Abstract
Autism spectrum disorder (ASD), a neurodevelopmental disorder affecting 1% of the global population, is increasingly associated with dysregulation of the microbiota–gut–brain axis. While genetic and environmental factors have been well-studied, the role of gut microbial metabolites in the pathogenesis of ASD remains underexplored. In this study, we integrated network pharmacology, molecular docking, and multi-database analysis to elucidate the molecular mechanisms by which gut microbiota-derived metabolites regulate ASD. Utilizing the gutMGene, GeneCards, and OMIM databases, we identified 51 core targets that intersect with ASD-related genes and gut metabolite targets. Validation of four topological algorithms (Degree, EPC, MCC, MNC) identified AKT1 and IL6 as key pivotal genes, as revealed by protein-protein interaction (PPI) network analysis. Functional enrichment highlighted important associations with the PI3K/Akt and IL-17 signaling pathways. The Microbiome-Metabolite-Target-Signaling (MMTS) network linked eight key metabolites (e.g., short-chain fatty acids, indole derivatives) to AKT1/IL6 regulation. Drug similarity and toxicity assessments confirmed the safety of short-chain fatty acids (acetate, butyrate, propionate) and indole derivatives of the selected metabolites. Molecular docking revealed a strong binding affinity between glycerylcholic acid (AKT1: − 10.2 kcal/mol) and 3-indolepropionic acid (IL6: − 4.9 kcal/mol), suggesting that they are closely related to ASD. This study provides a new research direction on the relationship between microbial metabolites and ASD and gives better help to future researchers.
Similar content being viewed by others
Introduction
ASD is a neurodevelopmental condition characterized by early-onset social communication deficits and restricted, repetitive patterns of behavior or interests1,2. Over the past decade, the prevalence of ASD has shown a gradual increase, currently affecting approximately 1% of the global population3. While the precise etiology remains unclear, traditional research has primarily focused on the interplay between genetic variations, synaptic dysfunction, and environmental factors1,4,5. Notably, accumulating evidence suggests that the microbiota-gut-brain axis plays a crucial role in the pathophysiology of ASD4,6.
The gut microbiota plays a pivotal role in bidirectional communication between the gut and the brain, suggesting its potential involvement in neurodevelopment, neurotransmission, and behavioral regulation, thereby influencing the pathogenesis of various neurodevelopmental, psychiatric, and neurological disorders4. Growing evidence indicates that gut dysbiosis significantly contributes to the development of ASD7,8,9. Clinical studies have found that children with ASD are often accompanied by disturbances in the composition of the intestinal flora (e.g., abnormal Firmicutes/Bacteroidetes ratio), increased intestinal barrier permeability, and changes in serum levels of microbial metabolites10,11. Animal studies have demonstrated that fecal microbiota transplantation from ASD patients can induce social behavior deficits in mice, while probiotic interventions or microbiota modulation may ameliorate certain ASD-like symptoms12. Notably, a clinical trial utilizing microbiota transfer therapy reported symptomatic improvement in 18 children with ASD13,14.These findings highlight the promising therapeutic potential of gut microbiota modulation in ASD. However, the precise mechanisms underlying the microbiota’s therapeutic effects require further investigation.
Network pharmacology, an interdisciplinary approach integrating systems biology and pharmacology15, enables comprehensive analysis through target identification, protein-protein interaction (PPI) network construction, and molecular docking validation16,17,18. Recent network pharmacology studies have successfully identified potential therapeutic metabolites derived from gut microbiota for related disorders19,20. Therefore, we propose to employ network pharmacology methodologies to systematically investigate the molecular mechanisms underlying ASD-gut microbiota interactions, providing novel insights for future ASD-microbiota research. The workflow is represented in Fig. 1.
Methods
Selection of gut microbial metabolites and targets
Using the gutMGene v2.0 database (accessed February 2025)21(http://bio-annotation.cn/gutmgene), we retrieved information on human gut microbiota, gut metabolites, and their associated targets. The SMILES representations of metabolites were obtained from PubChem (https://pubchem.ncbi.nlm.nih.gov/)22. To identify target genes for each metabolite, we utilized the Swiss Target Prediction (STP) database (accessed February 2025) (https://www.swisstargetprediction.ch/)23 and the Similarity Ensemble Approach (SEA) database (accessed February 2025) (https://sea.bkslab.org/)24. Finally, the Evenn online tool (http://www.ehbio.com/)25 was employed to integrate and intersect the metabolite target genes identified from both databases, yielding the final set of gut metabolite-associated target genes. All data were sourced from database versions released before March 2025.
Search for targets of ASD
We accessed the GeneCards database26 (https://www.genecards.org/) and the Online Mendelian Inheritance in Man (OMIM) database27 (https://www.omim.org/) on February 22, 2025, using “autism spectrum disorder” as the keyword to identify ASD genes. For GeneCards, we set a relevance score of ≥ 10 as the screening threshold. The two datasets were merged into a union set, and duplicate genes were removed to establish the ASD gene set. Next, we intersected the ASD gene set with the gut metabolite targets and further cross-referenced the overlapping genes with human intestinal targets in the gutMGene database to identify potential therapeutic targets for ASD based on gut metabolites. All data were sourced from database versions released before March 2025.
