Abstract
Understanding the mechanisms of endophytic fungal transmission is crucial for deciphering plant–microbe interactions and leveraging microbiomes in crop improvement. In this study, we examined the potential for intergenerational inheritance and external recruitment of endophytic fungi in common wheat. Fungal community structure was compared in generation G1 and parental G0 plants using ITS2 metabarcoding data and culture-based identification in four tissues (roots, stems, leaves, and seeds) in ten cultivars. A set of 27 operational taxonomic units (OTUs) was consistently detected in both generations, suggesting the presence of a core mycobiome dominated by genera such as Fusarium, Trichoderma, Penicillium, Cladosporium, Sarocladium and Lecanicillium. Experimental inoculation of axenic wheat seedlings with fungi Fusarium proliferatum, Penicillium expansum, Trichoderma hamatum originating from the wheat endosphere confirmed the ability of these species to recolonize host tissues and thus their role in stable association with plants. However, several other taxa (Engyodontium album, Sarocladium spinificis, Clonostachys candelabrum, and Nigrospora gorlenkoana) originating from internal wheat tissues failed to recolonize axenic plants, suggesting transient colonization via horizontal pathways. These findings highlight the dual contribution of vertical inheritance and environmental recruitment in shaping the wheat endomycobiome, offering a foundation for targeted manipulation of beneficial fungi in cereal crops.
Similar content being viewed by others
Introduction
The endosphere of common wheat (Triticum aestivum L.)—a cosmopolitan species from the Poaceae family that evolved through human selection and is now one of the most widely cultivated cereals worldwide—provides a habitat for various microorganisms, including fungi1. According to a recent literature review, fungi that “colonise the internal tissues of a plant for all or part of their life”2 and “do not cause disease in the plant under any known circumstances”3 are called endophytes. This rather general definition masks gaps in knowledge about the functions of fungi associated with internal plant tissues. There are indications that the role of an endophytic fungus in a plant is determined not only by its genome, but by the set of all genomes co-functioning in the plant holobiome4,5. Common wheat, due to its domestication in the early Holocene and subsequent dispersal of cultivation around the world, where a variety of breeding practices are used, exhibits considerable genetic diversity in forms and cultivars. This diversity likely has a significant impact on determining the composition of the endophytic fungal community. It has been documented that, in addition to the wheat domestication, breeding and genotype, its growth stage and plant compartment may also shape spatiotemporal changes in the structure of the endo-mycobiome6,7,8,9,10. In addition to host plant factors, environmental factors such as abiotic and biotic stress, geographic location and growing season can also account for differences in wheat mycobiome11,12,13. Also soil properties (edaphic factors), especially soil pH, drive changes in the structure of fungal communities12,14. The dynamics of changes in the composition of the endo-mycobiome are also influenced by anthropogenic factors, mainly the tillage intensity and management type10,14,15.
A key role in establishing the composition of the endophyte community is played by two modes of their transmission to plant tissues: horizontal and vertical. Horizontal transmission refers to endophyte fungi acquired via their airborne spores (air source), carried by insects, birds or other animals (animal source), through the rhizosphere soil (soil source) or via pollen from another plant that is the host of the fungus16. This is the most common way for endophytic fungi to spread to wider areas or entire ecosystems, within the same or another plant species. While vertical transmission spreads endophytic fungi exclusively in the genetic lineage from the parent plant to the offspring via host seeds or vegetative propagules, thus establishing a close and strong association with the host plants17,18. It was documented that the same fungus species can adopt both—vertical and horizontal—transmission strategies at the same time, which increases its potential to spread to entire populations in the ecosystem19.
Recently, we characterised a structure of endogenous fungal communities in four organs (root, stem, leaf, grain) of common wheat, five spring and five winter cultivars, grown in different conditions, i.e.: (i) field cultivation with conventional mineral fertilisation and herbicides, but without fungicides, (ii) no-till field cultivation without mineral fertilization, herbicides and fungicides, and (iii) cultivation under controlled greenhouse conditions with mineral fertilisation10. These studies have shown that the endomycobiome profile of wheat depends mainly on the type of plant organs and growth conditions, and the management strategy has a greater impact on the structure of the fungal community than on its abundance. Since the plant material consisted of Polish wheat cultivars with relatively low genetic diversity due to their common origin and breeding history, the influence of the host genotype, including the wheat form, on the formation of the endospheric mycobiome composition was not detectable. Finally, the endogenous core mycobiome of common wheat was defined, and 726 organ-, genotype- and growth condition-specific isolates were obtained from wheat tissues10. Despite this knowledge, questions remain regarding the recruitment and persistence of these fungi, particularly across generations.
It was therefore hypothesised that some fungal taxa, especially those constituting the core mycobiome, may exhibit intergenerational persistence under controlled conditions, potentially indicating vertical transmission mechanisms in wheat. However, there are also some that are recruited from the external environment in a horizontal manner and reside in wheat tissues temporarily. Accordingly, the aim of the current study was to: (i) characterise the structure of the fungal community colonizing the tissues of progeny plants (generation 1, G1) derived from seeds of parental plants (generation 0, G0) cultivated under controlled greenhouse conditions, (ii) compare the structure of the endomycobiome of parental G0 and progeny G1 plants growing in controlled greenhouse conditions, (iii) assess the ability of selected endophytic fungi, originally isolated from G0 plants, to recolonize axenic tissues of G1 seedlings under phytotron conditions.
Results
Endophytic fungi community structure in G1 plants
High-throughput cultivation-independent analyses—G1 plants
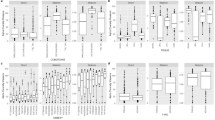
Based on high-throughput sequencing (HTS) data, we first quantified within-sample (alpha) diversity to assess differences in fungal richness and evenness across plant organs (tissues), forms and cultivars. Analysis of alpha diversity revealed statistically significant differences in fungal richness and diversity among tissues (Fig. 1a). Shannon diversity index varied notably between leaves and kernels (p = 0.009) as well as leaves and stems (p = 0.026). Similarly, the number of observed OTUs differed between kernels and roots (p = 0.03) and between leaves and roots (p = 0.04). Beta diversity analysis (Weighted UniFrac) showed significant (p < 0.002) variation in fungal community structure exclusively between tissues (Fig. 1b).
High-throughput cultivation-independent analyses showed that the fungal community composition of the studied cultivars (Figs. 2a and c, Fig. S1) and tissues (Fig. 2b and d, Fig. S2) differed significantly, with the core genera group remaining relatively stable.
The classes Sordariomycetes and Leotiomycetes were the most abundant among the studied tissues (Fig. 2). The members of class Sordariomycetes predominated in roots (93.5%), mainly Fusarium (73.7%) and Sarocladium (14.8%), but also Setophoma (3.6%) and Phialemonium (3.2%). In the aboveground tissues (kernels, leaves, and stems), communities were dominated by Leotiomycetes driven by Blumeria—leaf 99.5%, stem 95.7%. Sordariomycetes accounted for only 0.4% in leaves and 2.7% in stems. The mycobiome of kernels was characterised by very low diversity and strong dominance of Blumeria (92.1%); Sordariomycetes contributed modestly (e.g., Fusarium 3.35%, Sarocladium 0.64%, Setophoma 0.12%, Phialemonium 0.05%). In addition, Penicillium was detected in stems (1.02%) and kernels (0.78%), Aspergillus in kernels (0.51%), Cladosporium in leaves (0.03%), and Trichoderma in stems (0.32%). Other classes, such as Eurotiomycetes and Dothideomycetes occurred at smaller but consistent levels, especially in roots (1.3% and 3.6%) and kernels (1.5% and 1.3%).
