Abstract
There is an initiative driven by the carbon-neutrality nature of biochar in recent times, where various countries across Europe and North America have introduced perks to encourage the production of biochar for construction purposes. This objective aligns with the zero greenhouse emission targets set by COP27 for 2050. This research work seeks to assess the effectiveness of biochar in soils with varying grain size distributions in enhancing the soil–water characteristic curve (SWCC). This work further explores the effect of different combinations of biochar content (0 to 15 mass %) on the bioelectricity generation from biochar-improved plant microbial fuel cells (BPMFC). Additionally, different machine learning models such as the “Gradient Boosting (GB)”, “CN2 Rule Induction (CN2)”, “Naive Bayes (NB)”, “Support vector machine (SVM), “Stochastic Gradient Descent (SGD)”, “K-Nearest Neighbors (KNN)”, “Tree Decision (Tree)”, “Random Forest (RF)”, and “Response Surface Methodology” (RSM), have been developed to predict SWCC based on soil suction, electric current, electrical potential, volumetric water content, temperature, and bulk density. The newly established model demonstrates a reasonable ability to predict SWCC and a cheaper technology in predicting the suction of unsaturated soils in relation to the studied bioelectric factors of the BPMFC. Overall, in this research paper, the GB, SVM and CN2 outclassed the other regression techniques in this order thereby proposing the cheapest technology with the highest performance index to predict the SWCC behavior of unsaturated soils in a BPMFC system.
Similar content being viewed by others
Introduction
Unsaturated soil behavior, also referred to as unsaturated soil mechanics, is a field within geotechnical engineering dedicated to studying soils that are not fully saturated with water1. While traditional soil mechanics has primarily focused on saturated soils, unsaturated soils are critical in various engineering and environmental contexts1,2,3. These soils contain both air and water within their pore spaces1,4, with saturation levels that can range from partial to full1,3. Factors such as suction, matric potential, and water content significantly influence the behavior of unsaturated soils1,2,3,4,5,6. The integration of biochar with a Plant Microbial Fuel Cell offers a promising, comprehensive approach to tackling soil degradation, boosting agricultural productivity, and supporting sustainable energy solutions in unsaturated soils1. Continued research and experimentation are essential to fully explore these synergies and to optimize this combined system for practical applications, leveraging machine learning techniques. Figure 1 presents an illustrative mechanism of the PMFC system.
Chen et al.1 examined how plant microbial fuel cells (PMFCs) behave in terms of bio-hydrology and suggested using biochar amendment to enhance their bioelectricity production under drought. Biochar-enhanced Plant Microbial Fuel Cells were created by cultivating green-roof plants on sandy lean clay mixed with biochar at various mass ratios. Biochar significantly boosted the electrical power production of PMFCs by up to 30 times in unsaturated circumstances, with bioelectricity closely linked to suction. The study proposed utilizing bioelectricity to monitor soil-water properties and improve the drought resilience of PMFCs. Hussain et al.2 examined how plant microbial fuel cells behave in terms of bio-hydrology and suggested using biochar amendment to enhance their bioelectricity production under drought. Biochar-enhanced Plant Microbial Fuel Cells were created by cultivating green-roof plants in sandy lean clay mixed with biochar at various mass ratios. The study found that biochar significantly boosted the electrical power generated by PMFCs by up to 30 times under dry conditions, and bioelectricity was closely linked to suction. The study proposes utilizing bioelectricity to monitor soil-water properties and improve the drought resilience of PMFCs. Also, Cai et al.3 examined how biochar affects the soil shrinkage and water retention properties of kaolin and bentonite in salty environments. The findings indicated that biochar effectively reduced the shrinkage of clays under salt stress by 6-14% and 50-107%, respectively. Biochar’s porous structure and hydrophilic properties trap sodium ions by undergoing ion exchange and protonation processes. The research indicated that biochar-modified clays may aid in desalination and enhance resistance to shrinkage-induced harm in hydro-chemical barriers. Zhou et al.4 examined biochar synthesis, engineering techniques, uses, and future possibilities. The text emphasized the advancement of physical, chemical, and bio-engineering methods for producing biochar. Possible substitutes for biochar include the potential for creating carbon-based products, treating wastewater, storing energy, and repairing super-capacitors. The report also explored upcoming tactics and technology for the circular bio-economy. Guo et al.5 discovered that the use of peanut shell biochar can enhance the interaction between soil and grass in grassed plots. The use of biochar increased the grass leaf area index by 38% and root length density by 200%. The biochar decreased peak suction during evapo-transpiration