Abstract
Nesolagus netscheri, a Sumatran striped rabbit, is one of the rarest rabbits in the Leporidae family, and its genetic information is still limited. This study provides the first mitochondrial genome and molecular systematic characterization of the Sumatran striped rabbit, Nesolagus netscheri, Indonesia’s rarest rabbit. It consists of a circular double-stranded DNA of 16,709 bp. It showed that the mitochondrial genome structure of N. netscheri is similar to that of N. timminsi. The mitochondrial genome of N. netscheri contained 22 transfer RNA (tRNA) genes, and all tRNA except for trnS1 showed a characteristic cloverleaf secondary structure. Evidence was found that the atp8 gene of N. netscheri is under positive selection pressure. The phylogenetic analysis shows Leporidae was monophyletic, with Nesolagus at the basal. The study indicates a split between N. netscheri and N. timminsi in the Late Pleistocene around 0.43 million years ago. This research is a fundamental reference for the conservation of the rarest lagomorph species and provides important information for future evolutionary studies in the Leporidae family.
Similar content being viewed by others
Introduction
Nesolagus is a rabbit genus that comprises three distinct species: the Annamite striped rabbit (Nesolagus timminsi), the Sumatran rabbit (Nesolagus netscheri), and the extinct N. sinensis1. The N. netscheri is a lagomorph species found endemically in Sumatra, Indonesia2,3. Because insufficient information is available to estimate population size, range, and density, the International Union for Conservation of Nature (IUCN) has classified this species as Data Deficient4. The N. netscheri has been a legally protected species in Indonesia since then, and it continues to be protected under the recently updated Indonesian law. It has been classified as the rarest lagomorph owing to the small number of museum specimens and the rarity of historical observations5.
The relationship between N. netscheri and N. timminsi adds complexity to the genus’s evolutionary context because geographically they are separated and both have significant threats from habitat loss and fragmentation3,6. Despite its historical recognition and occasional sightings, there remain substantial gaps in understanding the evolutionary relationships of N. netscheri within the Nesolagus genus and the broader family Leporidae, indicating a need for molecular phylogenetic studies to elucidate the evolutionary history and genetic diversity of N. netscheri. A comprehensive understanding of these aspects is crucial, as it would provide insights into the evolutionary processes that have shaped the species and inform conservation strategies.
Mitochondrial (mtDNA) genomes provide a wealth of genetic markers that are crucial for conservation efforts, particularly for species like the Sumatran striped rabbit (Nesolagus netscheri). The complete sequencing of mtDNA can uncover specific genetic markers, such as cytochrome c oxidase subunit I (COI) and cytochrome b, which is widely recognized as a universal DNA barcode for animal species identification7,8,9. This marker is particularly valuable for assessing genetic diversity and population structure, allowing conservationists to monitor the health of populations and identify distinct genetic lineages that may require targeted conservation efforts10,11,12,13,14. For instance, in the case of the Sumatran striped rabbit, identifying unique mtDNA haplotypes could inform habitat management and restoration strategies, ensuring that genetic diversity is preserved in fragmented landscapes. Additionally, mtDNA can reveal information about historical population dynamics and gene flow, critical for understanding how habitat fragmentation affects species like the Sumatran striped rabbit. By analyzing variations in mtDNA, researchers can identify genetic bottlenecks and assess the impacts of habitat loss on gene flow between populations15,16. This information is particularly relevant in the context of Sumatra’s rapidly changing landscape, where deforestation and agricultural expansion have led to isolated populations that may be at risk of inbreeding17,18,19,20.
The use of mtDNA markers can thus help prioritize conservation actions, such as creating wildlife corridors to enhance connectivity between fragmented habitats21. Moreover, the high mutation rate and maternal inheritance of mtDNA make it an ideal candidate for studying evolutionary relationships and phylogenetic analyses within Leporidae9,10,22,23. For example, the identification of specific mitochondrial markers can aid in distinguishing between closely related species or subspecies, providing insights into their evolutionary history and informing conservation strategies that consider the unique genetic makeup of each population9.
This research is the first successful characterization of the mitochondrial genome of N. netscheri, which has not been previously published or available in any database. This study aims to characterize the mitogenome of N. netscheri and reconstruct the evolutionary tree in the family Leporidae using a novel mitochondrial genome sequence. This study provides valuable information, including molecular markers, which can be utilized to address conservation needs, considering the limited extent of studies on rabbits. These findings will provide valuable information for phylogenetic, evolutionary, and population genetic studies. Additionally, findings can potentially be important to conserving the rarest rabbit, N. netscheri.
