Abstract
The development of X-ray free electron lasers has driven significant progress in X-ray science. Given the broad range of their applications, implementing a new generation of this technology at the laboratory scale has been under consideration for several years. This initiative is now under commissioning and construction at Arizona State University, known as the Compact X-ray Light Source (CXLS) and the Compact X-ray Free Electron Laser (CXFEL). Alongside experimental advances in this direction, whether in large or compact X-ray free electron lasers, there is also a growing need for new algorithmic and analytical methods to process the data obtained from such facilities. This work introduces a novel approach for analyzing Small- and Wide-Angle X-ray Scattering (SWAXS) profiles using a data-driven machine learning algorithm. The method is proposed for application to SWAXS datasets collected at both compact and large-scale X-ray facilities. To evaluate the performance of this approach, we analyzed simulated time-resolved SWAXS data from a protein, generated based on the current CXLS experimental parameters, and compared the results with those from the standard singular value decomposition (SVD) technique. Despite the low photon counts in the data, the results demonstrate that our method achieves higher accuracy in extracting structural dynamics information compared to SVD.
Similar content being viewed by others
Introduction
X-ray Free-Electron Lasers (XFELs) offer the opportunity to study biological macromolecules under physiological conditions, since the incurred radiation damage could be ”outrun” due to the short femtosecond duration of the X-ray pulse itself1,2,3,4,5. While initial applications involved probing biomolecules in crystalline form5, the same “diffraction before destruction” principle also holds for biomolecules in solution. The ability to probe structure under physiological conditions is of great benefit for time-resolved investigations of biological processes. To this end, scientists have developed time-resolved Small- and Wide-angle X-ray Scattering (TR-SWAXS) at XFELs to study the time evolution of macromolecules in solution. A further benefit of the femtosecond-duration XFEL pulses is the unprecedented temporal resolution enabled by these short probe pulses. This has revolutionized structural biology, enabling the study of a protein quake in photosynthetic reaction center6, ultrafast reaction dynamics in myoglobin7, hemoglobin8 and most recently RNA9 and visual rhodopsin10. The technique has recently been expanded to include data collection at Megahertz pulse repetition rates11,12,13.
Current XFELs rely on the process of Self-Amplified Spontaneous Emission (SASE) for generating these unique X-ray pulses; they require linear accelerators14,15,16. This severely limits the number of experiments that can be conducted at any one time at an XFEL facility (unlike synchrotron radiation sources, where the accelerators are circular and dozens of instruments can take beam concurrently). Currently, there are five hard X-ray XFEL facilities operational in the world, namely the Linac Coherent Light Source (LCLS) in Stanford, USA17, the Spring-8 Angstrom compact free electron laser (SACLA) in Harima, Japan18, the Pohang Accelerator Laboratory XFEL (PAL-XFEL) in Pohang, Korea19, the European XFEL GmbH in Hamburg, Germany13, and most recently the SwissFEL in Villigen, Switzerland20. All five institutions implement a peer-reviewed application process for awarding beamtime to scientists, with experiments typically scheduled months to years after the initial application for beamtime. These kilometers-long accelerators are expensive to maintain and run and require a multitude of scientific, engineering, and administrative staff.
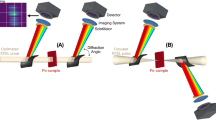
The prospect of building smaller, compact XFELs that may be operated by an academic institution has, therefore, gained traction recently. The first compact synchrotron light source went into operation in Munich (MuCLS) in 2016, and has shown remarkable results in X-ray science and source versatility21 (The reader is referred to this review for an overview of inverse Compton Scattering X-ray sources). In 2015, Arizona State University (ASU) proposed building a Compact X-ray Light Source (CXLS) to produce incoherent femtosecond X-ray pulses in a proof-of-principle instrument. Instead of using SASE for X-ray generation, CXLS uses Inverse Compton Scattering, shrinking the entire X-ray source down to fit within 600 sqft (Fig.1), while simultaneously providing superior beam stability characteristics22,23,24. Funded by ASU, philanthropic donations, and the National Science Foundation (NSF), the CXLS successfully generated X-rays for the first time in February 2023, and has commissioned its first structural biology studies. The fully lasing Compact X-ray Free Electron Laser (CXFEL) will produce coherent X-ray pulses and has begun construction in the same building.
While capable of producing attosecond to femtosecond duration X-ray pulses, one limitation of compact sources compared to their larger counterparts is the reduction in photon flux per pulse by 4-5 orders of magnitude. This leads to a substantial decrease in the signal-to-noise ratio of data collected at CXLS/CXFEL. Conventional signal processing techniques, such as singular value decomposition (SVD), may not be ideal for handling such low photon count data, highlighting the need for alternative algorithms. In the work presented here, we have used the current and expected operation parameters of the CXLS to investigate the information content that can be retrieved from simulated time-resolved SWAXS data, demonstrating the feasibility of performing these experiments at CXLS currently, as well when the machine reaches its full design parameters.
Here we present and evaluate a novel data analysis approach for time-resolved SWAXS data. We demonstrate that even with the current flux at CXLS, structural information can be retrieved from SWAXS data, often outperforming SVD. This method integrates concepts from scattering physics, graph theory, Riemannian geometry, and machine learning–particularly dimensionality reduction by diffusion maps25,26,27. The core algorithm has previously made it possible to study a variety of topics in structural biology and physics, such as extracting 3D structures of biomolecules28,29,30, 3D structural conformations and thermodynamics of macromolecules and molecular machines31,32,33, ultrafast structural dynamics at the atomic level34,35, gestational age estimation of human fetuses36, as well as identifying and mitigating data artifacts in XFEL experiments37. We additionally demonstrate a graphical user interface (GUI) to facilitate the long-term goal of becoming a user facility, increasing access to these powerful X-ray technologies.
Theory
Our approach is in the framework of manifold-based machine learning. This data analytic technique can be understood by considering diffraction snapshots or images, for instance, from a biological molecule, each composed of p pixels. Geometrically, each snapshot can be represented as a vector in a p-dimensional space, where each dimension corresponds to the intensity value of a single pixel. In this picture, the whole data set forms a manifold in the p-dimensional space, where each vector from the origin to the manifold surface represents a single data point. Therefore, any change in pixel values, whether due to changes in molecular structure or experimental artifacts, yields a new vector in the new space. This property enables the exploration of structural variability through the manifold, as well as the detection and reduction of stochastic or systematic errors in the data37. However, to make this goal practical, we represent the data in a low-dimensional feature space through a dimensionality reduction method, known as manifold embedding by diffusion map25. The dimensions in the new space reflect potential physical variations exercised by the system, along with contributions from noise or other extraneous artifacts. In the following section, we outline the manifold-based algorithm used in this work, called Nonlinear Laplacian Spectral Analysis (NLSA). Additional details and applications are available in Refs.34,35,38.
