Abstract
Some viruses encode proteins that mimic host chemokines and/or cytokines to modulate the host immune responses against infection. Human herpesvirus 8 (HHV-8), also termed Kaposi sarcoma-associated herpesvirus (KSHV), encodes three viral CC-type chemokines (vCCLs 1–3) and an interleukin-6 ortholog (vIL-6). Therapeutic targeting of the vCCLs and vIL-6 has yet to be established despite their importance in immune evasion and transformation by HHV-8. The present study aimed to design a robust and straightforward method to assay receptor binding and downstream signaling using a Nano-Luciferase Binary Technology reporter system (NanoBiT); with this approach, we verified the direct interactions of vCCLs 1–3 with their established receptors. Furthermore, screening with a comprehensive set of human chemokine receptors revealed additional vCCL-interacting receptors, including ACKR4. The vCCL-receptor interactions were validated using competition, antibody-mediated neutralization, or mutagenesis of vCCLs. We extended the utility of NanoBiT to assess the intracellular protein-protein interactions involved in vCCL-induced downstream signaling via b-arrestin. Likewise, we used NanoBiT to monitor vIL-6 binding to its receptor gp130 and downstream signaling via STAT3. In sum, we demonstrate the utility of NanoBiT for assessing viral chemo/cytokine-receptor interactions and signaling and its potential in screening for antibodies to treat HHV-8 infection and related diseases.
Similar content being viewed by others
Introduction
Viruses have evolved mechanisms to modulate the host immune system, allowing them to evade detection and persist in the body. Large DNA viruses, including herpesviruses and poxviruses, encode proteins that mimic chemokines and chemokine receptors (CKRs) or bind chemokines1,2,3,4. These viral immunomodulatory proteins can influence the host’s immune responses by disrupting or mimicking host immune cell trafficking and signaling pathways, ultimately facilitating viral replication and spread.
Human herpesvirus 8 (HHV-8), also known as Kaposi sarcoma-associated herpesvirus (KSHV), is the causative agent of endothelial cell-derived Kaposi sarcoma (KS), and at least two lymphoproliferative disorders, primary effusion lymphoma (PEL) and multicentric Castleman’s disease (MCD)5,6,7. HHV-8 encodes three viral CC chemokine ligands (vCCLs): vCCL1 (also known as viral macrophage inflammatory protein 1 (vMIP-I)), vCCL2 (vMIP-II), and vCCL3 (vMIP-III). vCCL1 is recognized as an agonist for the CC chemokine receptor 8 (CCR8)8,9,10,11. In addition, some research suggests that vCCL1 may act as an agonist ligand for CCR312 and CCR513,14. vCCL2 acts not only as a broad-spectrum chemokine antagonist of human CKRs10,15,16,17including CCR1, CCR2, CCR5, XCR1, CX3CR1, and CXCR4, but also as an agonist of CCR811,18, CCR319, and the atypical chemokine receptor ACKR320. vCCL3 is an agonist ligand for CCR421 and XCR110. As CCR8, CCR4, and CCR3 are preferentially expressed in Th2 lymphocyte22,23,24the viral chemokines are believed to drive the antiviral immune response toward Th2, while vCCL2 hinders the recruitment of Th1-biased (cytotoxic) lymphocytes as well as B cells and NK cells via its antagonistic function. vCCL1 and vCCL2 also promote the survival of infected cells via CCR8 by countering pro-apoptotic signaling during productive viral replication via autocrine signaling11. Moreover, the vCCLs can promote the growth and survival of the surrounding majority of latently infected cells in KS11.
Despite the potential functional importance of the vCCLs, further research is needed to fully understand their impact on the immune responses to HHV-8 infection and associated pathogenesis. With the recent discovery of new human atypical CKRs (ACKRs), a comprehensive screen of a full panel of human CKRs is warranted to verify and/or identify novel CKRs that interact directly with vCCLs. A highly sensitive, robust, and quantitative approach that can be applied to viable cells would greatly facilitate screening for such direct interactions with the vCCLs. Such a cell-based approach would also be helpful for the discovery and development of drugs and neutralizing antibodies (vaccines) to target vCCL-CKR interactions.
In the present study, we employed Nano-Luciferase (NLuc) Binary Technology (NanoBiT)25a protein fragment complementation assay using reconstituted NLuc enzyme activity as a reporter, to re-evaluate the interactions of HHV-8 vCCLs 1–3 with a comprehensive set of human CKRs. We validated the NanoBiT method by evaluating its performance in assessing previously established vCCL-CKR interactions. Subsequently, we uncovered novel interactions between vCCLs and CKRs, and utilized competition with host chemokine, neutralization of vCCLs by antibodies, and vCCL mutagenesis to validate these putative vCCL-CKR interactions. Moreover, we examined vCCL-induced interactions between CKR and b-arrestin 2, an adaptor crucial in regulating CKR signaling and receptor desensitization following agonist activation26. Finally, we employed NanoBiT to monitor gp130 receptor binding and consequent interaction of gp130 with STAT3 elicited by HHV-8 viral interleukin 6 (vIL-6), a cytokine critical for viral biology and pathogenesis27,28,29,30.
Results
NanoBiT assessment of the interactions between HHV-8 vCCLs 1–2 and CCR8
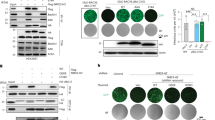
NanoBiT comprises two subunits derived from Nano-Luciferase (NLuc): a Large BiT (LgB; 17.6 kDa) and a Small BiT (SmB; 1.3 kDa)25. When chemokine and CKR fused with the complementary NanoBiT subunits interact, the subunits come into proximity and can then form an active enzyme, resulting in the emission of a bright luminescent signal in the presence of the NLuc substrate furimazine (Fig. 1A).
The protein fragment complementation assay of the interactions between HHV-8 viral chemokines and CCR8. (A) Illustration of the NanoBiT-based protein fragment complementation assay of chemokine-receptor interaction. When a chemokine binds to its receptor on the plasma membrane of intact cells, their fused Small BiT (SmB; 1.3 kDa) and Large BiT (LgB; 17.6 kDa) fragments of Nano Luciferase (NLuc) are brought into proximity, which allows structural complementation thus yielding a functional enzyme acting on NLuc substrate, furimazine. (B) Schematic diagram of NanoBiT-fused CCR8. CCR8 was tagged with the bovine prolactin signal sequence (PSS) and the V5 epitope for efficient translocation into the endoplasmic reticulum and immunological detection of expression, respectively. (C) Immunofluorescence detection of cell surface expression of the NanoBiT-fused CCR8 protein transfected into 293T cells. DAPI (4’,6-diamidino-2-pheynylindole) was used to visualize cell nuclei. Red asterisks denote untransfected cells. Scale bar, 10 μm. (D) Schematic diagram of the NanoBiT-fused viral chemokines vCCL1 (D1-23) and vCCL2(D1-20), which lack their signal sequences. (E) Immunoblot detection of the NanoBiT-fused viral chemokines in the culture media. Red asterisks indicate the vCCL bands, and black asterisks indicate non-specific bands. Supplementary Fig. S4 presents the original membranes of the immunoblots. (F) NanoBiT assays of vCCL-CCR8 interaction. The relative luminescence unit (RLU) was measured at 30 s after the mixture of the NanoBiT-fused vCCL and furimazine was added to the suspension of the NanoBiT-CCR8-transfected 293T cells. Each value represents the mean of triplicate samples from two independent experiments. Error bars represent standard deviations. The one-way ANOVA test was used to assess the statistical significance of differences between groups, and the t-test was used for post hoc comparisons. ***, p < 0.001.