Results and discussion
Mitogenome composition and organization
The complete assembly of N. netscheri’s circular mitochondrial genome was generated by the Oxford Nanopore Technolgy (ONT) long reads. The length of this final assembly was 16,709 bp, with a coverage of 5021×. It is 3 bp smaller than the other N. timminsi mitogenome (NC_063946.1). The complete mitochondrial genome sequence of Sumatran striped rabbit Nesolagus netscheri has been deposited in GenBank under the accession number PQ047138. This genome size is within the range of available Leporidae mitochondrial genome (Table 1.). As in other mammals24, the mitogenome of N. netscheri encodes 37 classical mitochondrial genes (13 protein-coding genes [PCGs], 22 tRNAs, 2 rRNAs, and 1 D-loop) (Fig. 1). The complete mitogenome had a significant A + T bias in its nucleotide composition (30.05% A, 31.8% T, 13.3% C, and 24.4% G). In this mitogenome, the H strand transcribed twelve PCGs, two rRNAs, and thirteen tRNAs, while the L strand transcribed the remaining ten genes (Table 2). This transcription pattern is the same as that of the genus N. timminsi. N. timminsi and N. netscheri belong to the same genus, and their gene orders are similar (Supplementary File 1).
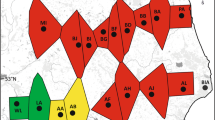
(a) Circular representation of the mitochondrial genome of Nesolagus netscheri. The colored blocks indicate different gene types, with genes encoded on the light and heavy strands in counterclockwise and clockwise orientations, respectively. The black inner circle represents the GC content of the mitogenome, while the pink and red regions indicate GC skew. The rabbit illustration was adapted from https://scentsindonesia.com; (b) Geographic distribution of N. netscheri (orange regions) within its habitat. The map was created using QGIS version 3.40.1 Bratislava (https://www.qgis.org).
Protein-coding genes
Standard initiation codons (ATN) were used for the initiation of all PCGs, and conventional stop codons (TAG or TAA) were used for the termination of 9 out of 13 PCGs. The genes ND3, ND4, and COX3 all terminated with incomplete stop codons (either T or TA; Table 2), while the CYTB gene had an alternate putative stop codon (AGG). The start codons found in the PCGs are consistent with the standard vertebrate mitochondrial start codons, except for ATT. There have been comparable reports of this terminal codon among various Leporidae28,31,33. The same phenomenon has been observed with the regular and common stop codons TAG and TAA in mammals24. AGG is an uncommon stop codon; however, it has been identified in other Leporidae species28,31,33 and other vertebrates in previous study34. The COX3, NAD3, and NAD4 genes in vertebrate mitochondrial genomes, including Leporidae28,31,33, frequently exhibit reduced stop codons. These reduced stop codons have been reported to be addressed through post-transcriptional polyadenylation35.
Relative synonymous codon usage values for N. netscheri are presented in Fig. 2. The PCGs contained 3,510 codons. The most prevalent codons in these mitogenomes encode Leu, Ile, Gly, and Thr. In contrast, the least frequently observed were those that encoded Trp. (Fig. 2).
Transfer and ribosomal RNA genes
The mitochondrial genome of N. netscheri had a total of 22 transfer RNA (tRNA) genes. The length of the tRNA genes varied from 54 bp (trnG) to 72 bp (trnN). A typical ‘cloverleaf’ secondary structure was inferred in all tRNA genes, with the exception of trnS1 (see Fig. 3). This is a commonly observed characteristic in genes for tRNA in mammals36,37 and vertebrates38. The lack of an arm in trnS1 may potentially have a functional purpose in the process of structural compensation with other structures39. The 16 S RNA (rrnL) is positioned between tnrL2 and trnV and has a length of 1,580 bp, whereas the 12 S RNA (rrnS) is placed between trnV and trnF and has a length of 950 bp. The positions of the control region and rRNA genes (rrnS and rrnL) in the N. netscheri mitochondrial genome are similar to those already identified for other leporid mitogenomes28,31,33.
