Abstract
This paper introduces a novel method for spleen segmentation in ultrasound images, using a two-phase training approach. In the first phase, the SegFormerB0 network is trained to provide an initial segmentation. In the second phase, the network is further refined using the Pix2Pix structure, which enhances attention to details and corrects any erroneous or additional segments in the output. This hybrid method effectively combines the strengths of both SegFormer and Pix2Pix to produce highly accurate segmentation results. We have assembled the Spleenex dataset, consisting of 450 ultrasound images of the spleen, which is the first dataset of its kind in this field. Our method has been validated on this dataset, and the experimental results show that it outperforms existing state-of-the-art models. Specifically, our approach achieved a mean Intersection over Union (mIoU) of 94.17% and a mean Dice (mDice) score of 96.82%, surpassing models such as Splenomegaly Segmentation Network (SSNet), U-Net, and Variational autoencoder based methods. The proposed method also achieved a Mean Percentage Length Error (MPLE) of 3.64%, further demonstrating its accuracy. Furthermore, the proposed method has demonstrated strong performance even in the presence of noise in ultrasound images, highlighting its practical applicability in clinical environments.
Similar content being viewed by others
Introduction
Diseases that affect the size of the spleen, such as splenomegaly, are significant clinical issues for several medical disciplines. Splenomegaly, defined as the enlargement of the spleen with respect to its standard size, can be due to various pathological causative agents: infectious diseases, such as malaria and mononucleosis; hematologic disorders, including leukemia and lymphoma; inflammatory affections, such as rheumatoid arthritis; and hepatic diseases, including cirrhosis. Accurate assessment of spleen size is essential in diagnosing and monitoring the disease, as splenomegaly is often a marker of both disease severity and progression. Also, since the accurate dimension of the spleen is essential, it is imperative for guiding appropriate treatment strategies and virtually monitoring treatment efficiency1.
Segmentation in ultrasound images plays a crucial role in an accurate measurement of the size of the spleen. The measurement can further help the physician to diagnose the spleen with its associated medical conditions. Automated segmentation, and more specifically segmentation powered by Deep Learning (DL) methodologies, has proven to be reliable when it comes to providing efficient and accurate delineation of the spleen for ultrasound images. This fully automated segmentation is very important toward the determination of splenic volume with great accuracy, as it allows one to detect spleen volumes that are abnormal and hence support clinical diagnosis2.
Several approaches have been presented using deep learning for automated splenic segmentation. These methods can quantify the volume of the spleen and can assess splenomegaly accurately. For example, Perez et al.3 demonstrated the effectiveness of deep learning-based approaches for spleen segmentation, highlighting the ability to provide quick and precise segmentation when contrasted with classical ones.
The use of artificial intelligence methods comes with immense benefits for disease diagnosis, especially in spleen images. These methods reduce the tedium of diagnosis because they mainly automate tasks, such as image segmentation processes that take a lot of analysts/evaluators. Besides, artificial intelligence algorithms can accurately analyze large quantities of data received from medical imaging, thus diagnosing and appropriately intervening through measures4.
Besides, AI-based diagnostic tools are exact and consistent ways of finding answers compared to usual methods. AI diagnostic tools, with the help of in-built advanced algorithms and machine learning techniques, can catch patterns in medical images and facts that generally go unnoticed as per human intervention. Due to this, the diagnostic reports generated are quite reliable and, therefore, can lead to a better outcome and more functional treatment by healthcare providers5. Despite these advancements, spleen segmentation in ultrasound images remains challenging due to the low contrast and noise inherent in ultrasound imaging ,anatomical variability of the spleen across patients and lack of spleen dataset. These challenges necessitate robust methods to ensure accurate and reliable segmentation results, particularly in clinical applications.
Our research proposes a novel method of segmenting ultrasound images of the spleen. Our proposed method consists of using a combination of SegFormerB0 and Pix2Pix. The proposed method corrects the lack of existing approaches, proposing a more efficient and accurate alternative for segmenting the spleen. The proposed method removes the noises in the segmentation and gives us a unified and better structure of the predicted mask.
Secondly, we have collected a dataset, the Spleenex dataset. This dataset consists of 450 ultrasound images and is used as a valuable tool in training and testing our segmentation model. The Spleenex dataset is a set of annotated images that guarantee accurate and reliable ground truth labels of the spleen. The Spleenex dataset is the first public dataset published in the field of Image Segmentation in spleen images.
We summarize our contributions as follows:
-
We Propose a novel method for ultrasound image segmentation of the spleen by leveraging a custom combination of SegFormerB0 and Pix2Pix networks to improve the quality of extracted segmentations.
