Abstract
Klebsiella pneumoniae is a major cause of bloodstream infections worldwide and is increasingly associated with multidrug resistance, particularly to carbapenems and colistin, which severely limits treatment options. This study investigated the genomic epidemiology of K. pneumoniae bloodstream infection (BSI) isolates collected in Thailand between 2020 and 2024. A total of 227 K. pneumoniae BSI isolates collected nationwide through the National Antimicrobial Resistance Surveillance Center, Thailand were subjected to antimicrobial susceptibility testing and whole-genome sequencing, followed by multilocus sequence typing, AMR gene detection, plasmid profiling, virulence scoring, phylogenetic analysis, and genotype–phenotype correlation. Among the isolates, 199 were resistant to carbapenems, seven were resistant only to colistin, and 21 were not resistant to either carbapenems or colistin. Three high-risk sequence types (ST) (ST16, ST147, and ST231) predominated in most regions, whereas the south showed greater clonal diversity. ST16 isolates commonly co-harbored blaNDM−1 and blaOXA−232 and had high rates of carbapenem and colistin co-resistance. ST231 isolates exhibited carbapenem resistance along with rmtF1-mediated aminoglycoside resistance, whereas ST147 isolates demonstrated high genomic plasticity and resistance diversity. Rare ST isolates carrying mcr genes and hypervirulent ST23 and ST268 lineages were also identified, with ST268 resistant to all carbapenems tested. These findings highlight the dissemination of high-risk clones in Thailand and underscore the need for real-time genomic surveillance and molecular diagnostics to inform infection control and national AMR strategies.
Similar content being viewed by others
Introduction
Klebsiella pneumoniae, a member of the Enterobacteriaceae family, is a major cause of healthcare-associated infections, including pneumonia, bloodstream infections (BSIs), and urinary tract infections1,2,3,4. BSIs are associated with high morbidity and mortality, especially in immunocompromised patients2,3. The global dissemination of multidrug-resistant (MDR) K. pneumoniae, particularly carbapenem-resistant K. pneumoniae (CRKP), is a serious clinical and public health challenge5. CRKP is designated as a critical priority pathogen by the WHO and a priority concern by the US Centers for Disease Control and Prevention, requiring urgent public health intervention6,7. The rise in CRKP has led to wider use of colistin in human and veterinary medicine. However, this is contributing to the development and spread of colistin-resistant CRKP (Col-CRKP), an alarming trend given that colistin is a last-line therapeutic agent. This rising resistance to both carbapenem and colistin increases the risk of treatment failure, mortality, and rapid nosocomial transmission, which can potentially lead to the emergence of pan-drug-resistant strains5,8,9.
In Thailand, the Department of Medical Sciences (DMSc) established the National Antimicrobial Resistance Surveillance Center (NARST) in 1997 to monitor antimicrobial resistance (AMR) in humans. NARST collects laboratory data and pathogenic bacterial isolates through a nationwide network, which currently includes 141 hospitals, for confirmatory AMR testing. NARST data indicate that the prevalence of CRKP in Thai hospitals has been steadily increasing, with meropenem resistance rising from 2.1% in 2015 to 15.5% in 202310. Moreover, in 2022, CRKP was linked to extremely high rates of in-hospital mortality (52.9% of patients with hospital-origin CRKP BSIs and 33.1% of patients with community-origin CRKP BSIs), making it one of the deadliest antimicrobial-resistant pathogens in the country11. Due to its growing clinical burden, CRKP has been designated as a priority pathogen for infection reduction in Thailand’s AMR National Action Plan (NSP on AMR 2023–2027).
Thailand is part of the Southeast Asian region where carbapenem-resistant Enterobacteriaceae (CRE) are considered endemic, and carbapenemase-producing K. pneumoniae clones carrying blaNDM and blaOXA−48-like genes have been reported to circulate widely in the country12,13. However, the genomic characteristics of these clones, including their sequence types (STs), AMR genes, and plasmids, as well as their transmission dynamics, are still largely unknown. This information is crucial for understanding the epidemiology of this high-risk pathogen at the regional, national, and international levels4,12. Expanding the genomic surveillance of CRKP will generate key information for the development of antimicrobial stewardship programs, infection prevention and control, and policies designed to address the regional and national threats posed by CRKP.
The existing knowledge gap can be reduced considerably by performing whole-genome sequencing (WGS), which provides deeper insights than phenotypic testing alone. Hence, we aimed to analyze the whole genomes of K. pneumoniae BSI isolates collected by the NARST between 2020 and 2024, including CRKP, colistin-resistant (ColR-KP), and colistin- and carbapenem-resistant (Col-CRKP) isolates, to identify the clonality, AMR genes, plasmid content, virulence potential, and genetic relatedness of isolates from different regions. Although real-time WGS was initially limited, integrating genomic and phenotypic surveillance revealed critical insights into resistance and transmission dynamics, thereby underscoring its value for broader implementation14,15. Our findings have implications for public health responses, infection control, and the clinical management of AMR in Thailand in accordance with the NSP on AMR (2023–2027). In ongoing efforts, we are investigating the implementation of near-real-time sequencing to detect emerging high-risk clones, with the objective of developing targeted onsite diagnostics for clinical surveillance and tracking of outbreak-related activity in the future.
Results
Antibiotic-resistant bacteria recorded in the NARST database between 2020 and 2024
Between 2020 and 2024, 5,783 presumptive antibiotic-resistant bacterial isolates were collected from NARST network hospitals for confirmatory AMR testing. Based on the regional classification of the National Statistical Office of Thailand, the majority originated in Central Thailand (49.8%), followed by Northern (22.5%), Northeastern (16.2%), and Southern (11.4%) Thailand (Table 1). This distribution reflects not only the number of provinces in each region but also regional differences in healthcare access, population density, and sample submission practices. K. pneumoniae was the most prevalent species (2,318 isolates), accounting for 40.1% of the isolates, followed by Escherichia coli (907 isolates, 15.8%) (Table 1). Other Enterobacteriaceae species, such as Enterobacter cloacae, Citrobacter freundii, and Klebsiella aerogenes, were also identified in several regions, albeit less frequently.
K. pneumoniae was the predominant species in all regions. Most of the K. pneumoniae isolates were obtained from sputum (n = 926) and urine (including catheterized; n = 817) samples, followed by blood (n = 248) and pus (n = 139) samples. Interestingly, a high prevalence (93.15%) of carbapenem resistance was observed among the K. pneumoniae BSI isolates, and it exceeded the prevalence recorded in the isolates from urine (91.31%) and sputum (91.79%) samples. Of the 248 K. pneumoniae BSI isolates, 227 culturable isolates were subjected to AST to confirm their resistance profiles, followed by WGS for genomic characterization. The distribution of the study isolates across Thailand is shown in Fig. 1A.
Geographical distribution and antimicrobial resistance characteristics of the K. pneumoniae bloodstream infection isolates included in this study. (A) Geographical distribution of the isolates. (B) The carbapenem and colistin resistance profiles of the isolates. (C) The proportions of colistin and aminoglycoside resistance among carbapenem-resistant isolates. The map was generated by the authors using Microsoft Excel (version 2507, build 16.0.19029.20136) with embedded geographic data derived from OpenStreetMap (© OpenStreetMap contributors, https://www.openstreetmap.org). OpenStreetMap data are made available under the Open Database License (ODbL) by the OpenStreetMap Foundation (OSMF).
Antimicrobial susceptibility of the K. pneumoniae BSI isolates
The antimicrobial susceptibility results of the study isolates are summarized in Supplementary Table S1. Among the 227 isolates, 199 were CRKP, seven were ColR-KP (resistant to colistin but remained susceptible to carbapenem), 21 were not resistant to either carbapenems or colistin which were included for comparison. Among the CRKP isolates, 38 were resistant to ertapenem only, while 158 showed pan-carbapenem resistance (i.e., resistance or intermediate resistance to all tested carbapenems). The remaining three isolates exhibited other carbapenem resistance patterns (Fig. 1B).