The Ka/Ks ratio is an important parameter for quantifying selection pressure40. To assess the effect on mitochondrial PCGs, we investigated the Ka/Ks ratios for N. netscheri and two leporid species, Lepus europaeus and Nesolagus timminsi, which were used as reference species. The 13 PCGs showed varying Ka/Ks ratios, which suggests that the genes are exposed to a range of functional constraints (Fig. 4). The data indicates that all the PCGs are undergoing evolutionary changes due to purifying selection (< 1), except for atp8. The lowest calculated value of Ka/Ks, which indicates selected pressure, was found for the nd4l gene in the Genus Nesolagus (Ka/Ks = 0.025). This suggests that nd4l faces the highest level of selective pressure and evolves at the slowest rate among the genes analysed. Purifying selection in mammalian mitochondrial DNA (mtDNA) is evidenced by studies that demonstrate a consistent pattern of selective pressure against deleterious mutations. For example, Stewart et al. highlighted that strong purifying selection operates during the transmission of mtDNA across generations, effectively maintaining the integrity of mitochondrial genomes by favoring the survival of less mutated variants41. Similarly, Tsai et al. reported that reducing the dosage of mitochondrial DNA polymerase enhances the elimination of defective mitochondrial genomes, indicating that purifying selection plays a critical role in maintaining mtDNA quality during oogenesis42. Furthermore, Ennis et al. found that the complete mitochondrial genome of Baird’s tapir exhibited patterns of purifying selection in its protein-coding genes, reinforcing the notion that mtDNA is subject to evolutionary constraints across various mammalian species43. These studies underscore the importance of purifying selection in preserving the functional integrity of mitochondrial genomes in mammals.
The atp8 gene in the genus Nesolagus exhibits a Ka/Ks ratio of 1.22, indicating positive selection and a rapid rate of evolution. This suggests its role in adaptation to ecological challenges. The atp8 gene exhibits the same pattern of high Ka/Ks ratios in various mammals, including Rattus44, Chiroptera45, caviomorph rodents46, as well as between Puma concolor47. It has been reported in other studies that positive selection also affects mitochondrial genes, particularly in animals that are adapted to harsh environments23,48,49,50. In extreme environments, positive selection in ATP synthase genes enhances metabolic efficiency and energy production, as seen in species inhabiting high-altitude or low-oxygen conditions51,52. For tropical mammals, atp8 has been identified as critical for adaptation to the metabolic demands of dense rainforest ecosystems, which are characterized by high biodiversity and fluctuating conditions53,54. Studies on bats, cetaceans, and tropical chickens highlight the evolutionary significance of atp8 in overcoming environmental stressors specific to their habitats, such as low oxygen, high energy demands, or thermal adaptation55,56. In the context of Nesolagus, which inhabits the tropical forests of Sumatra, the positive selection of atp8 may enhance its capacity to adapt to the unique challenges of its environment, including dense vegetation, resource competition, and fluctuating climatic conditions46,52,57,58,59,60. These findings collectively underscore the ecological and evolutionary importance of the atp8 gene in facilitating adaptation to harsh environments.
Phylogenetic of N. netscheri
Mitochondrial DNA is a commonly employed genetic marker for understanding molecular systematic in animals61. To better comprehend the evolutionary relationships within the Leporidae family, we collected concatenated nucleotide sequences of 13 PCGs and two ribosomal RNA (rRNA) genes from 13 leporids. The topologies of the phylogenetic trees obtained from the Bayesian inference (BI) and maximum likelihood (ML) analyses were nearly identical, as illustrated in Fig. 5. Both trees generated from the two models were supported by robust statistics values.
The phylogenetic analysis confirmed the monophyly of the family Leporidae (posterior probability = 0.99; bootstrap = 100), with strong support for the Nesolagus genus (1; 100), the genus Lepus (1; 100), and the group comprising Oryctolagus cuniculus, Brachylagus idahoensis, and Sylvilagus sp. (1; 91), forming a single single and well-supported clade. In the Leporidae family, Nesolagus is positioned basally as a sister group to a clade consisting of Oryctolagus cuniculus, Brachylagus idahoensis, Sylvilagus sp., Romerolagus diazi, and Lepus spp. The topology in our study is consistent with previous research that examined cranial morphology in Leporidae62,63 and molecular data64,65. The data analysis shows a split between N. netscheri and N. timminsi during the Late Pleistocene (95%HPD, 0.43 mya ± 0.19). This study’s divergence time matches the study of Nesolagus sinensis fossils from the Early Pleistocene period1. These fossils have been reported to be more primitive and directly related to the currently existing Nesolagus. In further studies, it is necessary to include the mitogenome of additional genera such as Pentalagus, Bunolagus, Poelagus, Caprolagus, and Pronolagus into phylogenetic analysis to perform a more comprehensive investigation of Leporidae phylogeny.
The phylogenetic tree of Leporidae is based on 13 PCGs (11,396 bp), which is estimated using the Bayesian relaxed-molecular clock method. Posterior probability values from Bayesian inference and percentages of bootstrap from maximum likelihood analyses are listed on each node, respectively. Purple bars were used to show the 95% HPD for each node, with each genus represented by a different color.