The classes Sordariomycetes and Leotiomycetes were also the most abundant among the studied cultivars (Fig. 2). The members of class Sordariomycetes predominated in Arabella, Bamberka, Bombona, Euforia, Kandela, Legenda, Ostroga, and Rusałka cultivars, whereas Leotiomycetes dominated in the varieties Arkadia and Rospuda. The most common fungal genera in the studied varieties were Fusarium (58.9%), Blumeria (20.9%), Sarocladium (11.8%), Setophoma (2.8%), Phialemonium (2.6%), and Trichocladium (0.6%). The genus Fusarium was detected in high abundance across nearly all wheat cultivars, with particularly elevated levels observed in Arabella (88.2%), Bombona (79.9%), Kandela (98.6%), Ostroga (77.9%), and Rusalka (65.4%). In contrast, Blumeria genus showed the highest relative abundance in the Arkadia (55.1%), Bamberka (41.3%), and Rospuda (93.8%) cultivars.
Other classes, such as Dothideomycetes and Eurotiomycetes, were detected in low abundance (2.9% and 1.1%, respectively), and restricted to a few cultivars (Fig. 2). Dothideomycetes were most pronounced in Euforia (16.2%) and Ostroga (10.3%), with lower levels in Arabella (3.2%) and Legenda (1.8%), and only trace amounts (≤ 0.8%) in Arkadia, Bamberka, Bombona, Kandela, Rospuda, and Rusalka. Eurotiomycetes occurred in every cultivar but remained ≤ 3.2%, peaking in Rospuda (3.19%), Bombona (3.07%), and Bamberka (2.58%), and staying ≤ 1.1% in the remaining cultivars.
Analysis of alpha and beta (Weighted UniFrac) diversity showed no significant difference between wheat cultivars or forms (spring vs. winter) (Fig. S3). Moreover, high-throughput cultivation-independent analyses revealed no significant differences in the fungal community composition between wheat forms (spring vs. winter) (Fig. S4).
The culture- based endophyte identification—G1 plants
The culture-dependent approach led to the isolation of 59 fungi representing 14 taxa from the endosphere of G1 progeny plants (Table 1, Fig. 3a–c). The dominant species in this generation of plants, regardless of the cultivar and plant organ, was F. proliferatum. In addition, the following fungal taxa were identified: Penicillium sp., Penicillium crustosum, Penicillium olsonii, Cladosporium sp., Trichoderma sp., Trichoderma hamatum, Trichoderma koningii, Trichoderma asperellum, Trichoderma atroviride, Trichoderma harzianum, Lecanicillium lecanii, Simplicillium chinense, Aspergillus westerdijkiae. The number of unique and common species with respect to form and plant organ was presented in Fig. 4.
Network graphs for fungal species isolated from G1 plants by: (a) variety; (b) organ; (c) wheat form. Numbers indicate species: 1. Acremonium sclerotigenum 2. Acremonium sp. 3. Alternaria sp. 4. Aspergillus sp. 5. Aspergillus westerdijkiae 6. Chrysosporium pseudomerdarium 7. Cladosporium cladosporioides 8. Cladosporium sp. 9. Clonostachys candelabrum 10. Engyodontium album 11. Fusarium proliferatum 12. Fusarium sp. 13. Geomyces pannorum 14. Lecanicilium lecanii 15. Lecanicillium lecanii 16. Nigrospora gorlenkoana 17. Penicilium crustosum 18. Penicilium olsonii 19. Penicilium sp. 20. Penicillium chrysogenum 21. Penicillium crustosum 22. Penicillium digitatum 23. Penicillium expansum 24. Penicillium olsonii 25. Phlebia sp. 26. Sarocadium spinificis 27. Sarocladium sp. 28. Sarocladium spinificis 29. Sarocladium strictum 30. Simplicilium chinense 31. Trichoderma asperellum 32. Trichoderma atroviride 33. Trichoderma hamatum 34. Trichoderma harzianum 35. Trichoderma koningii 36. Trichoderma sp. 37. Trichoderma viride. Asterisk* indicates species that were also identified in G0 generation.
Fungal community shifts across G0 and G1 plant generations—vertical transmission
High-throughput cultivation-independent analyses—G0 and G1 plants
Taxonomic classification of combined ITS metabarcoding reads obtained for two generations (G0 and G1) of ten cultivars (Arabella, Bombona, Kandela, Rospuda, Rusałka, Arkadia, Bamberka, Euforia, Legenda, Ostroga) and two forms (winter and spring) of common wheat by analyzing four plant organs (root, stem, leaves, seeds) revealed the occurrence of 27 OTUs common to both plant generations, 3 OTUs unique to G0 and 5 unique to G1.
Despite the generational change, 27 fungal OTUs were consistently detected in both parental (G0) and progeny (G1) wheat plants. These shared OTUs belonged to a diverse group of genera, including Alternaria, Aureobasidium, Chrysosporium, Colletotrichum, Conioscypha, Epicoccum, Fusarium, Gibberella, Lecanicillium, Malbranchea, Penicillium, Phialemonium, Puccinia, Sarocladium, Setophoma, Talaromyces, and Trichoderma, suggesting the existence of a potentially heritable or environmentally stable endophytic core community. However, the overall degree of tissue colonization by endogenous fungi was significantly lower in the G0 generation than in the G1 generation of wheat plants, hence, the abundance of those taxa recorded as common to both generations was significantly lower in G0 than in G1 (Fig. 5a). Moreover, G1 generation exhibited a higher diversity in the relative abundance of fungi across different plant tissues (Fig. 5b–d). These intergenerational differences in fungal community structure were statistically significant (Weighted UniFrac;p = 0.001; Fig. 5b–d).
(a) Relative abundance of fungal taxa in wheat tissues across two plant generations (G0 and G1); (b, c, d) Principal Coordinates Analysis (PCoA) based on Weighted UniFrac distances showing differences in endophytic fungal community composition between the two generations, considering tissue (b), variety (c) and type (d).
The culture- based endophyte identification—G0 and G1 plants
Comparing the two generations of plants based on DNA barcoding data, it was observed that only 7 fungal taxa were common to the G0 parental and G1 progeny plants (Fig. 6). These were: Penicillium sp., P. crustosum, P. olsonii, Cladosporium sp., Trichoderma sp., L. lecanii, T, hamatum, F. proliferatum and T. koningii.
Network graph (a) and venn diagram (b) comparing two (G0 and G1) wheat populations with respect to the fungi isolated from their endosphere. Numbers indicate species name: 1. A. sclerotigenum 2. Acremonium sp. 3. Alternaria sp. 4. Aspergillus sp. 5. A. westerdijkiae 6. Ch. pseudomerdarium 7. C. cladosporioides 8. Cladosporium sp. 9. C. candelabrum 10. E. album 11. F. proliferatum 12. Fusarium sp. 13. G. pannorum 14. L. lecanii 15. L. lecanii 16. N. gorlenkoana 17. P. crustosum 18. P. olsonii 19. Penicilium sp. 20. P. chrysogenum 21. P. crustosum 22. P. digitatum 23. P. expansum 24. P. olsonii 25. Phlebia sp. 26. S. spinificis 27. Sarocladium sp. 28. S. spinificis 29. S. strictum 30. S. chinense 31. T. asperellum 32. T. atroviride 33. T. hamatum 34. T. harzianum 35. T. koningii 36. Trichoderma sp. 37. T. viride. Asterisk* indicates species that were identified in both generations.
