Abstract
Vertebrates possess two condensins, I and II, that are essential for chromosome condensation and segregation. Condensin II has also been implicated in maintaining genome integrity outside of mitosis, though the underlying mechanisms are unclear. Here, we found that condensin II interacts with a short linear motif in the disordered N-terminal tail of the Bloom syndrome helicase BLM, contributing to BLM association with nascent DNA and genome stability. Disrupting the BLM-condensin II interaction reduced replication speed, increased fork stalling and sister-chromatid exchanges, delayed repair of DNA double-strand breaks, and led to micronuclei. In S phase, interactions of SMC2 with other condensin II subunits and with BLM weakened temporarily, suggesting a conformational change followed by phosphorylation-induced disruption of BLM interactions with TOP2A and RPA. Our findings suggest a new way by which BLM contributes to genome integrity and implicates condensin II in interphase functions linked to genome stability.
Similar content being viewed by others
Introduction
DNA replication and repair are highly coordinated mechanisms involving multiple protein complexes. Cells lacking factors in either are prone to genome instability and tumorigenesis, and some proteins have been demonstrated to play roles in both processes. One of these factors is the tumor suppressor BLM, a 3’ – 5’ DNA helicase of the highly conserved RecQ family1. Loss of BLM activity leads to a rare disorder known as Bloom syndrome (BSyn), which is characterized by extreme cancer incidence, abnormally short stature, hypersensitivity to sun exposure, immunodeficiency, and increased incidence of early onset type II diabetes2,3. At the cellular level, BLM deficiency causes defects in DNA damage repair and DNA replication, including an increased rate of sister chromatid exchanges (SCEs), slow response to and repair of DNA double-strand breaks (DSBs), accumulation of R-loops, and sensitivity to DNA-damaging agents, as well as reduced replication speed, increased fork stalling, and difficulty in restarting stalled forks4,5,6,7,8,9,10,11. While BLM’s role in DNA repair through the homologous recombination repair (HR) pathway is established and more recent studies have illuminated its functions in the response to replication stress, our understanding of BLM function in unperturbed cells is still incomplete.
Single-molecule studies have demonstrated that BLM, like its S. cerevisiae homolog Sgs1, is a highly processive DNA helicase capable of unwinding DNA at a rate of 70-80 bp per second in tracks as long at 8–10 kilobases12. A growing number of proteins predominantly bind to BLM’s long, intrinsically disordered N-terminal tail and affect BLM activity or recruit it to its substrates13,14,15,16. BLM binds topoisomerase IIIα and RMI1/2 to form the BTR complex, which is the primary dissolvase needed for Holliday junction dissolution during the final step of HR to prevent crossovers17,18. Replication protein A (RPA) increases the processivity of the BLM helicase, targets it to sites of DNA damage, and is important for fork restart19. BLM also binds to MCM to regulate normal replication speed, ensure timely repair of replication-associated DSBs, and suppress sensitivity to replication stress4. Other DNA repair and replication proteins that have been found in complexes with BLM include RAD51, DNA2, MLH1, FEN1, TOPBP1, TOP2A, POLδ, and two other members of the RecQ helicase family, RecQ4 and WRN, which are associated with Rothmund-Thomson syndrome and Werner syndrome, respectively 20,21,22,23,24,25,26,27,28. However, since exact binding sites on BLM have been identified for only a few of these interacting proteins, it has remained uncertain how most of these factors contribute to the cellular functions of BLM.
Condensin I and II are essential for chromosome condensation and segregation in vertebrates. Condensin II initiates chromosome condensation prior to nuclear envelop breakdown by establishing an initial chromosome axis by using its loop extrusion activity to create large loops that act as a scaffold for condensin I29,30,31,32. Following nuclear envelop breakdown in late prophase, condensin I then inserts smaller loops into the chromatin scaffold provided by condensin II through loop extrusion to impart axial chromosome rigidity30,31,32. Loss of either condensin I or II activity results in distorted metaphase chromosome architecture and chromosome segregation errors including chromatin bridges, ultra-fine bridges, and micronuclei formation, which are thought to be the source of the human condensin-associated disorders known as condensinopathies30,33. While the roles of both condensin I and II in chromatin compaction during mitosis have been known for some time, more recent studies are revealing their roles outside of mitosis in additional genome maintenance mechanism. Condensin I in complex with PARP1-XRCC1 has been shown to play a role in DNA single-strand break repair34,35 and condensin II has been shown to facilitate DSB repair through the HR pathway36. Moreover, condensin II loop extrusion activity has been implicated in interphase chromatin folding and spatial partitioning37,38,39 as well as in the resolution of newly replicated sister chromatids to prevent entanglements prior to mitosis40. However, while the functions of condensins outside of their established roles in mitosis have begun to come to light, the exact mechanisms by which they help maintain genome stability have yet to be identified.
Here, we characterize a novel interaction between the Bloom syndrome helicase BLM and the condensin II complex outside of mitosis. BLM utilizes a conserved N-terminal motif to interact with the non-SMC subunits of condensin II, which allows BLM to discriminate it from condensin I. Stable expression of BLM that is deficient for condensin II binding revealed that the BLM-condensin II interaction promotes normal replication speed, prevents fork stalling, and suppresses sensitivity to high levels of replication stress but, unlike BLM, does not play a role in replication fork restart. Moreover, our analysis reveals that the BLM-condensin II interaction aids in the timely repair of DSBs and in suppressing chromosome aberrations such as chromatin bridges, micronuclei, and SCEs. We also identify that while BLM and condensin II are present on newly synthesized DNA, BLM localization to nascent DNA is facilitated through interaction with condensin II but not vice versa.
