Abstract
Nailfold Capillaroscopy (NFC) is a simple, non-invasive diagnostic tool used to detect microvascular changes in nailfold. Chronic pathological changes associated with a wide range of systemic diseases, such as diabetes, cardiovascular disorders, and rheumatological conditions like systemic sclerosis, can manifest as observable microvascular changes in the terminal capillaries of nailfolds. The current gold standard relies on experts performing manual evaluations, which is an exhaustive time-intensive, and subjective process. In this study, we demonstrate the viability of a deep learning approach as an automated clinical screening tool. Our dataset consists of NFC images from a total of 225 participants, with normal images accounting for 6% of the dataset. This study introduces a robust framework utilizing cascade transfer learning based on EfficientNet-B0 to differentiate between normal and abnormal cases within NFC images. The results demonstrate that pre-trained EfficientNet-B0 on the ImageNet dataset, followed by transfer learning from domain-specific classes, significantly enhances the classifier’s performance in distinguishing between Normal and Abnormal classes. Our proposed model achieved superior performance, with accuracy, precision, recall, F1 score, and ROC_AUC of 1.00, significantly outperforming both models of single transfer learning on the pre-trained EfficientNet-B0 and cascade transfer learning on a convolutional neural network, which each attained an accuracy, precision, recall, and F1 score of 0.67 and a ROC_AUC of 0.83. The framework demonstrates the potential to facilitate early preventive measures and timely interventions that aim to improve healthcare delivery and patients’ quality of life.
Similar content being viewed by others
Introduction
Nailfold capillaroscopy (NFC) is a traditional method for health assessment that provides a direct and non-invasive means to evaluate the microvascular structure in nailfold capillaries. Microvascular integrity plays a vital role in maintaining microcirculation, facilitating material exchange in the body. Previous studies indicate that diseases such as diabetes1, rheumatoid arthritis2, Raynaud’s phenomenon, and systemic sclerosis (SSc)3,4,5,6 can lead to microcirculatory changes. Over the past three decades, NFC has demonstrated significant promise in diagnosing cardiovascular conditions such as hypertension7, hypotension8, pulmonary hypertension9, as well as auto-immune rheumatological conditions such as systemic sclerosis5, systemic lupus erythematosus10, idiopathic inflammatory myopathies11, juvenile dermatomyositis12, ankylosing spondylitis13, mixed connective tissue disease14, and Sjögren’s syndrome15. Studies have revealed meaningful variations in quantitative characteristics of microcirculations observed in NFC images among patients16. Figure 1 illustrates the approximate location and the microcirculatory pathways within these capillaries. The conventional manual approach to NFC analysis typically entails visual inspection and manual measurement of parameters such as the number of crossing capillaries, as well as their length and diameter5,17. This process heavily relies on the expertise of medical professionals to interpret the images and make diagnostic assessments based on their observations. However, this manual approach poses several challenges. Firstly, it is time-consuming and labor-intensive, requiring significant effort and attention to detail from skilled practitioners. Additionally, subjective interpretations can lead to variability in results, impacting the reliability and consistency of diagnoses. Moreover, the manual analysis may overlook subtle or complex patterns in the NFC images that could be indicative of underlying health conditions.
The emergence of machine learning, particularly deep learning techniques, has presented a promising alternative to NFC analysis due to its superior accuracy over traditional image processing techniques18. These techniques leverage datasets to automatically learn and extract meaningful features from NFC images, allowing for more objective and accurate analysis compared to traditional methods. Recent advancements in deep learning for NFC analysis have shown remarkable potential in improving diagnostic accuracy and efficiency. These methods can automatically detect and quantify various microcirculatory abnormalities, providing valuable insights for early disease detection and monitoring. Subsequently, an overview of recent advancements in NFC analysis employing deep learning techniques is provided.
Deep learning-based nailfold capillary analysis
In recent years, numerous studies have focused on deciphering NFC images to extract their morphology and blood flow patterns, which serve as crucial indicators for various medical conditions. Precise identification of these patterns enables accurate diagnosis, monitoring, and assessment of treatment effectiveness, providing valuable insights into microcirculation health and subsequently potential health issues. Some of these studies have primarily concentrated on segmentation techniques which provide doctors with specific measurements of capillaries, enabling them to assess their appearance more easily. In addition to segmentation, other studies have directly addressed classification techniques. These techniques aim to categorize images based on characteristic features, such as the presence of abnormalities or specific patterns indicative of particular medical conditions.