The distribution of endophytic fungi isolated from wheat plants with respect to cultivars, organs (roots, stems, leaves, seeds) and forms (spring, winter) for two (G0, G1) generations was illustrated in Figs. 7, 8 and 9, respectively. Some species were consistently associated with multiple cultivars in both generations—including Cladosporium sp., F. proliferatum, L. lecanii, Penicillium sp., T. hamatum, T. koningii, and Trichoderma sp.—others were unique to specific cultivars or appeared only in one generation.
Network graphs of the endophytic fungi structure isolated from plants of populations G0 (a) and G1 (b) and regardless of generation (c) with respect to wheat cultivar. Numbers indicate species name: 1. A. sclerotigenum 2. Acremonium sp. 3. Alternaria sp. 4. Aspergillus sp. 5. A. westerdijkiae 6. Ch. pseudomerdarium 7. C. cladosporioides 8. Cladosporium sp. 9. C. candelabrum 10. E. album 11. F. proliferatum 12. Fusarium sp. 13. G. pannorum 14. L. lecanii 15. L. lecanii 16. N. gorlenkoana 17. P. crustosum 18. P. olsonii 19. Penicilium sp. 20. P. chrysogenum 21. P. crustosum 22. P. digitatum 23. P. expansum 24. P. olsonii 25. Phlebia sp. 26. S. spinificis 27. Sarocladium sp. 28. S. spinificis 29. S. strictum 30. S. chinense 31. T. asperellum 32. T. atroviride 33. T. hamatum 34. T. harzianum 35. T. koningii 36. Trichoderma sp. 37. T. viride. Asterisk* indicates species that were identified in both generations.
Network graphs and venn diagrams for the structure of endophytic fungi isolated from plants of populations G0 (a) and G1 (b) and regardless of generation (c) with respect to wheat organ. Numbers indicate species name: 1. A. sclerotigenum 2. Acremonium sp. 3. Alternaria sp. 4. Aspergillus sp. 5. A. westerdijkiae 6. Ch. pseudomerdarium 7. C. cladosporioides 8. Cladosporium sp. 9. C. candelabrum 10. E. album 11. F. proliferatum 12. Fusarium sp. 13. G. pannorum 14. L. lecanii 15. L. lecanii 16. N. gorlenkoana 17. P. crustosum 18. P. olsonii 19. Penicilium sp. 20. P. chrysogenum 21. P. crustosum 22. P. digitatum 23. P. expansum 24. P. olsonii 25. Phlebia sp. 26. S. spinificis 27. Sarocladium sp. 28. S. spinificis 29. S. strictum 30. S. chinense 31. T. asperellum 32. T. atroviride 33. T. hamatum 34. T. harzianum 35. T. koningii 36. Trichoderma sp. 37. T. viride. Asterisk* indicates species that were identified in both generations.
Network graphs and venn diagrams for the structure of endophytic fungi isolated from plants of populations G0 (a) and G1 (b) and regardless of generation (c) with respect to wheat form. Numbers indicate species name: 1. A. sclerotigenum 2. Acremonium sp. 3. Alternaria sp. 4. Aspergillus sp. 5. A. westerdijkiae 6. Ch. pseudomerdarium 7. C. cladosporioides 8. Cladosporium sp. 9. C. candelabrum 10. E. album 11. F. proliferatum 12. Fusarium sp. 13. G. pannorum 14. L. lecanii 15. L. lecanii 16. N. gorlenkoana 17. P. crustosum 18. P. olsonii 19. Penicilium sp. 20. P. chrysogenum 21. P. crustosum 22. P. digitatum 23. P. expansum 24. P. olsonii 25. Phlebia sp. 26. S. spinificis 27. Sarocladium sp. 28. S. spinificis 29. S. strictum 30. S. chinense 31. T. asperellum 32. T. atroviride 33. T. hamatum 34. T. harzianum 35. T. koningii 36. Trichoderma sp. 37. T. viride. Asterisk* indicates species that were identified in both generations.
Tissue recolonization by wheat-derived endophytic fungi—horizontal transmission
Axenic wheat seedlings obtained for 10 cultivars were inoculated with 8 species (F. proliferatum, P. olsonii, P. expansum, T. hamatum, E. album, S. spinificis, C. candelabrum, N. gorlenkoana) of fungi individually and with a mixture of their spores. Re-isolation of fungi from wheat seedlings and their molecular identification showed that E. album, S. spinificis, C. candelabrum and N. gorlenkoana were unable to successfully colonize wheat tissues following spore inoculation. Meanwhile, P. olsoni inhabited seedlings of all wheat varieties, as did P. expansum, although the occurrence of the latter species was not confirmed in the Bamberka variety. Fusarium proliferatum was reisolated from seedlings of 8 cultivars, except for the cultivars Bombona and Ostroga. Meanwhile, T. hamatum demonstrated the ability to reconnect with the tissues of 7 tested varieties, namely: Arabela, Arkadia, Bamberka, Euforia, Legenda, Ostroga, Rospuda. Treatment of wheat seedlings with a mixture of spores of all selected species led to the colonization of the Arabella cultivars Arkadia, Bamberka, Legenda and Ostroga exclusively by P. expansum, the Kandela cultivar exclusively by T. hamatum, the Rospuda cultivar by two Penicillium species—P. expansum and P. olsoni, and the Bombona, Euforia and Rusałka cultivars simultaneously by P. expansum and F. proliferatum.
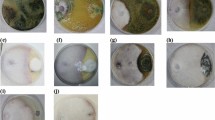
To confirm the presence of fungi in the tissues of inoculated wheat seedlings, microscopic observations were performed. As shown in Fig. 10, fungal hyphae were found on the root surface (Fig. 10B, C, F1) and were visible inside the rhizodermal cells (Fig. 10D–F). Furthermore, it was noted that fungi can produce spores on the roots (Fig. 10B, C, D2). Specific coiling structures were observed in the tissues of plants treated with all tested strains (Fig. 10F1). The observations were based on morphological differences between control plants (plants treated with sterile water) and plants inoculated with fungi.
Colonization of wheat roots by fungi. Micrographs were taken on light microscopy. The materials were dyed with trypan blue. A–A1—control, B–B1—P. olsonii, C—P. expansum, D–D1—T. hamatum, E–E1—F. proliferatum, F–F1—mix. Black arrows indicate hyphae of fungi, black dotted arrows indicate on B and C spores of Penicillium sp., on F1 coiling hyphae. Barr = 20 μm.
