}_{5}\) and \(\:{r}_{6}\) refer to randomly generated numbers between \(\:0\) and 1. \(\:{\beta\:}_{1}\) and \(\:{\beta\:}_{2}\) are dual in-between parameters that can specify the power of the extra group mating. \(\:u{b}_{j}\) and \(\:l{b}_{j}\) denote lower and upper limits in the \(\:jth\) size. \(\:{p}_{d}\) denotes directional likelihood.
Parameters’ updating.
The parameter’s updating is extremely significant for the BO model, making the BO performance efficient, stable, and flexible. Owing to whether the present iteration discovers a novel best, BO gives two types of parameter upgrading methods. When an improved \(\:{\alpha\:}_{bonobo}\) is discovered, the parameters should be adjusted as shown:
Whereas \(\:rcpp\:\)signifies the rate of change in the positive and negative stages. In the positive stage, \(\:{p}_{p}\) grows gradually to improve the exploitation capability of BO, which assists the bonobos in discovering optimal solutions. In other cases (for example, negative stage), better \(\:{\alpha\:}_{bonobo}\) is not designed; these parameters are upgraded as shown:
By utilizing Eq. (22), the exploration capability of BO is enhanced owing to the smaller \(\:{p}_{p}\) (lower than 0.5). Table 1 describes the hyper parameters of the BO technique.
The BO method generates a fitness function (FF) for attaining a heightened result of a classifier. It states a positive integer to suggest a better outcome of the candidate solution. Here, the classifier ratio of error minimization is evaluated as FF, as set in Eq. (23).
Performance analysis
The performance evaluation of the EBTS-BIFMFBO model is examined under the Figshare BT dataset36. This dataset contains 3093 images under three classes: meningioma, glioma, and pituitary, as exposed in Table 2. Figure 3 shows the sample brain images and Fig. 4 shows a sample of pre-processed brain images and segmented regions.
Table 3; Fig. 5 represent the segmented outcomes of the EBTS-BIFMFBO technique with existing models. The table values imply that the EBTS-BIFMFBO technique obtained greater outcomes of \(\:acc{u}_{y}\) of 99.95%, dice score of 97.32%, \(\:pre{c}_{n}\)of 97.44%, and\(\:\:rec{a}_{l}\) of 97.29%. Meanwhile, the SVM and Hyb-DCNN-ResnNet 152 TL models have obtained somewhat closer outcomes.AEDCNN + Inception ResnetV2, Capsule Networks, Extreme learning based on FCM, FCM clustering algorithm, Ensemble-DL, and Contrast Normalization-SVM techniques have the worst performance.
Figure 6 represents the classifier results of the EBTS-BIFMFBO technique below 80%TRPH and 20%TSPH. Figure 6a and b demonstrates the confusion matrices with correct recognition and identification of each class label. Figure 6c shows the PR values, demonstrating superior outcomes over every class. Figure 6d exemplifies the ROC values, signifying capable outcomes with maximum ROC analysis for diverse class labels.
Table 4; Fig. 7 established the BT detection of the EBTS-BIFMFBO methodology under 80%TRPH and 20%TSPH. The outcomes suggest that the EBTS-BIFMFBO methodology correctly recognized the samples. With 80%TRPH, the EBTS-BIFMFBO methodology presentedan average \(\:acc{u}_{y}\), \(\:pre{c}_{n}\), \(\:rec{a}_{l}\), \(\:{F1}_{score}\), and MCC of 99.16%, 98.68%, 98.72%, 98.70%, and 98.04%, respectively. Besides, with 70%TRPH, the EBTS-BIFMFBO approach presented an average \(\:acc{u}_{y}\), \(\:pre{c}_{n}\), \(\:rec{a}_{l}\), \(\:{F1}_{score}\), and MCC of 98.28%, 97.33%, 97.15%, 97.24%, and 95.87%, correspondingly.
