Abstract
Group B Streptococcus (GBS, Streptococcus agalactiae) is a leading cause of invasive disease in neonates, pregnant women, elderly, and immunocompromised individuals. The epidemiology of invasive GBS disease remains poorly documented in small, isolated populations such as the Faroe Islands. This nationwide study assessed invasive GBS cases from 2009 to 2024 in the Faroe Islands. A total of 42 GBS cases were identified. Serotyping was performed using phenotypic assays, with in silico whole-genome sequencing (WGS) based serotype confirmation available for 15 isolates collected between 2020 and 2024. Multilocus sequence typing (MLST), antimicrobial resistance profiling, and virulence gene were also performed. GBS incidence showed an increasing trend over the 15-year period, with the average annual incidence increasing from 3.69 (2009–2018) to 7.47 (2019–2024) per 100,000 population. Predominant serotypes were II, V, Ib, and Ia. All isolates carried genes coding for Alp-family proteins, which are immunogenic targets that may meaningfully influence the effectiveness of future GBS vaccines. MLST revealed a predominance of clonal complex (CC) 12, followed by CC452, CC1, and CC23. Phenotypic testing showed that all isolates were susceptible to penicillin, while four displayed resistance to erythromycin and clindamycin, with two carrying the erm(A) gene. Virulence genes including pilus islands (PI), cfb, sodA, and srr1 were detected. This study establishes a genomic and epidemiological baseline for invasive GBS in the Faroe Islands, highlighting an increase in incidence, data suggesting the likely efficacy of future GBS vaccine candidates, and preserved penicillin susceptibility. Continued genomic surveillance will be essential for informing public health and vaccine policy in small and remote populations.
Similar content being viewed by others
Introduction
Streptococcus agalactiae (Group B Streptococcus, GBS) is a beta-hemolytic, gram-positive bacterium commonly colonizing the gastrointestinal and genitourinary tracts of humans1. While in general asymptomatic, GBS is a cause of neonatal sepsis, meningitis, and invasive disease in pregnant women, elderly, and immunocompromised individuals1. Global colonization rates in pregnant women range from 18% to 35%, underscoring its public health significance1. In recent years, the incidence of invasive GBS infections has increased notably among elderly populations in developed countries2,3.
Ten capsular serotypes (Ia, Ib, II–IX) of GBS were identified to predominate globally; based on polysaccharide antigens, with serotypes Ia, Ib, II, III, and V accounting for the majority of invasive and colonizing isolates globally4,5. The distribution of these serotypes, however, varies geographically4,6.
In alignment with the World Health Organization’s (WHO) goal of eliminating meningitis by 20307, maternal vaccination against GBS is being actively explored as a preventive measure8,9. Currently, two vaccine candidates have advanced to phase III clinical trials: Pfizer’s GBS6 vaccine targets serotypes Ia, Ib, II–V, and Minervax’s GBS-NN/NN2 vaccine targets the alpha-like surface proteins (AlpNs; AlphaC, Rib, Alp1, Alp2/Alp3). Alp-family proteins (AlphaC, Rib, Alp1, Alp2, and Alp3) are surface expressed adhesins that contribute to epithelial binding and immune evasion, and are highly immunogenic. Their conserved N-terminal domains form the basis of several protein-based vaccine candidates10.
Additional vaccines are in early stages of development9
The Faroe Islands, a geographically isolated high-income archipelago with approximately 54000 inhabitants, provides a unique epidemiological setting to study invasive GBS and assess whether trends observed elsewhere apply similarly. In the Faroe Islands, antenatal and intrapartum management of GBS follows the Danish national guidelines. As described by Slotved et al.11, Denmark applies a risk-based strategy, with intrapartum PCR testing used in defined clinical risk situations to guide intrapartum antibiotic prophylaxis, rather than universal antenatal screening. To date, there are no published data on the incidence, serotype distribution, resistance profile or protein profiles of invasive GBS on the Faroe Islands. Consequently, local health authorities lack essential evidence for evaluating the potential implementation of a GBS vaccination program or assessing the adequacy of current preventive strategies.
This study investigates temporal trends and the population-level burden of invasive GBS disease in the Faroe Islands over a 15-year period (2009–2024). In addition, whole-genome sequencing (WGS) was performed on isolates collected between 2020 and 2024 to characterize capsular serotype genes, vaccine-relevant surface proteins, multilocus sequence types (MLST)/clonal complexes (CC), virulence-associated surface proteins, and antimicrobial resistance determinants. The study aims to provide information to the health authorities in the Faroe Islands in order to support future GBS surveillance and vaccine policy decisions.