Discussion
Despite increasing interest in plant-associated microbiota, there is still no clear definition of endophytic fungi, nor a comprehensive understanding of their functional roles within host plants. Similarly, mechanisms governing the transmission of fungi into the plant endosphere remain only partially understood. The lack of this knowledge is particularly noticeable in the case of crops such as wheat. Addressing the unresolved questions related to the transmission of endophytic fungi associated with common wheat is of critical importance. When combined with information on the structure of the endophytic fungi communities, their dynamics and functioning, such knowledge could enable targeted manipulation of the wheat mycobiome to qualitatively and quantitatively improve its yield.
Recently, Sharon et al.16 investigated the vertical transmission of endophytic fungi in common wheat, wild emmer wheat (Triticum turgidum ssp. dicoccoides), and wild barley (Hordeum spontaneum). In a controlled greenhouse experiment, they demonstrated that certain fungal taxa are capable of vertical transmission from parent to offspring via seeds. Their study focused on monitoring the composition of fungal endophytes in the original seeds and subsequently in the stems and seeds of the resulting progeny. In the present study, to assess the potential for intergenerational transmission of endophytic fungi, a comparative analysis of the endomycobiome was conducted across multiple tissues—roots, stems, leaves, and seeds—of both parental and progeny plants. Both generations were cultivated under strict sterile greenhouse conditions to minimize external fungal contamination. Importantly, the initial material for each G0 plant consisted of seeds obtained directly from breeding companies, while each G1 plant was grown from seeds produced by its corresponding G0 individual. This design enabled detailed tracking of fungal community composition along the following developmental pathway: G0 seeds → roots, stems, leaves → G1 seeds → roots, stems, leaves → G2 seeds. This observation aligns with the findings of Nelson33 and Samreen34, who emphasized the pivotal role of seed-borne endophytes in shaping plant microbiomes throughout different developmental stages. In the present study, several fungal taxa previously reported as wheat endophytes9,10,16,33,34,35,36,37,38,39 were detected, including Alternaria, Aureobasidium, Epicoccum, Chaetomium, Chrysosporium, Cladosporium, Colletotrichum, Fusarium, Lecanicillium, Microdochium, Penicillium, Puccinia, Sarocladium, Setophoma, Talaromyces, and Trichoderma. These taxa can therefore be considered potential candidates for intergenerational transmission via internal plant tissues. The presence of certain species from these genera—such as Penicillium sp., Penicillium crustosum, Penicillium olsonii, Cladosporium sp., Trichoderma sp., Lecanicillium lecanii, Trichoderma hamatum, Fusarium proliferatum, and Trichoderma koningii—was confirmed in both plant generations using a culture-dependent approach. Although this method is generally regarded as having limitations, particularly in capturing the full diversity of fungal communities, it nevertheless supports the persistence of selected endophytic taxa across generations40. A detailed discussion of these species in the context of their ecological importance as pathogens, symbionts, or saprophytes was previously provided by Salamon et al.10. In the current study, the consistent detection of these fungi across generations, combined with successful recolonization of axenic seedlings by selected isolates, provides strong support for their ecological integration within the wheat endosphere. Evidence has been presented that some fungi, such as those from the genus Fusarium, although known mainly as plant pathogens, may undergo numerous and frequent lifestyle transitions due to their highly diverse and dynamic genome40,41. It has been demonstrated that fungal lifestyles can switch in response to changes in their nutritional patterns and are influenced by various factors, including mutualistic relationships, host tolerance, and evasion of host immune responses40,41,42. Our experimental conditions, including the germination of surface-sterile seeds and growth in sterile sand, may have facilitated the survival and vertical transfer of certain fungal endophytes by eliminating environmental competitors and stabilising host–microbe interactions. Nonetheless, the molecular and ecological mechanisms enabling these fungi to persist across generations remain poorly understood and warrant further investigation.
Horizontal colonization of internal plant tissues is a more common mode of transmission of endophytic fungi than the vertical pathway40. Endophytes known to be associated with C3 subfamily Pooide grasses are mainly restricted to vertical transmission. Meanwhile, most known endophytes have evolved a horizontal or mixed horizontal-vertical method of penetrating the plant endosphere. Typically, horizontal transmission occurs by spores or hyphae carried passively by vectors (herbivores, insects) or by natural events (wind, rain). Tissue colonization by these endophytes occurs through wounds and stomatal infiltrates, and in some cases by active penetration of the cuticle. Endophytes are known to form infectious structures such as appressoria or penetrate plant tissues directly through hyphae40,43. Horizontal colonisation of roots by endophytes via mycelial fragments or spores is usually initiated by the formation of a superficial hyphal network and their penetration along the main root axis, followed by colonisation of the spaces between cortical and epidermal cells. Intracellular root expansion occurs via closely packed, thick, pseudoparenchymatous, melanized sclerotia or microsclerotia44. Endophytes are known to form Hartig-like net tissues or labyrinth tissues in root cells, and others form rounded chlamydospore-like cells in cortical cells45. To assess the potential for horizontal transmission under sterile conditions, eight fungal species (F. proliferatum, P. olsonii, P. expansum, T. hamatum, E. album, S. spinificis, C. candelabrum, N. gorlenkoana) previously isolated from G0 plants10 were reintroduced individually or as a mixture to axenic wheat seedlings representing 10 cultivars. The presence of the remaining four species (Engyodontium album, Sarocladium spinificis, Clonostachys candelabrum, and Nigrospora gorlenkoana) was not confirmed in axenic seedlings of any of the wheat varieties tested. Their inability to recolonise may reflect host incompatibility or limited ecological adaptation to wheat under controlled conditions. Interestingly, the three recolonising genera—Fusarium, Penicillium, and Trichoderma—were also recovered from G1 plants in the metabarcoding dataset, further confirming their capacity for both vertical and horizontal transmission. These species are commonly classified as Class 2 Non-clavicipitaceous (NC) endophytes, which rapidly colonise aboveground and belowground plant tissues, including shoots, roots, and rhizomes, and are transmitted both vertically and horizontally by seed coats and rhizomes46,47. Class 2 NC endophytes colonise plant tissues by forming infectious structures, such as appressoria, or by entering plant tissues directly via hyphae. Among the species for which the capacity for horizontal transmission was not documented in this study is E. album. It is believed that this is related to the lifestyle and nutrition of this fungus. Engyodontium album belongs to the Cordycipitaceae family and is a widespread pathogen that causes various types of dermatoses and respiratory diseases in humans and animals48. This species is considered a keratinophilic fungus. It is therefore assumed that its presence in G0 generation wheat plants was caused by the colonisation of grains originating from plants previously cultivated in an agricultural environment where feathers or animal fur could have been present. This species probably has no evolutionarily long-established interactions with wheat plants. Nigrospora gorlenkoana belongs to a genus of cosmopolitan fungi with a diverse host range. Species of this genus are pathogens, endophytes, and saprobionts. However, little is known about the lifestyle of the species N. gorlenkoana itself. Little is also known about S. spinificis, which has so far been isolated from the endosphere of the coastal grass Spinifex littoreus in Taiwan49. Clonostachys candelabrum is known as Sesquicillium candelabrum, Verticillium candelabrum or Clonostachys chuyangsinensis50. To date, this species has been isolated from forest soil, litter, leaves, and decaying wood, and there is limited knowledge about its interactions with plants and its existence as a plant endophyte [50]. The failure of these species to reintegrate under sterile conditions suggests they may represent transient or opportunistic colonisers, rather than core members of the wheat endosphere.