by 54%, therefore decreasing excessive water loss. The study suggested using 5% peanut shell biochar content for the long-term maintenance of vegetated earthen infrastructures to mitigate adverse effects on plant development. Chen et al.6 investigated how freezing-thawing cycles affect clay permeability and saturated permeability (Ksat) in clay. Clay samples were compressed at different moisture levels, and the amount of biochar applied was recorded. The study found that the hydraulic conductivity (Ksat) decreased significantly when the biochar application rate exceeded 4%. This indicated that biochar can be utilized as an environmentally acceptable additive to control clay permeability, particularly in cold regions, to reduce Ksat in geo-environmental constructions. Keeffe7 intended to include biochar in precision agriculture technology to enhance soil health, water retention, and crop output. A finite element analysis model was created and utilized on a conventional Palouse silt loam soil at various concentrations. The model demonstrated higher retention for both types of amendments and concentrations, resulting in increased moisture retention in and around the amendment zone. The study investigated the impact of biochar amendment on the hydrologic processes in the Palouse region, specifically targeting the argillic and frangipani layers. Chen et al.8 examined the molecular composition changes in paddy topsoil following biochar amendments using untreated, manured, and burned maize straw. Research findings indicate that biochar improves the retention of carbon in the soil and the durability of organic matter, although there is limited investigation into alterations in the molecular structure of organic compounds. The research revealed that biochar raised soil organic matter content by 12% and 36% in comparison to no amendment. Yet, biochar also reduced the quantity of n-alkanes, fatty acids, and free lipids in the study. The study proposed that reintroducing agricultural residue as biochar could be a sustainable method to improve soil organic matter and molecular diversity. Another research report, Quinn9 examined the movement of Escherichia coli pathogenic and nonpathogenic. Coli k12 strains in water-saturated sand columns treated with Magnesium and Nitrogen-doped biochar. Bacterial cell retention was improved by adding biochar at a 2% weight ratio. The study determined that biochar repels water, whereas sand and bacteria attract water. The optimal biochar was created at 600 degrees Celsius, indicating that while developing efficient biochar filters, all factors influencing bacterial movement should be taken into account. In other studies, Geuder10 created optimal management strategies for farmers and producers to mitigate these detrimental effects. Two components are discussed: a soil amendment experiment comparing greenhouse gas (GHG) reduction of various biochar application methods and the development and building of a passive multi-component bioreactor. Biochar additions have been proven to enhance soil health and crop yield, as well as reduce greenhouse gas emissions when used in conjunction with fertilizer on farmland. The study investigated the release of CO2 and N2O from four small areas in Winchester, southern Ontario, Canada, after applying varying amounts of liquid swine dung and biochar. The bioreactors were created to reduce N2O gas emissions and eliminate veterinary and pharmaceutical substances present in field amendments. Additional research is required to establish the ideal biochar co-amendment quantities and timing, as well as to evaluate bioreactor efficiency. Graves11 evaluated the effectiveness of various biochar types derived from Miscanthus grass for controlling ammonia levels in chicken houses. Acid-activated biochar reduced biochar pH and raised overall acidity levels. During a two-week lab-scale experiment, biochar produced at 400 °C, processed with acetic acid, and administered at a high addition rate decreased ammonia emissions by 19.7%. Future studies investigated novel biochar activation techniques to improve the preservation of acid functional groups to boost ammonia adsorption.
Methodology
Preamble
The preparation of materials for a biochar-enhanced plant microbial fuel cell experiment in unsaturated soil involved assembling the components required for a controlled and reproducible setup1. Unsaturated soil samples were collected from the study site, and moisture content was adjusted to maintain unsaturated conditions12,13,14. Containers were prepared to hold the soil and selected vegetation. Biochar was produced via a controlled pyrolysis process following the under-pressure-controlled heating method outlined by Onyelowe et al.15. Soil and biochar particle distributions are shown in Fig. 2. The soil was composed of 65% silt and 35% sand, with a plasticity index of 15%, specific gravity of 2.66, and a saturated hydraulic permeability (Ks) of 1.5E-6 m/s, classified as CL in the unified soil classification system1. The biochar feedstock was Prunus persica, processed at 600 °C, with a cation exchange capacity (CEC) of 85.0 cmol/kg, ash content of 21.6%, pH of 9.1, and a water absorption capacity (WAC) of 3.53 g/g1. Comprehensive characteristics of both soil and biochar are detailed in the literature1.