Figure 8 illustrates the training (TRA) \(\:acc{u}_{y}\) and validation (VAL) \(\:acc{u}_{y}\) analysis of the EBTS-BIFMFBO approach below 80%TRPH and 20%TSPH. The \(\:acc{u}_{y}\) analysis is computed within the range of 0–20 epochs. The figure highlights that the TRAand VAL \(\:acc{u}_{y}\) analysis exhibited an increasing trend, which indicates the capacity of the EBTS-BIFMFBO method with superior performance across multiple iterations. Followed by, the TRA and VAL \(\:acc{u}_{y}\)leftovers closer across the epochs, which indicates inferior overfitting and exhibits maximal performance of the EBTS-BIFMFBO method.
Figure 9 shows the TRA loss (TRALOS) and VAL loss (VALLOS) curve of the EBTS-BIFMFBO approach below 80%TRPH and 20%TSPH. The loss values are computed within the range of 0–20 epochs. The TRALOS and VALLOS analysis exemplify a reducing tendency, informing the capability of the EBTS-BIFMFBO approach in balancing a trade-off. The constant reduction also assures the developed performance of the EBTS-BIFMFBO method and tunes the forecast outcomes.
Figure 10 established the classifier outcomes of the EBTS-BIFMFBO methodology under 70%TRPH and 30%TSPH. Figure 10a and b illustrates the confusion matrices with perfect recognition of each class label. Figure 10c shows the PR values, specifying greater performance over every class label. Simultaneously, Fig. 10d reveals the values of ROC, which indicate the capability of better ROC analysis for dissimilar classes.
Table 5; Fig. 11 represent the BT detection of the EBTS-BIFMFBO method below 70%TRPH and 30%TSPH. The outcomes imply that the EBTS-BIFMFBO model accurately identifies the samples. With 70%TRPH, the EBTS-BIFMFBO model presents an average\(\:\:acc{u}_{y}\), \(\:pre{c}_{n}\), \(\:rec{a}_{l}\), \(\:{F1}_{score}\), and MCC of 97.75%, 96.38%, 96.47%, 96.43%, and 94.68%, correspondingly. In addition, with 30%TRPH, the EBTS-BIFMFBO approach provides average \(\:acc{u}_{y}\), \(\:pre{c}_{n}\), \(\:rec{a}_{l}\), \(\:{F1}_{score}\), and MCC of 97.63%, 96.42%, 96.28%, 96.34%, and 94.49%, respectively.
Figure 12 illustrates the TRA \(\:acc{u}_{y}\) and VAL \(\:acc{u}_{y}\)results of the EBTS-BIFMFBO methodology below 70%TRPH and 30%TSPH. The \(\:acc{u}_{y}\)analysis is intended over the range interval of 0–20 epochs. The graph states that TRA and VAL \(\:acc{u}_{y}\) analysis exhibit an increasing trend, which indicates the capacity of the EBTS-BIFMFBO methodology with superior performance across multiple iterations. Also, TRA and VAL \(\:acc{u}_{y}\:\)remains closer across the epochs, which signifies inferior overfitting and exhibits optimal performance of the EBTS-BIFMFBO method.
Figure 13 demonstrates the TRALOS and VALLOS analysis of the EBTS-BIFMFBO approach below 70%TRPH and 30%TSPH. The loss values are computed throughout 0–20 epochs. The TRALOS and VALLOS values demonstrate a diminishing trend, indicating the capacity of the EBTS-BIFMFBO technique in balancing a trade-off. The persistent decrease assures the maximum performance of the EBTS-BIFMFBO approach and tunes the prediction outcomes.