The results of this study, combined with previously published data on G0 plants10, provide a comprehensive picture of the endophytic fungal community associated with common wheat across generations. Previous work has shown that the structure of the wheat endomycobiome is largely shaped by plant organs and growing conditions, with roots showing the greatest diversity and leaves the least. Although the effects of wheat genotype and form were minimal, a consistent core of fungal taxa, such as Cladosporium, Penicillium, Sarocladium, and Fusarium, were detected across multiple cultivars and growing environments. The present G1-focused study built on these observations by showing that several of these core taxa are maintained across plant generations, suggesting intergenerational persistence via vertical transmission. Compared with G0, G1 plants exhibited higher fungal diversity and abundance, particularly in root tissues. Similarly, Sharon et al.16 observed a richer endophytic fungal community in the stems and seeds of progeny plants, with a much higher number of taxa than in the original seeds or the soil in which they were grown. These authors identified 78 taxa in the stems and 202 taxa in the new seeds, compared to only 35 taxa detected in the original seeds. The soil contained 83 taxa, of which 27 were found in the new plants. According to Sharon et al.16, rarer taxa could have appeared in the stems and new seeds because the plants were already grown in a greenhouse, where sterile conditions prevailed, and they were not exposed to the emergence of a single dominant taxon. In this study, plants from two consecutive generations were grown under strictly sterile conditions to minimise environmental contamination. Fungal species colonising seeds sourced from external agricultural environments may have penetrated G0 generation plants; notably, such fungi are often fast-growing and capable of rapidly establishing themselves in new habitats through competitive exclusion. Meanwhile, the growth of wheat plants and the production of new seeds under sterile conditions, with no access to environmental fungi, freed up endosphere space for other, slower-growing, and rarer taxa. Consequently, a greater degree of tissue colonisation by fungi and their greater biodiversity was noted in the G1 generation. According to Liao et al.40 the above issue may also be related to the methodology of sample preparation for high-throughput analyses. Surface sterilization technique is a significant step in preventing contamination by other microorganisms present on the host surface, including epiphytes, as well as tissue maceration and DNA isolation for metagenomic studies, especially for taxa that are unique or masked by rapidly growing species.
Conclusions
-
Both culture-dependent and culture-independent approaches revealed consistent patterns in the composition of endophytic fungal communities in G1 wheat plants. Fusarium proliferatum emerged as the dominant taxon across cultivars and tissues in both datasets. Several other genera—Penicillium, Trichoderma, Cladosporium, and Lecanicillium—were also detected by both methods, confirming their stable presence within the wheat endosphere. While metabarcoding highlighted broader taxonomic diversity and tissue-specific patterns, culture-based isolation provided complementary insights into viable and cultivable endophytes. Collectively, the results support the existence of a conserved core mycobiome in wheat plants, alongside tissue- and method-specific differences in fungal detectability and abundance.
-
Comparison of wheat G0 and G1 plants revealed the presence of a stable, transgenerational core of endophytic fungi, with 27 OTUs consistently detected in both generations. This core mycobiome, identified by combining metabarcoding and barcoding approaches, consisted mainly of the genera Fusarium, Trichoderma, Penicillium, Lecanicillium, and Cladosporium, with Sarocladium additionally present in the metabarcoding data. The persistence of these taxa suggests potential vertical transmission. Fungal abundance and diversity were significantly higher in G1, particularly in different tissues. The use of both culture-dependent and culture-independent methods provided a more comprehensive picture of the wheat endophytic mycobiome, encompassing both viable fungal isolates and broader taxonomic diversity, thus strengthening the evidence for a core but still dynamic fungal community structure across generations.
-
Experimental re-inoculation confirmed that several core endophytes, such as F. proliferatum, P. expansum, and T. hamatum, can recolonize axenic wheat tissues across multiple cultivars, supporting their active role in host association and persistence across generations.
-
Microscopy confirmed fungal presence both on root surfaces and within tissues, providing direct evidence of true endophytic colonization.
-
The observations highlight the combined influence of vertical transmission and environmental acquisition in shaping the wheat endomycobiome across generations
Material and methods
This study is a continuation of previously published work10 on the endophytic mycobiome of G0 wheat plants. All material, cultivation, sampling, and processing procedures for G1 plants followed the same protocols as for G0. Unless otherwise stated, the methodology used was identical to that previously described10 and is therefore not described in detail here. The present study includes a detailed analysis of the endophytic fungal community in the G1 generation and a comparative assessment of community structure, diversity, and transmission patterns between G0 and G1 plants.
Plant material
The plant material consisted of two forms of common wheat (Triticum aestivum L.): Arabella, Bombona, Kandela, Rospuda and Rusałka spring cultivars; Arkadia, Bamberka, Euforia, Legenda, and Ostroga winter cultivars. The wheat grains of Arkadia, Ostroga, Arabella, Bombona and Kandela cultivars were obtained kindly from Danko, Hodowla Roślin Sp. z o.o. in Chorynia. The wheat grains of Bamberka, Euforia, Rospuda and Rusałka cultivars were obtained kindly from Hodowla Roślin Strzelce Grupa IHAR. The grains of Legenda cultivars were obtained kindly from Poznańska Hodowla Roślin in Tulce. The outline of plant material preparation for the assessment of the horizontal and vertical transmission capacity of endophytic fungi is presented in Fig. 11.
Greenhouse cultivation of G1 progeny plants, sampling and processing
G1 progeny plants were cultivated under the same greenhouse conditions as described for G0 plants previously10. Plant material was collected at the same developmental stage, and the same sampling scheme was applied: root, stem, leaf, and kernel tissues were harvested, processed, and prepared for downstream analyses using identical protocols. Unless otherwise stated, all procedures mirrored those used for the G0 generation.
ITS2 metabarcoding—G1 progeny plants
To characterize the fungal endophytic community structure of wheat plants first-generation (G1), the same approach was used as for the parental plants (G0), using identical procedures for DNA isolation, library preparation, Illumina MiSeq sequencing of ITS2 amplicons10. As in the previous work, the second step of library preparation, sequencing of 40 samples and preliminary sequence processing were supported by the service company Genomed (Warsaw, Poland). High-throughput sequencing data were also processed using a standardised bioinformatics and statistics pipeline10. Maintaining methodological consistency was crucial due to the need to conduct further direct comparative analysis of the wheat endophytic fungal community in two generations of plants—parent G0 and progeny G1.
Taxonomic assignment of readings for two plant generations—G0 and G1
To compare the present results with the previously reported (BioProject PRJNA899455) G0 parental plants data, a joint taxonomic classification of reads from both experiments (G0 and G1) was performed using the QIIME platform, referencing the ITS UNITE v8.2 fungal database20. Adapter sequences were removed using Cutadapt21, followed by quality filtering to exclude low-quality reads (Q < 20) and sequences shorter than 30 nucleotides. Paired-end reads were merged with the SeqPrep algorithm to reconstruct full-length amplicons. Chimeric sequences were identified and removed using the USEARCH61 algorithm22, ensuring the reliability of downstream analyses. After quality control, sequences were clustered against the reference database using the UCLUST algorithm, and taxonomic assignments were made for representative sequences from each cluster using BLAST23.
