Abstract
The growing interest in using peptide molecules as therapeutic agents, driven by their high selectivity and efficacy, has become a significant trend in the pharmaceutical industry. However, their oral administration remains challenging due to their low bioavailability and vulnerability to proteases, which produce the cleavage of peptide bonds. To optimize peptide drug development, in silico tools based on machine learning algorithms have been developed for site of cleavage prediction. These tools, which rely on manual feature extraction, have limitations in capturing complex peptide structures, especially those involving non-natural amino acids or cyclic peptides. This study presents two novel in silico approaches for cleavage site prediction. The first approach uses protein language models, specifically ESM-2, which has been fine- tuned to leverage its learned peptide structure embeddings for accurate cleavage site prediction, eliminating the need for manual feature engineering. The second approach employs graph neural networks, representing peptides via hierarchical graphs at the atom and amino acid levels, effectively handling cyclic peptide structures, including those containing non-natural amino acids. The applicability of this second approach is shown through a case study on a set of four cyclic peptides containing non-natural amino acids, comparing in silico predictions with experimental data.
Similar content being viewed by others
Introduction
The growing interest in peptide molecules as therapeutic agents, owning to their high selectivity and efficacy, has emerged as a significant trend in the pharmaceutical industry1. Despite their promising therapeutic potential, the development of peptide-based drugs is limited by their pharmacokinetic properties, particularly their vulnerability to proteolytic degradation2. Proteases, which are enzymes widely distributed throughout the body, catalyze the hydrolysis of amide bonds in peptides, recognizing specific amino acids sequences3. Their high specificity is determined by the structure and characteristics of their active sites.
Several strategies have been developed to enhance stability against proteolytic degradation. These include structural chemical modifications such as peptide cyclization and the incorporation of non-natural amino acids4,5. These approaches aim to develop peptide analogues whose cleavage sites are either modified or masked, preventing recognition by proteases6. For this reason, characterizing proteolytic cleavage sites within a peptide sequence could help in the design of new peptide drugs with improved stability. However, current experimental methods for identifying cleavage sites are resource-intensive, and time-consuming, posing a significant barrier to peptide drug development7.
Numerous cost-effective computational methods for cleavage site prediction have been developed to address these challenges, with machine learning-based techniques emerging as the most promising in recent years8. However, these methods have certain limitations, such as relying on manual feature extraction, which fails to effectively represent the complexity of protein and peptide structures9. Additionally, they are typically restricted to linear sequences composed exclusively of the 20 standard amino acids, without accounting for chemical modification strategies – such as those previously discussed – that are commonly employed in peptide-drug design. A methodology introduced in a previous study by our research group tries to overcome some of these constraints by enabling applicability to non-standard amino acids10.
This work presents two novel in silico approaches for predicting cleavage sites on peptides, using data curated from the MEROPS database spanning 29 proteases implicated in peptide-drug degradation11. The first approach employs protein language models, which have shown significant potential for extracting meaningful representations from proteins and peptides12. Specifically, the study applies ESM-2, a state-of-the-art approach, which is based on the transformer encoder architecture13. The pre-trained model has been fine-tuned to leverage its generated peptide structure embeddings for accurate cleavage site prediction, eliminating the need for manual feature extraction. Despite its potential for superior performance, a key limitation lies in its inability to process non-natural amino acids and cyclic peptides14.
To address these limitations – which are also present in existing in silico tools for predicting peptide cleavage sites – a second approach has been developed using graph neural networks (GNNs). More specifically, this method employs a hierarchical graph for the representation of peptides at the atom and amino acid levels, enabling the representation of cyclic peptide structures, including those containing non-natural amino acids.
To demonstrate the applicability of this method to cyclic peptides containing non-canonical amino acids, a case study is presented. Cleavage site predictions, based on the availability of the experimental data, were made for four peptides using models developed for three proteases. These in silico results were then compared with experimentally obtained metabolite identification data from incubation studies analyzed via liquid chromatography-high-resolution mass spectrometry.
Methods
Data collection and pre-processing
Experimentally verified cleavage sites in protein substrates, produced by proteases involved in peptide-drug degradation, were collected from the MEROPS database (Release 12.5), a publicly available resource11. Missing FASTA sequences in some substrates were retrieved from UniProt15 using their unique identifiers, with only a few substrates excluded due to discrepancies with the cleavage site reported in MEROPS. To eliminate sequence redundancy, the CD-HIT program was applied with a 70% identity threshold between sequences within the same protease dataset16. The residue range specified in the MEROPS database for each cleavage site entry was used to define the corresponding substrate sequences in the datasets.
A detailed step-by-step description of the data extraction and processing pipeline, along with an illustrative example, is provided in the Supplementary Information. The full preprocessing script is also available via the Data Availability Statement.
To ensure that the models could learn reliable patterns, avoid overfitting, and generalize effectively to unseen data, datasets with fewer than 100 substrates were excluded. This filtering resulted in 29 independent datasets, each corresponding to a specific protease and comprising between 132 and 14,886 cleavage sites. These datasets, which are provided in the Supplementary Information, were used for training and evaluation of the developed methods. A statistical summary of the curated datasets is presented in Supplementary Table S1.
ESM-2 token classification
A major challenge in computational biology is encoding biological sequences into numerical vectors suitable for machine learning algorithms. Current in silico tools rely on manual feature extraction, which often fails to effectively represent the complexity and diversity of protein and peptide sequences, limiting the performance of the derived models9.