Results
BLM interacts with condensin II in S phase
In addition to its helicase activity, the function of BLM in the maintenance of genome integrity is determined by the numerous proteins that bind its exceptionally long, intrinsically disordered N-terminal tail. Recently, we performed a co-immunoprecipitation to identify factors that work with BLM to maintain genome integrity in unperturbed S phase4. Co-immunoprecipitates of BLM from a BLM-proficient skin fibroblast cell line (GM00637, BLM-WT) 3 h after release from G1/S arrest were analyzed by mass spectrometry and compared to the isogenic BLM-knockout (BLM-KO) cell line derived by CRISPR-Cas41, identifying the condensin subunit SMC2 as a putative novel interactor of BLM (Fig. 1a, Supplementary Table S1). Known BLM interacting proteins were also detected including the BTR complex members TOP3A and RMI1 (Fig. 1a) as well as a subunit of the replicative DNA helicase MCM, which was the focus of a previous study in the lab4 (Supplementary Table S1). An interaction between BLM and SMC2 was confirmed using reciprocal co-immunoprecipitation of endogenous BLM and SMC2 from nuclear extracts of cells synchronized at G1/S and released into S phase for 3 h (Fig. 1b). Inclusion of benzonase in all co-immunoprecipitations demonstrated that the BLM-SMC2 interaction is not mediated by DNA. Successful proximity ligation of BLM and SMC2 in wildtype (WT) and BLM-complemented BLM-KO cells (BLM-COMP) further validated the BLM-SMC2 interaction in vivo (Fig. 1c, Supplementary Fig. S1a).
a SMC2 was identified by mass spectrometry in a co-immunoprecipitate of endogenous BLM from nuclear extracts of WT cells 3 h after release from G1/S arrest4. Select known BLM interacting proteins are shown for comparison. Isogenic BLM-KO cells were subjected to the same analysis and yielded zero hits for the listed proteins. Experiment was performed in triplicate. b Reciprocal co-immunoprecipitation of endogenous BLM and SMC2 from nuclear extracts of WT and BLM-KO cells 3 h after release from G1/S arrest. c, Proximity ligation assay (PLA) using antibodies against endogenous BLM and SMC2 in WT, BLM-COMP, and BLM-KO cells 3 h after release from G1/S arrest and quantification of BLM-SMC2 PLA foci/nucleus (WT, n = 111 nuclei; BLM-COMP, n = 103 nuclei; BLM-KO, n = 104 nuclei). Scale bars, 10 μM. Significance of difference between WT and BLM-COMP was determined by a Mann–Whitney test (ns, not significant). For additional PLA controls, see Supplementary Fig. S1a. d Five-subunit condensin I and II complexes (H, NCAPH; D2, NCAPD2; G, NCAPG; H2, NCAPH2; D3, NCAPD3; G2, NCAPG2). e Co-immunoprecipitation of endogenous BLM from whole-cell extracts of WT cells 3 h after release from G1/S arrest was probed with antibodies against condensin I and II subunits. Condensin subunits are color-coordinated with their respective condensin from (d). f Co-immunoprecipitation of endogenous NCAPH from whole-cell extracts of WT cells 3 h after release from G1/S arrest was probed with antibodies against condensin I and II subunits and BLM. g Co-immunoprecipitation of endogenous NCAPH2 from whole-cell extracts of WT cells 3 h after release from G1/S arrest was probed with antibodies against condensin I and II subunits and BLM.
Since SMC2 is a core component of both condensin I and condensin II42 (Fig. 1d), we next determined to which complex BLM binds. Using whole-cell extract, which contains both the primarily cytoplasmic condensin I as well the nuclear condensin II, we determined that BLM specifically interacts with condensin II as evidenced by the ability for BLM to co-immunoprecipitate the NCAPH2 and NCAPD3 subunits of condensin II but not the NCAPH or NCAPG subunits of condensin I (Fig. 1e). Moreover, we performed reciprocal co-immunoprecipitation using antibodies against endogenous NCAPH and NCAPH2. While both NCAPH and NCAPH2 could co-immunoprecipitate SMC2 and SMC4, as expected, and NCAPH could co-immunoprecipitate the condensin I non-SMC subunit NCAPG, only NCAPH2 could co-immunoprecipitate BLM (Fig. 1f,g). Importantly, NCAPH and NCAPH2 failed to co-immunoprecipitate with each other, in line with previous reports42,43.
BLM residues W154, M157 and F160 specifically mediate the condensin II interaction
BLM contains two major domains: the intrinsically disordered N-terminal tail spanning residues 1-64744, which acts as a major protein-interaction hub, and the mostly ordered C-terminal half, which includes the helicase domain and spans residues 648-1417. Since the initial co-immunoprecipitation of BLM pulled down SMC2, we used SMC2 in the mammalian two-hybrid (M2H) system to identify the site on BLM required for association with condensin II. Having narrowed down the site to residues 160-184 of BLM (Fig. 2a, Supplementary Fig. S1c) we targeted the conserved sequence 153DWDDMDDFD161 for mutagenesis (Fig. 2b). In addition to F160, we selected W154 and M157 for mutagenesis as aromatic and hydrophobic residues are generally underrepresented in disordered regions and may be part of a molecular recognition motif for protein binding. Indeed, mutating W154, M157 and F160 to lysine, hereafter referred to as BLM-WMFmut, significantly reduced the ability of full-length BLM to interact with SMC2 in the M2H system (Fig. 2c, Supplementary Fig. S1d). To assess the effects of BLM-WMFmut on association with condensin II in vivo, we constructed two clones of the BLM-KO cell line stably expressing BLM-WMFmut at levels similar to BLM in the BLM-COMP and WT cell lines (Fig. 2d). BLM-WMFmut was unable to co-immunoprecipitate SMC2 and the condensin-II-specific subunit NCAPH2 (Fig. 2e) and prevented proximity ligation with NCAPH2 and SMC2 (Fig. 2f, Supplementary Fig. S1e), demonstrating that the WMF mutation disrupts the BLM interaction with condensin II in cells. The region surrounding the WMF site of BLM had previously also been implicated in binding RPA13,19, TOP3A45, RAD5120, TOP2A46, and MCM64, but none of these interactions were disrupted by BLM-WMFmut (Fig. 2e), indicating that BLM residues W154, M157 and F160 specifically mediate the interaction with condensin II.