PPI network construction and hub gene screening
Protein–protein interaction (PPI) analysis of the 51 ASD-related host genes of gut microbial metabolites (AHGGMM) was performed using the STRING database (https://string-db.org/)28,29. To identify hub genes within the PPI network, we utilized the CytoHubba plugin in Cytoscape30, applying four topological analysis algorithms: Degree, Edge Percolated Component (EPC), Maximal Clique Centrality (MCC), and Maximum Neighborhood Component (MNC). The Degree algorithm reflects the centrality of a node based on its number of direct connections. The EPC algorithm evaluates the stability of a node within the network, with higher EPC scores indicating inclusion in more robust subnetworks. MCC is a highly sensitive and accurate method for detecting key nodes in complex networks. MNC highlights nodes that occupy central positions in densely connected local regions31. The highest-scoring targets identified across all four algorithms were then selected as the most valuable protein-coding targets for ASD32.
Gene ontology (GO) and kyoto encyclopaedia of genes and genomes (KEGG) enrichment analysis
GO analysis characterizes gene sets through cellular components (CC), molecular functions (MF) and biological processes (BP) to uncover biological significance33. KEGG, an integrated database of genomic and chemical information, provides metabolic pathway maps for understanding gene functions34,35,36. KEGG enrichment reveals gene set involvement in metabolic pathways. We used the Sangerbox online tool37 (http://www.sangerbox.com/) to perform GO and KEGG enrichment analyses and visualize the results for the 51 overlapping genes. A threshold of P < 0.05 was considered statistically significant for the enrichment results.
Microbiota-metabolites-targets-signaling (MMTS) pathways network analysis
To comprehensively elucidate the relationships among gut microbiota, metabolites, the core targets (AKT1 and IL6), and their signaling pathways, we first utilized the gutMGene database to identify gut microbial communities and metabolites directly associated with the core targets AKT1 and IL6. We then further queried the gutMGene database for additional metabolites or microbes associated with these gut microbial and metabolites, thereby indirectly identifying potential gut-derived regulators of AKT1 and IL6. KEGG pathway enrichment analysis was performed on the final set of target genes, and the KEGG pathways involving AKT1 and IL6 were used to construct the MMTS network. The MMTS network was visualized using Cytoscape (version 3.10.3).
Prediction of drug similarity and toxicity parameters of metabolites
The drug-likeness is determined based on the Lipinski’s rule of five: (1) molecular weight < 500; (2) lipid-water partition coefficient < 5; (3) hydrogen bond acceptor count < 5; (4) hydrogen bond donor count < 5; (5) polar surface area < 140. Metabolites meeting Lipinski’s Rule of Five criteria were selected for further analysis through the SwissADME platform38. Thus, we confirmed the six parameters by using ADMETlab 3.0 platform39: hERG blockers obstruct potassium channels40, cause human hepatotoxicity41, ames mutagenicity42, skin sensitization43, Lethal Dose 50 (LD50) of acute toxicity44, and Drug Induced Liver Injury (DILI)45.
The docking testing of metabolites and targets for molecular
To evaluate the interaction and binding of intestinal metabolites with the core targets AKT1 and IL6, we obtained the protein structure files of the core targets AKT1 (PDB ID: 3O96) and IL6 (PDB ID: 4O9H) from the PDB46 database, and removed co-crystalline ligands, ions, and water molecules from the protein structures of the core targets using PyMOL software. PyMOL software was used to remove co-crystalline ligands, ions and water molecules from the protein structure of the core targets, and polar hydrogen and Kollman charges were added using AutoDock. Core metabolites were obtained as sdf format files from Pubchem and pre-docking processed using Chembio3D Ultra. The core metabolites were then analyzed to determine the binding pockets of the core target structures for molecular docking using AutoDock Vina47. The center coordinates of the two key targets are: AKT1: x = − 15.877, y = 9.362, z = 37.115 and IL-6: x = 11.214, y = 33.473, z = 11.159. The docking site was set in cubic box (x = 40 Å, y = 40 Å, and z = 40 Å) in a central point of each target. and the binding energies were calculated and finally combined with the Protein-Ligand Interaction Profiler online tool (https://plip-tool.biotec.tu-dresden.de/plip-web/plip/index) with PyMOL software to visualize them.
Results
Identification of ASD-Related host genes of gut microbial metabolites
We first retrieved 278 gut microbial metabolites and 238 human intestinal targets from the gutMGene v2.0 database. Using SEA and STP databases, we predicted the targets of the 278 metabolites and identified 1323 and 1040 targets, respectively. Taking the intersection of these two datasets yielded 755 gut microbial metabolite target genes (Fig. 2A). To identify ASD-related genes, we queried the GeneCards and OMIM databases with the term “autism spectrum disorder.” After merging and removing duplicates, a total of 4,576 ASD-related genes were obtained. Intersecting these with the 755 gut microbial metabolite targets resulted in 399 genes associated with both gut microbial metabolites and ASD (Fig. 2B). Further comparison of these 399 genes with the 238 host intestinal targets revealed 51 common genes, which were defined as ASD-related host genes of gut microbial metabolites (AHGGMM) (Fig. 2C) (Supplementary Table S1–S3).
(A) The 755 gut microbial metabolite target genes. (B) The 399 overlapping genes (ASD-related gut microbial metabolite target genes) between the 755 gut metabolite target genes and the 4576 ASD genes. (C) The 51 overlapping genes (ASD-related host genes of gut microbial metabolites) between the 399 ASD-related gut microbial metabolite target genes and the 238 host genes.