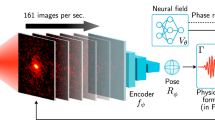
An overview of the pipeline we employ is shown in Fig. 1, consisting of five main steps as follows. After pre-processing the raw data, the primary step of NLSA is time-lagged embedding, in which the original time series is embedded into a high-dimensional space, resulting in a so-called concatenated dataset. This is followed by diffusion map embedding during which NLSA discovers the manifold from the concatenated dataset using a dimensionality reduction technique called diffusion map (DM). The curved structure of the manifold revealed by the diffusion map enables the extraction of nonlinear features hidden within data. Singular Value Decomposition (SVD) is then applied in the manifold space to uncover the fundamental characteristics of the system. During the final stages of NLSA, the data is reconstructed using a subset of the SVD modes and the results are projected back to the data space. A more detailed description is provided in “Methods”.
To simulate real-world conditions at the CXLS, the scattering profiles were calculated taking into account real-world artifacts and noise introductions. This was done using a combination of DENSS39,40 to calculate a contrast corrected SWAXS profile from an atomic model and the package reborn41 which includes experimental conditions when using the first Born approximation for far-field elastic X-ray scattering to calculate the profiles (see “Methods”). This enabled us to model experimental artifacts that we are anticipating (such as significant noise level) when performing SWAXS experiments at CXLS. A detailed description of the implementation in reborn is provided in “Methods”.
CXLS Accelerator and overview of the NLSA algorithm. The right panel shows the data analysis pipeline within the framework of manifold-based machine learning.
Results
Calmodulin data
To test our approach, we used Calmodulin (CaM) as a model system, which is a calcium-binding messenger protein found in eukaryotic cells. Its structure has been experimentally determined in different conformational states, making it of particular interest for the research presented here. Calmodulin undergoes very large structural changes upon calcium-binding, as illustrated in Fig. 2 for PDB entries 1CFD42 and 2BBM43. This large change should produce a significant change in scattering intensity in the small angle region, serving as a suitable test case for our analysis at the CXLS.
Calmodulin SWAXS simulation data. (a) Overlay of Calmodulin structures 1CFD (pink) and 2BBM (blue) in its open, unbound, and closed state with Calcium bound. (b) The ideal scattering intensity profiles for the two Calmodulin states used in the data simulation overlayed on the noised mixture intensity from a 10 mg/mL concentration of protein in water at a time delay of 2.5 ms. (c) The ground truth difference scattering intensity compared to the noisy simulated difference intensity for the same time delay. Both simulations were done with \(6\times 10^4\) exposures per time delay (60 s of exposure) with \(10^5\) incident photons per exposure.
As described in the Methods section, the time-resolved simulated SWAXS intensities were generated using the DENSS39,40 and reborn packages41 as a weighted mixture of the calcium-free and calcium-bound structures (see Fig. 2a). The concentrations of these states were set to evolve over time as sigmoid functions with a characteristic time of \(\tau =490\) \(\mu s\)44 and an inflection point at 3 ms. The concentrations of the two states sum to 1 at each time delay. We simulated difference intensity profiles at 100 time delays between 0-5 ms, so that the time-step was \(\Delta t= 50\) \(\mu s\), similar to the fluorometry data discussed by Park et. al.44. We then applied our NLSA algorithm to denoise the difference intensity profiles and to extract reaction kinetics. The lower flux at CXLS compared to larger XFELs results in weaker scattering intensities (e.g., see Fig. 2b). In addition, like in real experimental setups, the time delays in this simulation represent the relative delays between the activation and the X-ray probe. Figure 2c shows the simulated profiles of the same Calmodulin states in water with 10 mg/mL concentration of protein. See “Methods” for further simulation details.
Calmodulin analysis
To analyze the simulated Calmodulin data by NLSA, we began by performing a parameter search as outlined in “Methods”. To explore the parameter space for a dataset consisting of 100 time points, each containing an intensity profile from \(10^4\) radially averaged snapshots (10 seconds per time delay with a \(1\) kHz repetition rate) with \(10^5\) photon counts, we proposed \(\sigma _f \in \{0.5,1, 2, 4, 8\}\) and \(nN\in \{20,40,60,84\}\), and fixed the size of the delay-coordinate embedding window (or concatenation order) c to 16 (see “Methods” for more details about these parameters). Because the total number of time points is only \(N=100\), we considered a range of nN values up to N to ensure that local and global structures in such a small dataset are captured by the diffusion map embedding in the NLSA pipeline. However, for very large datasets, small nN values (e.g., up to \(10-20\%\) of N) are preferred to reduce computational complexity.
As detailed in “Methods” , the outcome of the above parameter search is presented as a heatmap. Based on this graph, we selected \(nN=84\) and \(\sigma _f=8\) for subsequent analyses. Choosing other contiguous hot areas of the heatmap is also allowed and will yield similar results. With the specified parameters, the manifold embedding by diffusion map (“Methods”) produced a set of eigenfunctions and their corresponding eigenvalues. Since the non-noise eigenfunctions generally appear at the beginning of the spectrum, using only the leading eigenfunctions is ideal for projecting the concatenated data matrix X onto the manifold, and reducing its dimensionality while doing so. For the current data, we used the first five and constructed the matrix A, as described by Eq. 3. SVD was then applied to this matrix on the curved manifold, producing singular values and vectors illustrated by Eq. 4. For the purpose of comparison, we also applied the standard SVD analysis on the same initial data.
Analysis of 100 TR-SAXS difference scattering profiles from Calmodulin using NLSA and SVD. (a) The singular values (normalized to the first) given by NLSA (red) and SVD (blue). (b) NLSA chronos. (c) SVD right singular vectors. (d) The sigmoid curve fit to \(V_{2,NLSA}\), and (e) the fit to \(V_{3,SVD}\). In both NLSA and SVD, mode 1 (\(V_1\)) shows the evolution of the averaged SAXS profiles across the entire time interval.
Figure 3b, c display the five leading NLSA chronos (modes), as well as the right singular vectors from SVD (hereafter also referred to as modes), along with their corresponding relative singular values (Fig. 3a). Like in the standard SVD, the first NLSA mode corresponds to the moving average of the signal in the SWAXS profiles, which is around \(\pm 1\) here due to a normalization step in the algorithm. The other modes reflect deviations from this average (see Methods, “NLSA algorithm”). In particular, \(V_{2,NLSA}\) represents the underlying dynamics in the data, which clearly follows a sigmoid function, as discussed in “Methods”. In contrast, the second SVD mode is dominated by noise and shows no clear trend. \(V_{3,SVD}\) is also very noisy, however, it reveals the sigmoidal time dependence from the simulation. SVD assigned a higher singular value to the noise-dominated second mode than to the signal-bearing third mode, despite the third mode actually indicating the expected dynamics. Such ambiguity is not present in NLSA, where non-noise and noisy eigenfunctions are clearly separated.