Materials and methods
Study population and clinical isolates
This study was based on invasive GBS isolates registered by the National Hospital of the Faroe Islands (NHFI) and Statens Serum Institut (SSI, Copenhagen, Denmark) between 2009 and 2024. In total, 42 invasive isolates were included, each representing a unique patient and a single episode of invasive GBS disease. Case identification was conducted through the digital microbiology registration system (MADS) at NHFI, followed by manual verification of invasiveness by a certified infectious disease specialist. Cases were classified as invasive when GBS was isolated from normally sterile sites, including blood, cerebrospinal fluid, ascitic fluid, deep tissue, or implanted medical devices. No recurrent invasive GBS episodes (isolates separated by more than 30 days or differed in serotype) were identified during the study period; accordingly, one isolate per patient was included4,12.
Viable isolates were available for 18 of the 42 cases (Table 1). This reflects that submission of clinical isolates from the Faroe Islands to SSI is voluntary, and consequently isolates were not consistently stored for all invasive GBS cases, particularly prior to 2020. The 18 isolates identification was confirmed phenotypically based on colony morphology on 5% horse blood agar plates (SSI Diagnostica, Hillerød, Denmark) and group-specific agglutination testing (Oxoid A/S, Greve, Denmark). Isolates were stored at − 80 °C in nutrient beef broth supplemented with 10% glycerol (SSI Diagnostica).
Serotyping
Isolates from 2005 to 2015 were serotyped by the capillary precipitation (Lancefield) method, preceded by screening using GBS latex agglutination tests (SSI Diagnostica, Denmark). From 2016 onward, isolates were primarily serotyped by latex agglutination, with inconclusive results confirmed by the Lancefield method. Non-typable isolates were classified accordingly13.
Whole-genome sequencing and molecular characterization
Fifteen clinical isolates collected between 2020 and 2024 were sequenced. The genomic data for these isolates are deposited at the European Nucleotide Archive (ENA) under Bioproject no. PRJEB89722. SSI only performed systematic sequencing of GBS isolates received from the Faroe Islands from the summer of 2019 to 2024. Before the summer of 2019, SSI performed only phenotypic testing. The two isolates obtained prior to 2020 were therefore not sequenced.
Genomic DNA was extracted using the DNeasy Blood & Tissue Kit (QIAGEN, Hilden, Germany). Sequencing libraries were prepared using the Nextera XT DNA Library Preparation Kit (Illumina, US) and sequenced on MiSeq or NextSeq 550 platforms (Illumina, US), generating 150–250 bp paired-end reads. De novo genome assembly was performed using SKESA (v.2.2)14. Quality control was ensured using the bifrost pipeline (https://github.com/ssi-dk/bifrost, accessed 24-06-2025), with accepted average coverage > 30 and absence of contamination.
Gene profile
Several surface protein and virulence-associated genes, including Alpha-like surface proteins, pilus islands, cfb, sodA, and serine-rich repeat genes, were selected for analysis because they are established markers of invasiveness, colonization potential, and vaccine relevance in global GBS populations. The selection of target genes was based on previously published studies15,16,17. Genotyping was performed using the Short Read Sequence Typer v2 (SRST2) tool (Inouye et al., 2014). In silico capsular serotyping (serotypes Ia–IX) was conducted using a validated reference database17, applying a minimum sequence coverage threshold of 95%.
Additional strain features were extracted from the genomic data, including the presence or absence of surface protein genes and virulence-associated determinants. These included:
-
1.
Hypervirulent GBS adhesin: hvga (AM051291).
-
2.
Serine-rich repeat proteins: srr1 (AL766851), srr2 (AY669067).
-
3.
Alpha-like surface proteins: alpha (M97256), rib (U58333), alp1 (AY461799), alp2/alp3 (AF245663).
-
4.
Pilus islands: PI-1 (EU929742, CP013908), PI-2a (EU929870, EU929881, EU929876, EU929899), PI-2b (EU929104, AM051289).
-
5.
Other conserved genes: cfb (JQ289578.1) and sodA (DQ232566.1).