It is well established that HHV-8 vCCL1 and vCCL2 function as agonists for CCR88,11,18. We first sought to identify optimal NanoBiT tags for observing the vCCL: CCR8 interaction. CCR8 was tagged at the N-terminus with either LgB or SmB, which was immediately followed by V5 epitope for the immunological detection of the CKR protein (Fig. 1B). The surface expression of LgB-V5-CCR8 and SmB-V5-CCR8 in transfected cells was verified using immunofluorescence assays on non-permeabilized cells (Fig. 1C). The mature vCCL1 (amino acids 24–95) and vCCL2 (amino acids 21–94) proteins lacking their signal sequences were fused at either the N- or C-terminus with the NanoBiT subunits (Fig. 1D), and their presence in the culture media was validated using immunoblotting with anti(a)-vCCL1 and a-vCCL2 antibodies (Fig. 1E). The bovine prolactin signal sequence (PSS) was added to the N-termini of the vCCLs and CCR8 proteins (Fig. 1B and D) to enable delivery to the endoplasmic reticulum and trafficking to the cell surface. An aliquot of the vCCL-containing culture media supplemented with furimazine was added to a suspension of 293T cells transfected with the variously NanoBiT-tagged CCR8 expression plasmids for 24 h. Consequently, we were able to find one binary combination that yielded significant NLuc activity: vCCL-SmB and LgB-V5-CCR8 (Fig. 1F). These results suggest that the NanoBiT system can be employed to assess the direct interaction between chemokines and their respective receptors in intact cells.
NanoBiT-based extensive analysis of chemokine receptors interacting with HHV-8 vCCLs
There are two families of CKRs: conventional CKRs (cCKRs) and atypical CKRs (ACKRs)31. To date, there are 19 known human cCKRs termed according to the predominant type of chemokine they bind (i.e., CC, CXC, CX3C, and XC), including 11 CCRs, 6 CXCRs, and a single CX3CR1 and XCR1. GPR35 was initially identified as a CXCL17 receptor32while subsequent studies failed to demonstrate that GPR35 is a bona fide receptor for CXCL1733,34. ACKRs are structurally related to cCKRs but do not couple to the classical (G protein-coupled) signaling pathways in response to chemokines35and five human ACKRs 1–5 have been identified to date.
While HHV-8 vCCLs are known to function as either an agonist or antagonist to certain CKRs, a comprehensive approach to characterize the binding properties of the vCCLs to human CKRs has not been undertaken. We aimed to apply the optimized NanoBiT system to examine the binding capabilities of the HHV-8 vCCLs to twenty-five human CKRs. The open reading frames (ORFs) of the human CKRs were PCR-amplified from specified cell lines (refer to the “Methods” section) and cloned into the LgB-V5 plasmid as performed with CCR8. We also included HHV-8 ORF74-encoded viral G-protein-coupled receptor (vGPCR), whose signaling is known to be inhibited by vCCL236. Our immunoblot analysis indicated that the expression of all the CKRs except CCR7 was detectable with anti-V5 antibody from whole cell extracts of transfected 293T cells (Fig. 2A). Moreover, immunofluorescence assays performed on non-permeabilized transfected cells verified the surface expression of all the CKRs (Fig. 2B). Note that immunofluorescence was detectable for LgB-V5-CCR7 but that the signal intensity was less than those of the other LgB-V5-CKRs, consistent with the immunoblot.
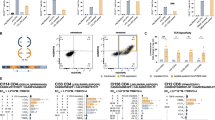
NanoBiT-based extensive analysis of human chemokine receptors interacting with HHV-8 chemokines. (A) V5-immunoblot analysis of the expression of twenty-five human chemokine receptors (CKRs) and the HHV-8-encoded viral chemokine receptor vGPCR. The CKRs were fused with PSS, LgB, and V5 as described in Fig. 1. 293T cells were transfected with the indicated CKR plasmids for 24 h, and whole-cell extracts were used for immunoblotting. EV indicates empty vector. The red asterisk indicates a non-specific immunoreactive band. Supplementary Fig. S4 presents the original membranes of the immunoblots. (B) V5-immunofluorescence detection of cell surface expression of LgB-fused V5-CKRs transfected into 293T cells. Scale bar, 10 μm. Supplementary Fig. S2 presents the original membranes of immunoblots. (C) NanoBiT analysis of vial chemokine-CKR interactions. The luminescence intensity was measured at 30 s after the mixture of SmB-fused vCCL (vCCL1, vCCL2, or vCCL3) and furimazine was added to the suspensions of cells expressing each LgB-fused V5-CKR. Each value in the heatmap indicates fold activation, determined by the ratio of RLU from treatment with each viral chemokine to the RLU from no treatment in CKR-transfected cells. Data represent the mean of triplicate samples from three independent experiments.
We subsequently conducted NanoBiT assays using the complete set of the engineered CKRs and HHV-8 vCCL1, vCCL2, and vCCL3 (amino acids 27–114); the latter also was tagged at the C-terminus with SmB as shown in Fig. 1D. The results of the experiment indicated, to our surprise, that vCCL1 exhibited a stronger binding affinity for CCR1, CCR3, ACKR2, and ACKR4 compared to CCR8 (Fig. 2C). As vCCL2 functions as an antagonist or agonist to a broad-spectrum of human CKRs16it is reasonable to expect that vCCL2 has the ability to interact directly with the corresponding CKRs. Contrary to our expectation, however, the results of the experiment revealed that vCCL2 bound exclusively to CCR8 (with the highest luminescence intensity) among the CKRs and to ACKR1 and ACKR4 (with medium intensity) (Fig. 2C). On the other hand, vCCL3 generally exhibited a lower luminescence intensity compared to vCCL1 and vCCL2 toward all the CKRs (Fig. 2C), including CCR4 and XCR1 that are known to respond to vCCL3 treatment10,21. It is noteworthy that vCCL3 could bind to ACKR4, also targeted by vCCL1 and vCCL2, with the highest signal among all of the CKRs tested (Fig. 2C). Together, our NanoBiT experiments confirmed the interactions of vCCL1 and vCCL2 with CCR8 and uncovered novel interactions of vCCLs 1–3 with ACKRs, including ACKR4. However, there are some inconsistencies with previous reports, and potential causes of the discrepancy are proposed in the “Discussion” section.