-
We collected a new dataset, named here the Spleenex dataset, which consists of 450 labeled ultrasound images of the spleen. The Spleenex dataset is the first public dataset published in spleen image segmentation.
-
Extensive experiments and comparisons against state-of-the-art segmentation models in the spleen image to demonstrate its effectiveness and efficiency in our proposed method.
Related works
Image segmentation
The development and improvement of segmentation methodologies with neural networks have achieved rapid progress. U-Net laid the first stone6, which utilized an unpadded convolution and pixel-wise soft-max function for segmentation in biomedical images. DeepLabV37 used mechanisms like atrous spatial pyramid pooling (ASPP), capturing multi-scale contextual information, which significantly improved the techniques of semantic segmentation.
U-Net++8 further improved the segmentation architectures with increased feature aggregation and more expansive receptive fields. Attention U-Net9 embedded attention mechanisms to increase feature representation in an enriched way and was able to capture long-range dependencies, hence providing an increase in segmentation accuracy. OCRNet (Object-contextual Representations Network)10 put a significant emphasis on context awareness in the segmentation tasks and further advanced segmentation capabilities in the semantic segmentation landscape.
HRFormer (High-Resolution Transformer)11 learns hierarchical representations by transformer-based frameworks and performs much better in complex segmentation tasks. SegFormer12 has a highly efficient transformer architecture and is thus a move toward more efficient segmentation models. Finally, Seaformer (Squeeze-Enhanced Axial Transformer)13 is one of the latest examples of continuous innovation in segmentation architectures.
Building on these advancements, recent methods have targeted the challenges of ultrasound image segmentation, such as blurred boundaries and tumor heterogeneity. ESKNet14, incorporating enhanced kernel convolutions with attention mechanisms, achieved superior Dice scores (up to 84.76%) on datasets like BUSI, the breast ultrasound dataset, and STU. Similarly, NU-Net,15 a nested U-net with shared-weight architecture, delivered strong performance on BUSI and the breast ultrasound dataset. Meanwhile, C-Net16 combines U-net with bidirectional attention and residual refinement, excelling in segmentation tasks, particularly on the BUSIS dataset.
Spleen segmentation
The field of spleen segmentation in medical imaging has undergone significant advancements, particularly with the adoption of deep-learning methodologies. A noteworthy study by17 focused on diagnosing hepatic fibrosis, utilizing an Artificial Neural Network (ANN) to predict liver cirrhosis non-invasively. This study showcased promising implications for clinical decision-making in chronic liver disease. In this paper, spleen size measurement was used to diagnose the disease.
A seminal contribution to this domain is the SSNet (Splenomegaly Segmentation Network), as detailed in18, which employs a conditional generative adversarial network (cGAN)19. This model, comprising a generator and discriminator, demonstrated substantial progress in spleen segmentation, surpassing benchmarks set by previous models such as GCN and U-Net. SSNet achieved a mean Dice coefficient of 0.9260, with particular attention to addressing spatial variations in spleen size and shape, thereby reducing false positives and negatives. However, limitations were observed in its 2D segmentation approach, potentially compromising the handling of 3D spatial data.
In a related endeavor,2 introduced a modified U-Net for spleen segmentation and direct regression to predict spleen length. Accompanied by a quality control (QC) model aimed at filtering out poor-quality images to enhance accuracy, this approach substantially reduced the mean percentage length error (MPLE) to 4.58%, showcasing marked improvement with QC implementation. The research primarily focused on precise spleen length measurements using ultrasound images.
Additionally,20 utilized a U-Net architecture for spleen segmentation, incorporating deep regression models to achieve a 7.42% length error comparable to human expert variability in estimating spleen length. Their study centered on 2D ultrasound images from pediatric Sickle Cell Disease patients. Departing from segmentation,4 introduced a model based on the MobileNetV2 architecture, surpassing individual doctors’ diagnoses in classifying splenic trauma. The model exhibited heightened sensitivity and specificity when compared to clinical diagnoses, utilizing animal models and clinical ultrasonic images from multiple centers. Expanding the scope,4 developed a dual-channel Convolutional Neural Network (CNN) specifically designed to enhance ultrasonic diagnosis precision for liver and spleen injuries, consistently outperforming existing methods across varying injury severities. The approach achieved recognition rates ranging between 92.40% and 94.96%.
Moreover,21 presented an innovative framework employing a variational autoencoder (VAE) for spleen volume estimation from 2D segmentations, offering integration potential into standard clinical workflows. This approach surpassed conventional methods and human expert levels, facilitating direct 3D spleen volume estimation from 2D segmentations. Additionally,22 introduced an end-to-end automated pipeline for spleen segmentation in CT scans utilizing a Deep Neural Network (DNN) based on ResNet, significantly reducing segmentation time while maintaining quality comparable to manual labeling.