A minimum sequence identity of ≥ 95% and ≥ 95% query coverage was required for gene detection.
Multi-locus sequence typing and clonal complex
In S. agalactiae, MLST assigns isolates to sequence types (STs) based on allelic profiles, and STs sharing six or seven identical alleles are grouped into a clonal complex (CC). The designation of a CC reflects its ancestral or predominant ST6.
MLST profiles were assigned by the PubMLST database (http://pubmlst.org/sagalactiae/, accessed 13-04-2025) to obtain information of which Clonal Complex the ST profiles belonged to.
Phenotypic susceptibility profile
The phenotypic and molecular analysis of susceptibility, were performed as described by Slotved et al.18. Briefly, the 18 invasive isolates (Table 1) underwent disc diffusion susceptibility testing for erythromycin (15 µg), clindamycin (2 µg), and from 2012, penicillin G (1 µg). Inducible clindamycin resistance was assessed via D-test, as described by EUCAST in the Reading guide for disk diffusion method, Version 11.0, January 2025 (https://www.eucast.org/ accessed 13-12-2025). Minimum inhibitory concentrations (MICs) for penicillin, erythromycin, and clindamycin in resistant isolates were determined by Etest® (bioMérieux, Denmark) following EUCAST recommendations (www.eucast.org, accessed 24-06-2025).
Genetic susceptibility profile
Of the 18 GBS isolates, the 15 isolates for which whole-genome sequencing (WGS) data were generated were analyzed for genes conferring resistance to erythromycin and clindamycin using ResFinder v3.1 (https://cge.cbs.dtu.dk/services/ResFinder/) with an identity threshold of 80% and a minimum gene length of 60%19.
Susceptibility to penicillin in GBS is associated with the sequence of penicillin-binding proteins (PBPs), particularly PBP2x. The 15 isolates were screened for their PBP profiles as described by Metcalf et al. and McGee et al.15,16, as specific amino acid substitutions in PBP genes may be linked to reduced susceptibility to beta-lactam antibiotics.
Data analysis
Descriptive analyses were performed using GraphPad Prism v.8.0.2. Population data were obtained from Statistics Faroe Islands, and annual averages were calculated to determine incidence rates per 100,000 population (https://hagstova.fo/fo/folk/folkatal/folkatal, accessed 03-03-2025).
Ethical considerations
Because SSI processes personal data under the authority of Article 6(1)(e) of the General Data Protection Regulation (GDPR), in conjunction with Sect. 222 of the Danish Health Act, the requirement for approval and informed consent was waived by the Scientific Ethics Committee. As part of its monitoring duties, SSI manages the national systems for reportable diseases, including invasive GBS cases, and since data were collected routinely for national surveillance purposes, no additional ethical approval or informed consent was required. Data from national administrative registers were linked with the invasive GBS cases and according to Danish law, ethical approval is not needed for this type of research. All methods were carried out in accordance with relevant guidelines and regulations, see also Schjørring et al.20.
Results
Incidence and case identification
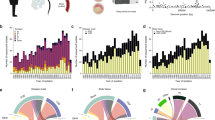
A total of 42 invasive GBS isolates were identified in the databases at NHFI from 2009 to 2024 (Fig. 1), predominantly from blood samples, with additional isolates from cerebrospinal fluid, deep tissues, peritoneal dialysis fluid, or ascites. The overall invasive GBS incidence was 5.11 per 100,000 population, increasing from an average of 3.69 per 100,000 (2009–2018) to 7.47 per 100,000 (2019–2024). Among isolates collected, 18 were initially stored at SSI, and additional 24 cases were identified directly by NHFI, confirming that not all invasive isolates had been submitted to SSI.
Serotype distribution
Viable isolates were available for 18 of the 42 invasive GBS cases, and these were subsequently serotyped (Fig. 2; Table 1). The most prevalent serotypes were serotype II (n = 4) and serotype V (n = 4), followed by serotype Ib (n = 3), Ia (n = 2), III (n = 2), IV (n = 1), VIII (n = 1), and IX (n = 1). Serotypes VI and VII were not detected. Notably, serotype III was not observed in any cases after 2018.
Whole-genome sequencing (WGS) was performed on 15 isolates collected from 2020 to 2024, representing the most recent cases. For these 15 isolates, in silico capsular serotyping based on WGS was used to confirm the phenotypic serotyping results (Table 1). The three earlier isolates (two from 2018 and one from 2009) were serotyped using phenotypic methods only.