Conclusions
Mitochondrial genomes are increasingly utilized as valuable markers in phylogenetic, population genetic, and evolutionary studies. This paper reports and annotates the first complete mitochondrial genome of the elusive Sumatran striped rabbit (Nesolagus netscheri). There was 16,709 bp of circular double-stranded DNA. The mitochondrial genome’s organization and gene arrangement is similar to the one described for its sister taxon, N. timminsi. According to our Bayesian-based phylogenetic tree, the genus Nesolagus is at the basal of the Leporidae tree, and N. netscheri diverged in the Late Pleistocene (0.43 Mya). This research provides vital genetic information essential for developing effective conservation strategies for this rabbit. In the context of Sumatra’s unique and threatened forests, the mitochondrial genome data is critical for assessing genetic diversity, identifying evolutionary adaptations, and informing strategies such as inbreeding or bottleneck detection and translocation. This new information can be used as a reference mitogenome for future mitochondrial diversity studies to diagnose the species’ population status. It is a crucial point for adaptation to environmental change because of its limited habitat and distribution. Other Leporidae mitochondrial genomes are still required to provide a more thorough evolutionary history, and population genetic research and monitoring are needed to support effective conservation management—particularly in the case of Sumatran striped rabbits.
Materials and methods
Sample collection and DNA extraction
A tissue sample from the leg muscular of Nesolagus netscheri was taken from a specimen from Setiawan et al. research2. The tissue sample was collected immediately after the animal’s death to reduce DNA degradation caused by post-mortem damage. The sample was promptly frozen to ensure its preservation. Additionally, sterilized tools and gloves were used during sample handling to minimize the risk of exogenous contamination. Permission to access genetic resources for the Sumatran rabbit sample has been obtained from the Ministry of Environment and Forestry (no. SK.154/KSDAE/SET.3/KSA.2/8/2023). No live animal was involved in this research. This individual was reportedly caught in a forest within the Dempo mountain, South Sumatra province, Indonesia. The tissue sample was extracted using the Qiagen Blood and Tissue Kit. Then the DNA samples were quantified using a Qubit Fluorometer.
Mitogenome sequencing and assembly
Following Genome DNA Extraction, genome DNA was sequenced using PromethION Flow Cell (Oxford Nanopore Technologies) and Ligation Sequencing Kit V14 SQK-LSK114 to prepare the genomic DNA library. A total of 1000 ng in 48µL of gDNA sample was used to produce the library. The first step is the DNA repair and end-prep step using NEBNext FFPE DNA Repair Mix and Ultra II End-prep Enzyme Mix. In the next step, the sample is given adapter ligation using Ligation Adapter reagent, Ligation Buffer, and NEBNext Quick Ligation Module. The genomic DNA library was sequenced on a PromethIon 24 device with super accuracy (SUP) base-calling mode and running for approximately 72 h. A total number of 14,512,100 reads were generated and made available in FASTQ. All these reads were employed to conduct de novo assembly of the mitogenome of N. netscheri.
Mitogenome annotation and analysis
The Guppy base caller ONT v.3.2.4 was utilized to determine the base sequence from the raw data accurately. Only sequences with a Phred score greater than 13, indicating good quality, were chosen for the de novo mitogenome assembly using Oxford Nanopore Technology66,67. The assembly was performed utilizing the Flye program v.2.568 in the mitogenome assembly mode and functioned as baits to identify all potential mitochondrial sequences using Minimap269. A subsequent set of ONT assemblies was generated and subsequently refined over four iterations applying the same dataset using racon70 and medaka (https://github.com/nanoporetech/medaka).
The mitogenome sequences that were assembled were annotated using MITOS271. The tRNA scan-SE 1.21 software was employed to detect tRNA genes, using the genetic code specific to vertebrate mitochondria72. A visual illustration of the mitogenome of N. netscheri was drawn using the Proksee73, an online tool for visualizing mitochondrial data. The Ka_Ks calculator was utilized to quantify the ratio of nonsynonymous to synonymous substitution rates (Ka/Ks) for all 13 protein genes in three leporids, using MEGA X74. Bayesian and Maximum Likelihood phylogenetic trees were constructed using the 13 concatenated protein-coding genes (PCGs) of Leporids mitogenomes, as shown in Table 1. The best evolutionary model for analysis has been identified using jModeltest 275 to be the GTR model of sequence evolution, which includes a discrete gamma distribution and a proportion of invariable sites to account for variation in rates among sites (GTR+ Г + I). The Maximum-Likelihood (ML) phylogenetic was inferred using IQ-TREE (version 2.1) via the IQ-TREE web server with76 model selection and ultrafast bootstrap approximation (1000 iterations) based on the alignment of 13 PCGs. Beast 2.7.6 77 was used to estimate divergence times based on PCGs. The time tree was constructed using two calibrated points: the divergence of Lepus americanus and Lepus europaeus, which occurred 8.6 mya78, and the divergence of L. americanus and Lepus tolai, which was estimated based on fossil evidence to have occurred 0.78 mya79. The divergence time analysis employed an uncorrelated lognormal relaxed molecular clock and Yule process, with a total of 50 million generations. The convergence of the analysis was corroborated using Tracer80. All parameters show an ESS greater than 200, confirming sufficient sampling and reliability. The software TreeAnnotator was used to discard the initial 25% of trees and calculate the nodes’ ages along with their 95% credible intervals. The produced tree was viewed using FigTree Version 1.4.4.