Validation of the NanoBiT method to evaluate vCCL-CKR interactions
To support the results of the NanoBiT experiments, we conducted additional experiments. Firstly, we utilized purified recombinant human CCL1 (I-309), the only chemokine known to interact with CCR8, for competition experiments. Indeed, CCL1 could reduce the luminescence from the vCCL1 interaction with CCR8, but not CCR1, XCR1 and ACKR4, in a dose-dependent manner (Fig. 3A). Next, we performed neutralization experiments using rabbit a-vCCL1 and a-vCCL2 antisera diluted 1:50. The results showed that a-vCCL1 significantly reduced the luminescence intensities induced by vCCL1-SmB addition to the LgB-CCR1, LgB-CCR8, and LgB-ACKR4 cultures compared to rabbit pre-immune (PI) serum (Fig. 3B). However, a modest increase in the luminescence induced by the addition of vCCL1-SmB to the LgB-XCR1 culture was not affected significantly by a-vCCL1 (Fig. 3B). Likewise, a-vCCL2 significantly decreased the luminescence intensities induced by vCCL2-SmB addition to LgB-CCR8 and LgB-ACKR4 compared to the PI serum (Fig. 3C). However, modest increases in the luminescence caused by vCCL2-SmB addition to the LgB-CCR1 and LgB-XCR1 cultures was not affected by the a-vCCL2 serum (Fig. 3C), implying the increases may result from non-specific binding. Together, these competition and neutralization experiments validate our NanoBiT assays for surveying true interactions between chemokine and CKRs in the intact cell.
Characterization of the CKR interactions of vCCLs 1–2 through competition and neutralization experiments. (A) CCL1 competition NanoBiT assays. The luminescence was measured at 30 s after the mixture of vCCL1-SmB and furimazine, together with purified recombinant human CCL1 at the indicated final concentrations, was added to the suspensions of cells expressing the indicated LgB-fused V5-CKRs. The percentage of RLU from CCL1 treatment to the RLU from no CCL1 treatment in CKR-transfected cells. Data represent the mean of triplicate samples from three independent experiments. Error bars represent standard deviations. (B-C) Viral chemokine neutralization NanoBiT assay. Rabbit antisera to vCCL1 and vCCL2, or pre-immune serum (PI), were added at a 1:50 dilution to the suspensions of the SmB-fused vCCL1 and vCCL2 proteins, respectively, and the mixtures were incubated at 4℃ for 3 h. The NanoBiT assays were performed as described above. The percentage of RLU from anti-vCCL serum treatment to the RLU from PI treatment in CKR-transfected cells. The one-way ANOVA test was used to assess the statistical significance of differences between groups, and the t-test was used for post hoc comparisons. *, p < 0.05; **, p < 0.01; ***, p < 0.001; ns, not significant.
Next, we examined vCCL3-receptor interactions further using mutagenesis of vCCL3. The mature vCCL3 secreted into the extracellular milieu contains six cysteine (C) residues. Among them, two cysteine residues at positions 37 and 38 align with the CC motif of CC-type chemokines, which is crucial for forming the internal disulfide bonds that ensure functional conformation for receptor binding. Thus, we predicted that a mutation of the CC motif would reduce the binding of vCCL3 to CKRs. Using site-directed mutagenesis, we generated the vCCL3 variant (CC/SS), in which cysteine residues at 37 and 38 were replaced with serine (S), and it was tagged at either the C-terminal or N-terminal with SmB (Fig. 4A). Immunoblotting using a-vCCL3 showed that the levels of the SmB-tagged native (WT) and CC/SS vCCL3 proteins released to the culture media from transfected 293T cells were comparable (Fig. 4B). Afterward, we conducted NanoBiT assays with the LgB-fused CKRs, including CCR1, CCR4, CCR8, XCR1, and ACKR4, and assessed the luminescence intensities at 30 s, the same incubation duration as in Fig. 2C, and again at 5 min following the addition of vCCL3. The findings showed that all the CKRs responded at 5 min, while some CKRs (CCR1, XCR1, and ACKR4) responded within 30 s, after the treatment with vCCL3-SmB, but not SmB-vCCL3, SmB-vCCL3(CC/SS), and vCCL3(CC/SS)-SmB(Fig. 4C). These results suggest that vCCL3 may function an agonist for several CKRs despite their low binding affinities. To investigate this possibility, we decided to assess the effect of vCCL3, along with the other vCCLs, on CKR downstream signaling.
Mutagenic assessment of vCCL3-CKR interactions. (A) Schematic diagram of the vCCL3 proteins (WT and CC/SS) fused with SmB at either their N- or C-termini. CC/SS denotes the vCCL3 variant (C37S/C38S) with a cysteine-to-serine substitution at positions 37 and 38 of vCCL3. WT indicates the native form of vCCL3. (B) Immunoblot detection of the SmB-fused vCCL3 proteins in the cell culture media using rabbit anti-vCCL3 antiserum. Supplementary Fig. S4 presents the original membrane of the immunoblot. (C) NanoBiT analysis of vCCL3-CKR interactions. The luminescence intensity was recorded at 30 s and for 5 min after the mixture of the SmB-fused vCCL3 protein and furimazine was added to the suspensions of cells expressing each LgB-fused V5-CKR, including CCR1, CCR4, CCR8, XCR1, and ACKR4. The ratio of RLU from treatment with each vCCL3 to the RLU from no treatment in CKR-transfected cells. Each value represents the mean of triplicate samples from three independent experiments. *, p < 0.05; **, p < 0.01; ***, p < 0.001.
NanoBiT assessments of vCCL-induced interactions between CKRs and b-arrestin 2
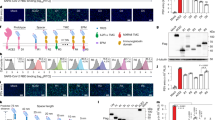
Upon chemokine treatment, CKRs interact with b-arrestins, an adaptor protein that regulates CKR desensitization, internalization, and signaling26. Besides vCCL-CKR interactions, we also sought to employ NanoBiT to investigate vCCL-induced interactions between CKRs and b-arrestins. b-arrestin 2 was tagged at its N-terminus with SmB, and the specified CKRs were tagged at their C-terminus with LgB (Fig. 5A). The NanoBiT experiments showed that vCCLs could trigger the binary interactions of b-arrestin 2 with the following CKRs: CCR8, ACKR1, and ACKR4 for vCCL1; ACKR1 for vCCL2; XCR1, ACKR1, and ACKR3 for vCCL3 (Fig. 5B). Interestingly, even though vCCL2 can bind and signal through CCR8, it did not affect the CCR8 interaction with b-arrestin 2 (Fig. 5B). To assess b-arrestin recruitment using an independent approach, we employed the PRESTO-Tango (parallel receptorome expression and screening via transcriptional output, with transcriptional activation following arrestin translocation) method37. The cDNA fragments of CCR4, XCR1, ACKR1, ACKR2, ACKR4, and ACKR5 genes were cloned into the Tango plasmid by replacing CCR8 in the CCR8-Tango plasmid using ClaI sites (Fig. 5C). ACKR3 was omitted due to a cloning problem. The results of the Tango assays (Fig. 5D) were comparable with those of the NanoBiT assays. Our approaches suggest that some CKRs, including XCR1, ACKR1, and ACKR4, may serve as functional receptors for vCCL3. Overall, our NanoBiT and subsequent experiments identified CKRs for vCCL-binding and signaling.
