Abstract
Little is known about how exclusive e-cigarette use affects exosomal microRNA (miRNA) expression, which is crucial in inflammation and disease processes like cancer. We compared exosomal miRNA profiles between exclusive e-cigarette users and non-users. We used plasma samples from 15 exclusive e-cigarette users and 15 non-users from the Population Assessment of Tobacco and Health (PATH) Wave 1 study (2013–2014) and sequenced miRNAs with Illumina NextSeq 500/550. We performed differential analyses using DESeq2 in R/Bioconductor, adjusting for race, and conducted gene enrichment analyses on target genes regulated by significant miRNAs. Further, molecular-based techniques using the miRNA mimics and inhibitors were applied for the validation of the expressions of the miRNAs in vitro. We identified four miRNAs that were upregulated in exclusive e-cigarette users compared to non-users: hsa-miR-100-5p, hsa-miR-125a-5p, hsa-miR-125b-5p, and hsa-miR-99a-5p, after adjusting for the confounding effects of race. However, none of the miRNAs remained statistically significant after controlling for the false discovery rate (FDR) at 5%. Subgroup analysis of White participants only identified four miRNAs (hsa-miR-100-5p, hsa-miR-125b-5p, hsa-miR-200b-3p, and hsa-miR-99a-5p) that were also upregulated in e-cigarette users with one miRNA hsa-miR-200b-3p remaining statistical significance after controlling for the FDR at 5%. GO enrichment analysis showed that these miRNAs are involved in processes like transcription regulation and cellular protein modification. KEGG pathway analysis indicated their involvement in cancer pathways, including small cell lung cancer, renal cell carcinoma, and signaling pathways (neurotrophin, ErbB, PI3K-Akt, FoxO, Hippo, MAPK, TGF-beta). Overexpression of hsa-miR-125b-5p promoted DNA damage in bronchial epithelial cells. These findings suggest an elevation of carcinogenic cellular signaling pathways in exclusive e-cigarette users.
Similar content being viewed by others
Introduction
An electronic cigarette (e-cigarette) is a device designed to simulate the feeling of traditional cigarette smoking to vaporize a liquid (often flavored) to generate aerosols for inhalation1. The e-liquid typically contains propylene glycol and/or vegetable glycerin as well as nicotine and flavoring agents/chemicals2. According to the 2023 National Youth Tobacco Survey, 10.0% of high school students and 4.6% of middle school students reported current use of e-cigarettes3. E-cigarette delivers nicotine to users, leading to addiction comparable to or even exceeding that of conventional cigarettes4. While e-cigarettes appear to reduce or eliminate certain carcinogens found in combustible cigarettes, the toxicological effects of inhaling e-cigarette aerosols containing flavorings remain unclear5,6,7.
Animal and cell line studies indicate that e-cigarette aerosols can trigger oxidative stress, cause DNA damage, and induce inflammatory responses in both human lung epithelial cells and mouse lungs8,9,10,11,12,13. Several cross-sectional national survey studies have indicated an association between e-cigarette use and respiratory symptoms and diseases such as wheezing, asthma, and chronic obstructive pulmonary disease (COPD)14,15,16,17,18. Additionally, e-cigarette use is associated with an increased risk of hypertension and cardiovascular diseases, including myocardial infarction, stroke, and coronary artery disease19,20,21,22,23. Mental health problems, depressions, and cognitive impairments have also been linked to e-cigarette use23,24,25,26.
Exosomes are small extracellular vesicles (EVs) released by various cell types, including immune and cancer cell, and are present in biofluids such as blood, urine, and other body fluids27,28. Exosomes play a crucial role in mediating cell-to-cell communication and influencing numerous physiological processes29,30,31,32. Exosomes are enriched with surface proteins, regulatory proteins, mRNAs, microRNAs and long non-coding RNAs33,34. Plasma exosomes, in particular, hold potential as biomarkers for immune dysfunction in cancer35.
MicroRNAs (miRNAs), a class of small non-coding RNAs consisting of 21–25 nucleotides, play vital regulatory roles in various cellular and biological process, including immune regulation and inflammatory responses36,37,38,39. miRNAs have been reported to play pivotal roles in the development of chronic inflammatory lung diseases and lung cancers40,41,42,43,44,45. Exosomal miRNAs, aside from revealing novel toxicological subtypes, show promise as potential biomarkers for flavor-related toxicities. Previous research has demonstrated that tobacco consumption modulates the expression levels of miRNAs through signaling pathways such as apoptosis, angiogenesis and inflammatory pathways46,47,48. These pathways play significant roles in the development of various human diseases, including lung disease, cardiovascular disease, liver disease, and cancer49,50,51. Exosomal microRNAs has emerged as diagnostic biomarkers and therapeutic targets for these diseases. Currently, there is a lack of information on the patterns of genome-wide and locus-specific miRNA expression in plasma samples when the subjects are exposed to e-cigarettes.
This study utilized plasma samples obtained from participants in the Population Assessment of Tobacco and Health (PATH) Study Wave 1 (2013–2014), including 15 exclusive e-cigarette users and 15 non-users52. Our aim is to identify plasma exosomal miRNAs that are differentially expressed between participants who exclusively use e-cigarettes and participants who don’t use any tobacco products. We seek to understand the functions of identified significant miRNAs in various signaling pathways. The identified plasma exosomal miRNAs and their associated targeted genes will significantly contribute to our understanding of the impact of e-cigarettes on cellular signaling pathways.
Materials and methods
Study participants
The PATH study is a nationally representative longitudinal cohort study initiated in September 2013 with 32,320 adults and 13,651 youth (12 years and older) participants52. In addition to collecting epidemiological information on tobacco use behaviors, attitudes, beliefs, exposures, and health, the PATH study includes biospecimens from a subsample of 11,500 adult participants from the Wave 1 study. Blood specimens were exclusively collected from PATH Wave 1 adults and aged-up adults from Wave 1 youth participants.
We obtained plasma samples through the PATH Study Biospecimen Access Program from a cohort of 15 current adult (aged 18 years or older) established exclusive e-cigarette users and 15 adult non-users who were not established any tobacco product users. Current established exclusive e-cig users are defined as individuals who have ever used an e-cigarette, have used regularly, and use every day or some days. Furthermore, these current established exclusive e-cig users do not regularly use any other tobacco products.
The study has been reviewed by the University of Rochester Research Subjects Review Board and was determined as not research involving human subjects (Study ID: STUDY00005750). All biospecimens obtained from the PATH biospecimen program were deidentified, and the linkage ID connecting demographic information with the PATH biospecimens were disclosed only after the completion of sequencing experiments on all biospecimens. All research in this study were performed in accordance with relevant guidelines and regulations.
Sample collection
Biospecimens, including urine and blood, are acquired from PATH Study participants who have provided informed consent, following a protocol reviewed and approved by the Westat Institutional Review Board (IRB) using validated PATH Study standard operating procedures (SOPs)52. The collected biospecimens were transported from the field to the PATH Study biorepository and subsequently processed into aliquots. The PATH Wave 1 blood sample was collected between 2013 and 2014, processed into plasma aliquots with EDTA additives, and has been stored at the PATH Study biorepository since then. Following approval from the PATH Study Biospecimen Access program, we received 30 plasma aliquots (200ul each) via mail in October 2021.