Treatment process and data collection
The biochar was incorporated into the PMFC system at mass ratios of 0%, 5%, and 10% relative to soil mass1,2. To ensure uniform mixing, variations in particle size and biochar source were taken into account, with shovels or mixing tools used to evenly distribute the biochar within the soil. Hydrocotyle vulgaris plants, suitable for the experiment, were selected for planting1. A watering system using watering cans provided consistent moisture control throughout the experiment. Carbon-based materials were selected for the anode and cathode electrodes in a PMFC setup, which were securely placed in containers to house the plants and electrodes16. Electrical connections were established with wires and connectors linked to measurement instruments, including a multimeter or data logger to record voltage, current, and power output from the PMFC. Soil moisture sensors were installed for real-time monitoring, and data recording tools like notebooks, spreadsheets, or data loggers were used to track all measurements. Safety protocols were followed, with appropriate personal protective equipment (PPE) such as gloves and safety glasses, depending on the materials and procedures. Soil samples were either collected from the study area or selected from a standard soil type, and moisture content was adjusted by air-drying or adding water to achieve unsaturated conditions. Containers or pots were filled with the prepared soil, and the biochar was mixed into the soil in precise amounts for each treatment level, ensuring even distribution16,17,18. The selected vegetation was planted according to recommended spacing and depth guidelines, ensuring consistency in plant type, growth stage, and health1. The PMFC system was completed by securely installing the chosen electrode materials into the soil containers19,20,21,22. Wires connected the electrodes to measuring devices, and soil moisture sensors were placed at different depths. Data recording tools were configured to continuously track soil moisture levels, PMFC performance, and other relevant data1. Experimental conditions such as temperature and lighting were maintained at consistent levels, with replicates set up for each treatment to support reliable results. The complete setup is illustrated in Fig. 31.
A total of ninety (90) records were collected from the experimentally tested silty sand samples mixed with different amount of biochar. Each record contains the following data:
-
Bio Biochar ratio;
-
I Electric current (µA);
-
U Electrical potential (mV);
-
θ Metric water content;
-
T Temperature (°C);
-
γb Bulk density (g/cm3);
-
Log ψ the logarithm of Suction (kPa) to base 10.
The collected records were divided into training set (70 records ≈ 75%) and validation set (20 records ≈ 25%). Tables 1 and 2 summarize their statistical characteristics and the Pearson correlation matrix. Finally, Fig. 4 shows the histograms for both inputs and outputs and the relations between the inputs and the outputs.
Sensitivity analysis
The suction of unsaturated granular soil treated with biochar plays a critical role in the performance of Plant Microbial Fuel Cells (PMFCs) for bioelectricity generation. Suction controls the soil moisture retention, hydraulic conductivity, microbial activity, and electrochemical interactions that influence both soil behavior and bioelectricity output. Conducting a sensitivity analysis helps to identify the most influential parameters affecting suction and their impact on the system’s efficiency. A preliminary sensitivity analysis was carried out on the collected database to estimate the impact of each input on the (Log Ψ) values. “Single variable per time” technique is used to determine the “Sensitivity Index” (SI) for each input using Hoffman a Gardener formula as follows:
A sensitivity index of 1.0 indicates complete sensitivity, a sensitivity index less than 0.01 indicates that the model is insensitive to changes in the parameter. Figure 5 shows the sensitivity analysis with respect to (Log Ψ). It can be shown that U: the Electric potential has the highest influence on the suction of the unsaturated soil and this is followed closely by θ: Metric water content and γb: Bulk density (g/cm3).
Research program
Eight different ML classification techniques were used to predict the suction of biochar incorporated into the PMFC system at mass ratios of 0%, 5%, and 10% relative to soil mass using the collected database. These techniques are “Gradient Boosting (GB)”, “CN2 Rule Induction (CN2)”, “Naive Bayes (NB)”, “Support vector machine (SVM), “Stochastic Gradient Descent (SGD)”, “K-Nearest Neighbors (KNN)”, “Tree Decision (Tree)” and “Random Forest (RF)”. The developed models were used to predict (Log Ψ) using the inputs (Bio, I, U, θ, T, γb). All the developed models were created using “Orange Data Mining” software version 3.36. The considered data flow diagram is shown in Fig. 6. The following section discusses the results of each model. The accuracies of developed models were evaluated by comparing SSE, MAE, MSE, RMSE, Error (%), Accuracy (%) and R2 between predicted and calculated suction parameters values. The definition of each used measurement is presented in Eq. (2) to (7).