Among the CRKP isolates, 65 (32.7%) were also resistant to colistin (referred to as ColR-CRKP). The pan-carbapenem-resistant isolates exhibited a higher rate of colistin resistance (35.4%) than those resistant to ertapenem only (21.1%; Fig. 1C). Regarding aminoglycoside resistance, 33.2% of the CRKP isolates were resistant to at least one of the tested aminoglycosides. Interestingly, the pan-carbapenem-resistant isolates had a slightly lower aminoglycoside resistance rate (31.6%) than those resistant to ertapenem only (42.1%) (Fig. 1C).
When we examined the resistance of the isolates to the other tested antibiotics, we found that most of the CRKP isolates (93.0%) exhibited resistance or intermediate resistance to all tested β-lactam/β-lactamase inhibitors, cephalosporins, and fluoroquinolones. In addition, 85.9% of the CRKP isolates were resistant to trimethoprim-sulfamethoxazole. In contrast, most of the non-CRKP isolates were susceptible to these antibiotics (Supplementary Table S1).
Distribution of STs among CRKP BSI isolates
The 227 K. pneumoniae BSI isolates were subjected to WGS, and the genome assembly quality metrics and taxonomic identification results are summarized in Supplementary Table S2. Using MLST, based on the Pasteur scheme, we identified 40 distinct STs (Supplementary Table S1). Notably, isolates P0097 (non-CRKP) and P0121 (CRKP) contained novel STs. Subsequent submission of the data to the BIGSdb-Pasteur database (https://bigsdb.pasteur.fr/) resulted in P0097 being classified as an ST8922 isolate, and P0121 as an ST8923 isolate.
Overall, 31 STs were identified in the CRKP isolates, and their regional distribution is shown in Fig. 2. ST16, ST147, and ST231 were widespread across all regions. In the central region, ST16, ST147, and ST231 predominated, with ST11 also highly represented. Several minor STs, including ST15, ST340, ST395, and ST394, occurred at lower frequencies, while ST15 was uniquely detected in the central region. In the northeastern region, ST16, ST147, and ST231 were also prevalent, alongside ST340, ST20, and ST395. The STs detected in the northern region exhibited less diversity, with ST16, ST147, and ST231 as the predominant lineages. ST231 was less common in the northern region than in the central and northeastern regions. Interestingly, ST37 was uniquely detected in the northern region and was found in 16% of the isolates. In contrast, the STs detected in the southern region displayed the greatest heterogeneity, with no single dominant ST. The detected STs included ST11, ST16, ST48, ST147, ST231, ST273, ST268, and ST337. Notably, a substantial proportion of the isolates (29.2%) from the southern region contained rare STs, with each rare ST detected in a single isolate. This finding indicates considerable genomic diversity among CRKP in the southern region. Overall, ST16, ST147, and ST231 were the dominant lineages nationwide. They accounted for the highest proportions of the STs detected in the isolates from the central, northeastern, and northern regions, whereas the STs detected in the isolates from the southern region showed more diversity and no dominant lineage.
Whole-genome SNP-based phylogenetic analysis, AMR gene and plasmid profiling, and determination of virulence potential
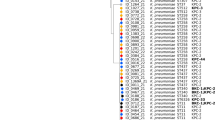
The whole-genome SNP-based phylogenetic tree of the 227 K. pneumoniae BSI isolates and the reference genome is shown in Fig. 3. Clustering consistent with the ST classification can be observed, with the major clusters dominated by high-risk clones (e.g., ST16, ST147, and ST231 isolates) and low SNP divergence within each cluster, suggesting clonal expansion. These clusters were geographically dispersed across multiple regions, implying inter-regional transmission.
The ST16 isolates had MDR profiles, with most showing resistance to all tested carbapenems. Additionally, 59% (23/39) of ST16 isolates were colistin-resistant, which is a markedly higher rate compared to other STs. Resistance to amikacin and gentamicin was relatively low (5.1% and 12.8%, respectively), whereas resistance to netilmicin was notably higher at 76.9% (Supplementary Table S1). Two carbapenemase genes, blaNDM−1 and blaOXA−232, were detected in nearly all isolates. Importantly, 66.7% (26/39) of the ST16 isolates harbored both blaNDM−1 and blaOXA−232. In addition, mutations in the ompK36 gene were identified in nearly all ST16 isolates. Despite the high rate of colistin resistance, only three ST16 isolates harbored mutations in crrB, mgrB, pmrA, or pmrB, and no known colistin resistance genes were found to be associated with the resistant phenotype. Regarding aminoglycoside resistance, all netilmicin-resistant isolates carried the aac(6’)-Ib gene, while all susceptible isolates lacked this gene, suggesting that it is involved in mediating netilmicin resistance. The ST16 isolates also exhibited conserved plasmid replicon profiles, frequently harboring Col440I, ColKP3, and variants of IncFIA, IncFIB, and IncFII (Fig. 3).
The ST231 isolates exhibited variations in their carbapenem resistance patterns and carbapenemase gene profiles, with blaOXA−232 and blaOXA−48 being predominant and associated with pan-carbapenem-resistant phenotypes. Additionally, mutations in the ompK36 gene were identified in most isolates. The colistin MIC varied among the ST231 isolates; however, no known colistin resistance genes were detected. Most of the ST231 isolates (87.5%) were resistant to all tested aminoglycosides, with most harboring both aac(6’)-Ib and rmtF1 genes. All isolates harboring rmtF1 showed resistance to all tested aminoglycosides, indicating a strong association between this gene and the aminoglycoside-resistant phenotype. ST231 isolates also harbored a conserved plasmid replicon set, which included ColKP3, and variants of IncFIA, IncFIB, and IncFII (Fig. 3).
The ST147 isolates exhibited considerable variation in their antibiotic resistance patterns, ranging from susceptible to all tested carbapenems, colistin, and aminoglycosides; to pan-resistance against all three antibiotic classes. The resistance genes and plasmids present in these isolates were also highly diverse. Most of the pan-carbapenem-resistant isolates (75%; 30/40) carried blaNDM−1, while blaNDM−5 was also found in certain isolates. In addition, blaOXA−48, blaOXA−181, and blaOXA−232 were detected at low frequencies. Interestingly, a subgroup of ST147 isolates carrying both blaNDM−1 and blaOXA−48 was observed. Most isolates within this subgroup exhibited pan-carbapenem resistance. Furthermore, a unique mutation in pmrB was universally present among ST147 isolates; however, this mutation was not associated with colistin resistance. For aminoglycosides, 29.4%, 35.3% and 56.9% of ST147 were resistant to amikacin, gentamicin, and netilmicin, respectively. aph(3’)-VI was the predominant aminoglycoside resistance gene identified in this ST. Interestingly, most of the isolates carrying armA exhibited resistance to all tested aminoglycosides. The plasmid analysis revealed high diversity, suggesting that plasmid-mediated dissemination of AMR genes may occur among ST147 strains (Fig. 3).
Other STs were detected at low frequencies and were often limited to specific geographic regions. The isolates with these STs displayed substantial diversity in their AMR gene profiles and plasmid content. In addition to blaNDM−1, blaNDM−4 and blaNDM−5 were also identified in rare ST isolates. Despite their low prevalence, five isolates with these uncommon STs harbored mcr genes, and this is of particular concern due to their strong association with phenotypic colistin resistance. Isolate P0273 (ST127) carried mcr-1 and displayed colistin resistance, along with an unusual carbapenem resistance profile, being resistant to meropenem only. Isolates P0188 (ST881) and P0264 (ST3167) harbored mcr-3 and were resistant to colistin; however, both were susceptible to all tested carbapenems. Two ST273 isolates (P0130 and P0192) carried mcr-8. P0130 was susceptible to all tested carbapenems and showed a moderate colistin MIC (2 µg/mL), whereas P0192 was resistant to ertapenem, doripenem, and meropenem, showed intermediate resistance to imipenem, and was resistant to colistin (Supplementary Table S1, Fig. 3). Four of the five isolates harboring mcr genes originated from the central region, and one was isolated from a northern province adjacent to the central region, suggesting potential regional dissemination.
