Abstract
Measurement techniques often result in domain gaps among batches of cellular data from a specific modality. The effectiveness of cross-batch annotation methods is influenced by inductive bias, which refers to a set of assumptions that describe the behavior of model predictions. Different annotation methods possess distinct inductive biases, leading to varying degrees of generalizability and interpretability. Given that certain cell types exhibit unique functional patterns, we hypothesize that the inductive biases of cell annotation methods should align with these biological patterns to produce meaningful predictions. In this study, we propose KIDA, Knowledge-based Inductive bias and Domain Adaptation. The knowledge-based inductive bias constrains the prediction rules learned from the reference dataset, composed of multiple batches, to functional patterns relevant to biology, thereby enhancing the generalization of the model to unseen batches. Since the query dataset also contains gaps from multiple batches, KIDA’s domain adaptation employs pseudo labels for self-knowledge distillation, effectively narrowing the distribution gap between model predictions and the query dataset. Benchmark experiments demonstrate that KIDA is capable of achieving accurate cross-batch cell type annotation.
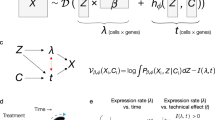
Similar content being viewed by others
Introduction
The rapid accumulation of cellular data creates opportunities for artificial intelligence (AI) in bioinformatics1, where cellular data annotation is an essential task that drives numerous related scientific fields2,3,4. Typically, we manually annotate cellular labels on a few batches from initial measurements (reference dataset), then train machine learning models and apply these models to predict cell labels for batches derived from subsequent measurements (query dataset). However, current machine learning methods for cell type annotation face multiple challenges.
The first challenge in cellular data annotation arises from batch effects5,6. Technical noise and biological artifacts result in observation offsets between multiple batches that make up the reference dataset or query dataset. Thus, when dealing with cells of the same type from various batches, there will be notable differences in the embeddings produced directly after reducing the dimensionality. The goal of batch integration is to eliminate gaps while preserving biological heterogeneity (clustering cells of the same type together). Batch effects can interfere with the discriminability of cell representations. The current standard pipeline involves first aligning cell representations from heterogeneous batches through batch integration and then identifying low-dimensional cell representations using classifiers7,8,9. In the standard pipeline, annotation, and batch integration are independently considered, making it difficult to track the contributions of inputs, resulting in loss of interpretability. Furthermore, we are often required to provide additional batch labels to achieve batch integration. Another challenge is the modality heterogeneity of cellular data. Taking two measurement technologies as examples, the feature space for the scRNA-seq modality is genes, while the feature space for the scATAC-seq modality is open chromatin peaks. There are already many machine learning methods available for annotating scRNA-seq modality. However, due to inconsistently defined feature spaces across modalities, methods specifically designed for annotating scATAC-seq modality are relatively scarce10. There is an urgent need to develop a unified annotation method that supports multi-modal cellular data.
For the first challenge, it is known that specific types of cells have specific functional patterns. However, existing machine learning methods achieve predictions by identifying patterns in the data that are related to cell types but may lack functional relevance. These patterns are referred to as non-causal bias (there is a correlation but no causation)11,12,13,14. The correlation and causality of patterns depend on inductive biases15,16. Inductive bias in machine learning refers to a set of explicit or implicit assumptions that learning algorithms adopt to make predictions on unseen inputs17,18. Essentially, inductive bias helps narrow down the hypothesis space, allowing learning algorithms to prioritize one solution over another19,20,21. Inspired by TOSICA22, the phenotype of a cell can be explained by pathways, and the gene sets of the pathways correspond to specific functional patterns23,24,25,26. Therefore, we have the hypothesis that the annotation model should convert genes into functional patterns (gene sets) and then predict labels. Therefore, function-based inductive bias guides the model to focus on interactions relevant to biological functions rather than observations susceptible to batch effects. We illustrate this motivation using image-based species classification27,28,29. The relationships and semantics between local regions of animals are discriminative information unaffected by camera angles. In cellular data, batch effects correspond to pixel differences caused by camera angles, functional patterns correspond to local semantic regions, and interactions between functional patterns correspond to relationships between local regions. For the second challenge, intuitively, unification of the feature space is necessary, and tools already exist to efficiently standardize it. For example, Seurat and UnpairReg can convert peaks corresponding to scATAC-seq into gene scores for scRNA-seq7,30. However, the conversion may further exacerbate batch effects on gene scores, thus reducing the robustness of annotation methods. Since function-based inductive bias is batch-insensitive and does not require batch information, it can naturally be applied to feature space transformation tools.
In this paper, we propose KIDA, Knowledge-based Inductive bias and Domain Adaptation, a method for cross-batch cell type annotation. Knowledge-based inductive bias is embodied in a set of parallel linear transformations. Each branch of the parallel linear transformations is used to extract a specific latent pattern from high-dimensional counts data. Furthermore, we utilize biological knowledge from web or local databases (such as pathway collections, ‘gmt’ files) to parameterize the linear transformations. Such an inductive bias not only reduces cell representations from tens of thousands of genes to hundreds of pathways but also constrains the classification rules to functional patterns relevant to biological knowledge. Therefore, knowledge-based inductive bias ensures that the model can generalize to unseen batches or datasets with similar functional pattern interactions. Functional patterns are represented as tokens by parallel linear transformations, and we use self-attention to learn the interactions between these functional patterns. Additionally, due to the overlay of multiple batch effects in query dataset, the distribution consistency between the reference dataset and the query dataset is very low. We use a two-round supervised learning method to achieve domain adaptation. We first train the model on the reference dataset and annotate the query dataset. On the query dataset, the model selects some cells (anchors) deemed to predict well to form a new training set. Then, the model utilizes pseudo labels of the anchors for self-knowledge distillation. Finally, the distilled model is used to predict the cell types in query dataset. Benchmark experiments demonstrate that KIDA achieves accurate cross-batch cell type annotation. The integration cellular atlas reveals the robustness of KIDA. This paper contains the following contributions: (1) We use an inductive bias based on biological knowledge to constrain machine learning models to correlate with cellular functional patterns, thereby improving model generalization on unseen batches. (2) We use a two-round supervised learning method to enhance domain adaptability between the model predictions and the query dataset. (3) KIDA is an annotation method that supports multi-modal cellular data.