a BLM truncations tested in mammalian-two-hybrid (M2H) assay against full-length SMC2. Interaction is indicated by secreted alkaline phosphatase (SEAP) activity as ++ (strong), + (mild), and - (absent). See Supplementary Fig. S1c for measurements of SEAP activity from three experiments. Red box indicates BLM region required for interaction with SMC2. b Alignment of BLM residues 150-184 with BLM from other vertebrates. Residues W154, M157, and F160 were mutated to lysine, referred to as BLM-WMFmut. c SEAP activity detected in M2H assay between two independently generated pVP16-BLM-WMFmut clones (#1, #2) and pM-SMC2. pM3-VP16 is a positive control from the manufacturer expressing a fusion of the GAL4 DNA-binding domain to the VP16 activation domain. Significance of differences between means ± SD was determined by a t-test and is reported as ****p ≤ 0.0001. d Plasmid expressing BLM-WMFmut was stably transfected into BLM-KO cells. Western blot of nuclear extracts from two independently generated BLM-WMFmut clones (#1, #2) and of WT, BLM-KO, and BLM-COMP cells was probed with BLM antibody. PCNA was used as a loading control. Expression levels were analyzed 3 h after release from G1/S arrest. e Co-immunoprecipitations of BLM and BLM-WMFmut #1 were probed with antibodies against SMC2, condensin II-specific subunit NCAPH2, and known BLM-interacting proteins. Whole-cell extracts were prepared from cells 3 h after release from G1/S arrest. f PLA using antibodies against endogenous BLM and NCAPH2 in BLM-COMP, BLM-WMFmut #1, and BLM-KO cells 3 h after release from G1/S arrest. Scale bars, 10 μm. BLM-NCAPH2 PLA foci/nucleus were quantified (BLM-COMP, n = 102 nuclei; BLM-WMFmut #1, n = 130 nuclei; BLM-KO, n = 100 nuclei). Significance of differences was determined by a Mann–Whitney test and is reported as ****p ≤ 0.0001. For additional PLA controls, see Supplementary Fig. S1b.
Regulation of BLM-condensin II interaction during S phase and G2 phase
We had identified the BLM-condensin II interaction in extracts prepared from early S phase cells. To determine the regulation of the BLM-condensin II interaction throughout S and G2 phases, we synchronized cells at the G1/S boundary by double-thymidine block, released the cells into S phase, collected samples every hour for a total of 12 h, and co-immunoprecipitated endogenous BLM from nuclear extracts. We assessed the synchrony and cell cycle progression of our double-thymidine block scheme by Western blotting for common cell cycle markers (Supplementary Fig. S2a). In the co-immunoprecipitation, BLM pulled down NCAPH2 for the entire 12-h time course but briefly showed the weakened ability to pull down SMC2 at hours 4 and 5 (Fig. 3a). We repeated the BLM co-immunoprecipitation and also probed for SMC4, showing that unlike SMC2, BLM pulled down SMC4 equally throughout the 12-h time course (Fig. 3c). Since condensin II subunits and BLM are phosphorylated24,47,48,49 we tested whether phosphorylation events regulate the BLM-condensin II interaction. We observed that λ-protein phosphatase (λPPase) treatment had no effect on the co-immunoprecipitation of NCAPH2 by BLM whereas SMC2 and SMC4 were not recovered (Fig. 3b, d). These observations suggest that BLM engages with condensin II through the non-SMC subunits, possibly NCAPH2, thereby allowing BLM to distinguish it from condensin I and that the interaction of BLM with the SMC subunits is dependent on phosphorylation.
Endogenous BLM was immunoprecipitated after release from G1/S arrest in (a) the absence of λ protein phosphatase (λPPase) and in (b) the presence of λPPase and probed with antibodies against SMC2, NCAPH2, RPA, and TOP2A. Cells were synchronized by double-thymidine, released, and BLM immunoprecipitated every hour for 12 h. IgG was used as a negative control for immunoprecipitation. Endogenous BLM was immunoprecipitated after release from G1/S arrest in (c) the absence of λ protein phosphatase (λPPase) and in (d) the presence of λPPase and probed with antibodies against SMC2, SMC4, and NCAPH2. Cells were synchronized by double-thymidine, released, and BLM immunoprecipitated every hour for 12 h. IgG was used as a negative control for immunoprecipitation. Experiment was performed at least twice with similar results. e Endogenous NCAPH2 was immunoprecipitated from WT, BLM-KO, and BLM-WMFmut #1 cells and probed with antibodies against BLM, SMC2, and SMC4. Cell cultures were arrested at G1/S by double-thymidine treatment, released, and NCAPH2 immunoprecipitated every hour for 7 h. IgG was used as a negative control for immunoprecipitation. Experiment was performed twice with similar results. f Endogenous NCAPH2 and SMC2 was immunoprecipitated from the same WT lysate used for hours 3–7 in (e) and probed with antibodies against NCAPH2, BLM, SMC2, and SMC4. IgG was used as a negative control for immunoprecipitation. NCAPH2 immunoprecipitation was performed three times with similar results and the SMC2 reciprocal immunoprecipitation once.
Reports that NCAPH2 and SMC2 disengage during condensin II-mediated loop extrusion50 prompted us to test whether BLM contributes to the change in the NCAPH2-SMC2 interaction at hours 4 and 5. However, NCAPH2 from whole-cell extracts displayed weakened interaction with SMC2 in S phase whether BLM was present (WT) or not (BLM-KO) or whether BLM-condensin II interaction was disrupted (BLM-WMFmut) (Fig. 3e), although the reduction in NCAPH2-SMC2 interaction shifted from hours 4 and 5 in WT and BLM-WMFmut cells to hours 5 and 6 in BLM-KO cells (Fig. 3e, Supplementary Fig. S2b). Reciprocal co-immunoprecipitation of SMC2 also displayed weakened interaction with NCAPH2, SMC4, and BLM at hours 4 and 5 in WT (Fig. 3f). A separate study found that NCAPH2 and SMC2 disengagement could also regulate chromatin binding of condensin II51. Therefore, we examined the presence of condensin II subunits in chromatin fractions of WT cells during the 4–5-h interval. Neither BLM nor condensin II showed major changes in chromatin association during 7 h after release from G1/S arrest (Supplementary Fig. S2c), indicating that the reduced interaction between BLM/NCAPH2 and SMC2 is not due to a marked chromatin loading/unloading event. These findings show that BLM does not affect the disengagement of NCAPH2 from SMC2; however, BLM deficiency, which slows DNA replication speed, delays the period of weakened NCAPH2-SMC2 interaction.