Enrichment analysis
GO and KEGG enrichment analyses were performed on the 51 AHGGMM, identifying a total of 122 signaling pathways, 210 biological processes (BP), 72 molecular functions (MF), and 24 cellular components (CC). GO enrichment analysis showed that the most significantly enriched BPs were response to stress, cellular response to chemical stimulus, and response to organic substance (Fig. 3A), suggesting the involvement of these genes in host adaptive responses modulated by gut microbial metabolites. The top enriched CC terms were protein-containing complex, nuclear part, and nucleoplasm (Fig. 3B), indicating a predominant localization of these genes in nuclear and protein complexes. For MF, the top terms included enzyme binding, transcription factor binding, and RNA polymerase II transcription factor binding (Fig. 3C), highlighting their regulatory roles in transcriptional processes. KEGG pathway analysis revealed that these genes were primarily enriched in signaling pathways such as the AGE-RAGE signaling pathway in diabetic complications, IL-17 signaling pathway, and PI3K/Akt signaling pathway. In addition, enrichment in human disease-related pathways, including pathways in cancer and Yersinia infection, indirectly reflected the strong association of these genes with inflammatory processes (Fig. 3D).
Protein–protein interaction network analysis
We subjected the 51 AHGGMM obtained to PPI network analysis, the total of 114 nodes and 144 edges were identified in the network (Fig. 4A). The top 10 genes in the four algorithms of “Degree”, “EPC”, “MCC” and “MNC” were ranked as hub genes, among which AKT1 and IL6 proteins had the highest scores in all four algorithms (Fig. 4B,E). Indicating that they play a pivotal role in the gene network of intestinal flora in the regulation of ASD. Therefore, we chose AKT1 and IL-6 as the core targets of ASD regulation by gut flora metabolites in our subsequent studies. (Table 1)
The MMTS network analysis
We queried the gutMGene database to identify gut microbes and metabolites associated with the two core targets, AKT1 and IL6. The results showed that IL6 was directly associated with six gut microbes and five gut-derived metabolites, whereas AKT1 was associated with three metabolites: vancomycin, glycocholic acid, and indole. We further explored other metabolites or microbes functionally linked to these directly associated factors in the gutMGene database, thereby indirectly identifying gut-derived factors with potential regulatory effects on AKT1 and IL6. KEGG pathway enrichment analysis of 51 AHGGMM revealed enrichment in 122 signaling pathways, including 28 pathways involving AKT1 and 10 pathways involving IL6. Based on these findings, we constructed a MMTS network centered on the two core targets, AKT1 and IL6, comprising 72 gut microbes, 12 gut metabolites, and 30 signaling pathways (Fig. 5). (Supplementary Table S1–S3).
The MMTS network. The orange nodes represent the two core genes, the pink nodes represent gut microbes, the blue nodes represent gut metabolites, and the green nodes represent signaling pathways. The edges indicate the associations between two nodes. hsa04151: PI3K-Akt signaling pathway, hsa04933: AGE-RAGE signaling pathway in diabetic complications, hsa04625: C-type lectin receptor signaling pathway, hsa04066: HIF-1 signaling pathway, hsa04620: Toll-like receptor signaling pathway, hsa04668: TNF signaling pathway, hsa04926: Relaxin signaling pathway, hsa04010: MAPK signaling pathway, hsa04660: T cell receptor signaling pathway, hsa04012: ErbB signaling pathway, hsa04722: Neurotrophin signaling pathway, hsa04919: Thyroid hormone signaling pathway, hsa04071: Sphingolipid signaling pathway, hsa04915: Estrogen signaling pathway, hsa04024: cAMP signaling pathway, hsa04014: Ras signaling pathway, hsa04370: VEGF signaling pathway, hsa04917: Prolactin signaling pathway, hsa04662: B cell receptor signaling pathway, hsa04068: FoxO signaling pathway, hsa04022: cGMP-PKG signaling pathway, hsa04062: Chemokine signaling pathway, hsa04910:Insulin signaling pathway, hsa04371: Apelin signaling pathway, hsa04072: Phospholipase D signaling pathway hsa04630: JAK-STAT signaling pathway, hsa04664: Fc epsilon RI signaling pathway, hsa04920: Adipocytokine signaling pathway, hsa04657: IL-17 signaling pathway, hsa04621: NOD-like receptor signaling pathway.
Prediction of drug similarity and toxicity parameters of metabolites
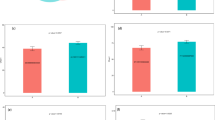
We further evaluated the drug-like properties and toxicity of the major metabolites. Among these eight metabolites, vancomycin violated Lipinski’s rule of five in its ADME parameters, particularly due to its topological polar surface area, which does not meet the characteristics of orally active drugs. The remaining Indole, glycocholic acid, 3- Indole, glycocholic acid, 3-Indolepropionic acid, acetate, butyrate, trimethylamine oxide, propionate, metabolites are all in accordance with the Lipinski’s five rules (Table 2). Toxicity testing results (Table 3) showed that acetate, butyrate, trimethylamine oxide, and propionate were positive for H-HT, suggesting potential hepatotoxicity.
