Abstract
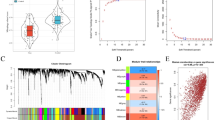
Alzheimer’s disease (AD) is a progressive neurodegenerative disorder characterized by complex neuroimmune interactions. Identifying reliable neuropathological markers and understanding immune cell infiltration in the brain are essential for improving our understanding of AD pathology. We integrated four temporal cortex gene expression datasets from the GEO database (GSE36980, GSE37263, GSE118553, GSE122063). Differentially expressed genes (DEGs) were identified using RobustRankAggreg (RRA) and batch correction. Functional enrichment was analyzed via GO and KEGG, and hub genes were identified through protein–protein interaction networks and comparative intersection analysis. Diagnostic performance was evaluated using ROC curves, and immune cell infiltration was profiled with CIBERSORT, with significant immune subsets identified via Wilcoxon tests and LASSO regression. Analysis revealed 98 robust DEGs, prominently enriched in pathways related to synaptic transmission and neuroactive ligand-receptor interactions. Two hub genes, CRH and GAD2, were identified and validated as being significantly downregulated in AD. ROC analysis affirmed their high discriminatory value (AUC ≥ 0.7), with a combined model demonstrating good performance. Immune infiltration profiling in the AD temporal cortex uncovered significant alterations in six immune cell populations: M2 macrophages, activated dendritic cells, and resting mast cells were increased, while plasma cells, regulatory T cells (Tregs), and activated NK cells were decreased. However, no significant correlation was found between the expression of CRH/GAD2 and these immune cell alterations. CRH and GAD2 are potential neuropathological markers for AD. The distinct immune infiltration patterns observed highlight the involvement of both innate and adaptive immunity in AD pathogenesis, offering new insights for understanding AD pathology and informing future therapeutic strategies. The lack of direct correlation suggests that neuronal gene dysregulation and immune alterations may represent parallel or independently regulated pathological dimensions in AD.
Similar content being viewed by others
Data availability
The gene expression datasets used in this study are publicly available from the GEO database (accession numbers: GSE36980, GSE37263, GSE118553, GSE122063, and GSE132903). The code supporting the findings is available at: https://github.com/caoyezi/analysis-code. Processed data are available from the corresponding author upon reasonable request.
Abbreviations
- AD:
-
Alzheimer’s disease
- RRA:
-
RobustRankAggreg
- Tregs:
-
Regulatory T cells
- NK:
-
Natural killer
- GO:
-
Gene ontology
- KEGG:
-
Kyoto encyclopedia of genes and genomes
- ROC:
-
Receiver operating characteristic
- RMA:
-
Robust multiarray average
- PPI:
-
Protein–protein interaction
- AUC:
-
The area under the ROC curve
- Ct:
-
The threshold cycle
- SEM:
-
Standard error of the mean
- CSF:
-
Cerebrospinal fluid
- PET:
-
Positron emission tomography
- CRH:
-
Corticotropin-releasing hormone
- DLPFC:
-
Dorsolateral prefrontal cortex
- BWS:
-
Bu-Wang San
References
Bhole, R. P., Chikhale, R. V. & Rathi, K. M. Current biomarkers and treatment strategies in Alzheimer disease: An overview and future perspectives. IBRO Neurosci. Rep. 16, 8–42. https://doi.org/10.1016/j.ibneur.2023.11.003 (2024).
McDade, E. M. Alzheimer disease. Continuum 28(3), 648–675. https://doi.org/10.1212/con.0000000000001131 (2022).
Day, G. S. Diagnosing Alzheimer disease. Continuum 30(6), 1584–1613. https://doi.org/10.1212/con.0000000000001507 (2024).
Jaqua, E. E., Tran, M. N. & Hanna, M. Alzheimer disease: Treatment of cognitive and functional symptoms. Am. Fam. Physician 110(3), 281–293 (2024).
Therriault, J. et al. Biomarker-based staging of Alzheimer disease: Rationale and clinical applications. Nat. Rev. Neurol. 20(4), 232–244. https://doi.org/10.1038/s41582-024-00942-2 (2024).
Mary, A., Mancuso, R. & Heneka, M. T. Immune activation in Alzheimer disease. Annu. Rev. Immunol. 42(1), 585–613. https://doi.org/10.1146/annurev-immunol-101921-035222 (2024).