A notable study referenced in23 focused on splenomegaly segmentation, employing a deep CNN that integrated multi-resource labeled cohorts, achieving a Dice similarity coefficient of 0.94. This work underscored adaptability beyond spleen segmentation, hinting at broader application potential, including within 3D networks. Lastly, a comprehensive survey outlined in24 extensively explored diverse deep learning techniques, including CNNs, for organ segmentation from CT scans and Magnetic Resonance Imaging (MRI). Leveraging public datasets like the Medical Segmentation Decathlon25, the survey highlighted the transformative potential of deep learning in revolutionizing radiology practices despite ongoing challenges in standardization and evaluation.
Methods
This study presents a novel method for accurately segmenting spleens in ultrasound images using a two-phase training approach. Initially, the SegFormerB0 model is employed, followed by refinement with the Pix2Pix generative adversarial network (GAN). This approach leverages powerful feature extraction and refinement capabilities, resulting in precise and detailed segmentation. The following sections provide detailed information about the training and inference processes.
Training procedure
The training process begins with preprocessing the ultrasound images. Each image is resized to a standardized resolution of 512x512 pixels and normalized to the [0, 1] range to ensure consistency in model input.
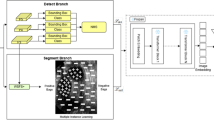
As shown in Fig. 1, during the initial training phase, the SegFormerB0 model is trained to generate initial segmentation mask. In the fine-tuning phase, both the discriminator and the generator (pre-trained SegFormerB0) are trained. This dual fine-tuning process enhances the generator’s ability to produce segmentation masks that closely align with the ground truth while simultaneously improving the discriminator’s ability to distinguish between ground truth and generated masks.
the SegFormerB0 model is employed as the initial segmentation network. Known for its transformer-based architecture, SegFormerB0 extracts rich semantic features from the input images through a hierarchical structure of encoders and decoders. The SegFormerB0 model is trained using a composite loss function that combines Binary Cross-Entropy (BCE) loss and Dice loss. The BCE loss is defined in Eq. (1):
where \(y_i\) denotes the ground truth label, \(p_i\) represents the predicted probability, and \(N\) is the total number of pixels. The Dice loss, defined in Eq. (2), which enhances overlap between the predicted and ground truth segmentations, is given by:
by combining Eqs. (1 and 2), the overall loss function for the SegFormerB0 model is defined in Eq. (3)
This combined loss ensures that the SegFormerB0 model outputs initial segmentation maps with accurate but coarse spleen boundaries.
After the SegFormerB0 model is fully trained, its output is used as the generator in the Pix2Pix GAN framework. The Pix2Pix framework consists of this pre-trained SegFormer generator and a discriminator. The generator’s role is to refine the initial segmentation maps, producing enhanced versions, while the discriminator ensures the realism of these outputs.
The objective of the conditional GAN (cGAN) used in Pix2Pix can be expressed as Eq. (4):
where \(G\) tries to minimize this objective against an adversarial \(D\) that tries to maximize it.
Formally, \(G^* = \arg \min _G \max _D \mathscr {L}_{\text {cGAN}}(G, D)\). Additionally, to encourage the generator to produce outputs that are near the ground truth, an L1 distance term is added. The L1 loss, defined in Eq. (5), is preferred over L2 as it encourages less blurring in the generated images.
The final objective for the generator combines the cGAN loss with the L1 loss defined in Eq. (6).
Here, \(\lambda\) is a hyperparameter that balances the importance of the L1 loss relative to the GAN loss. The discriminator’s loss function remains focused on distinguishing between real and generated segmentation maps. It loss function defined in Eq. (7).
These loss functions (4,7) guide the training of both the generator and discriminator to ensure the production of high-quality refined segmentation maps.
Figure illustrates the training method used in our paper. It shows the process where the SegFormerB0 network is pre-trained and then utilized as the generator in the Pix2Pix GAN framework. The input ultrasound image is first processed by the generator (pre-trained SegFormerB0), producing an initial segmentation map. This map is then refined, and the discriminator evaluates the realism of the generated segmentation compared to the real segmentation.
Inference process
During inference, the ultrasound images undergo the same preprocessing steps as during training, including resizing to 512x512 pixels and normalization.
As shown in Fig. 2, during the inference phase, the preprocessed images are directly passed to the trained Pix2Pix generator (which is the refined SegFormerB0) to obtain the final segmentation maps. This step enhances the accuracy and detail of the segmentation, leveraging the capabilities of the Pix2Pix generator to refine the initial maps.