Our first approach employs pre-trained protein language models to encode peptide sequences in the form of embeddings, eliminating the need for manual feature extraction. These pre-trained protein language models have emerged as powerful tools for encoding biological data, as they effectively capture both the functional and structural properties of proteins and peptides. They achieve this through being trained in a self-supervised manner in large corpora of protein sequences9.
In this study, we use the pre-trained ESM-2 model17 (version: esm2_t6_8M_UR50D) to predict proteolytic cleavage sites in peptide sequences, as shown in Fig. 1. This model is based on a transformer encoder architecture and consists of 6 layers, each with 20 attention heads, with approximately 8 million parameters. The model takes peptide sequences in FASTA format as input, which are tokenized using the standard 20-letter amino acid alphabet. Due to the inherent vocabulary and tokenizer, the model is limited to natural amino acids and cannot process sequences containing non-natural residues or cyclic structures.
The model represents each amino acid in the sequence as a 320-dimensional vector, capturing not only its identity but also its context within the entire sequence via a self-attention mechanism. These vectors, known as embeddings, are then used to predict whether the amino acid at a given position lies on the N-terminal side of a proteolytic cleavage site.
Notably, the model does not classify amino acid residues or bonds directly. Instead, it predicts whether the peptide bond immediately following each amino acid (i.e., on the C-terminal side) is a site of cleavage. To do this, a classifier layer, constisting of a dropout followed by a single linear layer projecting the 320-dimensional embeddings to two output values, uses the amino acid embeddings to estimate a probability score for each bond. If the score exceeds 0.5, the bond is predicted to be cleaved; otherwise, it is predicted to remain intact (see Fig. 1).
The model is fine-tuned using standard supervised learning. During training, each amino acid is labeled based on whether the bond following it is cleaved (positive) or not (negative), allowing the model to learn cleavage patterns from sequence context.
The learning process is guided by a binary cross-entropy loss function, which helps the model learn the patterns associated with cleavage sites. All models were trained for 20 epochs using a learning rate of 0.0003 with linear decay, a batch size of 4, and a weight decay of 0.00001. Hyperparameters were selected during early exploratory experiments on a small number of proteases, using an 80/20 split within the training data to set aside a validation portion. Based on these preliminary results, we selected fixed hyperparameter values chosen for simplicity and consistency.
In all the final experiments, these hyperparameters were applied uniformly across the datasets. No grid search or early stopping was performed during training. This decision was made to ensure consistent evaluation conditions and to avoid overfitting any specific dataset.
While more extensive tuning could potentially improve absolute performance, the primary focus of this study is on model comparison and applicability across diverse input types and protease scenarios.
The self-attention mechanism, the core of the transformer architecture, is implemented through scaled dot-product attention, Eq. (1), where the query (Q), key (K), and value (V) matrices encode representations of the input sequence. Attention scores are computed via the dot product of Q and KT and subsequently scaled by the inverse of the square root of the dimensionality of Q and K to stabilize the gradients. The softmax function is then applied to these scores, producing attention weights that determine the relative importance of each amino acid to others in the sequence. Finally, these weights are used to compute a weighted sum of V, generating refined contextual representations for each amino acid18.
Graph neural networks
The second approach is based on the idea that molecules can naturally be represented as graphs, with nodes corresponding to atoms and edges representing the bonds between them19.
GNNs use a message-passing mechanism to aggregate information from neighboring nodes, enabling the model to capture complex relationships within graphs. Each node and edge in the graph are assigned feature vectors that represent various attributes, including both structural and feature-based information.
During each iteration, the embedding \(\:{h}_{u}^{\left(k\right)}\)of node u is updated by aggregating messages (\(\:{m}_{N\left(u\right)}^{\left(k\right)}\) from its neighbors N(u). The update process follows the general form of Eq. (2):
The AGGREGATE function combines the embeddings of neighboring nodes\(\:\:{h}_{u}^{\left(k\right)}\) for all \(\:v\in\:N\left(v\right)\) producing an aggregated message \(\:{m}_{N\left(u\right)}^{\left(k\right)}\). The UPDATE function then combines this message with the current embedding \(\:{h}_{u}^{\left(k\right)}\) of the node u to generate an updated embedding \(\:{h}_{u}^{\left(k+1\right)}.\) This process is repeated for \(\:\:K>0\) iterations, allowing information to propagate through the graph and the representation long-distance relationships20.
After \(\:K\:\)iterations, the final node embeddings \(\:{h}_{u}^{\left(K\right)}\) are used to generate a graph-level representation via pooling of the node embeddings, using what is known as a readout function. This final representation is then employed for downstream tasks21,22.
Hierarchical graph
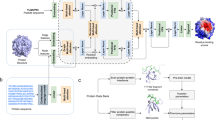
To represent peptides, we construct a hierarchical graph that captures features at both the atom and amino acid levels. As illustrated in Fig. 2, the lower level of the graph models each monomer by representing atoms as nodes and chemical bonds as edges. At a higher level, each amino acid in the peptide sequence is abstracted as a hypernode, with connections between them reflecting the peptide bonds in sequential order from the N-terminus to the C-terminus. This hierarchical design allows for the inclusion of all amino acids, including non-natural -including D-amino acids and chemically modified residues such as N-methylated, hydroxylated, or phosphorylated forms, among others- within a unified graph structure.