Heneka, M. T. et al. Neuroinflammation in Alzheimer’s disease. Lancet Neurol. 14(4), 388–405. https://doi.org/10.1016/s1474-4422(15)70016-5 (2015).
Kinney, J. W. et al. Inflammation as a central mechanism in Alzheimer’s disease. Alzheimer’s Dement. Transl. Res. Clin. Interv. 4, 575–590. https://doi.org/10.1016/j.trci.2018.06.014 (2018).
Jorfi, M. et al. Infiltrating CD8(+) T cells exacerbate Alzheimer’s disease pathology in a 3D human neuroimmune axis model. Nat. Neurosci. 26(9), 1489–1504. https://doi.org/10.1038/s41593-023-01415-3 (2023).
Chen, X. et al. Microglia-mediated T cell infiltration drives neurodegeneration in tauopathy. Nature 615(7953), 668–677. https://doi.org/10.1038/s41586-023-05788-0 (2023).
Lu, Y., Li, K., Hu, Y. & Wang, X. Expression of immune related genes and possible regulatory mechanisms in Alzheimer’s disease. Front. Immunol. 12, 768966. https://doi.org/10.3389/fimmu.2021.768966 (2021).
Newman, A. M. et al. Determining cell type abundance and expression from bulk tissues with digital cytometry. Nat. Biotechnol. 37(7), 773–782. https://doi.org/10.1038/s41587-019-0114-2 (2019).
Jin, F., Xi, Y., Xie, D. & Wang, Q. Comprehensive analysis reveals a 5-gene signature and immune cell infiltration in Alzheimer’s disease with qPCR validation. Front. Genet. 13, 913535. https://doi.org/10.3389/fgene.2022.913535 (2022).
Lai, Y. et al. Identification of immune microenvironment subtypes and signature genes for Alzheimer’s disease diagnosis and risk prediction based on explainable machine learning. Front. Immunol. 13, 1046410. https://doi.org/10.3389/fimmu.2022.1046410 (2022).
Zhuang, X. et al. Development of a novel immune infiltration-related diagnostic model for Alzheimer’s disease using bioinformatic strategies. Front. Immunol. 14, 1147501. https://doi.org/10.3389/fimmu.2023.1147501 (2023).
Tian, Y. et al. Identification of diagnostic signatures associated with immune infiltration in Alzheimer’s disease by integrating bioinformatic analysis and machine-learning strategies. Front. Aging Neurosci. 14, 919614. https://doi.org/10.3389/fnagi.2022.919614 (2022).
Duan, K., Ma, Y., Tan, J., Miao, Y. & Zhang, Q. Identification of genetic molecular markers and immune infiltration characteristics of Alzheimer’s disease through weighted gene co-expression network analysis. Front. Neurol. 13, 947781. https://doi.org/10.3389/fneur.2022.947781 (2022).
Vogelgsang, J. & Wiltfang, J. New biomarkers for Alzheimer’s disease in cerebrospinal fluid and blood. Nervenarzt 90(9), 907–913. https://doi.org/10.1007/s00115-019-0772-9 (2019).
Wang, Y. et al. Advance and prospect of positron emission tomography in Alzheimer’s disease research. Mol. Psychiatry 30(10), 4899–4909. https://doi.org/10.1038/s41380-025-03081-2 (2025).
Assfaw, A. D., Schindler, S. E. & Morris, J. C. Advances in blood biomarkers for Alzheimer disease (AD): A review. Kaohsiung J. Med. Sci. 40(8), 692–698. https://doi.org/10.1002/kjm2.12870 (2024).
Thijssen, E. H. et al. Plasma phosphorylated tau 217 and phosphorylated tau 181 as biomarkers in Alzheimer’s disease and frontotemporal lobar degeneration: A retrospective diagnostic performance study. Lancet Neurol. 20(9), 739–752. https://doi.org/10.1016/s1474-4422(21)00214-3 (2021).
Stevenson-Hoare, J. et al. Plasma biomarkers and genetics in the diagnosis and prediction of Alzheimer’s disease. Brain 146(2), 690–699. https://doi.org/10.1093/brain/awac128 (2023).
Koutsodendris, N., Nelson, M. R., Rao, A. & Huang, Y. Apolipoprotein E and Alzheimer’s disease: Findings, hypotheses, and potential mechanisms. Annu. Rev. Pathol. Mech. Dis. 17, 73–99. https://doi.org/10.1146/annurev-pathmechdis-030421-112756 (2022).