Data availability
The new complete mitochondrial genome sequence of the Sumatran striped rabbit Nesolagus netscheri has been deposited in the GenBank of NCBI (https://www.ncbi.nlm.nih.gov/) database under the accession number PQ047138.
References
Jin, C., Tomida, Y., Wang, Y. & Zhang, Y. First discovery of fossil Nesolagus (Leporidae, Lagomorpha) from Southeast Asia. Sci. China Earth Sci. 53, 1134–1140 (2010).
Setiawan, A. et al. First description of an immature Sumatran striped rabbit (Nesolagus netscheri), with special reference to the wildlife trade in South Sumatra. Mammalia 84, 250–252 (2020).
Smith, A. T., Johnston, C. H., Alves, P. C. & Hackländer, K. Lagomorphs: Pikas, Rabbits, and Hares of the World (JHU, 2018).
McCarthy, J., Holden, J., Martyr, D. & McCarthy, K. Nesolagus netscheri. (2019).
Chapman, J. A. & Flux, J. E. Rabbits, hares and Pikas: Status survey and conservation action plan (IUCN, 1990).
Nguyen, A. et al. New records and southern range extension of the Annamite striped rabbit Nesolagus timminsi in Vietnam. Mammalia 85, 417–421 (2021).
Dawnay, N., Ogden, R., McEwing, R., Carvalho, G. R. & Thorpe, R. S. Validation of the barcoding gene COI for use in forensic genetic species identification. Forensic Sci. Int. 173, 1–6 (2007).
Petit-Marty, N., Vázquez‐Luis, M. & Hendriks, I. E. Use of the nucleotide diversity in COI mitochondrial gene as an early diagnostic of conservation status of animal species. Conserv. Lett. 14, e12756 (2021).
Álvarez-Castañeda, S. T. & Lorenzo, C. Genetic evidence supports Sylvilagus mansuetus (Lagomorpha: Leporidae) as a subspecies of S. Bachmani. Zootaxa 4196, 289–295 (2016).
Mora, J. M. et al. Systematics, distribution, and conservation status of Dice’s cottontail, Sylvilagus dicei Harris, 1932 (Mammalia, Lagomorpha, Leporidae), in Central America. Syst. Biodivers. 19, 74–88 (2020).
Hsieh, H. M. et al. Cytochrome B gene for species identification of the conservation animals. Forensic Sci. Int. 122, 7–18 (2001).
Linacre, A. & Lee, J. C. I. Species determination: the role and use of the cytochrome b gene. Forensic DNA Typing Protocols 287–296 (2016).
Arif, I. A. & Khan, H. A. Molecular markers for biodiversity analysis of wildlife animals: A brief review. Anim. Biodivers. Conserv. 32, 9–17 (2009).
Hohenlohe, P. A. & Funk, W. C. Rajora, O. P. Population genomics for wildlife conservation and management. Mol. Ecol. 30, 62–82 (2021).
Knaus, B. J., Cronn, R., Liston, A., Pilgrim, K. L. & Schwartz, M. K. Mitochondrial genome sequences illuminate maternal lineages of conservation concern in a rare carnivore. BMC Ecol. 11 (2011).
Van Der Valk, T. et al. Significant loss of mitochondrial diversity within the last century due to extinction of peripheral populations in eastern gorillas. Sci. Rep. 8, 6551 (2018).
Lubis, M. I. Planning for megafauna recovery in the tropical rainforests of Sumatra. Front. Ecol. Evol. 11 (2023).
Gaveau, D. L., Wandono, H. & Setiabudi, F. Three decades of deforestation in southwest Sumatra: Have protected areas halted forest loss and logging, and promoted re-growth? Biol. Conserv. 134, 495–504 (2007).