Pattern recognition
Each hypernode (amino acid) in a given peptide sequence is constructed by applying a fragmentation algorithm to the input sequence, provided in formats such as SDF or SMILES. The algorithm utilizes SMARTS patterns to model two predefined reactions: amide hydrolysis and disulfide bond reduction. Amide hydrolysis is incorporated to simulate peptide bond cleavage, enabling the extraction of individual amino acids, including both natural and non-natural residues. Disulfide bond reduction is employed to simulate the breaking of disulfide bridges between cysteine residues (or other thiol-containing amino acids), a bond commonly found inside chain cyclized peptides. This process fragments the peptide into monomers, which are then neutralized.
This strategy can be extended to include additional reactions by introducing new SMARTS patterns that define new bond-breaking rules between amino acids, allowing adaptation as new cyclization strategies emerge.
Monomer representation
To build an initial representation for each monomer obtained in the previous step, atomic and bond-level features are extracted using the RDKit Python library, with a detailed list available in Table 123.
Model architecture
All models are trained for 30 epochs using a learning rate of 0.0003 and a batch size of 4. Hyperparameters were selected following the same approach used for the ESM-2 Token Classification model. The architecture, illustrated in Fig. 2, begins with batch normalization applied to the input atom features. This is followed by a single transformer convolutional layer that operates at the atomic level, leveraging self-attention to update each atom’s feature vector based on its neighboring atoms and the associated bond attributes. The dimensionality of the hidden channels in all the graph convolution layers was set to 600, and no dropout layers were used.
Next, four gated graph convolutional layers are applied to progressively refine the atomic representations. These layers expand the effective receptive field to a radius of 5, allowing the model to capture localized, node-specific information across multiple hops in the graph.
To transition from atomic to amino acid-level representations, the refined atom features within each amino acid are aggregated via mean pooling. Specifically, each atom is associated with a feature vector of dimensionality d (e.g., 128), and for each amino acid (treated as a hypernode), we compute the element-wise average of its constituent atoms’ feature vectors. This results in a single d-dimensional vector per amino acid, where each element reflects the mean of a specific feature (e.g., spatial, chemical, or learned) across all atoms in the residue.
To model interactions between neighboring amino acids, we apply four additional gated graph convolutional layers at the amino acid level. The number of layers determines the effective window size, controlling how many iterations allow information to propagate between neighboring amino acids in the graph within the peptide sequence.
The model performs binary classification at the level of amino acid bonds (edges), aiming to predict whether a given bond is a site of cleavage. During training, each bond is labeled as either positive (1) or negative (0) based on this criterion.
For classification, the model constructs a bond representation by concatenating the feature vectors of the two amino acids it connects, preserving the directional order from the N-terminus. This concatenated vector is passed through a final classification module (a multi-layer perceptron) that outputs a probability indicating the likelihood that the bond is positive.
Training is conducted using standard supervised learning with a binary cross-entropy loss, which compares the predicted probabilities to the ground truth labels. This loss function guides the optimization process, enabling the model to learn feature representations and interaction patterns that are predictive of the target bond property.
Graph neural network model architecture. The atoms in the amino acid structures are represented as spheres, with colors denoting atom types: grey for carbon, red for oxygen, blue for nitrogen, and yellow for sulfur. The black arrows indicate the direction of the message passing between neighboring atoms and bonds during graph convolution. In the Gated Graph Convolution Layer (II), dashed grey lines group atoms belonging to the same amino acid residue. The cyan and green spheres highlight two different amino acids to illustrate how amino acid-level representation is constructed.
Performance evaluation
To evaluate the predictive performance of the models, the following metrics were computed: accuracy, precision, specificity, recall, and F1 score. These metrics are based on the values of true positives (TP), false positives (FP), true negatives (TN), and false negatives (FN), where TP represents correctly predicted cleavage sites, FP denotes non-cleavage sites incorrectly classified as cleavage sites, TN corresponds to correctly identified non-cleavage sites, and FN refers to cleavage sites that were misclassified as non-cleavage sites.
The metrics are calculated as follows:
-
Accuracy is the proportion of correct predictions over all predictions (positive and negative) as presented by Eq. (3).
$$Accuracy =\frac{\text{T}\text{P}+\text{T}\text{N}}{\text{T}\text{P}+\text{T}\text{N}+\text{F}\text{P}+\text{F}\text{N}}$$(3)
-
Precision is the proportion of predicted cleavage sites (positive predictions) that are correct as presented by Eq. (4).
$$Precision =\frac{\text{T}\text{P}}{\text{T}\text{P}+\:\text{F}\text{P}}$$(4)
-
Specificity is the proportion of non-cleavage sites (negative samples) correctly identified by the model as presented by Eq. (5).
$$Specificity=\frac{\text{T}\text{N}}{\text{T}\text{N}+\text{F}\text{P}}$$(5)
-
Recall is the proportion of cleavage sites (positive samples) correctly identified by the model as presented by Eq. (6).
$$Recall=\frac{\text{T}\text{P}}{\text{T}\text{P}+\:\text{F}\text{N}}$$(6)
-
F1-Score is defined as the harmonic mean of the precision and the recall, and reflects the balance between the two, which proves particularly advantageous in the imbalanced data sets as presented by Eq. (7).
The number of positive samples (cleavage sites) was substantially lower than the number of negative samples (non-cleavage sites), with negative-to-positive sample ratios across datasets ranging from 36 to 592, and a median of 188 across the 29 datasets. This pronounced class imbalance presents a specific challenge when splitting data into training and testing sets: without careful handling, random splitting can lead to uneven class distributions, where one or both splits contain too few positive samples. This undermines both the learning process and the reliability of performance estimates.