Deming, Y. et al. Neuropathology-based APOE genetic risk score better quantifies Alzheimer’s risk. Alzheimers Dement. 19(8), 3406–3416. https://doi.org/10.1002/alz.12990 (2023).
Rajendran, K. & Krishnan, U. M. Biomarkers in Alzheimer’s disease. Clin. Chim. Acta 562, 119857. https://doi.org/10.1016/j.cca.2024.119857 (2024).
Jin, B., Cheng, X., Fei, G., Sang, S. & Zhong, C. Identification of diagnostic biomarkers in Alzheimer’s disease by integrated bioinformatic analysis and machine learning strategies. Front. Aging Neurosci. 15, 1169620. https://doi.org/10.3389/fnagi.2023.1169620 (2023).
Swaab, D. F., Bao, A. M. & Lucassen, P. J. The stress system in the human brain in depression and neurodegeneration. Ageing Res. Rev. 4(2), 141–194. https://doi.org/10.1016/j.arr.2005.03.003 (2005).
Chrousos, G. P. & Zoumakis, E. Milestones in CRH research. Curr. Mol. Pharmacol. 10(4), 259–263. https://doi.org/10.2174/1874467210666170109165219 (2017).
Zhang, Y. et al. Stress triggers gut dysbiosis via CRH-CRHR1-mitochondria pathway. NPJ Biofilms Microbiomes 10(1), 93. https://doi.org/10.1038/s41522-024-00571-z (2024).
Pedersen, W. A. et al. Corticotropin-releasing hormone protects neurons against insults relevant to the pathogenesis of Alzheimer’s disease. Neurobiol. Dis. 8(3), 492–503. https://doi.org/10.1006/nbdi.2001.0395 (2001).
Karalis, K., Muglia, L. J., Bae, D., Hilderbrand, H. & Majzoub, J. A. CRH and the immune system. J. Neuroimmunol. 72(2), 131–136. https://doi.org/10.1016/s0165-5728(96)00178-6 (1997).
May, C., Rapoport, S. I., Tomai, T. P., Chrousos, G. P. & Gold, P. W. Cerebrospinal fluid concentrations of corticotropin-releasing hormone (CRH) and corticotropin (ACTH) are reduced in patients with Alzheimer’s disease. Neurology 37(3), 535–538. https://doi.org/10.1212/wnl.37.3.535 (1987).
Tiwari, H. K., Bouchard, L., Pérusse, L. & Allison, D. B. Is GAD2 on chromosome 10p12 a potential candidate gene for morbid obesity?. Nutr. Rev. 63(9), 315–319. https://doi.org/10.1111/j.1753-4887.2005.tb00147.x (2005).
Davis, K. N. et al. GAD2 alternative transcripts in the human prefrontal cortex, and in schizophrenia and affective disorders. PLoS ONE 11(2), e0148558. https://doi.org/10.1371/journal.pone.0148558 (2016).
Lennartz, M. et al. GAD2 is a highly specific marker for neuroendocrine neoplasms of the pancreas. Am. J. Surg. Pathol. 48(4), 377–386. https://doi.org/10.1097/pas.0000000000002186 (2024).
Mosharaf, M. P., Alam, K., Gow, J., Mahumud, R. A. & Mollah, M. N. H. Common molecular and pathophysiological underpinnings of delirium and Alzheimer’s disease: Molecular signatures and therapeutic indications. BMC Geriatr. 24(1), 716. https://doi.org/10.1186/s12877-024-05289-3 (2024).
Wang, H. et al. Exploring the pharmacological mechanism of Bu-Wang San on Alzheimer’s disease through multiple GEO datasets of the human hippocampus, network pharmacology, and metabolomics based on GC-MS and UPLC-Q/TOF-MS. J. Ethnopharmacol. 350, 119994. https://doi.org/10.1016/j.jep.2025.119994 (2025).
Schwab, C., Yu, S., Wong, W., McGeer, E. G. & McGeer, P. L. GAD65, GAD67, and GABAT immunostaining in human brain and apparent GAD65 loss in Alzheimer’s disease. J. Alzheimers Dis. 33(4), 1073–1088. https://doi.org/10.3233/jad-2012-121330 (2013).