NanoBiT and Tango assays of vCCL-induced b-arrestin 2 recruitment to CKRs. (A) Schematic diagram of the NanoBiT-fused b-arrestin 2 (bAR2) and CKRs. bAR2 was tagged with SmB at its N-terminus. The CKRs CCR4, CCR8, XCR1, and ACKR1-5 were fused with LgB at their C-termini. (B) NanoBiT assays to detect the vCCL-induced recruitment of bAR2 to the CKRs. Fold activation was determined by the ratio of RLU from treatment with each viral chemokine to the RLU from no treatment (mock) in CKR-transfected cells. (C) Schematic illustration of the Tango construct. The diagram was adapted from the paper published by Kroeze WK et al.37. HAsignal, the influenza hemagglutinin signal peptide; V2 tail, the C terminus of the V2 vasopressin receptor; TEV, the tobacco etch virus endopeptidase cleavage site; tTA, the tetracycline transactivator (tTA) protein. (D) Tango assays were performed as described in the Method section. The Tango reporter cell line HTLA was transfected with the specified CKR plasmids. Fold activation was determined by the ratio of RLU from treatment with each viral chemokine to the RLU from no treatment (mock) in CKR-transfected cells. The one-way ANOVA test assessed the statistical significance of differences between groups, and the t-test was used for post hoc comparisons. *, p < 0.05; **, p < 0.01; ***, p < 0.001.
NanoBiT evaluations of vIL-6-receptor interaction and signaling
HHV-8 encodes viral interleukin 6 (vIL-6)13,27,38. While human IL-6 requires interaction with gp130 signal transducer and gp80 receptor subunits for signaling, vIL-6 does not require gp80 for active complex formation. Rather, vIL-6 can signal through tetrameric (gp1302:vIL-62) or hexameric (gp1302:gp802:vIL-62) complexes39,40,41,42. We examined whether NanoBiT can also assess the interaction between vIL-6 and gp130 in intact cells. The proteins were tagged at the N-terminus or C-terminus with the NanoBiT subunits (Fig. 6A). The PSS was added in place of the gp130 signal sequence (amino acids 1–22) to the N-terminus of the NanoBiT-gp130 constructs, and their expression was confirmed through gp130-immunoblotting of whole cell extracts (Fig. 6A). Also, the PSS was added in place of the predicted vIL-6 signal sequence (amino acids 1–22) to the N-terminus of the NanoBiT-vIL-6 constructs (Fig. 6A). We transfected 293T cells with the NanoBiT-vIL-6 plasmids to produce conditioned media, but immunoblotting with a-vIL-6 barely detected the vIL-6 proteins therein. This is consistent with the previous report that vIL-6 is poorly secreted and resides primarily within the endoplasmic reticulum43,44. Regardless of the technical issue of vIL-6 detection, we could find the NanoBiT binary combinations of vIL-6 and gp130 (the latter comprising fusions with the extracellular N-terminus of gp130) with strong luminescence intensities (Fig. 6B).
NanoBiT assessment of vIL6-gp130-STAT3 interactions. (A) Schematic diagram of the vIL6 and gp130 proteins fused with the NanoBiT subunits at either their N- or C-termini. Expression of NanoBiT-tagged gp130 in 293T cells was examined using immunoblotting with anti-gp130 antibody. Supplementary Fig. S4 presents the original membranes of the immunoblots. (B) NanoBiT assays of vIL6-gp130 interaction. The luminescence intensity was measured at 5 min after the mixture of NanoBiT-fused vIL6 and furimazine was added to the suspension of NanoBiT-gp130-transfected 293T cells. Each RLU value represents the mean of triplicate samples from a representative experiment. Error bars represent standard deviations. The one-way ANOVA test assessed the statistical significance of differences between groups, and the t-test was used for post hoc comparisons. **, p < 0.01; ***, p < 0.001. (C) Schematic diagram of the NanoBiT-fused FLAG-gp130 and mEGFP-STAT3 proteins. mEGFP refers to monomeric EGFP. (D) Co-immunoprecipitation assay of vIL-6-induced interaction of NanoBiT-fused gp130 and STAT3 proteins. 293T cells were single or double-transfected with plasmids encoding the indicated genes and, 24 h later, left untreated or treated with vIL-6-conditional medium for 5 min before cell lysis. FLAG-immunoprecipitates and cell extracts were immunoblotted with the indicated antibodies. The arrow indicates β-actin. Supplementary Fig. S4 presents the original membranes of the immunoblots. (E) NanoBiT assays of vIL6-induced interaction between gp130 and STAT3. The luminescence intensity was recorded every minute for 10 min after the addition of empty vector-control medium or vIL6-conditional medium, together with furimazine, to the suspension of 293T cells transfected with Flag-gp130-LgB and/or mEGFP-STAT3-SmB. Each RLU value represents the mean of triplicate samples from a representative experiment. Error bars represent standard deviations.
vIL-6 triggers the activation of STAT1 and STAT3 through gp130 complexes45. Mainly, co-immunoprecipitation (co-IP) methodology has been used to assess the intracellular interactions between gp130 and STAT proteins. We examined whether NanoBiT can be used to assess the gp130-STAT interactions. STAT3 was tagged at the N-terminus or C-terminus with SmB (Supplementary Fig. S1). For the NanoBiT experiments, 293T cells were co-transfected with plasmids expressing gp130-LgB and SmB-STAT3 or STAT3-SmB and, 24 h later, treated with the conditioned culture media containing vIL-6 in the presence of furimazine. The results showed that the relative luminescence intensity of gp130-LgB was higher with STAT3-SmB than with SmB-STAT3 (Supplementary Fig. S1). To confirm the NanoBiT-based binary interaction between gp130 and STAT3, we employed co-IP. To facilitate immunoprecipitation of gp130-LgB, a FLAG tag was added to the N-terminus of the protein (Fig. 6C). Also, to differentiate STAT3-SmB from endogenous STAT3, monomeric EGFP (mEGFP) was incorporated into the protein (Fig. 6C). As expected, the co-IP experiment confirmed the IL-6-induced interaction of the NanoBiT-fused gp130 and STAT3 (Fig. 6D). Consistent with this finding, immunoblotting of cell extracts using anti-phospho-STAT3 (Y705) antibody confirmed that vIL-6 significantly promoted phosphorylation of the STAT3-SmB protein in cells co-transfected with the gp130-LgB protein (Fig. 6D). The inclusion of FLAG and mEGFP tags did not interfere with the NanoBiT binary interaction (Fig. 6E). Together, our NanoBiT experiments identified the binary combinations of vIL-6:gp130 and gp130:STAT3 interactions that will be applied for future development of neutralizing vIL-6 antibodies and drugs.