Statistical and bioinformatic processing of two plants generations data
All statistical and bioinformatic analyses were performed in R (v.4.4.2), unless stated otherwise24. Alpha diversity was calculated and visualized using the plot_richness function from phyloseq (v.1.50.0) and compared between groups using pairwise Wilcoxon Rank Sum tests (pairwise.wilcox.test)25. Beta diversity analyses, including weighted UniFrac distances and principal coordinate analysis (PCoA), were conducted using phyloseq. The effect of tissues, wheat form, and cultivar on fungal community structure was assessed using PERMANOVA (adonis) from the vegan package (v. 2.6.10)26. Community structure was visualised with phyloseq and ggplot2 (v.3.5.1)27.
Preparation of axenic plant seedlings, cultivation in phytotrons and sampling
To study the horizontal transmission of endophytic fungi, axenic wheat seedlings were developed. Then, after treatment with selected fungal strains previously isolated from wheat plant tissues of the G0 generation, they were grown under sterile, controlled conditions in phytotrons. Axenic seedlings were obtained by isolating embryos from mature grains of wheat parent G0 plants28. Whole seeds were soaked overnight in sterilized distilled water at 4 °C. After incubation, seeds were rinsed in fresh sterilized distilled water, and the embryo was then carefully separated from each seed by cutting the scutellum, the connective tissue between the endosperm and the embryo. Isolated embryos were placed in sterile glass test tubes with a diameter of 2 cm and a height of 20 cm filled to 1/3 of the volume of Gamborg’s B5 Basal Medium (Merck KGaA, Darmstadt, Germany) where they grew for 7 days under optimized in vitro conditions. Seven-day-old axenic plants were then transplanted into twice-sterilized quartz sand filling the holes (3 × 3 × 12 cm) of seedling trays sterilized with 96% ethyl alcohol (Avanator Performance Materials Poland S.A.). For the next 7 days, until inoculation with endophytic fungi, the plants were grown under conditions of photoperiod (day/night) 12 h/12 h, temperature (day, night) 23⁰C/18⁰C and relative humidity 50%. The plants were watered manually. After inoculation, the plants were further grown in the growth chamber under the same conditions as above for another 7 days, but with physical separation (in separate boxes) for each treatment and control plants to avoid contamination. Twenty-one-day-old seedlings [5 plants/plant variety/treatment/replication], 7 days after inoculation, were harvested, divided into underground (roots) and aboveground (leaves and stems) parts, and stored at 4 °C or − 80 °C until further analysis. The experiment was arranged in completely randomized design with ten cultivars, 10 treatments (8 species of endophytes, 8 species of endophytic fungi spores mix, and sterile distilled water), three pot-replications and fifty plants per replications.
Preparation of fungi spores and axenic plant inoculation
Eight species/strains of endophytic fungi previously isolated from the internal tissues of wheat plants of the G0 generation10 were selected as follows: two strains originated from the seeds of the Euforia cultivar—Fusarium proliferatum E202 and Penicillium expansum E193; Clonostachys candelabrum E83 obtained from seeds and Trichoderma hamatum E81 obtained from the roots of Arabella cultivar; Penicillium olsonii E69 isolated from Kandela seeds; Engyodontium album E36 isolated from Bombona seeds; Nigrospora golenkoana E208 isolated from Euforia leaves; Sarocladium spinificis E11 acquired from roots of Rospuda cultivar. Pure cultures of 8 species of endophytic fungi were sterilely transferred from slants with SNA (Synthetic nutrient-poor agar)29 medium to sterile 8.5 cm diameter Petri dishes with PDA (Potato Dextrose Agar, Oxoid™ Thermo Fisher Scientific, Waltham, MA, USA) medium. Cultures were grown for 7 days at 25 ± 2 °C. The above-ground parts of the seedlings were sprayed with fungal spores at a concentration of approximately 5 × 105 mL−1. For this purpose, spore-forming fungi were purified in 10 mL of sterile distilled water and filtered through 2 layers of cheesecloth and then adjusted to the appropriate concentration using a hemocytometer. Control plants were sprayed with sterile water in the same volume as the spore mixture. The underground parts of the plants were inoculated by placing a 0.8 cm diameter agar disk cut with a corkscrew from a 7-day culture at a depth of about 0.5 cm near the root zone of each seedling. In the case of testing the effect of all selected fungi on the plant, disks cut from cultures of all tested fungal species were placed near the root ball. In the case of control plants, a clean agar disk not colonised by fungi was placed. The experiment was organised in a completely randomised design, with 10 cultivars (spring forms: Arabella, Bombona, Kandela, Rospuda, Rusałka; winter forms: Arkadia, Bamberka, Euforia, Legenda, Ostroga), 10 treatments (F. proliferatum, P. olsonii, P. expansum, N. gorlenkoana, S. spinificis, E. album, C. candelabrum, T. hamatum/ MIX of fungi/ control—sterile water), 3 plot replications, 5 plants per replication.
Microscopic detection of fungi in axenic plant tissues after inoculation
The microscopic observations were made based on the method proposed by Phillips and Hayman30 with slight modifications. Roots and leaves fragments were incubated for 24 h in 5% KOH (Avantor Performance Materials Poland S.A.). After this time, the material was washed three times with deionised water. The next plant samples were incubated for one hour with 50% lactic acid (Avantor Performance Materials Poland S.A., Gliwice, Poland) and treated with 0.1% trypan blue (Merck KGaA, Darmstadt, Germany, formerly Sigma, St. Louis, MO, USA) in a lactoglycerol solution [in equal parts of lactic acid: water: glycerol (Merck KGaA, Darmstadt, Germany, formerly Sigma, St. Louis, MO, USA)] for 2 h. Each step was performed at room temperature. A light microscope (Olympus CX-41–1 with UC-30 camera, Olympus, Japan) was used to conduct the observations.
Taxonomic assignment of isolated fungi
To access the taxonomic composition of cultivable fungal communities in G1 plant cultivars and organs and in treated axenic seedlings, the sequencing of selected phylogenetic markers was performed: internally transcribed spacer regions 1 and 2 (ITS1 and ITS2) of the rRNA gene cluster, a fragment of the translation-elongation factor 1-alpha (tef1) gene or partial beta tubulin 2 (tub2) gene and the second largest subunit of RNA-polymerase II (partial RPB2). Fungi were isolated/ re-isolated from the plant endosphere and identified at the genus/species level using the same approach as described in our previous study31.
Visualisation of DNA barcoding result data
Figures were created in Python (v. 3.13.3): venn diagrams were created utilizing the pyvenn module (https://github.com/tctianchi/pyvenn). Network graphs were created with Cytoscape.js (v 3.31.4)32 in a custom application written in Dash (v. 3.0.4) framework.
Statements
Experimental and field studies on crop plants, including collection and storage of plant material, were in accordance with the guidelines and legislation of the Institute of Plant Genetics of the Polish Academy of Sciences and Poland.
Breeding companies directly provided wheat seeds for experimental research, including field research, and granted permission to use the donated wheat seeds and the obtained plant progeny for further analyses.
Data availability
The datasets (DNA barcoding and metabarcoding) generated during the current study are available in the NCBI repository: MW713470—MW713524, MW720740-MW720743, OK309864-OK309871, OK258086, OK258087, OK180971-OK180973, MZ224007, OK258078-OK258085, BioProject PRJNA1292107.