To address this, a stratified splitting strategy is used to ensure that both training and testing sets maintain a similar proportion of cleavage and non-cleavage sites. Instead of randomly selecting individual samples, the split is performed at the level of substrates, while stratification ensures that approximately 30% of the cleavage sites are included in the test set. This approach preserves the original class imbalance in a controlled way, allowing models to be trained and evaluated under realistic conditions that reflect the skewed distribution typical of real-world data.
Results and discussion
Predictive performance
In this section, the predictive performance of both developed models is described and analyzed.
ESM-2 token classification
The ESM-2 Token classification model results are shown in Table 2. The results are presented for a selected subset of proteases from our available datasets, including caspase-1 (C14.004), caspase-3 (C14.003), caspase-7 (C14.004), caspase-6 (C14.005), MMP-2 (M10.003), MMP-3 (M10.005), granzyme B (S01.010), and thrombin (S01.217), in line with the state-of-the-art tools, which will be used for comparison in the section “Comparison with other in silico cleavage site prediction models”. The results for the remaining proteases can be found in Supplementary Table S2.
The specificity values for all the models are consistently high, as reported in Supplementary Table S2, with scores upwards of 0.99. This is expected from the inherent class imbalance in the datasets, with a predominance of negative samples. As a result, even a model that always outputs a negative prediction would yield similar results. Similarly, accuracy is not a particularly informative metric either, as the high proportion of the negative class can lead to inflated accuracy values, as observed in nearly all the models with scores of 0.99 (see Supplementary Table S2). In such cases, F1 scores, which represent the harmonic mean of precision and recall, provide a more balanced assessment of each model’s overall performance.
The F1 scores obtained show variability in predictive performance across the different proteases. Among those analyzed, C14.005 demonstrated the highest score of 0.8665, followed by S01.217 with an F1 of 0.7671. Additional higher F1 scores, such as 0.9487 for signal peptidase I (S26.010) and 0.8524 for caspase-2 (C14.006), are found in some datasets within Supplementary Table S2.
Graph neural networks
The results of the GNN model for the 8 selected proteases are presented in Table 3, with C14.005 achieving the highest F1 score of 0.7370. Performance metrics for the remaining proteases can be found in Supplementary Table S3, where C14.006 achieved the second highest F1 score of 0.6834.
To establish a reference point for evaluating F1 scores, the ESM-2 token classification and GNN models were compared against two baselines: a random model (which predicts that a bond is a site of cleavage with 50% probability) and one that predicts every bond as a cleavage site (referred to as Constant 1). As shown in Supplementary Table S4, both models considerably outperformed both baselines across all proteases. In the section “Comparison with other in silico cleavage site prediction models”, the performance of the developed models for both approaches are compared to the current state-of-the-art method for site of cleavage prediction.
Improving the predictive performance by restricting the sequence length
Upon preliminary analysis, as shown in Fig. 3, reducing the sequence length of the substrates appears to improve model performance, as measured by F1 scores. To further investigate this, we computed the F1 scores on the test sets of each protease only including substrates of a length below specified thresholds, for all possible thresholds.
To maintain robustness, only datasets with at least 50 substrates in the test set were included; thus, datasets C14.001 and S01.217 were excluded from this evaluation. For each remaining dataset, Spearman’s correlation coefficient (ρ) was calculated for both modeling approaches to quantify the relationship between the substrate sequence length (number of amino acids) and the F1 score.
Spearman correlation is a non-parametric measure of the strength and direction of the association between two ranked variables. It assesses how well the relationship between the variables can be described using a monotonic function. A negative Spearman’s ρ indicates that longer substrate sequences tend to correspond with lower model F1 scores, implying that restricting sequence length may improve predictive performance. This trend holds across all datasets except for S01.010 and C14.003 in the GNN model, where the correlation was not negative.
Since these models are intended for drug discovery case studies involving peptides rather than larger proteins, limiting the substrate sequence length helps better align them with their application. Restricting the sequence length of the substrates in the dataset to a maximum of 200 amino acids improved the F1 scores of the models, as shown in Table 4, with the exception of C14.001, where an insufficient number of substrates with this length prevented effective model training.
At first glance, reducing the substrate length appeared to lead to better performance gains for the GNN model compared to the ESM-2 token classification model. To investigate this further, we employed a one-sided paired permutation test, which is well-suited for this analysis as it avoids the distribution of the data assumptions and allows for a direct comparison of the mean relative improvements between the two models. Specifically, for each of the 29 protease models (detailed in the Supplementary Tables S5 and S6), we computed the relative improvement in F1 score under the reduced-length condition compared to the unrestricted setting, separately for both the GNN and ESM-2 models. We then calculated the paired differences by subtracting the ESM-2 improvement from the GNN improvement for each protease, resulting in 29 paired differences. These differences reflect the per-protease advantage (or disadvantage) of the GNN model over ESM-2 due to substrate length restriction. To construct a null distribution, which represents the scenario in which no systematic difference exists between the two models, we randomly shuffled the signs of these paired differences over multiple iterations. The p-value was then computed as the proportion of permutations in which the mean paired difference is at least as extreme as the observed value. This quantifies the likelihood that the observed improvement occurred by chance, providing statistical support for our hypothesis that GNN benefits more from substrate length reduction than ESM-2.
The mean relative improvement for the ESM-2 token classification model is 25.22%, whereas the GNN model shows a mean relative improvement of 125.30%. The paired permutation test with one million permutations revealed a p-value of 0.0017, indicating high statistical significance in the difference improvements between the GNN model and the ESM-2 token classification model. A histogram of the empirical null distribution for the mean of paired differences can be found in Supplementary Fig. S1.
