Abstract
The KCNE family (KCNE1-5) is a group of single transmembrane auxiliary subunits for the voltage-gated K+ channel KCNQ1. The KCNQ1-KCNE complexes are crucial for numerous physiological processes including ventricular repolarization and K+ recycling in epithelial cells. We identified a new member of the KCNE family, “KCNE6”, from zebrafish. We found that KCNE6 is expressed in the zebrafish heart and is involved in cardiac excitability. When co-expressed with KCNQ1, KCNE6 produces a slowly activating current like the slow delayed-rectifier K+ current (IKs) induced by KCNE1, despite the fact that the KCNE6 amino acid sequence has the highest similarity to that of KCNE3, which forms a constitutively open channel with KCNQ1. The kcne6 nucleotide sequences exist throughout vertebrates, including humans, although only the KCNE6 proteins of lower vertebrates, up to marsupials, can modulate KCNQ1, and it has become a pseudogene in eutherians. Our findings will facilitate a better understanding of how the KCNE family has evolved to modulate KCNQ1.
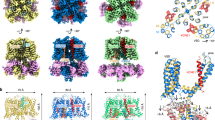
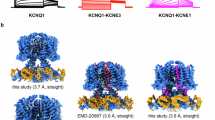
Similar content being viewed by others
Introduction
The functional properties of ion channels, including expression, localization, and gating, are modulated by auxiliary subunits. Auxiliary subunits are proteins that directly interact to modulate ion channels, diversifying physiological and biophysical functions. Altered expression or dysfunctions of auxiliary subunits caused by mutations have been linked to various channelopathies such as cardiac arrhythmias and epilepsy1,2.
The KCNE family is a group of single-transmembrane proteins serving as auxiliary subunits for the voltage-gated K+ (Kv) channel KCNQ1 (Kv7.1). The KCNQ1-KCNE complexes are expressed in both excitable and non-excitable cells and play a crucial role in numerous physiological processes including ventricular repolarization in the heart, maintenance of high K+ levels in the endolymph of the inner ear, and K+ recycling in epithelial cells in the trachea, stomach, and intestine1,2. The first isoform of the KCNE family (KCNE1/MinK) was cloned in 19883. It was initially identified as a putative pore-forming minimal K+ channel (MinK)3 and was later confirmed to co-assemble with KCNQ1 as an auxiliary subunit4,5. Five isoforms (KCNE1-5) were later characterized by 20026,7,8,9,10,11. The KCNE1 and KCNE3 isoforms have been more extensively studied than the other isoforms in terms of their modulatory effects on KCNQ1. KCNE1 upregulates the current amplitude, slows the activation/deactivation kinetics, and shifts the voltage dependence of KCNQ1 in a positive direction to produce the slow delayed-rectifier K+ (IKs) current. The KCNQ1-KCNE1 channel is involved in ventricular repolarization in the heart and in maintenance of high K+ levels in the endolymph of the inner ear1. In addition to KCNQ1, KCNE1 can modulate the channel properties of Kv2.112, Kv3.112, Kv3.413, Kv4.314, and Kv11.1 (also known as hERG)15 and possibly of the Ca2+-gated Cl- channel (TMEM16A)16,17 in heterologous expression systems. KCNE3 shifts the voltage dependence of KCNQ1 in the far-negative direction to produce a constitutively active current at physiological voltages9. The KCNQ1-KCNE3 channel is involved in K+ recycling in epithelial cells in the trachea, stomach, and intestine and is coupled with some ion transporters and ion channels to facilitate Cl- secretion1,2. In addition to KCNQ1, KCNE3 can modulate the channel properties of Kv2.112, Kv3.413, Kv4.218, and Kv4.319 in heterologous expression systems.
Since no kcne genes have been identified in invertebrates, it is believed that the first kcne gene arose in lower vertebrates20. The number of isoforms varies among vertebrates. For example, in zebrafish (Danio rerio), only two KCNE isoforms, KCNE1 and KCNE4, have been reported so far21,22. The zebrafish has been widely used as a popular animal model in various biological research fields including genetics, neurobiology, toxicology, and drug discovery. Therefore, understanding the extent of similarity in ion channel functions between zebrafish and humans is important for extrapolating the results obtained through experiments using zebrafish to humans23. Accordingly, we initially started a study to identify other homologous KCNE isoforms in zebrafish, particularly KCNE3, which unexpectedly resulted in the discovery of a previously unknown KCNE-like protein. Compared to the known KCNE1-5 isoforms, the amino acid sequence of this KCNE-like protein has the highest similarity to KCNE3. However, when co-expressed with KCNQ1, this KCNE-like protein shifts the KCNQ1 voltage dependence in a positive direction like KCNE1 rather than KCNE3, which shifts the G-V relationship in the far-negative direction. Promoter analysis and gene knockout (KO) experiments using zebrafish revealed that this KCNE-like protein is expressed in the heart and is involved in cardiac excitability. A homology search showed that the nucleotide sequences coding for this KCNE-like protein are conserved in vertebrates and that its genomic position is different from those for KCNE1 and KCNE3 isoforms. When co-expressed with KCNQ1, these KCNE-like proteins from lower vertebrates, including marsupials (e.g., koalas), can modulate KCNQ1 currents as in the case of zebrafish, while those from eutherians (e.g., dogs, orangutans, and humans) cannot do so, suggesting that these kcne-like genes in eutherians became pseudogenes at a certain point of evolution. Based on the results, we concluded that this KCNE-like protein is a new isoform of the KCNE family and we named it KCNE6.
Results
Identification of a new kcne gene in zebrafish
We initially searched for KCNE homologs in zebrafish using the tBLASTn program24,25 with the amino acid sequences of all KCNE isoforms from various species as queries. When we examined several KCNE3 sequences from different lower vertebrates, a 369-nucleotide sequence, which is located between the nucleotide sequences coding the putative long intergenic non-coding RNA (lincRNA) (ENSDARG00000117370) and those coding the transcription factor Paired Box3 (pax3) on zebrafish chromosome 2, was retrieved as a putative KCNE3-like protein-coding sequence (Fig. 1a). For clarity, we will hereafter refer to this putative KCNE3-like protein as KCNE6 because it was later shown that this protein is different from the five known KCNE isoforms, including KCNE3. When translated into the amino acid sequence and compared to the five KCNE isoforms, zebrafish KCNE6 (DrKCNE6; ‘Dr’ referring to Danio rerio) showed the highest similarity to the KCNE3 isoform (Fig. 1b, c; Supplementary Fig. 1). In humans, KCNE3 is responsible for shifting the conductance-voltage (G-V) relationship of KCNQ1 in the far-negative direction and making KCNQ1 a constitutively open channel2,9.