Exosome isolation and RNA extraction
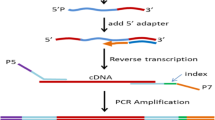
The plasma samples were processed together with library prepared in one batch. Exosome RNA was isolated from 175ul of plasma using Norgen’s plasma exosome purification and RNA Isolation Kit (Cat. 58300). The kit was used according to the manufacturer’s instructions. RNA quantification was then done on an Agilent Bioanalyzer, using the Agilent RNA 6000 Pico Kit kit (cat. 5067 − 1513) according to the manufacturer’s instructions. Small RNA library prep was done using Norgen’s Small RNA Library Prep Kit for Illumina (cat. 63620) according to the manufacturer’s instructions. Library QC was done on an Agilent Bioanalyzer, using the Agilent High Sensitivity DNA Kit (cat. 5067 − 4626).
Small RNA sequencing
The small RNA sequencing analysis was carried out using the Illumina NextSeq 500 sequencing platform with 6ul RNA from each sample. Finally, samples were pooled and sequenced on the Illumina NextSeq 550 platform, using the NextSeq 500/550 High Output Kit v2.5 (75 Cycles). The number of reads from the plasma exosomal miRNAs for the 30 participants ranged from 3,010 to 243,141 in the small RNA sequencing experiment.
Statistical analysis
Unique participant IDs were used to establish links between PATH demographic data and the sequencing outcomes of plasma exosomal miRNAs. Fisher’s exact tests were used to identify potential confounding variables for the e-cigarette user group. miRNAs with a total count below 10 across all samples were excluded from subsequent analyses. The DESeq2 package in Bioconductor (version 3.19) was used to identify statistically significant miRNAs differentially expressed between exclusive e-cigarette users and non-users, while accounting for significant confounding variable. miRNAs with a p-value less than 0.05 were considered statistically significant. The BonEV method was used to control the false discovery rate at 5%53.
To visually represent the results, enhanced volcano plot was used to show the fold change in log2 scales and p-values of these significant miRNAs. Dot plots were used to show the distributions of miRNAs in the e-cigarette and non-user groups. The pheatmap function in the pheatmap package in R (version 4.4.0) was used to generate the heatmap of significant miRNAs. Additionally, a receiver operating characteristic (ROC) curve was constructed using the results of a logistic regression model with predictor variables including significant miRNAs and covariate, employing the rocit function in the ROCit package in R version 4.4.0 (R Core Team, 2019). The area under curve (AUC) of the ROC curve was calculated using the auc function in the pROC package in R (version 4.4.0). For molecular experiments the statistical analysis was performed using Student’s t-test. Data were analyzed using Graphpad prism software (version 7.02).
Gene enrichment analysis
Predictions of gene targets for differentially expressed miRNAs were conducted using the EVmiRNA (http://bioinfo.life.hust.edu.cn/EVmiRNA)54, miRDB (http://mirdb.org/)55,56, and ExoCarta (http://exocarta.org)57,58,59database. The target genes predicted from these databases were subsequently imported into the enrichrichment function within the enrichR package in R60. This facilitated gene enrichment analysis using the Gene Ontology (GO) 2021 and the Kyoto Encyclopedia of Genes and Genomes (KEGG) 2021 databases61,62,63.
Cell culture
Human bronchial epithelial cells (BEAS-2B, ATCC Cat#: CRL-9609) were obtained from the American Type Culture Collection (ATCC) and maintained in DMEM/F12K medium supplemented with 5% fetal bovine serum (FBS) and 1% Penicillin-Streptomycin-Glutamine (Thermo Fisher Scientific Cat#: 103–78016). The cells were incubated at 5% CO₂ and 95% humidity.
BEAS-2B cells (PASSAGE 11–13) were seeded at a density of 30,000 cells/cm² in 6-well plates and allowed to adhere for 24 h. Following this, the cells were starved in 1% FBS medium overnight. After starvation, transfection was performed using Lipofectamine RNAiMAX reagent (Invitrogen, Carlsbad, CA, USA). The treatments applied were Normal/negative control (NC), miR-125b-5p mimics (Cat#: 4464066; Thermo Fisher Scientific), miR-125b-5p inhibitors (Cat#: AM17000; Thermo Fisher Scientific). For miR-125b-5p treatment, the concentrations used were Negative Control: 25 µL Lipofectamine, miRNA Mimic Negative Control: 25 ng/mL in nuclease-free water, miR-125b-5p mimics: 25 ng/mL and 50 ng/mL. After 24 h, cells were exposed to Eliquid (Eliq), PG/VG (50:50), Eliq + miR-125 Mimic, Eliq + miR-125 Inhibitors, PG/VG + miR-125 Mimic, PG/VG + miR-125 Inhibitors. After exposure, cells were either lysed for RNA isolation or fixed for flow-based assays. The dose was referred from literature64,65 .
E-liquid
The e-liquid used in this study was procured from a local vendor and consisted of a PG/VG mixture in a 50:50 ratio. A concentration of 0.25% of the e-liquid was applied to the cells for treatment. This in vitro exposure model aims to investigate the effects of acute exposure on miR-125 and its role in wound healing dysregulation.
Wound healing assay
Confluent BEAS-2B cells were “scratched” by dragging a 200 µL micropipette tip across the center of each well in a single motion. Only wells with cells displaying well-defined edges were included in the study. Immediately following the scratch, images were captured using the Cytation 5 imaging system. The cells were then exposed to e-liquid, PG/VG, and the respective groups of miR-125b-5p mimics and inhibitors. The same scratched area was imaged again at 24 h post-exposure. The wound or scratch area was calculated using ImageJ software.
DNA damage assay
The DNA damage assay was conducted using the FlowCellect™ DNA Damage Histone H2A.X Dual Detection Kit (Cat#: FCCS025153, Merck) according to the manufacturer’s instructions. DNA damage was analyzed using the Luminex Millipore Guava EasyCyte flow cytometer.
Results
Demographic characteristics of the participants providing plasma samples
Table 1 displays the demographic characteristics of the participants who contributed plasma samples for analysis. Ethnicity, gender, and age groups are balanced between the exclusive e-cigarette user and the non-user group with similar frequency distributions. However, there is a significantly difference in the frequency distribution of race, with exclusive e-cigarette users having a higher proportion of White individuals than the non-user group (86% vs. 67%). Consequently, race was included as a covariate in the generalized linear model for miRNA differential analyses.
We also checked the status of other tobacco products used by all 30 participants. No participants had used any other tobacco products (e.g., cigars, cigarillos, pipes filled with tobacco, snus pouches, dissolvable tobacco) except two participants who used hookah for some days. However, the distribution of hookah use was not significantly different between the exclusive e-cigarette users and non-users (P-value = 0.483). The frequency distribution of subjects who ever smoked a cigarette, even one or two puffs, was not significantly different between exclusive e-cigarette users and non-users (P-value = 0.682). Secondhand smoking exposure was also not significantly different between exclusive e-cigarette users and non-users, including those living with a smoker during childhood (P-value = 0.272) and currently living with a smoker (P-value = 1.000).