Figure 3a compares the singular values of the two methods, normalized to the same scale. The singular values from NLSA reach a distinct noise floor after the third component, whereas the SVD spectrum fails to level out even by the fifth singular value. This clear separation in the NLSA spectrum simplifies the identification of relevant modes for data reconstruction, particularly in low-flux conditions. Since CXLS and CXFEL are intended as user facilities, it is critical that the analysis tools provide clear guidance; NLSA simplifies the selection of relevant modes for reconstruction, whereas mode selection in standard SVD remains ambiguous for non-expert users. The second NLSA mode (\(V_{2,NLSA}\)) in Fig. 3b is in fact the reaction coordinate for the system35. Due to the simplicity of this two-state system, we fitted a sigmoid shape to \(V_{2,NLSA}\) and \(V_{3,SVD}\) modes and directly extracted the characteristic time (see Fig. 3d, e). Doing so yields a relaxation time of \(\tau _{NLSA}=540\pm 20 \upmu s\) with a coefficient of determination of \(R^2=0.990\). Applying the same approach to \(V_{3,SVD}\) reveals a relaxation time of \(\tau _{SVD}=450\pm 100 \upmu s\) with a coefficient of determination of \(R^2=0.803\). The relaxation time extracted from NLSA is more precise than that obtained from SVD. However, this overestimate arises from the relatively large concatenation window used for this data (16% of N). This is significant for the low-fidelity data analyzed here. In low-flux regimes, NLSA prioritizes signal recovery and temporal noise reduction. While the necessary use of larger concatenation windows in small datasets (N=100) introduces a systematic shift in the absolute relaxation time, it enables the high-precision identification of kinetic trends that are otherwise lost to the noise floor in standard SVD analysis. For higher-quality and larger datasets, a smaller concatenation window can be employed, thereby minimizing this effect, as demonstrated later in Fig. 4 .
As mentioned in Methods, when prior knowledge about the system is unavailable, as is often the case with experimental data, the NLSA reconstruction is done using modes displaying clear signals. Such modes are typically associated with the leading singular values that often appear before the flat part of the singular value spectrum or the so-called noise floor (see Fig. 3a). Due to the sigmoidal dynamics embedded in the simulated data, we expect one mode (beyond the average) represents this sigmoidal character. Therefore, in this simulation, we used the first two modes for the NLSA reconstruction (in SVD, we used modes 1, 2, and 3). The outputs of this step were a reconstructed time series of SAXS difference intensity profiles, which is assessed in Supplementary Information, Noise Estimation in Reconstructed Data.
Plots of kinetic data extracted using NLSA and SVD from five replicates of the Calmodulin simulated data for each pair of parameters. The relaxation rates (\(k=1/\tau\)) extracted from fitting a sigmoid to (a) NLSA modes (chronos) and (b) SVD modes (right singular vectors). Also plotted are the squared residuals scaled by the squared amplitude of the sigmoid fit to (c) NLSA modes and (d) SVD right singular vectors. Error bars for each plot are the 95% confidence interval given by the standard deviation of the replicates.
The above dataset demonstrates a minimum viable parameter set for extracting meaningful results from SWAXS data at CXLS, showing that extracting kinetic data from SWAXS data at the CXLS, with the current flux limitations which are still well below the design parameters, is already possible using both NLSA and SVD, an important finding for the practical feasibility of these experiments. As the equipment develops and improves, higher photon counts will become available so that less beam time is required for equivalent results. Figure 4 demonstrates the parameters we can extract given combinations of photon counts per pulse and exposure time per time delay. The results shown here are average values resulting from five replicate simulations with unique Poisson noise for each. The NLSA parameters were similarly chosen using the GUI and heatmap process (see Supplementary Information, Table S1). We see from Fig. 4a,b that NLSA extracts a reasonably accurate relaxation rate from the modes at even \(10^5\) photon counts, whereas SVD singular vectors struggle to do so with confidence. Higher photon counts and exposure times yield higher quality raw data such that each method converges to the actual value. However, NLSA data does so with more consistency and precision for nearly all cases. Further, Fig. 4c,d show that the time-lagged embedding in NLSA reduces noise in modes by at least an order of magnitude for the SAXS difference intensities analyzed here.
Photoactive yellow protein data
The calcium binding process of Calmodulin results in a large structural change over a few milliseconds. As such, the structural change seen with SAXS is fairly visible with low exposures. To challenge the capabilities of the CXLS, we simulated the beginning states of the Photoactive Yellow Protein (PYP) photocycle, which is a much smaller structural change involving isomerization of a chromophore leading to a cascade of small conformational changes in the protein, rather than a large domain alteration as in CaM. As done with CaM, we simulated two states evolving via sigmoidal concentration profiles, starting from a pure dark state (PDB: 6P5F) to a 30 ps activated state (PDB: 6P5D). This evolution occurred over 30 ps with a relaxation time of \(\tau =2\)ps centered around \(15\)ps. SAXS profiles were generated at 1000 time delays with a temporal spacing of \(\Delta t=30\) fs. These profiles were averaged over time using a Gaussian envelope with \(FWHM=300\) fs, mimicking the assumed probe pulse-width. In addition, Gaussian distributed timing jitter with \(\sigma =50\) fs was added to each timestamp before resorting the profiles to the new jittered time delays. This value was chosen based on the known timing jitter of the laser used at CXLS for X-ray generation. Inverse Compton Scattering based sources do not have the same spectral and timing jitter inherent to SASE-based sources, and demonstrate very low shot to shot variability. We therefore believe that a Gaussian pulse profile is an accurate model for the CXLS. Finally, the dark state was simulated (including noise) 50 times and averaged for use in calculating the SAXS difference scattering cross-section profiles.
Photoactive yellow protein analysis
Analysis of 1000 TR-SWAXS difference scattering profiles from PYP using NLSA and SVD. (a) The singular values (normalized to the first) given by NLSA (red) and SVD (blue). (b) NLSA modes. (c) SVD modes. (d,e) The sigmoid curve fit to \(V_{2,NLSA}\) and \(V_{10,SVD}\), respectively. In both NLSA and SVD, the first mode (\(V_1\)) shows the evolution of the averaged SAXS profiles across the entire time interval.
As done above for the CaM protein, NLSA was performed on these simulations from PYP to extract kinetics and denoise the difference profiles. Figure 5 shows an example of NLSA modes, SVD right singular vectors, and relative singular values from the PYP difference profiles generated from 60 seconds of exposure per time delay and \(10^5\) photons per X-ray pulse. Using the GUI package (“Methods”), NLSA parameters of \(c=180\), \(nN=820\), and \(\sigma _f=8\) were chosen. The standard SVD was additionally applied to the same data for comparison.