Hariyani et al.19 proposed an improved version of U-Net architecture using a dual attention module for nailfold capillary segmentation to precisely distinguish capillaries from the background in images.It achieved higher IoU (Intersection over Union) scores compared to the baselines, with increases of 0.43, 0.09, and 0.087 over the adaptive Gaussian algorithm, original U-Net, and SegNet, respectively. The dataset comprised NFC images from seven healthy participants, and a total of 40 images, 30 for training and 10 for testing were used in this study. Liu et al.8 addressed the challenges associated with poor-quality nailfold capillary images by proposing a neural network combining U-Net and ResNet, termed Res-Unet. The method involved preprocessing the images, data augmentation, and employing the Res-Unet architecture for capillary segmentation. The Res-Unet model demonstrated superior performance compared to U-Net, accurately segmenting capillary structures even in low-quality images. The dataset primarily consisted of images from healthy subjects, supplemented by a few cases of hypertension and hypotension. Nailfold capillary patterns associated with these conditions were analyzed. A total of 50 images were utilized, with 30 images designated for training and 20 for testing. Nitkunanantharajah et al.20 employed optoacoustic imaging (OAI) to compare nailfold capillaries of patients with SSc and healthy controls. A pre-trained model named Resnet18 was utilized and achieved an accuracy of 0.848, with a 78.3% sensitivity and 89.5% specificity in automatically differentiating NFC images from both groups. This study included a dataset of nailfold images from 23 SSc patients and 19 healthy controls, focusing on detecting capillary patterns unique to SSc. Shah et al.21 utilized nailfold capillary images acquired through video capillaroscopy for diagnosing diabetes. They employed different versions of the ResNet family to analyze the NFC images. The study found that the models could accurately predict the presence of diabetes with areas under the receiver operating characteristics curves (AUROC) 0.84 and a history of cardiovascular events in diabetic patients with AUROC 0.65. In this study, the dataset comprised 5236 nailfold images collected from 120 participants, including 60 individuals with diabetes (30 with type 1 diabetes and 30 with type 2 diabetes) and 60 without diabetes. K V et al.22 proposed an approach for detecting diabetic microvascular changes using NFC images. The dataset comprised 600 images collected from diabetic and healthy subjects.Their method involves employing YOLOv2 and YOLOv3 object detection algorithms, to classify capillary abnormalities indicative of diabetes. Their proposed model achieved an accuracy of 88.2% in differentiating between normal and diabetic images. Li et al.23 developed a method for the automatic measurement of vascular parameters in nailfold microcirculation vessels to aid medical diagnosis. The cohorts or the exact number of images used for training and testing are not specified. Their method involves using YOLOv5 algorithms for blood vessel detection and segmentation, and U-Net for image segmentation. The final results demonstrate the superior performance of their method compared with manual methods traditionally used for this purpose. Bharathi et al.16 developed a fully automated image analysis system for NFC to aid in the diagnosis of SSc. The study included a balanced dataset with a significant representation of SSc patients, focusing on detecting abnormalities such as giant capillaries, capillary tortuosity, and avascular zones to aid in distinguishing SSc from normal cases. They employed the image analysis pipeline to detect and segment capillaries using U-Nets, measure their morphology using ResNet34, and logistic regression to classify subjects based on microvascular abnormalities. The system achieved diagnostic performance comparable to SSc experts, with the area under the ROC curves of 97% and 95% for different imaging conditions which outperformed expert consensus which achieved sensitivity of 82% and specificity of 73%. Kassani et al.12 explored the potential of artificial intelligence (AI), specifically a lightweight and explainable deep neural network model called NFC-Net, to differentiate children with juvenile dermatomyositis (JDM) from healthy controls using NFC images, as well as to assess disease activity in JDM patients. NFC-Net achieved good performance in distinguishing JDM patients from healthy controls, with an AUROC of 0.93 and an accuracy of 0.91. Their dataset consisted of 1,441 images (1,120 from 111 JDM patients and 321 from 31 healthy controls), with features such as capillary dropout and irregular loops being prominent. Another study by Nguyen et al.24 proposed a system for detecting nailfold capillary morphology indicative of type 2 diabetes mellitus. Their dataset consisted of 613 images, capturing features such as tortuosity and capillary enlargement. They employed the YOLOv5 model with a Bi-directional Feature Pyramid Network (Bi-FPN) to detect and classify capillaries into different morphology types. The final results show that YOLOv5Bi-FPN achieves a mean Average Precision of 0.74, outperforming other variations of YOLO. Venkatapathiah et al.25 developed an object detection algorithm based on deep learning architectures for detecting and locating various capillary loops in the nailfold region. Their dataset comprised NFC images highlighting abnormalities such as tortuosity, hemorrhages, variations in capillary width, and other morphological changes commonly associated with SSc and related vascular conditions. However, the exact number of images included in the dataset was not specified. Their method involves creating a custom dataset, training an object detection network using YOLOv3, and evaluating the performance of detecting different capillary features such as width, tortuosity, and hemorrhages. The final results demonstrated that YOLOv3 outperforms YOLOv2 in terms of accuracy for various classes of capillaries. Gracia Tello et al.26 utilized deep convolutional neural networks (CNNs) to classify nailfold videocapillaroscopy images according to the presence/absence of structural abnormalities and/or microhemorrhages. The final results demonstrate high levels of agreement between the automated system and expert capillaroscopists. The dataset used in this study comprised 1,164 nailfold videocapillaroscopy (NVC) images from patients with Raynaud’s phenomenon (RP). These images included various categories of abnormalities such as dilations, giant capillaries, abnormal shapes, tortuosities, and microhaemorrhages, as well as normal capillaries. However, the exact division between healthy controls and capillary abnormalities was not specified in this study. In another study, Chen et al.27 developed a U-Net backbone network combined with attention mechanisms (CBAM, CA, ECA, SE) and residual blocks, along with contrast enhancement using CLAHE. Their dataset consisted of 613 nailfold capillary images, focusing on abnormalities such as capillary dropout, hemorrhages, and tortuosity. The proposed method, Res_CBAM_Unet with CLAHE (clipLimit = 2), achieves an overall accuracy of 94.31%, Dice score of 0.8297, and IoU score of 0.7090, demonstrating effectiveness in nailfold capillary segmentation. In addition to the aforementioned studies, our research team reviewed state-of-the-art studies utilizing AI in nailfold capillary analysis, which highlighted the need for additional investigation in this field28.