Theoretical frameworks for the selected machine learning techniques used in this study
Gradient boosting
Gradient boosting is an enhanced machine learning technique, which generates a strong predictive model by combining a series of decision trees5. The gradient boosting has a framework that iteratively minimizes loss function by using gradient descent, to become e effective for both classification and regression analysis. It starts with an initial model, f0(x), often a simple prediction like the mean of the target values for regression or a uniform class probability for classification. In each step, the model is improved by training a new weak learner to focus on the remaining errors, or residuals, from previous predictions. At each iteration, the algorithm calculates the negative gradient of the loss function with respect to the model’s predictions and essentially finding the direction in which the model should adjust to reduce error. This gradient guides the training of a new weak learner, hm(x), which is then added to the existing model. The updated model can be written as:
where α = learning rate and it controls the influence of each weak learner on the overall model.
CN2 rule induction
The CN2 Rule Induction operates as a rule-based classification algorithm that has been designed to generate a set of if-then rules, which differentiate between classes within a dataset24. There set of instances as the starting point and each instance contains a class label and a attribute values collection. The algorithm’s objective is to iteratively create simple, interpretable rules that classify data accurately by sequentially optimizing rules for maximum coverage and accuracy. Each rule in CN2 takes the general form: IF Condition→THEN Class. Where the Condition is a conjunction of attribute-value pairs that define a subset of instances for which the rule is valid, and Class is the predicted class label for instances satisfying the condition. CN2 evaluates candidate rules using a heuristic measure, commonly based on the entropy or likelihood ratio of the rule’s coverage and accuracy in differentiating a specific class. For a given rule R, the information gain can be calculated using entropy to measure the quality of the rule. The entropy H for a rule’s distribution over classes is defined as:
where P(c∣R)is the conditional probability of class ccc given that an instance matches the conditions of rule RRR. The information gain IGIGIG of a rule is then:
where H(C) represents the entropy of the class distribution in the dataset, and H(R) is the entropy of instances covered by the rule RRR. A higher information gain implies a more effective rule in separating instances of different classes. The coverage of a rule R, denoted as Cov(R), refers to the proportion of instances in the dataset that satisfy the rule’s conditions. This is mathematically represented as:
where ∣X∣ is the total number of instances. Higher coverage indicates that the rule applies to a larger portion of the dataset, though there may be a trade-off between coverage and precision.
Naive Bayes
Naive Bayes is based on Bayes’ theorem of probabilistic classifier and leveraging the assumption that class features are conditionally independent. For instance, a given class C and feature vector X = (x1,x2,…,xn), will have a posterior probability as follow:
Thus the class \(\:\widehat{C}\) that maximizes this posterior probability is predicted by the classifier, such that:
Where P(C) = prior probability of each class, estimated as the relative frequency of instances in that class. P(xi∣C) = conditional probability of each feature given the class, which can be calculated as frequency counts for categorical features or approximated by a Gaussian distribution for continuous features.
Support vector
Support Vector Machines (SVMs) are supervised machine learning techniques mainly used for classification projects25. In SVMs, finding the optimal hyperplane that maximally separates data points from different classes is achieved. For instance, in linearly separable data, SVM can identify this hyperplane, through maximizing the distance or margin between the each data closest data points or support vectors. Considering dataset of labeled instances (xi,yi)) where xi∈Rn and yi∈{−1,1}, the decision boundary becomes a hyperplane w⋅x + b = 0, where w = weight vector perpendicular to the hyperplane, and b = bias term. The optimization problem to maximize the margin is formulated as:
Subject to the constraints:
In the case of non-linearly separable data, SVM applies the kernel functions to project data into a higher-dimensional space, where a linear separation is possible. Common kernels include the linear, polynomial, and radial basis function (RBF) kernels. The decision function for classification is then:
where αi = Lagrange multipliers, and K(x, xi) = chosen kernel function.