The relative singular values in Fig.5a are dramatically lower for \(S_{2,NLSA}\) and beyond compared to \(S_{2,SVD}\). This again suggests the ability for NLSA to clearly identify true signal in the data and separate it from the noise. Looking at Fig. 5b, we see a clear trend in \(V_{1,NLSA}\) and \(V_{2,NLSA}\) reminiscent of the sigmoidal concentrations imparted in the simulations despite the inclusion of timing jitter. The same cannot be said for any of the first five SVD right singular vectors (Fig. 5c). Applying a sigmoidal fit to \(V_{2,NLSA}\) in Fig. 5d resulted in a relaxation time of \(\tau _{NLSA}=4.5\pm 0.2 ps\) (\(R^2=0.987\)). Despite the systematic shift in absolute relaxation time caused by pulse broadening and the necessary use of a large concatenation window (18% of N), the method remains highly effective for the qualitative identification of structural dynamics, however with quantitative limitations in extreme noise environments. In contrast, attempts to fit any of the first five SVD right singular vectors were unsuccessful at yielding a physically useful relaxation time. In fact, the first mode to potentially exhibit the expected kinetics is \(V_{10,SVD}\) (\(R^2=0.026\)), shown in Fig. 5e. The relaxation time extracted from the fit is \(\tau _{SVD}=3.2\pm 2.3\) ps. Although this time is relatively near what we expect, the fit would provide no confidence to an experimentalist without prior knowledge of the system. This further showcases the power of NLSA over SVD at isolating time-domain information from weak and noisy datasets.
Given the simplicity of this two-state system and the linear nature of the concentration mixing, the spatial components from NLSA and SVD are similar (see S.I. Figure S2). However, since the modes of interest are clear using NLSA, reconstructions can be made using only the first two modes. For SVD in this and many other cases analyzed in this work, the modes of interest are not distinguished as easily. Thus, more modes must be kept in an SVD reconstruction to reach the noise floor and have confidence that all real signal is preserved. In doing so, there is a larger likelihood of returning some spatial (q-domain) noise into the dataset (see Supplementary Information, Noise Estimation in Reconstructed Data and Fig. S4).
Plots of kinetic data extracted using NLSA and SVD from 5 replicates of simulated PYP TR-SAXS difference intensity data for each pair of parameters. The relaxation rates extracted from fitting (a) NLSA and (b) SVD modes. Also plotted are the squared residuals scaled by the squared amplitude of the sigmoid fit to (c) NLSA and (d) SVD modes. Error bars for each plot are the 95% confidence interval given by the standard deviation of the replicates.
To assess the capabilities more broadly, we simulated 5 replicates of each parameter set as done with CaM, and analyzed the data with NLSA and SVD accordingly. The results are shown in Fig. 6. For NLSA, the parameters were chosen according to Table S2 in Supplementary Information. Figure 6a shows that NLSA is usable for kinetics extraction with \(10^5\) photon counts per pulse and long exposure times. Short exposure times are enough when increasing the photon flux to \(10^6\) photons per pulse, which is achievable at CXLS when operating the Laser responsible for generating X-rays at full flux. The extracted times also converge to the imparted value with high precision, effectively removing any of the pulse broadening effects. In contrast, it is not possible to extract relaxation times from \(10^5\) photon count data using SVD (see Fig. 6b). However, it is possible with \(10^6\) photon counts, but not very consistently. Further, these times are on the longer side of the expected value, indicating the impacts of pulse broadening. Without any other preprocessing steps, SVD requires at least \(10^7\) photons per pulse to produce a relaxation time with both accuracy and confidence. Notably, panels Fig. 6c,d indicate that NLSA generates temporal modes with squared residuals around three orders of magnitude smaller than those obtained from the right singular vectors of SVD. This demonstrates substantial temporal noise reduction from NLSA compared to SVD.
Discussion
We have developed and demonstrated a machine learning approach based on the Nonlinear Laplacian Spectral Analysis (NLSA) algorithm that effectively extracts structural dynamics from time-resolved SWAXS data under the low-flux conditions characteristic of compact X-ray sources. By applying this method to simulated datasets for Calmodulin and Photoactive Yellow Protein, we established that NLSA provides a significant advantage over standard Singular Value Decomposition (SVD) in identifying signal where noise is extreme. This capability is vital for the commissioning and early operation of facilities like the CXLS, where initial photon counts may be several orders of magnitude lower than those found at large-scale X-ray free electron lasers.
The primary advantage of the NLSA framework is its ability to perform unsupervised signal discovery in noisy datasets. In our simulations at lower photon counts, the signals associated with structural transitions were clearly isolated in the first few NLSA modes. On the other hand, the standard SVD did not clearly separate these dynamical components and often assigned higher singular values to noise-dominated modes, burying the intrinsic dynamics deep in the spectrum. While SVD can effectively reduce pixel noise through its linear decomposition, it does not adequately capture the nonlinear temporal structure of the data and may therefore mix dynamical modes when the signal is hidden in noisy observations. As a result, identifying the modes associated with the signal using SVD often requires prior knowledge of the expected kinetics. By contrast, NLSA uses time-lagged embedding and nonlinear manifold learning algorithms to exploit the intrinsic geometry of time series data, thereby improving the separation of dynamical components from noise and enabling more accurate identification of underlying biological processes. Additionally, we expect that this method will isolate any unpredicted states that are captured within the dataset. This would depend on the rate of occurrence, i.e. the fraction of population transitioning to this extra state.
A notable feature of this approach is the trade-off between temporal noise reduction and absolute accuracy. In the most challenging low-flux scenarios, the use of a larger delay-coordinate embedding (concatenation) window in NLSA is necessary to resolve the signal from the background noise. While this introduces a systematic shift or smoothing effect in the extracted relaxation times, it allows for the high-precision recovery of the kinetic trend. For an experimentalist working at the limits of detection, the ability to qualitatively confirm a sigmoidal transition is far more valuable than an ambiguous result that remains indistinguishable from noise. As photon flux increases toward design parameters, this systematic bias is mitigated because smaller concatenation windows can be employed without losing the signal.
Based on the results of our simulations, NLSA becomes a powerful tool for strategizing beamtime by fundamentally shifting the requirements for signal detection. In traditional time-resolved experiments, researchers often rely on brute-force fluence to overcome the noise floor and resolve small structural changes. However, our results demonstrate that NLSA can extract clear kinetic trends from as few as 100, 000 photons per pulse, a regime where standard SVD analysis remains effectively blind. This computational capability acts as a signal amplifier in the SWAXS region, allowing scientists to make significant progress even under the current flux limitations of compact sources. By lowering the total photon requirement for a successful experiment, NLSA enables a more efficient use of limited beamtime, as users can screen a larger number of samples or experimental conditions within the same timeframe without sacrificing the ability to identify critical structural transitions.
Furthermore, this methodological shift has direct implications for the feasibility of complex experiments at the laboratory scale. Because NLSA can recover structural dynamics from low-fidelity data that would otherwise require much longer total exposure times to reach a usable signal-to-noise ratio with linear methods, it bridges the gap between the hardware constraints of a compact source and the high-quality temporal resolution required for biological discovery. Ultimately, by integrating NLSA into the data analysis pipeline, facilities like the CXLS can transition from being seen merely as lower-flux versions of national facilities to becoming efficient engines for structural discovery, where sophisticated algorithms compensate for hardware constraints to yield high-impact results.