Results
KIDA overview
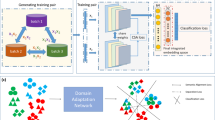
The workflow of KIDA is illustrated in Fig. 1(a) to (f), with each subplot representing a step. Step (a) is used to unify the feature space of cellular data from arbitrary modalities into genes. Steps (b) to (d) cover biological knowledge from web or local databases into the parameters of the model, transforming features from genes to functional patterns, corresponding to the knowledge-based inductive bias proposed in this paper. Details of these steps are provided in section 4.1. Step (e) utilizes self-attention to learn the interactions between functional patterns on the reference dataset and adds a token to represent cells. In step (f), the model from step (e) predicts pseudo labels on the query dataset, and then performs self-knowledge distillation using these pseudo labels. Steps (e) and (f) correspond to the two rounds of supervised training proposed in this paper. Details of these steps are provided in section 4.2. Interpretability of KIDA is provided in section 4.3.
a Unified feature space as genes. b Retrieve pathway names based on gene names. c Mask the parameters of the linear transformation using gene-pathway relationships. d Transform features from genes to functional patterns. e First round of supervised learning: learn the interaction of functional patterns on the reference dataset and add a token as cell embedding. f Second round of supervised learning: on the query dataset, the model from step (e) selects anchors (cells with high confidence) to form a new training set. Then, the model utilizes pseudo labels of the anchors for self-knowledge distillation. Orange arrows are training, gray arrows are inference.
Datasets and baselines
BMMC31: This dataset comprises single-cell RNA sequencing (scRNA-seq) data collected from bone marrow mononuclear cells, encompassing a total of 13 batches, 22 distinct cell types, and an overall cell count of 69,249. PBMC32: This dataset includes single-cell ATAC sequencing (scATAC-seq) data obtained from peripheral blood mononuclear cells, consisting of 2 batches, 6 cell types, and a total cell count of 9058. Pan-cancer33,34: This dataset features scRNA-seq data comprising 71,113 myeloid cells, categorized into 23 cell types. It encompasses a total of 13 cancer types, with each cancer type treated as a separate batch. In the spatial transcriptome dataset, each sample represents a spot (containing multiple cells), and the objective of the annotation task is to identify the tissue region type corresponding to each spot. DLPFC35: This dataset contains 12 batches (slices) of the adult dorsolateral prefrontal cortex. Each slice includes mRNA expression data and 2D spatial coordinates for each spot. KIDA utilizes only the mRNA expression data to identify the tissue type of each spot. Additionally, we employ multimodal integration to validate the advantages of KIDA’s cellular representations. The multimodal datasets utilized in the experiments include: 10x-Multiome36, Chen-201937, Ma-202038, Muto-202139, and Human 15 Organs40,41. Each multimodal dataset comprises two modalities: scRNA-seq and scATAC-seq, with further details provided in Supplementary Text A.1. Statistical information for all datasets is available in Supplementary Table B.1.
Data preprocessing follows the standard scanpy, we extract highly variable features, then normalize and scale the dataset. For BMMC, we select the first 6 batches to form the reference dataset and the remaining 7 batches to form the query dataset. For PBMC, we select the first batch as the reference dataset and the remaining batch as the query dataset. Pan-cancer was used to validate the performance of KIDA on cross-disease cell type annotation. We use three batches (ESCA: esophageal carcinoma, THCA: thyroid carcinoma, and UCEC: uterine corpus endometrial carcinoma) as the reference dataset, and the remaining diseases (batches) as the query dataset. For DLPFC, we select the first 6 slices as the reference dataset and the remaining slices as the query dataset. For the datasets of integration, we randomly select 50% to form the reference dataset and the rest as the query dataset.
We use follow methods for annotation as baselines: Seurat7, CellTypist42, ACTINN43, TOSICA22, Cellcano44, MetaTiME45, Geneformer46, scBERT47, CellLM48, LangCell49, scGPT50. We use follow methods for multimodal integration as baselines: Seurat7, GLUE51, harmony52, LIGER53, bindSC54, iNMF55, CoVEL56, unioncom5, scButterfly57. We describe these baselines in the Supplementary Text A.2. For cell type annotation, we use the following evaluation metrics: Acc (Accuracy), F1 score. For integration, we use the following evaluation metrics: NMI (Normalized Mutual Information), ARI (Adjusted Rand Index), Overall score. Details are provided in the Supplementary Text A.3.
Annotation benchmarks
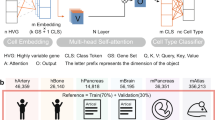
We evaluated KIDA and comparative methods on four datasets related to cell type annotation. These datasets (BMMC, PBMC, DLPFC, Pan-cancer) are modal heterogeneous, with only BMMC and Pan-cancer coming from scRNA-seq. The evaluation results are shown in Fig. 2(a). We divided all methods into two sections, with the first section including all comparative methods and the second section including KIDA and KI (KIDA without domain adaptation). Note that all comparative methods can be further divided into large model-based methods and lightweight models. Geneformer, scBERT, CellLM, LangCell and scGPT are large model-based methods. In Fig. 2(a), including large model-based methods, all comparative methods used the same settings to train or fine-tune on the reference dataset and then test on the query dataset. None of the comparative methods could cover cell data annotation in multiple heterogeneous modalities. Although large models overall exhibit excellent annotation accuracy, their architecture is always complex. Tosiac has better annotation performance on Pan-cancer, which reflects the advantage of training from scratch in the cross-disease scenario (models focus on specific diseases). However, compared to KIDA, we found KIDA to achieve more robust performance. Furthermore, KI also showed competitiveness on four datasets, validating the effectiveness of the proposed knowledge-based inductive bias.
a Comparison of accuracy and F1 on four datasets. There are n = 8 repeats with different random seeds. The error bars indicate mean ± s.d. b Inference time of various methods on the different data sizes, ranging from small to large. The comparison methods here do not include methods based on large models. c UMAP visualizations on BMMC (first row) and PBMC (second row). TOSICA is a SOTA lightweight model dedicated to RNA modality. Cellcano is a SOTA lightweight model dedicated to ATAC modality. d KIDA prediction results on 6 query batches (slices). Source data are provided in Supplementary Data 1.