Besides chromosome condensation in mitosis, condensin II is important for interphase chromatin maintenance, likely through its loop extrusion activity52, and for sister chromatid resolution in S phase40, and its actions have been linked to topoisomerase IIα52,53,54. Since BLM physically interacts with topoisomerase IIα46 we tested if the temporary disruption of the interaction of BLM/NCAPH2 with SMC2 coincided with changes in BLM-TOP2A interaction. Interestingly, BLM interaction with TOP2A was disrupted during hours 5 and 6 by a phosphorylation event (Fig. 3b), partially overlapping with the weakened interaction with SMC2 (Fig. 3a, b), raising the possibility that BLM-TOP2A interaction in S phase is functionally linked to the BLM-condensin II interaction.
Notably, the site at which BLM interacts with condensin II overlaps with one of the sites at which BLM has been reported to bind RPA19, and condensin II subunits SMC2-SMC4 have been shown to interact with ssDNA and are capable of reannealing RPA-coated ssDNA in vitro55,56,57. Thus, we probed for RPA in BLM co-immunoprecipitates in the presence and absence of λPPase. Intriguingly, RPA interacted with BLM throughout S phase and G2 phase except for the same 2-h period where BLM-TOP2A interaction was disrupted (Fig. 3a). In another parallel to TOP2A, the disruption of BLM-RPA interaction at hours 4 and 5 was induced by a phosphorylation event (Fig. 3b).
BLM-WMFmut cells exhibit decreased replication speed and increased fork stalling but normal fork restart after replication stress
BSyn cells show markedly reduced replication speed, increased fork stalling, impaired fork restart, and increased doubling time6,58,59,60. In contrast to BLM-KO cells, we did not detect a significant difference in doubling time between BLM-proficient cells and BLM-WMFmut cells; however, similar to BLM-KO cells, BLM-WMFmut cell populations showed a small accumulation of cells in the S phase (Supplementary Fig. S3a-c). Single-molecule analysis of CIdU/IdU-labeled DNA fibers determined that cells expressing BLM-WMFmut displayed a 14% reduction in replication speed as compared to a 28% reduction in BLM-KO cells (Fig. 4a). Analysis of sister forks revealed that BLM-WMFmut cells had a lower IdU/IdU ratio than BLM-COMP cells (Fig. 4b), indicating increased fork stalling upon uncoupling of BLM from condensin II.
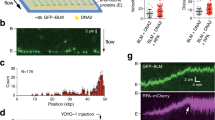
a Median replication speed as determined by DNA fiber assay. Cells were labeled for 30 min in CIdU followed by 30 min in IdU. Representative images of DNA fibers are shown below the graph. At least 100 DNA fibers per cell line were analyzed. Significance of differences was determined by a Mann–Whitney test and is reported as ***p ≤ 0.001. Experiment was performed twice with similar results. b Sister fork asymmetry was determined by determining the ratio of IdU track lengths of sister forks from (a). Number of sister forks analyzed: BLM-COMP, n = 32; BLM-KO, n = 25; BLM-WMFmut #1, n = 31; BLM-WMFmut #2, n = 32. Significance of differences was determined by Mann–Whitney test and is reported as *p ≤ 0.05; **p ≤ 0.01; ***p ≤ 0.001. c Replication fork restart efficiency following exposure to 4 mM hydroxyurea (HU) for 4 h. Median IdU/CIdU track length ratios were calculated. Representative images of DNA fibers are shown below the graph. At least 100 DNA fibers were analyzed for each cell line. Significance of differences was determined by a Mann-Whitney test and is reported as ***p ≤ 0.001; ns, not significant. d Clonogenic assay following 48-h treatment with varying concentrations of HU. Average percentage of survival from three biological replicates is reported ± SD. Significance of differences was determined by a t test and is reported as *p ≤ 0.05; **p ≤ 0.01; ***p ≤ 0.001. Experiment was performed 3 times with similar results. e Immunofluorescence microscopy showing examples of sister chromatid exchanges (SCEs) in DAPI-stained metaphase chromosome spreads of BLM-KO cells probed with antibody against BrdU. Quantification of SCEs per metaphase is reported. SCEs in chromosomes from 5 metaphases per cell line were analyzed and significance of differences was determined by a Mann–Whitney test (**p ≤ 0.01). f Western blots of γ-H2AX clearance during release from 1-h exposure to 0.5 μM camptothecin (CPT) in WT, BLM-KO, BLM-WMFmut #1 cells, as well as BLM-COMP cells treated with mock siRNA or NCAPH2 siRNA. Experiment was performed 3 times with similar results and representative blots are shown. Quantification of γ-H2AX abundance relative to the histone H2A loading control shows the average ± SD of the three experiments. γ-H2AX/H2A was normalized to 1 for the 0-h time point when CPT was removed. Significance of differences between WT BLM-KO, and BLM-WMFmut #1 as well as between BLM-COMP siMock and siNCAPH2 was determined by a Mann–Whitney test (*p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, ****p ≤ 0.0001). g Clonogenic assay following 48-h treatment with varying concentrations of camptothecin (CPT). Average survival of three replicates is reported ± SD. Significance of differences was determined by a t test and is reported as *p ≤ 0.05; **p ≤ 0.01; ***p ≤ 0.001; **** p ≤ 0.0001. Experiment was performed 2 times with similar results. h Microscopy images of DAPI-stained asynchronous cell cultures showing anaphase bridges and lagging chromosomes (laggards). Percentage of anaphases with either bridges or laggards; at least 100 anaphases from four experiments were analyzed to calculate the average ± SD. Significance of differences was determined by a t-test and is reported as *p ≤ 0.05; **p ≤ 0.01; ***p ≤ 0.001; **** p ≤ 0.0001; ns, not significant. i Micronuclei occurrence in untreated cells. To detect micronuclei, cells were stained with DAPI. Images were pseudo-colored with ImageJ and micronuclei are indicated by arrows. Percentage of cells with one (1) or multiple (>1) micronuclei per interphase nucleus was calculated. At least 100 nuclei from three experiments were used to calculate the average ± SD. Significance of differences was determined by a t-test and is reported as *p ≤ 0.05; **p ≤ 0.01; ***p ≤ 0.001; ns, not significant. j Analysis of CREST-staining micronuclei in untreated cells. Representative image of a micronucleus positive for CREST antibody staining. Quantification of micronuclei positive for CREST staining. At least 100 nuclei from three experiments were used to calculate the average ± SD. Significance of differences between means were analyzed by a t test (ns, not significant).