After the Pix2Pix generator produces the refined segmentation maps, postprocessing steps are applied. These steps include thresholding to convert the probability maps into binary segmentation maps and morphological operations to remove noise and enhance boundary definitions. The final segmentation maps are thus precise and suitable for clinical use.
This combined approach leverages the strengths of both the SegFormerB0 and Pix2Pix models, providing a robust solution for accurate spleen segmentation in ultrasound imaging. The methodology ensures that the final segmentation maps are of high quality, addressing the inherent complexities and variabilities in medical ultrasound imaging.
Figure depicts the inference process. During inference, the preprocessed ultrasound image is directly passed to the trained generator (pre-trained SegFormerB0), which produces the final refined segmentation map. The post processed output shows the precise and detailed segmentation of the spleen in the ultrasound image.
Dataset
The dataset utilized in this study, named Spleenex, comprises 450 ultrasound images of the spleen. No patient metadata has been retained, adhering strictly to ethical guidelines. Each image has been anonymized through the assignment of coded names and IDs. All images have been shared with explicit consent from the respective patients, ensuring full compliance with ethical standards. Patient anonymity has been rigorously maintained throughout the dataset. Also the images were acquired using two distinct ultrasound devices (Philips Affinity 70 and Supersonic Aixplorer ). This dataset is the first public dataset in spleen image segmentation.
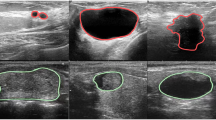
As illustrated in Fig. 3, the segmentation results are presented in three columns. The “Ultrasound” column contains the original ultrasound images. The “Mask” column displays the segmentation labels indicating the spleen. The “Image With Mask” column overlays the segmentation labels on the original ultrasound images.
Data labeling
In this section, the images were initially stored in DICOM format, and labeling was performed using the 3DSlicer software. The labeling was a collaborative effort involving two radiologists and four computer engineering master’s students. They segmented the spleen in the ultrasound images, clearly identifying and isolating it. Additionally, they measured the spleen length.
All labels and measurements were verified by the radiologist to ensure their accuracy. This collaborative approach ensured the reliability and precision of the labeled dataset.
Data partitioning
The dataset was divided into training and testing sets with an 80:20 ratio, ensuring no individual’s images were included in both sets simultaneously. To ensure robust experimental results, 5-fold cross-validation was applied, utilizing different training and testing datasets in each experiment. This careful division and validation process ensured unbiased model evaluation and reliable research outcomes.
Results
The performance of the proposed spleen segmentation method was evaluated using multiple metrics, including Precision, Recall, Intersection over Union (IoU), Kappa, and mean Dice (mDice). Our method was compared against several state-of-the-art models, including High-Resolution Net (HrNet-48)26, Attention-U-Net9, DeepLab-V37 (ResNet-101 and ResNet-50), LSPNet27 (Large, Medium, Small), OCRNet10 ,HrFormer-Small11, Real-Time Transformer (RtFormer)28, Yolov829 (Small), SeaFormer13 (Tiny), and SegFormer12 (B0, B1, B2, B3).
Performance evaluation metrics
In this study, an assessment of the efficacy of all segmetntation models, in estimating spleen length was conducted. The evaluation employed two primary metrics: the Mean Percentage Length Error (MPLE) and Intersection over Union (IoU). These metrics are defined in Eq. (8) and Eq. (10).
Where \(\textrm{L}^{\prime }=\left\{ l_1^{\prime }, l_2^{\prime }, l_3^{\prime }, \ldots , l_n^{\prime }\right\}\) represents the estimated lengths, \(L=\left\{ l_1, l_2, l_3, \ldots , l_n\right\}\) denotes the ground truth lengths, and n indicates the number of test images.
Distinct quantitative metrics were adopted to assess the performance of the models for segmentation tasks. For the segmentation-based model evaluation, the Dice Similarity Coefficient (Eq. 9) and Intersection over Union (IoU) were computed to evaluate the overlap between the predicted segmentation (A) and the ground truth segmentation (B). iou metrics are defined as Eq. (10):
Both measures yield values within the range of 0 to 1, where 0 signifies no overlap between the two segmentations, while 1 indicates perfect overlap.
Precision in image segmentation describes the accuracy of positive predictions. Precision equation’s defined in Eq. (11). It illustrates the proportion of pixel classifications that the model correctly identified as positive out of all the pixels it labeled as positive. The mathematical expression for precision defined in Eq. (11).
where \(TP\) is the number of true positives, and \(FP\) is the number of false positives. High precision indicates that the model made fewer false positive errors, essential in applications like medical diagnostics where such errors can have severe implications.