As shown in Fig. 2(b), the inference time of the compared methods are all within acceptable limits. KIDA’s speed is completely acceptable. For three different modalities (BMMC: RNA, PBMC: ATAC, DLPFC: ST), we visualize the atlas respectively. We use uniform manifold approximation and projection (UMAP) to visualize the cell embeddings of BMMC and PBMC. As shown in Fig. 2(c), the cell embeddings of KIDA not only remove batch effects but also preserve biological heterogeneity. For DLPFC, the prediction results of KIDA on 6 query slices are shown in Fig. 2(d). KIDA predictions are nearly close to the ground truth.
We achieve novel cell recognition using the same settings as scBERT47: for a cell input, when the recognition probability of all known categories is less than 70%, the cell is identified as a novel cell, i.e., assigned the label “unknown”. On Pan-cancer dataset, we take each cell type as novel cell type in turn, KIDA is trained on the reference dataset without current novel cell type, tested on the query dataset with current novel cell type. We record the F1 score of KIDA for identifying different novel cell types. As shown in Supplementary Fig. S1, KIDA is able to successfully identify current novel cells with “unknown” labels on query dataset.
Annotation consistency
We compare KIDA and baselines’ annotation results on Ma-2020 and Muto-2021. These datasets are chosen additionally because they provide batch labels for both their RNA and ATAC modalities. Thus, for any given modality, we consider the first two batches as the reference dataset and the remaining batches as the query dataset, totaling four datasets for annotation. Since Ma-2020 and Muto-2021 serve as datasets for evaluating integration experiments, comparing annotation performance between RNA and ATAC modalities allows us further investigation into the consistency of annotation methods in modal-nexus scenarios. Supplementary Table B.2 illustrates that TOSICA, specialized for the RNA modality, performs better on the RNA datasets, while Cellcano, specialized for the ATAC modality, performs better on the ATAC datasets. Cellular large models are generally effective on the RNA modality. In contrast, KIDA achieve similar accuracy and F1 score on both RNA and ATAC modalities.
Case study
In real-world applications, cellular data may come from different diseases (biological states). We test the cross-disease cell type annotation of KIDA on Pan-cancer dataset. The different batches are not only across diseases, they are also across tissues. Therefore, this task increases the difficulty of cell type annotation. We use UMAP to visualize all ‘cls tokens’. As shown in Fig. 3(a) to (c), compared with the PCA-based atlas, KIDA’s atlas completely presents biological heterogeneity (cell clusters are not affected by disease-related batch effects). Then, we filtered out all ‘cDC’ cell embeddings and obtained a graph through PAGA58, as shown in Fig. 3(e). The graph reveals two potential origins of cDC3_LAMP3, namely cDC1_CLEC9A and cDC2_CXCL9 (Fig. 3(e) blue circle), which is consistent with the situation in the measured trajectory (Fig. 3(d) black arrows)22. Through interpretability analysis (details are provided in section 4.3), we can discover the cDC3_LAMP3-specific pathway ‘TOLL RECEPTOR’, which is differentially expressed between inflammation-related cDCs (cDC2_FCN1, cDC2_IL1B) and mature cDC subset (cDC3_LAMP3), as shown in Fig. 3(f). This result is consistent with previous study34. We further use the subset of cell type cDC3_LAMP3 and perform differential analysis based on the disease. We can obtain the disease-specific pathways differential expressions in target cell type. As shown in Fig. 3(g), ‘MEMBRANE TRAFFICKING’ is differentially expressed between OV-FTC (ovarian or fallopian tube carcinoma) and other diseases.
a PCA embeddings labeled by disease. b PCA embeddings labeled by cell type. c KIDA embeddings labeled by cell type. d Measured trajectory, black arrows indicate the direction of cell differentiation. e KIDA's trajectory, blue circle represents connections between target cells. f Cell-specific pathway differential expressions. g Disease-specific pathway differential expressions. Source data are provided in Supplementary Data 1.
Integration
Since the performance of cellular data annotation is closely related to the cell representations in the latent space. Thus, we use integration experiments (batch integration and multimodal integration) to analyze the cell representations of KIDA. We select two datasets, BMMC and PBMC, corresponding to RNA and ATAC modalities, respectively. We compare KIDA with baselines on batch integration performance on these two datasets. The integration results are shown in Fig. 4(a) and (b). KI and KIDA both achieve the best batch integration performance on these independent modalities. These results demonstrate knowledge-based inductive bias can learn the functional patterns and their interactions across batches. Heterogeneous modalities have larger gaps than batches. Therefore, we examine KIDA with the more challenging multimodal integration. KIDA supports multimodal cellular data annotation, naturally achieving multimodal integration by all sample embeddings in the joint space. Multimodal integration comparison is shown in Fig. 4(c). For KIDA integration, cell type labels can guide better alignment of the two modalities.
a Batch integration comparison on BMMC dataset, the metrics are NMI and ARI. b Batch integration comparison on PBMC dataset, the metrics are NMI and ARI. c Multimodal integration comparison on multiple datasets. Source data are provided in Supplementary Data 1.
Most of the existing methods integrate the entire latent space unsupervisedly, so large-scale dataset is a challenge. KIDA only needs to merge cell embeddings to achieve integration. We use KIDA to integrate Human 15 organs (large-scale). The UMAP visualization results of glue, covel and KIDA are shown in Fig. 5. In Fig. 5, we have selected two cell types, Hepatoblasts and Cardiomyocytes, with blue and black dashed circles, respectively. For Hepatoblasts, KIDA aligned the RNA modality and ATAC modality, and the two modalities corresponding to glue and covel are still separated, see the first row of Fig. 5. For Cardiomyocytes, compared with glue and covel, the cell clusters corresponding to KIDA are better separated from other cell clusters, see the second row of Fig. 5. The results show that KIDA’s cell embedding will not be interfered by cross-batch or even cross-modality.