One possible reason for the observed improvements in model performance relates to the structural differences between peptide and protein structures. The small size of peptides is one of the primary factors contributing to their poor in vivo stability, which prevents them from benefiting from the structural stability provided by well-defined secondary or tertiary structures24,25,26. The ESM-2 token classification model uses a transformer architecture capable of capturing long-range dependencies and obtaining a global receptive field, making it highly effective for analyzing large sequence data, such as proteins. In contrast, GNN models may struggle to accurately represent very long sequences, particularly due to the increased complexity of secondary tertiary structures, as they can lose critical long-range information.
Figure 4 shows the distribution of substrate sequence lengths across the 8 protease datasets. GNN models tend to achieve higher predictive performance on datasets with shorter sequences, such as C14.005.
As done previously with the unrestricted substrate lengths, the F1 scores were also compared against two baselines: the random model and the Constant 1 model. Both the ESM-2 token classification and GNN models considerably outperformed these baselines across all proteases, as shown in Supplementary Table S7.
The current evaluation is based on a single stratified training/test split which preserves class balance. However, this may not fully capture variability across data partitions. More robust evaluation strategies, such as k-fold cross-validation or repeated subsampling, could provide more stable performance estimates and help generalize conclusions more effectively.
Comparison with other in silico cleavage site prediction models
Several in silico cleavage site prediction tools evaluate the predictive performance of their models using multiple independent datasets which correspond to distinct proteases. Among these tools, ProsperousPlus demonstrated superior predictive accuracy compared with the other predictors27.
To ensure a fair comparison, this study exclusively compares against ProsperousPlus, as both utilize the same MEROPS database release, data curation process, and dataset splitting strategy – allocating 70% of the data corresponds to the training and 30% to the test based on the number of cleavages sites on the dataset.
ProsperousPlus is a tool for predicting protease-specific substrate cleavage sites and constructing custom machine-learning models. Its predictors have been benchmarked against other existing methods, showing differences in accuracy, ROC AUC, and other metrics for proteases such as caspases and metallopeptidases.
In ProsperousPlus, cleavage sites are represented using 8-amino-acid windows, with 4 residues upstream and downstream of the cleavage point. Sequence data are encoded through eight scoring functions that produce 17 feature scores, including position-specific matrices (PPM, PWM, PSSM-derived IC50) for evolutionary and structural information, conservation-based metrics (WebLogo), similarity-based scores (Nearest Neighbor Similarity, K-Nearest Neighbors), and substitution matrix indices (SMIs) based on BLOSUM/PAM matrices. These features capture positional amino acid frequencies, sequence conservation, and evolutionary constraints for use in predictive modeling.
The AutoML framework in ProsperousPlus automates model development. It trains nine base machine-learning algorithms, including Logistic Regression, SVM, Random Forest, XGBoost, and LightGBM, using stratified k-fold cross-validation to ensure robust evaluation. From these, the three models with the highest Area Under the Receiver Operating Characteristic Curve (ROC AUC) and accuracy are selected. These models undergo optimization through incremental feature selection (IFS), which iteratively refines the feature set to improve performance. The framework then constructs ensemble models by applying stacking, blending, and bagging techniques to combine the strengths of the selected models. One of the current limitations of this tool is that it requires the FASTA sequence of the peptide as input, as well as the fact that it generates probability score predictions by applying a window size of 8 amino acids to the compound. Consequently, for a peptide substrate consisting of 9 amino acids, only two prediction outputs would be generated, corresponding to positions 4 and 5, as only two windows of 8 amino acids can be formed. This approach misses potential cleavage sites at the start and end positions of the peptide sequence. In turn, this can result in the incorrect calculation of certain metrics, as these positions are wrongly excluded from their calculation.
Based on the originally reported metrics27, the models achieved precision values greater than 90% in nearly all the cases, indicating that the model likely has a very low number of false positives. This is particularly surprising given the highly imbalanced nature of the datasets, with a significantly greater number of non-cleavage sites, which would typically result in a large proportion of negative samples potentially misclassified by the model. Using their published metrics, the ratio of negative to positive samples in the test datasets was calculated, using the formula provided in the Supplementary Equation S1, which expresses this ratio as a function of the accuracy, sensitivity and precision metrics. These ratios differ significantly from those calculated using our entire datasets, as shown in Table 5.
The calculated ratios suggest that an undersampling balancing technique was applied to the test set, using a 1:1 ratio of negative to positive samples, meaning that non-cleavage sites were randomly removed from the test set to match cleavage sites in number, closely in line with what the authors report using a previous study28.
To compare the metrics without applying balancing techniques to the test set, new models were trained using the publicly available ProsperousPlus code. This approach ensured that the test sets substrates did not overlap with those in the training sets of the pre-existing models from the webserver, as the ProsperousPlus datasets and split data are not publicly available. Consistent with the published methodology, undersampling with a 1:1 ratio was applied, except only in the training set, without restricting the maximum length of the input substrates. Undersampling was performed by randomly removing peptide bonds from the negative class until the number of positive and negative samples was equal. The retrained models were then used to generate predictions on the test set and then calculate the actual performance metrics, as presented in Table 6.
To evaluate the alignment between the training methodology using the available code and the published approach, predictions were generated on the test set after undersampling was applied at a 1:1 ratio. This adjustment aimed to assess whether the observed performance trends were consistent with the originally reported metrics under balanced conditions. As presented in Supplementary Table S8, the models exhibited strong performance under these conditions. Furthermore, a paired permutation test comparing the mean F1 score from the reported results with those from the undersampled test set yielded a p-value of 0.1250 (one million permutations), indicating no statistically significant difference. These findings suggest that the training approach used is comparable to that of the ProsperousPlus models.