Discussion
This study explores the suitability of the NanoBiT technology for re-evaluating the protein-protein interactions involved in viral chemokine and cytokine signaling. Our NanoBiT assays confirmed the established protein-protein interactions, including the chemo/cytokine-receptor interactions between vCCL1 and CCR8, vCCL2 and CCR8, vCCL3 and XCR1, and vIL-6 and gp130. In addition, our screening of the complete set of known human CKRs identified novel and notable chemokine-receptor interactions, including vCCLs 1–3 interactions with ACKR4. Moreover, we demonstrated receptor interactions with signaling molecules involved in the viral chemo/cytokine responses. Therefore, our NanoBiT approach provides a practical way to measure protein interactions in viral chemo/cytokine signaling with high sensitivity and simplicity, thus being applicable for the development of neutralizing antibodies and small molecule drugs in preventing these interactions and the host immune and cell trafficking dysfunction they cause.
Despite the successful NanoBiT detection of the viral chemo/cytokine-receptor interactions, we could not verify some known interactions between vCCL2 and CKRs. vCCL2 is known to act as a broad-spectrum antagonist of many CKRs10,16,46, including CCR1, CCR2, CCR5, XCR1, CX3CR1, CXCR4, and vGPCR36 and as an agonist of CCR319 and ACKR320. Furthermore, a crystal structure analysis proposed the direct interaction between vCCL2 with CXCR447. However, the expected interactions of vCCL2 with these CKRs were hardly detectable in the NanoBiT system. These weak interactions might be accounted for by a steric hindrance that may result from tagging with the NanoBiT subunits; however, the subunits appeared to have no impact on the vCCL2:CCR8 interaction. Further studies are warranted to tackle the differences between functional and physical interactions of vCCL2 with the CRKs. On the other hand, vCCL2 exhibited a strong affinity for ACKR1 and ACKR4; these and other ACKRs have not previously been investigated for vCCL2 binding and signaling. ACKRs are known to have the ability to form hetero-oligomerization with classical CKRs, which may influence signaling. For instance, ACKR1 inhibits CCR5 signaling through hetero-oligomerization with CCR548. ACKR4 also inhibits CXCR3-mediated chemotaxis by hetero-oligomerizing with CXCR349. Thus, the promiscuous functions of vCCL2 as an antagonist toward diverse CKRs might be explained by the possibility that the vCCL2-bound ACKRs are prone to form oligomerization with classical CKRs and thereby inhibit chemokine signaling.
Among the CKRs newly identified as interacting with vCCLs, ACKR4 is of particular interest due to its shared binding properties toward all the vCCLs. ACKR4, previously called CCX-CKR1 or CCRL1, functions primarily as a decoy receptor by binding and scavenging the chemokines CCL19, CCL20, CCL21, CCL22, and CCL25, thereby regulating their availability and modulating the CC-type CKR signaling50. The precise roles of ACKR4 in tumorigenesis and tumor immunity are poorly understood, although ACKR4 expression is inversely correlated with the growth and metastasis of certain tumors51. Thus, ACKR4 will likely capture and scavenge vCCLs and potentially counteract their roles in HHV-8 infection. ACKR4 is expressed in lymphatic vessels and endothelial cells lining the ceiling of the subcapsular sinus in lymph nodes52. This postulated viral chemokine scavenging function of ACKR4 in lymph nodes might modulate HHV-8 lytic and latent biology and associated disease within this compartment. Perhaps vCCL1 and vCCL3, which facilitate the interaction between ACKR4 and b-arrestin 2, might downregulate ACKR via its internalization. This could contribute to the development of HHV-8-related cancers, such as KS and PEL. In the future, exploring the functional significance of ACKR4 and its interplay with the vCCLs 1–3 in the pathogenesis of HHV-8 diseases will be important.
HHV-8 utilizes vCCLs to disrupt the recruitment of key immune cell subsets and their complex interplay and to re-polarize the dampened response away from an antiviral Th1 immunity53. vCCL1 and vCCL2 also promote the survival of infected cells by countering pro-apoptotic signaling during productive viral replication via autocrine signaling and the growth and survival of neighboring latently infected cells via paracrine signaling11. Thus, the induction of antibodies that neutralize the activities of vCCLs could compromise productive replication and HHV-8-associated pathogenesis. Our NanoBiT-based approach for characterizing the vCCL-receptor interactions and their downstream signaling protein-protein interactions will benefit the development of neutralizing antibodies to vCCLs. Moreover, this approach can help identify new viral ligand-receptor pairings among the atypical chemokine receptor family and facilitate the discovery and development of drugs targeting such interactions critical for HHV-8 persistence and host cell transformation.
Methods
Cell culture
Human embryo kidney 293T (ATCC) and HeLa (ATCC) cell lines were cultured in DMEM media supplemented with 10% fetal bovine serum and 1% antibiotics of penicillin, streptomycin, and 1.5 µg/ml plasmocin (Invivogen) in a humidified incubator at 5% CO2 at 37 ℃. HTLA cells stably expressing a tTA-dependent luciferase reporter and a b-arrestin-TEV protease fusion gene (a gift of Bryan Roth) were maintained in DMEM supplemented with 10% FBS, 2 µg/ml puromycin, and 100 µg/ml hygromycin. THP-1 (ATCC), BCBL-1 (ATCC), Jurkat (ATCC), and TL-oM1 (a gift of Edward Harhaj) cell lines were cultured in RPMI1640 media supplemented with 10% fetal bovine serum and 1% antibiotics of penicillin, streptomycin, and 1.5 µg/ml plasmocin in a humidified incubator at 5% CO2 at 37 ℃.
Transfection
Transient transfection with plasmids was performed using GenJet DNA transfection reagent (SignaGen Laboratories) or Lipofectamine 2000 (Invitrogen) according to the manufacturer’s instructions.
RNA and DNA manipulation
Total RNAs were extracted from cultured cells using an RNA isolation kit (Zymo Research) and reverse transcribed into cDNA using a Superscript IV synthesis kit (Thermo Fisher Scientific). The cDNAs of b-arrestin 2 and STAT3 were derived from HeLa cells; CCR1, CCR2, CCR7, CXCR2, CXCR6, ACKR2, and CX3CR1 from THP1 cells; CCR3 from BCBL1 cells; CCR5, CCR9, CXCR1, ACKR1, ACKR3, ACKR5, and GPR35 from Jurkat cells; CCR6, CCR11, CXCR3, CXCR4, CXCR5, and ACKR4 from TIME cells; CCR10 from TL-oM1 cells; XCR1 (HG19858-UT) and CCR4 (HG13064-UT) from Sino Biologicals; CCR8 and gp130 from John Nicholas11. The cDNA fragment of the bovine prolactin signal sequence (PSS) was synthesized by Integrated DNA Technologies. All polymerase chain reaction amplification and site-directed mutagenesis, including point and deletion mutations, were performed using Platinum SuperFi II DNA polymerase (Thermo Fisher Scientific). Subcloning of cDNAs and their derivatives into expression plasmids, including pICE (a gift from Steve Jackson, Addgene plasmid #46960) and NanoBiT system vectors (Promega), was performed using appropriate restriction enzyme sites. CCR8-Tango was a gift from Bryan Roth (Addgene plasmid # 66245). Other CKR genes were inserted into the Tango plasmid by replacing the CCR8 gene using the restriction enzyme ClaI. The cloning information and the list of oligonucleotides can be found in Supplementary Fig. S2 and Supplementary Table S1 online. After cloning, subcloned DNA fragments were verified by DNA sequencing to ensure accuracy.