Recall, or sensitivity, measures the model’s ability to detect all relevant instances (true positives) within the dataset. For segmentation, recall calculates the proportion of actual positives the model correctly identified, important for ensuring no potential areas of interest are missed in the analysis. Equation defined in Eq. (12)
Lastly, Cohen’s Kappa provides a more nuanced view of model performance by measuring agreement adjusted for chance. Equation defined in Eq. (13)
where \(P_o\) is the observed agreement between the rater and the model, and \(P_e\) is the expected agreement by chance.
Collectively, these metrics provide a detailed picture of a segmentation model’s performance, highlighting aspects such as boundary accuracy, classification correctness, and the ability to handle imbalanced data.
Scatter plots illustrate the fine-tuning of hyperparameters for the spleen segmentation model using the Spleenex dataset. Each plot shows the relationship between the selected hyperparameter and the model performance over 300 training sessions, each consisting of 100 epochs. The cross markers indicate the final selected values for each hyperparameter: initial learning rate (lr0), final learning rate (lrf), momentum, weight decay, warmup momentum, HSV hue (hsv_h), translation fraction, scaling factor, HSV saturation (hsv_s), HSV value (hsv_v), flip left to right probability (fliplr), and mosaic augmentation probability.
Implementation details
The hyperparameters specified in Table 1 were fine-tuned through extensive experimentation using the Spleenex dataset. Specifically, the model underwent 300 distinct training sessions, each consisting of 100 epochs. This training was designed to iteratively refine the model’s parameters, ultimately optimizing its performance for spleen segmentation tasks. Each iteration provided valuable insights into the model’s behavior, allowing us to adjust the hyperparameters for maximum efficiency and accuracy.
The implementation of our spleen segmentation model involved a combination of SegFormer(b0) and the Pix2Pix discriminator, optimized for high performance and accuracy. The input image size was set to 512x512 pixels to maintain a balance between resolution and computational efficiency. The model was trained using the Stochastic Gradient Descent (SGD) optimizer with momentum, which helps in accelerating gradients vectors in the right directions, thus leading to faster convergence. The initial learning rate was set to 0.00834, gradually increasing to a final learning rate of 0.08933, allowing the model to adapt progressively during training. The momentum value used was 0.903, and a weight decay of 0.00042 was applied to prevent overfitting. Additionally, a warmup period of 3.21 epochs with a warmup momentum of 0.727 was used to stabilize the initial training phase.
Data augmentation played a crucial role in enhancing the robustness and generalizability of our model. We applied various augmentation techniques such as hue, saturation, and value adjustments (HSV_h: 0.0179, HSV_s: 0.9, HSV_v: 0.317), translations (0.11), and scaling (0.494). The probability of flipping images left to right was set at 0.34, while the mosaic augmentation, which merges four training images into one, had a high probability of 0.856. These augmentations helped in creating a diverse set of training samples, improving the model’s ability to generalize to different scenarios.
The scatter plots in Fig. 4 illustrate the relationship between the selected hyperparameters and the model performance, highlighting the distribution of results and the final selected values for each hyperparameter.
Experiments
In our experiments, we evaluated the performance of the proposed hybrid deep learning model for spleen segmentation in ultrasound images. We compared it with several state-of-the-art segmentation models, including HrNet-48, Attention-U-Net, DeepLab-V3 with ResNet backbones, LSPNet variants, OCRNet, RtFormer, Yolov8, SeaFormer, SegFormer variants, and others. The evaluation metrics included Precision, Recall, Intersection over Union (IoU), Kappa, and mean Dice (mDice) scores. Additionally, we assessed model complexity in terms of the number of parameters and floating-point operations per second (FLOPs).
Initially, we reviewed various segmentation models as shown in Tables 2 and 3. These tables provide a detailed comparison of different models based on multiple metrics. Among these models, HrNet-48, DeepLab-V3 (ResNet-101), and SegFormer-B3 demonstrated superior performance in terms of IoU. Specifically, HrNet-48 achieved an IoU of 93.421%, and SegFormer-B3 achieved an IoU of 93.276%, indicating their effectiveness in accurate segmentation tasks.
Our proposed method demonstrated superior performance, achieving a higher mean IoU and mean Dice score compared to existing methods. Specifically, our model attained a mean IoU of 94.17% and a mean Dice score of 96.82%, outperforming the SSNet and other models that achieved mean IoU scores ranging from 89.26% to 93.15% as shown in Table 4. The precision and recall metrics indicated a high level of accuracy and sensitivity, showcasing the model’s ability to identify and segment the spleen from ultrasound images correctly.