Stochastic gradient descent
Stochastic Gradient Descent (SGD) is a machine learning models that performs iterative optimization for data training26. For instance, in high-dimensional spaces, SGD minimizes a given objective function, J(θ), which typically determines the model error with parameters θ. In each iteration, SGD computes the gradient using a single randomly chosen instance or a small batch rather than computing the gradient over the entire dataset. Thus, this technique speeds up convergence by updating the parameters, such that:
where η is the learning rate, controlling the step size and \(\:\eta{\nabla\:}_{\theta\:}J(\theta\:;{x}^{\left(i\right)},{y}^{\left(i\right)})\) is the gradient of the objective function with respect to θ, evaluated at a training example\(\:{x}^{\left(i\right)},{y}^{\left(i\right)}\).
k-Nearest neighbours
The k-Nearest Neighbors algorithm, also denoted as k-NN, is a non-parametric and instance-based classification technique, which predicts the class of a query instance based on the majority class among its k closest neighbors in the population27. Figure 12 shows the illustration of the K-nearest neighbours. It operates by estimating the distance between the query instance and all other points in the dataset, commonly using Euclidean distance for continuous variables:
where x and x′ are two instances in n-dimensional space.
Tree decision
Decision Trees are supervised learning algorithms, which are used for classification and regression projects. They are able to split data recursively using feature values to create a tree structure, having each internal node, branches and leaf nodes representing feature test, outcomes, and predicted values, respectively. For example, considering a dataset D with classes C, the tree grows by selecting features that maximize the information gain or minimize the impurity. Hence, information gain IG for a split on feature X is respected as:
where H(D) is the entropy or impurity of dataset D, and Dv is the subset of D for each value v of feature X.
Random forest
The random forest algorithm is an ensemble learning approach, which builds multiple decision trees for regression or classification project, and it improves the robustness and accuracy by reducing single trees overfitting25. Each tree in the forest is trained on a different bootstrap sample of the dataset, with random subsets of features selected at each split, introducing diversity among trees. For a training dataset D with n samples, for instance, Random Forest will construct m decision trees T1, T2, …, Tm. Thus, each of the trees is trained on a bootstrap sample Di (random sample with replacement) from D, and at each node, a random subset of k features is selected to find the best split. For classification, the output is determined by a majority vote across all trees:
For regression, the output is the average prediction from all trees:
Response surface methodology
The Response Surface Methodology (RSM) comprises of both mathematical and statistical methods for modelling and optimizing processes by exploring the relationships between response variables and multiple input factors28. In its operation, RSM approximates the underlying process using mostly a second-order polynomial equation that is suitable for capturing response surface curvature. For a process with input variables x1,x2,…,xk and response y, the second-order response surface model is:
where β0 is the intercept, Βi, βii, and βij are coefficients for linear, quadratic, and interaction terms, respectively, \(\epsilon\) is the error term. Also, RSM utilizes Box-Behnken Design and Central Composite Design (CCD) to gather data and fit model efficiently. Moreover, RSM also identifies the optimal settings of the input factors by analyzing the fitted response surface, often using gradient-based methods to locate the maximum or minimum response or desired regions.
Results presentation and discussion
GB model
The developed (GB) model was based on (Scikit-learn) method with learning rate of 0.1 and minimum tensile subset of 2. Six trials were conducted for each model started with two trees and three tree levels and increased to five tree levels but 4 and 5 trees produced negative errors invoking distortions. The reduction of the prediction Error (%) for each trail is presented in Fig. 7. Accordingly, the models with four trees and four tree levels are considered the optimum ones. Performance metrics of the developed model for both training and validation dataset are listed in Table 2. The average achieved accuracy was (98%). The relations between calculated and predicted values are shown in Fig. 8.
CN2 model
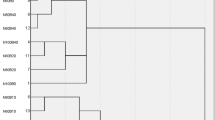
Similarly, five (CN2) models were developed considering “Laplace accuracy” as evaluation measurement with beam width of 1.0 and minimum rule coverage of 1.0. The maximum rule length was started by 1.0 and increased up to 5.0. Figure 9 shows the reduction in Error % with increasing the rule length. Accordingly, rule length of 5.0 is considered. The developed models contains 54 “IF condition” rules, Fig. 10 presents some of these rules. Performance metrics of the developed model for both training and validation dataset are listed in Table 2. The average achieved accuracy was (95%). The relations between calculated and predicted values are shown in Fig. 11.