Two novel STs, ST8922 (isolate P0097) and ST8923 (isolate P0121), were also identified. ST8923 clustered within a clade containing the reference lineage ST38, although it remained clearly separated from the major high-risk clones. This isolate exhibited pan-carbapenem resistance and intermediate resistance to colistin, while remaining susceptible to all tested aminoglycosides, and was characterized by the presence of blaNDM−4 alongside multiple plasmid replicons. In contrast, ST8922 grouped with ST882 within the broader MDR-associated clade that includes ST231; however, this isolate was susceptible to all tested antibiotics (Fig. 3). A sub-tree showing the phylogenetic placement of these novel STs, along with the allelic profiles used for ST designation, is provided in Supplementary Figure S1.
The virulence potential of all K. pneumoniae isolates based on the presence of specific virulence genes was evaluated using Kleborate virulence score (from 0 to 5). The ST23 and ST268 isolates were classified as hypervirulent (virulence score: 5). Both isolates carried the full repertoire of hvKp-associated loci, including yersiniabactin (ybt), aerobactin (iuc), salmochelin (iro), and colibactin (clb), as well as rmpA2. Capsule locus typing revealed that the ST23 isolate belonged to the K1 type, whereas the ST268 isolate corresponded to the K20 type. The ST231 and ST3167 isolates were considered virulent (virulence score: 4). All remaining isolates were classified as nonvirulent (virulence score ≤ 1; Supplementary Table S3).
The single-nucleotide polymorphism-based phylogenetic tree, antimicrobial susceptibility, antimicrobial resistance genes, and plasmid profiles of the K. pneumoniea bloodstream infection isolates collected in Thailand between 2020 and 2024. Antibiotic abbreviations: ETP, ertapenem; DOR, doripenem; MEM, meropenem; IMP, imipenem; AMK, amikacin; GEN, gentamicin; NET, netilmicin; COL, colistin. AST = antimicrobial susceptibility testing. Susceptibility categories: S, susceptible; I, intermediate; R, resistant. ST = Sequence type,.
Association between AMR genes and phenotypes
We performed a binary logistic regression analysis to evaluate the associations between the presence of carbapenemase genes and phenotypic resistance in the K. pneumoniae isolates. Multicollinearity among predictor variables was evaluated using variance inflation factors (VIF). All included gene predictors demonstrated VIF values < 3, indicating no significant multicollinearity (Supplementary Table S4). All logistic regression models included 227 isolates with complete genotypic and phenotypic data. The numbers of resistant versus susceptible isolates for each antibiotic were: ertapenem (202/25), doripenem (160/67), meropenem (168/59), imipenem (159/68), and pan-carbapenem resistance (158/69). Significant correlations were observed between the presence of specific carbapenemase genes and carbapenem-resistant phenotypes (Table 2). blaNDM−1 was strongly associated with resistance to imipenem (aOR = 283.44, q < 0.0001), meropenem (aOR = 103.34, q = 0.0002), doripenem (aOR = 240.82, q < 0.0001), ertapenem (aOR = 14.19, q = 0.0298), and the pan-carbapenem-resistant phenotype (aOR = 304.24, q < 0.0001). Additionally, perfect associations were identified between blaNDM−4 and blaNDM−5 and resistance to all tested carbapenems (i.e., present in all resistant isolates and absent in all susceptible isolates). When all blaNDM variants were pooled, these associations remained consistent. blaOXA−232 was significantly associated with resistance to all tested carbapenems, most notably to ertapenem (aOR = 14.96, q = 0.0298). blaOXA−48 and blaOXA−181 were also significantly associated with resistance to all carbapenems except ertapenem. When blaOXA variants were pooled, consistent associations were observed across all individual carbapenems and the pan-carbapenem-resistant phenotype.
When we analyzed the associations between AMR genes and colistin resistance (72 resistant and 155 susceptible isolates), only one mutation in pmrB (pmrB_T157P) showed a significant association (aOR = 5.20, q = 0.0388). Among the seven isolates carrying pmrB_T157P, six were colistin resistant. All isolates that harbored mcr-1 or mcr-3 were resistant to colistin. Of the two isolates that carried mcr-8, one was resistant and one was susceptible to colistin. In most cases (62/72 isolates), resistance to colistin could not be attributed to known resistance genes or mutations.
Discussion
In this nationwide genomic study, we analyzed the epidemiology and molecular mechanisms of the resistance of K. pneumoniae BSI isolates collected in Thailand between 2020 and 2024. The high proportion of CRKP (93.15%) found in K. pneumoniae BSI specimens compared with other specimen types in the NARST dataset underscores the threat that this pathogen poses in Thai healthcare settings and highlight the growing prevalence of MDR bacteria in Southeast Asia12. The prevalence of co-resistance to colistin in the CRKP isolates (32.7%) is particularly alarming, as colistin is a last-line antimicrobial agent used to treat infections with MDR bacteria16.
Notably, the CRKP isolates exhibited moderate rates of resistance to amikacin and gentamicin (33.2% and 45.2%, respectively), which were lower than the rates of resistance to the other antibiotic classes. These findings are consistent with national antibiogram data10. Furthermore, there are also previous reports of CRKP exhibiting moderate resistance to amikacin and gentamicin, with rates below 50% reported in several countries17,18,19. This suggests that aminoglycosides are still partially effective against CRKP and are a therapeutic option when treatment choices are limited20,21.
Our results demonstrate that high-risk ST16, ST147, and ST231 clones, which appear to be endemic in Thailand, predominate and are widely distributed across the country, particularly in the central, northern, and northeastern regions. These observations are consistent with the classification of such clones as global epidemic lineages associated with MDR and hospital outbreaks13,22,23,24. In contrast, the southern region exhibited highly diverse lineages, suggesting that a distinct epidemiological dynamic exists in this region. The absence of dominant clones in the south may reflect the occurrence of localized outbreaks or multiple introductions rather than sustained clonal expansion25. Several factors may contribute to this elevated lineage diversity in southern Thailand. The region shares an active land border and substantial population movement with Malaysia, and cross-border healthcare utilization, travel, and trade may introduce additional K. pneumoniae lineages26,27,28. Ecological factors may also play a role, as southern Thailand has extensive coastal and aquaculture activities, and K. pneumoniae can persist in brackish water, marine sediments, and aquaculture systems. Human interaction with these settings may provide opportunities for transmission of environmentally adapted or rare STs into clinical settings29,30,31,32. In addition, lower sequencing density relative to other regions indicates that additional sampling may be needed to fully characterize the population structure in the south. Collectively, such diversity and the unique influencing factors that differ from other regions highlight the importance of continuous genomic surveillance in the south, as it may serve as a reservoir for emerging resistance determinants or facilitate horizontal gene transfer events33.
After exploring regional differences in lineage distribution, we next examined the genomic and antimicrobial resistance profiles of all sequence types, with particular focus on the dominant high-risk clones. Most of the ST16 isolates exhibited pan-carbapenem resistance, which was mediated primarily by the co-occurrence of blaNDM−1 and blaOXA−23234,35. The co-occurrence of these genes in ST16 isolates has been reported in multiple countries, including Thailand36,37,38. The co-existence of these genes appears to be driven by the co-introduction of different plasmids, each carrying a distinct carbapenemase gene37,38. Most of the ST16 isolates also harbored mutations in ompK36, which contributed to porin loss39. Furthermore, ST16 emerged as the predominant Col-CRKP clone, with 59% of the ST16 isolates exhibiting colistin resistance despite the absence of acquired colistin resistance genes such as mcr genes40, or chromosomal mutations in pmrAB, crrB, or mgrB41. These findings underscore the complexity of colistin resistance and the limitations of current genotypic predictors, suggesting the involvement of uncharacterized genetic factors and, hence, the need for further investigation. Moreover, the ST16 isolates exhibited extensive resistance profiles, as well as conserved AMR genes and plasmid replicons (e.g., Col440I, ColKP3, IncFIA, IncFIB, and IncFII). These findings, which are consistent with previous reports42,43,44, indicate that this lineage has undergone clonal expansion and achieved broad geographic dissemination.