Antibodies
V5 tag antibodies were purchased from Cell Signaling Technology (#13202) and Thermo Fisher Scientific (R960-25). Phospho-STAT3 (Tyr705) (#9131) and gp130 (#3732) antibodies was purchased from Cell Signaling Technology. GFP (FL) antibody was purchased from Santa Cruz Biotechnology. Anti-Flag (M2) antibody was purchased from Sigma-Aldrich. b-Actin (60008-1) antibody was purchased from Proteintech. The antibodies to vCCL1 and vCCL2 were kindly provided by John Nocholas. The vCCL3 antibody was custom-generated with the purified full-length protein of vCCL3 (Thermo Fisher Scientific). In brief, New Zealand white rabbits were injected with 500 µg of the purified, bacterially expressed HHV-8 multi-antigen fusion protein vCCL123-vIL-6 with a C-terminal 6xHIS tag. The ability of the polyclonal serum (#886) to recognize the vCCL3 protein expressed in 293T cells was confirmed through immunoblotting, shown in Supplementary Fig. S3.
Immunological assays
For the preparation of whole cell extracts, cells were resuspended in RIPA buffer (50 mM Tris [pH 7.4], 150 mM NaCl, 1% Igepal CA-630, 0.25% deoxycholate, and 0.1% sodium dodecyl sulfate(SDS)) containing a protease inhibitor cocktail and protein phosphatase inhibitors (10 mM sodium fluoride and 5 mM sodium orthovanadate). For immunoblotting, cell lysates were boiled in 1x SDS sample buffer and separated by SDS-PAGE, transferred to polyvinylidene difluoride membranes, and immunoblotted with appropriate primary antibodies diluted in SuperBlock (phosphate-buffered saline (PBS)) blocking buffer (Thermo Fisher Scientific). Following incubation with horse radish peroxidase-labeled appropriate secondary antibody, immunoreactive bands were visualized by enhanced chemiluminescent (ECL) reagents, Clarity (Bio-Rad) or SuperSignal West Femto (Thermo Fisher Scientific) on an ECL film. For immunoblot of the membrane proteins CKRs, whole cell extracts were heated at 75℃ for 5 min before loading on SDS-PAGE. For immunoprecipitation of FLAG-tagged proteins, cells were lysed in SDS-free RIPA buffer supplemented with protease inhibitor cocktail on ice. The lysates cleared by centrifugation at 12,000 × g for 10 min at 4 °C were incubated with anti-DYKDDDDK tag (L5) affinity gel (Biolegend) at 4 °C overnight. Immunoprecipitates were washed with RIPA buffer, followed by boiling elution of bound proteins in 1× SDS sample buffer. For immunofluorescence assay (IFA) of CRKs, cells grown on a glass chamber slide were transfected and fixed in Image-iT fixative solution (Thermo Fisher Scientific) for 10 min. Following incubation with SuperBlock-PBS blocking buffer for 1 h at room temperature, cells were incubated with V5 antibody, washed with PBS, and then incubated with appropriate Alexa 488-conjugated secondary antibody. Glass slides were covered with rectangular coverslips using DAPI-containing ProLong Gold Antifade Mountant (Thermo Fisher Scientific). Cells were imaged by Zen software on a Zeiss confocal laser scanning microscope 700 with 20x objective, and images were processed using Image J (National Institutes of Health).
Protein fragment complementation NanoBiT assay
293T cells transfected with LgB-tagged receptors for 24 h were harvested and resuspended at 5 × 105 cells/ml in Opti-MEM I medium. For SmB-tagged vCCLs and vIL-6 production, 293T cells were transfected with the expression plasmids in an appropriate volume of Opti-MEM I medium for 48 h, and the conditioned culture media were collected. For chemo/cytokine-receptor interaction assays, 50 µl of NanoBiT-tagged receptor cell suspension was treated with 25 µl of the conditioned culture media supplemented with 1% furimazine (N1110, Promega) for set times, and luminescence intensities were measured using GloMax luminometers (Promega). NanoBiT assays for CKR: b-arrestin 2 and gp130:STAT3 interactions were performed as previously described54,55. In brief, 24 h after co-transfection of 293T cells in a 6-well plate with the indicated genes in the NanoBiT binary plasmids, cells were harvested and resuspended in 1 ml of Opti-MEM I medium, and 100 µl of the cell suspension was transferred to a 96-well opaque plate in triplicate. Furimazine was prepared in PBS at a dilution of 1:100, and 25 µl of the diluted substrate was added to each well. Following timed incubations, luminescence was measured by a GloMax 96-well microplate luminometer. Fold activation was determined by the ratio of RLU from treatment with each viral chemokine to the RLU from no treatment in each CKR-transfected culture.
PRESTO-Tango b-arrestin-recruitment assay
HTLA cells were plated in a 24-well plate in triplicate for transfection of each CKR-Tango plasmid. At 24 h after transfection, culture media were replaced by Opti-MEM I medium, and cells were incubated for another 24 h. Cells were then stimulated by the addition of viral chemokine-conditional media, two-fold diluted in fresh Opti-MEM, and incubated overnight at 37 °C. As a mock control, only vector-transfected conditional medium was used. The following day, the media and viral chemokines were removed, and 100 µl of passive lysis buffer was added. The plate was kept for 20 min at room temperature before luminescence was measured.
Statistical analyses
The one-way ANOVA test assessed the statistical significance of differences between groups, and Student’s t-test was used for post hoc pairwise comparisons. The statistical significance was considered significant when the p-value was less than 0.05 (p < 0.05).
Data availability
Data is available on request from the corresponding author.
References
Alcami, A. & Lira, S. A. Modulation of chemokine activity by viruses. Curr. Opin. Immunol. 22, 482–487. https://doi.org/10.1016/j.coi.2010.06.004 (2010).
Alcami, A. Viral mimicry of cytokines, chemokines and their receptors. Nat. Rev. Immunol. 3, 36–50. https://doi.org/10.1038/nri980 (2003).
Pontejo, S. M. & Murphy, P. M. Chemokines encoded by herpesviruses. J. Leukoc. Biol. 102, 1199–1217. https://doi.org/10.1189/jlb.4RU0417-145RR (2017).
Hernaez, B. & Alcami, A. Poxvirus immune evasion. Annu. Rev. Immunol. 42, 551–584. https://doi.org/10.1146/annurev-immunol-090222-110227 (2024).