Identification of differentially expressed miRNAs in exclusive adult electronic cigarette users
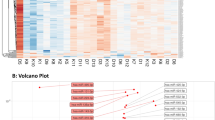
Utilizing a cutoff of two times fold change and a p-value less than 0.05, we identified four significant miRNAs that were differentially expressed between exclusive e-cigarette users and non-users, as illustrated in the volcano plot (Fig. 1). Further examination of the distributions of these four significant miRNAs revealed significantly higher expression levels in the exclusive e-cigarette user group than the non-user group (Fig. 2). The heatmap of the four significant miRNAs, differentially expressed between exclusive e-cigarette users and non-users, highlighted their upregulation in the exclusive e-cigarette user group (Fig. 3). The identified significant miRNAs include hsa-miR-100-5p, hsa-miR-125a-5p, hsa-miR-125b-5p, and hsa-miR-99a-5p. After adjusting for the multiplicity using the BonEV method53 to control for the false discovery rate (FDR), no significant miRNAs remain significant at the 5% level.
Volcano plot showing significant differences in miRNA expression between exclusive e-cigarette users and non-users. The grey dots mean no significant difference (NS) between exclusive e-cigarette users and non-users. The green dots indicate the log2 fold change is beyond the cutoff value (−1 or 1), so the fold change in miRNA expression levels is either larger than 2 or smaller than 0.5. The red dots indicate the miRNA expression levels between exclusive e-cigarette users and non-users are both significant (p-value < 0.05) and have a folder change smaller than 0.5 when comparing non-users with exclusive e-cigarette users. Four miRNAs (hsa-miR-100-5p, hsa-miR-125a-5p, hsa-miR-125b-5p, and hsa-miR-99a-5p) were identified based on the two-fold change and p-value less than 0.05 cutoff.
Identification of differentially expressed miRNAs in exclusive White adult electronic cigarette users
Considering the majority of the participants in our study are White, we conducted a sensitivity analysis to identify differentially expressed miRNAs in exclusive White adult electronic cigarette users. With the same cutoff of two times fold change and a p-value less than 0.05, we identified four significant miRNAs (hsa-miR-100-5p, hsa-miR-125b-5p, hsa-miR-200b-3p, and hsa-miR-99a-5p) that were differentially expressed between exclusive e-cigarette users and non-users in White adults, as illustrated in the volcano plot (S1 Fig). All four miRNAs had higher expression levels in the exclusive White adult electronic cigarette users group, with three of the identified miRNAs overlapping with our previously identified miRNAs using all participants (S2 Fig). S3 Fig shows the heatmap of the four significant miRNAs. One miRNA (hsa-miR-200b-3p) remains significant after the FDR is controlled for at 5% using the BonEV method (adjusted P-value = 0.008).
Sensitivity analysis of miRNA expression profiles between exclusive electronic cigarette users and non-users
We observed that two participants (P000055469 and P000436666) showed much higher miRNA expression levels than others in Fig. 3 and S3, suggesting that they might be potential outliers. To ensure the reliability of our findings, we conducted a sensitivity analysis of the miRNA expression profiles between exclusive electronic cigarette users and non-users, excluding these two participants. In the analysis involving all races, we found three significant miRNAs (hsa-miR-100-5p, hsa-miR-125a-5p, hsa-miR-125b-5p). However, none of them remained significant at the 5% level after adjusting for multiplicity using the BonEV method to control the false discovery rate (FDR). When analyzing only White participants, we identified four significant miRNAs (hsa-miR-100-5p, hsa-miR-125a-5p, hsa-miR-125b-5p, hsa-miR-200b-3p). After controlling for the FDR at 5% using the BonEV method, the miRNA hsa-miR-200b-3p remained significant (adjusted P-value = 0.024). Therefore, we arrived at the same conclusion as when we included the two participants (P000055469 and P000436666), ensuring the robustness of our findings.
Gene enrichment analysis of differentially expressed miRNAs in exosomes of exclusive adult electronic cigarette users
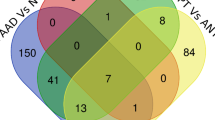
The target genes of the four significantly upregulated miRNAs in exclusive e-cigarette users were predicted using the EVmiRNA (http://bioinfo.life.hust.edu.cn/EVmiRNA), miRDB (http://mirdb.org/), and ExoCarta (http://exocarta.org) databases. The Venn diagrams illustrate the overlap of target genes identified by all three methods for each significant miRNA (Fig. 4). Notably, variations of the number of target genes were observed among different miRNA target gene prediction methods, as well as variations of the number of target genes regulated by different miRNAs. EVmiRNA identified a greater number of gene targets than the other two methods. The union of target genes from all three methods was used for gene enrichment analysis.
Target genes of significant miRNAs differentially expressed between exclusive e-cigarette users and non-users predicted by three different miRNA target prediction methods (EVmiRNA, miRDB, ExoCarta). (A) has-miR-100-5p; (B) has-miR-125a-5p; (C) has-miR-125b-5p; (D) has-miR-99a-5p; (E) has-miR-200b-3p.
Figure 5 showed the gene enrichment analysis results from the GO functional enrichment analysis of significant miRNA target genes and the systematic analysis of target gene functions using the KEGG 2021 Human database. The GO molecular function results highlighted the involvement of significant miRNA target genes in sequence-specific DNA binding, RNA-binding, ubiquitin protein ligase binding, and protein kinase binding. Genes related to intrinsic and integral component of mitochondrial and Golgi membrane were implicated in the GO cellular component analysis. The GO biological process revealed involvement in processes such as extrinsic apoptotic signaling pathway, miRNA catabolic process, miRNA metabolic process, embryonic appendage morphogenesis, and more. The KEGG analysis results identified target genes associated with various cancer pathways, including non-small cell lung cancer, small cell lung cancer, hepatocellular carcinoma, pancreatic cancer, Hippo signaling pathway, HIF-1 signaling pathway, p53 signaling pathway, and MAPK signaling pathway.
Gene enrichment analysis results using identified common target genes of significant miRNAs differentially expressed between exclusive e-cigarette users and non-users predicted by at least one of the three different miRNA target prediction methods (EVmiRNA, miRDB, ExoCarta). (A) GO Molecular Function 2021 database; (B) GO Cellular Component 2021 database; (C) GO Biological Process 2021 database; (D) KEGG 2021 Human database.
Gene enrichment analysis of the significant miRNA has-miR-200b-3p also showed similar GO molecular function (e.g., transcription regulatory region nucleic acid binding, GDP, mRNA, RNA, DNA binding, protein kinase binding and activities), GO cellular components (e.g., intracellular membrane-bounded organelle, adherence junction, cell-cell junction, cell-substrate junction, cytoskeleton), GO biological process (regulation of transcription and gene expression, cellular protein modification process, regulation of intracellular signal transduction and cell differentiation, and protein phosphorylation), and various disease and cancer pathways from the KEGG analysis results (Fig. 6). The miRNA has-miR-200b-3p regulated genes involved in a neurotrophin signaling pathway that regulates neural cell survival, growth, and differentiation, ErbB signaling pathway that regulates cell proliferation, differentiation, migration, apoptosis, and motility, renal cell carcinoma, PI3K-Akt, FoxO, Hippo, MAPK, and TGF-beta signaling pathway, longevity regulating pathway, small cell lung cancer, and prostate cancer.