The docking testing of metabolites and targets for molecular
We evaluated the binding affinities of AKT1 with two metabolites, glycocholic acid and indole, as well as IL6 with five metabolites, including 3-indolepropionic acid, acetate, butyrate, propionate, and trimethylamine oxide, in co-crystallized complexes. For AKT1 (PDB ID: 3O96), glycocholic acid exhibited the highest binding energy (− 10.2 kcal/mol), followed by indole (− 5.9 kcal/mol). Glycocholic acid promoted the formation of the glycocholic acid–AKT1 complex (Fig. 6) through hydrogen bonding interactions with GLN79, VAL83, ASP292, and GLY294, as well as hydrophobic interactions with GLN79, TRP80, THR82, VAL270, TYR272, and ARG273. In contrast, for IL6 (PDB ID: 4O9H), 3-indolepropionic acid showed the strongest binding affinity, with a binding energy of − 4.9 kcal/mol (Table 4).
Discussion
In recent years, the complex interactions between gut microbial metabolites and the pathogenesis of ASD have attracted increasing research attention, with the gut-brain axis emerging as a crucial regulatory mechanism in ASD48. While existing studies have demonstrated the therapeutic potential of gut microbiota in ASD treatment11,49, the specific metabolites involved and their molecular targets remain unclear. Recent advances in network-based systems pharmacology have enabled the investigation of relationships between gut microbial metabolites and disease mechanisms50. Our study systematically elucidates the molecular mechanisms underlying ASD regulation by gut microbial metabolites through an integrated approach combining multi-database analysis, network pharmacology, and molecular docking.
During the data collection process, we collected microbial and disease gene targets through three different platforms, gutMGene21, Genecard26 and OMIM27 three different platforms collected microbial and disease gene targets, and 51 key targets were obtained after analysis. GO enrichment analysis of these target genes revealed that ASD-related targets were significantly enriched in processes such as stress response and chemical stimulus response, emphasizing the role of gut-derived metabolites in regulating host adaptation to environmental challenges. And KEGG pathway analysis further identified PI3K/Akt and IL-17 as the core pathways of ASD.
We then performed a PPI network analysis, in which we used four algorithms to identify AKT1 and IL6 as key targets of gut microbial metabolites regulating ASD. Notably, AKT1 is a core component of the PI3K/Akt pathway, and impairment of its PI3K/Akt signaling pathway may lead to defective synaptic pruning and abnormal neuronal connectivity to affect ASD51. Its downstream AKT-mTOR signaling pathway is a common therapeutic target in ASD52. AKT1 is a core member of the AKT/PKB (protein kinase B) signaling pathway, which plays an important role in the regulation of neural development, synaptic plasticity and metabolic homeostasis53,54,55. In addition, AKT1 plays a key role in the PI3K-AKT-mTOR signaling pathway, which is involved in the regulation of a variety of biological processes56. Abnormalities in the mTOR signaling pathway are considered to be one of the potential pathogenic mechanisms of ASD, and a large body of evidence suggests that the mTOR inhibitor rapamycin is effective in alleviating social dysfunction in a variety of animal models of ASD52,57,58,59. Dysregulation of AKT1 is associated with neurodevelopmental disorders including ASD. In addition, the expression level of AKT1 in the amygdala correlates with emotional memory learning, suggesting that it may be involved in social interaction disorders in ASD patients60.
IL6, a pro-inflammatory cytokine, has also been identified as another core target. In the IL-17 signaling pathway, transactivation of the IL-6 receptor promotes the release of IL-17 A, and IL-6 is an essential cytokine for the polarization of Th0 cells into Th17 cells, which makes it involved in ASD expression61. IL-6 levels were significantly elevated in frozen brain tissue from ASD patients compared to healthy subjects62. In a study of ASD, Weili Wu et al. found that IL-6 activation in the maternal placenta is a necessary mediator that affects fetal brain development and impairs their ability to explore socially63. Similarly, in patients with ASD, elevated levels of IL6 are associated with neuroinflammation, microglia activation, and behavioral abnormalities61. Recent studies have demonstrated that IL-6 enhances activation of the PI3K-AKT/mTOR-GSK-3β pathway in hippocampal neurons of ASD mice by upregulating GRPR64. In summary, AKT1 and IL6 play important roles in the pathogenesis of autism, and the present study found that intestinal microbial metabolites could exert therapeutic effects on autism mainly by affecting AKT1 through network pharmacological analysis.
In MMTS network analysis, we identified three metabolites (indole, glycylcholic acid, and vancomycin) associated with AKT1, and five metabolites (3-indolepropionic acid, acetic acid, butyric acid, trimethylamine oxide, and propionic acid) associated with IL6. We hypothesized that these eight metabolites may activate the related key genes (AKT1 and IL6) through different mechanisms, which in turn are directly or indirectly involved in the pathological process of ASD through signaling pathways. Through research reports, we found that the levels of the metabolite TMAO were associated with cardiovascular diseases65, NAFLD66,67 and metabolic disorders (e.g., type 2 diabetes mellitus)68, etc. In addition, a recent study reported that TMAO was observed in the urine of children with ASD69,70. In contrast, butyrate, acetate, and propionate—classified as short-chain fatty acids (SCFAs)—have been reported to alleviate ASD-related symptoms71,72 and reduce inflammation73. 3-indolylpropionic acid and indoles are neuroprotective and enhance intestinal barrier function, helping to reduce neuroinflammation and decrease intestinal permeability, thereby alleviating ASD symptoms74,75. Based on the above findings, we hypothesize that these metabolites may be potentially involved in the pathogenesis of ASD. Their key genes may affect the development of ASD by activating specific signaling pathways. This hypothesis needs to be further verified in future studies.