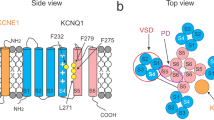
a Genomic region containing the kcne6 ORF sequence in zebrafish. b Sequence alignment around the TM segments of KCNE1, KCNE3, KCNE4, and KCNE6 proteins. c Phylogenetic tree of KCNE1-6 proteins. Sequence alignment was performed and the phylogenic tree was prepared using the EMBL-EBI Job Dispatcher sequence analysis tools framework63. Sequence alignment is shown using ESPript364. Residues corresponding to “the triplet” 51,52 are highlighted with an orange square. For sequence alignment, human KCNE proteins (HsKCNE1, NCBI Accession Number: NP_000210; HsKCNE2, NP_751951.1; HsKCNE3, NP_005463.1; HsKCNE4, NP_542402.4; and HsKCNE5, NP_036414.1), mouse KCNE proteins (MmKCNE1, NP_001349385.1; MmKCNE2, NP_001345301.1; MmKCNE3, NP_001177798, MmKCNE4, NP_067317.1; and MmKCNE5, NP_067462.1), and zebrafish KCNE proteins (DrKCNE1, XP_017213386.1; DrKCNE4, NP_001076366.1; and DrKCNE6) were used. Representative current traces (d–h) and G-V relationships (i) of DrKCNQ1 WT alone and DrKCNE6 WT alone as well as DrKCNQ1 WT co-expressed with DrKCNE1 WT, MmKCNE3 WT, and DrKCNE6 WT. Representative current traces (upper row) and fluorescence traces (lower row) (j–m) and G-V and F-V (n) of DrKCNQ1vcf WT alone as well as DrKCNQ1vcf WT co-expressed with DrKCNE1 WT, MmKCNE3 WT and DrKCNE6 WT. Error bars indicate ± s.e.m. for n = 5 in (i, n).
To investigate whether the DrKCNE6 protein can modulate KCNQ1 gating like KCNE3, we cloned the protein-coding sequence (CDS) of DrKCNQ1 and that of DrKCNE6 from a zebrafish heart cDNA library (Supplementary Fig. 2) and co-expressed their cRNAs in Xenopus oocytes to record currents. Strikingly, it shifted the G-V relationship of KCNQ1 in a positive direction like KCNE1 rather than in the far-negative direction like KCNE3, although the G-V shift induced by DrKCNE6 was milder than that induced by DrKCNE1 (Fig. 1d–g, i). To exclude the possibility that DrKCNQ1 cannot shift its G-V relationship in the far-negative direction by KCNE3, we co-expressed DrKCNQ1 with mammalian KCNE3 from mouse (MmKCNE3; ‘Mm’ referring to Mus musculus) and confirmed that DrKCNQ1 became a constitutively open channel by KCNE3 (Fig. 1f, i). These results suggest that KCNE6 modulates KCNQ1 gating in a manner like KCNE1 rather than KCNE3. KCNE1 was originally identified as a minimal potassium channel subunit (MinK) that, when expressed alone in Xenopus oocytes, produces a significant K+ current by co-assembling with endogenous Xenopus KCNQ13. To investigate whether KCNE6 could similarly increase the activity of endogenous KCNQ1, we expressed DrKCNE6 alone. We found that it generates small but measurable currents, similar to those produced by the KCNQ1-KCNE6 complex, suggesting that KCNE6 enhances the current of endogenous KCNQ1 (Fig. 1h).
We next performed voltage-clamp fluorometry (VCF) analysis to investigate how KCNE6 affects the voltage sensor domain (VSD) movement of KCNQ1. We created a DrKCNQ1 construct for VCF (DrKCNQ1 G187C; this construct hereafter being referred to as ‘DrKCNQ1vcf WT’) with a cysteine mutation at position G187, equivalent to G219 in human KCNQ1 used for the VCF analyses26,27. Then, we labeled it with Alexa Fluor 488 maleimide. DrKCNQ1vcf WT alone showed two components in the fluorescence signals, as previously reported28,29. However, since the first component was dominant, the fluorescence-voltage (F-V) relationship was fitted to a single Boltzmann function and closely overlapped with its G-V relationship (Fig. 1j, n). In contrast, when co-expressed with DrKCNE6, DrKCNE1, and MmKCNE3, DrKCNQ1vcf WT showed a prominent split F-V relationship that fitted to a double Boltzmann function (Fig. 1j–n; Supplementary Table 1). According to previous VCF studies, the prominent split F-V relationship of KCNQ1, which was observed only when co-expressed with KCNE1 and KCNE3, means that KCNE1 and KCNE3 directly affect the VSD movement of KCNQ1 and that the VSD can occupy another position (intermediate) in addition to the two positions (down and up) observed as in the case of KCNQ1 alone28,29,30,31,32,33. Moreover, it was concluded from those VCF studies that the most significant difference between KCNE1 and KCNE3 arises when the VSD is in the intermediate position: KCNE1 restricts the pore opening of KCNQ1, resulting in the KCNQ1-KCNE1 channel occupying the intermediate-closed (IC) state, while KCNE3 promotes the pore opening of KCNQ1, resulting in the KCNQ1-KCNE3 channel occupying the intermediate-open (IO) state29. When co-expressed with DrKCNE6, the G-V relationship of DrKCNQ1vcf WT follows the second fluorescence component of the F-V relationship as in the case of when co-expressed with DrKCNE1 (Fig. 1j–n; Supplementary Table 1), suggesting that the pore opening of KCNQ1 may be inhibited when the VSD is in the intermediate position as in the case of KCNE1. Therefore, results of the VCF analyses also support the idea that KCNE6 modulates the KCNQ1 gating like KCNE1 rather than KCNE3.
Promoter analysis suggested that KCNE6 is expressed in the zebrafish heart
To investigate whether and where the kcne6 gene is expressed in vivo, we performed promoter analysis to identify the target genomic regions that regulate kcne6 gene expression in zebrafish. We created a transgenic (Tg) zebrafish line, Tg(kcne6:eGFP), that harbors the nucleotide sequences containing a 3.5K-bp genomic region upstream of the kcne6 gene open reading frame (ORF) connected to the enhanced green fluorescent protein (eGFP) CDS. In Tg(kcne6:eGFP), eGFP expression became prominent at 48 hours postfertilization (hpf) and was sustained in the adult zebrafish heart (Fig. 2a; Supplementary Fig. 3), suggesting that the kcne6 gene is expressed in both larval and adult zebrafish hearts. Expression of the kcne6 gene in the adult zebrafish heart was further confirmed by RT-PCR. In addition, weaker kcne6 expression was detected in the brain, intestine, testis, and kidney of adult zebrafish (Supplementary Fig. 2). In immunofluorescence staining using the adult Tg(kcne6:eGFP) line, eGFP expression controlled by the kcne6 promoter were broadly observed in atrial and ventricular myocytes but not in the bulbus arteriosus (Fig. 2b–d; Supplementary Fig. 3b). In humans, the KCNQ1-KCNE1 channel is involved in ventricular repolarization in the heart and in maintenance of a high K+ level in the endolymph of the inner ear1. Accordingly, for comparison, we also created a transgenic line, Tg(kcne1:eGFP), that harbors the nucleotide sequences containing a 4K-bp genomic region upstream of the kcne1 gene ORF region connected to the eGFP CDS. In Tg(kcne1:eGFP) at 72 hpf, eGFP expression controlled by the kcne1 promoter was observed in various organs, including the inner ear and lateral line, whereas its expression in the heart was weak or almost invisible at that stage (Fig. 2e). In immunofluorescence staining using the adult Tg(kcne1:eGFP) line, eGFP expression controlled by the kcne1 promoter was also observed in the atrial and ventricular myocytes, as in the case of the Tg(kcne6:eGFP) line (Fig. 2f–i; Supplementary Fig. 3b). However, eGFP signals of the Tg(kcne1:eGFP) line differ from those of the Tg(kcne6:eGFP) line. Although eGFP signals of the Tg(kcne1:eGFP) line are also broadly observed in these myocytes, some regions show higher signal levels than others. These results suggest that the KCNE6 protein may play a more specific role in the heart, while the KCNE1 protein in zebrafish may have multiple physiological roles, as is known in the human body1.
