Abstract
Development and severity of alcohol use disorder (AUD) has been linked to variations in gut microbiota and their associated metabolites in both animal and human studies. However, the involvement of the gut microbiome in alcohol consumption of individuals with AUD undergoing treatment remains unclear. To address this, stool samples (n = 32) were collected at screening (baseline) and trial completion from a double-blind, placebo-controlled trial of zonisamide in individuals with AUD. Alcohol consumption was measured both at baseline and endpoint of 16-week trial period. Fecal microbiome was analyzed via 16 S rRNA sequencing and metabolome via untargeted LC-MS. Both sex (p = 0.003) and psychotropic medication usage (p = 0.025) are associated with baseline microbiome composition. The relative abundance of 11 genera at baseline was correlated with percent drinking reduction (p.adj < 0.1). Overall microbiome community structure at baseline differed between high and low reducers of alcohol drinking (67–100% and 0–33% drinking reduction, respectively; p = 0.034). A positive relationship between baseline fecal GABA levels and percent drinking reduction (R = 0.43, p.adj < 0.07) was identified by microbiome function prediction and confirmed by ELISA and metabolomics. Metabolomics analysis also found 3-hydroxykynurenine, a neurotoxic intermediate metabolite of tryptophan, was negatively correlated with drinking reduction (p.adj = 0.047), and was over-represented in low reducers. These findings highlight importance of baseline microbiome and amino acid metabolites in drinking reduction in AUD participants undergoing zonisamide treatment. It may hold significant value as a predictive tool in clinical settings to better personalize intervention and improve reduction in alcohol consumption in future.
Similar content being viewed by others
Introduction
Alcohol use disorder (AUD) persists as a major global public health challenge. AUD causes a high disease burden accounting for over 5% of annual deaths globally1,2. Chronic heavy alcohol consumption has been associated with higher risks of developing other chronic health conditions such as cancer, dementia, digestive disorders, and liver disease3,4,5,6. Recently, there has been growing interest in the bidirectional relationship between the development, progression, and treatment of AUD and the gut microbiome via the gut-brain axis.
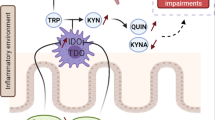
Both acute and chronic alcohol consumption are linked to a shift in microbiome composition7,8,9,10,11,12. These shifts in composition also affect the microbial production of bioactive compounds. Microbially-produced compounds such as kynurenic & quinolinic acid, short chain fatty acids (SCFAs), and γ-aminobutyric acid (GABA) influence AUD as well as commonly co-occurring biological phenotypes, such as depression, anxiety, and craving13,14,15,16,17,18,19,20. Bacterial composition in the gut of patients with AUD has been associated with plasma concentrations of tryptophan-kynurenine pathway metabolites21, which have in turn been associated with AUD16,22. The reciprocal nature of the relationship between AUD and the gut microbiome is an area of great interest, as the gut microbiome is an key mediator of not only AUD development and severity but also overall health. Restoring beneficial microbiota that are depleted by alcohol overconsumption or supplementing with compounds produced by these depleted microbes has emerged as a potential therapeutic target in AUD.
Supplementing specific compounds produced by the gut microbiota such as SCFAs or GABA has been investigated as a potential route to reduce drinking behavior but not necessarily restore the microbiota that produce these compounds. The administration of SCFAs has been shown to reduce drinking behavior in rodent models23,24,25. Gabapentin, a structural analog of GABA, has also been proposed as a potential therapeutic treatment for people with AUD26,27,28,29. While there is not a clear consensus on the nature of the connection between GABA produced in the gut and the CNS, increased expression of GABA-related metabolic pathways has been observed in the gut microbiome of patients with AUD17. Beyond compensating for depleted metabolic functionality, wholesale replacement of the gut microbiome with gut microbiota taken from a healthy donor has been shown to reduce alcohol consumption in both human and animal models30,31. While there is currently a great deal of interest in developing novel microbiome-based treatments to improve AUD outcomes, there remains a lack of understanding on the role of the gut microbiome in existing AUD treatments and intervention strategies. Understanding the influence of the gut microbiome on ability to reduce alcohol consumption could enable a more personalized and predictive approach to intervention strategies.
The baseline gut microbiome has been found to play an important role in response to cancer therapy32,33, nutritional intervention34, and efficacy of SARS-CoV-2 vaccines35. The importance of baseline gut microbiome in alcohol drinking behavior in AUD patients undergoing medication-based treatment has not been investigated. We hypothesize that the gut microbiome composition and metabolites at baseline are important factors influencing alcohol drinking behavior in participants undergoing zonisamide/placebo treatment. To test this hypothesis, we leverage a randomized, placebo-controlled, double-blind clinical trial recently completed by our research team testing the effect of zonisamide treatment of heavy drinking adults with AUD. Here, we present a secondary analysis and are not reporting direct effects of the trial intervention. Using 16 S rRNA gene sequencing and untargeted metabolomics, we showed reduction in alcohol consumption between baseline and endpoint visits of the clinical trial was significantly correlated with composition of the baseline microbiome and levels of specific gut metabolites such as GABA and tryptophan-kynurenine pathway intermediates. Participants who achieved a high level of drinking reduction during the trial (high reducers) had a distinct gut microbiome profile and specific gut metabolic signatures related to tryptophan metabolism and GABA, compared to participants who achieved a lower level of drinking reduction (low reducers). Our findings highlight importance of the baseline gut microbiome composition and metabolites in alcohol consumption and the potential for development of a microbiome signature to predict treatment response for AUD.
Results
Study participants
The demographic and clinical characteristics and relevant alcohol-related measures of the 32 participants with AUD included in the analysis are reported in Table 1. These participants were enrolled at UConn Health, one site of the multi-site trial (ClinicalTrials.gov identifier: NCT02900352, Zonisamide Treatment of Alcohol Use Disorder: an Evaluation of Efficacy and Mechanism of Action (Z-Comp), first registered 14/09/2016). No significant differences in demographic and clinical variables were observed between the zonisamide and placebo treatment groups. Participants were overall 44% female. Participant age ranged from 23 to 70 years with a median age of 56. More than half of the participants identified as white/non-Hispanic. In total, all 32 participants provided baseline stool samples. Twenty-one participants provided an endpoint stool sample. It is vital to note that we have performed a separate secondary analysis and are not reporting any effects of the trial intervention.