In this study, we further identified AKT1 and IL6 as key targets within the network of ASD-related host genes influenced by gut microbial metabolites through PPI network analysis. These targets are likely to play significant roles in the regulation of autism spectrum disorder (ASD) onset and progression by gut metabolites. Correlation analysis revealed direct associations between AKT1/IL6 and eight metabolites. To further assess the potential regulatory capacity of these metabolites, we employed molecular docking to predict their binding strength to the two target genes. The results demonstrated that glycodeoxycholic acid, produced by Escherichia coli, Prevotella, Wickerhamomyces, and Akkermansia, exhibited the strongest binding affinity to AKT1 (− 10.2 kcal/mol), indicating its potential as a key regulatory metabolite for AKT1 function. Notably, Akkermansia, recognized as a probiotic genus, exerts functions that enhance intestinal barrier integrity and modulate the immune system76,77. Previous studies have reported that Akkermansia ameliorates aberrant behaviors and intestinal dysfunction in mouse models of autism78,79. Consistent with the results of studies showing that the bile acid derivative glycocholic acid can attenuate gastrointestinal tract via metabolic and inflammatory pathways, its effects on ASD should be investigated and it could be very interesting80. The molecular docking results not only provide a structural basis for the predicted microbe–host interactions but also lay the foundation for subsequent screening of candidate therapeutic agents. In conclusion, integrating network pharmacology with molecular docking not only enhances the reliability of target prediction but also provides a theoretical framework for developing microbiota-based intervention strategies for ASD. This finding provides new clues to the mechanism of association between intestinal metabolites and ASD, and future studies could focus on their potential role in the development of ASD.
Furthermore, network pharmacology leverages extensive omics and public databases, enabling it to capture and visualize multidimensional interactions between metabolites and multiple host targets/pathways. This approach overcomes the limitations inherent in traditional single-target research and aligns better with the complex pathophysiology of ASD. Through topological network analysis, it also facilitates the identification of critical regulatory nodes (AKT1 and IL6) from numerous potential targets and predicts potential relationships among metabolites, targets, and pathways. This provides a theoretical basis for subsequent experimental validation81. These advantages underscore the utility of network pharmacology in revealing mechanisms underlying gut microbiota-host interactions in ASD.
Future perspectives
Although this study elucidates the potential mechanisms by which gut microbial metabolites regulate ASD-associated pathways, several challenges and opportunities remain. Firstly, the identified core genes and metabolites require experimental validation in cellular and animal models to confirm their roles in ASD pathogenesis. Secondly, longitudinal clinical studies are needed to evaluate whether modulating specific gut microbiota or supplementing targeted metabolites can ameliorate ASD symptoms. Furthermore, the potential hepatotoxicity risks associated with certain metabolites warrant careful evaluation in therapeutic applications. Future research should also explore how inter-individual variations in gut microbial composition and metabolism impact treatment efficacy. Integrating multi-omics data, including metagenomics, metabolomics, and transcriptomics, holds promise for refining our understanding of the gut-brain axis in ASD. These efforts will ultimately advance the development of precise and safe gut microbiota-based management strategies for ASD.
Conclusion
This study employed a network pharmacology approach to systematically identify and analyze the potential regulatory mechanisms of gut microbial metabolites in autism spectrum disorder (ASD). We identified a total of 51 ASD-associated host genes targeted by gut microbial metabolites, which were enriched in pathways such as PI3K/Akt and IL-17. Further protein-protein interaction (PPI) network analysis highlighted AKT1 and IL6 as core hub genes. Centered on these two key targets, we constructed a Microbe-Metabolite-Target-Signaling (MMTS) network, revealing complex interactions among gut microbiota, metabolites, and host signaling pathways in ASD pathogenesis. Molecular docking results further validated the high binding affinity between key metabolites and these core targets, suggesting their potential functional significance. Collectively, this study provides novel insights into the molecular interactions between gut microbiota and the host in ASD, and proposes AKT1 and IL6 as potential targets for microbiota-based interventions.
Data availability
The database described in the article is available upon reasonable request from the corresponding author.
References
Lai, M. C., Lombardo, M. V. & Baron-Cohen, S. Autism . Lancet. 383(9920), 896–910 (2014).
Lord, C., Elsabbagh, M., Baird, G. & Veenstra-Vanderweele, J. Autism spectrum disorder. Lancet 392 (10146), 508–520 (2018).
Zeidan, J. et al. Global prevalence of autism: A systematic review update. Autism Res. 15 (5), 778–790 (2022).
Socała, K. et al. The role of microbiota-gut-brain axis in neuropsychiatric and neurological disorders. Pharmacol. Res. 172, 105840 (2021).
Modabbernia, A., Velthorst, E. & Reichenberg, A. Environmental risk factors for autism: an evidence-based review of systematic reviews and meta-analyses. Mol. Autism. 8, 13 (2017).
Mehra, A. et al. Gut microbiota and autism spectrum disorder: from pathogenesis to potential therapeutic perspectives. J. Tradit. Compl. Med. 13 (2), 135–149 (2023).
Fattorusso, A., Di Genova, L., Dell’Isola, G. B., Mencaroni, E. & Esposito, S. Autism spectrum disorders and the gut microbiota. Nutrients. 11(3). (2019).