Chang, Y. et al. Identification of herpesvirus-like DNA sequences in AIDS-associated kaposi’s sarcoma. Science 266, 1865–1869 (1994).
Arvanitakis, L. et al. Establishment and characterization of a primary effusion (body cavity-based) lymphoma cell line (BC-3) harboring kaposi’s sarcoma-associated herpesvirus (KSHV/HHV-8) in the absence of Epstein-Barr virus. Blood 88, 2648–2654 (1996).
Uldrick, T. S., Polizzotto, M. N. & Yarchoan, R. Recent advances in Kaposi sarcoma herpesvirus-associated multicentric Castleman disease. Curr. Opin. Oncol. 24, 495–505. https://doi.org/10.1097/CCO.0b013e328355e0f3 (2012).
Dairaghi, D. J., Fan, R. A., McMaster, B. E., Hanley, M. R. & Schall, T. J. HHV8-encoded vMIP-I selectively engages chemokine receptor CCR8. Agonist and antagonist profiles of viral chemokines. J. Biol. Chem. 274, 21569–21574 (1999).
Endres, M. J., Garlisi, C. G., Xiao, H., Shan, L. & Hedrick, J. A. The kaposi’s sarcoma-related herpesvirus (KSHV)-encoded chemokine vMIP-I is a specific agonist for the CC chemokine receptor (CCR)8. J. Exp. Med. 189, 1993–1998. https://doi.org/10.1084/jem.189.12.1993 (1999).
Luttichau, H. R., Johnsen, A. H., Jurlander, J., Rosenkilde, M. M. & Schwartz, T. W. Kaposi sarcoma-associated herpes virus targets the lymphotactin receptor with both a broad spectrum antagonist vCCL2 and a highly selective and potent agonist vCCL3. J. Biol. Chem. 282, 17794–17805. https://doi.org/10.1074/jbc.M702001200 (2007).
Choi, Y. B. & Nicholas, J. Autocrine and paracrine promotion of cell survival and virus replication by human herpesvirus 8 chemokines. J. Virol. 82, 6501–6513. https://doi.org/10.1128/JVI.02396-07 (2008).
Wan, Y. et al. Identification of full, partial and inverse CC chemokine receptor 3 agonists using [35S]GTPgammaS binding. Eur. J. Pharmacol. 456, 1–10. https://doi.org/10.1016/s0014-2999(02)02621-3 (2002).
Moore, P. S., Boshoff, C., Weiss, R. A. & Chang, Y. Molecular mimicry of human cytokine and cytokine response pathway genes by KSHV. Science 274, 1739–1744 (1996).
Nakano, K. et al. Kaposi’s sarcoma-associated herpesvirus (KSHV)-encoded vMIP-I and vMIP-II induce signal transduction and chemotaxis in monocytic cells. Arch. Virol. 148, 871–890. https://doi.org/10.1007/s00705-002-0971-7 (2003).
Szpakowska, M. & Chevigne, A. vCCL2/vMIP-II, the viral master keymokine. J. Leukoc. Biol. 99, 893–900. https://doi.org/10.1189/jlb.2MR0815-383R (2016).
Kledal, T. N. et al. A broad-spectrum chemokine antagonist encoded by kaposi’s sarcoma-associated herpesvirus. Science 277, 1656–1659 (1997).
Luttichau, H. R., Lewis, I. C., Gerstoft, J. & Schwartz, T. W. The herpesvirus 8-encoded chemokine vMIP-II, but not the poxvirus-encoded chemokine MC148, inhibits the CCR10 receptor. Eur. J. Immunol. 31, 1217–1220 (2001).
Sozzani, S. et al. The viral chemokine macrophage inflammatory protein-II is a selective Th2 chemoattractant. Blood 92, 4036–4039 (1998).
Weber, K. S.et al. Selective recruitment of Th2-type cells and evasion from a cytotoxic immune response mediated by viral macrophage inhibitory protein-II. Eur J Immunol 31, 2458–2466 (2001).
Szpakowska, M. et al. Human herpesvirus 8-encoded chemokine vCCL2/vMIP-II is an agonist of the atypical chemokine receptor ACKR3/CXCR7. Biochem. Pharmacol. 114, 14–21. https://doi.org/10.1016/j.bcp.2016.05.012 (2016).
Stine, J. T. et al. KSHV-encoded CC chemokine vMIP-III is a CCR4 agonist, stimulates angiogenesis, and selectively chemoattracts TH2 cells. Blood 95, 1151–1157 (2000).
Sallusto, F., Mackay, C. R. & Lanzavecchia, A. Selective expression of the eotaxin receptor CCR3 by human T helper 2 cells. Science 277, 2005–2007. https://doi.org/10.1126/science.277.5334.2005 (1997).
Imai, T. et al. Selective recruitment of CCR4-bearing Th2 cells toward antigen-presenting cells by the CC chemokines thymus and activation-regulated chemokine and macrophage-derived chemokine. Int. Immunol. 11, 81–88. https://doi.org/10.1093/intimm/11.1.81 (1999).
Zingoni, A. et al. The chemokine receptor CCR8 is preferentially expressed in Th2 but not Th1 cells. J. Immunol. 161, 547–551 (1998).
Dixon, A. S. et al. NanoLuc complementation reporter optimized for accurate measurement of protein interactions in cells. ACS Chem. Biol. 11, 400–408. https://doi.org/10.1021/acschembio.5b00753 (2016).
Kee, T. R. et al. The multifaceted functions of beta-arrestins and their therapeutic potential in neurodegenerative diseases. Exp. Mol. Med. 56, 129–141. https://doi.org/10.1038/s12276-023-01144-4 (2024).
Nicholas, J. et al. Kaposi’s sarcoma-associated human herpesvirus-8 encodes homologues of macrophage inflammatory protein-1 and interleukin-6. Nat. Med. 3, 287–292 (1997).
Cannon, J. S. et al. Heterogeneity of viral IL-6 expression in HHV-8-associated diseases. J. Infect. Dis. 180, 824–828. https://doi.org/10.1086/314956 (1999).
Nicholas, J. Human herpesvirus 8-encoded proteins with potential roles in virus-associated neoplasia. Front. Biosci. 12, 265–281. https://doi.org/10.2741/2063 (2007).
Choi, Y. B., Cousins, E. & Nicholas, J. Novel functions and Virus-Host interactions implicated in pathogenesis and replication of human herpesvirus 8. Recent. Results Cancer Res. 217, 245–301. https://doi.org/10.1007/978-3-030-57362-1_11 (2021).
Hughes, C. E. & Nibbs, R. J. B. A guide to chemokines and their receptors. FEBS J. 285, 2944–2971. https://doi.org/10.1111/febs.14466 (2018).
Maravillas-Montero, J. L. et al. Cutting edge: GPR35/CXCR8 is the receptor of the mucosal chemokine CXCL17. J. Immunol. 194, 29–33. https://doi.org/10.4049/jimmunol.1401704 (2015).