Interpretability
KIDA can output cell type-specific gene co-expression networks and key genes (details are provided in section 4.3). On BMMC dataset, KIDA’s key genes indeed exhibit concentrated expression in target cell clusters. Compared with CellMarker59, these genes are marker genes for target cell type. In BMMC dataset, we obtained the top 10 important patterns for CD8+ T cells and extracted the top 2 important gene embeddings from each pattern. Finally, we obtained 20 gene embeddings specific to CD8+ T cells. By computing the similarity of these 20 gene embeddings, we constructed a graph, which was used as the CD8+ T cell-specific gene co-expression network, as shown in Supplementary Fig. S2(a). Based on the gene co-expression network, we computed five metrics (degree, betweenness, eigenvector centrality, pagerank, closeness) for each gene and integrated these five metrics using Q statistics to identify key genes specific to CD8+ T cells: ‘CD8A’, ‘CD8B’, ‘GZMK’, ‘CCL5’. As shown in Supplementary Fig. S2(c) to (f), these genes indeed exhibit concentrated expression in CD8+ T cell cluster (Supplementary Fig. S2(b) pink circle). We can also obtain the embeddings of all genes based on the top 10 important patterns specific to cell types (details are provided in section 4.3). Then, similar to the above method, we use these embeddings to construct similarity graph and obtain important genes. As shown in Supplementary Fig. S3, we found cell type-specific marker genes in NK cells and B1 B cells respectively. These genes are verified in CellMarker.
For integration experiments, we validate the biological interpretability of key genes. As shown in Supplementary Fig. S4, we select three cell types (oligodendrocytes, astrocytes, and neural progenitors) from the integration results of Human 15 organs (UMAP visualization of the latent space), represent by black, orange, and green circles, respectively. We identify four key genes for each cell type and visualize their expressions in the integration results. These genes exhibit concentrate expressions in their corresponding cell types. This demonstrates that the cell representations output by KIDA are biologically interpretable, enabling meaningful predictions in cellular data annotation tasks.
Ablation study
We conduct ablation study on Pan-cancer to test KIDA separately. For inductive bias, we employ different knowledge databases for KIDA, where ‘Random’ represents no knowledge. For domain adaptation, we adjust KIDA’s two components by setting α and β: distillation and adaptive clustering. As shown in Supplementary Fig. S5(a) and (b), different knowledge is robust, while ‘Random’ performs poorly, demonstrating the advantage of our inductive bias for both annotation and clustering. Moreover, prediction performance improves with an increase in the number of patterns. In Supplementary Fig. S5(c), distillation is more important than adaptive clustering, and our domain adaptation is advantageous for the model when the number of anchors is sufficient: KIDA surpasses KI. Additionally, in Supplementary Fig. S5(d), Adaptive clustering benefits the grouping of cell representations. Overall, knowledge-based inductive bias is an advantageous approach, and both distillation and adaptive clustering can improve the performance of the model.
Discussion
We introduce KIDA to enhance cellular annotation across batches. Experiments demonstrate that KIDA outperforms popular methods and accommodates heterogeneous modalities. We validate the practicality of KIDA using a pan-cancer case, achieving accurate annotation in challenging cross-disease scenarios, with interpretable results consistent with previous biological studies. Additionally, integration experiments reveal that KIDA’s cell representations remain unaffected by cross-batch or cross-modality scenarios. Furthermore, analysis based on marker genes confirms the biological interpretability of KIDA for heterogeneous modalities. Finally, in the ablation study, we verify that knowledge-based inductive bias enables the self-attention to focus on biologically relevant interactions rather than gene-level expressions susceptible to batch effects, while domain adaptation narrows the gap between model predictions and query datasets, thereby improving model performance. KIDA supports multimodal annotation, but a limitation is that the reference dataset and the query dataset come from the same modality (RNA, ATAC, or ST). In the future, we will explore more generalizable approach to extend KIDA for cross-modal annotation (the reference dataset and the query dataset belong to different modalities).
Methods
Knowledge-based inductive bias
In order for KIDA to support multimodal cellular data as input, a unified feature space is necessary. We use Seurat to convert the feature space of heterogeneous modalities into common genes. This step corresponds to Fig. 1(a). For scRNA-seq, Seurat selects highly variable genes and normalizes the raw count values. For scATAC-seq, we use Seurat to convert the peak values into gene scores. For spatial transcriptomics, unlike scRNA-seq, each sample corresponds to a spot, and the task is to annotate the tissue type of each spot.
For cell or spot i, it contains n genes. The gene expressions of this cell is \({x}_{i}\in {{\mathbb{R}}}^{n}\). We use a set of linear transformations \(\{{W}_{j}\in {{\mathbb{R}}}^{n\times h}| j=1,\ldots ,m\}\) to convert the cell into m pattern tokens, h is dimension of latent space. Each linear matrix Wj represents a biologically relevant functional pattern (pathway). In order to achieve it, we create a list consisting of all gene names in reference dataset, and obtain m pathway names corresponding to this gene list through GSEA (Gene Set Enrichment Analysis) combined with pathway databases (‘gmt’ file on the web or locally). This step corresponds to Fig. 1(b). Then, we add mask to each linear matrix according to the relationship between genes and pathways. In Fig. 1(c), assuming that gene 1 is not in pathway 1, the first row (gene 1) of linear matrix \({W}_{1}\in {{\mathbb{R}}}^{n\times h}\) (pathway 1) is all masked to value 0. On the contrary, assuming that gene n is in pathway 1, the nth row of linear matrix \({W}_{1}\in {{\mathbb{R}}}^{n\times h}\) will not have a mask added. The linear transformations after initialization in this way convert genes into m pattern tokens \(\{{v}_{j}\in {{\mathbb{R}}}^{h}| j=1,\ldots ,m\}\). This step corresponds to Fig. 1(d). Each token represents a gene set with specific pathway. For token generation, we can use TOSICA’s initialization method if memory is limited.