Iglesias-Vázquez, L., Van Ginkel, R. G., Arija, V. & Canals, J. Composition of gut microbiota in children with autism spectrum disorder: A systematic review and meta-analysis. Nutrients. 12(3). (2020).
Mangiola, F. et al. Gut microbiota in autism and mood disorders. World J. Gastroenterol. 22 (1), 361–368 (2016).
Wan, Y. et al. Underdevelopment of the gut microbiota and bacteria species as non-invasive markers of prediction in children with autism spectrum disorder. Gut 71 (5), 910–918 (2022).
Strati, F. et al. New evidences on the altered gut microbiota in autism spectrum disorders. Microbiome. 5 (1), 24 (2017).
Sharon, G. et al. Human gut microbiota from autism spectrum disorder promote behavioral symptoms in mice. Cell. 177(6), 1600–1618 (2019).
Kang, D. W. et al. Long-term benefit of microbiota transfer therapy on autism symptoms and gut microbiota. Sci. Rep.-UK. 9 (1), 5821 (2019).
Kang, D. W. et al. Microbiota transfer therapy alters gut ecosystem and improves Gastrointestinal and autism symptoms: an open-label study. Microbiome. 5 (1), 10 (2017).
Noor, F. et al. Integrating network pharmacology and molecular docking approaches to decipher the multi-target pharmacological mechanism of Abrus precatorius L. acting on diabetes. Pharmaceut.-Base 15(4). (2022).
Noor, F. et al. Recent advances in diagnostic and therapeutic approaches for breast cancer: A comprehensive review. Curr. Pharm. Des. 27 (20), 2344–2365 (2021).
Noor, F. et al. Designing a multi-epitope vaccine against chlamydia pneumoniae by integrating the core proteomics, subtractive proteomics and reverse vaccinology-based immunoinformatics approaches. Comput. Biol. Med. 145, 105507 (2022).
Zhai, Z., Tao, X., Alami, M. M., Shu, S. & Wang, X. Network pharmacology and molecular docking combined to analyze the molecular and pharmacological mechanism of Pinellia Ternata in the treatment of hypertension. Curr. Issues Mol. Biol. 43 (1), 65–78 (2021).
Cao, M., Huang, P., Xu, L. S. & Zhang, Y. H. Analysis of gut microbiota-derived metabolites regulating pituitary neuroendocrine tumors through network Pharmacology. Front. Pharmacol. 15, 1403864 (2024).
Yao, W., Huo, J., Ji, J., Liu, K. & Tao, P. Elucidating the role of gut microbiota metabolites in diabetes by employing network pharmacology. Mol. Med. 30 (1), 263 (2024).
Qi, C. et al. GutMGene v2.0: an updated comprehensive database for target genes of gut microbes and microbial metabolites. Nucleic Acids Res. 53 (D1), D783–D788 (2025).
Oh, K. K. et al. The identification of metabolites from gut microbiota in NAFLD via network Pharmacology. Sci. Rep.-UK. 13 (1), 724 (2023).
Daina, A., Michielin, O. & Zoete, V. SwissTargetPrediction: updated data and new features for efficient prediction of protein targets of small molecules. Nucleic Acids Res. 47 (W1), W357–W364 (2019).
Wang, Z., Liang, L., Yin, Z. & Lin, J. Improving chemical similarity ensemble approach in target prediction. J. Cheminformatics. 8, 20 (2016).
Yang, M., Chen, T., Liu, Y. X. & Huang, L. Visualizing set relationships: evenn’s comprehensive approach to Venn diagrams. Imeta 3 (3), e184 (2024).
Stelzer, G. et al. The genecards suite: from gene data mining to disease genome sequence analyses. Curr. Protoc. Bioinf. 54, 1–30 (2016).
Hamosh, A., Scott, A. F., Amberger, J. S., Bocchini, C. A. & McKusick, V. A. Online Mendelian inheritance in man (OMIM), a knowledgebase of human genes and genetic disorders. Nucleic Acids Res. 33 (Database issue), D514–D517 (2005).
Sikić, M., Tomić, S. & Vlahovicek, K. Prediction of protein-protein interaction sites in sequences and 3D structures by random forests. Plos Comput. Biol. 5 (1), e1000278 (2009).
Szklarczyk, D. et al. The STRING database in 2021: customizable protein-protein networks, and functional characterization of user-uploaded gene/measurement sets. Nucleic Acids Res. 49 (D1), D605–D612 (2021).
Otasek, D., Morris, J. H., Bouças, J., Pico, A. R. & Demchak, B. Cytoscape automation: empowering workflow-based network analysis. Genome Biol. 20 (1), 185 (2019).
Chin, C. H. et al. CytoHubba: identifying hub objects and sub-networks from complex interactome. BMC Syst. Biol. 8 (Suppl 4), S11 (2014).
Tang, Y., Li, M., Wang, J., Pan, Y. & Wu, F. X. CytoNCA: a cytoscape plugin for centrality analysis and evaluation of protein interaction networks. BIosystems 127, 67–72 (2015).
Mi, H., Muruganujan, A., Ebert, D., Huang, X. & Thomas, P. D. PANTHER version 14: more genomes, a new PANTHER GO-slim and improvements in enrichment analysis tools. Nucleic Acids Res. 47 (D1), D419–D426 (2019).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28 (1), 27–30 (2000).