Beyond its performance in extreme noise, the integration of this algorithm into a user-friendly graphical user interface makes it accessible to a broader community of structural biologists. By simplifying the selection of relevant modes and providing an interactive parameter search via heatmaps, the GUI ensures that non-expert users can maximize the information content of their collected data. This approach is not limited to compact light sources but is also applicable to other evolving mixtures, such as those found in size-exclusion chromatography coupled SWAXS experiments at synchrotron beamlines. Ultimately, this methodology enhances scientific throughput by enabling meaningful results to be obtained from fewer exposures or lower flux, reducing beamtime requirements and improving the likelihood of successful applications of these new X-ray sources.
Methods
NLSA algorithm
NLSA is essentially a nonlinear counterpart of the well-established Singular Spectrum Analysis (SSA) technique45, capable of extracting meaningful information from noisy time series data without requiring prior knowledge of the underlying system’s dynamics. The standard SSA relies on linear principal components (or empirical orthogonal functions) of data, whereas NLSA integrates SSA with advanced graph theory-based machine learning algorithms, making it more suitable for extracting signals generated by nonlinear and multidimensional dynamical systems35,38,46. The fundamental elements of the NLSA algorithm are summarized as follows.
Time-lagged embedding
The primary step of NLSA is called time-lagged (or delay-coordinate) embedding34,38, where the original time series of N observations \(x_i\) from an object is embedded in a D-dimensional data space. Given N data points \(\{x_1,x_2,...,x_N \}\) ordered in time and a concatenation parameter c, this procedure constructs a so-called trajectory (or Hankel) matrix X whose columns are supervectors \(X_i=[x_i, x_{i-1}, x_{i-2},...,x_{i-c}]\), each consisting of c consecutive data points within the concatenation window c. The trajectory matrix has a number of rows given by \(D'\equiv cD\) and columns given by \(N'\equiv N-c+1\). According to Takens’ theorem47, this embedding process enables access to the dynamics of the system (e.g. a protein) from which the time series data was obtained. In SSA, this goal can be achieved by applying an appropriate matrix factorization method to the matrix X, such as the singular value decomposition (SVD) technique.
Diffusion map embedding
Classical SSA has limited power to identify complex and nonlinear features hidden in the original data. To resolve this issue, in its second step, NLSA discovers the manifold from the concatenated dataset X. This is done through a dimensionality reduction technique, called diffusion map (DM). As a graph-theoretic algorithm, DM projects the high-dimensional data manifold of X onto a lower-dimensional manifold in the diffusion space whose components are the eigenfunctions of a normalized graph Laplacian, which typically approximates the Laplace-Beltrami operator25. The key steps of the DM algorithm are described as follows25.
-
Calculate the pairwise Euclidean distances \(S_{ij}=\Vert X_i-X_j \Vert\) between the column vectors \(X_i\) and \(X_j\) of the matrix X.
-
Construct a real and symmetric similarity kernel matrix \(K=[K_{ij}]\) (dim. \(N'\times N'\)), where
$$\begin{aligned} K_{ij}=\exp (-S_{ij}^2/2\sigma ^2). \end{aligned}$$(1)The Gaussian bandwidth \(\sigma\) determines the locality of the similarity measure and is practically set based on Euclidean distances of each data point to a fixed number of its nearest neighbors (nN).
-
Regularize the kernel matrix by the manifold sampling density:
$$\begin{aligned} \tilde{K}_{ij} = \frac{K_{ij}}{(\sum _m K_{im})^\alpha (\sum _m K_{mj})^\alpha } \end{aligned}$$(2)where \(\alpha\) is a user selected parameter discussed below.
-
Normalize the diffusion kernel and generate the transition probability matrix \(P = D^{-1}\tilde{K}\), where D is a diagonal degree matrix with elements given by \(D_{ii}=\sum _j \tilde{K}_{ij}\).
-
Build \(L=I-P\), where I is the identity matrix. For \(\alpha =1\), the effects of sampling density are minimized and L approximates the Laplace-Beltrami operator on the manifold. Else, if \(\alpha =0\), L is the standard graph Laplacian25.
-
Find the eigenvalues \(\{\lambda _i\}\) and eigenfunctions \(\{\Phi _i\} (i=1,..,N')\) of the matrix L (or P). These orthogonal eigenfunctions define a new space of diffusion distances in which the data manifold is embedded and corresponds to intrinsic features in the data25. Selecting \(m< N\) eigenfunctions, guided by the eigenvalue spectrum, defines the reduced dimensionality by the diffusion map algorithm.
Projecting data onto manifold
The curved structure of the manifold revealed by the diffusion map enables the extraction of nonlinear features hidden within data. Therefore, in this step, we construct a new matrix operator A by projecting the concatenated data matrix X onto the subspace \(\Phi\) of the manifold obtained from X as
Here, the columns of matrix \(\Phi\) (dim. \(N\times M\)) are the leading eigenfunctions of the diffusion map and matrix \(\mu\) denotes the Riemannian measure of the manifold, which has the elements of the first (trivial) eigenvector obtained from DM squared along the diagonal (see Ref.38 for more details).
SVD on manifold
To uncover the fundamental characteristics of the system, singular value decomposition is applied to the matrix A in the new manifold space. As in the standard SVD, this produces three components, namely U, S, and V, so that
where M stands for the number of SVD modes. Here, we use the terms “topos” and “chronos” for the left and right eigenvectors U and V, respectively. The singular values in the diagonal matrix S quantify the strength of SVD modes.
NLSA reconstruction
The final stage of NLSA involves the reconstruction of data using a selected subset of SVD modes and projecting the results back to the data space. This is performed through the matrix equation
where m stands for the number of selected modes and \(\Phi ^T\) is the transpose of the matrix \(\Phi\). The matrix of NLSA chronos thus becomes \(V\Phi\) in the data space, returning the reaction coordinates associated with the system’s dynamics. The number of modes to be retained is typically determined by the number of singular values that appear before the flat tail of the singular value spectrum. This step is followed by a diagonal averaging (unwrapping) of the matrix \(\tilde{X}\), similar to the SSA algorithm48. The output is a time series \(\{\tilde{x}_i\}\) of NLSA reconstructed data, ready for interpretation or further analysis. Figure1 summarizes the major steps of the whole data analysis pipeline, including data pre-processing of raw data, the NLSA, as well as post-processing and interpretation of the NLSA reconstructed data. It should be noted that including time-lagged embedding, SVD, and diffusion map in the NLSA algorithm allows filling gaps in case of data sparsity and incompleteness, reducing pixel and temporal noise or other data anomalies. It also enables easier access to important and meaningful signals hidden in the original data through the curved manifold.
Here we should also note that due to the time-lagged embedding, a set of \(c-1\) snapshots is discarded from the beginning of the original time series in NLSA. In addition, during the unwrapping processes described above, an additional number of \(nCopy-1\) snapshots are excluded from the end of the dataset. This reduction is particularly important when comparing the NLSA output with the original data or the results obtained from other methods, such as SVD, where the full length of the time series is usually preserved. For example, for the CaM data in this work with \(N=100\) time points, we selected \(c=8\) and \(nCopy=2\), thus 7 time points were surrendered from the beginning of the time series as well as one from the end.