NB model
Traditional Naive Bayes classifier technique considering the concept of “Maximum likelihood” was used to develop the nine models. Although this type of classifier is highly scalable and are used in many applications, but it showed a low performance as shown in Table 2. The relations between calculated and predicted values are shown in Fig. 12. The achieved average accuracy was 76%.
SVM model
The developed (SVM) model was based on “polynomial” kernel with cost value of 100, regression loss of 0.10 and numerical tolerance of 1.0. The kernel started with one-degree polynomial (linear) and increased up to two-degree polynomial (quadratic). The reduction in the error % with increasing the polynomial degree is illustrated in Fig. 13. Performance metrics of the three developed models for both training and validation dataset are listed in Table 2. The average achieved accuracy was (97%). The relations between calculated and predicted values are shown in Fig. 14.
SGD model
These three models were developed considering modified Huber classification function and “Elastic net” re-generalization technique with mixing factor of 0.01 and strength factor of 0.001.The learning rate starts with 0.01, then gradually decreased to 0.001. The reduction in error (%) with reducing the learning rate is presented in Fig. 15. Performance metrics of the three developed models for both training and validation dataset are listed in Table 2. The average achieved accuracy was (85%). The relations between calculated and predicted values are shown in Fig. 16.
KNN model
Considering number of neighbors of 1.0, Euclidian metric method and weights were evaluated by distances, the developed (KNN) models showed the best accuracy. (KNN) model showed the best performance where the average error (%) was (95%). The relations between calculated and predicted values are shown in Fig. 17.
Tree model
These Four models were developed considering minimum number of instants in leaves of 2.0 and minimum split subset of 5.0. The models began with only two tree levels and gradually increased to five levels. Figure 18 illustrates the reduction in error with increasing the number of layers. The generated trees layouts are shown in Fig. 19. Performance metrics of the last developed model for both training and validation dataset are listed in Table 2. The average achieved accuracy was (95%). The relations between calculated and predicted values are shown in Fig. 20.
RF model
Finally, six (RF) models were generated. The models began with only two trees and three levels and increased up to three trees and five levels. Figure 21 shows the reduction in Error (%) with increasing number of Tress and layers. Accordingly, the models with four trees and four layers are considered. The developed models are graphically presented using Pythagorean Forest in Fig. 22. These arrangements leaded to a good average accuracy of (91%). The relations between calculated and predicted values are shown in Fig. 23.
Overall, the performance summary of the suction models is presented in Table 2showing the selected indices of performance evaluation such as SSE, MAE, MSE, RMSE, Error, Accuracy and R2 utilized in this research paper. Figure 24 presents the Taylor’s chart for comparing the accuracies of the developed models.
RSM model
The application used the actual factor coding Type III - Partial Sum of squares. The Model F-value of 18.04 implies that the model is significant. There is only a 0.01% chance that an F-value this large could occur due to noise. The P-values less than 0.0500 indicate model terms are significant. In this case B, C, AB, AC, BF, CF, A2, C2 are significant model terms. Values greater than 0.1000 indicate the model terms are not significant. These indications are shown in Tables 3 and 4. If there are many insignificant model terms (not counting those required to support hierarchy), model reduction may improve your model. The Predicted R2 of 0.9812 is not as close to the Adjusted R2 of 0.8289 as one might normally expect; i.e. the difference is more than 0.2. This may indicate a large block effect or a possible problem with your model and/or data. The optimized model plots for the RSM prediction of the suction pressure are presented in Figs. 25, 26, 27 and 28. Things to consider are model reduction, response transformation, outliers, etc. All empirical models should be tested by doing confirmation runs. The Adeq Precision measures the signal to noise ratio. A ratio greater than 4 is desirable. Your ratio of 19.797 indicates an adequate signal. This model can be used to navigate the design space. The Eq. (23) in terms of actual factors can be used to make predictions about the response for given levels of each factor. Here, the levels should be specified in the original units for each factor. This equation should not be used to determine the relative impact of each factor because the coefficients are scaled to accommodate the units of each factor and the intercept is not at the center of the design space.