Gene enrichment analysis results using identified common target genes of significant miRNA hsa-miR-200b-3p differentially expressed between exclusive e-cigarette users and non-users predicted by at least one of the three different miRNA target prediction methods (EVmiRNA, miRDB, ExoCarta). (A) GO Molecular Function 2021 database; (B) GO Cellular Component 2021 database; (C) GO Biological Process 2021 database; (D) KEGG 2021 Human database.
Inhibiting miR-125b-5p impairs the wound-healing ability of bronchial epithelial cells upon exposure to e-liquid
To investigate whether inhibiting miR-125b-5p could impair the wound-healing ability of bronchial epithelial cells upon exposure to e-liquid, we treated the cells with e-liquid and PGVG (control for e-liquid) in combination with the miR-125b mimic and inhibitors. A schematic representation of the workflow is depicted in Fig. 7A. To understand the mechanism of action of the miR-125b mimic, we assessed cell viability. We observed reduced viability following treatment with the miR-125b mimic and the combination of e-liquid and the miR-125b mimic, indicating the effectiveness of the miR-125b mimic (Fig. 7B). Next, we performed a wound healing assay, summarized in Fig. 7C. After transfections, we generated a scratch and treated the cells with e-liquid, PGVG, and their combinations. The combination of miR-125b mimic treatment and e-cig treatment significantly slowed down the migration of bronchial epithelial cells into the wounded area. In contrast, a significant increase in migration was observed following treatment with the inhibitor and PGVG (Fig. 7D).
Inhibition of miR-125b-5p Impairs Wound Healing Mediated by Bronchial Epithelial Cells. A: Schematic representation of the workflow for the treatment of bronchial epithelial cells. Cells were seeded in six-well plates and transfected with miR-125b-5p mimic, inhibitors, or a nonspecific control. Following transfection, the cells were treated with e-liquid and PGVG in combination with the mimic and inhibitor. B: Cell number and viability were assessed using AO/PI staining. C: Schematic of the wound healing assay. D: The scratched wounds were monitored by capturing images under a microscope at 0 h and 24 h post-scratch.Statistical significance was determined as follows: ***p < 0.001 was considered significant compared to the negative control; #p < 0.05 and ##p < 0.001 were considered significant compared to the miR-125b-5p mimic treatment.
Overexpression of miR-125b-5p promotes DNA damage in bronchial epithelial cells
miR-125 is a microRNA known to play a significant role in various cellular processes, including the response to DNA damage. When DNA damage occurs, the expression of certain microRNAs, including miR-125, can be altered as part of the cellular stress response66,67. To assess the effect of the miR-125 mimic during DNA damage, we measured the levels of the DNA damage marker γH2AX following treatments. An enhanced level of DNA damage was observed with the miR-125 mimic and e-liquid treatments, which was alleviated upon treatment with the miR-125 inhibitor. No significant effects were noted in the control group (PGVG) with either the mimic or inhibitor treatment (Fig. 8).
Receiver operating characteristic (ROC) curve for differentiation of exclusive e-cigarette users from non-users
A ROC curve was constructed using the four significant miRNAs differentially expressed between exclusive e-cigarette users and non-users, along with race, to predict whether a participant falls into the category of exclusive e-cigarette users or non-users (Fig. 9). At the optimal Youden Index point, the sensitivity of the four significant miRNA markers and race reached 100%, with a specificity of 60%, effectively distinguishing exclusive e-cigarette users from non-users. The calculation of the Area Under Curve (AUC) yielded a value of 0.8571, indicating a highly reliable predictive capability for determining the e-cigarette use status using the four significant miRNAs and race. Using the four significant miRNAs differentially expressed between exclusive e-cigarette users and non-users within White adults, we found the e-cigarette use status can be predicted reliably from the expression levels of the four miRNAs with AUC value of 0.9167 (S6 Fig).
ROC curve of the four significant miRNAs differentially expressed between exclusive e-cigarette users and non-users and race for prediction of whether the participant is exclusive e-cigarette users or non-users. ROC curve showed that the sensitivity of the four miRNA markers and race for distinguishing non-users from exclusive e-cigarette users can reach 100% and the specificity can reach to 60% based on the optimal Youden Index point. The Area Under Curve (AUC) of the ROC is 0.8571, signifying excellent predictive capability when utilizing the four significant miRNAs in conjunction with race. TRP: true positive rate; FPR: false positive rate.
Discussion
Exosomal miRNAs play pivotal roles in intercellular communication and various biological processes68,69. Dysregulation of exosomal miRNAs could link to tumorigenesis through regulation changes in central cellular process including cell proliferation, cell survival, and apoptosis. Current research has emphasized the potential of exosomal miRNAs as prognostic and diagnostic biomarkers for diseases70. While short-term adverse effects of e-cigarettes on human health have been well-documented, the long-term health effects of e-cigarette use remains unknown due to the relatively short period these products have been on the market71,72. Few studies have examined the impact of e-cigarette use, particularly exclusive e-cigarette use, on exosomal miRNAs.
Using plasma samples from 30 participants in the PATH Wave 1 study, we investigated differences in exosomal miRNA expression levels between exclusive e-cigarette users and non-users. We identified four significant exosomal miRNAs (hsa-miR-100-5p, hsa-miR-125a-5p, hsa-miR-125b-5p, and hsa-miR-99a-5p) that were upregulated in exclusive e-cigarette users compared to non-users, after adjusting the confounding effect from race. However, none of them remain statistically significant after we control the FDR at 5%. A subgroup analysis on White adult participants identified four significant miRNAs (hsa-miR-100-5p, hsa-miR-125b-5p, hsa-miR-200b-3p, and hsa-miR-99a-5p) between exclusive e-cigarette users and non-users and one (hsa-miR-200b-3p) remain statistically significant after controlling the FDR at 5%. Further gene enrichment analysis of predicted target genes regulated by these five significant miRNAs revealed their involvement in various cancer pathways. Our results suggest a pro-carcinogenic cellular signaling pathways affected by exclusive e-cigarette use.
The involvement of hsa-miR-100-5p in numerous pathological processes associated with diseases has been documented. Overexpression of hsa-miR-100-5p in human ovarian endometriotic stromal cells has been linked to the promotion of invasion, contributing to the pathogenesis of endometriosis73. Additionally, hsa-miR-100-5p has been identified as an independent risk factor and a prognostic signature for patients with stomach adenocarcinoma74. Elevated expression levels of hsa-miR-100-5p have been associated with extracapsular extension and poorer survival in patients with oral squamous cell carcinoma compared to their normal counterparts75. In Alzheimer’s disease, hsa-miR-100-5p is known to regulate neuron survival by targeting the Mammalian Target of Rapamycin (mTOR) pathway, a central player in regulating many fundamental cell processes and a critical factor in tumor metabolism76,77,78. The upregulation of hsa-miR-100-5p in exclusive e-cigarette users, when compared to non-users, indicates potential elevated risks of various cancers.