Data simulation
Diffraction intensities (mean photon counts) were simulated using the equation
where n is the index of the area-detector pixel, \(J_0\) is the incident fluence (photons per area), \(r_e\) the classical electron radius (\(2.818 \times 10^{-15}\) m), N the number of proteins in the illuminated volume of liquid (determined by sample thickness and concentration), \(\Delta \Omega _n\) is the solid angle of the detector pixel, \(P_n\) is the polarization factor (assuming linear polarization), \({\bf q} = {\bf k}_n - {\bf k}_0\) is the wavevector transfer between the incoming wavevector \({\bf k}_0\) and the outgoing wavevector \({\bf k}_n\) directed at pixel n. Both wavevectors have a magnitude of \(2\pi /\lambda\), where \(\lambda\) is the photon wavelength. The terms \(\mathcal {P}(q_n)\) and \(\mathcal {W} (q_n)\) correspond to protein and bulk water scattering profiles, respectively, which we describe below.
A single-protein scattering profile for the structural configuration m is computed with the equation
where \(\rho _m({\bf r})\) is the scattering density contrast generated using the denss-pdb2mrc tool in DENSS39,40 and the angle brackets \(\left\langle \cdot \right\rangle\) represent a uniform average over rotations of coordinates \({\bf r}\). Critically, DENSS includes an accurate estimate of the solvent contributions that include both the effect of displaced bulk solvent (“excluded volume”) and the added scattering from the hydration shell of partially ordered solvent molecules at the particle surface. DENSS has been shown to generate accurate scattering profiles in both the SAXS and WAXS regions, closely matching experimental data40. Note that t denotes the time, and in our case the time series is constructed by a superposition of two different scattering densities representing two different protein states:
where \(c_1(t)\) is the time-varying concentration fraction of component 1. The water scattering profile \(\mathcal {W} (q_n)\) is computed using carefully calibrated experimental measurements49,50 via reborn.
We note that a helium gas background was not included in the simulations since the signals are much lower than the protein/water signals under our expected vacuum environment of \(10^{-5}\) mbar pressure, and a 500 \(\upmu\)m sample thickness. We also omitted detector noise since the CXLS is equipped with an Eiger 4M photon-counting detector assuming that the signals closely follow an ideal Poisson distribution. The most significant omission was parasitic background signals that were dependent upon the beamline slits and their optimization.
In order to create the 1D scattering profiles from the 2D area detector simulations, we first summed all of the intensities according to q bins:
where the notation \(\{n\}_k\) represents the set of pixel indices for the kth bin in the scattering profile. Then we divided the sum by the weights \(w_i = J_0 \, r_e^2 \Delta \Omega _i P_i\),
to produce the simulated measurement of the differential scattering cross section of the target, including all proteins and water molecules illuminated by the x-ray beam. We included noise by sampling a Poisson distribution with means equal to \(T_k(t)\).
Protein simulations
To prepare SWAXS profiles for the NLSA analysis, the scattering sum was first calculated as the total number of photons in each bin for a given X-ray pulse. The bins (500 in our CaM simulation; 300 for PYP) were defined by a set of pixels lying within circular annuli on the 2D detector plane and were sorted by scattering angle (wavevector transfer) q. The magnitude of q assigned to each bin represented the value at the center of its annulus.
In the next step, the scattering sum was divided by the total solid angle associated with each bin, as shown in Eq. 10. This normalization is beneficial because, in real experiments, the solid angles can vary from shot to shot. The resulting quantity is often referred to as the “differential scattering cross section”, and should be used when interpreting SWAXS profiles in terms of electron density pair distribution functions. An average dark state measured from several replicate measurements (10 for CaM; 50 for PYP) were then subtracted from the differential scattering cross sections of each time delay. This resulted in difference intensity profiles which were used in our analysis. The cited normalization and differencing were the only preprocessing steps used before the NLSA or SVD steps.
As a validation check for the NLSA algorithm, a control trial was performed with PYP simulations. A faux-time-series of 1000 noisy profiles were created from the same dark state; no actual dynamics were present. As is described in Supplementary Information, NLSA did not discover any artificial trends across the series of profiles, matching our expectations. Additionally, these profiles were reconstructed with a subset of leading modes. Both methods denoised the reconstructions to an equal extent, thus showing the benefit of using these methods despite a lack of any time evolution.
To evaluate the current experimental setup and measurements at CXLS, we examined a range of photon counts, including \(10^5\), \(10^6\), \(10^7\), and \(10^8\) photons per snapshot. Additionally, in line with traditional time-resolved SWAXS experiments6,8,10,12, we considered a limited number of time points, such as 100 for CaM and 1000 for PYP. Also, multiple datasets were generated for the analysis by varying the number of X-ray exposures per time point via the exposure time in a 1 kHz repetition rate beam (pulse energy of 740 nJ). In general, the NLSA algorithm performs better with higher photon counts and exposures and more time points. We present results for many combinations of parameters, but we focus on the results from the dataset with \(10^5\) photon counts and low exposure times to evaluate our method under the most challenging conditions anticipated for SWAXS data collection at CXLS.
NLSA GUI main window displaying heat map results. The interactive heatmap generated after running the parameter search. The colors indicate the number of eigenfunctions (derived from the parameter sets on the \(x-\) and \(y-\)axes) that have a correlation coefficient of 0.9 or higher. The lower triangle shows correlations grouped first by nN, then by \(\sigma _f\), while the upper triangle groups first by \(\sigma _f\), then by nN. The main diagonal is excluded because it reflects self-correlation. The pop-up window displaying the parameters of the selected cell (outlined in red) is also shown in this figure.
User-friendly GUI
The algorithm described in this paper is based on an unsupervised machine learning method, which is nonlinear dimensionality reduction via diffusion map. Therefore, it requires the user’s advice to select the embedding parameters such as the concatenation order c, the number of nearest-neighbors nN, the \(\sigma _f\) factor, as well as the number of DM eigenfunctions \(\{\Phi _i\}\) to construct the matrix A in Eq. (3). Choosing these parameters typically requires basic experience and knowledge in the manifold learning fields. Consequently, to simplify usage, we have developed a graphical user interface (GUI) for the main part of the code. With CXLS/CXFEL becoming user facilities, it is of paramount importance that inexperienced users can make use of the tools described here to maximize the information output of their collected tr-SWAXS data. We outline the key steps here, but additional details for running the program can be found in the online documentation available on the GitHub repository of the code (see Ref.51).