Kanehisa, M. Toward understanding the origin and evolution of cellular organisms. Protein Sci. 28 (11), 1947–1951 (2019).
Kanehisa, M., Furumichi, M., Sato, Y., Kawashima, M. & Ishiguro-Watanabe, M. KEGG for taxonomy-based analysis of pathways and genomes. Nucleic Acids Res. 51 (D1), D587–D592 (2023).
Chen, D. et al. Sangerbox 2: enhanced functionalities and update for a comprehensive clinical bioinformatics data analysis platform. Imeta 3 (5), e238 (2024).
Daina, A., Michielin, O. & Zoete, V. SwissADME: a free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci. Rep.-UK. 7, 42717 (2017).
Fu, L. et al. ADMETlab 3.0: an updated comprehensive online ADMET prediction platform enhanced with broader coverage, improved performance, API functionality and decision support. Nucleic Acids Res. 52 (W1), W422–W431 (2024).
Lamothe, S. M., Guo, J., Li, W., Yang, T. & Zhang, S. The human Ether-a-go-go-related gene (hERG) potassium channel represents an unusual target for Protease-mediated damage. J. Biol. Chem. 291 (39), 20387–20401 (2016).
Mulliner, D. et al. Computational models for human and animal hepatotoxicity with a global application scope. Chem. Res. Toxicol. 29 (5), 757–767 (2016).
Xu, C. et al. In silico prediction of chemical Ames mutagenicity. J. Chem. Inf. Model. 52 (11), 2840–2847 (2012).
Alves, V. M. et al. Predicting chemically-induced skin reactions. Part I: QSAR models of skin sensitization and their application to identify potentially hazardous compounds. Toxicol. Appl. Pharm. 284 (2), 262–272 (2015).
Lei, T. et al. ADMET evaluation in drug discovery: 15. Accurate prediction of rat oral acute toxicity using relevance vector machine and consensus modeling. J. Cheminformatics. 8, 6 (2016).
Xu, Y. et al. Deep learning for drug-induced liver injury. J. Chem. Inf. Model. 55 (10), 2085–2093 (2015).
Berman, H. M. et al. The protein data bank. Nucleic Acids Res. 28 (1), 235–242 (2000).
Trott, O. & Olson, A. J. AutoDock vina: improving the speed and accuracy of Docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 31 (2), 455–461 (2010).
Chen, P. et al. Brain-gut axis and psychiatric disorders: A perspective from bibliometric and visual analysis. Front. Immunol. 13, 1047007 (2022).
Li, Q. & Zhou, J. M. The microbiota-gut-brain axis and its potential therapeutic role in autism spectrum disorder. Neuroscience. 324, 131–139 (2016).
Gocho, Y. et al. Network-based systems pharmacology reveals heterogeneity in LCK and BCL2 signaling and therapeutic sensitivity of T-cell acute lymphoblastic leukemia. Nat. Cancer. 2 (3), 284–299 (2021).
Zhou, J. et al. Pharmacological Inhibition of mTORC1 suppresses anatomical, cellular, and behavioral abnormalities in neural-specific Pten knock-out mice. J. Neurosci. 29 (6), 1773–1783 (2009).
Xing, X. et al. Suppression of Akt-mTOR pathway rescued the social behavior in Cntnap2-deficient mice. Sci. Rep.-UK. 9 (1), 3041 (2019).
Lee, C. C., Huang, C. C. & HsuKS Insulin promotes dendritic spine and synapse formation by the PI3K/Akt/mTOR and Rac1 signaling pathways. Neuropharmacology 61 (4), 867–879 (2011).
Lee, H. G., Zhao, N., Campion, B. K., Nguyen, M. M. & Selleck, S. B. Akt regulates glutamate receptor trafficking and postsynaptic membrane elaboration at the drosophila neuromuscular junction. Dev. Neurobiol. 73 (10), 723–743 (2013).
Dudek, H. et al. Regulation of neuronal survival by the serine-threonine protein kinase Akt. Science 275 (5300), 661–665 (1997).
Woodgett, J. R. Recent advances in the protein kinase B signaling pathway. Curr. Opin. Cell. Biol. 17 (2), 150–157 (2005).
Ehninger, D. & Silva, A. J. Rapamycin for treating tuberous sclerosis and autism spectrum disorders. Trends Mol. Med. 17 (2), 78–87 (2011).
Burket, J. A., Benson, A. D., Tang, A. H. & Deutsch, S. I. Rapamycin improves sociability in the BTBR T(+)Itpr3(tf)/J mouse model of autism spectrum disorders. Brain Res. Bull. 100, 70–75 (2014).
Zhang, J., Liu, L. M. & Ni, J. F. Rapamycin modulated brain-derived neurotrophic factor and B-cell lymphoma 2 to mitigate autism spectrum disorder in rats. Neuropsych Dis. Treat. 13, 835–842 (2017).
Todd, R. M. & Anderson, A. K. Six degrees of separation: the amygdala regulates social behavior and perception. Nat. Neurosci. 12 (10), 1217–1218 (2009).
Nadeem, A. et al. Dysregulation in IL-6 receptors is associated with upregulated IL-17A related signaling in CD4 + T cells of children with autism. Prog. Neuro-Psychoph. 97, 109783 (2020).
Li, X. et al. Elevated immune response in the brain of autistic patients. J. Neuroimmunol. 207 (1–2), 111–116 (2009).