Table 6; Fig. 14 inspect the comparative results of the EBTS-BIFMFBO technique under existing techniques37,38,39,40. The outcomes reported that the Deep-CNN, DRLBP + CNN, DenseNet- 161, and GoogleNettechniques reported worse performance. Meanwhile, DCNN-SGD, 3D ConvNet, and VGG Net techniques have accomplished closer outcomes. Furthermore, the EBTS-BIFMFBO method reported superior performance with better\(\:\:pre{c}_{n}\), \(\:rec{a}_{l},acc{u}_{y},\:\)and \(\:{F1}_{score}\) of 98.68%, 98.72%, 99.16%, and 98.70%, correspondingly.
Table 7; Fig. 15 demonstrate the ablation study of the EBTS-BIFMFBO methodology. The EBTS-BIFMFBO methodology attains an \(\:acc{u}_{y}\) of 99.16%, \(\:pre{c}_{n}\) of 98.68%, recall of 98.72%, and an \(\:{F1}_{score}\) of 98.70%. These outputs exhibit enhancement over DenseNet201, which had an \(\:acc{u}_{y}\) of 97.82% and an \(\:{F1}_{score}\) of 97.54%. Additionally, CSAE illustrates an \(\:acc{u}_{y}\) of 98.38%, but its \(\:{F1}_{score}\) was lower at 98.05%. The other models, such as MobileNet, InceptionResNetV2, and DeepLabV3, performed slightly worse across all metrics, with \(\:acc{u}_{y}\) values ranging from 95.99 to 97.31%. The superior performance of the EBTS-BIFMFBO technique highlights its efficiency in improving gesture recognition accuracy for visually impaired individuals.
Conclusion
In this research paper, an EBTS-BIFMFBO model is proposed. The main intention of the EBTS-BIFMFBO model relies on enhancing the segmentation and classification model of BTs utilizing advanced models. Initially, the EBTS-BIFMFBO model follows BF-based noise removal and CLAHE-based contrast enhancement. Followed by, the proposed EBTS-BIFMFBO model involves a segmentation process by the DeepLabV3 + architecture to identify tumour regions for accurate diagnosis precisely. Furthermore, the fusion models such as InceptionResNetV2, MobileNet, and DenseNet201 are employed for the feature extraction. Besides, the proposed EBTS-BIFMFBO technique utilizes the CSAE method for the classification procedure of BT. Finally, the BO technique performs the hyper parameter selection of the CSAE model. A vast experiment is conducted to highlight the performance of the EBTS-BIFMFBO approach under the Figshare BT dataset. The comparison results of the EBTS-BIFMFBO approach portrayed a superior accuracy value of 99.16% over existing models. The limitations of the EBTS-BIFMFBO approach comprise reliance on the quality and diversity of the input data, which may affect generalization across diverse clinical environments. The model’s performance also varies with low-resolution or noisy MRI scans not represented in the training set. The model also lacks real-time integration with clinical decision systems, which is crucial for practical deployment. Limited explain ability of the DL outputs can affect clinician trust and utilization. Computational needs may pose challenges in resource-constrained settings. Future studies should focus on developing lightweight models for real-time evaluation, improving the interpretability of the model through explainable AI, and validating the system with larger, multi-institutional datasets. Integrating electronic health records and multimodal data fusion can also improve diagnostic accuracy and clinical relevance.
Data availability
The data supporting this study’s findings are openly available in the Kaggle repository.
References
Mostafa, N. N. Human brain tumors detection using neutrosophic c-means clustering algorithm. J. Neutrosophic Fuzzy Syst. 1 (1), 55–58 (2021).
Sarhan, A. M. Brain tumor classification in magnetic resonance images using deep learning and wavelet transform. JBiS 13, 102–112 (2020).
Ghaffari, M., Sowmya, A. & Oliver, R. Automated brain tumor segmentation using multimodal brain scans: a survey based on models submitted to the brats 2012–2018 challenges. IEEE Rev. Biomed. Eng. 13, 156–168 (2019).
Hashemzehi, R., Mahdavi, S. J. S., Kheirabadi, M. & Kamel, S. R. Detection of brain tumors from MRI images based on deep learning using hybrid model CNN and NADE. Biocybernetics Biomedical Eng. 40, 1225–1232 (2020).