The miRNA has-miR-125a-5p has been recognized as a tumor suppressor in various malignancies, including those affecting the breast, ovary, lung, and central nervous system79,80. It plays a role in cell proliferation through cell cycle regulation and has potential as a therapeutic target for treating squamous cell carcinoma of the head and neck79,80. Prior study has indicated that has-miR-125a-5p can induce apoptosis through a p53-dependent pathway in human lung cancer cells81. Furthermore, its associated with the pathological stage or lymph node metastasis in non-small cell lung cancer has been reported82.
The miRNA hsa-miR-125b-5p has been identified as a regulator of inflammatory genes, targeting MAPKs and NF-kB signaling pathways in human osteoarthritic chondrocytes83,84. Additionally, miR-125b-5p inhibits the expression of TNFR2, demonstrating immunosuppressive activity to enhance the antitumor efficacy in human colon adenocarcinoma patients85. In liver cancer, miR-125b-5p has been shown to inhibit cell proliferation, migration, and invasion86. The miRNA hsa-miR-99a-5p, recognized as a tumor suppressor in tumors like bladder cancer and breast cancer, suppresses cell proliferation, migration, and invasion87. Interestingly, a previous study found that the hsa-miR-99a-5p expression level in breast cancer tissues were significantly lower than healthy breast tissue, while the expression level in plasma samples were significantly higher in breast cancer patients than in healthy controls88.
Although has-miR-125a-5p, miR-125b-5p, and hsa-miR-99a-5p are known tumor suppressors, it remains uncertain whether the upregulation of these exosomal miRNAs in plasma samples indicates an elevated risk in exclusive e-cigarette users. The experimental validation assay showed that overexpression of miRNA hsa-miR-125b-5p along with e-Liquid treatments, slowed wound healing and contributed to DNA damage in bronchial epithelial cells. miR-125b is one miRNA among the freely circulating microRNAs and no significant differences were found between plasma miRNAs and plasma-derived exosomal miRNAs89. Further research is necessary to evaluate the role of exosomal miRNAs in the association between exclusive e-cigarette use and adverse health effects.
Sensitivity analysis of the miRNA expression levels between exclusive e-cigarette users and non-users within White adults identified three overlapped miRNAs (hsa-miR-100-5p, hsa-miR-125b-5p, and hsa-miR-99a-5p). This indicated the importance of these three miRNAs for all races, although they are not statistically significant after controlling the FDR at 5% due to the relatively small sample size. The extra miRNA identified in the White adult only (hsa-miR-200b-3p) remains significant after adjusting the multiple testing error rate to control the FDR at 5%. Gene enrichment analysis of all identified miRNAs showed their involvement in cancer-related pathways, which indicates the potential harm of exclusive e-cigarette use on human health.
Limited research has explored the impact of e-cigarette use on exosomal miRNA profiles. A prior study in 2019 investigated plasma exosomal miRNA expression levels, revealing a different set of miRNAs significantly differentially expressed between exclusive e-cigarette users and non-smokers47. This variation may stem from differences in e-liquid and e-cigarette devices. Our samples were collected during the PATH Wave 1 study from 2013 to 2014, aligning with the era of the first generation of e-cigarette devices90. During this period, most e-cigarettes were usually disposable single units containing natural nicotine in the e-liquid. In contrast, the predominant e-cigarette devices on the market in 2019 were fourth generation devices, largely disposable with nicotine salt in the e-liquid to mitigate nicotine harshness90. Given the large differences in e-liquid and e-cigarette devices, it is highly plausible that distinct exosomal miRNAs are influenced by e-cigarette use.
The activity of microRNA 125b and 100 was found to be differentially expressed in the participant cohort of e-cigarette users. We wanted to study the impact of these microRNAs on bronchial epithelial cells, which are considered to be one of the main targets when inhaling e-cigarettes. Prolonged inhalation of e-cigarettes may directly or indirectly cause dysfunction in the airway epithelial barrier, which could then potentially allow respiratory pathogens to invade, leading to the development and progression of respiratory comorbidities. This may also activate an indirect mechanism of signaling cascades, such as mediating paracrine and endocrine pathways. Our future research will focus on investigating the mechanisms of how these microRNAs interact with their targets to affect pro-carcinogenic cellular signaling pathways leading to damaging/injurious responses in exclusive e-cigarette users and subjects who have been exposed to e-cigarette use.
Our current study has several limitations. First, no biomarker of exposure (BOE) data is available for the 30 participants. The absence of BOE data limited our ability to verify the accuracy of the self-reported tobacco use history data, which might bring potential bias to our analysis results. Second, the relatively small sample size of 30 participants limited the power to identify more significant miRNAs associated with exclusive e-cigarette use. Third, our study lacked a positive control group for cigarette use, and we could not assess the expression profiles of those significant miRNAs in smokers. Considering the ample evidence of less toxicant exposure and potential adverse health effects of e-cigarettes compared to cigarette smoking, we anticipate that the changes in miRNA expression levels from exclusive e-cigarette users will be less pronounced than those in smokers, as e-cigarette users have a significantly lower risk than smokers91,92,93,94. Finally, with the evolution of e-cigarette products, results from our current study might not be applicable to current e-cigarette users, given the differences in nicotine form and devices.
Conclusions
In summary, our analysis of blood samples collected during the PATH Wave 1 study from 2013 to 2014 identified five plasma exosomal miRNAs including hsa-miR-100-5p, hsa-miR-125a-5p, hsa-miR-125b-5p, hsa-miR-200b-3p, and hsa-miR-99a-5p. All five miRNAs were upregulated in exclusive e-cigarette users compared to non-users. Gene enrichment analysis of predicted target genes regulated by these five significant miRNAs uncovered numerous target genes associated with cancer-related signaling pathways. Overexpression of miRNA hsa-miR-125b-5p can promote DNA damage in bronchial epithelial cells, suggesting an activation of pro-carcinogenic signaling pathways. To validate these results, further investigations into the impact of exclusive e-cigarette use on plasma exosomal miRNAs and their correlation with cancer risks are warranted. The AUC of the ROC curve indicated a robust predictive ability of the four significant miRNAs when combined with race in distinguishing exclusive e-cigarette users from non-users.
Data availability
The datasets generated and analyzed during the current study are available in the Gene Expression Omnibus (GEO) repository with access number GSE253603 [https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi? acc=GSE253603]. The code used for the microRNA sequencing data analysis could be found from the GitHub website: https://github.com/DongmeiLi2017/plasma-exosomal-microRNA/tree/main.