a Representative images of the Tg(kcne6:eGFP) line at 72 hpf under eGFP fluorescence. The scale bar indicates 0.5 mm. The inset shows a close-up view of the heart. b–d Immunofluorescence staining of a heart section from the adult Tg(kcne6:eGFP) line using the anti-GFP antibody. The scale bars indicate 0.5 mm in (b) and 0.1 mm in (c, d). AT atrial myocyte. V ventricular myocyte. BA bulbus arteriosus. e Representative images of the Tg(kcne1:eGFP) line at 72 hpf under eGFP fluorescence. The scale bar indicates 0.5 mm. The inset shows a close-up view of the heart. f–i Immunofluorescence staining of a heart section from the adult Tg(kcne1:eGFP) line using the anti-GFP antibody. The scale bars indicate 0.5 mm in (f, h) and 0.1 mm in (g, i). j Heart rates in the WT and kcne6 KO lines. In each line, ten healthy embryos from three independent pairs (n = 30 in total) were collected at 24 hpf and monitored from 24 hpf to 120 hpf. Student’s t-test was used to compare the WT and kcne6 KO lines and significance was assigned at p < 0.05 (*p < 0.05, **p < 0.01, and ***p < 0.001). The exact p-values were 0.0013 at 48 hpf and 0.0087 at 96 hpf.
To characterize the KCNE6 protein function in the zebrafish heart, we created a kcne6 knockout (KO) (kcne6−/−) line by CRISPR/Cas9 and examined its cardiac development (Supplementary Fig. 4a). The survival rate of the kcne6−/− line was similar to that of the WT line from spawning to the initiation of feeding (120 hpf) (Supplementary Fig. 4b). However, the heart rates in the kcne6−/− line were mildly but significantly reduced up to 96 hpf and then recovered at 120 hpf to a similar extent as the WT line (Fig. 2j). These results suggest that the KCNE6 protein is functionally expressed in the heart and is involved in cardiac excitability, at least in the early stages in zebrafish.
The kcne6 genes exist throughout vertebrates
We next investigated whether the kcne6 gene is conserved in other species. Using the nucleotide and amino acid sequences of DrKCNE6 as queries, we searched the KCNE6 sequences using BLAST and tBLASTn programs24,25. Putative kcne6 ORF sequences exist in all vertebrates we surveyed (Fig. 3a, b; Supplementary Fig. 5). In most cases, the kcne6 ORF sequence resides near the locus coding the pax3 gene, suggesting that the genome sequence around the kcne6 ORF is conserved throughout vertebrates. The lengths of nucleotide sequences of the putative kcne6 CDSs in vertebrates are variable. Among them, when translated to amino acid sequences, human (HsKCNE6) and zebrafish (DrKCNE6) have the shortest (29 AA) and longest (122 AA) sequences, respectively (Fig. 3a). Except for HsKCNE6, all putative KCNE6 proteins contain complete single transmembrane segments.
Sequence alignments (a) and phylogenetic tree (b) of KCNE6 proteins from human (HsKCNE6), Sumatran orangutan (Pongo abelii) (PaKCNE6), cow (Bos taurus) (BtKCNE6), dog (Canis lupus familiaris) (ClfKCNE6), koala (Phascolarctos cinereus) (PcKCNE6), chicken (Gallus gallus) (GgKCNE6), green anole (Anolis carolinensis) (AcKCNE6), tropical clawed frog (Xenopus tropicalis) (XtKCNE6), coelacanth (Latimeria chalumnae) (LcKCNE6), and zebrafish (DrKCNE6). Representative current traces (c–g) and G-V relationships (h) of HsKCNQ1 WT alone as well as HsKCNQ1 co-expressed with HsKCNE6 WT, PaKCNE6 WT, BtKCNE6 WT, and ClfKCNE6 WT. Representative current traces (i–n) and G-V relationships (o) of HsKCNQ1 co-expressed with PcKCNE6 WT, GgKCNE6 WT, AcKCNE6 WT, XtKCNE6, LcKCNE6 WT, and DrKCNE6 WT. Error bars indicate ± s.e.m. for n = 5 in (h,o).
To explore whether the putative KCNE6 proteins can modulate KCNQ1 gating like DrKCNE6, we synthesized putative kcne6 CDSs and co-expressed their cRNAs with HsKCNQ1 in Xenopus oocytes. The KCNE6 proteins from eutherians, which have relatively short in N and C termini, failed to shift the G-V relationship (Fig. 3c–h). However, the KCNE6 proteins from lower vertebrates, including marsupials, shifted the G-V relationship of KCNQ1 in a positive direction, like DrKCNE6 WT (Fig. 3i–o). Confocal microscopy analysis revealed that DrKCNE6 was localized to the membrane, whereas HsKCNE6 and dog KCNE6 (ClfKCNE6), both of which failed to shift the G-V relationship, were not (Supplementary Fig. 6). To identify the regions of KCNE6 required for modulating KCNQ1 gating, we created a series of DrKCNE6 constructs with truncated N and C termini and co-expressed them with DrKCNQ1 (Fig. 4a). In the constructs with truncated N termini, the ability to modulate DrKCNQ1 decreased as the N terminus was progressively shortened, except for the construct with the first 20 residues removed (DrKCNE6ΔN20). The modulatory ability was almost completely lost in the construct with 40 residues removed (DrKCNE6ΔN40) (Fig. 4b–h). Similarly, in the constructs with truncated C termini, the modulation of DrKCNQ1 was attenuated as the C terminus was shortened, except in the construct with the last 30 residues removed (DrKCNE6ΔC30). The ability to modulate was almost completely lost in the construct with 45 residues removed (DrKCNE6ΔC45) (Fig. 4i–o).
a Diagram of N and C termini truncation constructs of DrKCNE6. Representative current traces (b–g) and G-V relationships (h) of DrKCNQ1 WT alone, DrKCNQ1 WT co-expressed with DrKCNE6 WT, and DrKCNQ1 WT co-expressed with each of the N-terminally truncated DrKCNE6 constructs. Representative current traces (i–n) and G-V relationships (o) of DrKCNQ1 WT alone, DrKCNQ1 WT co-expressed with DrKCNE6 WT, and DrKCNQ1 WT co-expressed with each of the C-terminally truncated DrKCNE6 constructs. Error bars indicate ± s.e.m. for n = 5 in (h,o).