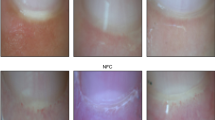
In addition to the promising results achieved by applying intelligent methods to identify abnormalities in NFC images, several challenges persist due to the inherent nature of these images. Factors such as variations in brightness, light reflections from the surrounding environment or the microscope, and diverse patient characteristics—including variations in skin thickness, pigmentation, and the presence of artifacts like dust particles—further complicate the interpretability of the images, as illustrated in Fig. 2. Therefore, it is imperative to design models that can effectively address the complexity and challenges inherent in these images.
Emerging approaches, building upon manual methods of NFC image analysis, utilize deep learning capabilities to segment images into distinct capillary types and extract their key measurements. These extracted features serve as the input of classification models or as intermediary reports for medical professionals to evaluate patients’ health. This study aims to introduce a straightforward, robust, and lightweight classification model, built on the capabilities of EfficientNets, to effectively differentiate between normal and abnormal NFC images. Our objective extends beyond catering solely to expert doctors; we seek to equip non-expert medical professionals as well as non-specialists with the ability to accurately discern between normal and abnormal cases. The proposed methodology delineating our approach to achieving this objective will be elaborated in the subsequent section.
Materials and methods
To enhance the reporting standards in clinical scientific literature utilizing AI, quality assessment tools are designed for the medical domain to mitigate bias and enhance publication quality. Hence, we integrated the CAIR29 and MINIMAR30 quality assessment tools during the manuscript preparation.
Dataset
The dataset was collected from participants at the Hospital General Zona Este de Tijuana, Mexico, following approval from the Ethics Committee (reference number 2023-AJF-32B). Informed consent was obtained from participants after ensuring their comprehension of the study details. All methods were performed in accordance with the relevant guidelines and regulations.
Participant selection and demographics
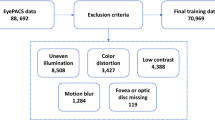
Participants were recruited through convenience sampling via prospective enrollment, with population demographics detailed in Table 1. Participant selection adhered to the inclusion and exclusion criteria outlined in Table 2.
Subject preparation
Participants underwent a series of preparatory steps to maximize precision and reliability in the obtained results. Prior to the examination day, they were advised to manage stress and anxiety, and it was made sure that they had avoided the application of any cosmetic procedures to their nails such as nail polish for at least 3 weeks prior. This is to avoid unnecessary misinterpretation of results, since nail polish will impair visualization directly, and stress might result in increased sympathetic activity resulting in vasoconstriction of blood vessels. On the day of image acquisition, they were advised to skip breakfast, and not to drink coffee, tea, or smoke to avoid any interaction with the microcirculation due to the vasoconstrictive effects of caffeine or nicotine. They were also asked to avoid cosmetics such as creams, ointments, or lotions to avoid blurring and to ensure the transparency and clarity of the image.
Apparatus setup
The apparatus was maintained at room temperature of 20–23℃ without any external temperature control. Participants were allowed to sit in the room for a while (5–10 min) to acclimate to room temperature before image acquisition. Next, they were instructed to clean their hands, and after being seated, a few drops of immersion oil was applied to the cuticle area of the nailfold to minimize light reflections and improve visualization.
Image acquisition
The Dino-Lite Edge medical microscope was used to acquire images at x200 magnification as recommended by Cutolo et al.31. It was operated by a trained technician, who underwent a 15-day training period to standardize image acquisition and minimize operator bias. One image was typically captured from the middle zone of the index finger (second finger) per participant. In case it was deemed of low quality because of inadequate focus, motion artifacts, or excessive light reflections, then a second image was taken. The images were captured right away instead of filming a video and then extracting images later. To ensure accurate measurements, the distance between the microscope and the nailfold was calibrated and a scale was included in the captured images. Consequently, a total of 361 NFC images were collected from a cohort of 225 participants.
Pre-analysis processing
Pre-analysis processing was conducted manually by three internal medicine specialist physicians. This process involved sorting and labeling the images by their class (ground truth). Images featuring evident foreign objects, or deemed of low quality due to blurring, motion artifacts, light reflections, or lack of focus as deemed by experts were excluded. Therefore 73 images were excluded from the original 361 images to yield the final dataset of 288 images.
Dataset description
After pre-analysis processing, the final dataset comprised 288 NFC images including 16 images (6%) in the normal group and 272 (94%) in the abnormal group. The abnormal group consisted of 112 dermatomycosis images, 143 tortuosity images, and 17 general abnormality images representing abnormal features or non-specific changes on NFC such as microhemorrhages, dilatation, giant capillaries, and ramifications.