Abbreviations
- E-cigarette:
-
Electronic cigarette
- EV:
-
Extracellular vesicles
- PATH:
-
Population Assessment of Tobacco and Health
- miRNA:
-
microRNA
- IRB:
-
Institutional Review Board
- SOP:
-
Standard operating procedures
- GO:
-
Gene Ontology
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- ROC:
-
Receiver operating characteristic
- AUC:
-
Area under the curve
References
Kaisar, M. A. et al. A decade of e-cigarettes: limited research & unresolved safety concerns. Toxicology 365, 67–75 (2016).
Lestari, K. S., Humairo, M. V. & Agustina, U. Formaldehyde Vapor Concentration in Electronic Cigarettes and Health Complaints of Electronic Cigarettes Smokers in Indonesia. J Environ Public Health, 2018: p. 9013430. (2018).
Birdsey, J. et al. Tobacco Product Use among U.S. Middle and High School Students - National Youth Tobacco Survey, 2023. MMWR Morb Mortal. Wkly. Rep. 72 (44), 1173–1182 (2023).
Jankowski, M. et al. E-Cigarettes are more addictive than traditional Cigarettes-A study in highly educated Young people. Int. J. Environ. Res. Public. Health, 16(13), 2279 (2019). https://doi.org/10.3390/ijerph16132279
Lerner, C. A. et al. Vapors produced by electronic cigarettes and e-juices with flavorings induce toxicity, oxidative stress, and inflammatory response in lung epithelial cells and in mouse lung. PLoS One. 10 (2), e0116732 (2015).
Chun, L. F. et al. Pulmonary toxicity of e-cigarettes. Am. J. Physiol. Lung Cell. Mol. Physiol. 313 (2), L193–L206 (2017).
Shields, P. G. et al. A review of pulmonary toxicity of electronic cigarettes in the context of smoking: a focus on inflammation. Cancer Epidemiol. Biomarkers Prev. 26 (8), 1175–1191 (2017).
Dimitrov, S., Hulteng, E. & Hong, S. Inflammation and exercise: Inhibition of monocytic intracellular TNF production by acute exercise via β2-adrenergic activation. Brain, Behavior, and Immunity, 61: pp. 60–68. (2017).
Gerloff, J. et al. Inflammatory response and barrier dysfunction by different e-Cigarette Flavoring Chemicals identified by Gas Chromatography-Mass Spectrometry in e-Liquids and e-Vapors on Human Lung epithelial cells and fibroblasts. Appl. Vitro Toxicol. 3 (1), 28–40 (2017).
Muthumalage, T. et al. Inflammatory and oxidative responses Induced by exposure to commonly used e-Cigarette flavoring chemicals and flavored e-Liquids without Nicotine. Front. Physiol. 8, 1130 (2017).
Solleti, S. K. et al. MicroRNA expression profiling defines the impact of electronic cigarettes on human airway epithelial cells. Sci. Rep. 7 (1), 1081 (2017).
Kaur, G., Muthumalage, T. & Rahman, I. Mechanisms of toxicity and biomarkers of flavoring and flavor enhancing chemicals in emerging tobacco and non-tobacco products. Toxicol. Lett. 288, 143–155 (2018).
Sundar, I. K. et al. Genetic ablation of histone deacetylase 2 leads to lung cellular senescence and lymphoid follicle formation in COPD/emphysema. FASEB J. 32 (9), 4955–4971 (2018). https://doi.org/10.1096/fj.201701518R
Li, D. & Xie, Z. Cross-sectional Association of Lifetime Electronic Cigarette use with wheezing and related respiratory symptoms in U.S. adults. Nicotine Tob. Res. 22 (Suppl 1), S85–S92 (2020).
Li, D. et al. Association of smoking and electronic cigarette use with wheezing and related respiratory symptoms in adults: cross-sectional results from the Population Assessment of Tobacco and Health (PATH) study, wave 2. Tob. Control. 29 (2), 140–147 (2020).
Schweitzer, R. J. et al. E-cigarette use and asthma in a multiethnic sample of adolescents. Prev. Med. 105, 226–231 (2017).
Osei, A. D. et al. The association between e-cigarette use and asthma among never combustible cigarette smokers: behavioral risk factor surveillance system (BRFSS) & 2017. Bmc Pulmonary Medicine, 2019. 19(1). (2016).
Xie, Z. et al. Use of Electronic cigarettes and Self-reported chronic obstructive Pulmonary Disease diagnosis in adults. Nicotine Tob. Res. 22 (7), 1155–1161 (2020).
Shi, H. et al. Sex Differences in the Association of E-cigarette and Cigarette Use and Dual Use with Self-reported Hypertension Incidence in US Adults (Nicotine Tob Res, 2022).
Miller, C. R. et al. Cross-sectional associations of Smoking and E-cigarette Use with Self-reported diagnosed hypertension: findings from Wave 3 of the Population Assessment of Tobacco and Health Study. Toxics, 9(3), 52 (2021). https://doi.org/10.3390/toxics9030052
Mohammadi, L. et al. Chronic E-Cigarette Use impairs endothelial function on the physiological and Cellular levels. Arterioscler. Thromb. Vascular Biology. 42 (11), 1333–1350 (2022).
Vindhyal, M. R. et al. Cardiovascular outcomes Associated with Adult Electronic cigarette use. Cureus J. Med. Sci., 12(8), e9618 (2020). https://doi.org/10.7759/cureus.9618
Vindhyal, M. R. et al. Impact on Cardiovascular outcomes among E-Cigarette users: a review from National Health interview surveys. J. Am. Coll. Cardiol. 73 (9), 11–11 (2019).
Grant, J. E. et al. E-cigarette use (vaping) is associated with illicit drug use, mental health problems, and impulsivity in university students. Ann. Clin. Psychiatry. 31 (1), 27–35 (2019).
Xie, C., Xie, Z. & Li, D. Association of electronic cigarette use with self-reported difficulty concentrating, remembering, or making decisions in US youth. Tob. Induc. Dis. 18, 106 (2020).
Xie, Z. et al. Electronic cigarette use and subjective cognitive complaints in adults. PLoS One. 15 (11), e0241599 (2020).
Thakur, A. et al. Exosomes: small vesicles with big roles in cancer, vaccine development, and therapeutics. Bioactive Mater. 10, 281–294 (2022).
Xie, F. et al. Extracellular vesicles in Cancer Immune Microenvironment and Cancer Immunotherapy. Adv. Sci., 6(24), 1901779 (2019). https://doi.org/10.1002/advs.201901779
Nolte-‘t Hoen, E. et al. Extracellular vesicles and viruses: are they close relatives? Proc. Natl. Acad. Sci. U.S.A. 113 (33), 9155–9161 (2016).
Balducci, E. et al. Extracellular vesicles from T cells overexpress miR-146b-5p in HIV-1 infection and repress endothelial activation. Sci. Rep. 9 (1), 10299 (2019). https://doi.org/10.1038/s41598-019-44743-w
Han, L., Lam, E. W. F. & Sun, Y. Extracellular vesicles in the tumor microenvironment: old stories, but new tales. Mol. Cancer 18 (1), 59 (2019). https://doi.org/10.1186/s12943-019-0980-8
Li, N. et al. Recent advances of exosomes in immune-mediated eye diseases. Stem Cell Res. Ther., 10 (1), 278 (2019). https://doi.org/10.1186/s13287-019-1372-0
Seo, Y., Kim, H. S. & Hong, I. S. Stem Cell-Derived Extracellular Vesicles as Immunomodulatory Therapeutics. Stem Cells Int, 2019: p. 5126156. (2019).