These results suggest that a specific length of the N- and C-terminal regions is required for the KCNE6 protein to properly localize to the membrane and modulate KCNQ1 gating, and this may be the reason why the eutherian KCNE6s are not functional. This is consistent with the results of previous studies showing that the N and C termini of KCNE proteins are important for the KCNE-dependent KCNQ1 modulation34,35,36. Notably, these putative kcne6 ORF sequences, which exhibited modulation activity on KCNQ1 when translated to proteins, are annotated as a long non-coding RNA (lncRNA) region in koalas (Phascolarctos cinereus) as well as unknown function protein-coding regions in chickens (Gallus gallus), green anoles (Anolis carolinensis), and tropical clawed frogs (Xenopus tropicalis) (Supplementary Fig. 5). KCNE6 proteins may be expressed in these species, as suggested in zebrafish.
The KCNE6 protein modulates Kv4 channels
While the KCNE family proteins have been extensively studied as auxiliary subunits for KCNQ1, they are also known to modulate other Kv channels, at least in heterologous expression systems1,2. That tempted us to investigate whether KCNE6 can modulate other Kv channels. We chose Kv4 and Kv11 because they are cardiac potassium channels known to be modulated by one or more of the five KCNE isoforms6,15,37,38,39,40. We expressed the zebrafish orthologs of these Kv channels with or without DrKCNE6. When co-expressed with zebrafish Kv4.2 (DrKCND2) and Kv4.3 (DrKCND3), DrKCNE6 did not significantly affect the voltage dependence but modulated current kinetics by decreasing the transient and increasing the sustained components (Fig. 5a–f), being in agreement with results of previous studies showing that the KCNE3 isoform modulates current kinetics of the Kv4 family18. In contrast, DrKCNE6 did not affect the gating properties of zebrafish KCNH6a (DrKCNH6a), which underlies the hERG-like current in zebrafish (Fig. 5g–i)41. These results suggest that KCNE6 may modulate other Kv channels, as in the case of other KCNE proteins.
Representative current traces (a–d), current-voltage (I-V) relationships (e), and ratios of the sustained to transient currents (f) of DrKCND2 WT and DrKCND3 WT alone as well as them co-expressed with DrKCNE6 WT. Student’s t-test was used to compare KCNDs alone and with DrKCNE6 WT and significance was assigned at p < 0.05 (*p < 0.05, **p < 0.01, and ***p < 0.001) in (f). The exact p-values were 0.0003 for DrKCND2 and 0.0017 for DrKCND3. Representative current traces (g, h) and G-V relationships (g) of DrKCNH6a WT alone and co-expressed with DrKCNE6 WT. Error bars indicate ± s.e.m. for n = 5 in (e, f, i).
Discussion
Here, we report the identification of KCNE6 as a sixth member of the single-transmembrane KCNE family of proteins that functions as an auxiliary subunit for KCNQ1 and possibly for other Kv channels. Remarkably, it has been more than two decades since the identification of KCNE5, the most recently discovered and characterized isoform of the KCNE family of proteins7,11. Using two-electrode voltage clamp (TEVC) and VCF recordings, we showed that the zebrafish KCNE6 protein modulates the VSD of KCNQ1 and thereby shifts the G-V relationship of KCNQ1 in a positive direction (Fig. 1; Supplementary Fig. 1). We also showed that KCNE6 can modulate Kv4.2 and Kv4.3 (Fig. 5).
The results of promoter analysis using the transgenic line Tg(kcne6:eGFP) and RT-PCR suggested that the KCNE6 protein is highly expressed in the heart (Fig. 2a-d; Supplementary Fig. 2). Furthermore, the heart rate of kcne6−/− zebrafish was slightly reduced in the early stages but recovered at 120 hpf (Fig. 2j). This is similar to the results of previous studies with KCNE1-5 isoform KO mice showing that the KO was not lethal but that the mice showed abnormalities to some extent39,42,43,44,45. Although the reason why the heart rates of the kcne6−/− line were temporarily reduced and recovered is not clear, a recent biophysical study using long QT syndrome (LQTS) type 5 (LQT5) KCNE1 variants in humans, which cause cardiac arrhythmia by inhibiting Iks current amplitudes, showed that the reduced IKs current amplitudes caused by LQT5 KCNE1 variants can be restored by co-expressing KCNE3 WT46. Considering this case, we speculate that KCNE6 in the heart might be replaced by other proteins, likely by KCNE1, in the kcne6−/− line. However, the physiological roles of the KCNE6 protein and its relationship to cardiac functions are yet to be determined. Further analysis using the kcne6−/− line is required for a better understanding of the role of KCNE6 in vivo. In addition, while we only examined the effect of kcne6−/− under normal conditions in this study, previous studies suggested that another KCNE protein, KCNE4, serves as an emergency protein: its expression is upregulated when the cardiovascular system is impaired, such as impairment caused by cardiac arrhythmia47 and hepatic ischemia-reperfusion injury48. Therefore, the evaluation of KCNE6 protein expression and localization in various zebrafish models of heart failure23 may help to further dissect the role of KCNE6 protein.
Homology searches from various vertebrates and subsequent TEVC recordings showed that KCNE6 is conserved in vertebrates and functions as an auxiliary subunit for KCNQ1 in lower vertebrates, up to marsupials. Since the kcne6 ORF sequences in lower vertebrates, including marsupials, overlap with the nucleotide sequences annotated as lncRNA or protein-coding regions of unknown function in the Ensembl genome browser49, KCNE6 proteins may be expressed in these species, as was observed in zebrafish (Fig. 2a–d; Supplementary Figs. 2 and 3). In contrast, KCNE6 proteins from eutherians, including humans, did not modulate KCNQ1 gating, suggesting that kcne6 genes in eutherians became pseudogenes at a certain point of evolution, probably after the divergence of marsupials and eutherians (Fig. 3; Supplementary Fig. 5). Considering that the effect of kcne6−/− on zebrafish heart rate was significant but mild (Fig. 2j) and considering that KCNE1 serves as a primary auxiliary subunit in the human heart1, it is likely that the KCNE6 function has been replaced by KCNE1, resulting in the KCNE6 protein being dispensable in eutherians. However, since additional protein-coding regions in exon 1 of KCNE3 and KCNE4, which extend their N-terminal extracellular regions by about 50 amino acids, were discovered in 201650, there is still a possibility that KCNE6 proteins exist as a longer variant and modulate KCNQ1 gating in eutherians.