Table 7 compares the performance of the ESM-2 Token Classification and GNN models with that of the ProsperousPlus approach. The ProsperousPlus model was evaluated both with and without 1:1 undersampling (applied only to the training set). Both the ESM-2 Token Classification and the GNN models outperform ProsperousPlus across all evaluated proteases.
The models were trained on an Intel Core i7-4790 CPU (4 cores, 8 threads, 3.6 GHz) with 32 GB RAM and SSD storage, using Python 3.7.16 on Arch Linux (kernel 6.12.3). ProsperousPlus models required significantly longer training times than neural-network-based models did, primarily due to the computational complexity of their scoring functions for representing training data. Without undersampling, training averaged over 4 days per protease, compared with less than 30 min for neural-network models. Additionally, some proteases encountered errors after several days of training when undersampling was not applied.
In addition to our primary evaluation metrics, we compute the ROC-AUC to enable comparison with ProsperousPlus, who use this metric in their model evaluations. ROC-AUC is a widely recognized measure of classifier performance that captures the trade-off between true positive rate and false positive rate across all thresholds. As a threshold-independent metric, it provides a comprehensive view of a model’s ability to distinguish between the positive and negative classes, making it especially useful in binary classification settings29. The corresponding ROC curves are available in the Supplementary Fig. S2.
In imbalanced datasets, where cleavage sites are much less frequent than the non-sites are, the ROC curve is less informative. As the ROC curve gives equal importance to both classes, a model can achieve high AUC scores even if it does not perform well on the positive class, because the large number of true negatives can dominate the curve. Hence, the ROC curve may not be the best tool to help us understand the model’s performance in detecting actual cleavage sites. Instead, the precision-recall curve provides a clearer assessment of model performance.
The average precision (AP) values, defined as the area under precision-recall curve (PR AUC), provides a more informative assessment of model performance in imbalanced classification settings than ROC-AUC, by focusing on the positive class without being influenced by the large number of negative samples so strongly, while retaining the advantage of being threshold-independent29,30. AP values range from 0 to 1, with values closer to 1 indicating better precision-recall trade-offs. Figure 5 illustrates the precision-recall curves of both developed models in comparison with the state-of-the-art method for cleavage site prediction across 8 proteases.
The ESM-2 token classification model demonstrated the best performance in terms of the PR AUC across all 8 proteases, outperforming both the GNN model and the ProsperousPlus approach. Notably, the C14.005 model has the highest AP value of 0.950. The GNN model demonstrated superior AP values compared with the state-of-the-art method for all proteases, except for M10.005. In summary, the results highlight that both developed approaches achieved higher predictive performance than the current state-of-the-art approach.
For C14.004, M10.005, and S0.010, ProsperousPlus achieved higher ROC-AUC values (Figure S2), likely due to its optimization specifically targeting this metric. While this demonstrates its contextual strengths, our models maintained superior accuracy and precision-recall performance (Fig. 5), reinforcing their overall advantage in practically-relevant metrics.
Precision-recall curves of the ESM-2 Token Classification, GNN models, and state-of-the-art approach (ProsperousPlus) for the cleavage site prediction of caspase-1 (C14.004), caspase-3 (C14.003), caspase-7 (C14.004), caspase-6 (C14.005), MMP-2 (M10.003), MMP-3 (M10.005), granzyme B (S01.010), and thrombin (S01.217) in the test datasets.
Case studies
To demonstrate the applicability of the GNN model, predictions were computed on a selection of peptide substrates with cyclic structures containing unnatural amino acids. The cleavage sites of these peptides were determined experimentally in prior studies31,32 by incubating each substrate with various protease enzymes, such as elastase-2 (S01.131), cathepsin G (S01.133) and chymotrypsin A (S01.001), and subsequently monitoring the resulting metabolites using liquid chromatography-high-resolution mass spectrometry. Detailed experimental conditions for this dataset are provided in the literature31,32.
To more closely reflect practical conditions, given that the experimental data consisted of peptides with a maximum length of 20 amino acids, the models were trained with a substrate length limit of 200 amino acids. A shorter length was not chosen because it would not provide enough training samples. Under this constraint, the models achieved F1 scores of 0.3604 for elastase-2, 0.4000 for cathepsin G, and 0.3937 for chymotrypsin A.
The peptides used in this study are all cyclized via disulfide bridges between distant cysteine residues (side chain cyclized). To perform the prediction, the complete peptide structure is utilized in its native, cyclic form without prior linearization. The top prediction for all peptides is shown in Fig. 6. For elastase-2, a prediction was made for somatostatin, a 14-amino acid peptide. The predicted cleavage site occurs between Thr(12) and Ser(13), which corresponds to one of the two first-generation metabolites previously determined experimentally and reported in the literature31.
For cathepsin G, the prediction was made on iseganan, a 17-amino acid peptide containing two disulfide linkages and a C-terminal modification with the non-standard amino acid L-argininamide. The predicted cleavage site between Phe(11) and Cys(12) corresponds to one of the two first-generation metabolites reported in prior research32.
The chymotrypsin A model was utilized to predict the cleavage sites for octreotide, and oxytocin. Octreotide is an 8-amino acid peptide containing two unnatural amino acids, D-phenylalanine and D-tryptophan, as well as a C-terminal modification with the non-standard amino acid L-threoninol. For octreotide, the cleavage site was predicted to occur between DTrp(4) and Lys(5), which aligns with one of the three first-generation metabolites reported in the literature32.