Change history
05 December 2025
A Correction to this paper has been published: https://doi.org/10.1038/s41598-025-31507-y
References
Błaszczyk, L., Salamon, S. & Mikołajczak, K. Fungi inhabiting the wheat endosphere. Pathogens 10, 1288. https://doi.org/10.3390/pathogens10101288 (2021).
Hardoim, P. R. et al. The hidden world within plants: ecological and evolutionary considerations for defining functioning of microbial endophytes. Microbiol. Mol. Biol. Rev. 79, 293–320. https://doi.org/10.1128/MMBR.00050-14 (2015).
Le Cocq, K., Gurr, S. J., Hirsch, P. R. & Mauchline, T. H. Exploitation of endophytes for sustainable agricultural intensification. Mol. Plant Pathol. 18, 469–473. https://doi.org/10.1111/mpp.12483 (2017).
Balfourier, F., et.al International Wheat Genome Sequencing Consortium; BreedWheat Consortium; Paux E. Worldwide phylogeography and history of wheat genetic diversity. Sci. Adv. 5(5):eaav0536 (2019); https://doi.org/10.1126/sciadv.aav0536.
Compant, S., Samad, A., Faist, H. & Sessitsch, A. A review on the plant microbiome: Ecology, functions, and emerging trends in microbial application. J. Adv. Res. 19, 29–37. https://doi.org/10.1016/j.jare.2019.03.004 (2019).
Emmett, B. D., Buckley, D. H. & Drinkwater, L. E. Plant growth rate and nitrogen uptake shape rhizosphere bacterial community composition and activity in an agricultural field. New Phytol. 225, 960–973. https://doi.org/10.1111/nph.16171 (2020).
Zheng, Y. et al. The assembly of wheat-associated fungal community differs across growth stages. Appl. Microbiol. Biotechnol. 105, 7427–7438. https://doi.org/10.1007/s00253-021-11550-1 (2021).
Kavamura, V. N., Mendes, R., Bargaz, A. & Mauchline, T. H. Defining the wheat microbiome: Towards microbiome-facilitated crop production. Comput. Struct. Biotechnol. J. 19, 1200–1213. https://doi.org/10.1016/j.csbj.2021.01.045 (2021).
Latz, M. A. C. et al. Succession of the fungal endophytic microbiome of wheat is dependent on tissue-specific interactions between host genotype and environment. Sci. Total Environ. 759, 143804. https://doi.org/10.1016/j.scitotenv.2020.143804 (2021).
Salamon, S., Mikołajczak, K. & Błaszczyk, L. Constellation of the endophytic mycobiome in spring and winter wheat cultivars grown under various conditions. Sci. Rep. 13, 6064. https://doi.org/10.1038/s41598-023-33195-y (2023).
Kerdraon, L., Barret, M., Laval, V. & Suffert, F. Differential dynamics of microbial community networks help identify microorganisms interacting with residue borne pathogens: the case of Zymoseptoria tritici in wheat. Microbiome 7, 125. https://doi.org/10.1186/s40168-019-0736-0 (2019).
Schlatter, D. C., Yin, C., Hulbert, S. & Paulitz, T. Core rhizosphere microbiomes of dryland wheat are influenced by location and land use history. Appl. Environ. Microbiol. 86, e02135-e2219. https://doi.org/10.1128/AEM.02135-19 (2020).
Schlatter, D. C., Hansen, J. C., Schillinger, W. F., Sullivan, T. S. & Paulitz, T. C. Common and unique rhizosphere microbial communities of wheat and canola in a semiarid Mediterranean environment. Appl. Soil Ecol. 144, 170–181. https://doi.org/10.1016/j.apsoil.2019.07.010 (2019).
Hartman, K. et al. Cropping practices manipulate abundance patterns of root and soil microbiome members paving the way to smart farming. Microbiome 6, 14. https://doi.org/10.1186/s40168-017-0389-9 (2018).
Gdanetz, K. & Trail, F. The wheat microbiome under four management strategies, and potential for endophytes in disease protection. Phytobiomes J. 1, 158–168. https://doi.org/10.1094/PBIOMES-05-17-0023-R (2017).
Sharon, O. et al. Transmission mode and assembly of seed fungal endophyte communities in wheat and wheat wild relatives. Phytobiomes J. 7, 113–124. https://doi.org/10.1094/PBIOMES-11-22-0084-R (2023).
Shearin, Z. R. C. et al. Fungal endophytes from seeds of invasive, non-native Phragmites australis and their potential role in germination and seedling growth. Plant Soil https://doi.org/10.1007/s11104-017-3241-x (2017).
Vujanovic, V. & Germida, J. J. Seed endosymbiosis: A vital relationship in providing prenatal care to plants. Can. J. Plant Sci. 97, 972–981. https://doi.org/10.1139/cjps-2016-0261 (2017).
Wiewióra, B., Żurek, G. & Pańka, D. Is the vertical transmission of Neotyphodium lolii in perennial ryegrass the only possible way to the spread of endophytes?. PLoS ONE 10, e0117231. https://doi.org/10.1371/journal.pone.0117231 (2015).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336. https://doi.org/10.1038/nmeth.f.303 (2010).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet Journal 17, 10–12. https://doi.org/10.14806/ej.17.1.200 (2011).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461. https://doi.org/10.1093/bioinformatics/btq461 (2010).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410. https://doi.org/10.1016/S0022-2836(05)80360-2 (1990).
R Core Team. R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing (2024); https://www.R-project.org.
McMurdie, P. J. & Holmes, S. phyloseq: An R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 8, e61217. https://doi.org/10.1371/journal.pone.0061217 (2013).
Oksanen, J. et al. vegan: Community Ecology Package. R package version 2.6–10 (2025); https://CRAN.R-project.org/package=vegan.
Robinson RJ, Fraaije BA, Clark IM, Jackson RW, Hirsch PR, Mauchline TH. (2016). Wheat seed embryo excision enables the creation of axenic seedlings and Koch’s postulate testing of putative bacterial endophytes. Sci. Rep. 6:25581 (2016); https://doi.org/10.1038/srep2558.
Nirenberg, H. I. Untersuchungen iiber die morphologische und biologische Differenziemng in der Fusarium- SektionLiseola. Mitt. Biol. Bundesanst. LandForstwirtsch. Berlin-Dahlem 169, 1–117 (1976).
Phillips, J. M. & Hayman, D. A. Improved procedures for clearing roots and staining parasitic and vesicular-arbuscular mycorrhizal fungi for rapid assessment of infection. Trans. Br. Mycol. Soc. 55, 158–161. https://doi.org/10.1016/S0007-1536(70)80110-3 (1970).
Salamon, S. et al. Changes in root-associated fungal communities in Triticum aestivum ssp spelta and Triticum aestivum ssp vulgare under drought stress and in various soil processing. PLoS One 15, e0240037 (2020); https://doi.org/10.1371/journal.pone.0240037.
Franz, M., Kucera, M. & Mondal, A. cytoscape/cytoscape.js: v3.31.4 (v3.31.4). Zenodo (2025); https://doi.org/10.5281/zenodo.15312292.
Nelson, E. B. The seed microbiome: Origins, interactions, and impacts. Plant Soil 422, 7–34. https://doi.org/10.1007/s11104-017-3289-7 (2018).