ST231 was the most prevalent ST in this study and was detected in multiple regions. In ST231 clones, carbapenem resistance is primarily mediated by blaOXA−48 or blaOXA−23235. The rate of aminoglycoside resistance was notably high among the ST231 isolates, with 87.5% showing resistance to all tested aminoglycosides. The copresence of aac(6’)-Ib and rmtF1 genes in the majority of isolates suggests that the resistance was mediated by plasmids via both the acetylation and methylation of ribosomal targets45. The perfect correlation between rmtF1 and aminoglycoside resistance underscores the clinical importance of this gene46. Furthermore, the ST231 isolates contained conserved plasmid replicons (e.g., IncFIA, IncFIB, and ColKP3), indicating the presence of stable mobile elements47. The high AMR rate, substantial clonal expansion, and widespread geographic distribution of the ST231 clones analyzed in this study indicate that ST231 is an established epidemic lineage in Thailand, and this is consistent with findings from previous surveillance studies13.
ST147 was the second most common ST detected in this study. The ST147 isolates were characterized by remarkable genomic diversity and a broader range of resistance profiles compared to the ST16 and ST231 isolates. These isolates displayed variations in their carbapenem, colistin, and aminoglycoside resistance, with many exhibiting a pan-resistant phenotype. The majority of the carbapenem-resistant isolates harbored blaNDM−1, and blaNDM−5, whereas blaOXA−48, blaOXA−181 and blaOXA−232 were occasionally detected. The copresence of blaNDM−1 and blaOXA−48 was observed, particularly in pan-carbapenem-resistant isolates. A pmrB mutation (R256G) was detected in all ST147 isolates; however, its role in colistin resistance remains uncertain, as only 31.37% of isolates of this ST were resistant. Previous studies have reported an association between pmrB mutations and polymyxin B resistance48; however, other investigations have shown that the presence of pmrB mutations does not always correspond to colistin resistance49,50. Aminoglycoside resistance in the ST147 isolates was driven by a range of genetic factors. The most commonly detected gene was aph(3’)-VI, while a subset of isolates harbored armA, which showed a strong correlation with high-level resistance to all tested aminoglycosides45. The heterogeneity of the AMR genes and plasmid replicons within the ST147 isolates reflects the genomic plasticity of this lineage and suggests that it may serve as a reservoir for AMR genes51.
In addition to the dominant STs, several rare STs were detected, with some isolates carrying mcr genes that confer plasmid-mediated colistin resistance40. Specifically, one ST127 isolate carried mcr-1, one ST881 isolate and one ST3167 isolate each carried mcr-3, and two ST273 isolates carried mcr-8. Although these STs were detected at low frequencies, and the isolates were mostly susceptible to carbapenems, the presence of these mcr variants in BSI isolates underscores the potential for horizontal gene transfer of colistin resistance within and across bacterial species52. Notably, STs associated with mcr genes have also been found in animals. For example, ST127 has been detected in horses53, and ST881 was previously reported in pigs in Europe54. Interestingly, ST3167 has been found in pigs from Thailand55. This overlap indicates that mcr-harboring clones can be spread across different ecological niches and may facilitate the spread of colistin resistance between bacteria that infect animals and those that infect humans56. Such findings reinforce the need for integrated genomic surveillance across human, animal, and environmental settings to monitor the evolution and transmission of mcr genes56,57. In contrast, ST273 appears to be a hospital-associated lineage58,59. Notably, a study in China also identified an ST273 isolate carrying the mcr-8 gene on a plasmid from a patient with bacteremia60. In this study, we identified an ST273 isolate harboring mcr-8 that exhibited resistance to colistin, ertapenem, doripenem, and meropenem, as well as intermediate resistance to imipenem. These findings raise concern that ST273 may serve as a reservoir for plasmid-mediated colistin and carbapenem resistance and could emerge as a high-risk clone in clinical settings.
The ST23 and ST268 isolates analyzed in this study were classified as hvKp based on their Kleborate virulence score of 5. Both isolates carried the full repertoire of hvKp-associated loci, including rmpA2, which is associated with the hypermucoid phenotype61. Capsule locus typing identified the ST23 isolate as K1 and the ST268 isolate as K20. ST23 has been recognized as a globally disseminated hvKp lineage, and it has been detected in Southeast Asia24,62. While carbapenem-resistant and MDR ST23 isolates have been reported in several countries63,64,65, the ST23 isolate identified in this study was susceptible to all tested carbapenems, and no known carbapenemase genes were detected. ST268 has also been reported as a global hvKp lineage, with multiple studies describing its association with extended-spectrum β-lactamase and carbapenemase genes66,67,68. In this study, the ST268 isolates exhibited pan-carbapenem resistance, harboring either blaNDM−4 or blaNDM−5. Moreover, an ST268 isolate obtained from the southern region was also nonsusceptible to all tested aminoglycosides, which underscores clinical concerns about this lineage. The ST231 and ST3167 isolates were classified as virulent (virulence score: 4). ST231 is recognized as a globally distributed resistant lineage and was the most prevalent ST detected in our CRKP isolates. Although ST231 clones harbor multiple virulence genes, several studies have suggested that they generally exhibit low to moderate virulence69,70. In this study, the ST3167 isolate was susceptible to carbapenems but carried mcr-3 and displayed colistin resistance. Little is currently known about the virulence potential of this lineage, and further investigation is required to clarify its pathogenic and epidemiological significance. The isolates with other STs were classified as nonvirulent, which is consistent with the findings of previous studies suggesting that MDR K. pneumoniae strains generally exhibit low virulence71. We determined the virulence potential of the isolates by assessing the presence of virulence-associated genes in the genome; however, virulence phenotypes and clinical outcome data were not assessed. Future genomic surveillance, combined with virulence phenotypes and clinical outcome analysis, will provide a more comprehensive understanding of the virulence and AMR of K. pneumoniae in clinical settings.
We identified two novel STs in this study. The ST8922 isolate exhibited susceptibility to all tested antibiotics, and no plasmid replicons were detected in this isolate. In contrast, the ST8923 isolate exhibited resistance to all tested carbapenems and carried blaNDM−4 along with multiple plasmids. Notably, the ST8923 isolate clustered separately from the other isolates, indicating an evolutionary distinction. It is possible that the ST8923 lineage has a nonclinical origin, such as an animal or environmental reservoir72,73. The detection of novel STs highlights the importance of performing continuous genomic surveillance, which enables the early detection and tracking of previously unrecognized high-risk lineages and provides insights into the evolutionary dynamics of AMR74.
Our binary logistic regression analysis showed strong associations between specific carbapenemase genes and phenotypic resistance. blaNDM−1 showed high aORs for resistance to all tested carbapenems. This finding is consistent with global reports highlighting blaNDM−1 as a key driver of high-level carbapenem resistance75. Moreover, blaNDM−4 and blaNDM−5 showed perfect associations with carbapenem resistance. A pooled analysis of blaNDM variants yielded similarly robust associations across all carbapenems. Among the OXA-48-like genes, blaOXA−232 was significantly associated with resistance to all tested carbapenems, particularly ertapenem, whereas blaOXA−48 and blaOXA−181 were significantly associated with resistance to doripenem, imipenem, and meropenem. Pooled OXA-48-like variants also demonstrated consistent associations with resistance to each carbapenem and with the pan-carbapenem resistant phenotype. Although OXA-48-like genes showed significant associations with carbapenem resistance in our regression models, this association likely reflects co-existing mechanisms in the genetic background of the isolates particularly the frequent co-harboring of blaNDM−1 and ompK36 mutations rather than the effect of OXA-48-like enzymes alone. Previous studies have demonstrated that OXA-48–like enzymes exhibit limited carbapenemase activity and typically confer low-level carbapenem resistance76,77. Thus, the high-level resistance observed in these isolates is expected to result from the combined effects of carbapenemase co-carriage and porin loss78,79.