Conclusions
This research aims to predict the logarithmic of suction pressure (Log Ψ) of granular soil treated with biochar using biochar ratio (Bio), Electric current (I), Electrical potential (U), Metric water content (θ),Temperature (T) and Bulk density (γb). Eight ML classification techniques namely GB, CN2, NB, SVM, SGD, KNN, Tree and RF and one symbolic regression technique such as the RSM were considered in this research. The outcomes of this study could be concluded as follows:
-
GB, SVM, and CN2 models showed an excellent accuracy of about 97%, while KNN, and Tree models showed very good accuracies of about 93%, SGD, and RF models showed fair accuracy level of about (90–83%) and at last NB with poor accuracy of 74%.
-
Sensitivity analysis indicated that all inputs had almost the same level of influence on the suction pressure (20–24%) except the temperature which showed less influence level (about 10%).
-
All the developed models are too complicated to be used manually, which may be considered as the main disadvantage of the ML classification techniques compared with other symbolic regression ML techniques such as GP and EPR.
-
The RSM symbolic model produced an R2 of 0.9812 with adequate precision of 19.797 which indicated an adequate signal from the model interface.
-
The developed models are valid within the considered range of parameter values, beyond this range; the prediction accuracy should be verified.
Data availability
The data supporting this research work will be made available upon reasonable request from the corresponding author.
References
Chen, B., Cai, W. & Garg, A. Relationship between bioelectricity and soil–water characteristics of biochar-aided plant microbial fuel cell. Acta Geotech. 18, 3529–3542. https://doi.org/10.1007/s11440-022-01787-z (2023).
Hussain, R., Garg, A. & Ravi, K. Soil-biochar-plant interaction: differences from the perspective of engineered and agricultural soils. Bull. Eng. Geol. Environ. 79, 4461–4481. https://doi.org/10.1007/s10064-020-01846-3 (2020).
Cai, W., Bordoloi, S., Zhu, C. & Gupt, C. B. Influence of plasticity and porewater salinity on shrinkage and water retention characteristics of biochar-engineered clays. Soil. Sci. Soc. Am. J. 87, 1285–1303 (2023).
Zhou, Y. et al. Production and beneficial impact of biochar for environmental application: a comprehensive review. Bioresour. Technol. 337, 125451. https://doi.org/10.1016/j.biortech.2021.125451 (2021).
Guo, H., Charles Wang Wai, N. G., Ni, J., Zhang, Q. & Wang, Y. Three-year field study on grass growth and soil hydrological properties in biochar-amended soil. J. Rock. Mech. Geotech. Eng. https://doi.org/10.1016/j.jrmge.2023.08.025 (2023).
Chen, Z., Kamchoom, V., Leung, A. K., Xue, J. & Chen, R. Influence of biochar on the water permeability of compacted clay subjected to freezing–thawing cycles. Acta Geophys. https://doi.org/10.1007/s11600-023-01141-1 (2023).
O’Keeffe, A. Targeted Application of Biochar in the Palouse: Modeling Hydrologic Processes in Undulating Topographies (2021).
Chen, S. et al. Amendment of straw biochar increased molecular diversity and enhanced preservation of plant-derived organic matter in extracted fractions of a rice paddy. J. Environ. Manag. 285, 112104. https://doi.org/10.1016/j.jenvman.2021.112104 (2021).
Quinn, K. Quantifying the Transport of Pathogenic and Nonpathogenic Escherichia coli in Magnesium and Nitrogen-Doped Biochar Amended Sand Columns (2021).
Geuder, D. Mitigating Agricultural Impacts from Field to STREAM: Optimizing Biochar Field Amendments and Novel Denitrifying Bioreactors for Mitigating Greenhouse Gas Emissions (2021).
Graves, C. L. L. Evaluating Acidified Miscanthus Biochar as a Broiler Litter Amendment for Ammonia Control (2023).
Onyelowe, K. C. et al. Selected AI optimization techniques and applications in geotechnical engineering. Cogent Eng. 10, 1 (2022).
Onyelowe, K. C. et al. Innovative overview of SWRC application in modeling geotechnical engineering problems. Designs 6, 69 (2022).
Aneke, F. I. et al. Predictive models of swelling stress—a comparative study between BP- and GRG-ANN. Arab. J. Geosci. 15, 1438 (2022).
Onyelowe, K. C. et al. Recycling and reuse of solid wastes; a hub for ecofriendly, ecoefficient and sustainable soil, concrete, wastewater and pavement reengineering. Int. J. Low-Carbon Technol. 14 (3), 440–451 (2019).
Bui Van, D. 2018. Adsorbed complex and laboratory geotechnics of Quarry Dust (QD) stabilized lateritic soils. Environ. Technol. Innov. 10, 355–368 (2018).