Kaur, G. et al. Differential plasma exosomal long non-coding RNAs expression profiles and their emerging role in E-cigarette users, cigarette, waterpipe, and dual smokers. PLoS One. 15 (12), e0243065 (2020).
Whiteside, T. L. The emerging role of plasma exosomes in diagnosis, prognosis and therapies of patients with cancer. Contemp. Oncol. (Pozn). 22 (1A), 38–40 (2018).
Shivdasani, R. A. MicroRNAs: regulators of gene expression and cell differentiation. Blood 108 (12), 3646–3653 (2006).
Ambros, V. & Chen, X. The regulation of genes and genomes by small RNAs. Development 134 (9), 1635–1641 (2007).
Wahid, F. et al. MicroRNAs: synthesis, mechanism, function, and recent clinical trials. Biochim. Biophys. Acta. 1803 (11), 1231–1243 (2010).
Yang, P. et al. Identification of key microRNAs and Genes Associated with Abdominal Aortic Aneurysm Based on the gene Expression Profile (Exp Physiol, 2019).
Zhang, W., Liu, J. & Wang, G. The role of microRNAs in human breast cancer progression. Tumour Biol. 35 (7), 6235–6244 (2014).
Peng, Y. & Croce, C. M. The role of MicroRNAs in human cancer. Signal. Transduct. Target. Ther. 1, 15004 (2016).
Zheng, Q. et al. Prognostic role of microRNAs in human gastrointestinal cancer: a systematic review and meta-analysis. Oncotarget 8 (28), 46611–46623 (2017).
Lamichhane, S. R. et al. Prognostic Role of MicroRNAs in Human Non-Small-Cell Lung Cancer: A Systematic Review and Meta-Analysis. Dis Markers, 2018: p. 8309015. (2018).
Sundar, I. K., Li, D. & Rahman, I. Small RNA-sequence analysis of plasma-derived extracellular vesicle miRNAs in smokers and patients with chronic obstructive pulmonary disease as circulating biomarkers. J. Extracell. Vesicles. 8 (1), 1684816 (2019).
Kaur, G. et al. Distinct exosomal miRNA profiles from BALF and Lung tissue of COPD and IPF patients. Int. J. Mol. Sci., 22(21), 11830 (2021). https://doi.org/10.3390/ijms222111830
Willinger, C. M. et al. MicroRNA signature of cigarette smoking and evidence for a putative causal role of MicroRNAs in Smoking-related inflammation and target organ damage. Circ. Cardiovasc. Genet., 10(5), e001678 (2017). https://doi.org/10.1161/CIRCGENETICS.116.001678
Singh, K. P. et al. Exosomal microRNAs are novel circulating biomarkers in cigarette, waterpipe smokers, E-cigarette users and dual smokers. BMC Med. Genomics. 13 (1), 128 (2020).
Karabegovic, I. et al. Smoking-related dysregulation of plasma circulating microRNAs: the Rotterdam study. Hum. Genomics. 17 (1), 61 (2023).
Soares do Amaral, N. et al. Noncoding RNA profiles in Tobacco- and Alcohol-Associated diseases. Genes (Basel), 8(1), 6. (2016). https://doi.org/10.3390/genes8010006
Li, Y. et al. The roles of exosomal miRNAs and lncRNAs in lung diseases. Signal. Transduct. Target. Ther. 4, 47 (2019).
Xie, Y. et al. The role of exosomal noncoding RNAs in cancer. Mol. Cancer. 18 (1), 37 (2019).
Hyland, A. et al. Design and Methods of the Population Assessment of Tobacco and Health (PATH) Study (Tobacco Control, 2016).
Li, D. et al. Bon-EV: an improved multiple testing procedure for controlling false discovery rates. BMC Bioinform. 18 (1), 1 (2017).
Liu, T. et al. EVmiRNA: a database of miRNA profiling in extracellular vesicles. Nucleic Acids Res. 47 (D1), D89–D93 (2019).
Chen, Y. & Wang, X. miRDB: an online database for prediction of functional microRNA targets. Nucleic Acids Res. 48 (D1), D127–D131 (2020).
Liu, W. & Wang, X. Prediction of functional microRNA targets by integrative modeling of microRNA binding and target expression data. Genome Biol. 20 (1), 18 (2019).
Keerthikumar, S. et al. ExoCarta: a web-based compendium of Exosomal Cargo. J. Mol. Biol. 428 (4), 688–692 (2016).
Mathivanan, S. et al. ExoCarta 2012: database of exosomal proteins, RNA and lipids. Nucleic Acids Res. 40 (Database issue), D1241–D1244 (2012).
Mathivanan, S. & Simpson, R. J. ExoCarta: a compendium of exosomal proteins and RNA. Proteomics 9 (21), 4997–5000 (2009).
Kuleshov, M. V. et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 44 (W1), W90–W97 (2016).
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28 (1), 27–30 (2000).
Kanehisa, M. Toward understanding the origin and evolution of cellular organisms. Protein Sci. 28 (11), 1947–1951 (2019).
Kanehisa, M. et al. KEGG for taxonomy-based analysis of pathways and genomes. Nucleic Acids Res. 51 (D1), D587–D592 (2023).
Zhang, X. et al. The expression of miR-125b in Nrf2-silenced A549 cells exposed to hyperoxia and its relationship with apoptosis. J. Cell. Mol. Med. 24 (1), 965–972 (2020).
Jiang, L. et al. Upregulation of endothelial cell-derived exosomal microRNA-125b-5p protects from sepsis-induced acute lung injury by inhibiting topoisomerase II alpha. Inflamm. Res. 70 (2), 205–216 (2021).
Hu, H. & Gatti, R. A. MicroRNAs: new players in the DNA damage response. J. Mol. Cell. Biol. 3 (3), 151–158 (2011).
Liu, G. et al. Silencing miR-125b-5p attenuates inflammatory response and apoptosis inhibition in mycobacterium tuberculosis-infected human macrophages by targeting DNA damage-regulated autophagy modulator 2 (DRAM2). Cell. Cycle. 19 (22), 3182–3194 (2020).
Nail, H. M. et al. Exosomal miRNA-mediated intercellular communications and immunomodulatory effects in tumor microenvironments. J. Biomed. Sci. 30 (1), 69 (2023).
Hu, G., Drescher, K. M. & Chen, X. M. Exosomal miRNAs: Biological properties and therapeutic potential. Front. Genet. 3, 56 (2012).
Negrete-Garcia, M. C. et al. Exosomal Micro-RNAs as intercellular communicators in idiopathic pulmonary fibrosis. Int. J. Mol. Sci., 23(19), 11047. (2022). https://doi.org/10.3390/ijms231911047
Daynard, R. Public health consequences of e-cigarettes: a consensus study report of the National Academies of Sciences, Engineering, and Medicine. J. Public Health Policy. 39 (3), 379–381 (2018).