For colistin resistance, only one mutation in pmrB (pmrB_T157P) showed a significant association, and this variant was present in only a small subset of isolates. Notably, the majority of colistin-resistant isolates did not carry any known resistance determinants, such as mcr genes or mutations in pmrAB, mgrB, or crrB80,81. This observation suggests that additional or uncharacterized mechanisms may contribute to colistin resistance in our collection. Several factors may account for the absence of known resistance markers. Colistin resistance can emerge through alternative Lipid A modification pathways, potentially involving genes or mutations that remain unannotated81. The phenomenon of heteroresistance, in which only a subset of cells displays increased tolerance, may also contribute and is not readily identified through genomic analysis alone82. Furthermore, incomplete annotation of regulatory networks, including two-component systems involved in lipid A modification, may limit detection of relevant mutations81,83,84. Additional factors, such as efflux pumps or other poorly defined stress response pathways, may also underlie colistin resistance in these isolates84. Together, these observations highlight the complexity of colistin resistance mechanisms and emphasize the need for more comprehensive genomic and multiomics analyses to fully elucidate the factors contributing to this phenotype.
This study offers a detailed genomic overview of CRKP and ColR-KP circulating in Thailand; however, several limitations should be acknowledged. This study was based exclusively on short-read sequencing, which limited the ability to resolve complete plasmid structures and distinguish plasmid-borne from chromosomal elements. The absence of patient-level clinical information also restricted interpretation of how specific genomic features relate to disease severity or treatment outcomes. In addition, the cross-sectional sampling approach provides a snapshot of circulating lineages but does not capture temporal or directional transmission events. Future studies that incorporate long-read sequencing, clinical metadata, and longitudinal sampling will enable a more comprehensive view of CRKP and ColR-KP dissemination and its clinical consequences.
Despite these limitations, our study demonstrates the utility of integrating WGS into routine AMR surveillance. The capacity to analyze genomic data not only increased our understanding of resistance mechanisms and transmission dynamics but also enabled the detection of high-risk clones with epidemic potential. While retrospective in nature, our findings provide a strong rationale for implementing prospective, near-real-time genomic surveillance to inform infection control and antimicrobial stewardship strategies in Thailand.
Conclusion
This nationwide genomic surveillance study revealed that high-risk CRKP is widespread in Thailand. The predominance of ST16, ST147, and ST231 clones, coupled with their extensive AMR, underscores the critical threat posed by these lineages. The detection of mcr genes in isolates with rare STs and the emergence of a hypervirulent, carbapenem-resistant clone further emphasizes the evolving complexity of AMR in this pathogen. Integrating WGS into national surveillance provides vital insights into resistance mechanisms, clonal dynamics, and evolutionary trends. Furthermore, expanding the use of genomic surveillance and molecular diagnostics is necessary to control further dissemination and to reach the goals outlined in Thailand’s AMR National Strategic Plan (2023–2027).
Methods
K. pneumoniae isolates and laboratory data collection
All K. pneumoniae isolates included in this study were recovered from glycerol stocks stored at − 80 °C as part of routine AMR surveillance performed by NARST. Between 2020 and 2024, the nationwide surveillance program collected 248 antibiotic-resistant K. pneumoniae isolates from blood cultures of hospitalized patients. A total of 227 isolates were successfully recovered and included in this study. Non-viable isolates were excluded, and no additional selection was applied. All isolates were re-identified by standard biochemical methods following the Manual of Clinical Microbiology85. The location at which each isolate was obtained, and the year of isolation are summarized in Supplementary Table S1.
Antimicrobial susceptibility testing
Antimicrobial susceptibility testing (AST) was performed via broth microdilution using customized THAN2F Sensititre plates (Thermo Fisher Scientific, Waltham, MA, USA). The panel included 21 antibiotics: amikacin, amoxicillin-clavulanate, ampicillin, ampicillin-sulbactam, cefepime, cefotaxime, cefoxitin, ceftazidime, ceftriaxone, cefuroxime, ciprofloxacin, colistin, doripenem, ertapenem, gentamicin, imipenem, levofloxacin, meropenem, netilmicin, piperacillin-tazobactam, and trimethoprim-sulfamethoxazole. Briefly, isolated colonies cultured on sheep blood agar plates were suspended in 5 mL of cation-adjusted Mueller–Hinton broth (Thermo Fisher Scientific, Waltham, MA, USA). The bacterial suspension was then adjusted to a 0.5 McFarland standard using a nephelometer. A 30 µL aliquot of this suspension was added to 11 mL of Mueller–Hinton broth to prepare the inoculum. Using the AIM Sensititre system (Thermo Fisher Scientific, Waltham, MA, USA), 50 µL of the inoculum was dispensed into each well of the THAN2F plates. The plates were incubated at 35 °C ± 2 °C for 18–24 h and read using the Sensititre ARIS 2X system (Thermo Fisher Scientific, Waltham, MA, USA) according to the manufacturer’s instructions. Minimum inhibitory concentrations (MICs) were calculated and interpreted using the Clinical and Laboratory Standards Institute (CLSI) M100 guidelines for AST86.
DNA extraction and whole genome sequencing
All K. pneumoniae isolates were freshly cultured on sheep blood agar plates and incubated overnight at 37 °C. Genomic DNA was extracted using the Presto™ Mini gDNA Bacteria Kit (Geneaid Biotech, New Taipei City, Taiwan) according to the manufacturer’s protocol. DNA purity and concentration were initially assessed using a NanoDrop™ 1000 spectrophotometer (Thermo Scientific, Waltham, MA, USA) and further quantified using a Qubit™ 3.0 Fluorometer (Invitrogen, Carlsbad, CA, USA). DNA integrity was evaluated by gel electrophoresis. Sequencing libraries were prepared from 100 ng of genomic DNA using Illumina DNA Prep kit (M) Tagmentation (Illumina, San Diego, CA, USA), following the manufacturer’s instructions. Libraries were sequenced on the Illumina NextSeq platform with the P1 XLEAP-SBS High Output Reagent Kit (Illumina, San Diego, CA, USA), generating 150-bp paired-end reads.
Genomic assembly and annotation
The raw sequencing reads were initially assessed for their quality and trimmed using Trim Galore v0.6.5 and then assembled using the SPAdes genome assembler v4.0.0 via the BV-BRC web resources using default parameters87. The genome assemblies were evaluated using QUAST v5.0.288, and only those with a minimum coverage of 30× were included in further analyses. Taxonomic classification based on the average nucleotide identity and alignment fraction was performed using GTDB-Tk v2.1.189. Multilocus sequence typing (MLST) of seven housekeeping genes (gapA, infB, mdh, pgi, phoE, rpoB, and tonB) was conducted using MLST v2.23.090. AMR genes and plasmid replicons were identified using AMRFinderPlus v3.12.891 and PlasmidFinder v 2.1.692, respectively. Virulence potential was assessed using Kleborate v2.3.2, which assigns a virulence score (0–5) based on the presence of specific virulence genes61.
Phylogenetic analysis and visualization
A phylogenetic tree was constructed using the 227 assembled genomes and the reference genome K. pneumoniae MGH 78,578 (GCF_000016305.1). Core single-nucleotide polymorphisms (SNPs), including substitutions, deletions, and insertions, were identified using Snippy v4.6.0 (https://github.com/tseemann/snippy). Recombinant regions were detected and masked using Gubbins93. Maximum likelihood phylogenetic analysis was then performed in IQ-TREE using the GTR + G nucleotide substitution model with 1,000 bootstrap replicates94. The resulting phylogenetic tree was visualized and annotated using the iTOL v6 web tool95.