Among the five known KCNE isoforms, KCNE1 and KCNE3 have gained more attention than the others due to their contrasting effects on KCNQ1 gating and their potential as tools for understanding and manipulating KCNQ1 gating, along with their physiological importance2,32. As for the mechanism by which KCNE1 and KCNE3 exhibit contrasting effects, some biophysical studies suggested that a specific region termed the “triplet”, which consists of three consecutive amino acid residues located in the middle of the transmembrane segment (“FTL” in KCNE1 and “TVG” in KCNE3) (orange box in Fig. 1b), serves as the structural element responsible for their modulation properties on KCNQ132,51,52. However, the structure of the KCNQ1-KCNE3 complex and our following biophysical study showed that a broad range of the KCNE3 transmembrane segment interacts with the VSD of KCNQ1 and is required for the constitutive activity of the KCNQ1-KCNE3 channel33,53, suggesting that the “triplet” is not the only region responsible for the modulation property of each KCNE isoform. In KCNE6, the “triplet” is composed of “LAL” or “MAL”, suggesting that the “FTL triplet” in KCNE1 might not be essential for KCNE1-like modulations, such as the positive shift in voltage dependence and slow current activation. Given that KCNE6 shifts the G-V relationship of KCNQ1 in a positive direction similar to KCNE1, despite having the highest amino acid sequence similarity to KCNE3, it may serve as a valuable new tool for understanding the distinct modulation mechanisms of KCNE1 and KCNE3.
Overall, in this study, we unexpectedly identified a new KCNE isoform in zebrafish and showed that a trace of the gene exists throughout vertebrates, including humans. The possible reasons why KCNE6 was not discovered until now are that it is a small gene and became a pseudogene during evolution. This is different from the recently characterized auxiliary subunits for ion channels54,55,56,57, as many of them were identified on the basis of prior reports of genes or physiological functions. Therefore, the process for identification of KCNE6 in this study is unique and can lead to the discovery of more genes, including auxiliary subunits, that once appeared but were hidden during evolution.
Materials and methods
Zebrafish maintenance and lines
Zebrafish adults, embryos, and larvae (RIKEN-WT, kcne6−/−, Tg(kcne6:eGFP), and Tg(kcne1:eGFP)) used in this study were maintained in self-circulating aquarium systems at Jichi Medical University (Japan). All experiments using zebrafish were approved by the Animal Care Committee of Jichi Medical University under protocol no. 21049-01 and were performed according to guidelines. We have complied with all relevant ethical regulations for animal use.
Generation of genetically modified zebrafish lines
For generating the kcne6 KO line (kcne6−/−), a frameshift mutant of the kcne6 gene was generated by the CRISPR/Cas9 system. A CRISPR/Cas9 target site in kcne6 was searched for using the CRISPRdirect program58 (Supplementary Fig. 4a). The RIKEN WT strain was injected at the 1-cell stage in the cell body for CRISPR/Cas9 injections with 25 ng/µl gRNA and 50 ng/µl Cas9 mRNA. The genomic DNA of F1 fish was extracted from the caudal fin. An amplicon that includes the target region of each gene was generated by PCR and sequenced by a commercial company (Eurofins Genomics). F1 fish that had mutations were crossed. Homozygous KO F2 fish were selected by the genome sequence.
For generating Tg(kcne6:eGFP) and Tg(kcne1:eGFP) lines, the standard Tol2 transgenesis protocol was used59. Nucleotide sequences containing a 3.5K-bp genome region upstream of the kcne6 ORF region and those containing a 4K-bp genome region upstream of the kcne1 ORF region were respectively inserted into the pT2AL200R150G expression vector, which contains eGFP CDS60. The RIKEN WT strain was injected at the 1-cell stage in the cell body with 25 pg of each of the recombinant plasmids and 25 pg of Tol2 RNA. At 72-120 hpf, injected fish were screened for eGFP expression. eGFP-positive fish were raised and outcrossed to establish the lines.
Heart rate measurement
Heart rates of zebrafish embryos and larvae were manually counted using an MVX10 microscope (Olympus) at room temperature. Before the counting, the zebrafish embryos and larvae were acclimated to the room temperature environment under visible light.
RNA isolation, cDNA synthesis, and reverse transcription-PCR
Total RNA was extracted and purified from the heart of the adult RIKEN WT strain using NucleoSpin RNA Plus (Takara Bio, 740984) according to the manufacturer’s instructions. Reverse transcription was performed using PrimeScriptTM RT Master Mix (Takara Bio, RR036A). The obtained cDNA was stored at −20 °C and later used for PCR as a template. cDNAs of DrKCNQ1, DrKCNE6, and Drβ-actin were amplified using the primers listed below.
DrKCNQ1-Fw: ATGACCTCCCCAGCTCACAGTACAGCCACCGAGGC
DrKCNQ1-Rv: GCTGTCCTCATGAATAGCAGTTGTTGTGGACAGTT
DrKCNE6-Fw: CCTCATTGTTAGCATCGATATGCCTC
DrKCNE6-Rv: TCACACACTGTTCTCCTCCTC
β-actin-Fw: TAATACACAGCCATGGATGAGG
β-actin-Rv: GGGAGCAATGATCTTGATCTTC
Immunostaining of the zebrafish heart
We used mature WT as well as Tg(kcne6:eGFP) and Tg(kcne1:eGFP) lines (more than 12 months post-hatching) for our study. The zebrafish were deeply anesthetized using tricaine (Sigma-Aldrich, E10521). Then the hearts were dissected and fixed in phosphate-buffered saline (PBS) containing 4% paraformaldehyde (PFA) for several hours at 4 °C. GFP fluorescence was observed using an MVX10 microscope (Olympus) at room temperature, and photographs were captured with a DS-Fi3 digital camera (Nikon). The fixed hearts after observations were used for immunohistochemistry to analyze the localization of kcne1- or kcne6-expressing cells in detail. The fixed hearts were immersed in PBS containing 30% sucrose, embedded in O.C.T. compound (Sakura Finetek), frozen at −80 °C, cryosectioned at 20 μm using a CM 3050S cryostat (Leica Microsystems), and mounted onto micro slide glass CREST (Matsunami Glass). Sections were incubated overnight with an anti-GFP polyclonal antibody (Thermo Fisher Scientific, A6455) diluted at a 1:500 ratio with a PBS buffer containing 0.3% Tween 20 (PBST). After rinsing twice with PBST, the sections were incubated with CoraLite488-conjugated Goat Anti-Rabbit IgG (Proteintech, SA00013-2) diluted at a 1:500 ratio with PBST for 2 h at room temperature. Then the sections were washed with PBST and coverslipped with CC/Mount (Diagnostic BioSystems). GFP fluorescence was observed using an MVX10 microscope (Olympus), and photographs were captured with a DS-Fi3 digital camera (Nikon).