In the current analysis, treatment (placebo group vs. zonisamide group) did not influence percent drinking reduction reported at study endpoint, calculated from average drinks per week at baseline and average drinks per week over the last four weeks of the study (t-test, p = 0.27). Separating participants into male and female groups did not change the results for percent drinking reduction (t-test, male p = 0.40, female p = 0.41) or endpoint average drinks per week over the last four weeks of the study (t-test, male p = 0.24, female p = 0.46). Percent drinking reduction was not significantly correlated with average drinks per week at baseline by Pearson correlation (p = 0.09) but was significantly correlated with average drinks per week during the last four weeks of the study (p = 6.9E-10). No other significant correlations were found between percent drinking reduction and other study variables.
Baseline microbiome composition is sex- and medication-linked
We first examined whether any demographic or clinical factors are associated with the baseline microbiome. Clear separation was observed in a PCoA plot of male and female microbiome (Fig. 1a and c). This difference was confirmed by PERMANOVA analysis (p = 0.003, R2 = 0.051, Fig. 1a). Linear discriminant analysis effect size analysis (LEfSe)36 detected 10 genera showing significant differences in relative abundance between female and male participants (Fig. 1b). Bacteroides (LDA Score = 5.44, p = 0.013) and Blautia (LDA Score = 5.52, p = 0.030) were the most overrepresented genera in female participants at baseline. Male participant microbiomes showed an overrepresentation of Enterococcus (LDA Score = 4.67, p = 0.043), UCG-003 (LDA Score = 3.01, p = 0.049), and Veillonella (LDA Score = 2.9, p = 0.025).
Associations between baseline gut microbiome and clinical characteristics of the participants in the study. (a) Baseline microbiome composition varies based on sex with male and female participants clustering separately in the PCoA plot determined from Bray-Curtis dissimilarity (p = 0.003, R2 = 0.051 by PERMANOVA). (b) Ten genera are overrepresented in male or female participant microbiomes. (c) Baseline microbiome composition also varies based on psychotropic medication usage controlling for sex (PERMANOVA, p = 0.025, R2 = 0.034). (d) Ten genera are overrepresented in psychotropic medication users and non-users.
We also found the microbiome in psychotropic medication users clustered separately from non-users (Fig. 1c). PERMANOVA analysis indicated overall microbiome difference between these two groups (p = 0.025, R2 = 0.034) controlling for sex, as female participants were more likely to report using psychotropic medication (Fisher’s exact test, p = 0.040). LEfSe analysis revealed that psychotropic medication promoted overrepresentation of 8 genera (Fig. 1d). Only 2 genera were overrepresented in psychotropic medication non-users. Given that both sex and psychotropic medication usage were associated with significant changes in microbiome composition, subsequent analysis controlled for both sex and psychotropic medication in addition to treatment groups.
Baseline Microbiome is associated with drinking reduction
Results from the 32 participants that completed the study were used to explore correlation between baseline microbiome and alcohol consumption at both baseline and endpoint as well as drinking reduction over the course of the study after controlling for treatment, sex, psychotropic medication and adjusting for multiple comparisons (Fig. 2a). Eubacterium brachy group (p.adj = 0.06), Collinsella (p.adj = 0.07), Eisenbergiella (p.adj = 0.09), and Marvinbryantia (p.adj = 0.09) were positively correlated with percent drinking reduction. Lachnospiraceae ND3007 group (p.adj = 0.02), Faecalibaterium (p.adj = 0.02), Eubacterium ventriosum group (p.adj = 0.07), Fusicatenibacter (p.adj = 0.09), Roseburia (p.adj = 0.03), Monoglobus (p.adj = 0.03), and CAG-56 (p.adj = 0.03) were all negatively correlated with percent drinking reduction. Notably, a majority of identified taxa had a consistent correlation direction with both drinking reduction and average drinks per week at baseline, although the correlation may not reach to statistical significance. This trend in correlation directionality was opposite with average drinks per week at endpoint.
Baseline gut microbiome composition is associated with drinking reduction. (a) Genus-level relative abundance is correlated with percent drinking reduction over the study duration, baseline and endpoint alcohol consumption. Colors of the heatmap represent correlation coefficients derived from the multiple linear regression analysis. An adjusted p value is denoted in the cell of the heatmap if the adjusted p value is less than 0.1 for a given correlation analysis. (b) Percent drinking reduction is largely varied across participants and can be broken into tertiles. (c) The highest (percent drinking reduction 67–100%) and lowest (percent drinking reduction 0–33%) tertiles cluster separately in PCoA plot determined from Bray-Curtis dissimilarity (PERMANOVA p = 0.034). (d) 11 genera are significantly overrepresented in the high- and low reducer groups.
Baseline microbiome composition differs between high and low reducers
Percent drinking reduction showed a high inter-subject variation (Fig. 2b). To have a better understanding of the involvement of the microbiome in the “high vs. low reducers”, the spread of percent drinking reduction across participants was broken into tertiles. Of the 32 completing participants, 10 had a “high reduction” classified henceforth as percent drinking reduction of 67–100% (Fig. 2b, blue points). Seven participants had a “low reduction” classified henceforth as percent drinking reduction of 0–33% (Fig. 2b, orange points). Fifteen participants fell between 34 and 66% drinking reduction and were considered “medium reducers.” Compared to the medium reducer group, the high reducer group and low reducer group had significantly fewer female participants (Fisher’s test, p = 0.01) and significantly lower BMI (Fisher’s test, p = 0.008). No other clinical variables differed between different reducer groups (Supporting Table 1).
Baseline microbiome composition as described by Bray-Curtis dissimilarity varied significantly between the high and low reducers (PERMANOVA, p = 0.018) (Fig. 2c). Alpha diversity was consistent across reduction tertiles (Supporting Fig. 1a). LefSE identified 7 genera that were significantly overrepresented in low reducers and 4 genera that were significantly overrepresented in high reducers (Fig. 2d). Many of the genera identified have previously been associated with alcohol consumption. CAG-56 was significantly overrepresented in low reducers (p.adj = 0.034, Fig. 2d) and showed a negative association trend with percent drinking reduction (p.adj = 0.03, Fig. 2a). Eisenbergiella was overrepresented in high reducer baseline microbiome (p.adj = 0.015, Fig. 2d) and positively associated with percent drinking reduction (p.adj = 0.09, Fig. 2a). Additional comparisons between low vs. medium reducers and medium vs. high reducers identified differential genera with consistent trends in terms of their relative abundance and drinking reduction mostly (Supporting Fig. 1b, c).