Model architecture and development
The primary objective of this study is to develop a model capable of effectively distinguishing between normal and abnormal NFC images. The dataset comprised 6% normal images compared to 94% abnormal images, reflecting a significant class imbalance. This can be explained by the fact that NFC image collections predominantly occur in clinical settings, where abnormal cases are more prevalent. Therefore, it becomes crucial to develop a model capable of effectively addressing such disparate class distributions while maintaining robust classification performance. Furthermore, the intricate nature of capillary morphology in NFC images and the nuanced criteria used by both AI models and medical professionals to assess image normality or abnormality highlight the necessity of a model capable of deciphering these complexities. The limited number of normal cases further increases the risk of overfitting which necessitates a model that can generalize well from a small and imbalanced dataset. In light of these considerations, our model was built on the foundation of EfficientNets. This choice was strategic, as EfficientNet offers a balanced trade-off between model complexity and computational efficiency which leads to a reduced parameter count and achieving faster processing compared to other state-of-the-art architectures32. Its lightweight design makes it particularly suitable for tasks with limited data, as it can help mitigate overfitting risks by employing fewer parameters while maintaining strong feature extraction capabilities32,33, making it well-suited for our objectives. The architecture of EfficientNet-B032 utilized in this study is depicted in Table 3.
The main architectural component of the EfficientNet-B0 baseline network is the mobile inverted bottleneck MBConv, as proposed by Sandler et al.34 in 2018 and further refined by Tan et al.35 in 2019. This building block combines depthwise separable convolutions with a bottleneck structure, aimed at reducing computational complexity while maintaining performance. Additionally, the network incorporates squeeze-and-excitation optimization techniques introduced by Hu et al.36 in 2018, which enhance feature representation by adaptively recalibrating channel-wise feature responses.
Cascade transfer learning
One persistent challenge in supervised learning is the scarcity of labeled data, which is common in various domains. This scarcity is particularly notable in the medical domain, where labeling data requires the expertise of professional specialists. Transfer learning can mitigate this challenge by transferring the knowledge extracted from one task, problem, or domain to another. Innovative perspectives on transfer learning can extend to the transfer of knowledge across diverse classes within a given problem domain. To optimize resource utilization and address dataset imbalances, this approach facilitates the simultaneous utilization of all available resources. Methods must be developed to comprehensively extract knowledge from accessible classes, encompassing not only the target class but also other available classes within the dataset. Our innovative cascade transfer learning framework builds on this objective by progressively transferring knowledge across different classes within the dataset. This approach aims to comprehensively utilize the available data, mitigate the effects of class imbalance, and minimize the risk of overfitting. To optimize the learning process and implement cascade transfer learning the dataset was strategically partitioned into three subsets of Dataset A, Dataset B, and a Test set. Dataset A includes 255 images from the dermatomycosis and tortuosity classes, used for domain-specific pretraining to enable the model to learn NFC-specific features. Dataset B contains 21 images, consisting of 10 normal and 11 general abnormality images, used for fine-tuning the model for the target classification of Normal versus Abnormal. A separate test set of 12 completely independent and randomly selected images was reserved exclusively for final performance evaluation. The cascade transfer learning process in this study involved two sequential steps. Step A - Domain-Specific Pretraining on Dataset A: In the first step, the EfficientNet-B0 model, pre-trained on the ImageNet dataset, is fine-tuned on Dataset A, which consists of domain-specific non-target classes of dermatomycosis and tortuosity images. By treating dermatomycosis and tortuosity as separate classes, the model is able to adapt its generalized visual features learned from ImageNet to the specific morphological patterns seen in NFC images. During this stage, deeper layers of the model are fine-tuned to focus on NFC-specific details, enhancing the representation of domain-relevant features. This pretraining establishes a foundation for the next step, which focuses on the target classification task. Step B - Fine-Tuning on Target Classes with Dataset B: After pretraining, the model is fine-tuned on Dataset B, which includes target classes. This step fine-tunes the model for the target classification task of distinguishing between Normal and Abnormal classes. By using the refined feature representations developed in Step A, this stage allows the model to better handle the small target dataset.
Data augmentation
To further mitigate the risk of overfitting due to the small dataset size, we employed data augmentation techniques during the training process. These included random rotations, where each image was randomly rotated by an angle up to 20% of a full rotation, horizontal and vertical flipping, and brightness and contrast adjustments to simulate variations in lighting conditions. These augmentation strategies artificially increased the diversity of the training data which enables the model to generalize more effectively to unseen images.
In the subsequent section, we provide a detailed explanation of our proposed network architecture, tailored to accomplish these objectives.
Model building
To implement the cascade transfer learning framework described earlier, we developed a deep learning pipeline based on the EfficientNet-B0 architecture. This section provides a detailed explanation of the model-building process, including the rationale for layer selection, regularization techniques, hyperparameter optimization, and training configurations.