Margono, B. A. et al. Mapping and monitoring deforestation and forest degradation in Sumatra (Indonesia) using landsat time series data sets from 1990 to 2010. Environ. Res. Lett. 7, 034010 (2012).
Smith, O., Wang, J. & Carbone, C. Evaluating the effect of forest loss and agricultural expansion on Sumatran tigers from scat surveys. Biol. Conserv. 221, 270–278 (2018).
Sloan, S., Alamgir, M., Campbell, M. J., Setyawati, T. & Laurance, W. F. Development corridors and remnant-forest conservation in Sumatra, Indonesia. Trop. Conserv. Sci. 12, 1940082919889509 (2019).
Ding, L. et al. Complete mitochondrial DNA sequence of Lepus sinensis (Leporidae: Lepus). Mitochondrial DNA Part. A. 27, 1711–1712 (2016).
Shan, W., Tursun, M., Zhou, S., Zhang, Y. & Dai, H. Complete mitochondrial genome sequence of Lepus yarkandensis Günther, 1875 (Lagomorpha, Leporidae): Characterization and phylogenetic analysis. ZooKeys 1012, 135 (2021).
Rackham, O. & Filipovska, A. Organization and expression of the mammalian mitochondrial genome. Nat. Rev. Genet. 23, 606–623 (2022).
Huang, Y. L. et al. The complete mitochondrial genome sequence of Yarkand hare (Lepus yarkandensis). Mitochondrial DNA Part. B. 4, 3727–3728 (2019).
Saha, A. et al. The first complete mitochondrial genome data of the pygmy rabbit Brachylagus idahoensis, the world’s smallest leporid. Data Brief. 42, 108314 (2022).
Hisgen, L. et al. Genetic diversity of Treponema paraluisleporidarum isolates in European lagomorphs. Authorea Preprints (2023).
López-Cuamatzi, I. L., Ortega, J. & Baeza, J. A. The complete mitochondrial genome of The’zacatuche’Volcano rabbit (Romerolagus diazi), an endemic and endangered species from the Volcanic Belt of Central Mexico. Mol. Biol. Rep. 49, 1141–1149 (2022).
Zhang, X., Fu, L. & Guo, S. The sequence and characterization of mitochondrial of Lepus oiostolus (Lagomorpha: Leporidae). Mitochondrial DNA Part. B. 5, 2135–2136 (2020).
Melo-Ferreira, J. et al. The elusive nature of adaptive mitochondrial DNA evolution of an arctic lineage prone to frequent introgression. Genome Biol. Evol. 6, 886–896 (2014).
Yao, C. Y. et al. The complete mitochondrial DNA sequence of Yimeng wool rabbit. Mitochondrial DNA Part. B. 4, 3858–3859 (2019).
Saha, A. et al. The first complete mitochondrial genome data of the Afghan pika Ochotona rufescens (Lagomorpha, Ochotonidae), near the type locality. Data Brief. 110246 (2024).
Shan, W. J. & Liu, Y. G. The complete mitochondrial DNA sequence of the cape hare Lepus capensis pamirensis. Mitochondrial DNA Part. A. 27, 4572–4573 (2016).
Osawa, S., Ohama, T., Jukes, T. H. & Watanabe, K. Evolution of the mitochondrial genetic code I. Origin of AGR serine and stop codons in metazoan mitochondria. J. Mol. Evol. 29, 202–207 (1989).
Nagaike, T., Suzuki, T. & Ueda, T. Polyadenylation in mammalian mitochondria: Insights from recent studies. Biochim. et Biophys. Acta (BBA)-Gene Regul. Mech. 1779, 266–269 (2008).
Helm, M. et al. Search for characteristic structural features of mammalian mitochondrial tRNAs. RNA 6, 1356–1379 (2000).
Watanabe, Y., Suematsu, T. & Ohtsuki, T. Losing the stem-loop structure from metazoan mitochondrial tRNAs and co-evolution of interacting factors. Front. Genet. 5, 83261 (2014).
Pereira, S. L. Mitochondrial genome organization and vertebrate phylogenetics. Genet. Mol. Biol. 23, 745–752 (2000).
Steinberg, S. & Cedergren, R. Structural compensation in atypical mitochondrial tRNAs. Nat. Struct. Biol. 1, 507–510 (1994).
Hurst, L. D. The Ka/Ks ratio: Diagnosing the form of sequence evolution. Trends Genet. 18, 486–487 (2002).
Stewart, J. B. et al. Strong purifying selection in transmission of mammalian mitochondrial DNA. PLoS Biol. 6, e10 (2008).