Prochaska, J. J. The public health consequences of e-cigarettes: a review by the National Academies of Sciences. A call for more research, a need for regulatory action. Addiction 114 (4), 587–589 (2019).
Takebayashi, K. et al. Hsa-miR-100-5p, an overexpressed miRNA in human ovarian endometriotic stromal cells, promotes invasion through attenuation of SMARCD1 expression. Reprod. Biol. Endocrinol. 18 (1), 31 (2020).
Wang, G. et al. Comprehensive Analysis of the Prognostic significance of hsa-mir-100-5p and its related gene signature in stomach adenocarcinoma. Front. Cell. Dev. Biol. 9, 736274 (2021).
Jakob, M. et al. MicroRNA expression patterns in oral squamous cell carcinoma: hsa-mir-99b-3p and hsa-mir-100-5p as novel prognostic markers for oral cancer. Head Neck. 41 (10), 3499–3515 (2019).
Ye, X. et al. MicroRNAs 99b-5p/100-5p regulated by endoplasmic reticulum stress are involved in Abeta-Induced pathologies. Front. Aging Neurosci. 7, 210 (2015).
Saxton, R. A. & Sabatini, D. M. mTOR Signaling in Growth, Metabolism, and Disease. Cell 169 (2), 361–371 (2017).
Zou, Z. et al. mTOR signaling pathway and mTOR inhibitors in cancer: progress and challenges. Cell. Biosci. 10, 31 (2020).
Nishida, N. et al. MicroRNA-125a-5p is an independent prognostic factor in gastric cancer and inhibits the proliferation of human gastric cancer cells in combination with trastuzumab. Clin. Cancer Res. 17 (9), 2725–2733 (2011).
Vo, D. T. et al. miR-125a-5p functions as tumor suppressor microRNA and is a marker of Locoregional recurrence and poor prognosis in Head and Neck Cancer. Neoplasia 21 (9), 849–862 (2019).
Jiang, L. et al. MicroRNA HSA-miR-125a-5p induces apoptosis by activating p53 in lung cancer cells. Exp. Lung Res. 37 (7), 387–398 (2011).
Jiang, L. et al. Hsa-miR-125a-3p and hsa-miR-125a-5p are downregulated in non-small cell lung cancer and have inverse effects on invasion and migration of lung cancer cells. BMC Cancer. 10, 318 (2010).
Rasheed, Z. et al. Author correction: MicroRNA-125b-5p regulates IL-1beta induced inflammatory genes via targeting TRAF6-mediated MAPKs and NF-kappaB signaling in human osteoarthritic chondrocytes. Sci. Rep. 9 (1), 14729 (2019).
Rasheed, Z. et al. MicroRNA-125b-5p regulates IL-1beta induced inflammatory genes via targeting TRAF6-mediated MAPKs and NF-kappaB signaling in human osteoarthritic chondrocytes. Sci. Rep. 9 (1), 6882 (2019).
Jiang, M. et al. MiR-125b-5p modulates the function of regulatory T cells in tumor microenvironment by targeting TNFR2. J. Immunother Cancer, 10(11), e005241 (2022). https://doi.org/10.1136/jitc-2022-005241
Hua, S. et al. miR-125b-5p inhibits cell proliferation, migration, and invasion in hepatocellular carcinoma via targeting TXNRD1. Cancer Cell. Int. 19, 203 (2019).
Sun, X. & Yan, H. MicroRNA-99a-5p suppresses cell proliferation, migration, and invasion by targeting isoprenylcysteine carboxylmethyltransferase in oral squamous cell carcinoma. J. Int. Med. Res. 49 (5), 300060520939031 (2021).
Garrido-Cano, I. et al. Circulating miR-99a-5p expression in plasma: a potential biomarker for early diagnosis of breast Cancer. Int. J. Mol. Sci., 21(19), 7427 (2020). https://doi.org/10.3390/ijms21197427
Tian, F. et al. No significant difference between plasma miRNAs and plasma-derived exosomal miRNAs from healthy people. Biomed. Res. Int. 2017, p1304816 (2017).
Li, D. & Xie, Z. E-cigarettes and Vaping, in Reference Module in Biomedical Sciences (Elsevier, 2023).
Glantz, S. A. & Bareham, D. W. E-Cigarettes: use, effects on Smoking, Risks, and Policy implications. Annu. Rev. Public. Health. 39, 215–235 (2018).
Marques, P., Piqueras, L. & Sanz, M. J. An updated overview of e-cigarette impact on human health. Respir Res. 22 (1), 151 (2021).
Balfour, D. J. K. et al. Balancing consideration of the risks and benefits of E-Cigarettes. Am. J. Public. Health. 111 (9), 1661–1672 (2021).
Holt, N. M. et al. Comparison of biomarkers of exposure among US adult smokers, users of electronic nicotine delivery systems, dual users and nonusers, 2018–2019. Sci. Rep. 13 (1), 7297 (2023).
Hyland, A. et al. Design and methods of the Population Assessment of Tobacco and Health (PATH) study. Tob. Control. 26 (4), 371–378 (2017).
Acknowledgements
The authors would like to thank Md Faizan Imam, PhD, and Calla Fahey for their technical help during the revision of the manuscript.
Funding
Research reported in this publication was supported by the National Institute of Environmental Health Sciences of the National Institutes of Health under Award Number R21ES032159, the Western New York–based Center for Research on Flavored Tobacco Products under cooperative agreement U54CA228110 from the National Cancer Institute of the National Institutes of Health and the US Food and Drug Administration, and the National Institute on Aging under Award Number U54AG075931. This work was also partly supported by the University of Rochester’s Clinical and Translational Science Award (CTSA) number UL1 TR002001 from the National Center for Advancing Translational Sciences of the National Institutes of Health. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
DL, ZX, and IR designed the study. DL obtained data from PATH Biospecimen Access Program. DL analyzed the exosomal miRNA sequencing data. SS conducted the wound healing assay and DNA damage assay. DL wrote the original manuscript. DL, ZX, SS, and IR reviewed and edited the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
The PATH Study design and protocol was approved by the Westat Institutional Review Board to protect the rights, welfare, and well-being of all humans involved in this study. All participants provided verbal informed consent before participating in the PATH Study95. Using validated PATH Study standard operating procedures, blood biospecimens were collected from consenting PATH Study participants under a protocol reviewed and approved by the Westat Institutional Review Board. Our applications to the collected plasma biospecimens have been approved by the PATH Study Biospecimen Access Program. The study has been reviewed by the University of Rochester Research Subjects Review Board and was determined as not research involving human subjects. All biospecimens obtained from the PATH biospecimen program were deidentified, and the linkage ID connecting demographic information with the PATH biospecimens were disclosed only after the completion of sequencing experiments on all biospecimens.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, D., Xie, Z., Shaikh, S.B. et al. Altered expression profile of plasma exosomal microRNAs in exclusive electronic cigarette adult users. Sci Rep 15, 2714 (2025). https://doi.org/10.1038/s41598-025-85373-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-85373-9