Yargicoglu, E. N. & Reddy, K. R. Biochar-amended soil cover for microbial methane oxidation: effect of biochar amendment ratio and cover profile. J. Geotech. Geoenviron. Eng. 144 (3), 04017123 (2018).
Zhu, H. et al. Assessment of the coupled effects of vegetation leaf and root characteristics on soil suction: an integrated numerical modeling and probabilistic approach. Acta Geotech. 15 (5), 1331–1339 (2020).
Liu, Y., Zhang, S. & Liu, H-H. The relationship between fingering flow fraction and water flux in unsaturated soil at the laboratory scale. J. Hydrol. 622, 129695 (2023).
Haruzi, P. & Moreno, Z. Modeling water flow and solute transport in unsaturated soils using physics-informed neural networks trained with geoelectrical data. Water Resour. Res. 59, e2023WR034538 (2023).
Lu, L., Xing, D. & Ren, Z. J. Microbial community structure accompanied with electricity production in a constructed wetland plant microbial fuel cell. Bioresour. Technol. 195, 115–121 (2015).
Hoffman, F. O. & Gardner, R. H. Chapter 11 of radiological assessment: a textbook on environmental dose analysis. In Evaluation of Uncertainties in Radiological Assessment Models (eds Till, J. E. & Meyer, H. R.) (NRC Office of Nuclear Reactor Regulation, (1983).
Gong, W. et al. Gradient boosting decision tree algorithms for accelerating nanofiltration membrane design and discovery. Desalination 592, 118072. https://doi.org/10.1016/j.desal.2024.118072 (2024).
Heymann, F. et al. Scarcity events analysis in adequacy studies using CN2 rule mining. Energy AI 8, 100154. https://doi.org/10.1016/j.egyai.2022.100154 (2022).
Zou, Z. M., Chang, D. H., Liu, H. & Xiao, Y. D. Current updates in machine learning in the prediction of therapeutic outcome of hepatocellular carcinoma: what should we know? Insights Imaging 12, 31. https://doi.org/10.1186/s13244-021-00977-9 (2021).
Lee, C. Y., Shon, J. G. & Park, J. S. An edge detection-based eGAN model for connectivity in ambient intelligence environments. J. Ambient Intell. Humaniz. Comput. 13, 4591–4600. https://doi.org/10.1007/s12652-021-03261-2 (2022).
Musolf, A. M., Holzinger, E. R., Malley, J. D. & Bailey-Wilson, J. E. What makes a good prediction? Feature importance and beginning to open the black box of machine learning in genetics. Hum. Genet. 141, 1515–1528. https://doi.org/10.1007/s00439-021-02402-z (2022).
Ahmadi, A. A., Arabbeiki, M., Ali, H. M., Goodarzi, M. & Safaei, M. R. Configuration and optimization of a minichannel using water–alumina nanofluid by non-dominated sorting genetic algorithm and response surface method. Nanomaterials 10, 901. https://doi.org/10.3390/nano10050901 (2020).
Garg, A., Onyelowe, K. C., Chen, B. & Ebid, A. Forecasting Soil-Water (Suction) Behavior of Unsaturated Soil from Bioelectricity Generated from Biochar-Improved Plant Microbial Fuel Cell (BPMFC) (2024).
Acknowledgements
The fourth author (VK) would like to acknowledge the Grant (RE-KRIS/FF68/12) from King Mongkut’s Institute of Technology Ladkrabang (KMITL) and National Science, Research and Innovation Fund (NSRF).
Funding
The fourth author (VK) would like to acknowledge the Grant (RE-KRIS/FF68/12) from King Mongkut’s Institute of Technology Ladkrabang (KMITL) and National Science, Research and Innovation Fund (NSRF).
Author information
Authors and Affiliations
Contributions
KCO Conceptualized, KCO, AME, RBRJ, VK, KPA, and MV wrote the main manuscript text and prepared the figures. RBRJ, VK, and KPA reviewed the manuscript and all authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Onyelowe, K.C., Ebid, A.M., Ramos Jiménez , R.B. et al. Modeling suction of unsaturated granular soil treated with biochar in plant microbial fuel cell bioelectricity system. Sci Rep 15, 1439 (2025). https://doi.org/10.1038/s41598-025-85701-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-85701-z