Burbaeva, G. et al. A role of glutamate decarboxylase in Alzheimer’s disease. Zh. Nevrol. Psikhiatr. Im. S S Korsakova 114(4), 68–72 (2014).
Tian, X. et al. Identification of vascular dementia and Alzheimer’s disease hub genes expressed in the frontal lobe and temporal cortex by weighted co-expression network analysis and construction of a protein-protein interaction. Int. J. Neurosci. 132(10), 1049–1060. https://doi.org/10.1080/00207454.2020.1860966 (2022).
Xu, W. et al. Identification of autophagy-related biomarkers and diagnostic model in Alzheimer’s disease. Genes https://doi.org/10.3390/genes15081027 (2024).
Liu, J., Yan, M., Chen, L., Yu, W. & Lü, Y. Construction and evaluation of a diagnostic model for Alzheimer’s disease based on mitophagy-related genes. Sci. Rep. 15(1), 10632. https://doi.org/10.1038/s41598-025-89980-4 (2025).
Fan, Y. et al. Identification and validation of biomarkers in Alzheimer’s disease based on machine learning algorithms and single-cell sequencing analysis. Comput. Biol. Chem. 118, 108475. https://doi.org/10.1016/j.compbiolchem.2025.108475 (2025).
Darwish, S. F., Elbadry, A. M. M., Elbokhomy, A. S., Salama, G. A. & Salama, R. M. The dual face of microglia (M1/M2) as a potential target in the protective effect of nutraceuticals against neurodegenerative diseases. Front. Aging 4, 1231706. https://doi.org/10.3389/fragi.2023.1231706 (2023).
Sackmann, V. et al. Anti-inflammatory (M2) macrophage media reduce transmission of oligomeric amyloid beta in differentiated SH-SY5Y cells. Neurobiol. Aging 60, 173–182. https://doi.org/10.1016/j.neurobiolaging.2017.08.022 (2017).
Théry, C. & Amigorena, S. The cell biology of antigen presentation in dendritic cells. Curr. Opin. Immunol. 13(1), 45–51. https://doi.org/10.1016/s0952-7915(00)00180-1 (2001).
Brezovakova, V., Valachova, B., Hanes, J., Novak, M. & Jadhav, S. Dendritic cells as an alternate approach for treatment of neurodegenerative disorders. Cell. Mol. Neurobiol. 38(6), 1207–1214. https://doi.org/10.1007/s10571-018-0598-1 (2018).
Yang, H. et al. Adoptive therapy with amyloid-β specific regulatory T cells alleviates Alzheimer’s disease. Theranostics 12(18), 7668–7680. https://doi.org/10.7150/thno.75965 (2022).
Patel, A. B., de Graaf, R. A., Martin, D. L., Battaglioli, G. & Behar, K. L. Evidence that GAD65 mediates increased GABA synthesis during intense neuronal activity in vivo. J. Neurochem. 97(2), 385–396. https://doi.org/10.1111/j.1471-4159.2006.03741.x (2006).
Acknowledgements
We extend our profound gratitude to the patients, their families, and caregivers for their invaluable participation and trust in this study.
Funding
This work was supported by the National Natural Science Foundation of China (Grant 82302026).
Author information
Authors and Affiliations
Contributions
Pan Liu and Chenglong Huang contributed equally as co-first authors. Their contributions included: conceptualization and study design, data collection, software utilization and formal analysis, methodology implementation and validation, and writing—original draft preparation. Lin Lu contributions included: data collection and software utilization. Zhaoyang Huang and Yilin Pang contributed equally as co-corresponding authors. Their contributions included: supervision and project administration, funding acquisition, resource provision, writing—review and editing, and final approval of the version to be published. All authors have read and approved the final manuscript and take full responsibility for the integrity and accuracy of all aspects of the work.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
The Institutional Review Board of the Institutional Ethics Committee at Renmin Hospital of Wuhan University, granted approval for this study (approval No.2025K-K290). The research was conducted in compliance with the principles outlined in the Declaration of Helsinki. Informed consent for participation in this study was obtained from the next of kin or legal guardians of the postmortem tissue donors.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liu, P., Huang, C., Lu, L. et al. Integrated transcriptomic analysis of the temporal cortex identifies CRH and GAD2 as neuropathological markers and reveals altered immune microenvironment in Alzheimer’s disease. Sci Rep (2026). https://doi.org/10.1038/s41598-026-40762-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-026-40762-6