Domain adaptation
We use self-attention to model the pattern-pattern interactions. This step corresponds to Fig. 1(e). We add a ‘cls token’ \(c\in {{\mathbb{R}}}^{h}\) to represent the global information of current cell. We concat all tokens as \({I}_{0}=concat({v}_{1},\ldots ,{v}_{m},c)\in {{\mathbb{R}}}^{(m+1)\times h}\). The query, key and value are Q = I0WQ, K = I0WK and V = I0WV, respectively. \({W}^{Q},{W}^{K},{W}^{V}\in {{\mathbb{R}}}^{h\times h}\) are learnable parameters. The self-attention output \({I}_{1}\in {{\mathbb{R}}}^{(m+1)\times h}\) is \({I}_{1}=softmax((Q{K}^{\top })/\sqrt{h})V\). The last row tensor of I1 is updated ‘cls token’ \({c}_{r}\in {{\mathbb{R}}}^{h}\). We use cr as the embedding of the cell. With the cr, we use UMAP60 to visualize all cells’ embedding. For the type annotation, we use MLP61 as the classifier. We map the embedding cr of cell i to the category, and record the classification logits output by the classifier as \({y}_{i}^{teacher,r}\in {{\mathbb{R}}}^{A}\), where A represents the total number of labels. We train the model (linear transformations, self-attention and MLP) on the reference dataset (N cells). The one-hot label of cell i is \({g}_{i}^{r}\). The model is trained with cross entropy loss, which is:
We perform the second round of supervised training on the query dataset (M cells). This step corresponds to Fig. 1(f). We use the model trained on the reference dataset as the Teacher and predict on the query dataset. The predicted probability is denoted as qia for cell i being in cell type a. The entropy \({E}_{i}\in {\mathbb{R}}\) for cell i is \({E}_{i}=-{\sum }_{a = 1}^{A}{q}_{ia}\log ({q}_{ia})\). When a cell label is more confidently assigned, its entropy over the predicted probabilities is lower, and the prediction is in general more accurate. For each cell type, we selected 40% cells with the lowest entropies as anchors, we use all anchors to form the new training set. Therefore, the number of anchors accounts for 40% of the total number of cells in the query dataset.
Since some anchors will be mistakenly predicted, we apply the knowledge distillation to deal with the issue. We use the logits output by Teacher on anchor i as pseudo label \({y}_{i}^{teacher,q}\in {{\mathbb{R}}}^{A}\). Then, we initialize a Student with the same architecture as the Teacher. We use the logits output by Student on anchor i as \({y}_{i}^{student,q}\in {{\mathbb{R}}}^{A}\). Let KL(P, Q) be the KL divergence of two probability distributions P and Q, T = 3 be the temperature parameter (to make the label “softer”), and the knowledge distillation loss is:
We employ adaptive clustering loss for Students to minimize the correlation among the various clusters. We represent correlation by the similarity between clusters. Define a similarity matrix \(S\in {{\mathbb{R}}}^{(40 \% M)\times (40 \% M)}\), element si,j = exp( − ∣∣cr,i − cr,j∣∣2) represents the similarity between two cells, cr,i and cr,j represent the embedding (‘cls token’) of cell i and cell j respectively. Define Qa = [q1a, …, q(40%M)a] (qia: probability of cell i being in type a). Minimizing \({Q}_{a}S{Q}_{b}^{\top }\) means that we want to make Qa and Qb orthogonal. Adaptive clustering loss is:
The sum of Teacher’s loss is \({L}^{teacher}={L}_{ce}^{teacher}\). The sum of Student’s loss is \({L}^{student}=\alpha {L}_{kd}^{student}+\beta {L}_{ada}^{student}\). α and β are two hyper-parameters for balancing the two losses.
Interpretability of KIDA
The inductive bias in KIDA has natural interpretability. During model initialization, a linear matrix \({W}_{j}\in {{\mathbb{R}}}^{n\times h}\) corresponds to a pathway name. We sum it up to output a vector \({P}_{j}\in {{\mathbb{R}}}^{n}\). Pj reflects the set of genes corresponding to functional pattern j. For the ‘cls token’ of cell i, we extract the attention values of m patterns about ‘cls token’ in the self-attention layer. Through the difference analysis function in scanpy, we can treat m attention values as scanpy input of cell i, and obtain the cell type-specific top-10 important patterns. For pattern j, we select the top-2 genes in Pj based on its values. Assuming that the indices of these two genes are x and y, we retain \({W}_{j}[x]\in {{\mathbb{R}}}^{h}\) and \({W}_{j}[y]\in {{\mathbb{R}}}^{h}\) as gene embeddings. Finally, we obtained cell type-specific embeddings of 20 genes. Then, we constructed a graph of these important genes based on embedding similarity. We use the graph as a cell type-specific gene co-expression network. Based on the gene co-expression network, we calculate 5 indicators for each gene (degree, betweenness, eigenvalue, pagerank, proximity), and use Q statistics to integrate the 5 indicators to further narrow the scope and obtain cell type-specific key genes62.
To consider all genes, we provide an alternative interpretable method. First, through the difference analysis function in scanpy, we can treat m attention values as scanpy input of cell i, and obtain the cell type-specific top-10 important patterns. Pattern j has a linear matrix \({W}_{j}\in {{\mathbb{R}}}^{n\times h}\). We sum the 10 matrices into one matrix \({W}_{sum}\in {{\mathbb{R}}}^{n\times h}\). The row x of the matrix serves as the embedding of gene x. Then, we constructed a graph based on the gene embedding similarity, and calculate 5 indicators (degree, betweenness, eigenvalue, pagerank, proximity) for each gene to obtain key genes.