-
(i)
The analysis begins with loading the preprocessed data (with h5 as default format) into the GUI, and the first step is the parameter search for the manifold analysis. This helps users identify the optimal values of nN and \(\sigma _f\). The concatenation order c is typically set to a number around or below \(10\%\) of the total number of snapshots (time points in time-resolved SWAXS data here). Given a specified c value, as well as a user-defined range of nN and \(\sigma _f\), the time-lagged and manifold embedding processes are performed for each set of these parameters. Pearson’s correlations are then calculated among DM eigenfunctions obtained from each combination of the parameters \(\sigma _f\) and nN. The outcome of this systematic search is visualized through an interactive heatmap, which is displayed in the main window of the GUI (see Fig. 7). Higher correlation coefficients reflect stronger agreement between the resulting eigenfunctions and indicate the robustness of the DM embedding output against the variations in parameters. This is the primary criterion for choosing a cell in the heatmap. The above procedure needs to be repeated for a few additional values of c (e.g., by searching around the initial selection) until the robustness of the DM eigenfunctions against change in c is observed through the heatmap. Then the corresponding c is considered as the optimal concatenation order.
GUI NLSA results tab. This tab of the GUI displays the singular values and chronos resulting from NLSA. These plots appear after the “Run NLSA” process is complete. Chronos selected for reconstruction are outlined in red, and once selected, pressing “NLSA Reconstruct” will generate the reconstructed dataset and save it in a file.
-
(ii)
Selecting any cell in the heatmap opens a pop-up menu that provides information about the associated parameters and the number of matching eigenfunctions. This window also includes two buttons, each corresponding to a specific set of nN and \(\sigma\). Clicking either button loads the corresponding parameter values into the left panel of the GUI for visualization of DM eigenvalues, and then eigenfunctions appear in a new tab (not shown here) to make a decision about the next part of the analysis.
-
(iii)
The next step is the calculation of the NLSA components, including topos U, chronos V, and singular values S, as demonstrated in “Methods”. This operation is initiated through the“Run NLSA” button in the main window of the GUI, and upon completion, the chronos and singular values are displayed in a new tab, as shown in Fig. 8.
-
(iv)
The last stage of the main program requires selecting the appropriate NLSA chronos (modes). This can be done by clicking on the desired chronos in the new tab, using singular values as guidance. Typically, chronos corresponding to the leading singular values are selected, as in the standard SVD. This is followed by the NLSA reconstruction using a selected number of copies associated with time-lagged embedding, triggered via the last button in the GUI (Run NLSA Reconstruction). The reconstructed and denoised data are then stored in a file for post-processing and additional analysis determined by users.
Data availability
All simulation code is available through the respective GitLab repositories. The NLSA program is also available on GitHub. All data will be made available upon reasonable request to the corresponding authors.
References
Barends, T. R. M., Stauch, B., Cherezov, V. & Schlichting, I. Serial femtosecond crystallography. Nat. Rev. Methods Prim. 2, 59. https://doi.org/10.1038/s43586-022-00141-7 (2022).
Botha, S. & Fromme, P. Review of serial femtosecond crystallography including the COVID-19 pandemic impact and future outlook. Structure 31, 1306–1319. https://doi.org/10.1016/j.str.2023.10.005 (2023).
Neutze, R., Wouts, R., van der Spoel, D., Weckert, E. & Hajdu, J. Potential for biomolecular imaging with femtosecond X-ray pulses. Nature 406, 752–757. https://doi.org/10.1038/35021099 (2000).
Chapman, H. N. et al. Femtosecond diffractive imaging with a soft-X-ray free-electron laser. Nat. Phys. 2, 839. https://doi.org/10.1038/nphys461 (2006).
Chapman, H. N. et al. Femtosecond X-ray protein nanocrystallography. Nature 470, 73–77. https://doi.org/10.1038/nature09750 (2011).
Arnlund, D. et al. Visualizing a protein quake with time-resolved X-ray scattering at a free-electron laser. Nat. Methods 11, 923–926. https://doi.org/10.1038/nmeth.3067 (2014).
Levantino, M. et al. Ultrafast myoglobin structural dynamics observed with an X-ray free-electron laser. Nat. Commun. 6, 6772. https://doi.org/10.1038/ncomms7772 (2015).
Lee, Y. et al. Ultrafast coherent motion and helix rearrangement of homodimeric hemoglobin visualized with femtosecond X-ray solution scattering. Nat. Commun. 12, 3677. https://doi.org/10.1038/s41467-021-23947-7 (2021).
Zielinski, K. A. et al. RNA structures and dynamics with Å resolution revealed by X-ray free-electron lasers. Sci. Adv. 9, eadj3509. https://doi.org/10.1126/sciadv.adj3509 (2025).
Karpos, K. et al. Pump-probe TR-WAXS distinguishes activation from heating effects in ultrafast rhodopsin dynamics. In SPring-8/SACLA Research Report. Vol. 13. 175–182. https://doi.org/10.18957/rr.13.3.175 (2025) .
Blanchet, C. E. et al. Form factor determination of biological molecules with X-ray free electron laser small-angle scattering (XFEL-SAS). Commun. Biol. 6, 1057. https://doi.org/10.1038/s42003-023-05416-7 (2023).
Konold, P. E. et al. Microsecond time-resolved X-ray scattering by utilizing MHz repetition rate at second-generation xfels. Nat. Methods 21, 1608–1611. https://doi.org/10.1038/s41592-024-02344-0 (2024).
Decking, W. et al. A MHz-repetition-rate hard X-ray free-electron laser driven by a superconducting linear accelerator. Nat. Photon. 14, 391–397. https://doi.org/10.1038/s41566-020-0607-z (2020).
Emma, P. et al. First lasing and operation of an ångstrom-wavelength free-electron laser. Nat. Photon. 4, 641–647. https://doi.org/10.1038/nphoton.2010.176 (2010).
Bonifacio, R., Pellegrini, C. & Narducci, L. M. Collective instabilities and high-gain regime in a free electron laser. Opt. Commun. 50, 373–378. https://doi.org/10.1016/0030-4018(84)90105-6 (1984).
Pellegrini, C., Marinelli, A. & Reiche, S. The physics of X-ray free-electron lasers. Rev. Mod. Phys. 88, 15006. https://doi.org/10.1103/RevModPhys.88.015006 (2016).
Emma, P. et al. First lasing and operation of an ångstrom-wavelength free-electron laser. Nat. Photon. 4, 641–647. https://doi.org/10.1038/nphoton.2010.176 (2010).
Ishikawa, T. et al. A compact X-ray free-electron laser emitting in the sub-ångström region. Nat. Photon. 6, 540–544. https://doi.org/10.1038/nphoton.2012.141 (2012).
Kang, H.-S. et al. Hard X-ray free-electron laser with femtosecond-scale timing jitter. Nat. Photon. 11, 708–713. https://doi.org/10.1038/s41566-017-0029-8 (2017).
Prat, E. et al. A compact and cost-effective hard X-ray free-electron laser driven by a high-brightness and low-energy electron beam. Nat. Photon. 14, 748–754. https://doi.org/10.1038/s41566-020-00712-8 (2020).
Günther, B. et al. The versatile X-ray beamline of the Munich compact light source: Design, instrumentation and applications. J. Synchrotron Radiat. 27, 1395–1414. https://doi.org/10.1107/S1600577520008309 (2020).