Logistic regression analysis of AMR genes and phenotypic characteristics
Multicollinearity among predictor variables was evaluated using variance inflation factors. Associations between antimicrobial susceptibility and the presence of AMR genes were evaluated using multivariable binary logistic regression. For each antibiotic, phenotypic resistance (resistant vs. susceptible) was modeled as the dependent variable, with AMR gene presence as independent predictors. All 227 isolates with complete genotype and phenotype data were included in each model, with non-resistant isolates serving as controls. Adjusted odds ratios with 95% confidence intervals were calculated, and multiple testing correction was applied using the Benjamini–Hochberg false discovery rate (FDR), with significance defined as q < 0.05. All statistical analyses were performed in R (v4.3.1) (https://www.R-project.org/).
Data availability
All sequence data generated in this study have been deposited in the GenBank database (BioProject accession number: PRJNA1309027). The data presented in this study are also provided in the supplementary materials. Further inquiries can be directed to the corresponding author.
References
Paczosa, M. K. & Mecsas, J. Klebsiella pneumoniae: going on the offense with a strong defense. Microbiol. Mol. Biol. Rev. 80, 629–661 (2016).
Kochan, T. J. et al. Genomic surveillance for multidrug-resistant or hypervirulent Klebsiella pneumoniae among united States bloodstream isolates. BMC Infect. Dis. 22, 603 (2022).
Mejía-Limones, I. et al. Whole-genome sequencing of Klebsiella pneumoniae MDR Circulating in a pediatric hospital setting: a comprehensive genome analysis of isolates from Guayaquil, Ecuador. BMC Genom. 25, 928 (2024).
On, Y., Kim, J. W., Lee, J. & Yoo, J. S. Genomic analysis of carbapenem-resistant Klebsiella pneumoniae blood isolates from nationwide surveillance in South Korea. Front Microbiol 16, 1–12 (2025).
Murray, C. J. L. et al. Global burden of bacterial antimicrobial resistance in 2019: a systematic analysis. Lancet 399, 629–655 (2022).
Centers for Disease Control and Prevention (U.S.). Antibiotic Resistance Threats in the United States, 2019. (2019). https://doi.org/10.15620/cdc:82532
World Health Organization. WHO Bacterial Priority Pathogens List, 2024: Bacterial Pathogens of Public Health Importance to Guide Research, Development and Strategies to Prevent and Control Antimicrobial Resistance. (2024).
Kempf, I., Jouy, E. & Chauvin, C. Colistin use and colistin resistance in bacteria from animals. Int. J. Antimicrob. Agents. 48, 598–606 (2016).
Hatrongjit, R. et al. Genomic analysis of carbapenem- and colistin-resistant Klebsiella pneumoniae complex harbouring mcr-8 and mcr-9 from individuals in Thailand. Sci. Rep. 14, 16836 (2024).
NARST & Antibiograms (2025). https://narst.dmsc.moph.go.th/antibiogram
Tuamsuwan, K. et al. Frequency of antimicrobial-resistant bloodstream infections in 111 hospitals in Thailand, 2022. J. Infect. 89, 106249 (2024).
Wyres, K. L. et al. Genomic surveillance for hypervirulence and multi-drug resistance in invasive Klebsiella pneumoniae from South and Southeast Asia. Genome Med. 12, 11 (2020).
Takeuchi, D. et al. Nationwide surveillance in Thailand revealed genotype-dependent dissemination of carbapenem-resistant enterobacterales. Microb Genom 8, 1–12 (2022).
Mellmann, A. et al. Real-time genome sequencing of resistant bacteria provides precision infection control in an institutional setting. J. Clin. Microbiol. 54, 2874–2881 (2016).
Raabe, N. J. et al. Real-time genomic epidemiologic investigation of a multispecies plasmid-associated hospital outbreak of NDM-5-producing enterobacterales infections. Int. J. Infect. Dis. 142, 106971 (2024).
Binsker, U., Käsbohrer, A. & Hammerl, J. A. Global colistin use: a review of the emergence of resistant enterobacterales and the impact on their genetic basis. FEMS Microbiol. Rev 46, 1–37 (2022).
Galani, I. et al. Nationwide epidemiology of carbapenem resistant Klebsiella pneumoniae isolates from Greek hospitals, with regards to Plazomicin and aminoglycoside resistance. BMC Infect. Dis. 19, 167 (2019).
Khoshnood, S. et al. Molecular evaluation of aminoglycosides resistance and biofilm formation in Klebsiella pneumoniae clinical isolates: A cross-sectional study. Health Sci. Rep 6, 1–12 (2023).
Shi, H. et al. Molecular characteristics and antimicrobial susceptibility of carbapenem-resistant Klebsiella pneumoniae in a multicenter study in Ningbo, China. Front Microbiol 16, 1–16 (2025).
Shields, R. K., Clancy, C. J., Press, E. G. & Nguyen, M. H. Aminoglycosides for treatment of bacteremia due to carbapenem-resistant Klebsiella pneumoniae. Antimicrob. Agents Chemother. 60, 3187–3192 (2016).
Thy, M., Timsit, J. F. & de Montmollin, E. Aminoglycosides for the treatment of severe infection due to resistant gram-negative pathogens. Antibiotics 12, 860 (2023).
Wyres, K. L., Lam, M. M. C. & Holt, K. E. Population genomics of Klebsiella pneumoniae. Nat. Rev. Microbiol. 18, 344–359 (2020).
Argimón, S. et al. Rapid genomic characterization and global surveillance of Klebsiella using pathogenwatch. Clin. Infect. Dis. 73, S325–S335 (2021).
Hinthong, W. et al. Genomic insights into Klebsiella pneumoniae: Virulence, resistance, and transmission in South and Southeast Asia. Int. J. Med. Microbiol. 320, 151666 (2025).
Palmieri, M. et al. Genomic evolution and local epidemiology of Klebsiella pneumoniae from a major hospital in Beijing, China, over a 15 year period: dissemination of known and novel high-risk clones. Microb Genom 7, 1–14 (2021).
Phua, K. L. The promotion of cross-border medical tourism in developing countries: economic growth at the expense of healthcare system efficiency and cost containment? Open. Public. Health J. 9, 98–105 (2016).
Wangbenmad, C. Factors affecting Malaysia tourists’ destination loyalty behavior: a case study of Hatyai, Thailand. AMAR (Andalas Manage. Review). 7, 79–90 (2023).
Linkevicius, M. et al. Cross-border spread of a mosaic resistance (OXA-48) and virulence (aerobactin) plasmid in Klebsiella pneumoniae: a European Antimicrobial Resistance Genes Surveillance Network investigation, Europe, February 2019 to October 2024. Eurosurveillance 30, (2025).
Barati, A. et al. Isolation and characterization of aquatic-borne Klebsiella pneumoniae from tropical estuaries in Malaysia. Int. J. Environ. Res. Public. Health. 13, 426 (2016).
Xu, Y. et al. First report of potentially pathogenic Klebsiella pneumoniae from serotype K2 in mollusk Tegillarca Granosa and genetic diversity of Klebsiella pneumoniae in 14 species of edible aquatic animals. Foods 11, 4058 (2022).
Cornacchia, A. et al. Bathing seawater and sand as reservoirs of clinically relevant and antimicrobial resistant Klebsiella pneumoniae strains. Sci. Total Environ. 1006, 180930 (2025).
Natrah, I. et al. Antimicrobial resistance in Malaysian shrimp aquaculture and strategies to reduce its occurrence. Rev Aquac 17, 1–21 (2025).