Statistics and reproducibility
In KIDA, we set m = 300, h = 128, and the self-attention network has 4 layers and 8 heads. Let α = 0.8, β = 0.2. If α = β = 0, KIDA only uses Teacher, which we call KI. The knowledge database required by KIDA when generating functional patterns comes from63 and64. Enrichment analysis tool is GSEApy65, preprocessing and differential analysis tool is scanpy66. KIDA is trained using NVIDIA GeForce RTX A6000 with 48 GB memory. Adam67 optimizer with 0.001 learning rate is used to update model parameters. The batch size is set to 16. The epochs for Teacher and Student are 6 and 3, respectively.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Related work
Annotation
Most cellular data annotation methods are designed for the single-cell transcriptome modality (scRNA-seq), where the feature space corresponds to genes. These methods utilize gene expression counts to identify cell types. Seurat is currently the most popular cellular annotation framework, but the analysis results depend on additional preprocessing steps such as feature selection7. Essentially, cellular annotation is a repetitive task, transferring cell types from the reference dataset to the query dataset aligns with supervised deep learning. Actinn and TOSICA respectively employ fully connected networks and Transformers68 to learn features from gene expression counts and then predict cell types on the query dataset22,43. Corresponding to the scRNA-seq modality, Cellcano first converts the feature space of scATAC-seq from open chromatin peaks to gene scores using ArchR69, and then trains a fully connected network based on knowledge distillation to achieve cell-type annotation for the scATAC-seq modality44. For spatial transcriptomics, each sample represents a spot in space, and the features are genes. Similar to cell type prediction, we can use the deep learning methods mentioned above to predict the tissue types of each spot70. Recently, large-scale language models have gained popularity, with models like Geneformer46, scBERT47, CellLM48, LangCell49, and scGPT50. Note that for unknown diseases or biological tissues, large models still need fine-tuning on reference datasets to achieve acceptable predictions42,71,72. We find that existing methods follow the trend of AI model development but lack consideration for the inductive bias and batch effects.
Integration
Integration is divided into batch integration and multimodal integration. For batch integration, the goal is to eliminate batch effects in the reference dataset or query dataset. Seurat can integrate multiple batches using a mutual nearest neighbor approach7. Harmony projects the raw data into a unified space and groups them by cell type52, requiring additional batch labels. For Seurat and Harmony, batch integration and cell type annotation are separate, making it difficult to track the contributions of inputs, leading to a loss of interpretability. Recently, TOSICA uses attention values as cell representations and experimentally validates its batch integration capabilities22. scGPT introduces additional batch labels and fine-tunes according to the rules of classification50. Our KIDA does not require batch labels and simultaneously satisfies batch integration and cellular annotation. As for multimodal integration, the goal is to bridge the gap between modalities. Multimodal integration is similar to batch integration, with the additional requirement of unifying the multimodal joint space7,51,52,53,54,55,56,73,74. Our KIDA supports multimodal cellular data annotation, naturally achieving multimodal integration by all sample embeddings in the joint space.
Data availability
All datasets used in this study are already published and were obtained from public data repositories. The BMMC dataset is available at NCBI Gene Expression Omnibus (GEO) with the accession number GSE19412231 [https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE194122]. The PBMC dataset is available at NCBI Gene Expression Omnibus (GEO) with the accession number GSE12978532 [https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE129785]. The Pan-cancer dataset is available from GSE154763 and GSE15672833,34 [https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE154763,https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE156728]. The DLPFC dataset is available in the OpenNeuro database under accession code ds00207635 [https://openneuro.org/datasets/ds002076/versions/1.0.1]. The 10x-Multiome dataset is available at https://support.10xgenomics.com/single-cell-multiome-atac-gex/datasets/1.0.0/pbmc_granulocyte_sorted_10k. The Chen-2019 dataset is available at NCBI Gene Expression Omnibus (GEO) with the accession number GSE12607437 [https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE126074]. The Ma-2020 dataset is available at GEO with the accession number GSE14020338 [https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE140203]. The Muto-2021 dataset is available at GEO with the accession number GSE15130239 [https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE151302]. The Human 15 organs dataset is collected from GEO with the accession number GSE15679340 [https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE156793] and GEO with the accession number GSE14968341 [https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE149683]. Source data are provided with this paper.
Code availability
The code of this study is available at https://github.com/shapsider/cellannotationand https://doi.org/10.5281/zenodo.1397029475.
References
Heumos, L. et al. Best practices for single-cell analysis across modalities. Nat. Rev. Genet. 24, 550–572 (2023).
Chen, G. et al. Vaerhnn: Voting-averaged ensemble regression and hybrid neural network to investigate potent leads against colorectal cancer. Knowl.-Based Syst. 257, 109925 (2022).
Chen, S., Li, Q., Zhao, J., Bin, Y. & Zheng, C. Neuropred-clq: incorporating deep temporal convolutional networks and multi-head attention mechanism to predict neuropeptides. Brief. Bioinform. 23, 319 (2022).
Lv, Q., Chen, G., Yang, Z., Zhong, W. & Chen, C.Y.-C. Meta-molnet: A cross-domain benchmark for few examples drug discovery. IEEE Trans. Neural Netw. Learn. Syst. (2024).
Cao, K., Bai, X., Hong, Y. & Wan, L. Unsupervised topological alignment for single-cell multi-omics integration. Bioinformatics 36, i48-i56 (2020).
Yu, X., Xu, X., Zhang, J. & Li, X. Batch alignment of single-cell transcriptomics data using deep metric learning. Nat. Commun. 14, 960 (2023).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 (2019).
Xu, J. et al. Graph embedding and gaussian mixture variational autoencoder network for end-to-end analysis of single-cell rna sequencing data. Cell Rep. Methods 3, 100382 (2023).
Lin, X., Tian, T., Wei, Z. & Hakonarson, H. Clustering of single-cell multi-omics data with a multimodal deep learning method. Nat. Commun. 13, 7705 (2022).
Clarke, Z. A. et al. Tutorial: guidelines for annotating single-cell transcriptomic maps using automated and manual methods. Nat. Protoc. 16, 2749–2764 (2021).