Mohan, G. & Subashini, M. M. MRI based medical image analysis: survey on brain tumor grade classification. Biomed. Signal Process. Control. 39, 139–161 (2018).
Toğaçar, M., Ergen, B., Cömert, Z. & BrainMRNet Brain tumor detection using magnetic resonance images with a novel convolutional neural network model. Med. Hypotheses. 134, 109531 (2020).
Alam, M. S. et al. Automatic human brain tumor detection in mri image using template-based k means and improved fuzzy c means clustering algorithm. BDCC 3, 27 (2019).
Amin, J., Sharif, M., Yasmin, M. & Fernandes, S. L. Big data analysis for brain tumor detection: deep convolutional neural networks. Future Generation Comput. Syst. 87, 290–297 (2018).
Chattopadhyay, A. & Maitra, M. MRI-based brain tumour image detection using CNN-based deep learning method. Neurosci. Inf. 2, 100060 (2022).
Mostafa, N. N. Human brain tumor detection using a neutrosophic c-means clustering algorithm. JNFS 1, 50–58 (2021).
Rohilla, S. & Jain, S. Detection of Brain Tumor Employing Residual Network-based Optimized Deep Learning. CAD. 21, 15–27 (2025).
Ashimgaliyev, M., Matkarimov, B., Barlybayev, A. & Li, R. Y. M. &Zhumadillayeva, A. Accurate mri-based brain tumor diagnosis: integrating segmentation and deep learning approaches. Appl. Sci. 14, 7281 (2024).
Mansur, Z., Talukdar, J., Singh, T. P. & Kumar, C. J. Deep learning-based brain tumor image analysis for segmentation. SN COMPUT. SCI. 6, 42 (2025).
Lata, K., Singh, P., Saini, S. & &Cenkeramaddi, L. R. Deep learning-based brain tumor detection in privacy-preserving smart health care systems. IEEE Access. 12, 140722–140733 (2024).
Schwehr, Z. & Achanta, S. Brain tumor segmentation based on deep learning, attention mechanisms, and energy-based uncertainty predictions. Multimed Tools Appl. https://doi.org/10.1007/s11042-024-20443-0 (2025).
Natha, S. et al. Automated brain tumor identification in biomedical radiology images: a multi-model ensemble deep learning approach. Appl. Sci. 14, 2210 (2024).
Tiwari, R. G. et al. Adaptive neuro-FUZZY inference system-fusion-deep belief network for brain tumor detection using MRI images with feature extraction. Biomed. Signal Process. Control. 103, 107387 (2025).
Awudong, B. & Li, Q. Brain tumor automated diagnosis from MR images via attention-guided semi-supervised deep learning. In International Conference on Optics and Machine Vision (ICOMV 2024). 131791SPIE (2024).
Sharif, M. I., Li, J. P., Khan, M. A., Kadry, S. & Tariq, U. M3BTCNet: multi model brain tumor classification using metaheuristic deep neural network features optimization. Neural Comput. Appl. 36 (1), 95–110 (2024).
Hekmat, A., Zhang, Z., Khan, S. U. R. & Bilal, O. Brain tumor diagnosis redefined: Leveraging image fusion for MRI enhancement classification. Biomedical Signal Processing and Control, 109, p.108040. (2025).
Zahid, U. et al. BrainNet: optimal deep learning feature fusion for brain tumor classification. Computational Intelligence and Neuroscience, 2022(1), p.1465173. (2022).
Rastogi, D. et al. Brain tumor detection and prediction in MRI images utilizing a Fine-Tuned transfer learning model integrated within deep learning frameworks. Life, 15(3), p.327. (2025).
Nawaz, M. et al. Melanoma segmentation: A framework of improved DenseNet77 and UNET convolutional neural network. Int. J. Imaging Syst. Technol. 32 (6), 2137–2153 (2022).