BLM-KO cells are defective in replication fork restart after HU-induced replication stress59. Notably, BLM residues 150–163 where the WMF mutation resides had previously been implicated in a BTR-RPA interaction involving two sites in BLM and one in RMI1, and this BTR-RPA interaction was also required for fork restart following HU treatment19. While BLM-WMFmut cells were sensitive to high concentrations of HU, albeit not to the same level as BLM-KO cells (Fig. 4d), they restarted stalled forks as efficiently as BLM-COMP cells (Fig. 4c). Thus, BLM-WMFmut separates BLM function in fork restart after replication stress from BLM functions in fork progression under unperturbed conditions.
Since uncoupling BLM and condensin II resulted in replication defects, we hypothesized that depletion of endogenous condensin II by siRNA should phenocopy the observed replication defects in BLM-WMFmut cells. BLM-COMP cells treated with siNCAPH2, which depletes endogenous NCAPH2 (Supplementary Fig. S5d), exhibited a mild replication speed defect compared to cells treated with mock siRNA and produced a fork-stalling phenotype similar to BLM-WMFmut (Supplementary Fig. S4a, b). Thus, condensin II promotes normal replication and prevents fork stalling, and its interaction with BLM aids in this Sphase function.
The WMF motif contributes to BLM’s role in DSB repair
A hallmark of cells lacking BLM is a sharp increase in the exchange of genetic material between sister chromatids7. To determine if the BLM-condensin II interaction contributes to the suppression of sister chromatid exchanges (SCEs), we immunofluorescently labeled mitotic chromosome spreads after two rounds of BrdU incorporation61, revealing a significant (~3.5-fold) increase in SCEs in both BLM-WMFmut clones compared to cells expressing wildtype BLM (Fig. 4e, Supplementary Fig. S3d). To gain further insight into the role of the BLM-condensin II interaction in HR, we induced replication-dependent DSBs using 0.5 µM camptothecin (CPT) and followed DSB repair kinetics by assessing γ-H2AX clearance after CPT removal. In WT cells, γ-H2AX was efficiently cleared around 24 h post-release, indicative of completion of DSB repair, whereas BLM-KO and BLM-WMFmut cells exhibited a slow DSB repair phenotype with γ-H2AX still detectable 60 h after release from CPT (Fig. 4f). This failure of BLM-WMFmut cells to repair CPT-induced DNA damage in a timely manner prompted us to test whether they were hypersensitive to CPT-induced DSBs. Indeed, disrupting the BLM-condensin II interaction resulted in a significant reduction in survival after CPT exposure, albeit not to the extent seen in BLM-KO cells (Fig. 4g).
Furthermore, disruption of condensin II by treatment with siNCAPH2 resulted in a similar (~3-fold) increase in SCEs (Supplementary Fig. S4c,d) as that seen in BLM-WMFmut cells (Fig. 4e). Similar to BLM-WMFmut cells, BLM-COMP cells treated with siNCAPH2 also failed to effectively repair DSBs by 60 h after release from CPT (Fig. 4f). Taken together, these findings indicate that condensin II and the BLM-condensin II interaction significantly contribute to the regulation and efficiency of HR-mediated DSB repair.
BLM-WMFmut promotes mitotic chromosome instability
Both BLM deficiency and condensin II deficiency cause elevated rates of lagging anaphase chromosomes, anaphase bridges, and micronuclei, which are all precursors to genetic loss and genome instability33,62,63,64,65. We sought to determine the extent to which the BLM-condensin II interaction contributes to the prevention of these mitotic aberrations and the resulting genome instability. We observed that mitoses of both BLM-WMFmut clones exhibited a significant increase in anaphase bridges compared to cells expressing wildtype BLM. But in contrast to BLM-KO cells, which show a slight increase in lagging chromosomes, BLM-WMFmut cells exhibited the same frequency of lagging chromosomes as BLM-proficient cells (Fig. 4h). BLM is also known to localize to a sub-class of anaphase bridges that do not stain with DAPI known as ultrafine anaphase bridges (UFBs)63. We determined that BLM-WMFmut was able to bind UFBs and that UFBs coated with wildtype BLM or with BLM-WMFmut were found at similar rates in the respective cell lines (Supplementary Fig. S3e-g).
Unlike WT cells, BLM-WMFmut cells showed increased micronuclei formation, which arise from chromosomes that failed segregation into daughter cell nuclei66, demonstrating mitotic chromosome instability in BLM-WMFmut cells (Fig. 4i). Nuclei of BLM-WMFmut cells mostly had a single micronucleus as opposed to BLM-KO cells, which frequently had multiple micronuclei (Fig. 4i). Increased micronuclei formation is inherently harmful; however, micronuclei that stain with CREST antibody, which binds to human centromeres, are thought to be more detrimental as they are an indicator of entire, rather than partial, chromosome loss67,68. We found that micronuclei in WT, BLM-COMP, and BLM-KO cells all had CREST foci in approximately one-third of their micronuclei and BLM-WMFmut cells were not significantly different (Fig. 4j). These results suggest a requirement of the BLM-condensin II interaction for maintaining mitotic chromosome stability.