Tsai, P. S., Korotkevich, E. & O’Farrell, P. H. Mitigation of age-dependent accumulation of defective mitochondrial genomes. Proceedings of the National Academy of Sciences 119, (2022).
Ennis, C. C., Ortega, J. & Baeza, J. A. First genomic resource for an endangered neotropical mega-herbivore: The complete mitochondrial genome of the forest-Dweller (Baird’s) Tapir (Tapirus bairdii). Peerj 10, e13440 (2022).
Wei, H. et al. The characterization of complete mitochondrial genome and phylogenetic relationship within Rattus genus (Rodentia: Muridae). Biochem. Syst. Ecol. 71, 179–186 (2017).
Meganathan, P., Pagan, H. J., McCulloch, E. S., Stevens, R. D. & Ray, D. A. Complete mitochondrial genome sequences of three bats species and whole genome mitochondrial analyses reveal patterns of codon bias and lend support to a basal split in Chiroptera. Gene 492, 121–129 (2012).
Tomasco, I. H. & Lessa, E. P. The evolution of mitochondrial genomes in subterranean caviomorph rodents: Adaptation against a background of purifying selection. Mol. Phylogenet. Evol. 61, 64–70 (2011).
Ochoa, A., Onorato, D. P., Fitak, R. R., Roelke-Parker, M. E. & Culver, M. Evolutionary and functional mitogenomics associated with the genetic restoration of the Florida panther. J. Hered. 108, 449–455 (2017).
Luo, Y. et al. Mitochondrial genome analysis of Ochotona curzoniae and implication of cytochrome C oxidase in hypoxic adaptation. Mitochondrion 8, 352–357 (2008).
Ben Slimen, H., Schaschl, H., Knauer, F. & Suchentrunk, F. Selection on the mitochondrial ATP synthase 6 and the NADH dehydrogenase 2 genes in hares (Lepus capensis L., 1758) from a steep ecological gradient in North Africa. BMC Evol. Biol. 17, 1–16 (2017).
Jin, Y. et al. Evolutionary analysis of mitochondrially encoded proteins of toad-headed lizards, Phrynocephalus, along an altitudinal gradient. BMC Genom. 19, 1–11 (2018).
Fonseca, R. R., Johnson, W. E., O’Brien, S. J., Ramos, M. J. & Antunes, A. The adaptive evolution of the mammalian mitochondrial genome. BMC Genom. 9, (2008).
Ramos, B. et al. Landscape Genomics: natural selection drives the evolution of Mitogenome in Penguins. BMC Genom. 19, (2018).
Shen, Y. et al. Adaptive Evolution of Energy Metabolism Genes and the Origin of Flight in Bats. Proceedings of the National Academy of Sciences 107, 8666–8671 (2010).
Shang, Y. et al. Adaptability and evolution of Gobiidae: A genetic exploration. Animals 12, 1741 (2022).
Li, X. et al. Positive selection drove the adaptation of mitochondrial genes to the demands of Flight and High-Altitude environments in Grasshoppers. Front. Genet. 9, (2018).
Zhou, T., Shen, X., Irwin, D. M., Shen, Y. & Zhang, Y. Mitogenomic analyses propose positive selection in mitochondrial genes for high-altitude adaptation in Galliform Birds. Mitochondrion 18, 70–75 (2014).
Amorim, C. E. G., Daub, J. T., Salzano, F. M., Foll, M. & Excoffier, L. Detection of convergent genome-wide signals of adaptation to tropical forests in humans. Plos One. 10, e0121557 (2015).
Gutiérrez, E. G. The mitochondrial genome of the Mountain Wooly Tapir, Tapirus pinchaque and a formal test of the Effect of Altitude on the adaptive evolution of mitochondrial protein coding genes in Odd-Toed ungulates. BMC Genom. 24, (2023).
Bergey, C. M. et al. Polygenic Adaptation and Convergent Evolution on Growth and Cardiac Genetic Pathways in African and Asian Rainforest Hunter-Gatherers. Proceedings of the National Academy of Sciences 115, (2018).
Shen, X., Pu, Z., Chen, X., Murphy, R. W. & Shen, Y. Convergent evolution of mitochondrial genes in Deep-Sea fishes. Front. Genet. 10, (2019).
Zardoya, R. & Meyer, A. Phylogenetic performance of mitochondrial protein-coding genes in resolving relationships among vertebrates. Mol. Biol. Evol. 13, 933–942 (1996).