Wang, J. et al. Generalizing to unseen domains: a survey on domain generalization. IEEE Trans. Knowl. Data Eng. 35, 8052–8072 (2022).
Schölkopf, B. et al. Toward causal representation learning. Proc. IEEE 109, 612–634 (2021).
Nguyen, T., Tong, A., Madan, K., Bengio, Y. & Liu, D. Causal discovery in gene regulatory networks with gflownet: towards scalability in large systems. In NeurIPS 2023 Generative AI and Biology (GenBio) Workshop (2023).
Atanackovic, L. et al. Dyngfn: Towards Bayesian inference of gene regulatory networks with gflownets. Adv. Neural Inf. Process. Syst. 36, 74410–74428 (2023).
Satorras, V.G., Hoogeboom, E. & Welling, M. E (n) equivariant graph neural networks. In Proc. International Conference on Machine Learning, 9323–9332 (PMLR, 2021).
Dong, T., Yang, Z., Zhou, J. & Chen, C. Y.-C. Equivariant flexible modeling of the protein–ligand binding pose with geometric deep learning. J. Chem. Theory Comput. 19, 8446–8459 (2023).
Goyal, A. & Bengio, Y. Inductive biases for deep learning of higher-level cognition. Proc. R. Soc. A 478, 20210068 (2022).
Yang, Z. et al. Interaction-based inductive bias in graph neural networks: enhancing protein-ligand binding affinity predictions from 3d structures. IEEE Trans. Pattern Anal. Mach. Intell. (2024).
Tang, Z., Chen, G., Yang, H., Zhong, W. & Chen, C.Y.-C. Dsil-ddi: A domain-invariant substructure interaction learning for generalizable drug–drug interaction prediction. IEEE Trans. Neural Netw. Learn. Syst. 35, 10552–10560 (2023).
Chen, S., Tang, Z., You, L. & Chen, C. Y.-C. A knowledge distillation-guided equivariant graph neural network for improving protein interaction site prediction performance. Knowl. Based Syst. 300, 112209 (2024).
Lv, Q., Chen, G., Yang, Z., Zhong, W. & Chen, C.Y.-C. Meta learning with graph attention networks for low-data drug discovery. IEEE Trans. Neural Netw. Learn. Syst. 35, 11218–11230 (2023).
Chen, J. et al. Transformer for one stop interpretable cell type annotation. Nat. Commun. 14, 223 (2023).
Liu, T., Wang, Y., Ying, R. & Zhao, H. Muse-gnn: Learning unified gene representation from multimodal biological graph data. Adv. Neural Inf. Process. Syst. 36, 24661–24677 (2023).
Dai, C. et al. scimc: a platform for benchmarking comparison and visualization analysis of scrna-seq data imputation methods. Nucleic Acids Res. 50, 4877–4899 (2022).
Huang, X. et al. scgrn: a comprehensive single-cell gene regulatory network platform of human and mouse. Nucleic Acids Res. 52, 293–303 (2024).
Bereket, M. & Karaletsos, T. Modelling cellular perturbations with the sparse additive mechanism shift variational autoencoder. Adv. Neural Inf. Process. Syst. 36, 1–12 (2023).
Chen, C. et al. This looks like that: deep learning for interpretable image recognition. Adv. Neural Inf. Process. Syst. 32, 1–12 (2019).
Tang, Z., Yang, H. & Chen, C.Y.-C. Weakly supervised posture mining for fine-grained classification. In Proceedings of the 2023 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 23735–23744 (2023).
Stevens, S. et al. Bioclip: A vision foundation model for the tree of life. In Proceedings of the 2024 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 19412–19424 (2024).
Yuan, Q. & Duren, Z. Integration of single-cell multi-omics data by regression analysis on unpaired observations. Genome Biol. 23, 160 (2022).
Luecken, M. D. et al. A sandbox for prediction and integration of dna, rna, and proteins in single cells. In Thirty-fifth Conference on Neural Information Processing Systems Datasets and Benchmarks Track (Round 2) (2021).
Satpathy, A. T. et al. Massively parallel single-cell chromatin landscapes of human immune cell development and intratumoral t cell exhaustion. Nat. Biotechnol. 37, 925–936 (2019).
Zheng, L. et al. Pan-cancer single-cell landscape of tumor-infiltrating t cells. Science 374, 6474 (2021).
Cheng, S. et al. A pan-cancer single-cell transcriptional atlas of tumor infiltrating myeloid cells. Cell 184, 792–809 (2021).
Maynard, K. R. et al. Transcriptome-scale spatial gene expression in the human dorsolateral prefrontal cortex. Nat. Neurosci. 24, 425–436 (2021).
PBMC from a Healthy Donor, Single Cell Multiome ATAC Gene Expression Demonstration Data by Cell Ranger ARC 1.0.0. 10X Genomics https://support.10xgenomics.com/single-cell-multiome-atac-gex/datasets/1.0.0/pbmc_granulocyte_sorted_10k (2020)
Chen, S., Lake, B. B. & Zhang, K. High-throughput sequencing of the transcriptome and chromatin accessibility in the same cell. Nat. Biotechnol. 37, 1452–1457 (2019).
Ma, S. et al. Chromatin potential identified by shared single-cell profiling of rna and chromatin. Cell 183, 1103–1116 (2020).
Muto, Y. et al. Single cell transcriptional and chromatin accessibility profiling redefine cellular heterogeneity in the adult human kidney. Nat. Commun. 12, 2190 (2021).
Cao, J. et al. A human cell atlas of fetal gene expression. Science 370, 7721 (2020).
Domcke, S. et al. A human cell atlas of fetal chromatin accessibility. Science 370, 7612 (2020).
Domínguez Conde, C. et al. Cross-tissue immune cell analysis reveals tissue-specific features in humans. Science 376, 5197 (2022).
Ma, F. & Pellegrini, M. Actinn: automated identification of cell types in single cell rna sequencing. Bioinformatics 36, 533–538 (2020).