Condensin II contributes to BLM localization to nascent DNA
BLM physically interacts with several replisome components and may be present at nascent DNA, including in unstressed cells4,26. Prompted by our finding that the BLM-condensin II interaction prevents fork stalling and contributes to normal replication speed, we analyzed the association of BLM and condensin II with nascent DNA and tested whether their interaction aids in this localization. Using immunofluorescence microscopy, we determined that ~20% of endogenous BLM and ~50% of condensin II subunits SMC2 and NCAPH2 readily overlapped with EdU-labeled nascent DNA (Fig. 5a). To determine whether the interaction between BLM and condensin II aids in their localization to nascent DNA, we performed in situ protein interaction with nascent replication forks (SIRF)69. First, following EdU labeling and EdU:biotin co-Click with biotin azide and Alexa Fluor 488 azide, we performed PLA using antibodies against biotin and endogenous SMC2 or NCAPH2. SMC2 and NCAPH2 localized to nascent DNA equally in cells proficient (BLM-COMP) or deficient (BLM-KO) for BLM or expressing BLM-WMFmut, demonstrating that the association of condensin II with nascent DNA is not dependent on BLM or interaction with BLM (Fig. 5b, c). However, the SIRF signal for BLM-WMFmut was significantly reduced compared to the SIRF signal for BLM (Fig. 5d), suggesting that condensin II contributes to BLM association with nascent DNA. To validate this finding, we used siNCAPH2 to deplete NCAPH2 in BLM-COMP cells, which produced a SIRF signal for BLM that was similar to that for BLM-WMFmut (Fig. 5e). These findings demonstrate that subsets of BLM and condensin II localize to nascent DNA, with condensin II increasing the BLM association on nascent DNA but not vice versa.
a Immunofluorescence microscopy images of endogenous BLM, NCAPH2, SMC2 and EdU-labeled nascent DNA in WT cells. Nucleus is outlined in merged image and box represents ROI for magnified image. Scale bars, 10 μM and 1 μM (magnified image). Quantification of BLM, NCAPH2, and SMC2 colocalization with EdU in WT cells. Colocalization percentage was determined by counting the number of BLM, NCAPH2, or SMC2 foci that overlapped with EdU in comparison to the total number of BLM, NCAPH2, or SMC2 foci in an ROI for each nucleus. 20 nuclei were analyzed for each protein and median percentage of colocalization is reported. b SIRF assay of SMC2 in BLM-COMP, BLM-WMFmut #1, and BLM-KO cells. Following nascent DNA labeling with EdU and EdU:biotin co-Click with biotin azide and Alexa Fluor 488 azide, PLA was performed using antibodies against biotin and endogenous SMC2. Nucleus is outlined in the merged image. Scale bars, 10 μM. Corrected total Cellular Fluorescence (ctCF) of the SMC2 SIRF signal/EdU ctCF is shown for individual nuclei (BLM-COMP, n = 150 nuclei; BLM-WMFmut #1, n = 156 nuclei; BLM-KO, n = 79 nuclei) and the median presented by a horizontal line. Significance of differences between cell lines was determined using a Mann-Whitney test (ns, not significant). c SIRF assay of NCAPH2 in BLM-COMP, BLM-WMFmut #1, and BLM-KO cells. Following nascent DNA labeling with EdU and EdU:biotin co-Click with biotin azide and Alexa Fluor 488 azide, PLA was performed using antibodies against biotin and endogenous NCAPH2. Nucleus is outlined in the merged image. Scale bars, 10 μM. Corrected total Cellular Fluorescence (ctCF) of the NCAPH2 SIRF signal/EdU ctCF is shown for individual nuclei (BLM-COMP, n = 99 nuclei; BLM-WMFmut #1, n = 85 nuclei; BLM-KO, n = 85 nuclei) and the median presented by a horizontal line. Significance of differences between cell lines was determined using a Mann-Whitney test (ns, not significant). d SIRF assay of BLM in BLM-COMP, BLM-WMFmut #1, and BLM-KO cells. Following nascent DNA labeling with EdU and EdU:biotin co-Click with biotin azide and Alexa Fluor 488 azide, PLA was performed using antibodies against biotin and endogenous BLM. Nucleus is outlined in the merged image. Scale bars, 10 μM. Corrected total Cellular Fluorescence (ctCF) of the BLM SIRF signal/EdU ctCF is shown for individual nuclei (BLM-COMP, n = 120 nuclei; BLM-WMFmut #1, n = 125 nuclei; BLM-KO, n = 140 nuclei) and the median presented by a horizontal line. Significance of differences between cell lines was determined using a Mann–Whitney test and is reported as ****p ≤ 0.0001. e BLM SIRF assay in the BLM-COMP and BLM-WMFmut #1 cell lines following transfection with NCAPH2 siRNA or mock siRNA. Following nascent DNA labeling with EdU and EdU:biotin co-Click with biotin azide and Alexa Fluor 488 azide, PLA was performed using antibodies against biotin and endogenous BLM. Nucleus is outlined in the merged image. Scale bars, 10 μM. Corrected total Cellular Fluorescence (ctCF) of the BLM SIRF signal/EdU ctCF was calculated for each nucleus (BLM-COMP siNCAPH2, n = 102 nuclei; BLM-COMP siMock, n = 89 nuclei; BLM-WMFmut #1 siMock n = 94 nuclei) with the horizontal line representing the median. Significance of differences was determined by a Mann-Whitney test and is reported as ****p ≤ 0.0001. For additional controls for Fig. 5b–e see supplementary Fig. S5b. f Immunofluorescence microscopy of calyculin-induced premature chromosome condensation (PCC) in BLM-COMP, BLM-KO, and BLM-WMFmut #1 cells. Following nascent DNA labeling with EdU and chromosome spreading, EdU:biotin Click-iT was performed with biotin azide and slides were hybridized with biotin antibody and stained with DAPI. Box represents ROI for magnified image. Scale bars, 10 μM. The chromosomes of three spreads from a single experiment for each cell line were analyzed to quantify the percentage of resolved and unresolved sister chromatids.
Finally, prematurely condensing chromosomes in S phase by calyculin-induced histone H3S10 phosphorylation showed that cells expressing BLM-WMFmut or lacking BLM did not show the severe defect in the resolution of newly synthesized sister chromatids (Fig. 5f) previously reported for condensin II-defective cells40 and as seen when we depleted NCAPH2 (Supplementary Fig. S5c,d), suggesting that BLM does not significantly contribute to this critical S phase function of condensin II.