Kraatz, B. P., Sherratt, E., Bumacod, N. & Wedel, M. J. Ecological correlates to cranial morphology in Leporids (Mammalia, Lagomorpha). PeerJ 3, e844 (2015).
Wood-Bailey, A. P., Cox, P. G. & Sharp, A. C. The evolution of unique cranial traits in leporid lagomorphs. PeerJ 10, e14414 (2022).
Sparwel, M. et al. The volcano rabbit in the phylogenetic network of Lagomorphs. Genome Biol. Evol. 11, 11–16 (2019).
Cano-Sanchez, E. et al. Using ultraconserved elements to unravel lagomorph phylogenetic relationships. J. Mammalian Evol. 29, 395–411 (2022).
Franco-Sierra, N. D. & Díaz‐Nieto, J. F. Rapid mitochondrial genome sequencing based on Oxford Nanopore Sequencing and a proxy for vertebrate species identification. Ecol. Evol. 10, 3544–3560 (2020).
Slapnik, B. et al. The quality and detection limits of mitochondrial heteroplasmy by long read nanopore sequencing. Sci. Rep. 14, 26778 (2024).
Kolmogorov, M., Yuan, J., Lin, Y. & Pevzner, P. A. Assembly of long, error-prone reads using repeat graphs. Nat. Biotechnol. 37, 540–546 (2019).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
Vaser, R., Sović, I., Nagarajan, N. & Šikić, M. Fast and accurate de novo genome assembly from long uncorrected reads. Genome Res. 27, 737–746 (2017).
Donath, A. et al. Improved annotation of protein-coding genes boundaries in metazoan mitochondrial genomes. Nucleic Acids Res. 47, 10543–10552 (2019).
Chan, P. P., Lin, B. Y., Mak, A. J. & Lowe, T. M. tRNAscan-SE 2.0: Improved detection and functional classification of transfer RNA genes. Nucleic Acids Res. 49, 9077–9096 (2021).
Grant, J. R. et al. Proksee: in-depth characterization and visualization of bacterial genomes. Nucleic Acids Res. 51, W484–W492 (2023).
Kumar, S., Stecher, G., Li, M., Knyaz, C. & Tamura, K. MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 35, 1547–1549 (2018).
Darriba, D., Taboada, G. L., Doallo, R. & Posada, D. jModelTest 2: More models, new heuristics and parallel computing. Nat. Methods. 9, 772 (2012).
Trifinopoulos, J., Nguyen, L. T., von Haeseler, A. & Minh, B. Q. W-IQ-TREE: A fast online phylogenetic tool for maximum likelihood analysis. Nucleic Acids Res. 44, W232–W235 (2016).
Bouckaert, R. et al. BEAST 2: A Software platform for Bayesian evolutionary analysis. PLoS Comput. Biol. 10, e1003537 (2014).
Ge, D. et al. Evolutionary history of lagomorphs in response to global environmental change. PLoS One. 8, e59668 (2013).
Erbajeva, M. A. & Alexeeva, N. V. Pliocene and Pleistocene biostratigraphic succession of Transbaikalia with emphasis on small mammals. Quatern. Int. 68, 67–75 (2000).
Rambaut, A., Drummond, A. J., Xie, D., Baele, G. & Suchard, M. A. Posterior summarization in bayesian phylogenetics using Tracer 1.7. Syst. Biol. 67, 901–904 (2018).
Acknowledgements
We would like to express our sincere gratitude to the Ministry of Environment and Forestry of Indonesia for providing the necessary permissions to access genetic resources for the Sumatran rabbit sample used in this study (no. SK.154/KSDAE/SET.3/KSA.2/8/2023). We also thank all the local authorities and field assistants for their invaluable help during the sample collection process.
Author information
Authors and Affiliations
Contributions
D.S Conceptualized and designed the study, performed the genetic analysis, and wrote the the manuscript., N.R contributed to the sequencing and annotation of the mitochondrial genome, T.A Assisted with the bioinformatics analysis and data visualization. I.L performed genetic data analysis and sequencing. I.Y visualized the data and coordinated the fieldwork and sample collection. , A.S coordinated the fieldwork and sample collection and wrote the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Priyono, D.S., Rafina, N., Arisuryanti, T. et al. The first complete mitochondrial genome of Sumatran striped rabbit Nesolagus netscheri (Schlegel, 1880), and its phylogenetic relationship with other Leporidae. Sci Rep 15, 2002 (2025). https://doi.org/10.1038/s41598-025-85212-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-85212-x
Keywords
This article is cited by
-
Adaptive evolution and phylogeny of cerithioid gastropods with six new mitogenomes
Scientific Reports (2025)