Protein type distribution
All 15 WGS-analyzed isolates carried one Alpha-like protein (Alp) gene (Fig. 2; Table 1). The most common was alphaC (n = 7), followed by alp1 (n = 5) and alp2/alp3 (n = 3). Rib gene was not detected in any of the sequenced isolates.
MLST and clonal complexes
Multilocus sequence type (MLST) was determined for all 15 sequenced invasive GBS isolates. The most prevalent clonal complex (CC) was CC12 (n = 7), comprising sequence types ST8, ST10, and ST12. The second most common was CC452 (n = 3), which included ST24 and ST498.
Two isolates were assigned to CC459 (ST196 and ST2216), while one isolate each belonged to CC1 (ST1) and CC23 (According to the MLST database, the novel sequence type is closest to ST-88 and 11 other profiles, each differing by one locus). One isolate was identified as ST130, for which a clonal complex has not yet been defined.
Genomic features of invasive isolates
A selection of virulence-associated and conserved genes was screened across the 15 sequenced GBS isolates to evaluate molecular features relevant to invasiveness and vaccine development (Table 1).
The CAMP factor gene (cfb) and the housekeeping gene sodA were detected in all 15 isolates. The serine-rich repeat gene srr1 was detected in 14 of the 15 sequenced isolates and was absent only in one serotype II/ST12 genome. The srr2 gene was not detected in any isolate (Table 1).
The genes encoding pilus islands were detected in the majority of isolates. PI-1 was present in 11 out of 15 isolates, and PI-2a in 14 isolates. Both PI-1 and PI-2a were co-detected in 10 isolates. PI-2b was identified in only one isolate (serotype VIII/ST1), together with PI-1.
The hypervirulent adhesin gene hvgA was detected in only one isolate, associated with serotype Ia/ST24.
Susceptibility profile
Phenotypic resistance to erythromycin and clindamycin was observed in 4 out of 18 tested isolates, with two of these confirmed genetically as erm(A) positive (Table 1), demonstrating concordance between phenotypic and genotypic resistance. The remaining 14 isolates were phenotypically susceptible to both erythromycin and clindamycin. Among the phenotypically susceptible isolates that underwent WGS (n = 13), no macrolide–lincosamide resistance genes were detected (Table 1).
All isolates were phenotypically susceptible to penicillin. Whole-genome analysis of penicillin-binding proteins (PBPs) identified only two PBP2X profiles (PBP2X-1 and PBP2X-5) (Table 1), none of which have previously been associated with reduced susceptibility or resistance to beta-lactams.
Discussion
This study presents the first long-term epidemiological and genomic analysis of invasive Streptococcus agalactiae (GBS) infections in the Faroe Islands. Despite the limited population size, our findings indicate an overall trend toward increasing GBS incidence over the 15-year period (2009–2024), consistent with trends reported across Northern and Western Europe (Fig. 1). This rising burden is likely associated with aging populations and increasing prevalence of comorbidities, as observed in comparable high-income settings2,4,21.
The potential influence of the COVID-19 pandemic on GBS incidence remains uncertain. While restrictive measures during the pandemic led to reduced transmission of respiratory pathogens such as Streptococcus pneumoniae, no similar reduction in GBS infections was observed (Fig. 1).
This pattern is consistent with the view that invasive GBS may be less influenced by short-term changes in respiratory transmission dynamics and may therefore function as a relatively stable comparator organism in surveillance analyses22. Following the easing of pandemic restrictions, increases in invasive bacterial infections have been reported in several settings, and a similar post-pandemic increase in GBS incidence is plausible23,24.
Given the suggested increase in incidence, vaccination represents a promising strategy. The hexavalent polysaccharide conjugate vaccine (GBS6) targets the six most prevalent serotypes globally (Ia, Ib, II–V) and would have covered 16 of the 18 serotyped isolates in this study. However, serotype VIII and IX were also detected—highlighting the risk of serotype replacement, as seen in pneumococcal disease following PCV introduction25. The protein-based GBS-NN/NN2 vaccine candidate targets conserved N-terminal domains of the Alp-family proteins (AlphaC, Rib, Alp1, Alp2/Alp3). All sequenced isolates expressed at least one Alp-family protein, suggesting that, based on the circulating serotypes and Alp-family genes identified, the strains observed in this study would likely be largely covered by current candidate GBS vaccines, even in older adults.