During training, the loss curves for various segmentation models were plotted to analyze the convergence behavior and stability. Our proposed model showed a steady convergence with minimal oscillations, indicating a stable training process as shown in Fig. 5. The comparison of training loss curves highlighted that our model not only converged faster but also achieved lower final loss values compared to other models. This demonstrates the efficiency and effectiveness of integrating SegFormer and Pix2Pix networks, leveraging the strengths of both transformer-based feature extraction and generative adversarial refinement.
Furthermore, the training curves for the mean Intersection over Union (mIoU) across iterations were analyzed for different models as shown in Fig. 6. The plot illustrated the progression and convergence of each model’s performance over time. Our proposed method consistently achieved higher mIoU values, indicating its robustness and efficiency in learning the segmentation task.
Additionally, we conducted a qualitative analysis by visually comparing the segmentation results on various test samples. The results included the original ultrasound image, the ground truth segmentation by a human expert, and the segmentation results from DeepLabV3, SeaFormer, U-Net++, SegFormer, and our proposed method as shown in Fig. 7. Our method consistently produced segmentation maps that closely matched the ground truth annotations, demonstrating clear and accurate delineation of the spleen boundaries. This visual comparison highlighted the effectiveness and robustness of our approach in challenging scenarios, minimizing noise and improving boundary adherence. The presence of high noise in the spleen ultrasound image makes some areas of the spleen difficult to see. The proposed method helps to improve the masks produced by segformer. The proposed method trains the network with the help of the Pix2Pix method in such a way that the production mask becomes similar to the original mask.
Table 5 presents the Mean Percentage Length Error (MPLE) for different models, including the proposed method, when compared to the ground truth for major axis length using the Spleenex dataset. The proposed method achieves the lowest MPLE of 3.64%, outperforming Yuan et al. (VAE Based) (4.21%) and SSNet (4.03%). Yuan et al. (U-Net Based) shows the highest error with an MPLE of 6.40%. These results suggest that the proposed method offers the most accurate performance for spleen segmentation within this dataset.
This improved accuracy in spleen size estimation directly enhances the ability to diagnose spleen-related conditions, such as splenomegaly, by providing precise measurements critical for clinical evaluation and decision-making.
Overall, our experiments demonstrated that the proposed hybrid model significantly enhances spleen segmentation performance in ultrasound images. It outperforms several state-of-the-art models across various quantitative metrics and provides clear and accurate visual results. These findings indicate the potential of our method for practical application in medical diagnostics, offering a robust and efficient tool for clinical use.
Training loss curves for various segmentation models over iterations. The y-axis represents the loss value, while the x-axis represents the number of iterations. The plot demonstrates the convergence behavior and stability of each model during training, highlighting the differences in performance and training efficiency across various architectures.
Training curves showing the Mean Intersection over Union (mIoU) across iterations for various segmentation models. The y-axis indicates the mIoU value, and the x-axis represents the number of iterations. This plot illustrates the progression and convergence of each model’s performance over time, allowing for a comparative analysis of their training stability and efficiency.
Qualitative comparison of spleen segmentation results on ultrasound images using various segmentation methods. From left to right, the columns show the original ultrasound image, the ground truth segmentation by a human expert, and the results from DeepLabV3, SeaFormer, U-Net++, SegFormer, and our proposed method. The rows represent different test samples, showcasing the performance of each method across a variety of ultrasound images. The red overlay indicates the segmented spleen area. Our proposed method consistently demonstrates more accurate and precise segmentation, closely matching the human expert’s annotations, and outperforms other methods in challenging scenarios with less noise and better boundary adherence. This visual comparison highlights the effectiveness and robustness of our approach in spleen segmentation tasks.
Conclusion
Our study introduced a novel approach to ultrasound image segmentation of the spleen by employing a two-phase training process involving SegFormerB0 and Pix2Pix networks. This method has demonstrated a high IoU score, achieving 94.17% on the Spleenex dataset. This performance surpasses existing state-of-the-art methods, such as SSNet and SegFormerB3, which achieved IoU scores of 92.96% and 93.27%, respectively. Additionally, our model is notably more efficient, requiring only 4 million parameters compared to the substantially higher counts in SegFormerB3 (47.23 million) and HrNet-48 (70.08 million). This significant reduction in model complexity not only facilitates quicker training times but also enhances the feasibility of deploying this model in clinical settings where computational resources may be limited.
The dataset gathered during this research, comprising 450 ultrasound images of the spleen, constitutes a valuable resource for future studies. Spleenex dataset is the first dataset in spleen image segmentation. This dataset not only enriches the available data for training segmentation models but also serves as a benchmark for evaluating the efficacy of novel segmentation techniques. By providing open access for non-commercial use of this dataset, we aim to foster further research and collaboration in the field, encouraging advancements in both methodology and clinical applications.