Expression in Xenopus laevis oocytes
The cDNAs for KCNQ1 (human, NCBI Accession Number NP_000209.2; zebrafish, NP_001116714.1), KCNE1 (zebrafish, XP_017213386.1), KCNE3 (mouse, NP_001177798), and KCNE6 as well as KCNE6 cDNAs with a C-terminal HRV-3C cleavage site followed by an eGFP-Flag tag were inserted into the pGEMHE expression vector61. All KCNE6 cDNAs except those for zebrafish were synthesized by Genewiz (Azenta Life Sciences). The cRNAs were transcribed using mMESSAGE mMACHINETM T7 Transcription Kits (Thermo Fisher Scientific, AM1344). Oocytes were surgically removed from adult female Xenopus laevis frogs and anesthetized in water containing 0.1% tricaine (Sigma-Aldrich, E10521) for 15-30 min. The oocytes were treated with collagenase (Sigma-Aldrich, C0130) for 6-7 h at room temperature to remove the follicular cell layer. Defolliculated oocytes of similar sizes at stage V or VI were selected, microinjected with 50 nl of cRNA solution (10 ng for KCNQ1 and 1 ng for KCNEs) using a NANOJECT II (Drummond Scientific Co.), and incubated until use at 18 °C in Barth’s solution (88 mM NaCl, 1 mM KCl, 2.4 mM NaHCO3, 10 mM HEPES, 0.3 mM Ca(NO3)2, 0.41 mM CaCl2, and 0.82 mM MgSO4, pH 7.6) supplemented with 0.1% penicillin-streptomycin solution (Sigma-Aldrich, P4333). All experiments using Xenopus laevis were approved by the Animal Care Committee of Jichi Medical University under protocol no. 21030-03 and were performed according to guidelines. We have complied with all relevant ethical regulations for animal use.
Two-electrode voltage clamp
cRNA-injected oocytes were incubated for 1-3 days. Ionic currents were recorded with a two-electrode voltage clamp using an OC-725C amplifier (Warner Instruments) at room temperature. The bath chamber was perfused with Ca2+-free ND96 solution (96 mM NaCl, 2 mM KCl, 2.8 mM MgCl2, 5 mM HEPES, pH 7.6) supplemented with 100 µM LaCl3 to block endogenous hyperpolarization-activated currents26,33. The microelectrodes were drawn from borosilicate glass capillaries (Harvard Apparatus, GC150TF-10) using a P-1000 micropipette puller (Sutter Instrument) to a resistance of 0.2–1.0 MΩ and filled with 3 M KCl. Currents were elicited from the holding potential of −90 mV to steps ranging from −100 to +60 mV in +20 mV steps each for 2 sec with 10-sec intervals. Oocytes with a holding current larger than −0.2 µA at -90 mV were excluded from the analysis. Generation of voltage-clamp protocols and data acquisition were performed using a Digidata 1550 interface (Molecular Devices) controlled by pCLAMP 10.7 software (Molecular Devices). Data were sampled at 10 kHz and filtered at 1 kHz.
Voltage dependence analysis
G–V relationships were taken from tail current amplitude at −30 mV fitted using pCLAMP 10.7 software (Molecular Devices) to a single Boltzmann equation:
where Gmax and Gmin are the maximum and minimum tail current amplitudes, respectively, z is the effective charge, V1/2 is the half-activation voltage, T is the temperature in degrees Kelvin, F is Faraday’s constant, and R is the gas constant. G/Gmax, which is the normalized tail current amplitude, was plotted against membrane potential for presentation of the G–V relationships.
Voltage-clamp fluorometry
cRNA-injected oocytes were incubated for 2-4 days, labeled for 30 min with 5 µM Alexa FluorTM 488 C5 maleimide (Thermo Fisher Scientific, A10254) in high potassium KD98 solution (98 mM KCl, 1.8 mM CaCl2, 1 mM MgCl2, 5 mM HEPES, pH 7.6)26,33, and washed with Ca2+-free ND96 solution to remove unreacted Alexa probes. The bath chamber was filled with Ca2+-free ND96 solution supplemented with 100 µM LaCl3. The microelectrodes were drawn from borosilicate glass capillaries (Harvard Apparatus, GC150TF-15). Currents were elicited from the holding potential of −90 mV to steps ranging from +80 to −180 mV or +60 to −180 mV in −20 mV steps each for 2 sec with 20-sec intervals. Oocytes with a holding current larger than −0.4 µA at −90 mV were excluded from the analysis. Generation of voltage-clamp protocols and data acquisition were performed using a Digidata 1440 A interface (Molecular Devices) controlled by pCLAMP 10.7 software (Molecular Devices). Data were sampled at 10 kHz and filtered at 1 kHz. Fluorescence recordings were performed with an MVX10 microscope (Olympus) equipped with a 2x objective lens (MVPLAPO 2XC, NA = 0.5, Olympus), 2x magnification changer (MVX-CA2X, Olympus), GFP filter cube (U-MGFPHQ/XL, Olympus), and an XLED1 LED light source with a BDX (450-495 nm) LED module (Excelitas Technologies). Fluorescence signals were obtained by using a photomultiplier (H10722-110; Hamamatsu Photonics) and digitized at 1 kHz through Digidata1440, filtered at 50 Hz, and recorded using pClamp10 simultaneously with ionic currents. The shutter for the excitation was open during the recording, which induced a continuous decrease in fluorescence due to photobleaching. Therefore, we calculated the bleaching rate for each experiment using the baseline levels of the initial 1100 ms before test pulses of each trace and compensated the fluorescence traces by subtracting the bleached component calculated from each trace’s bleaching rate (R), assuming that the fluorescence was linearly decreased. Arithmetic operations were performed with Clampfit software from pClamp10.
\([{{\rm{Compensated}}}\; {{\rm{trace}}}]=[{{\rm{recorded}}}\; {{\rm{trace}}}] \times \left(\right.1{{\mbox{-}}}({{\rm{R}}}\; \times[{{\rm{time}}}])\)33
where [time] is the time value of the point given by Clampfit. We then normalized the fluorescence traces by setting each baseline level to 1.
VCF analysis
F–V relationships were taken from the fluorescence change from the baseline (ΔF) plotted against membrane potential. ΔF values were then normalized by ΔF80mV for the normalized F-V relationships shown in Fig. 1. For KCNQ1 alone, F–V relationships were fitted using Igor Pro software (WaveMatrices Co.) to a single Boltzmann equation:
where Fmin and Fmax are the maximum and baseline fluorescence components, z is the effective charge for the fluorescence component, V1/2 is the half-activation voltage for the fluorescence component, T is the temperature in degrees Kelvin, F is Faraday’s constant, and R is the gas constant. For KCNQ1 co-expressed with each KCNE subtype, F–V relationships were fitted using Igor Pro software (WaveMatrices Co.) to a double Boltzmann equation:
where F1, F2, and Fmin are the first, second, and baseline fluorescence components, respectively, z1 and z2 are the effective charges for each fluorescence component, V1/2(F1) and V1/2(F2) are the half-activation voltage for each fluorescence component, T is the temperature in degrees Kelvin, F is Faraday’s constant, and R is the gas constant.