Discussion
In this study, we provide evidence for an interaction between BLM and condensin II that significantly enhances replication fork progression, suppression of crossover outcomes of DSB repair, efficiency of DSB repair, and mitotic chromosome stability. Based on interactions identified in this study, BLM associates with condensin II through its non-SMC subunits, possibly NCAPH2 (kleisin β), in a phosphorylation-independent manner. Interaction with the condensin-II-specific non-SMC proteins allows BLM to discriminate between condensin I and condensin II. BLM also co-immunoprecipitated with SMC2 and SMC4, however, in a phosphorylation-dependent manner, consistent with findings that phosphorylation stabilizes SMC-kleisin interactions49,70,71. Further, we observed transient weakening of SMC2 association with BLM-NCAPH2-SMC4 in S phase. The functional significance of this event is unknown. It does not appear to be related to chromatin loading of condensin II as we observed no gross changes of SMC2 levels in the chromatin fraction during this period of S phase. Instead, it could signify a conformational change in condensin II that is related to its function in S phase. For example, in models of loop extrusion by condensin, NCAPH2 dissociates from the coiled-coil neck of SMC2 to provide a gate for new DNA to enter the growing (extruding) loop32,50,51,72. A more recent model of loop extrusion additionally proposes that the SMC hinges could transiently dissociate73. Notably, NCAPH2 interaction with SMC2 during S phase was also reduced in the absence of BLM but delayed by 1 h. A possible explanation for this delay is that condensin II activity could be linked to progress of DNA replication, which is slowed in BLM-deficient cells. In support, BLM-WMFmut cells, which exhibit a less severe reduction in replication speed than BLM-KO cells and do not show the increased doubling time of BLM-KO cells, did not show this delay.
Functions of condensin II outside of mitotic chromosome condensation are only emerging, but they likely include its function in loop extrusion where the hinge region of SMC2-SMC4 grabs DNA at a distance and reels it into a growing DNA loop that is encircled by the SMC2-SMC4 globular domains and the non-SMC proteins at its base74 (Fig. 6a). Association of the WMF site in the flexible tether of BLM with condensin II could allow the BLM helicase domain to scan the newly reeled-in DNA for its substrates, such as G4 structures, R-loops, and DNA breaks that could impede replication forks as they move through the loops (Fig. 6a). In this context, it should be noted that replisome components do not interfere with loop extrusion by condensin75. Since all DNA of the newly synthesized sister chromatid will eventually need to be extruded into loops prior to mitosis, BLM association with condensin II could be a highly efficient mechanism for BLM to scan the sister chromatid for DNA breaks and illegitimate DNA structures prior to mitosis.
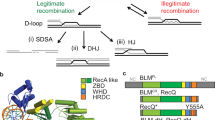
a Model of how BLM binding to condensin II increases efficiency of detection and repair of BLM substrates. Current models of loop extrusion posit that the hinge domain reels in new DNA through the SMC2-SMC4 lumen creating an intermediate loop with gate opening potentially allowing the intermediate loop to join the main loop50,72,74,106. BLM, through its interaction with condensin II at the WMF site, can scan this intermediate loop for its substrates such as R-loops, DNA breaks, or G-quadruplexes, leading to their resolution prior to becoming an impediment to the replication or transcription machinery, thus mediating normal DNA transactions and genome stability. The many DNA repair and replication factors that bind to BLM’s disordered tail likely contribute to resolution and repair. In the absence of BLM from condensin II due to the WMF mutation, the DNA lesions and replisome blocks are not resolved efficiently, thus increasing risk of collisions with the replisome or transcriptional machinery, leading to impaired replisome progression and fork stalling, impaired DNA-damage response, delayed DSB repair and, eventually, mitotic chromosome aberrations. Created in BioRender. Rodemoyer, B. (2025) https://BioRender.com/h40u746. b Model of how BLM-condensin II interaction may contribute to suppression of crossovers. Extrusion of adjacent loops may aid BLM helicase-mediated branch migration of Holliday junctions between sister chromatids. BLM-condensin II interaction puts BLM in close proximity to the HR intermediate, increasing efficiency of dissolution as noncrossovers (NCO) in cooperation with interacting proteins. Loss of BLM-condensin II interaction due to BLM-WMFmut reduces dissolution of HR intermediates by BLM complexes and increases resolution via cleavage by structure-specific endonucleases, causing crossovers (CO). Created in BioRender. Rodemoyer, B. (2025) https://BioRender.com/k17g890.
DNA structures that condensin II could encounter during loop extrusion of the new sister-chromatid would also include physical linkages between the sister chromatids that could result from repair of DSBs formed at ssDNA nicks or other DNA lesions during replication (Fig. 6b). As replication and loop extrusion near completion and sister-chromatid linkages accumulate between loops, BLM can dissolve such HR intermediates as noncrossovers (Fig. 6b). BLM’s association with condensin II in S/G2 phase could significantly increase the efficiency of this dissolution, thereby preventing the crossovers (i.e. SCEs) that result from nuclease cleavage in the absence of BLM or in BLM-WMFmut cells (Fig. 6b). Notably, condensin II had previously been implicated in HR repair efficiency through an interaction with the DNA damage response and cell cycle control protein Microcephalin 1 (MCPH1)36, although the exact mechanism is not clear.
Similar to BLM, MCPH1 discriminates between condensin I and II by interacting with the non-SMC subunits, specifically NCAPD3, to help regulate condensin II function in interphase. It is hypothesized that MCPH1 interaction with condensin II in G1 and G2 phases may help to facilitate ATP-dependent SMC2-SMC4 head dimerization51, which is a key step in condensin II-mediated loop extrusion. While this hypothesis has yet to be tested, the mechanism of MCPH1-condensin II and BLM-condensin II interaction are likely to be different since the proposed condensin II gate opening observed in S phase occurred in both, BLM-WT and BLM-KO cells. Intriguingly, a hallmark of patients with a wide range of condensinopathies is a reduced cranial head size characteristic of primary microcephaly33, which is uncoincidentally the main consequence of faulty MCPH1 activity76. The primary mechanism of microcephaly in patients with condensinopathies, it is thought to result from reduced neural stem cell proliferation potentially due to the aneuploidy caused by increased micronuclei frequency33. Bloom syndrome patients may also display microcephaly77,78 and BLM loss and BLM-WMFmut cause micronuclei; however, the exact mechanism of microcephaly in BSyn has remained elusive.