Ma, W., Lu, J. & Wu, H. Cellcano: supervised cell type identification for single cell atac-seq data. Nat. Commun. 14, 1864 (2023).
Zhang, Y. et al. Metatime integrates single-cell gene expression to characterize the meta-components of the tumor immune microenvironment. Nat. Commun. 14, 2634 (2023).
Theodoris, C. V. et al. Transfer learning enables predictions in network biology. Nature 618, 616–624 (2023).
Yang, F. et al. scbert as a large-scale pretrained deep language model for cell type annotation of single-cell rna-seq data. Nat. Mach. Intell. 4, 852–866 (2022).
Zhao, S., Zhang, J. & Nie, Z. Large-scale cell representation learning via divide-and-conquer contrastive learning. Preprint at https://arxiv.org/abs/2306.04371 (2023).
Zhao, S., Zhang, J., Luo, Y., Wu, Y. & Nie, Z. Langcell: Language-cell pre-training for cell identity understanding. Preprint at https://arxiv.org/abs/2405.06708 (2024).
Cui, H. et al. scgpt: toward building a foundation model for single-cell multi-omics using generative ai. Nat. Methods 21, 1470–1480 (2024).
Cao, Z.-J. & Gao, G. Multi-omics single-cell data integration and regulatory inference with graph-linked embedding. Nat. Biotechnol. 40, 1458–1466 (2022).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with harmony. Nat. Methods 16, 1289–1296 (2019).
Welch, J. D. et al. Single-cell multi-omic integration compares and contrasts features of brain cell identity. Cell 177, 1873–1887 (2019).
Dou, J. et al. Bi-order multimodal integration of single-cell data. Genome Biol. 23, 1–25 (2022).
Gao, C. et al. Iterative single-cell multi-omic integration using online learning. Nat. Biotechnol. 39, 1000–1007 (2021).
Tang, Z., Huang, J., Chen, G. & Chen, C. Y.-C. Comprehensive view embedding learning for single-cell multimodal integration. Proc. AAAI Conf. Artif. Intell. 38, 15292–15300 (2024).
Cao, Y. et al. scbutterfly: a versatile single-cell cross-modality translation method via dual-aligned variational autoencoders. Nat. Commun. 15, 2973 (2024).
Wolf, F. A. et al. Paga: graph abstraction reconciles clustering with trajectory inference through a topology preserving map of single cells. Genome Biol. 20, 1–9 (2019).
Zhang, X. et al. Cellmarker: a manually curated resource of cell markers in human and mouse. Nucleic Acids Res. 47, 721–728 (2019).
McInnes, L., Healy, J. & Melville, J. Umap: Uniform manifold approximation and projection for dimension reduction. Preprint at https://arxiv.org/abs/1802.03426 (2018).
Tolstikhin, I. O. et al. Mlp-mixer: An all-mlp architecture for vision. Adv. Neural Inf. Process. Syst. 34, 24261–24272 (2021).
Zhou, S. et al. Single-cell rna-seq dissects the intratumoral heterogeneity of triple-negative breast cancer based on gene regulatory networks. Mol. Ther. Nucleic Acids 23, 682–690 (2021).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. 102, 15545–15550 (2005).
Mootha, V. K. et al. Pgc-1α-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat. Genet. 34, 267–273 (2003).
Fang, Z., Liu, X. & Peltz, G. Gseapy: a comprehensive package for performing gene set enrichment analysis in python. Bioinformatics 39, 757 (2023).
Wolf, F. A., Angerer, P. & Theis, F. J. Scanpy: large-scale single-cell gene expression data analysis. Genome Biol. 19, 1–5 (2018).
Kingma, D. P. & Ba, J. Adam: a method for stochastic optimization. Preprint at https://arxiv.org/abs/1412.6980 (2014).
Vaswani, A. et al. Attention is all you need. Adv. Neural Inf. Process. Syst. 30, 1–11 (2017).
Granja, J. M. et al. Archr is a scalable software package for integrative single-cell chromatin accessibility analysis. Nat. Genet. 53, 403–411 (2021).
Tang, X. et al. Explainable multi-task learning for multi-modality biological data analysis. Nat. Commun. 14, 2546 (2023).
Liu, T., Li, K., Wang, Y., Li, H. & Zhao, H. Evaluating the utilities of large language models in single-cell data analysis. Preprint at bioRxiv https://www.biorxiv.org/content/10.1101/2023.09.08.555192v1 (2023).
Wang, S. et al. scfed: federated learning for cell type classification with scrna-seq. Brief. Bioinform. 25, 507 (2024).
Cao, K., Hong, Y. & Wan, L. Manifold alignment for heterogeneous single-cell multi-omics data integration using pamona. Bioinformatics 38, 211–219 (2022).
Tang, Z. et al. Modal-nexus auto-encoder for multi-modality cellular data integration and imputation. Nat. Commun. 15, 9021 (2024).
Tang, Z. et al. Source code for “Knowledge-Based Inductive Bias and Domain Adaptation: Enhancing Cell Type Annotation Across Batches”. Zenodo, https://doi.org/10.5281/zenodo.13970294 (2024).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (Grant No. 62176272, to C.Y.-C.C.), Research and Development Program of Guangzhou Science and Technology Bureau (No. 2023B01J1016, to C.Y.-C.C.), and Key-Area Research and Development Program of Guangdong Province (No. 2020B1111100001, to C.Y.-C.C.).
Author information
Authors and Affiliations
Contributions
Z.T. designed research. Z.T., G.C., S.C., and H.H. worked together to complete the experiment and analyze the data. L.Y. and C.Y.-C.C. contributed to analytic tools. Z.T. and G.C. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Yu-An Huang, and the other, anonymous, reviewers for their contribution to the peer review of this work. Primary Handling Editors: Laura Rodriguez Perez and Aylin Bircan.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Tang, Z., Chen, G., Chen, S. et al. Knowledge-based inductive bias and domain adaptation for cell type annotation. Commun Biol 7, 1440 (2024). https://doi.org/10.1038/s42003-024-07171-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-024-07171-9