Furthermore, the implications of this research extend beyond mere technical superiority. By achieving higher accuracy with a less complex model, our approach demonstrates the potential for practical application in medical diagnostics, where robustness and efficiency are paramount. The successful application of our model in segmenting spleen ultrasound images can lead to more precise and timely medical interventions, ultimately improving patient outcomes. However, our study has certain limitations that we aim to address in future work. One such limitation is the relatively small size of the Spleenex dataset, which, although unique, consists of only 450 images. Expanding the dataset with more diverse samples from additional sources would improve the model’s generalizability. Another limitation is that the dataset was collected using only two specific ultrasound devices, which might restrict the model’s performance on data from other devices. Including data from a broader variety of ultrasound machines could enhance the model’s robustness. Future work will focus on refining this model further, exploring its adaptability to other types of medical imaging, and enhancing its real-world applicability to support healthcare professionals in their diagnostic and therapeutic tasks.
Data Availability
The Spleenex dataset is available from the corresponding author on reasonable request. Interested researchers are invited to submit an application through the designated request form available at https://www.ariameditech.com/datasets/spleenex/. It should be noted that these datasets are provided solely for academic and non-commercial research purposes. Prior to access, requestors are required to agree to the terms and conditions specified in the request form, ensuring the data will be used in accordance with ethical standards and regulations.
References
Chapman, J., Goyal, A. & Azevedo, A. M. Splenomegaly (2024).
Yuan, Z. et al. Deep learning-based quality-controlled spleen assessment from ultrasound images. Biomed. Signal Process. Control 76, 103724 (2022).
Perez, A. A. et al. Automated deep learning artificial intelligence tool for spleen segmentation on ct: Defining volume-based thresholds for splenomegaly. Am. J. Roentgenol. 221, 611–619 (2023).
Jiang, X. et al. Development and validation of the diagnostic accuracy of artificial intelligence-assisted ultrasound in the classification of splenic trauma. Ann. Transl. Med.10 (2022).
Zhang, Q. et al. Deep learning models for diagnosing spleen and stomach diseases in smart Chinese medicine with cloud computing. Concurr. Comput.: Pract. Exp. 33, 1–1 (2021).
Ronneberger, O., Fischer, P. & Brox, T. U-net: Convolutional networks for biomedical image segmentation. In Medical image computing and computer-assisted intervention–MICCAI 2015: 18th international conference, Munich, Germany, October 5-9, 2015, proceedings, part III 18, 234–241 (Springer, 2015).
Chen, L.-C., Papandreou, G., Schroff, F. & Adam, H. Rethinking atrous convolution for semantic image segmentation. arXiv preprint arXiv:1706.05587 (2017).
Zhou, Z., Rahman Siddiquee, M. M., Tajbakhsh, N. & Liang, J. Unet++: A nested u-net architecture for medical image segmentation. In Deep Learning in Medical Image Analysis and Multimodal Learning for Clinical Decision Support: 4th International Workshop, DLMIA 2018, and 8th International Workshop, ML-CDS 2018, Held in Conjunction with MICCAI 2018, Granada, Spain, September 20, 2018, Proceedings 4, 3–11 (Springer, 2018).
Oktay, O. et al. Attention u-net: Learning where to look for the pancreas. arXiv preprint arXiv:1804.03999 (2018).
Yuan, Y., Chen, X. & Wang, J. Object-contextual representations for semantic segmentation. In Computer Vision–ECCV 2020: 16th European Conference, Glasgow, UK, August 23–28, 2020, Proceedings, Part VI 16, 173–190 (Springer, 2020).
Yuan, Y. et al. Hrformer: High-resolution transformer for dense prediction. arXiv preprint arXiv:2110.09408 (2021).
Xie, E. et al. Segformer: Simple and efficient design for semantic segmentation with transformers. Adv. Neural. Inf. Process. Syst. 34, 12077–12090 (2021).
Wan, Q., Huang, Z., Lu, J., Yu, G. & Zhang, L. Seaformer: Squeeze-enhanced axial transformer for mobile semantic segmentation. arXiv preprint arXiv:2301.13156 (2023).
Chen, G. et al. Esknet: An enhanced adaptive selection kernel convolution for ultrasound breast tumors segmentation. Expert Syst. Appl. 246, 123265 (2024).
Chen, G., Li, L., Zhang, J. & Dai, Y. Rethinking the unpretentious u-net for medical ultrasound image segmentation. Pattern Recogn. 142, 109728 (2023).