Tofteland, S., Naseer, U., Lislevand, J. H., Sundsfjord, A. & Samuelsen, Ø. A long-term low-frequency hospital outbreak of KPC-producing Klebsiella pneumoniae involving intergenus plasmid diffusion and a persisting environmental reservoir. PLoS One. 8, e59015 (2013).
Sakamoto, N. et al. Genomic characterization of carbapenemase-producing Klebsiella pneumoniae with chromosomally carried blaNDM–1. Antimicrob Agents Chemother 62, 1–6 (2018).
Meng, L. et al. The distribution characteristics of global blaOXA-carrying Klebsiella pneumoniae. BMC Infect. Dis. 23, 182 (2023).
Espinal, P. et al. Genomics of Klebsiella pneumoniae ST16 producing NDM-1, CTX-M-15, and OXA-232. Clin. Microbiol. Infect. 25, 385e1–385e5 (2019).
Lee, H. et al. Co-introduction of plasmids harbouring the carbapenemase genes, blaNDM–1 and blaOXA–232, increases fitness and virulence of bacterial host. J. Biomed. Sci. 27, 8 (2020).
Abe, R. et al. Clonal dissemination of carbapenem-resistant Klebsiella pneumoniae ST16 co-producing NDM-1 and OXA-232 in Thailand. JAC Antimicrob. Resist 4, 1–5 (2022).
Tsai, Y. K. et al. Klebsiella pneumoniae outer membrane porins OmpK35 and OmpK36 play roles in both antimicrobial resistance and virulence. Antimicrob. Agents Chemother. 55, 1485–1493 (2011).
Hussein, N. H., AL-Kadmy, I. M. S., Taha, B. M. & Hussein, J. D. Mobilized colistin resistance (mcr) genes from 1 to 10: a comprehensive review. Mol. Biol. Rep. 48, 2897–2907 (2021).
Nirwan, P. K., Chatterjee, N., Panwar, R., Dudeja, M. & Jaggi, N. Mutations in two component system (PhoPQ and PmrAB) in colistin resistant Klebsiella pneumoniae from North Indian tertiary care hospital. J. Antibiot. (Tokyo). 74, 450–457 (2021).
Nguyen, T. N. T. et al. Emerging carbapenem-resistant Klebsiella pneumoniae sequence type 16 causing multiple outbreaks in a tertiary hospital in Southern Vietnam. Microb Genom 7, 1–11 (2021).
Sanikhani, R. et al. Outbreak of colistin and carbapenem-resistant Klebsiella pneumoniae ST16 co-producing NDM-1 and OXA-48 isolates in an Iranian hospital. BMC Microbiol. 24, 59 (2024).
Sousa-Carmo, R. R. et al. Tracking pandemic pathogens from wastewater surveillance in international airports: Klebsiella pneumoniae ST16 coproducing NDM-4 and OXA-181 arriving in South America. Lancet Microbe. 6, 101071 (2025).
Zhang, Y. et al. The prevalence and distribution of aminoglycoside resistance genes. Biosaf. Health. 5, 14–20 (2023).
Galimand, M., Courvalin, P. & Lambert, T. RmtF, a new member of the aminoglycoside resistance 16S rRNA N7 G1405 methyltransferase family. Antimicrob. Agents Chemother. 56, 3960–3962 (2012).
Ikhimiukor, O. O. et al. Clonal background and routes of plasmid transmission underlie antimicrobial resistance features of bloodstream Klebsiella pneumoniae. Nat. Commun. 15, 6969 (2024).
Rodrigues, A. C. S. et al. Non-clonal occurrence of PmrB mutations associated with polymyxin resistance in carbapenem-resistant Klebsiella pneumoniae in Brazil. Mem Inst. Oswaldo Cruz 114, 1–6 (2019).
Bolourchi, N. et al. Comparative genome analysis of colistin-resistant OXA-48-producing Klebsiella pneumoniae clinical strains isolated from two Iranian hospitals. Ann. Clin. Microbiol. Antimicrob. 20, 74 (2021).
Li, Z. et al. Genetic diversity of polymyxin-resistance mechanisms in clinical isolates of carbapenem-resistant Klebsiella pneumoniae: a multicenter study in China. Microbiol Spectr 11, 1–14 (2023).
Zhang, J. et al. Mobilizable plasmids drive the spread of antimicrobial resistance genes and virulence genes in Klebsiella pneumoniae. Genome Med. 15, 106 (2023).
Quan, J. et al. Prevalence of mcr-1 in Escherichia coli and Klebsiella pneumoniae recovered from bloodstream infections in china: a multicentre longitudinal study. Lancet Infect. Dis. 17, 400–410 (2017).
Garcia-Fierro, R. et al. Comparative phylogenomics of ESBL-, AmpC- and carbapenemase-producing Klebsiella pneumoniae originating from companion animals and humans. J. Antimicrob. Chemother. 77, 1263–1271 (2022).
Thorpe, H. et al. One Health or Three? Transmission Modelling of Klebsiella Isolates Reveals Ecological Barriers to Transmission between Humans, Animals and the Environment. (2021). https://doi.org/10.1101/2021.08.05.455249
Leangapichart, T. et al. Characterization of Klebsiella pneumoniae complex isolates from pigs and humans in farms in thailand: population genomic structure, antibiotic resistance and virulence genes. J. Antimicrob. Chemother. 76, 2012–2016 (2021).
Liu, M. C., Jian, Z., Liu, W., Li, J. & Pei, N. One health analysis of Mcr -carrying plasmids and emergence of Mcr-10.1 in three species of Klebsiella recovered from humans in China. Microbiol Spectr 10, 1–10 (2022).
Touati, A. et al. One health at risk: plasmid-mediated spread of mcr-1 across clinical, agricultural, and environmental ecosystems. Antibiotics 14, 506 (2025).
Chou, A. et al. Emergence of Klebsiella pneumoniae ST273 carrying blaNDM–7 and ST656 carrying blaNDM–1 in Manila, Philippines. Microb. Drug Resist. 22, 585–588 (2016).
Liu, L., Feng, Y., Long, H., McNally, A. & Zong, Z. Sequence type 273 carbapenem-resistant Klebsiella pneumoniae carrying blaNDM–1 and blaIMP–4. Antimicrob Agents Chemother 62, 1–6 (2018).
Ge, H. et al. First report of Klebsiella pneumoniae co-producing OXA-181, CTX-M-55, and MCR-8 isolated from the patient with bacteremia. Front Microbiol 13, 1–7 (2022).
Lam, M. M. C. et al. A genomic surveillance framework and genotyping tool for Klebsiella pneumoniae and its related species complex. Nat. Commun. 12, 4188 (2021).
Das, M. Global update on hypervirulent Klebsiella pneumoniae. Lancet Infect. Dis. 24, e621 (2024).
Cheong, H. S. et al. Emergence of serotype K1 Klebsiella pneumoniae ST23 strains co-producing the plasmid-mediated AmpC beta-lactamase DHA-1 and an extended-spectrum beta-lactamase in Korea. Antimicrob. Resist. Infect. Control. 5, 50 (2016).
Shankar, C. et al. Emergence of multidrug resistant hypervirulent ST23 Klebsiella pneumoniae: multidrug resistant plasmid acquisition drives evolution. Front Cell. Infect. Microbiol 10, 1–13 (2020).
Gálvez-Silva, M. et al. Carbapenem-resistant hypervirulent ST23 Klebsiella pneumoniae with a highly transmissible dual-carbapenemase plasmid in Chile. Biol. Res. 57, 7 (2024).
Kiaei, S., Moradi, M., hosseini-nave, H., Ziasistani, M. & Kalantar-Neyestanaki, D. Endemic dissemination of different sequence types of carbapenem-resistant Klebsiella pneumoniae strains harboring blaNDM and 16S rRNA Methylase genes in Kerman hospitals, Iran, from 2015 to 2017. Infect. Drug Resist. 12, 45–54 (2018).