The absence of the rib gene in our dataset is noteworthy, as rib is uniformly present among serotype III/CC17 (ST17) strains, which are disproportionately associated with neonatal meningitis and sepsis16. Consistent with this, no CC17/serotype III isolates were identified in the present study, and the dominance of AlphaC supports findings from other high-income settings16. Cross-protection among Alp variants remains under investigation, particularly for Alp1 and Alp2/Alp3 26.
Our MLST data revealed a predominance of CC12 among recent invasive GBS isolates in the Faroe Islands, followed by CC452, CC1, CC23, and CC456 (Table 1). Notably, no isolates from the hypervirulent CC17 lineage were detected. Along with the near absence of serotype III, this suggests that most infections likely occurred in adults rather than neonates. However, because age data were not available, it is not possible to determine which age groups constitute the dominant burden of invasive GBS infections in this study. This interpretation is therefore based on established strain disease associations rather than patient level metadata. In other studies, CC17 and serotype III are consistently associated with neonatal disease, particularly late-onset meningitis27,28. These findings reflect a moderate degree of genetic diversity among circulating strains. This pattern is consistent with broader GBS population studies, which typically identify CC1, CC10, CC17, CC19, and CC23 as the most common human-associated clonal complexes29,30. CC10 and CC12 represent the same broader lineage, as ST10 and ST12 are single-locus variants. In the current PubMLST database (https://pubmlst.org/organisms), both ST10 and ST12 are assigned to CC12, which is now considered the predominant founder type of this clonal cluster. Accordingly, only CC12 and not CC10 was detected in our dataset.
At the molecular level, all sequenced isolates carried the cfb and sodA genes (Table 1). These conserved genes are widely used as molecular markers for species confirmation and identification of S. agalactiae in both diagnostic and genomic analyses18,31. The srr1 gene was present in nearly all isolates, reflecting its widespread distribution across GBS lineages6,16.
The detection of the hypervirulence-associated hvgA gene in the isolate (serotype Ia, CC452) (Table 1) is rare, as it is typically associated with serotype III and the CC17 lineage, although it has also been linked to other serotypes and to CC452.16. The hvgA gene is classically associated with the hypervirulent CC17 lineage and has been strongly linked to neonatal meningitis16.
PI-1, PI-2a, PI-2b are GBS surface structures involved in colonization and invasiveness, with PI-1 linked to innate immune evasion, PI-2a to adherence/biofilm formation, and PI-2b to enhanced invasiveness and intracellular survival, including blood–brain barrier penetration6. Genes for the Pilus islands (PI) proteins were detected in all isolates in this study (Table 1), with PI-1 and PI-2a found to be the dominant Pilus, while PI-2b was identified in only one isolate.
Phenotypic and genotypic antimicrobial susceptibility testing confirmed universal susceptibility to penicillin among the tested isolates. This finding is consistent with surveillance data from Denmark, where beta-hemolytic streptococci have remained susceptible to penicillin since national monitoring began in the DANMAP reports (https://www.danmap.org/, accessed 25-06-2025). Although sporadic cases of penicillin non-susceptibility have been reported in Europe32, GBS is generally considered penicillin-susceptible16,33.
The observation that four GBS isolates exhibited resistance to both erythromycin and clindamycin, with two carrying the erm(A) gene (Table 1), aligns with global trends. While resistance to first-line therapy remains uncommon, increasing resistance to second-line agents such as macrolides and lincosamides is a growing concern worldwide33. In Denmark, erythromycin and clindamycin resistance levels are reported at approximately 20–25%, as documented in recent DANMAP reports (accessed 25-06-2025).
This study is limited by the small population size and relatively low number of cases, which constrain its statistical power. The exclusion of patient-level demographic data, such as age, sex, comorbidities, and education level of the patients, due to both the small population and General Data Protection Regulation (GDPR) restrictions, further limits the ability to perform stratified analyses. Additionally, not all isolates underwent whole-genome sequencing or complete phenotypic testing, which may introduce selection bias. In particular, isolates available for genomic analysis were primarily from later years, potentially underrepresenting earlier cases or specific clinical presentations.