Graves, W. S. et al. Asu compact xfel. In Proceedings of the 38th International Free-Electron Laser Conference, FEL 2017. 225–228. https://doi.org/10.18429/JACoW-FEL2017-TUB03 (JACoW Publishing, 2017).
Spence, J. et al. The Asu Compact Xfel Project. Bull. Am. Phys. Soc. (2020).
Graves, W. The CXFEL Project at Arizona State University. In Proceedings of the 67th ICFA Advanced Beam Dynamics Workshop Future Light Sources (FLS’23). Vol. 67 . 54–57. https://doi.org/10.18429/JACoW-FLS2023-TU1C4 (JACoW Publishing, 2024).
Coifman, R. R. & Lafon, S. Diffusion maps. Appl. Comput. Harmonic Anal. 21, 5–30 (2006).
Coifman, R. R. & Lafon, S. Geometric harmonics: A novel tool for multiscale out-of-sample extension of empirical functions. Appl. Comput. Harmonic Anal. 21, 31–52 (2006).
Coifman, R. R., Kevrekidis, I. G., Lafon, S., Maggioni, M. & Nadler, B. Diffusion maps, reduction coordinates, and low dimensional representation of stochastic systems. Multiscale Model. Simul. 7, 842–864 (2008).
Giannakis, D., Schwander, P. & Ourmazd, A. The symmetries of image formation by scattering. I. Theoretical framework. Opt. Exp. 20, 12799–12826 (2012).
Schwander, P., Giannakis, D., Yoon, C. H. & Ourmazd, A. The symmetries of image formation by scattering. II. Applications. Opt. Exp. 20, 12827–12849 (2012).
Hosseinizadeh, A. et al. High-resolution structure of viruses from random diffraction snapshots. Philos. Trans. R. Soc. B Biol. Sci. 369, 20130326 (2014).
Hosseinizadeh, A. et al. Conformational landscape of a virus by single-particle X-ray scattering. Nat. Methods 14, 877–881 (2017).
Dashti, A. et al. Trajectories of the ribosome as a Brownian nanomachine. Proc. Natl. Acad. Sci. 111, 17492–17497 (2014).
Dashti, A. et al. Retrieving functional pathways of biomolecules from single-particle snapshots. Nat. Commun. 11, 4734 (2020).
Fung, R. et al. Dynamics from noisy data with extreme timing uncertainty. Nature 532, 471 (2016).
Hosseinizadeh, A. et al. Few-fs resolution of a photoactive protein traversing a conical intersection. Nature 599, 697–701 (2021).
Fung, R. et al. Achieving accurate estimates of fetal gestational age and personalised predictions of fetal growth based on data from an international prospective cohort study: A population-based machine learning study. Lancet Digit. Health 2, e368–e375 (2020).
Hosseinizadeh, A., Dashti, A., Schwander, P., Fung, R. & Ourmazd, A. Single-particle structure determination by X-ray free-electron lasers: Possibilities and challenges. Struct. Dyn. 2, 041601 (2015).
Giannakis, D. & Majda, A. J. Nonlinear Laplacian spectral analysis for time series with intermittency and low-frequency variability. Proc. Natl. Acad. Sci. 109, 2222–2227 (2012).
Grant, T. D. Ab initio electron density determination directly from solution scattering data. Nat. Methods 15, 191–193. https://doi.org/10.1038/nmeth.4581 (2018).
Chamberlain, S. R., Moore, S. & Grant, T. D. Fitting high-resolution electron density maps from atomic models to solution scattering data. Biophys. J. 122, 4567–4581 (2023).
Kirian, R. Reborn. https://gitlab.com/kirianlab/reborn. Accessed 09 Jul 2025 (2025).
Kuboniwa, H. et al. Solution structure of calcium-free calmodulin. Nat. Struct. Biol. 2, 768–776. https://doi.org/10.1038/nsb0995-768 (1995).
Ikura, M. et al. Solution structure of a calmodulin-target peptide complex by multidimensional NMR. Science 256, 632–638. https://doi.org/10.1126/science.1585175 (1992).
Park, H. Y. et al. Conformational changes of calmodulin upon Ca<sup>2+</sup> binding studied with a microfluidic mixer. Proc. Natl. Acad. Sci. 105, 542–547. https://doi.org/10.1073/pnas.0710810105 (2008).
Vautard, R. & Ghil, M. Singular spectrum analysis in nonlinear dynamics, with applications to paleoclimatic time series. Phys. D Nonlinear Phenom. 35, 395–424 (1989).
Giannakis, D. & Majda, A. J. Nonlinear Laplacian spectral analysis: Capturing intermittent and low-frequency spatiotemporal patterns in high-dimensional data. Stat. Anal. Data Min. ASA Data Sci. J. 6, 180–194 (2013).
Takens, F. Dynamical systems and turbulence, Warwick 1980. In Lecture Notes in Mathematics. Vol. 898. 366–381 (Springer, 1981).
Hassani, H. Singular spectrum analysis: Methodology and comparison. J. Data Sci. 5, 239–257 (2007).
Hura, G., Sorenson, J. M., Glaeser, R. M. & Head-Gordon, T. A high-quality X-ray scattering experiment on liquid water at ambient conditions. J. Chem. Phys. 113, 9 (2000).
Clark, G. N. I., Hura, G. L., Teixeira, J., Soper, A. K. & Head-Gordon, T. Small-angle scattering and the structure of ambient liquid water. Proc. Natl. Acad. Sci. 107, 14003–14007. https://doi.org/10.1073/pnas.1006599107 (2010).
Fung, R., Golla, S., Huang, S. & Hosseinizadeh, A. NLSA-SWAXS. https://github.com/UWM-CXFEL/NLSA_SWAXS (2025). Accessed 30 Sep 2025.
Funding
This work was supported by the U.S. National Science Foundation Bio Directorate under midscale research infrastructure Grants 2153503 and 1935994, and by the US Department of Energy, Basic Energy Sciences, under Award No. DE-SC0002164. Thomas D. Grant further acknowledges support from the U.S. National Institute of Health, under Award No. R35GM158053 and Richard A. Kirian acknowledges support from the U.S. National Science Foundation, under Award No. 1943448.
Author information
Authors and Affiliations
Contributions
A.H., R.A.K., T.D.G and S.B. conceived the experiment(s), T.D.G. and R.A.K. developed the simulation code, A.O. and S.H. performed simulations and the NLSA analysis, R.F., A.H., and S.N.G. contributed to the development of the NLSA code, S.N.G. and S.H. developed the NLSA GUI, A.K.O., S.H., S.B., K.E.S., T.D.G., R.A.K., and A.H. analysed the results. A.K.O., S.B., T.D.G., R.A.K., and A.H. drafted the manuscript, A.K.O. made the figures. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Opperman, A.K., Huang, S., Botha, S. et al. Signal extraction in SWAXS data for the compact X-ray light sources: a machine learning approach. Sci Rep 16, 11712 (2026). https://doi.org/10.1038/s41598-026-47265-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-47265-4