Karthik, A. et al. Unified approach for accurate brain tumorMulti-Classification and segmentation through fusion of advanced methodologies. Biomedical Signal Processing and Control, 100, p.106872. (2025).
Mathivanan, S. K. et al. Employing deep learning and transfer learning for accurate brain tumor detection. Scientific Reports, 14(1), p.7232. (2024).
Gao, H. et al. SwinBTC: transfer learning to brain tumor classification for healthcare electronics using augmented MR images. IEEE Trans. Consumer Electronics 71(1), 2297–2308 (2025).
Asiri, A. A. et al. Optimized brain tumor detection: a dual-module approach for mri image enhancement and tumor classification. IEEE Access. 12, 42868–42887 (2024).
Semwal, T., Jain, S., Mohanta, A. & Jain, A. A hybrid CNN-SVM model optimized with PSO for accurate and noninvasive brain tumor classification. Neural Comput. Applications 37, 1–30 (2025).
Brianna, D. F., Kesuma, L. I., Geovani, D. & Sari, P. Combination of Image Enhancement and Double U-Net Architecture for Liver Segmentation in CT-Scan Images. (2025). J. electron.electromedical.eng.med.inform. 7,208–219.
Sundaresan, A. A. & Solomon, A. A. Post-Disaster flooded region segmentation using DeepLabv3 + and unmanned aerial system imagery. Nat. Hazards Res. https://doi.org/10.1016/j.nhres.2024.12.003 (2024).
Kuntiyellannagari, B. & Dwarakanath, B. Glioma segmentation using hybrid filter and modified African Vulture optimization. Bull. EEI. 14, 1447–1455 (2025).
Korkmaz, M. & Kaplan, K. Effectiveness analysis of deep learning methods for breast Cancer diagnosis based on histopathology images. Appl. Sci. 15, 1005 (2025).
Butt, N. et al. Citrus disease detection using innovative deep learning approach and hybrid Meta-Heuristic. PloS One. 20, e0316081 (2025).
Banerjee, S. et al. Accurate prediction of Piperine content in black pepper using combined CNN and regression modelling with PDMAM@G electrode and Cyclic voltammetry. J. Food Compos. Anal. 141, 107355 (2025).
Zheng, R. et al. A new approach for fire and non-fire aerosols discrimination based on multilayer perceptron trained by modified bonobo optimizer. Cluster Comput. 28, 167 (2025).
Figshare brain tumor dataset. https://www.kaggle.com/datasets/ashkhagan/figshare-brain-tumor-dataset
Amjad, U. et al. Context-aware machine learning techniques for brain tumor classification and detection–A review. Heliyon 11, e41835 (2025).
Mithun, M. S. & Jawhar, S. J. Detection and classification on MRI images of brain tumor using YOLO NAS deep learning model. J. Radiation Res. Appl. Sci. 17, 101113 (2024).
Alqhtani, S. M. et al. Improved brain tumor segmentation and classification in brain MRI with FCM-SVM: A diagnostic approach. IEEE Access. 12, 61312–61335 (2024).
Zaitoon, R., Syed, H. &RU-Net2+: A deep learning algorithm for accurate brain tumor segmentation and survival rate prediction. IEEE Access. 11, 118105–118123 (2023).
Author information
Authors and Affiliations
Contributions
Dr. Pradeep Gurunathan : Formal Analysis, Design, Algorithm part, Results and Proof CheckingPreethi Saroj Srinivasan : Problem selection, Algorithm, Implementation, Coding and Testing, Results, EditingDr. Ravimaran S: Formal Analysis, Design, Algorithm part, Results and Proof Checking.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Gurunathan, P., Srinivasan, P.S. & S, R. Deep transfer learning based feature fusion model with Bonobo optimization algorithm for enhanced brain tumor segmentation and classification through biomedical imaging. Sci Rep 15, 34030 (2025). https://doi.org/10.1038/s41598-025-12973-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-12973-w