Matsuda, N. et al. Prevalence, clonal diversity, and antimicrobial resistance of hypervirulent Klebsiella pneumoniae and Klebsiella variicola clinical isolates in Northern Japan. J. Glob Antimicrob. Resist. 35, 11–18 (2023).
Chen, C. et al. Genomic characteristics of two strains of ESBL-producing Klebsiella pneumoniae ST268 isolated from different samples of one patient. J. Glob Antimicrob. Resist. 36, 319–325 (2024).
Chen, T. et al. Identification and characterization of OXA-232-producing sequence type 231 multidrug resistant Klebsiella pneumoniae strains causing bloodstream infections in China. Microbiol Spectr 11, 1–11 (2023).
Li, Z. et al. Characterization of OXA232-producing carbapenem-resistant Klebsiella pneumoniae: genomic analysis and virulence assessment. Pol. J. Microbiol. 74, 82–94 (2025).
Kochan, T. J. et al. Klebsiella pneumoniae clinical isolates with features of both multidrug-resistance and hypervirulence have unexpectedly low virulence. Nat. Commun. 14, 7962 (2023).
Runcharoen, C. et al. Whole genome sequencing reveals high-resolution epidemiological links between clinical and environmental Klebsiella pneumoniae. Genome Med. 9, 6 (2017).
Akinyemi, M. O. et al. Genomic characterisation of an extended-spectrum β-Lactamase-producing Klebsiella pneumoniae isolate assigned to a novel sequence type (6914). Gut Pathog. 16, 69 (2024).
Zhao, D. et al. The emergence of novel sequence type strains reveals an evolutionary process of intraspecies clone shifting in ICU-spreading carbapenem-resistant Klebsiella pneumoniae. Front Microbiol 12, 1–9 (2021).
Dadashi, M. et al. Prevalence of blaNDM–1 -producing Klebsiella pneumoniae in Asia: A systematic review and meta-analysis. Journal des Anti-infectieux 19, 58–65 (2017).
Poirel, L., Potron, A. & Nordmann, P. OXA-48-like carbapenemases: the Phantom menace. J. Antimicrob. Chemother. 67, 1597–1606 (2012).
Potron, A. et al. Genetic and biochemical characterisation of OXA-232, a carbapenem-hydrolysing class D β-lactamase from Enterobacteriaceae. Int. J. Antimicrob. Agents. 41, 325–329 (2013).
Doi, Y. et al. Co-Production of NDM-1 and OXA-232 by Klebsiella pneumoniae. Emerg. Infect. Dis. 20, 163–165 (2014).
Lumbreras-Iglesias, P., Rodicio, M. R., Valledor, P., Suárez-Zarracina, T. & Fernández, J. High-level carbapenem resistance among OXA-48-producing Klebsiella pneumoniae with functional OmpK36 alterations: maintenance of ceftazidime/avibactam susceptibility. Antibiotics 10, 1174 (2021).
Jayol, A. et al. High-level resistance to colistin mediated by various mutations in the CrrB gene among carbapenemase-producing Klebsiella pneumoniae. Antimicrob Agents Chemother 61, 1–4 (2017).
Mondal, A. H. et al. A review on colistin resistance: an antibiotic of last resort. Microorganisms 12, 772 (2024).
Sánchez-León, I., Pérez-Nadales, E., Marín-Sanz, J. A. & García-Martínez, T. & Martínez-Martínez, L. Heteroresistance to colistin in wild-type Klebsiella pneumoniae isolates from clinical origin. Microbiol Spectr 11, 1–15 (2023).
Kim, S. Y., Choi, H. J. & Ko, K. S. Differential expression of two-component systems, PmrAB and phoPQ, with different growth phases of Klebsiella pneumoniae in the presence or absence of colistin. Curr. Microbiol. 69, 37–41 (2014).
Naha, S., Sands, K., Mukherjee, S., Dutta, S. & Basu, S. A 12 year experience of colistin resistance in Klebsiella pneumoniae causing neonatal sepsis: two-component systems, efflux pumps, lipopolysaccharide modification and comparative phylogenomics. J. Antimicrob. Chemother. 77, 1586–1591 (2022).
Carroll, K. C. et al. Manual of Clinical Microbiology (ASM, 2019).
CLSI. CLSI M100 Performance Standards for Antimicrobial Susceptibility Testing. (2025).
Olson, R. D. et al. Introducing the bacterial and viral bioinformatics resource center (BV-BRC): a resource combining PATRIC, IRD and vipr. Nucleic Acids Res. 51, D678–D689 (2023).
Gurevich, A., Saveliev, V., Vyahhi, N. & Tesler, G. QUAST: quality assessment tool for genome assemblies. Bioinformatics 29, 1072–1075 (2013).
Chaumeil, P. A., Mussig, A. J., Hugenholtz, P. & Parks, D. H. GTDB-Tk v2: memory friendly classification with the genome taxonomy database. Bioinformatics 38, 5315–5316 (2022).
Jolley, K. A., Bray, J. E. & Maiden, M. C. J. Open-access bacterial population genomics: BIGSdb software, the PubMLST.org website and their applications. Wellcome Open. Res. 3, 124 (2018).
Feldgarden, M. et al. AMRFinderPlus and the reference gene catalog facilitate examination of the genomic links among antimicrobial resistance, stress response, and virulence. Sci. Rep. 11, 12728 (2021).
Carattoli, A. & Hasman, H. PlasmidFinder and in silico pMLST: identification and typing of plasmid replicons in whole-genome sequencing (WGS). in Horizontal Gene Transfer Methods and Protocols (ed. de la Cruz, F.) 285–294Springer Nature, (2020). https://doi.org/10.1007/978-1-4939-9877-7_20
Croucher, N. J. et al. Rapid phylogenetic analysis of large samples of Recombinant bacterial whole genome sequences using Gubbins. Nucleic Acids Res. 43, e15–e15 (2015).
Nguyen, L. T., Schmidt, H. A., von Haeseler, A. & Minh, B. Q. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 32, 268–274 (2015).
Letunic, I. & Bork, P. Interactive tree of life (iTOL) v6: recent updates to the phylogenetic tree display and annotation tool. Nucleic Acids Res. 52, W78–W82 (2024).
Acknowledgements
We thank the 15 regional medical science centers for facilitating sample collection. We thank the NARST hospital network for collaborating on confirmatory AMR testing. We thank Dr. Kristen Sadler from Scribendi (www.scribendi.com) for editing a draft of this manuscript.
Funding
This study was funded by the Department of Medical Sciences, Ministry of Public Health, Thailand. The funder had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
Conceptualization: KK, WK, OS, NT, and JP. Data curation: KK, WK, PL, AU, and JP. Methodology & Experiments: KK, WK, OS, NC, NT, and CK. Funding acquisition: PAO. Investigation: KK, WK, PL, AU, OS, NC, NT, CK, PAO, and JP. Resources: KK, WK, OS, PL, and AU. Data analysis: KK, WK, NT, and JP. Supervision: KK, AU, PAO, and JP. Validation: KK, WK, NT, and JP. Writing – original draft: KK and JP. Writing – review & editing: KK, WK, NT, and JP. All authors approved the final version for submission.
Corresponding author
Ethics declarations
Ethics statement
This study was approved by the Institute for the Development of Human Research Protections (IHRP) on January 21, 2025 (Project No. IHRP2025005; COA No. 6-2568). The study used re-identified clinical isolates collected as part of routine national antimicrobial resistance surveillance, and no patient-identifying information was included. Informed consent was waived by the IHRP. All methods were performed in accordance with the relevant standard guidelines and regulations.
Competing interests
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this er.pap
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Krobanan, K., Kamjumphol, W., Leethongdee, P. et al. Nationwide genomic surveillance of carbapenem- and colistin-resistant Klebsiella pneumoniae bloodstream isolates in Thailand (2020–2024). Sci Rep 16, 5853 (2026). https://doi.org/10.1038/s41598-026-36228-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-36228-4