The base model is initialized with pre-trained weights from ImageNet, which provides it with an immediate ability in image recognition by leveraging generalized features relevant to diverse image recognition tasks. However, to effectively capture the nuanced characteristics of the microvascular patterns in nailfold capillaries and optimize performance for the specific task of distinguishing between normal and abnormal NFC images, further fine-tuning of the final layers becomes essential. Our objective was to progressively guide the model in learning these patterns, addressing the class imbalance issue while simultaneously improving its discriminatory capabilities. To achieve this goal, we developed a two-step cascade model, termed CE-NFCNet (Cascade EfficientNet), illustrated in Fig. 3. Initially, an instance of the EfficientNet-B0 model was initialized with pre-trained weights from the ImageNet dataset. Fine-tuning was performed in two steps, corresponding to the cascade framework. In Step A, the fine-tuning process started with the base model on Dataset A including dermatomycosis and tortuosity samples. A 70/30 split was applied for training and validation within Dataset (A) Layers 6d, 7a, and the top layer from stages 6, 7, and 8, respectively, were fine-tuned, as determined through a grid search to optimize performance while minimizing overfitting (refer to Table 3 for stage details). To further enhance generalization and mitigate overfitting, stage 8 was augmented with additional regularization layers, as depicted in Fig. 3. These included dropout layers with a rate of 0.3, global average pooling to aggregate information across channels, and batch normalization to stabilize training and improve generalization. The configuration of these layers was determined through hyperparameter tuning, using a grid search across various regularization layer configurations to identify the optimal settings. Additionally, the optimal values for key hyperparameters were determined as a learning rate of 1e-3, a batch size of 16, and the Adam optimizer. The model was trained for up to 100 epochs, with early stopping applied based on validation loss, utilizing a patience threshold of 10 epochs to mitigate overfitting. Although this step does not directly target our intended classification, it helps the model capture the intricacies of the nailfold microvascular patterns from NFC images within the dermatomycosis and tortuosity classes, which can subsequently be transferred to the Normal versus Abnormal classification task. In Step B, the fine-tuned model from the previous step, serving as the base model, underwent further fine-tuning in stages 7a, and 8 on Dataset B, which encompasses the Normal and Abnormal classes. To enhance robustness and mitigate overfitting, 3-fold cross-validation was applied during training and validation for Dataset (B) No additional regularization layers were applied in Step B, as grid search results indicated that they were unnecessary for achieving optimal performance at this stage. The optimal configuration included a batch size of 8, a learning rate of 1e-4, and the Adam optimizer, with early stopping implemented using a patience threshold of 7 epochs and a maximum training limit of 100 epochs.
Schematic of the CE-NFCNet process. The pre-trained EfficientNet-B0 model from the ImageNet dataset serves as the base model. In Step A, specific final layers in stages 6,7, and 8 are fine-tuned to capture the characteristics of NFC images from non-target classes. In Step B, utilizing the fine-tuned model from Step A as the base, further fine-tuning in Stages 7 and 8 is done to learn distinct characteristics between Normal and Abnormal classes.
Evaluation metrics
The performance of the models in classifying NFC images was assessed using both quantitative metrics and visual analysis. The evaluation metrics selected for this study include Accuracy, Precision, Recall (Sensitivity), F1 Score, Area Under the ROC Curve (AUC-ROC), and the Precision-Recall (PR) curve, providing a comprehensive assessment of the model’s classification capabilities. Accuracy serves as a general measure of the model’s overall correctness but may be less informative for imbalanced datasets. Precision evaluates the model’s ability to minimize false positives, which is critical in medical diagnostics where incorrect positive predictions can lead to unnecessary interventions. Recall measures the model’s ability to identify all abnormal images, emphasizing its reliability in detecting clinically significant conditions by minimizing false negatives. The F1 Score, as the harmonic mean of Precision and Recall, provides a balanced evaluation of both false positives and false negatives. AUC-ROC quantifies the model’s discriminatory power, reflecting its capability to distinguish between normal and abnormal images across various thresholds. The PR curve emphasizes the trade-off between Precision and Recall. Additionally, confusion matrices were used to provide further insights into classification performance. Table 4 summarizes the formulas and meanings of each metric, while Table 5 defines the confusion matrix elements.
Results and discussion
In this study, in addition to evaluating the performance of CE-NFCNet, we assessed the performance of two additional models: SE-NFCNet (Single EfficientNet) and CC-NFCNet (Cascade CNN). SE-NFCNet was initialized with ImageNet pre-trained weights and directly fine-tuned on Dataset B without intermediate domain-specific pretraining. Furthermore, CC-NFCNet, a convolutional neural network trained from scratch on Dataset A and was subsequently fine-tuned on Dataset B. The comparative performance results of all three models are detailed in Table 6.