Wu, W. L., Hsiao, E. Y., Yan, Z., Mazmanian, S. K. & Patterson, P. H. The placental interleukin-6 signaling controls fetal brain development and behavior. Brain Behav. Immun. 62, 11–23 (2017).
Li, H. et al. IL-6 enhances the activation of PI3K-AKT/mTOR-GSK-3β by upregulating GRPR in hippocampal neurons of autistic mice. J. Neuroimmune Pharm. 19 (1), 12 (2024).
Zhang, H. et al. Intestinal flora metabolite trimethylamine oxide is inextricably linked to coronary heart disease. J. Cardiovasc. Pharm. 81 (3), 175–182 (2023).
He, X., Ji, G., Jia, W. & Li, H. Gut microbiota and nonalcoholic fatty liver disease: insights on mechanism and application of metabolomics. Int. J. Mol. Sci. 17 (3), 300 (2016).
Chu, H., Duan, Y., Yang, L. & Schnabl, B. Small metabolites, possible big changes: a microbiota-centered view of non-alcoholic fatty liver disease. Gut 68 (2), 359–370 (2019).
Wu, Q. et al. Gut microbiota, host lipid metabolism and regulation mechanism of high-fat diet induced mice following different probiotics-fermented wheat Bran intervention. Food Res. Int. 174 (Pt 1), 113497 (2023).
Osredkar, J. et al. Relationship between excreted uremic toxins and degree of disorder of children with ASD. Int J. Mol. Sci 24(8). (2023).
Açıkel, S. B., Kara, A., Bağcı, Z. & Can, Ü. Serum trimethylamine N-oxide and lipopolysaccharide binding protein levels among children diagnosed with autism spectrum disorder. Int. J. Dev. Neurosci. 83 (6), 571–577 (2023).
Bauer, E. K. et al. Synbiotics of encapsulated Limosilactobacillus fermentum K73 promotes in vitro favorable gut microbiota shifts and enhances short-chain fatty acid production in fecal samples of children with autism spectrum disorder. Food Res. Int. 209, 116227 (2025).
He, Y. et al. Synbiotic combination of 2’-fucosyllactose and bifidobacterium mitigates neurodevelopmental disorders and ASD-like behaviors induced by valproic acid. Food Funct. 16 (7), 2703–2717 (2025).
Wu, H. et al. Anemoside B4 alleviates ulcerative colitis by attenuating intestinal oxidative stress and NLRP3 inflammasome via activating Aryl hydrocarbon receptor through remodeling the gut Microbiome and metabolites. Redox Biol. 85, 103746 (2025).
Peralta-Marzal, L. N. et al. The Impact of Gut Microbiota-Derived Metabolites in Autism Spectrum Disorders. Int. J. Mol. Sci. 22(18). (2021).
Wei, W. et al. Psychological stress-induced microbial metabolite indole-3-acetate disrupts intestinal cell lineage commitment. Cell. Metab. 36 (3), 466–483 (2024).
Hu, Y., Zhou, J. & Lin, X. Akkermansia muciniphila helps in the recovery of lipopolysaccharide-fed mice with mild intestinal dysfunction. Front. Microbiol. 16, 1523742 (2025).
Wu, S. et al. Lactic acid bacteria target NF-κB signaling to alleviate gastric inflammation. Food Funct. (2025).
Fang, J., Kang, S. G., Huang, K. & Tong, T. Integrating 16S rRNA gene sequencing and metabolomics analysis to reveal the mechanism of L-proline in preventing autism-like behavior in mice. Nutrients. 17(2). (2025).
Miao, Z., Chen, L., Zhang, Y., Zhang, J. & Zhang, H. Bifidobacterium animalis subsp. Lactis Probio-M8 alleviates abnormal behavior and regulates gut microbiota in a mouse model suffering from autism. MSystems 9 (1), e101323 (2024).
Oktar, B. K. et al. Beneficial effects of glycocholic acid (GCA) on gut mucosal damage in bile duct ligated rats. Inflammation 25 (5), 311–318 (2001).
Ding, Y. et al. Integrating pharmacology and microbial network analysis with experimental validation to reveal the mechanism of composite sophora colon-soluble capsule against ulcerative colitis. Evid-Based Compl. Alt. 2020, 9521073 (2020).
Funding
The authors thank the funding support from the Program of Hunan Human Biobank (2020TP3003), the grants from National Natural Science Foundation of China(8247054292) and Hunan Province Graduate Education Innovative Practice Base (CSU—Shanghai Treatgut Pharmaceutical Technology Co.LTD).
Author information
Authors and Affiliations
Contributions
Conceptualization, W.Y.X. and F.S.Z.; Methodology, W.Y.X.; Formal analysis, F.S.Z.; resources, F.S.Z.; investigation, F.S.Z.; visualization, W.Y.X.; software, W.Y.X.; validation, F.S.Z; funding acquisition, J.F.H.; supervision, J.F.H.; project administration, J.F.H.; writing—original draft, W.Y.X. and F.S.Z.; writing—review and editing, W.Y.X. and Q.T.; All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, F., Xu, W., Tang, Q. et al. The identification of metabolites from gut microbiota in autism spectrum disorder via network pharmacology. Sci Rep 15, 31765 (2025). https://doi.org/10.1038/s41598-025-15921-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-15921-w
Keywords
This article is cited by
-
Gut microbiota analysis in children with autism spectrum disorder and their family members
Scientific Reports (2025)