Oxytocin is a 9-amino acid peptide containing a C-terminal modification with the non-standard amino acid L-glycinamide. For oxytocin, the model predicts cleavage between Tyr(2) and Ile(3), corresponding to one of the two first-generation metabolites identified experimentally32.
These examples showcase the model’s ability to effectively handle peptides with cyclic structures and unnatural amino acids, despite not having been previously trained with such data, a domain not addressed in prior works. While these case study results demonstrate promising performance of the GNN model on cyclic peptides with unnatural amino acids, they represent a limited evaluation. Further studies involving a larger and more diverse set of cyclic peptides are necessary to fully establish the model’s generalizability. However, such efforts are currently constrained by the limited availability of publicly accessible data on such compounds.
Conclusions
This study presents two novel approaches for predicting peptide cleavage sites. The first uses the pre-trained protein language model ESM-2, which has been fine-tuned to use its generated structure embeddings for accurate cleavage site prediction on linear peptides with natural amino acids. This approach eliminates the need for traditional, manual feature extraction methods employed by current in silico tools.
The second approach is the first method developed for predicting cleavage sites in cyclic peptides, including those with non-natural amino acids, by employing GNNs. This methodology is compatible with the strategies used for enhancing the therapeutic potential of peptides. To demonstrate its applicability, models trained on publicly available data were used to predict cleavage sites in a focused case study involving side chain cyclized peptides incorporating non-natural amino acids and C-terminal modifications, with in silico predictions matching experimental metabolite identification data.
Despite the evident class imbalance in the collected datasets both approaches exhibit promising predictive performance compared with the state-of-the-art methods, achieving improved F1 scores for all studied proteases. Importantly, these results were obtained without applying any balancing techniques to the test set, thereby reflecting a realistic evaluation scenario. To further strengthen the reliability and generalizability of the findings, future work should consider adopting more robust evaluation strategies, such as k-fold cross-validation or repeated subsampling.
While the performance of the GNN model is promising, further validation on external datasets containing a broader and more diverse range of cyclic peptides is necessary to fully assess its generalizability. Progress in this area will rely heavily on the expansion of publicly accessible datasets that include structurally complex and modified peptides. In parallel, future research could explore the development of an enhanced GNN architecture to more effectively capture long-range dependencies and the inherent structural complexity of proteins.
Data availability
The ESM-2 token classification code is freely available at the repository: https://anonymous.4open.science/r/2ae195dac097002e030618/. The GNN code is freely available at the repository: https://anonymous.4open.science/r/merops-soc-gnn-633 A/. All datasets used in this study are provided in the following repository: https://anonymous.4open.science/r/Supplementary-Information-Files-188 F/.
Change history
26 November 2025
The original online version of this Article was revised: The original version of this Article contained an error in the Acknowledgements section where one of the grant numbers was inadvertently omitted. The Acknowledgements section now reads: “This research project was financially supported by the Generalitat de Catalunya and Lead Molecular Design S.L through the Industrial Doctorate (Grant numbers: 00002/2023 and 00006/2023).” The original Article has been corrected.
References
Rossino, G. et al. Peptides as therapeutic agents: challenges and opportunities in the green transition era. Molecules 28, 7165. https://doi.org/10.3390/molecules28207165 (2023).
Lamers, C. Overcoming the shortcomings of peptide-based therapeutics. Future Drug Discov. 4 (2). https://doi.org/10.4155/fdd-2022-0005 (2022).
Yao, J. F. et al. Metabolism of peptide drugs and strategies to improve their metabolic stability. Curr. Drug Metab. 19, 892–901. https://doi.org/10.2174/1389200219666180628171531 (2018).
Deb, P. K. et al. Protein/peptide drug delivery systems: practical considerations in pharmaceutical product development. In Advances in Pharmaceutical Product Development and Research (ed. Tekade, R. K.) 651–684 (Academic Press, 2019). https://doi.org/10.1016/B978-0-12-817909-3.00016-9.
Gentilucci, L. et al. Chemical modifications designed to improve peptide stability: incorporation of non-natural amino acids, pseudo-peptide bonds, and cyclization. Curr. Pharm. Des. 16, 3185–3203. https://doi.org/10.2174/138161210793292555 (2010).
Evans, B. et al. (ed, J.) Methods to enhance the metabolic stability of peptide-based PET radiopharmaceuticals. Molecules 25 2314 https://doi.org/10.3390/molecules25102314 (2020).
Song, J. et al. iProt-Sub: a comprehensive package for accurately mapping and predicting protease-specific substrates and cleavage sites. Brief. Bioinform. 20, 638–658. https://doi.org/10.1093/bib/bby028 (2019).
Li, F. et al. Twenty years of bioinformatics research for protease-specific substrate and cleavage site prediction: a comprehensive revisit and benchmarking of existing methods. Brief. Bioinform. 20, 2150–2166. https://doi.org/10.1093/bib/bby077 (2019).
Chen, J. Y. et al. Evaluating the advancements in protein Language models for encoding strategies in protein function prediction: a comprehensive review. Front. Bioeng. Biotechnol. 13, 1506508. https://doi.org/10.3389/fbioe.2025.1506508 (2025).
Radchenko, T., Fontaine, F., Morettoni, L. & Zamora, I. Software-aided workflow for predicting protease-specific cleavage sites using physicochemical properties of the natural and unnatural amino acids in peptide-based drug discovery. PLoS One. 14, e0199270. https://doi.org/10.1371/journal.pone.0199270 (2019).