Perfect classification performance was achieved by CE-NFCNet, with accuracy, precision, recall, F1 score, and ROC_AUC all equal to 1.0. This indicates the capability of the model to accurately classify all samples in the test set with a perfect balance between precision and recall while maintaining flawless discrimination capability. We recognize that this perfect score is due to the ability to fine-tune the model on a small dataset. However, It demonstrates that this approach outperforms other models. In comparison, SE-NFCNet and CC-NFCNet both demonstrated lower performance compared to CE-NFCNet, with accuracy, precision, recall, and an F1 score of 0.67. However, their ROC_AUC scores remained relatively high at 0.83, indicating good discrimination ability. For a more detailed analysis of the models’ performance, the PR curves are shown in Fig. 4a, where the Break-Even Point (BEP) serves as an evaluation metric. BEP occurs when precision and recall are equal37, which offers a balanced assessment of the trade-off between the two metrics. As evident from Fig. 4a, CE-NFCNet significantly outperforms SE-NFCNet and CC-NFCNet in terms of BEP. Additionally, the Average Precision (AP) scores, which provide a comprehensive evaluation of the models’ performance across all thresholds, are presented in Fig. 4b. The AP scores were calculated as 1.00 for CE-NFCNet, 0.863 for SE-NFCNet, and 0.836 for CC-NFCNet. These results highlight the superior performance of CE-NFCNet, further confirming its ability to generalize and classify samples under varying conditions, while SE-NFCNet slightly outperforms CC-NFCNet in distinguishing between normal and abnormal images over the entire range of thresholds. As shown in the confusion matrices in Fig. 5, the proposed model, CE-NFCNet, achieved 100% accuracy, correctly classifying all samples. In contrast, SE-NFCNet and CC-NFCNet both correctly identified 8 out of the 12 samples while misclassifying the remaining 4 images, accounting for 33% misclassification rate. Although the overall performance of SE-NFCNet and CC-NFCNet was identical, some of the samples they misclassified differed which reflects variations in their feature learning and decision-making processes.
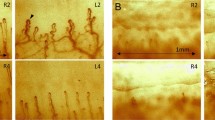
To better evaluate the models’ performance, we curated the test set to encompass various scenarios, including variations in brightness, light reflections from the surrounding environment or microscope, and diverse patient characteristics such as skin pigmentation. We categorized images based on different characteristics and randomly selected samples from each group to ensure a comprehensive testing set. Figure 6 presents some of the test samples incorrectly classified by CC-NFCNet and SE-NFCNet. Remarkably, all samples misclassified by these two models were accurately classified by CE-NFCNet.
The observations in Fig. 6 underscore the challenges encountered by both models in correctly identifying the true classes across different scenarios. For instance, in Fig. 6 (a) the challenge of blurring, low resolution, and dark images with poor contrast affected the SE-NFCNet model performance negatively. Similarly, in Fig. 6 (b) the performance of the CC-NFCNet model was impacted by blurring and light reflection, darker skin tone with the presence of light reflections, and variable microvascular distribution. However, the proposed CE-NFCNet, empowered by the capabilities of EfficientNet-B0 and cascade transfer learning, demonstrates remarkable effectiveness in distinguishing between Normal and Abnormal classes under various conditions and overcomes the aforementioned challenges the other models could not address. We note that despite both models achieving statistical equivalence in their overall performance Fig. 6 shows how they differ in misclassification.
Limitations and future work
While the proposed CE-NFCNet model achieved perfect classification metrics (accuracy, precision, recall, F1 score, and AUC-ROC) on the independent test set, we acknowledge that the small size of the test set (12 images) presents statistical limitations. This constraint stems from the limited availability of normal cases in our dataset, a challenge often encountered in clinical data collection. We mitigated this limitation by employing the cascade transfer learning framework to transfer knowledge from ImageNet and the domain-specific dataset to the target classification, as well as through cross-validation on the target classification task and careful selection of the test set. Despite the practical limitations of the dataset, the model demonstrated the viability of this approach.
Future work should focus on validating the model on larger datasets to further confirm its generalizability and robustness. Incorporating datasets with broader variability in both normal and abnormal cases will better capture the diversity seen in clinical practice. By addressing these limitations, CE-NFCNet can be refined into a reliable, clinically applicable tool for the automated screening of NFC images, contributing to earlier interventions.
Conclusion
In conclusion, this study presents a robust framework utilizing cascade transfer learning on EfficientNet-B0 to effectively distinguish between normal and abnormal NFC images. The utilization of a pre-trained EfficientNet-B0 model on the ImageNet dataset, followed by a two-step fine-tuning to transfer knowledge between classes within the NFC images, significantly enhanced the classifier’s performance in detecting abnormalities and addressing the challenge of class imbalance. By evaluating various performance metrics and analyzing confusion matrix elements, it became apparent that the CE-NFCNet model incorporating cascade transfer learning on EfficientNet-B0 surpasses both the SE-NFCNet model employing single transfer learning and one-step fine-tuning on Dataset B, as well as the convolutional neural network model trained on Dataset A and fine-tuned on Dataset B (CC-NFCNet). CE-NFCNet achieved perfect classification performance, striking a balance between precision and recall. Furthermore, the PR curves and incorporation of the BEP metric and AP score provided clear insights into the model’s performance, highlighting the superiority of CE-NFCNet in achieving a harmonious balance between precision and recall.
Overall, the findings of this study underscore the effectiveness of cascade transfer learning approaches along with the potential of EfficientNet-B0 models in NFC image classification tasks and highlight addressing challenges associated with small and imbalanced datasets, especially in healthcare applications. Encouraged by these promising findings, our next step involves expanding our study to detect various diseases in abnormal images.