Baseline GABA levels in the stool are correlated with drinking reduction
The gut microbiome has been shown to modulate production of many neurotransmitters involved in AUD17,21,37,38. To understand the potential involvement of neuroactive potential of the gut microbiome in drinking response, we predicted neuroactive compound production or degradation process based on the gut microbiome using the gut-brain module (GBM) analysis39,40. We found a positive linear correlation between percent drinking reduction and potential for γ-aminobutyric acid (GABA) metabolism in the baseline microbiome (Fig. 3a, R = 0.439, p.adj = 0.08). GABA has been implicated in mediating craving and severity of alcohol use disorder and identified as a potential target for intervention2,41. Baseline stool GABA concentration was also directly quantified via ELISA and was found to be positively correlated with drinking reduction controlling for sex, psychotropic medication usage, and treatment (Fig. 3b, R = 0.42, p.adj = 0.07), recapitulating findings from the predicted metagenome functionality.
Baseline gut metabolites are associated with percent drinking reduction. (a) GABA metabolism potential as identified from predictive gut-brain module analysis shows a positive linear relationship with percent drinking reduction controlling for sex, psychotropic medication usage, and treatment (R = 0.439, p.adj = 0.08). (b) This relationship is recapitulated by direct measurements of stool GABA by an ELISA assay (R = 0.42, p.adj = 0.07). (c) Stool metabolites are correlated with percent drinking reduction. Colors of the heatmap represent correlation coefficients derived from the multiple linear regression analysis controlling for sex, psychotropic medication usage, and treatment. An adjusted p value is denoted in the cell of the heatmap if the adjusted p value is less than 0.1 for a given correlation analysis. (d) Baseline stool metabolic pathways associated with high (n = 7) and low (n = 6) reducers. Log2-fold change of metabolites that are significantly different between high and low reducers were used to construct metabolic pathways using Qiagen IPA software. Dot color indicates the group: a blue dot indicates that the pathway is overrepresented in high reducers, orange indicates overrepresentation in low reducers. Dot size corresponds to pathway z-score magnitude (i.e. larger dot size corresponds to higher pathway representation).
Baseline stool amino acid metabolites are correlated with drinking reduction
Given the strong association between baseline stool GABA concentration and drinking reduction, we next analyzed the baseline stool metabolome to determine additional correlations with alcohol consumption. Baseline stool aliquots from 21 participants were subjected to untargeted metabolomics via LC-MS. Annotated stool metabolite levels were used to explore correlations between baseline stool metabolites and drinking-related measures. LC-MS confirmed the positive correlation between GABA and percent drinking reduction (p.adj = 0.002, Supporting Fig. 3). In addition, 2-pyrrolidone, a biologically relevant cyclization product of GABA42, was also positively correlated with percent drinking reduction (p.adj = 0.002, Supporting Fig. 3). However, correlations become non-significant after controlling for sex, psychotropic medication usage, and treatment (GABA, p.adj = 0.18; 2-pyrrolidone, p.adj = 0.24). Twenty-eight metabolites were significantly correlated with percent drinking reduction after controlling for sex, psychotropic medication usage, and treatment (Fig. 3c). The majority of identified metabolites were negatively correlated with percent drinking reduction and belonged to indeterminate metabolic pathways. Notably, tryptophan-kynurenine pathway intermediate 3-hydroxykynurenine (p.adj = 0.043, Fig. 3c) was negatively correlated with percent drinking reduction after controlling for sex, psychotropic medication usage, and treatment.
Stratifying the metabolome results based on percent drinking reduction allowed us to determine metabolites that varied between high and low reducers. Among 1,788 metabolites that had CV < 20 and corresponding PubChem database entries in the analysis, seventy metabolites were found to differ between high and low reducers (p.adj < 0.05, Wilcoxon rank-sum). We next performed metabolic pathway analysis using the log2-fold change of significantly differing metabolites in high reducers with respect to low reducers (Fig. 3d). Pathways differed significantly between high (n = 7) and low (n = 6) reducers were related to degradation and biosynthesis of several amino acids including tryptophan, histidine, glycine, and alanine. Tryptophan catabolism was highly overrepresented in the low reducer group (p.adj = 6.6E-5), consistent with the predicted metagenome functional analysis (Supporting Fig. 2, p.adj = 0.08). In addition, glutaminergic receptor signaling was overrepresented in the low reducer metabolome (p.adj = 0.083) as compared with high reducers. The directionality of pathway representation is largely consistent when comparing across other reduction groups (medium vs. high and low vs. medium), (Supporting Fig. 4).
Zonisamide treatment has minor impact on the microbiome composition
Six zonisamide-treated participants and 15 placebo-treated participants provided stool samples at both baseline and the endpoint visit, which allows us to explore whether there was any effect of zonisamide treatment on the gut microbiome. The mean relative abundance of top 20 most abundant genera is similar between groups or across the study durations (Supporting Fig. 5). PCoA analysis did not show distinct clustering within treatment groups across the study duration (Supporting Fig. 6a, b) or between treatment groups at baseline and endpoint (Supporting Fig. 6c, d). PERMANOVA analysis supported no overall microbiome difference between the two groups (p > 0.1). Alpha and beta diversity also show no difference between groups (p > 0.1 for all comparisons, Supporting Fig. 7). Specific genera analysis by linear mixed regression after controlling for sex, psychotropic medication usage, and average number of drinks per week at baseline identified several genera that are influenced by treatment, including Eisenbergiella, Sellimonas, and Intestinibacter which were in low abundance in general (Supporting Fig. 8).
Discussion
There is great interest to better understand the varied response in AUD interventions and to understand the influence of the gut microbiome in AUD. Baseline gut microbiome (before treatment) has been found to be a powerful influence on outcome in major depressive disorder treatment43, cancer therapy32,33, lifestyle intervention in obesity34, rheumatoid arthritis treatment44, and efficacy of SARS-CoV-2 vaccines35. Given the deep connections between microbiome and mental health via the gut-brain-axis and the growing body of evidence for the strength of the microbiome’s influence in determining treatment outcome, we sought to characterize the connections between the microbiome and changes in drinking behavior. Our study revealed important associations between baseline microbiome composition and function with percent drinking reduction in participants from a zonisamide clinical trial. These findings are a sub-study and are separate from the effects of the trial intervention, though we have accounted for medication effects on microbiome composition and function.