Confocal microscopy
cRNA-injected oocytes were incubated for 2-3 days. Imaging of the oocytes was performed under an FV1000-D confocal laser scanning fluorescent microscope (Olympus).
Statistics and reproducibility
The data were expressed as means ± SEM (n = 5). Statistical analysis was performed with Student’s t-test with EZR software62, and significance was assigned at p < 0.05 (*p < 0.05, **p < 0.01, and ***p < 0.001). For electrophysiological recordings, data were collected from two or more independent batches of oocytes. For experiments using zebrafish, data were collected from three independent pairs.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data were collected as described in the Methods section and are presented in the paper and/or the supplementary materials. The source data is available in the Supplementary Data file.
References
Abbott, G. W. Kv Channel Ancillary Subunits: Where Do We Go from Here? Physiology 37, 225–241 (2022).
Abbott, G. W. KCNE1 and KCNE3: The yin and yang of voltage-gated K+ channel regulation. Gene 576, 1–13 (2016).
Takumi, T., Ohkubo, H. & Nakanishi, S. Cloning of a membrane protein that induces a slow voltage-gated potassium current. Science 242, 1042–1045 (1988).
Barhanin, J. et al. KVLQT1 and lsK (minK) proteins associate to form the IKs cardiac potassium current. Nature 384, 78–80 (1996).
Sanguinetti, M. C. et al. Coassembly of KVLQT1 and minK (IsK) proteins to form cardiac IKs potassium channel. Nature 384, 80–83 (1996).
Abbott, G. W. et al. MiRP1 forms IKr potassium channels with HERG and is associated with cardiac arrhythmia. Cell 97, 175–187 (1999).
Piccini, M. et al. KCNE1-like gene is deleted in AMME contiguous gene syndrome: identification and characterization of the human and mouse homologs. Genomics 60, 251–257 (1999).
Tinel, N., Diochot, S., Borsotto, M., Lazdunski, M. & Barhanin, J. KCNE2 confers background current characteristics to the cardiac KCNQ1 potassium channel. EMBO J 19, 6326–6330 (2000).
Schroeder, B. C. et al. A constitutively open potassium channel formed by KCNQ1 and KCNE3. Nature 403, 196–199 (2000).
Grunnet, M. et al. KCNE4 is an inhibitory subunit to the KCNQ1 channel. J. Physiol. 542, 119–130 (2002).
Angelo, K. et al. KCNE5 induces time- and voltage-dependent modulation of the KCNQ1 current. Biophys. J. 83, 1997–2006 (2002).
McCrossan, Z. A. et al. MinK-related peptide 2 modulates Kv2.1 and Kv3.1 potassium channels in mammalian brain. J. Neurosci. 23, 8077–8091 (2003).
Abbott, G. W. et al. MiRP2 forms potassium channels in skeletal muscle with Kv3.4 and is associated with periodic paralysis. Cell 104, 217–231 (2001).
Deschênes, I. & Tomaselli, G. F. Modulation of Kv4.3 current by accessory subunits. FEBS Lett 528, 183–188 (2002).
McDonald, T. V. et al. A minK-HERG complex regulates the cardiac potassium current IKr. Nature 388, 289–292 (1997).
Ávalos Prado, P. et al. KCNE1 is an auxiliary subunit of two distinct ion channel superfamilies. Cell 184, 534–544.e11 (2021).
Talbi, K., Ousingsawat, J., Centeio, R., Schreiber, R. & Kunzelmann, K. KCNE1 does not shift TMEM16A from a Ca2+ dependent to a voltage dependent Cl- channel and is not expressed in renal proximal tubule. Pflugers Arch 475, 995–1007 (2023).
Wang, W. et al. Functional significance of K+ channel β-subunit KCNE3 in auditory neurons. J. Biol. Chem. 289, 16802–16813 (2014).
Delpón, E. et al. Functional effects of KCNE3 mutation and its role in the development of Brugada syndrome. Circ. Arrhythm. Electrophysiol. 1, 209–218 (2008).
Okamura, Y. et al. Comprehensive analysis of the ascidian genome reveals novel insights into the molecular evolution of ion channel genes. Physiol. Genomics 22, 269–282 (2005).
Abramochkin, D. V., Hassinen, M. & Vornanen, M. Transcripts of Kv7.1 and MinK channels and slow delayed rectifier K+ current (IKs) are expressed in zebrafish (Danio rerio) heart. Pflugers Arch 470, 1753–1764 (2018).
Whitesell, T. R. et al. foxc1 is required for embryonic head vascular smooth muscle differentiation in zebrafish. Dev. Biol. 453, 34–47 (2019).
Narumanchi, S. et al. Zebrafish Heart Failure Models. Front Cell Dev Biol 9, 662583 (2021).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25, 3389–3402 (1997).
Osteen, J. D. et al. KCNE1 alters the voltage sensor movements necessary to open the KCNQ1 channel gate. Proc. Natl. Acad. Sci. USA. 107, 22710–22715 (2010).
Osteen, J. D. et al. Allosteric gating mechanism underlies the flexible gating of KCNQ1 potassium channels. Proc. Natl. Acad. Sci. USA. 109, 7103–7108 (2012).
Zaydman, M. A. et al. Domain–domain interactions determine the gating, permeation, pharmacology, and subunit modulation of the IKs ion channel. Elife 3, e03606 (2014).
Taylor, K. C. et al. Structure and physiological function of the human KCNQ1 channel voltage sensor intermediate state. Elife 9, 1–31 (2020).
Barro-Soria, R. et al. KCNE1 divides the voltage sensor movement in KCNQ1/KCNE1 channels into two steps. Nat. Commun. 5, 3750 (2014).
Barro-Soria, R., Perez, M. E. & Larsson, H. P. KCNE3 acts by promoting voltage sensor activation in KCNQ1. Proc. Natl. Acad. Sci. USA. 112, E7286–E7292 (2015).
Barro-Soria, R. et al. KCNE1 and KCNE3 modulate KCNQ1 channels by affecting different gating transitions. Proc. Natl. Acad. Sci. USA. 114, E7367–E7376 (2017).
Kasuya, G. & Nakajo, K. Optimized tight binding between the S1 segment and KCNE3 is required for the constitutively open nature of the KCNQ1-KCNE3 channel complex. Elife 11, 1–19 (2022).
Nakajo, K. & Kubo, Y. KCNE1 and KCNE3 Stabilize and/or Slow Voltage Sensing S4 Segment of KCNQ1 Channel. J. Gen. Physiol. 130, 269–281 (2007).
Chen, J., Zheng, R., Melman, Y. F. & McDonald, T. V. Functional interactions between KCNE1 C-terminus and the KCNQ1 channel. PLoS One 4, e5143 (2009).