Data availability
The datasets utilized during the current study are not publicly available due to the hospital’s privacy policy restrictions but are available from the corresponding author upon reasonable request.
References
Niraj, B. et al. Nailfold capillary changes in diabetes mellitus and their correlation with diabetic retinopathy. MAMC J. Med. Sci. 8, 153–157. https://doi.org/10.4103/mamcjms.mamcjms_5_22 (2022).
Rajaei, A., Dehghan, P. & Amiri, A. Nailfold capillaroscopy in 430 patients with rheumatoid arthritis. Casp. J. Intern. Med. 8, 269–274. https://doi.org/10.22088/cjim.8.4.269 (2017).
Del Galdo, F. et al. Nailfold capillaroscopy in rheumatic diseases: which parameters should be evaluated? Biomed. Res. Int. 2015(974530). https://doi.org/10.1155/2015/974530 (2015).
Cutolo, M. & Matucci Cerinic, M. Nailfold capillaroscopy and classification criteria for systemic sclerosis. Clin. Exp. Rheumatol. 25, 663–665 (2007).
Smith, V. et al. Standardisation of nailfold capillaroscopy for the assessment of patients with Raynaud’s phenomenon and systemic sclerosis. Autoimmun. Rev. 19, 102458. https://doi.org/10.1016/j.autrev.2020.102458 (2020).
Ingegnoli, F. & Gualtierotti, R. A systematic overview on the use and relevance of capillaroscopy in systemic sclerosis. Expert Rev. Clin. Immunol. 9, 1091–1097. https://doi.org/10.1586/1744666X.2013.849198 (2013).
Mishra, A., Grover, C., Singal, A., Narang, S. & Das, G. K. Nailfold capillary changes in newly diagnosed hypertensive patients: an observational analytical study. Microvasc Res. 136, 104173. https://doi.org/10.1016/j.mvr.2021.104173 (2021).
Liu, S. et al. Segmenting nailfold capillaries using an improved U-net network. Microvasc Res. 130, 104011. https://doi.org/10.1016/j.mvr.2020.104011 (2020).
Arvanitaki, A. et al. Peripheral microangiopathy in precapillary pulmonary hypertension: a nailfold video capillaroscopy prospective study. Respir Res. 22, 27. https://doi.org/10.1186/s12931-021-01622-1 (2021).
Cutolo, M. et al. Nailfold capillaroscopy in systemic lupus erythematosus: a systematic review and critical appraisal. Autoimmun. Rev. 17, 344–352. https://doi.org/10.1016/j.autrev.2017.11.025 (2018).
Piette, Y. et al. Standardised interpretation of capillaroscopy in autoimmune idiopathic inflammatory myopathies: a structured review on behalf of the EULAR study group on microcirculation in Rheumatic diseases. Autoimmun. Rev. 21, 103087. https://doi.org/10.1016/j.autrev.2022.103087 (2022).
Kassani, P. H. et al. Artificial intelligence for nailfold capillaroscopy analyses – a proof of concept application in juvenile dermatomyositis. Pediatr. Res. 95, 981–987. https://doi.org/10.1038/s41390-023-02894-7 (2024).
Batko, B. et al. Microvascular dysfunction in ankylosing spondylitis is associated with disease activity and is improved by anti-TNF treatment. Sci. Rep. 8, 13205. https://doi.org/10.1038/s41598-018-31550-y (2018).
Granier, F., Vayssairat, M., Priollet, P. & Housset, E. Nailfold capillary microscopy in mixed connective tissue disease: comparison with systemic sclerosis and systemic lupus erythematosus. Arthritis Rheum. 29, 189–195 (1986).
Tektonidou, M., Kaskani, E., Skopouli, F. N. & Moutsopoulos, H. M. Microvascular abnormalities in Sjögren’s syndrome: Nailfold Capillaroscopy. Rheumatology 38, 826–830. https://doi.org/10.1093/rheumatology/38.9.826 (1999).
Bharathi, P. G. et al. A deep learning system for quantitative assessment of microvascular abnormalities in nailfold capillary images. Rheumatology 62, 2325–2329 (2023).
Smith, V. et al. Nailfold Capillaroscopy. Best Pract. Res. Clin. Rheumatol. 37, 101849. https://doi.org/10.1016/j.berh.2023.101849 (2023).
Gracia Tello, B. et al. The challenge of comprehensive nailfold videocapillaroscopy practice: a further contribution. Clin. Exp. Rheumatol. 40, 1926–1932 (2022).
Hariyani, Y. S., Eom, H. & Park, C. DA-Capnet: dual attention deep learning based on U-Net for nailfold capillary segmentation. IEEE Access. 8, 10543–10553. https://doi.org/10.1109/ACCESS.2020.2965651 (2020).
Nitkunanantharajah, S. et al. Three-dimensional optoacoustic imaging of nailfold capillaries in systemic sclerosis and its potential for disease differentiation using deep learning. Sci. Rep. 10, 16444. https://doi.org/10.1038/s41598-020-73319-2 (2020).
Shah, R. et al. Nailfold capillaroscopy and deep learning in diabetes. J. Diabetes 15, 145–151. https://doi.org/10.1111/1753-0407.13354 (2023).