While reviews and meta-analyses have identified many genera that are commonly altered by AUD8,37,45, the implications of these microbes in treatment outcome are not fully characterized. Collinsella was found to be positively correlated with drinking reduction in this study and has been found to be both enriched17 and depleted45 in AUD. Akkermansia and Faecalibacterium are both largely agreed to be depleted in the AUD microbiome relative to healthy controls17,46. While Akkermansia and Faecalibacterium are most often been discussed as a beneficial microbe and target for probiotic supplementation47, recent study has indicated that Akkermansia may be detrimental in the context of neuropsychiatric disorders48,49. In this study, we found that Akkermansia and Faecalibacterium are negatively correlated with percent drinking reduction suggesting that a lower abundance of either or both genera may be beneficial for reduced alcohol consumption.
The ideal composition of the gut microbiome is very difficult to conceptualize given the highly individual and fluctuating nature of the gut microbiome. However, microbial metabolism of the collective gut microbiome is relatively conserved due to multiplicity of function and can lend insight into the various metabolic niches that individual microbes may occupy. GABA is produced by the gut microbiome50,51 and is also vital in the context of AUD19,28. Existing evidence suggests that the microbial GABA production may contribute to circulating GABA levels, though it is believed that GABA itself does not cross the blood brain barrier. Microbial GABA more likely indirectly influences the brain through the vagus nerve without entering circulation50. Microbial GABA production has an unclear correlation to brain GABA concentrations40,51. The GABA present in the gut may also arise from dietary sources, but dietary GABA is relatively low52. In our study, we consistently identified the positive correlation between baseline GABA and drinking reduction using three independent approaches, highlighting importance of gut-derived GABA in association with alcohol consumption. In this cohort, the relationship between stool GABA and drinking reduction was influenced strongly by sex (p < 0.01). This relationship may arise from the sex-dependence of the baseline microbiome composition (Fig. 1a), especially given that GABA is produced in the gut microbiome. It is critical to reaffirm that the GABA levels were measured in baseline stool prior to the start of GABAergic zonisamide treatment and, thus, were not influenced by treatment. Looking at the baseline microbiome production of GABA prior to starting an intervention can provide valuable information on treatment response.
In addition to GABA, many of the metabolic changes that were correlated with drinking reductions in this study involve amino acids. Amino acid absorption and metabolism is known to shift in AUD53,54. Imbalance of amino acids can be due to malnutrition, liver dysfunction, or alcohol-induced metabolic disturbances. Tryptophan, the precursor of serotonin, can be degraded to kynurenine. The latter can be further catabolized to produce neurotoxic quinolinic acid and neuroprotective kynurenic acid. 3-hydroxykynurenine is an intermediate metabolite of quinolinic acid and endogenous neurotoxin that causes neuronal cell death55. Tryptophan‐kynurenine metabolism is disrupted in AUD21. Meta analysis revealed a down-regulation kynurenic acid in individuals with AUD22. We found that tryptophan degradation was different between high and low reducer groups, and 3‐hydroxykynurenine was negatively correlated with drinking reduction, suggesting these metabolites are not only altered in AUD but can potentially impact drinking reduction in individuals with AUD. Future studies are needed to characterize each metabolite involved in tryptophan metabolic pathway to comprehensively understand the role of tryptophan metabolism in drinking behavior in individuals with AUD.
Another significantly changed metabolic pathway is alanine metabolism. While alanine concentration in serum was increased in AUD54, intestinal absorption of alanine is inhibited by ethanol53, which is consistent with our observations. Overrepresentation of two alanine degradation-related pathways in low reducers may be in response to an increased alanine concentration. Conversely, the overrepresentation of an alanine biosynthesis pathway in high reducers may be in response to a comparatively lower alanine concentration. It is noteworthy that the shared nature of stool metabolites complicates separation of host and microbial metabolic pathways. The changes in amino acid metabolism between reduction groups may be due to differing intestinal permittivity, host metabolism, microbial metabolism, or most likely a combination of these factors.
The gut microbiome is often influenced by medication usage56. However, there was no net change in microbiome alpha- or beta-diversity with zonisamide treatment across the study. Our analysis identified several individual genera that appeared to be affected by zonisamide treatment, but the effects were dominated by only a few individual participants and were not consistent across participants. Zonisamide is rapidly absorbed in the gastrointestinal tract and very little is eliminated in feces57. While the exact location of zonisamide absorption is not known, it is most likely absorbed in the small intestine like other oral medications. Together the reported rapid and near-complete absorption, low fecal levels, and likely absorption site, the microbiome in the colon may not be exposed to high concentrations of zonisamide and, thus, may not have a strong shift in response to administration. Zonisamide treatment may have an indirect effect on microbiome composition due to the commonly reported gastrointestinal side effects. However, as only six zonisamide-treated participants provided stools at endpoint, these findings need to be taken as exploratory.
This study presents several limitations. Small sample size is a consistent issue in microbiome-related work given the large intra-individual variation in microbiome composition. Future studies with larger sample size from multiple clinical sites are warranted to verify the relationship between baseline microbiome and alcohol consumption. Additionally, collecting detailed information on diet and antibiotic use would provide additional control variables. In this study, daily diet recall and antibiotic use were not collected, and smoker status was limited to nicotine and does not include marijuana. The use of other drugs was not quantified, though opioid or benzodiazepine dependence were exclusion criteria. Lastly, the study is an association study, and we cannot conclude a causal relationship between the baseline microbiome and alcohol drinking reduction. Animal studies such as fecal microbiota transplantation from stools of high and low reducers will allow us to gain mechanistic understanding on contribution of the baseline microbiome in reducing alcohol consumption. Manipulation of transplanted human microbiome through probiotic supplementation could enable the development of personalized interventional treatment based on microbiome characteristics.
In conclusion, our study identified important associations between baseline gut microbiome composition, amino acid metabolites, GABA, and tryptophan with alcohol consumption reduction in a zonisamide clinical trial. Stool collection is simple and non-invasive. Screening baseline gut microbiome composition and metabolites may hold significant value as a predictive tool in clinical settings to better personalize intervention and improve reduction in alcohol consumption, durability of behavioral changes, and ultimately participant outcome.