Binti Mohd Amir, N. A. S. et al. Evidence for the existence of a CXCL17 receptor distinct from GPR35. J. Immunol. 201, 714–724. https://doi.org/10.4049/jimmunol.1700884 (2018).
Park, S. J., Lee, S. J., Nam, S. Y. & Im, D. S. GPR35 mediates lodoxamide-induced migration inhibitory response but not CXCL17-induced migration stimulatory response in THP-1 cells; is GPR35 a receptor for CXCL17? Br. J. Pharmacol. 175, 154–161. https://doi.org/10.1111/bph.14082 (2018).
Comerford, I. & McColl, S. R. Atypical chemokine receptors in the immune system. Nat. Rev. Immunol. 24, 753–769. https://doi.org/10.1038/s41577-024-01025-5 (2024).
Geras-Raaka, E., Varma, A., Clark-Lewis, I. & Gershengorn, M. C. Kaposi’s sarcoma-associated herpesvirus (KSHV) chemokine vMIP-II and human SDF-1alpha inhibit signaling by KSHV G protein-coupled receptor. Biochem. Biophys. Res. Commun. 253, 725–727. https://doi.org/10.1006/bbrc.1998.9557 (1998).
Kroeze, W. K. et al. PRESTO-Tango as an open-source resource for interrogation of the druggable human GPCRome. Nat. Struct. Mol. Biol. 22, 362–369. https://doi.org/10.1038/nsmb.3014 (2015).
Neipel, F. et al. Human herpesvirus 8 encodes a homolog of interleukin-6. J. Virol. 71, 839–842 (1997).
Aoki, Y., Narazaki, M., Kishimoto, T. & Tosato, G. Receptor engagement by viral interleukin-6 encoded by Kaposi sarcoma-associated herpesvirus. Blood 98, 3042–3049 (2001).
Chen, D. & Nicholas, J. Structural requirements for gp80 independence of human herpesvirus 8 interleukin-6 (vIL-6) and evidence for gp80 stabilization of gp130 signaling complexes induced by vIL-6. J. Virol. 80, 9811–9821. https://doi.org/10.1128/JVI.00872-06 (2006).
Chow, D., He, X., Snow, A. L., Rose-John, S. & Garcia, K. C. Structure of an extracellular gp130 cytokine receptor signaling complex. Science 291, 2150–2155. https://doi.org/10.1126/science.1058308 (2001).
Boulanger, M. J. et al. Molecular mechanisms for viral mimicry of a human cytokine: activation of gp130 by HHV-8 interleukin-6. J. Mol. Biol. 335, 641–654 (2004).
Chen, D., Sandford, G. & Nicholas, J. Intracellular signaling mechanisms and activities of human herpesvirus 8 interleukin-6. J. Virol. 83, 722–733. https://doi.org/10.1128/JVI.01517-08 (2009).
Meads, M. B. & Medveczky, P. G. Kaposi’s sarcoma-associated herpesvirus-encoded viral interleukin-6 is secreted and modified differently than human interleukin-6: evidence for a unique autocrine signaling mechanism. J. Biol. Chem. 279, 51793–51803. https://doi.org/10.1074/jbc.M407382200 (2004).
Hu, F. & Nicholas, J. Signal transduction by human herpesvirus 8 viral interleukin-6 (vIL-6) is modulated by the nonsignaling gp80 subunit of the IL-6 receptor complex and is distinct from signaling induced by human IL-6. J. Virol. 80, 10874–10878. https://doi.org/10.1128/JVI.00767-06 (2006).
Zhou, N., Luo, Z., Luo, J., Hall, J. W. & Huang, Z. A novel peptide antagonist of CXCR4 derived from the N-terminus of viral chemokine vMIP-II. Biochemistry 39, 3782–3787. https://doi.org/10.1021/bi992750v (2000).
Qin, L. et al. Structural biology. Crystal structure of the chemokine receptor CXCR4 in complex with a viral chemokine. Science 347, 1117–1122. https://doi.org/10.1126/science.1261064 (2015).
Chakera, A., Seeber, R. M., John, A. E., Eidne, K. A. & Greaves, D. R. The Duffy antigen/receptor for chemokines exists in an oligomeric form in living cells and functionally antagonizes CCR5 signaling through hetero-oligomerization. Mol. Pharmacol. 73, 1362–1370. https://doi.org/10.1124/mol.107.040915 (2008).
Vinet, J. et al. Inhibition of CXCR3-mediated chemotaxis by the human chemokine receptor-like protein CCX-CKR. Br. J. Pharmacol. 168, 1375–1387. https://doi.org/10.1111/bph.12042 (2013).
Szpakowska, M. et al. Inclusion of ACKR5 in the systematic nomenclature of atypical chemokine receptors. Nat. Rev. Immunol. 25, 225–226. https://doi.org/10.1038/s41577-025-01135-8 (2025).
Samus, M. & Rot, A. Atypical chemokine receptors in cancer. Cytokine 176, 156504. https://doi.org/10.1016/j.cyto.2024.156504 (2024).
Friess, M. C. et al. Mechanosensitive ACKR4 scavenges CCR7 chemokines to facilitate T cell de-adhesion and passive transport by flow in inflamed afferent lymphatics. Cell. Rep. 38, 110334. https://doi.org/10.1016/j.celrep.2022.110334 (2022).
Nicholas, J. Human herpesvirus 8-encoded cytokines. Future Virol. 5, 197–206. https://doi.org/10.2217/fvl.10.2 (2010).
Choi, C. Y., Vo, M. T., Nicholas, J. & Choi, Y. B. Autophagy-competent mitochondrial translation elongation factor TUFM inhibits caspase-8-mediated apoptosis. Cell. Death Differ. https://doi.org/10.1038/s41418-021-00868-y (2021).
Vo, M. T., Smith, B. J., Nicholas, J. & Choi, Y. B. Activation of NIX-mediated mitophagy by an interferon regulatory factor homologue of human herpesvirus. Nat. Commun. 10, 3203. https://doi.org/10.1038/s41467-019-11164-2 (2019).
Acknowledgements
We appreciate John Nicholas for proofreading the paper.
Funding
This work was supported by the National Cancer Institute (NCI) grant R01CA214131 to Y.B.C. and by the National Institute of Allergy and Infectious Diseases (NIAID) grant R21Al177806 to Y.B.C. The funders had no role in study design, data collection and analysis, publication decisions, or manuscript preparation.
Author information
Authors and Affiliations
Contributions
V.P.: Investigation; formal analysis; writing – review and editing. R. R.: Investigation; methodology; writing – review and editing. Y.B. C.: Funding acquisition; investigation; methodology; formal analysis; project administration; supervision; validation; writing – original draft, review, and editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Palande, V., Roden, R.B.S. & Choi, Y.B. Nano-Luciferase complementation assay of human herpesvirus 8 chemo/cytokine-receptor interactions. Sci Rep 15, 35365 (2025). https://doi.org/10.1038/s41598-025-19281-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-19281-3