Samreen, T. et al. Seed associated bacterial and fungal endophytes: Diversity, life cycle, transmission, and application potential. Appl. Soil Ecol. https://doi.org/10.1016/j.apsoil.2021.104191 (2021).
Rojas, E. C. et al. Selection of fungal endophytes with biocontrol potential against Fusarium head blight in wheat. Biol. Control. 144, 104222. https://doi.org/10.1016/j.biocontrol.2020.104222 (2020).
Lenc, L., Kwaśna, H., Sadowski, C. & Grabowski, A. Microbiota in wheat roots, rhizosphere and soil in crops grown in organic and other production systems. J. Phytopathol. 163, 245–263. https://doi.org/10.1111/jph.12313 (2015).
Larran, S., Perelló, A., Simón, M. R. & Moreno, V. The endophytic fungi from wheat (Triticum aestivum L.). World J. Microbiol. Biotechnol. 23, 565–572. https://doi.org/10.1007/s11274-006-9266-6 (2007).
Vujanovic, V., Mavragani, D. & Hamel, C. Fungal communities associated with durum wheat production system: A characterization by growth stage, plant organ and preceding crop. Crop Prot. 37, 26–34. https://doi.org/10.1016/j.cropro.2012.02.006 (2012).
Nicolaisen, M., Justesen, A. F., Knorr, K., Wang, J. & Pinnschmidt, H. O. Fungal communities in wheat grain show significant co-existence patterns among species. Fungal Ecol. 11, 145–153. https://doi.org/10.1016/j.funeco.2014.06.002 (2014).
Liao, C. et al. Challenges and update on fungal endophytes: Classification, definition, diversity, ecology, evolution and functions. Fungal Divers. 131, 301–367. https://doi.org/10.1007/s13225-025-00550-5 (2025).
Hill, R., Buggs, R. J., Vu, D. T. & Gaya, E. Lifestyle transitions in fusarioid fungi are frequent and lack clear genomic signatures. Mol. Biol. Evol. 39, msac085 (2022).
Hu, Y. et al. Endophytic fungi: Tracing the evolutionary roots and exploring the diversity of plant-fungal symbioses. Curr. Res. Environ. Appl. Mycol. 14, 1–48 (2024).
Rodriguez, R. J., White, J. F. Jr., Arnold, A. E. & Redman, A. R. Fungal endophytes: Diversity and functional roles. New Phytol. 182, 314–330. https://doi.org/10.1111/j.1469-8137.2009.02773.x (2009).
Jumpponen, A. & Trappe, J. M. Dark septate endophytes: A review of facultative biotrophic root colonizing fungi. New Phytol. 140, 295–310. https://doi.org/10.1046/j.1469-8137.1998.00265.x (1998).
Odell, T. E., Massicotte, H. B. & Trappe, J. M. Root colonization of Lupinus latifolius AGARDH and Pinus contorta DOUGL. by Phialocephala fortinii WANG & WILCOX. New Phytol. 124, 93–100. https://doi.org/10.1111/j.1469-8137.1993.tb03800.x (1993).
Rodriguez, R. & Redman, R. More than 400 million years of evolution and some plants still can’t make it on their own: Plant stress tolerance via fungal symbiosis. J. Exp. Bot. 59, 1109–1114. https://doi.org/10.1093/jxb/erm342 (2008).
Chitnis, V. R. et al. Fungal endophyte-mediated crop improvement: The way ahead. Front. Plant. Sci. 11, 561007. https://doi.org/10.3389/fpls.2020.561007 (2020).
Yuan, X. L. et al. The complete mitochondrial genome of Engyodontium album and comparative analyses with Ascomycota mitogenomes. Genet. Mol. Biol. 40, 844–854. https://doi.org/10.1590/1678-4685-GMB-2016-0308 (2017).
Yeh, Y. H. & Kirschner, R. Sarocladium spinificis, a new endophytic species from the coastal grass Spinifex littoreus in Taiwan. Bot. Stud. 55, 25. https://doi.org/10.1186/1999-3110-55-25 (2014).
Zhao, L. et al. Revising Clonostachys and allied genera in Bionectriaceae. Stud. Mycol. 105, 205–266. https://doi.org/10.3114/sim.2023.105.03 (2023).
Wang, Y. et al. Phylogeny and systematics of the genus Clonostachys. Front. Microbiol. 14, 1117753. https://doi.org/10.3389/fmicb.2023.1117753 (2023).
Acknowledgements
We wish to thank the graduate student, Agata Górka, for their technical support during the microscopic analysis.
Funding
This study was supported by the National Science Center in Poland Grant No 2017/27/B/NZ9/01591.
Author information
Authors and Affiliations
Contributions
L.B., S.S. and K.M. wrote the main text of the manuscript; S.S. was responsible for high-throughput and standard sequencing analyses, performed data processing, prepared Figs. 1, 2, 5 and Fig. S1, S2, S3, S4. K.M. was responsible for the isolation and molecular identification of endophytic fungi, processed the Sanger sequencing data, deposited the nucleotide sequences of phylogenetic markers in the NCBI database, prepared Supplement material (table). P.B processed and visualized DNA barcoding data, deposited the ITS metabarcoding data, completed the text of the manuscript, prepared Figs. 3, 4 and 6, 7, 8 and 9. A.B. performed the microscopic analyses, micrographs and prepared Fig. 10, completed the text of the manuscript. L.B., K.M. and A.B. were responsible for the phytotron experiment using axenic seedlings. L.B. is the Principal Investigator of the project, led the implementation of the research and conducted substantive supervision, prepared Fig. 11. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this Article was revised: In the original version of this Article, Piotr Banachewicz was erroneously listed as a corresponding author. The sole corresponding author is Lidia Błaszczyk. All correspondence and requests for materials should be addressed to lbla@igr.poznan.pl.
Supplementary Information
Below is the link to the electronic supplementary material.
41598_2025_26280_MOESM1_ESM.docx
Supplementary Material 1: Table S1. List of endophytic fungi obtained from G1 generation wheat plants identified by DNA barcoding along with NCB GenBank accession numbers.
41598_2025_26280_MOESM2_ESM.jpg
Supplementary Material 2: Figure S1. Relative abundance of endophytic fungi in wheat across variety on the Phyla (a), Order (b) and Family (c) level.
41598_2025_26280_MOESM3_ESM.jpg
Supplementary Material 3: Figure S2. Relative abundance of endophytic fungi in wheat across tissue on the Phyla (a), Order (b) and Family (c) level.
41598_2025_26280_MOESM4_ESM.jpg
Supplementary Material 4: Figure S3. Fungal community diversity across wheat variety (a, c) and type (b, d). (a) Alpha diversity metrics (Observed OTUs, Chao1, and Shannon index) compared across different variety (Arabella, Arkadia, Bamberka, Bombona, Euforia, Kandela, Legenda, Ostroga, Rospuda, Rusałka); (b) Alpha diversity metrics (Observed OTUs, Chao1, and Shannon index) compared across different wheat type (spring, winter); (c, d) Principal Coordinate Analysis (PCoA) based on Weighted UniFrac distances.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Salamon, S., Mikołajczak, K., Basińska-Barczak, A. et al. Intergenerational and horizontal transmission of wheat endophytic fungi. Sci Rep 15, 42275 (2025). https://doi.org/10.1038/s41598-025-26280-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-26280-x