Materials and methods
Human trial
This study was conducted with approval from the IRB of Virginia Commonwealth University and UConn Health in accordance with the relevant guidelines and regulations. Participants were recruited from the community as part of a double-blind, randomized, placebo-controlled study investigating the use of zonisamide in reducing drinking (ClinicalTrials.gov identifier: NCT02900352, Zonisamide Treatment of Alcohol Use Disorder: an Evaluation of Efficacy and Mechanism of Action (Z-Comp), first registered 14/09/2016). The study protocol was published online. Inclusion criteria limited participants to ages 21–70 who had regular heavy drinking, a current DSM-5 diagnosis of alcohol use disorder, and a desire to reduce or stop drinking. Potential participants who were currently lactating or with clinically significant physical disease, seizure disorder, use of any medications that could affect drinking or cause harm, schizophrenia, bipolar disorder, substantial risk of suicide or violence, opioid or benzodiazepine dependence were excluded. Women of child-bearing age were required to practice a reliable method of birth control. Participants gave written informed consent to participate in the study and were financially compensated. Participants were randomized into treatment and control groups matching for sex and current psychotropic drug usage. Medications such as serotonin selective reuptake inhibitors, serotonin norepinephrine reuptake inhibitors, norepinephrine dopamine reuptake inhibitors, and low dose benzodiazepines were considered psychotropic drugs in this study. The treatment group received flexibly titrated zonisamide over 7 weekly visits starting at 100 mg daily and increasing over the 8 weeks to a 500 mg daily maximum for the remaining 8 weeks of the study. Medical management58 served as a psychosocial intervention platform. Timeline Follow-back Method59 was used to measure self-reported drinking including number of drinking days during the 90-day pretreatment period and at each visit.
Fisher’s exact test was used to test the difference in demographic and clinical categorical variables between zonisamide and placebo groups (Table 1). Two-sample t-test was used for testing continuous demographic and clinical variables. Percent drinking reduction was quantified as the difference between self-reported drinks per week during the last 90 days at baseline and average drinks per week over the last four weeks of the study as calculated at study endpoint divided by the baseline self-reported drinks per week during the last 90 days.
16 S rRNA sequencing of stool samples
Stools were collected by participants at UConn Health site and stored on ice up to 24 h prior to baseline and 16-week (endpoint) visits. Stool aliquots were prepared upon receipt at the clinical research center at UConn Health and stored at -80℃ until the time of analysis. Microbial DNA was isolated from stool samples using the PowerSoil DNA Isolation kit (Qiagen) following manufacturer’s instructions. Bacterial 16 S ribosomal RNA (rRNA) gene sequencing was performed on V4 hypervariable regions using 515 F (5’- GTGYCAGCMGCCGCGGTAA-3’) and 806R (5’- GGACTACNVGGGTWTCTAAT-3’) primers to prepare an amplicon library that was purified using Zymo Select-a-Size MagBeads (Zymo), quantified (Qubit 2.0 fluorimeter, Invitrogen), and pooled with equal masses added from each sample. Two additional cleanup steps were performed on the initial pool again using Zymo Select-a-Size MagBeads (Zymo). The pooled and purified library was sequenced on the Illumina MiSeq platform (Illumina) using 2 × 250 bp, 500 cycles kits.
16 S rRNA data processing and analysis
Raw 16 S rRNA sequencing reads were initially processed by bcl2fastq2 (v2.20) and RTA (v1.18.54.4) software (Illumina). Mean read count was 60,460. Minimum read count was 12,241. Maximum read count was 60,813. Demultiplexed fastQ files were imported into the QIIME2 pipeline (version 2022.11)60. The DADA261 plugin was used to denoise reads and remove chimeras using the consensus method. Forward and reverse reads were truncated at position 250. All other parameters were set to default. The phylogeny was inferred using the align-to-tree-mafft-fasttree pipeline in QIIME2. Taxonomy was assigned with pre-trained naïve Bayesian classifier based on the SILVA reference database V138.1 using the q2-feature-classifier plugin with a 0.5 confidence value cut-off.
Subsequent analysis of 16 S rRNA sequencing data was done in R (version 4.3.1) using RStudio interface (version 2023.06.1) and Qiime2R62 and phyloseq63 packages. ASV counts were aggregated at various taxonomic levels (i.e. genus-level) and converted to relative abundance using phyloseq63 and MicroViz64 packages. PERMANOVA (permutational multivariate analysis of variance) was performed using the Adonis function in vegan package65 to evaluate differences in beta diversity across metadata variables. Principal coordinate analysis and visualization with 90% confidence intervals (stat_ellipse, ggplot266) were generated using microViz64, ggplot266, and tidyverse67 packages in R. Correlations between genus-level relative abundance and alcohol drinking measures were tested using multiple linear regression controlling for sex, psychotropic medication usage, treatment group (zonisamide vs. placebo), and adjusting p-values for multiple comparisons using false discovery rate. Linear discriminant analysis Effect Size (LEfSe)36 was performed using the microbial package in R. Percent change in genus-level relative abundance with treatment was performed using linear mixed regression controlling for sex, psychotropic medication usage, and adjusting p-values for multiple comparisons using false discovery rate. An adjusted p-value < 0.1 was considered statistically significant.
Gut-Brain module analysis based on 16 S data
The ASVs for samples of interest were exported from R and used for subsequent phylogenetic investigation of communities by reconstruction of unobserved states (PICRUSt) using PICRUSt239. The PICRUSt2 pipeline was run using picrust2_pipeline.py and add_descriptions.py. The resultant unstratified KO metagenome predictions and their associated descriptions was subsequently used for predictive functional analysis via gut-brain modules40. Correlations between number of hits in each module and clinical variables were tested using multiple linear regression with a significance cutoff of p-value < 0.1 after controlling for sex, psychotropic medication usage, treatment, and adjusting for multiple comparisons. ANCOVA was used to compare the differences number of hits in each module between high reducer (67–100% drinking reduction) and low reducer (0–33% drinking reduction) participants with a significance cutoff of p-value < 0.1 after controlling for sex, psychotropic medication usage, treatment, and adjusting for multiple comparisons.
Untargeted LC-MS analysis of stool metabolome
~ 20 mg aliquots of stool samples from 21 participants that provided both baseline and endpoint stools were subjected to untargeted LC-MS metabolomic analysis. Stool samples were homogenized in homogenization buffer (80% methanol in PBS with 1.8105 mM 13C3-lactate and 142 µM 13C5-glutamic acid) prior to the addition of 800 µL homogenization buffer. Homogenized samples were incubated 30 min at -20℃ and subsequently sonicated for 30 min on ice. Debris was pelleted via centrifugation and 800 µL supernatant was dried under vacuum (CentriVap Concentrator, Labconco). The dried residue was suspended in 150 µL 40% PBS/60% acetonitrile. A quality control sample was pooled from all study samples.