Wang, Y. et al. Probing the structural basis for differential KCNQ1 modulation by KCNE1 and KCNE2. J. Gen. Physiol. 140, 653–669 (2012).
Finley, M. R. et al. Expression and coassociation of ERG1, KCNQ1, and KCNE1 potassium channel proteins in horse heart. Am. J. Physiol. Heart Circ. Physiol. 283, H126–H138 (2002).
Radicke, S. et al. Functional modulation of the transient outward current Ito by KCNE β-subunits and regional distribution in human non-failing and failing hearts. Cardiovasc. Res. 71, 695–703 (2006).
Roepke, T. K. et al. Targeted deletion of kcne2 impairs ventricular repolarization via disruption of IK,slow1 and Ito,f. FASEB J 22, 3648–3660 (2008).
Niwa, N. & Nerbonne, J. M. Molecular determinants of cardiac transient outward potassium current (Ito) expression and regulation. J. Mol. Cell. Cardiol. 48, 12–25 (2010).
Langheinrich, U., Vacun, G. & Wagner, T. Zebrafish embryos express an orthologue of HERG and are sensitive toward a range of QT-prolonging drugs inducing severe arrhythmia. Toxicol. Appl. Pharmacol. 193, 370–382 (2003).
Charpentier, F., Merot, J., Riochet, D., Le Marec, H. & Escande, D. Adult KCNE1-knockout mice exhibit a mild cardiac cellular phenotype. Biochem. Biophys. Res. Commun. 251, 806–810 (1998).
Hu, Z. et al. Kcne3 deletion initiates extracardiac arrhythmogenesis in mice. FASEB J 28, 935–945 (2014).
Hu, Z., Wei, W., Zhou, L., Chen, M. & Abbott, G. W. Kcne4 deletion sex-specifically predisposes to cardiac arrhythmia via testosterone-dependent impairment of RISK/SAFE pathway induction in aged mice. Sci. Rep. 8, 8258 (2018).
David, J.-P. et al. Deletion in mice of X-linked, Brugada syndrome- and atrial fibrillation-associated Kcne5 augments ventricular KV currents and predisposes to ventricular arrhythmia. FASEB J 33, 2537–2552 (2019).
Déri, S. et al. A Possible Explanation for the Low Penetrance of Pathogenic KCNE1 Variants in Long QT Syndrome Type 5. Pharmaceuticals 15, 1550 (2022).
Kelemen, K., Greener, I. D., Wan, X., Parajuli, S. & Donahue, J. K. Heterogeneous repolarization creates ventricular tachycardia circuits in healed myocardial infarction scar. Nat. Commun. 13, 830 (2022).
Hu, Z. et al. Kcne4 deletion sex dependently inhibits the RISK pathway response and exacerbates hepatic ischemia-reperfusion injury in mice. Am. J. Physiol. Regul. Integr. Comp. Physiol. 316, R552–R562 (2019).
Martin, F. J. et al. Ensembl 2023. Nucleic Acids Res 51, D933–D941 (2023).
Abbott, G. W. Novel exon 1 protein-coding regions N-terminally extend human KCNE3 and KCNE4. FASEB J 30, 2959–2969 (2016).
Melman, Y. F., Domènech, A., De la Luna, S. & McDonald, T. V. Structural Determinants of KvLQT1 Control by the KCNE Family of Proteins. J. Biol. Chem. 276, 6439–6444 (2001).
Melman, Y. F., Krumerman, A. & McDonald, T. V. A single transmembrane site in the KCNE-encoded proteins controls the specificity of KvLQT1 channel gating. J. Biol. Chem. 277, 25187–25194 (2002).
Sun, J. & MacKinnon, R. Structural Basis of Human KCNQ1 Modulation and Gating. Cell 180, 340–347 (2020).
Han, W. et al. Shisa7 is a GABAA receptor auxiliary subunit controlling benzodiazepine actions. Science 366, 246–250 (2019).
Dudem, S. et al. LINGO1 is a regulatory subunit of large conductance, Ca2+-activated potassium channels. Proc. Natl. Acad. Sci. USA. 117, 2194–2200 (2020).
Peters, C. H. et al. Isoform-specific regulation of HCN4 channels by a family of endoplasmic reticulum proteins. Proc. Natl. Acad. Sci. USA. 117, 18079–18090 (2020).
Zhou, Z. et al. MyoD-family inhibitor proteins act as auxiliary subunits of Piezo channels. Science 381, 799–804 (2023).
Naito, Y., Hino, K., Bono, H. & Ui-Tei, K. CRISPRdirect: software for designing CRISPR/Cas guide RNA with reduced off-target sites. Bioinformatics 31, 1120–1123 (2015).
Suster, M. L., Kikuta, H., Urasaki, A., Asakawa, K. & Kawakami, K. Transgenesis in zebrafish with the tol2 transposon system. Methods Mol. Biol. 561, 41–63 (2009).
Urasaki, A., Morvan, G. & Kawakami, K. Functional dissection of the Tol2 transposable element identified the minimal cis-sequence and a highly repetitive sequence in the subterminal region essential for transposition. Genetics 174, 639–649 (2006).
Liman, E. R., Tytgat, J. & Hess, P. Subunit stoichiometry of a mammalian K+ channel determined by construction of multimeric cDNAs. Neuron 9, 861–871 (1992).
Kanda, Y. Investigation of the freely available easy-to-use software “EZR” for medical statistics. Bone Marrow Transplant 48, 452–458 (2013).
Madeira, F. et al. The EMBL-EBI Job Dispatcher sequence analysis tools framework in 2024. Nucleic Acids Res 52, W521–W525 (2024).
Robert, X. & Gouet, P. Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res 42, 320–324 (2014).
Acknowledgements
We thank the members of the Nakajo laboratory. This work was supported by the Japan Society for the Promotion of Science (JSPS) KAKENHI (Grant Nos. 23H02666 and 23K27357 to G.K. and 21K06786 to K.N.) and by The Nakajima Foundation to G.K..
Author information
Authors and Affiliations
Contributions
K.N. started the project. G.K. and K.N. conceived and designed the project. G.K., K.R., and K.N. performed electrophysiological experiments. G.K., Y.Y., F.O., and K.N. generated genetically modified zebrafish lines. G.K., B.Z., and K.R. performed the experiments using zebrafish. G.K. and K.N. wrote the manuscript and supervised all of the research. All authors approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Geoffrey Abbott, David Fedida and the other, anonymous, reviewers for their contribution to the peer review of this work. Primary Handling Editors: Janesh Kumar and Laura Rodríguez Pérez. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Kasuya, G., Zempo, B., Yamamoto, Y. et al. Identification of KCNE6, a new member of the KCNE family of potassium channel auxiliary subunits. Commun Biol 7, 1662 (2024). https://doi.org/10.1038/s42003-024-07352-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-024-07352-6
This article is cited by
-
Voltage-dependent potassium channel regulatory subunits in the immune system
Biophysical Reviews (2026)