Chen, G., Dai, Y. & Zhang, J. C-net: Cascaded convolutional neural network with global guidance and refinement residuals for breast ultrasound images segmentation. Comput. Methods Progr. Biomed. 225, 107086 (2022).
Zhang, L. et al. Artificial neural network aided non-invasive grading evaluation of hepatic fibrosis by duplex ultrasonography. BMC Med. Inform. Decis. Mak. 12, 1–6 (2012).
Huo, Y. et al. Splenomegaly segmentation using global convolutional kernels and conditional generative adversarial networks. Med. Imaging 2018: Image Process., 10574, 45–51 (SPIE, 2018).
Mirza, M. & Osindero, S. Conditional generative adversarial nets. arXiv preprint arXiv:1411.1784 (2014).
Yuan, Z. et al. Deep learning for automatic spleen length measurement in sickle cell disease patients. In Medical Ultrasound, and Preterm, Perinatal and Paediatric Image Analysis: First International Workshop, ASMUS 2020, and 5th International Workshop, PIPPI 2020, Held in Conjunction with MICCAI 2020, Lima, Peru, October 4-8, 2020, Proceedings 1, 33–41 (Springer, 2020).
Yuan, Z., Puyol-Antón, E., Jogeesvaran, H., Inusa, B. & King, A. P. Deep learning framework for spleen volume estimation from 2d cross-sectional views. arXiv preprint arXiv:2308.08038 (2023).
Moon, H. et al. Acceleration of spleen segmentation with end-to-end deep learning method and automated pipeline. Comput. Biol. Med. 107, 109–117 (2019).
Tang, Y. et al. Improving splenomegaly segmentation by learning from heterogeneous multi-source labels. In Med. Imaging 2019: Image Process., 10949, 53–60 (SPIE, 2019).
Altini, N. et al. Liver, kidney and spleen segmentation from CT scans and MRI with deep learning: A survey. Neurocomputing 490, 30–53 (2022).
Antonelli, M. et al. The medical segmentation decathlon. Nat. Commun. 13, 4128 (2022).
Wang, W. et al. Exploring cross-image pixel contrast for semantic segmentation. In Proceedings of the IEEE/CVF International Conference on Computer Vision, 7303–7313 (2021).
Zhang, Y., Yao, T., Qiu, Z. & Mei, T. Lightweight and progressively-scalable networks for semantic segmentation. Int. J. Comput. Vision 131, 2153–2171 (2023).
Wang, J. et al. Rtformer: Efficient design for real-time semantic segmentation with transformer. Adv. Neural. Inf. Process. Syst. 35, 7423–7436 (2022).
Jocher, G., Chaurasia, A. & Qiu, J. Ultralytics yolov8 (2023).
Guo, M.-H. et al. Segnext: Rethinking convolutional attention design for semantic segmentation. Adv. Neural. Inf. Process. Syst. 35, 1140–1156 (2022).
Zhang, W. et al. Topformer: Token pyramid transformer for mobile semantic segmentation. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 12083–12093 (2022).
Qin, X. et al. U2-net: Going deeper with nested u-structure for salient object detection. Pattern Recogn. 106, 107404 (2020).
ghattan kashani, h. & Shariat Panahi, M. Intelligent detection, measurement and segmentation of ultrasound images of internal organs using neural networks. Ministry of Health and Medical Education1402 (2024). https://ethicsresearch.ut.ac.ir/article_96212_e01372fa3cd2d1bbbb74ec651851a8dc.pdf.
Author information
Authors and Affiliations
Contributions
A.K. and F.M. wrote the manuscript, designed and conducted the experiments, and also developed and implemented the proposed method. J.S., A.S., K.F., P.E., A.S.M., and H.S. contributed to data collection and labeling. K.F. and J.S. were involved in assessing the project’s feasibility. H.G., A.S.M., and M.S. managed and supervised the project. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Ethical approval
In accordance with ethical guidelines, studies involving human participants were reviewed and approved by the Physical Education and Sport Sciences at University of Tehran (ID: IR.UT.SPORT.REC.1402.127)33. Prior to participation, all participants provided informed written consent. Additionally, explicit consent was obtained from each individual for the publication of any images in this manuscript.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Karimi, A., Seraj, J., Mirzadeh Sarcheshmeh, F. et al. Improving spleen segmentation in ultrasound images using a hybrid deep learning framework. Sci Rep 15, 1670 (2025). https://doi.org/10.1038/s41598-025-85632-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-85632-9
This article is cited by
-
Deep learning framework for automated frame selection in kidney ultrasound
Scientific Reports (2025)