The untargeted LC-MS metabolomic method was adapted from previously published methods68,69,70,71. In summary, each sample was injected twice (10 µL for negative ionization mode, 4 µL for positive ionization mode) onto an XBridge BEH Amide column (150 × 2.1 mm, 2.5 μm particle size, Waters) maintained at 40℃. Samples were maintained in an autosampler at 4℃. Mobile phase flow rate was 0.3 mL/min and was composed of MP A (5% acetonitrile in water, 10 mM ammonium acetate and ammonium hydroxide) and MP B (95% acetonitrile in water, 10 mM ammonium acetate and ammonium hydroxide). The mobile phase gradient is as follows: 1 min isocratic elution, 90% MP B; 10 min ramp to 40% MP B; 4 min hold at 40% MP B; ramp to 90% MP B prior to next injection. Untargeted data was collected from 70 to 1050 m/z using Thermo Vanquish UPLC-Exploris 240 Orbitrap MS instrument (Thermo Scientific) equipped with an electrospray ionization source.
Data were processed using Thermo Compound Discover 3.3 software (Thermo Scientific) for peak picking, alignment, and normalization. Only peaks with CV < 20% across quality control pools appearing in > 80% of all samples were included in all subsequent analysis. Identifications and annotations used available data for retention time, exact mass, and fragmentation & isotopic patterns. Data extraction absolute intensity threshold was 1,000 and mass accuracy limit was 5 ppm. Peaks in the obtained MS spectra were annotated using an extensive in-house library of ~ 600 aqueous metabolites in addition to the HMDB library, LIPID MAPS database72, METLIN database73, ChemSpider database74 and commercial databases (mzCloud (HighChem LLC), Metabolika (Thermo Scientific)). Metabolites were scored using the tiering system75. In total 130 metabolites were considered tier 1, 3290 considered tier 2, and 994 considered tier 3. Annotated metabolites were used for downstream analysis in R and Ingenuity Pathway Analysis (Qiagen). Correlations between normalized peak intensity and alcohol drinking measures were tested using multiple linear regression with a significance cutoff of p-value < 0.1 after controlling for sex, psychotropic medication usage, and treatment group (zonisamide vs. placebo), and adjusting p-values for multiple comparisons using false discovery rate. A subset of annotated metabolites with CV > 20 that had PubChem database ID numbers (n = 1789) were used for pathway analysis. Log2-fold change of metabolites in high reducers with respect to low reducers, high- with respect to medium reducers, medium with respect to low reducers, and corresponding p-values (Wilcoxon rank-sum test) were provided to Qiagen IPA software.
Stool GABA quantitation
~ 100 mg aliquots of stool samples from 31 participants that completed the study were analyzed for GABA content using an ELISA kit (LDN, Nordhorn, Germany) following manufacturer’s instructions. Stool was thawed and homogenized in 300 µL lysis solution (0.01 N HCl, 1 mM EDTA, & 4 mM sodium metabisulfite) (Thermo Fisher Scientific). Homogenized fecal slurry was clarified by centrifugation at 5000 g for 10 min at 4℃ prior to subsequent use. In brief, clarified fecal slurry and standards (provided by LDN) were extracted, derivatized, and incubated with antiserum. Derivatized samples and standards were subjected to a quantitative ELISA read at 450 nm in a 96-well plate reader (iMark, Biorad). Absorbance of derivatized standards was used to generate a standard curve that was used to quantify the experimental samples. Stool GABA content was normalized to initial input sample weight. Correlation between percent drinking reduction and stool GABA content was tested using multiple linear regression with a significance cutoff of p-value < 0.1 after controlling for sex, psychotropic medication usage, treatment, and adjusting for multiple comparisons.
Data availability
Raw sequencing reads from the 16 S amplicon sequencing are available via the National Center for Biotechnology Information (NCBI) Short Read Archive (SRA) under accession #PRJNA1065830.
References
WHO. Global status report on alcohol and health 2018. World Health Organisation. 478 (2018).
Carvalho, A. F., Heilig, M., Perez, A., Probst, C. & Rehm, J. Alcohol use disorders. Lancet 394, 781–792 (2019).
Schwarzinger, M., Thiébaut, S. P., Baillot, S., Mallet, V. & Rehm, J. Alcohol use disorders and associated chronic disease – a National retrospective cohort study from France. BMC Public Health 18, 43 (2018).
Rehm, J. et al. The relationship between different dimensions of alcohol use and the burden of disease—an update. Addiction 112, 968–1001 (2017).
Wu, L. T., Zhu, H. & Ghitza, U. E. Multicomorbidity of chronic diseases and substance use disorders and their association with hospitalization: results from electronic health records data. Drug Alcohol Depend. 192, 316–323 (2018).
Shield, K. D., Parry, C. & Rehm, J. Chronic diseases and conditions related to alcohol use. Alcohol Res. 35, 155–173 (2013).
Leclercq, S. et al. Intestinal permeability, gut-bacterial dysbiosis, and behavioral markers of alcohol-dependence severity. Proc. Natl. Acad. Sci. 111, E4485–E4493 (2014).
Day, A. W. & Kumamoto, C. A. Gut Microbiome dysbiosis in alcoholism: consequences for health and recovery. Front. Cell. Infect. Microbiol. 12, 198 (2022).
Litwinowicz, K. & Gamian, A. Microbiome alterations in alcohol use disorder and alcoholic liver disease. Int. J. Mol. Sci. 24, 2461 (2023).
Engen, P. A., Green, S. J., Voigt, R. M., Forsyth, C. B. & Keshavarzian, A. The Gastrointestinal microbiome: alcohol effects on the composition of intestinal microbiota. Alcohol Res. 37, 223 (2015).
Mutlu, E. A. et al. Colonic Microbiome is altered in alcoholism. Am. J. Physiology-Gastrointestinal Liver Physiol. 302, G966–G978 (2012).
Bjørkhaug, S. T. et al. Characterization of gut microbiota composition and functions in patients with chronic alcohol overconsumption. Gut Microbes 10, 663–675 (2019).
Taylor, A. M. & Holscher, H. D. A review of dietary and microbial connections to depression, anxiety, and stress. Nutr. Neurosci. 23, 237–250 (2020).
Meckel, K. R. et al. Microbial short-chain fatty acids regulate drug seeking and transcriptional control in a model of cocaine seeking. Neuropsychopharmacology 49, 386–394 (2023).
Davidson, M. et al. Tryptophan and substance abuse: mechanisms and impact. Int. J. Mol. Sci. 2023. 24, 2737 (2023).
Vidal, R. et al. Plasma Tryptophan and kynurenine pathway metabolites in abstinent patients with alcohol use disorder and high prevalence of psychiatric comorbidity. Prog Neuropsychopharmacol. Biol. Psychiatry. 102, 109958 (2020).