Conclusion
This study provides the first long-term overview of invasive GBS disease in the Faroe Islands, incorporating epidemiological surveillance and genomic characterization. The findings align with trends in other high-income regions, showing increasing incidence and broad coverage of circulating strains by candidate vaccines. The absence of CC17 and serotype III suggests a predominantly adult disease burden, however, in the absence of age specific clinical data, this inference is based on strain disease associations rather than patient level information.
Penicillin remains effective, however rising resistance to erythromycin and clindamycin highlights the need for awareness. These data form a baseline for evaluating future vaccine implementation and underscore the importance of continued genomic surveillance in small, geographically isolated populations.
Data availability
The datasets used and/or analysed during the current study are available from the corresponding author on reasonable request. Sequences data from all isolates are deposited under the BioProject accession id (Bioproject no. PRJEB89722).
References
Pannaraj, P. S., Flores, A. R. & Baker, C. J. Group B Streptococcal Infections. In Feigin and Cherry’s Textbook of Pediatric Infectious Diseases 830–842.e7 https://doi.org/10.1016/B978-0-323-82763-8.00083-2 (Elsevier, 2025).
Uggen, E. et al. Incidence of invasive infections with group B Streptococcus in adults in Norway 1996–2019: a nationwide registry-based case-control study. Infection 52, 1745–1752 (2024).
Balta, V. A. et al. Invasive group B Streptococcal disease among nonpregnant adults: Alaska, 2004–2023. J. Infect. Dis. 232, e580–e584 (2025).
Slotved, H. C. & Hoffmann, S. The epidemiology of invasive group B Streptococcus in Denmark from 2005 to 2018. Front. Public. Heal. 8, 40 (2020).
Lohrmann, F. et al. Maternal Streptococcus agalactiae colonization in europe: data from the multi-center DEVANI study. Infection 53, 373–381 (2025).
Shabayek, S. & Spellerberg, B. Group B Streptococcal Colonization, molecular Characteristics, and epidemiology. Front. Microbiol. 9, 437 (2018).
World Health Organization (WHO). Defeating meningitis by 2030: a global road map. (2021).
Oliveira, L. M. A. et al. WHO defeating meningitis symposium, 3rd international symposium on Streptococcus agalactiae disease (ISSAD) in Rio de Janeiro, Brazil: State-of-the-art overview of S. agalactiae meningitis. Vaccine 52, 126895 (2025).
Pena, J. M. S., Lannes-Costa, P. S. & Nagao, P. E. Vaccines for Streptococcus agalactiae: current status and future perspectives. Front. Immunol. 15, 1430901 (2024).
Dangor, Z. et al. Association of infant rib and Alp1 surface protein N-terminal domain Immunoglobulin G and invasive group B Streptococcal disease in young infants. Vaccine 41, 1679–1683 (2023).
Slotved, H. C., Møller, J. K., Khalil, M. R. & Nielsen, S. Y. The serotype distribution of Streptococcus agalactiae (GBS) carriage isolates among pregnant women having risk factors for early-onset GBS disease: a comparative study with GBS causing invasive infections during the same period in Denmark. BMC Infect. Dis. 21, 4–11 (2021).
Lambertsen, L. M., Ingels, H., Schønheyder, H. C. & Hoffmann, S. Nationwide laboratory-based surveillance of invasive beta-haemolytic Streptococci in Denmark from 2005 to 2011. Clin. Microbiol. Infect. 20, O216–O223 (2014).
Slotved, H. C. & Hoffmann, S. Evaluation of procedures for typing of group B streptococcus: a retrospective study. PeerJ 5, e3105 (2017).
Souvorov, A., Agarwala, R. & Lipman, D. J. SKESA: strategic k-mer extension for scrupulous assemblies. Genome Biol. 19, 153 (2018).
Metcalf, B. J. et al. Short-read whole genome sequencing for determination of antimicrobial resistance mechanisms and capsular serotypes of current invasive Streptococcus agalactiae recovered in the USA. Clin. Microbiol. Infect. 23, 574.e7-574.e14 (2017).
McGee, L. et al. Population-Based distributions of candidate vaccine Targets, clonal Complexes, and resistance features of invasive group B Streptococci within the united States, 2015–2017. Clin. Infect. Dis. 72, 1004–1013 (2021). Multistate.
Tiruvayipati, S., Tang, W. Y., Barkham, T. M. S. & Chen, S. L. GBS-SBG - GBS serotyping by genome sequencing. Microb Genomics 7, (2021).