Selvi, K. V. S., Nanda, S., Shetty, P., Awasthi, K. & M., M, V. & Deep learning approach to nailfold capillaroscopy-based diabetes mellitus detection. Int. J. Online Biomed. Eng. 18, 95–109. https://doi.org/10.3991/ijoe.v18i06.27385 (2022).
Li, H., Li, M. & Zhang, H. Detection and segmentation of nailfold microcirculation vessels based on deep learning. In Proceedings of ECITech 2022: The 2022 International Conference on Electrical, Control and Information Technology, 1–4 (2022).
Nguyen, H. T. P., Ko, S. & Jeong, H. Deep-learning-based capillary detection. In 2023 IEEE International Conference on Bioinformatics and Biomedicine (BIBM) 4932–4934. https://doi.org/10.1109/BIBM58861.2023.10385650 (2023).
Venkatapathiah, K. et al. Deep learning-based object detection in nailfold capillary images. IAES Int. J. Artif. Intell. (IJ-AI) 12, 931–942. https://doi.org/10.11591/ijai.v12.i2.pp931-942 (2023).
Gracia Tello, B. C. et al. External clinical validation of automated software to identify structural abnormalities and microhaemorrhages in nailfold videocapillaroscopy images. Clin. Exp. Rheumatol. 41, 1605–1611. https://doi.org/10.55563/clinexprheumatol/m6obl3 (2023).
Chen, S., Wei, D., Gu, S. & Yang, Z. Nailfold capillary segmentation based on U-Net and attention mechanism. In Proceedings of the 2023 3rd International Conference on Bioinformatics and Intelligent Computing (BIC ‘23), 337–343. https://doi.org/10.1145/3592686.3592747 (2023).
Emam, O. S., Ebadi Jalal, M., Garcia-Zapirain, B. & Elmaghraby, A. Artificial intelligence algorithms in nailfold capillaroscopy image analysis: a systematic review. medRxiv 2024.07.28.24311154. https://doi.org/10.1101/2024.07.28.24311154 (2024).
Olczak, J. et al. Presenting artificial intelligence, deep learning, and machine learning studies to clinicians and healthcare stakeholders: an introductory reference with a guideline and a Clinical AI Research (CAIR) checklist proposal. Acta Orthop. 92, 513–525. https://doi.org/10.1080/17453674.2021.1918389 (2021).
Hernandez-Boussard, T., Bozkurt, S., Ioannidis, J. P. A. & Shah, N. H. MINIMAR (Minimum Information for Medical AI reporting): developing reporting standards for artificial intelligence in health care. J. Am. Med. Inf. Assoc. 27, 2011–2015. https://doi.org/10.1093/jamia/ocaa088 (2020).
Cutolo, M. et al. Automated assessment of absolute nailfold capillary number on videocapillaroscopic images: Proof of principle and validation in systemic sclerosis. Microcirculation 25, e12447. https://doi.org/10.1111/micc.12447 (2018).
Tan, M., Le, Q. & EfficientNet Rethinking model scaling for convolutional neural networks. In International Conference on Machine Learning, PMLR, 6105–6114 (2019).
Rahman, Z. et al. Enhancing image-based diagnosis of gastrointestinal tract diseases through deep learning with EfficientNet and advanced data augmentation techniques. BMC Med. Imaging 24, 306. https://doi.org/10.1186/s12880-024-01479-y (2024).
Sandler, M., Howard, A., Zhu, M., Zhmoginov, A. & Chen, L. C. MobileNetV2: Inverted residuals and linear bottlenecks. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, 4510–4520 (2018).
Tan, M. et al. MnasNet: Platform-aware neural architecture search for mobile. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 2820–2828 (2019).
Hu, J., Shen, L. & Sun, G. Squeeze-and-excitation networks. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, 7132–7141 (2018).
Zhang, K. et al. Early gastric cancer detection and lesion segmentation based on deep learning and gastroscopic images. Sci. Rep. 14, 7847. https://doi.org/10.1038/s41598-024-58361-8 (2024).
Author information
Authors and Affiliations
Contributions
M.E.J. conducted the methodology and programming, wrote the main manuscript text, and prepared the figures. B.G.Z. and C.C.O. were responsible for data acquisition. M.E.J. and O.S.E. conducted data analysis, with O.S.E. performing nailfold capillary visual analysis and M.E.J. conducting deep learning-based analysis. M.E.J., O.S.E., B.G.Z., and A.E. contributed to the result interpretations. M.E.J., O.S.E., B.G.Z., and A.E. reviewed the final draft. A.E. supervised the entire work. All authors have approved the submitted version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ebadi Jalal, M., Emam, O.S., Castillo-Olea, C. et al. Abnormality detection in nailfold capillary images using deep learning with EfficientNet and cascade transfer learning. Sci Rep 15, 2068 (2025). https://doi.org/10.1038/s41598-025-85277-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-85277-8
Keywords
This article is cited by
-
Enhancing nail disease diagnosis: a capsule network with SE attention and dual backbone models
Naunyn-Schmiedeberg's Archives of Pharmacology (2026)