Addolorato, G. et al. Gut microbiota compositional and functional fingerprint in patients with alcohol use disorder and alcohol-associated liver disease. Liver Int. 40, 878–888 (2020).
Liang, J. & Olsen, R. W. Alcohol use disorders and current Pharmacological therapies: the role of GABAA receptors. Acta Pharmacol. Sinica 2014. 35, 981–993 (2014).
Dharavath, R. N. et al. GABAergic signaling in alcohol use disorder and withdrawal: pathological involvement and therapeutic potential. Front. Neural Circuits 17, 1218737 (2023).
Addolorato, G., Leggio, L., Abenavoli, L. & Gasbarrini, G. Neurobiochemical and clinical aspects of craving in alcohol addiction: A review. Addict. Behav. 30, 1209–1224 (2005).
Leclercq, S., Schwarz, M., Delzenne, N. M., Stärkel, P. & de Timary, P. Alterations of kynurenine pathway in alcohol use disorder and abstinence: a link with gut microbiota, peripheral inflammation and psychological symptoms. Transl Psychiatry 11, 503 (2021).
Wang, Z., Huang, S., Li, L., Wen, Y. & Shang, D. Kynurenine metabolite changes in individuals with alcohol use disorder: A systematic review and meta-analysis. Drug Alcohol Depend. 249, 110821 (2023).
Gao, L., Davies, D. L. & Asatryan, L. Sodium butyrate supplementation modulates neuroinflammatory response aggravated by antibiotic treatment in a mouse model of Binge-like ethanol drinking. Int. J. Mol. Sci. 23, 15688 (2022).
Bokoliya, S. C. et al. Short-chain fatty acid valerate reduces voluntary alcohol intake in male mice. Microbiome 12, 108 (2024).
Reyes, R. E., Gao, L., Zhang, Z., Davies, D. L. & Asatryan, L. Supplementation with sodium butyrate protects against antibiotic-induced increases in ethanol consumption behavior in mice. Alcohol 100, 1–9 (2022).
Mariani, J. J. et al. Pilot randomized placebo-controlled clinical trial of high-dose Gabapentin for alcohol use disorder. Alcohol Clin. Exp. Res. 45, 1639–1652 (2021).
Prisciandaro, J. J. et al. Effects of Gabapentin on dorsal anterior cingulate cortex GABA and glutamate levels and their associations with abstinence in alcohol use disorder: A randomized clinical trial. Am. J. Psychiatry. 178, 829–837 (2021).
Cheng, Y. C., Huang, Y. C. & Huang, W. L. Gabapentinoids for treatment of alcohol use disorder: A systematic review and meta-analysis. Hum. Psychopharmacology: Clin. Experimental. 35, 1–11 (2020).
Anton, R. F. et al. Efficacy of Gabapentin for the treatment of alcohol use disorder in patients with alcohol withdrawal symptoms: A randomized clinical trial. JAMA Intern. Med. 180, 728–736 (2020).
Bajaj, J. S. et al. A randomized clinical trial of fecal microbiota transplant for alcohol use disorder. Hepatology 73, 1688–1700 (2021).
Wolstenholme, J. T. et al. Reduced alcohol preference and intake after fecal transplant in patients with alcohol use disorder is transmissible to germ-free mice. Nature Communications 13, 6198 (2022).
Chaput, N. et al. Baseline gut microbiota predicts clinical response and colitis in metastatic melanoma patients treated with ipilimumab. Ann. Oncol. 28, 1368–1379 (2017).
Heshiki, Y. et al. Predictable modulation of cancer treatment outcomes by the gut microbiota. Microbiome 8, 1–14 (2020).
Hughes, R. L., Kable, M. E., Marco, M. & Keim, N. L. The role of the gut Microbiome in predicting response to diet and the development of precision nutrition models. Part II: results. Adv. Nutr. 10, 979–998 (2019).
Daddi, L. et al. Baseline gut Microbiome signatures correlate with immunogenicity of SARS-CoV-2 mRNA vaccines. Int. J. Mol. Sci. 24, 11703 (2023).
Segata, N. et al. Metagenomic biomarker discovery and explanation. Genome Biol. 12, R60 (2011).
Hofford, R. S. & Kiraly, D. D. Clinical and preclinical evidence for gut Microbiome mechanisms in substance use disorders. Biol. Psychiatry. 95, 329–338 (2024).
O’Mahony, S. M., Clarke, G., Borre, Y. E., Dinan, T. G. & Cryan, J. F. Serotonin, Tryptophan metabolism and the brain-gut-microbiome axis. Behav. Brain. Res. 277, 32–48 (2015).
Douglas, G. M. et al. PICRUSt2 for prediction of metagenome functions. Nat. Biotechnol. 38, 685–688 (2020).
Valles-Colomer, M. et al. The neuroactive potential of the human gut microbiota in quality of life and depression. Nat. Microbiol. 4, 623–632 (2019).
Marinkovic, K., Alderson Myers, A. B., Arienzo, D., Sereno, M. I. & Mason, G. F. Cortical GABA levels are reduced in young adult binge drinkers: association with recent alcohol consumption and sex. Neuroimage Clin. 35, 103091 (2022).
Petroff, O. A. C. Book review: GABA and glutamate in the human brain. Neuroscientist 8, 562–573 (2002).
Dong, Z. et al. Gut microbiome: A potential indicator for predicting treatment outcomes in major depressive disorder. Front. Neurosci. 16, 813075 (2022).
Artacho, A. et al. The pretreatment gut Microbiome is associated with lack of response to methotrexate in New-Onset rheumatoid arthritis. Arthritis Rheumatol. 73, 931–942 (2021).
Litwinowicz, K., Choroszy, M. & Waszczuk, E. Changes in the composition of the human intestinal Microbiome in alcohol use disorder: a systematic review. Am. J. Drug Alcohol Abuse. 46, 4–12 (2020).
Gurwara, S. et al. Alcohol use alters the colonic mucosa–associated gut microbiota in humans. Nutr. Res. 83, 119–128 (2020).
Cani, P. D., Depommier, C., Derrien, M., Everard, A. & de Vos, W. M. Akkermansia muciniphila: paradigm for next-generation beneficial microorganisms. Nat. Rev. Gastroenterol. Hepatol. 19, 625–637 (2022).
Lei, W. et al. Akkermansia muciniphila in neuropsychiatric disorders: friend or foe? Front. Cell. Infect. Microbiol. 13, 1224155 (2023).