Slotved, H. C., Fuursted, K., Kavalari, I. D. & Hoffmann, S. Molecular identification of invasive Non-typeable group B Streptococcus isolates from Denmark (2015 to 2017). Front. Cell. Infect. Microbiol. 11, 1–9 (2021).
Zankari, E. et al. Identification of acquired antimicrobial resistance genes. J. Antimicrob. Chemother. 67, 2640–2644 (2012).
Schjørring, C. B. et al. Increasing incidence of serotype 38 invasive Pneumococcal disease driven by the ST393 clone among children, Denmark 2022–2024. Sci. Rep. 15, 15446 (2025).
Navarro-Torné, A., Curcio, D., Moïsi, J. C. & Jodar, L. Burden of invasive group B Streptococcus disease in non-pregnant adults: A systematic review and meta-analysis. PLoS One. 16, e0258030 (2021).
Shaw, D. et al. Trends in invasive bacterial diseases during the first 2 years of the COVID-19 pandemic: analyses of prospective surveillance data from 30 countries and territories in the IRIS consortium. Lancet Digit. Heal. 1–12. https://doi.org/10.1016/S2589-7500(23)00108-5 (2023).
Serra, G. et al. Group B Streptococcus colonization in pregnancy and neonatal outcomes: a three-year monocentric retrospective study during and after the COVID-19 pandemic. Ital. J. Pediatr. 50, 175 (2024).
Cocchio, S. et al. Surveillance of pediatric invasive bacterial diseases in the Veneto region: epidemiological trends and outcomes over 17 years (2007–2023). Vaccines 13, 1–17 (2025).
Løchen, A., Croucher, N. J. & Anderson, R. M. Divergent serotype replacement trends and increasing diversity in Pneumococcal disease in high income settings reduce the benefit of expanding vaccine valency. Sci. Rep. 10, 18977 (2020).
Brokaw, A. et al. A Recombinant Alpha-Like protein subunit vaccine (GBS-NN) provides protection in murine models of group B Streptococcus infection. J. Infect. Dis. 226, 177–187 (2022).
Jamrozy, D. et al. Increasing incidence of group B Streptococcus neonatal infections in the Netherlands is associated with clonal expansion of CC17 and CC23. Sci. Rep. 10, 9539 (2020).
Hsu, J. F. et al. Genomic characterization of serotype III/ST-17 group B Streptococcus strains with antimicrobial resistance using whole genome sequencing. Biomedicines 9, 1–13 (2021).
Bianchi-Jassir, F. et al. Systematic review of group B Streptococcal capsular types, sequence types and surface proteins as potential vaccine candidates. Vaccine 38, 6682–6694 (2020).
Zeng, Z. et al. Strain-level genomic analysis of serotype, genotype and virulence gene composition of group B Streptococcus. Front. Cell. Infect. Microbiol. 14, 1396762 (2024).
Rosa-Fraile, M. & Spellerberg, B. Reliable detection of group B Streptococcus in the clinical laboratory. J. Clin. Microbiol. 55, 2590–2598 (2017).
McGuire, E. et al. A case of penicillin-resistant group B Streptococcus isolated from a patient in the UK. J. Antimicrob. Chemother. 80, 399–404 (2025).
Hsu, C. Y. et al. Global patterns of antibiotic resistance in group B streptococcus: a systematic review and meta-analysis. Front. Microbiol. 16, 1541524 (2025).
Acknowledgements
The authors thank all healthcare professionals for their invaluable contributions to data collection and patient care in the Faroe Islands. KAK is a member of the Center of PandemiX, which is supported by the Danish National Research Foundation (DNRF-170).
Author information
Authors and Affiliations
Contributions
HCS designed the study. ÓJ and HCS analysed the data and drafted the manuscript. KAK and SG analysed and reviewed the data, contributed to the manuscript and critically revised the manuscript. All authors have approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Declaration of generative AI and AI-assisted technologies in the writing process
We acknowledge the use of ChatGPT-4, developed by OpenAI, for assisting in enhancing the readability and language of this manuscript.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Joensen, Ó., Krogfelt, K.A., Gaini, S. et al. A nationwide study of invasive Streptococcus agalactiae in the Faroe Islands from 2009 to 2024. Sci Rep 16, 5090 (2026). https://doi.org/10.1038/s41598-026-35025-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-35025-3