Rawlings, N. D. & Bateman, A. How to use the MEROPS database and website to help understand peptidase specificity. Protein Sci. 30, 83–92. https://doi.org/10.1002/pro.3948 (2021).
Ieremie, I. et al. Protein Language models Meet reduced amino acid alphabets. Bioinformatics 40, btae061. https://doi.org/10.1093/bioinformatics/btae061 (2024).
Lin, Z. et al. Language models of protein sequences at the scale of evolution enable accurate structure prediction. bioRxiv https://doi.org/10.1101/2022.07.20.500902 (2022).
Zhang, R. et al. PepLand a large-scale pre-trained peptide representation model for a comprehensive landscape of both canonical and non-canonical amino acids. arXiv https://doi.org/10.48550/arXiv.2311.04419 (2023).
UniProt Consortium UniProt: the universal protein knowledgebase. Nucleic Acids Res. 46, 2699. https://doi.org/10.1093/nar/gky092 (2018).
Fu, L. et al. CD-HIT: accelerated for clustering the next-generation sequencing data. Bioinformatics 28, 3150–3152. https://doi.org/10.1093/bioinformatics/bts565 (2012).
Rives, A. et al. Biological structure and function emerge from scaling unsupervised learning to 250 million protein sequences. Proc. Natl. Acad. Sci. U.S.A. 118, e2016239118. https://doi.org/10.1073/pnas.2016239118 (2021).
Vaswani, A. et al. Attention is all you need. ArXiv [cs CL]. https://doi.org/10.48550/arXiv.1706.03762 (2017).
Kensert, A., Desmet, G. & Cabooter, D. MolGraph a Python package for the implementation of molecular graphs and graph neural networks with TensorFlow and Keras. arXiv https://doi.org/10.48550/arXiv.2208.09944 (2022).
Scarselli, F., Gori, M., Tsoi, A. C., Hagenbuchner, M. & Monfardini, G. The graph neural network model. IEEE Trans. Neural Networks. 20, 61–80. https://doi.org/10.1109/TNN.2008.2005605 (2009).
Reiser, P. et al. Graph neural networks for materials science and chemistry. Commun. Mater. 3, 93. https://doi.org/10.1038/s43246-022-00315-6 (2022).
Khemani, B. et al. A review of graph neural networks: concepts, architectures, techniques, challenges, datasets, applications, and future directions. J. Big Data 11, 18. https://doi.org/10.1186/s40537-023-00876-4 (2024).
RDKit: Open-source cheminformatics. Available at: https://www.rdkit.org.
Wang, L. et al. Therapeutic peptides: current applications and future directions. Signal Transduct. Target. Ther. 7, 48. https://doi.org/10.1038/s41392-022-00904-4 (2022).
Gupta, S., Azadvari, N. & Hosseinzadeh, P. Design of protein segments and peptides for binding to protein targets. Biodes. Res. 2022, 9783197. https://doi.org/10.34133/2022/9783197 (2022) (PMCID: PMC10521657).
Jiang, Y. et al. PHAT: interpretable prediction of peptide secondary structures using hypergraph multi-head attention network and transfer learning. bioRxiv 2022.06.09.495580. https://doi.org/10.1101/2022.06.09.495580 (2022).
Li, F. et al. ProsperousPlus: a one-stop and comprehensive platform for accurate protease-specific substrate cleavage prediction and machine-learning model construction. Brief. Bioinform 24, bbad372. https://doi.org/10.1093/bib/bbad372 (2023).
Song, J. et al. PROSPER: an integrated feature-based tool for predicting protease substrate cleavage sites. PLoS One. 7, e50300. https://doi.org/10.1371/journal.pone.0050300 (2012).
Brodersen, K. H. et al. The binormal assumption on precision-recall curves. In Proceedings of the 20th International Conference on Pattern Recognition, 4263–4266 (IEEE, 2010). https://doi.org/10.1109/ICPR.2010.1036.
Davis, J. & Goadrich, M. The relationship between precision-recall and ROC curves. In Proc. 23rd Int. Conf. Mach. Learn., 233–240 (Association for Computing Machinery, 2006). https://doi.org/10.1145/1143844.1143874.
Wesche, F. et al. Automated high-throughput in vitro assays to identify metabolic hotspots and protease stability of structurally diverse, Pharmacologically active peptides for inhalation. J. Pharm. Biomed. Anal. 211, 114518. https://doi.org/10.1016/j.jpba.2021.114518 (2022).
Radchenko, T. et al. Software-aided approach to investigate peptide structure and metabolic susceptibility of amide bonds in peptide drugs based on high resolution mass spectrometry. PLoS ONE. 12, e0186461. https://doi.org/10.1371/journal.pone.0186461 (2017).
Acknowledgements
This research project was financially supported by the Generalitat de Catalunya and Lead Molecular Design S.L through the Industrial Doctorate (Grant numbers: 00002/2023 and 00006/2023).
Author information
Authors and Affiliations
Contributions
Paula Cifuentes and Ramon Adàlia wrote the main manuscript text, designed the study, coordinated the study, carried out the data analysis, and interpreted the results. Ismael Zamora supervised this work. All the authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Cifuentes, P., Adàlia, R. & Zamora, I. Prediction of peptide cleavage sites using protein language models and graph neural networks. Sci Rep 15, 38048 (2025). https://doi.org/10.1038/s41598-025-21801-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-21801-0