Si, J., Kang, H., You, H. J. & Ko, G. Revisiting the role of Akkermansia muciniphila as a therapeutic bacterium. Gut Microbes 14, (2022).
Mazzoli, R. & Pessione, E. The neuro-endocrinological role of microbial glutamate and GABA signaling. Front. Microbiol. 7, 210784 (2016).
Strandwitz, P. et al. GABA-modulating bacteria of the human gut microbiota. Nat. Microbiol. 4, 396–403 (2018).
Hou, D. et al. Gamma-aminobutyric acid (GABA): a comprehensive review of dietary sources, enrichment technologies, processing effects, health benefits, and its applications. Crit. Rev. Food Sci. Nutr. 64, 8852–8874 (2023).
Butts, M., Sundaram, V. L., Murughiyan, U., Borthakur, A. & Singh, S. The influence of alcohol consumption on intestinal nutrient absorption: A comprehensive review. Nutrients 15, 1571 (2023).
Voutilainen, T. & Kärkkäinen, O. Changes in the human metabolome associated with alcohol use: A review. Alcohol Alcohol. 54, 225–234 (2019).
Okuda, S., Nishiyama, N., Saito, H. & Katsuki, H. 3-Hydroxykynurenine, an endogenous oxidative stress generator, causes neuronal cell death with apoptotic features and region selectivity. J. Neurochem. 70, 299–307 (1998).
Vich Vila, A. et al. Impact of commonly used drugs on the composition and metabolic function of the gut microbiota. Nature Communications 11, 362 (2020).
Leppik, I. E. Zonisamide: chemistry, mechanism of action, and pharmacokinetics. Seizure 13, S5–S9 (2004).
Pettinati, H. M. et al. A structured approach to medical management: a psychosocial intervention to support pharmacotherapy in the treatment of alcohol dependence. J. Stud. Alcohol. Drugs. Suppl. 66, 170–178 (2005).
Sobell, L. C. & Sobell, M. B. Timeline Follow-Back. Measuring Alcohol Consumption 41–72 (1992).
Bolyen, E. et al. Reproducible, interactive, scalable and extensible Microbiome data science using QIIME 2. Nat. Biotechnol. 37, 852–857 (2019).
Callahan, B. J. et al. DADA2: High-resolution sample inference from illumina amplicon data. Nat. Methods. 13, 581–583 (2016).
Bisanz, J. E. & qiime 2R: Importing QIIME2 artifacts and associated data into R sessions. Preprint at (2018). https://github.com/jbisanz/qiime2R
McMurdie, P. J. & Holmes, S. Phyloseq: an R package for reproducible interactive analysis and graphics of Microbiome census data. PLoS One. 8, e61217 (2013).
Barnett, D. J., Arts, I. C. & Penders, J. MicroViz: an R package for Microbiome data visualization and statistics. J. Open. Source Softw. 6, 3201 (2021).
Jari et al. vegan: Community Ecology Package. Preprint at https://CRAN.R-project.org/package=vegan (2022).
Wickham, H. Ggplot2 (Springer International Publishing, 2016).
Wickham, H. et al. Welcome to the tidyverse. J. Open. Source Softw. 4, 1686 (2019).
Jin, Y., Chi, J., LoMonaco, K., Boon, A. & Gu, H. Recent review on selected xenobiotics and their impacts on gut Microbiome and metabolome. TRAC Trends Anal. Chem. 166, 117155 (2023).
Wei, Y. et al. Early breast Cancer detection using untargeted and targeted metabolomics. J. Proteome Res. 20, 3124–3133 (2021).
Gu, H., Zhang, P., Zhu, J. & Raftery, D. Globally optimized targeted mass spectrometry: reliable metabolomics analysis with broad coverage. Anal. Chem. 87, 12355–12362 (2015).
Yao, W. et al. Integrated plasma and urine metabolomics coupled with HPLC/QTOF-MS and chemometric analysis on potential biomarkers in liver injury and hepatoprotective effects of Er-Zhi-Wan. Anal. Bioanal Chem. 406, 7367–7378 (2014).
Conroy, M. J. et al. LIPID MAPS: update to databases and tools for the lipidomics community. Nucleic Acids Res. 1, 13–14 (2013).
Xue, J., Guijas, C., Benton, H. P., Warth, B. & Siuzdak, G. METLIN MS2 molecular standards database: a broad chemical and biological resource. Nat. Methods. 17, 953–954 (2020).
Pence, H. E. & Williams, A. ChemSpider: an online chemical information resource. J. Chem. Educ. 87, 1123–1124 (2010).
Schymanski, E. L. et al. Identifying small molecules via high resolution mass spectrometry: communicating confidence. Environ. Sci. Technol. 48, 2097–2098 (2014).
Acknowledgements
This work was supported by the National Institute on Alcohol Abuse and Alcoholism (NIAAA) under grants AA027858, AA12722313, AA007290, and AA027055and by the National Institute of Health (NIH) General Clinical Research Centers Program under grant M01 RR06192. We thank Pam Fall and Judy Kalinowski at the UConn Health Clinical Research Center for stool and serum sample management.
Funding
This work was supported by the National Institute on Alcohol Abuse and Alcoholism (NIAAA) under grants AA027858, AA12722313, AA007290, and AA027055 and by the National Institute of Health (NIH) General Clinical Research Centers Program under grant M01 RR06192.
Author information
Authors and Affiliations
Contributions
AJA, JMC, and YZ were responsible for study design. LRD, HY, JC, and HG performed data analysis. LRD interpreted findings and drafted the manuscript. All authors critically reviewed the manuscript and agree to publication of the final version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Conflict of interest disclosure
AJA is a consultant to Sobrera Pharmaceuticals. All other authors report no conflict of interest.
Clinical trial registration
Participants were recruited from the community at three sites (two in Connecticut and one in Virginia) as part of a double-blind, randomized, placebo-controlled study investigating the use of zonisamide in reducing drinking (ClinicalTrials.gov identifier: NCT02900352, Zonisamide Treatment of Alcohol Use Disorder: an Evaluation of Efficacy and Mechanism of Action (Z-Comp), first registered 14/09/2016
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Dedon, L.R., Yuan, H., Chi, J. et al. Baseline gut microbiome and metabolites are correlated with changes in alcohol consumption in participants in a randomized Zonisamide clinical trial. Sci Rep 15, 10486 (2025). https://doi.org/10.1038/s41598-025-92313-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-92313-0