In addition to HR defects, BLM-WMFmut resulted in reduced replication speed and increased fork stalling, which is characteristic for BLM-deficient cells4,5,8, but did not impair BLM’s role in replication fork restart after replication stress. This suggests that the interaction with condensin II contributes to BLM function in unperturbed replication, whereas BLM interactions with factors at the fork, such as FEN179, POLδ26 and RPA19, may contribute to fork restart after replication stress. We note that BLM-WMFmut cells were only sensitive to high levels of HU, whereas the BLM-KO was sensitive at every concentration tested. Prolonged exposure of cells to HU has been reported to generate collapsed replication forks, which result in DSBs80. Given that BLM-WMFmut causes sensitivity to replication-dependent DSBs but does not impair fork restart, it is plausible that the lower concentrations of HU (0.2 and 0.4 mM) do not result in significant fork collapse, but the higher concentrations do so after 24-h exposure, leading to sensitivity of BLM-WMFmut cells not observed at lower HU concentrations.
Interestingly, the 13-amino-acid motif in BLM that includes the WMF residues necessary for condensin II binding has previously been implicated with one other site in BLM and one in RMI1 in RPA binding and fork restart19. The absence of a restart defect in the BLM-WMFmut cells shows that the interaction of BLM-WMFmut with TOP3A/RMI1/RMI2 and RPA is preserved, which was further supported by preservation of the BLM-RPA and BLM-TOP3A co-immunoprecipitation upon mutation of the WMF residues. It is possible that different residues in the 13-residue motif participate in binding at least two proteins; the RPA interaction likely depends on the abundance of DNA-mimicking acidic residues of the motif whereas condensin II interaction requires the embedded hydrophobic residues. In contrast to BLM-WMFmut, deleting a RPA binding site in BLM in the previous study did not lead to elevated SCEs19; this could be due to differences in the cell lines (RPE-1, GM00637) and SCE detection method used (Hoechst/Leishman’s differential staining versus immunofluorescence), which yielded ~3-fold higher SCE levels in this study.
That the BLM-condensin II interaction is present throughout S phase and G2 phase with structural changes in S phase, consistent with proposed condensin II activity, indicates roles of condensin II in genome stability in interphase outside of its established roles in mitosis33,81,82,83. This is further supported by our findings that depletion of condensin II by NCAPH2 siRNA resulted in several of the same defects observed in BLM-WMFmut cells, including a slight reduction in replication speed, an increase in fork stalling and SCEs, and delayed repair of replication-dependent DSBs. Intriguingly, cells from persons with BSyn and persons with condensinopathies share several defects, including increased chromatin bridging in anaphase, ultrafine bridges, and micronuclei, which have been linked to decatenation failures33 and increases in fork stalling84. Telomere stability and HR repair efficiency are also compromised in cells lacking either BLM or condensin and evidence for a centromere structural defect is growing36,85,86,87,88,89,90,91.
In addition to BLM helicase activity detecting and dissolving HR intermediates, G4s, and R-loops while bound to condensin II, BLM may also support genome maintenance by supplying a number of genome maintenance factors through interaction with its N-terminal tail. In addition to TOP3A/RMI1/RMI2, which participate with BLM in HJ dissolution, we showed that BLM physically interacts with TOP2A and RPA in S and G2 except for the same, brief phosphorylation-induced dissociation in S phase. Topoisomerase II is best known for its functional interaction with condensin II in sister chromatid resolution and chromosome condensation in mitosis53,92,93,94,95,96. Topoisomerase II has also been shown to decatenate and unknot DNA in C. elegans to support long-range loop extrusion by condensin54. Moreover, the ability of SMC2-SMC4 to reanneal RPA-covered ssDNA in vitro has been suggested to remove ssDNA, RPA and other ssDNA-binding proteins, as well as RNA from replicating chromosomes in vivo to prepare for mitosis97. Our observation that BLM interacts with TOP2A and RPA when BLM-NCAPH2 loses contact with SMC2, and BLM-TOP2A and BLM-RPA interactions are lost when BLM-NCAPH2 regains contact with SMC2, raises the possibility that BLM links functions of condensin II, RPA, and TOP2A during sister chromatid replication in S phase.
In summary, our study identifies a BLM-condensin II interaction in S phase and G2 phase that contributes to BLM association with nascent DNA and prevents genome instability. Furthermore, we provide insight into how condensin II promotes genome stability in interphase, and HR specifically, through interaction with a bona fide HR factor, BLM, contributing to crossover suppression and timely repair and survival of replication-dependent DSBs. Our study also uncovers a structural change in condensin II and its timing in S phase that might be linked to its functional state and to roles in the maintenance of genome stability outside of mitosis. We propose that condensin II targets BLM to sites where its substrates are most likely to occur during DNA replication, thereby increasing the efficiency by which BLM encounters and repairs its substrates in newly replicated sister chromatids prior to entry into mitosis.
Methods
Cell culture and cell lines
Human skin fibroblast cell line GM00637 from the NHGRI Sample Repository for Human Genetic Research at the Coriell Institute for Medical Research was used as the BLM-proficient wildtype cell line (WT). The isogenic BLM-KO cell line (KSVS1452) was generated by CRISPR/Cas9-mediated biallelic disruption of BLM exon 8 in GM0063741. The BLM-KO cell line was complemented with wildtype BLM cDNA by stable transfection of pcDNA3-BLM and selection of a clone with BLM expression similar to that of BLM in GM00637 to yield the BLM-COMP cell line (KSVS1454)41. For this study, cell lines expressing BLM-WMFmut were generated by stable transfection of BLM cDNA where the sequence for BLM residues 153DWDDMDDFD161 was mutated to 153DKDDKDDKD161. BLM-WMFmut expression was verified in multiple clones and two independent clones with BLM-WMFmut expression similar to BLM expression levels in the BLM-COMP cell line were selected for this study (KSBR1505, KSBR1506). All cell lines were grown in Modified Eagle’s Medium (Corning) supplemented with 10% Fetal Bovine Serum (Sigma-Aldrich), 2 mM L-glutamine and 1% penicillin–streptomycin (Corning) at 37 °C in the presence of 5% CO2.
Antibodies
Commercially available antibodies against the following proteins are as follows with the application and concentration used: β-actin (Santa Cruz, cat. no. sc-69879, WB, 1:5000), Biotin (Jackson Labs, cat. no. 200-002-211, Immunofluorescence (IF), 1:1000; Bethyl, cat. no. A150-109A, IF, 1:1000), BLM (Santa Cruz, cat. no. sc-365753, WB and IF, 1:200 and 1:250; Bethyl, cat. no. A300-110A, IP, 5 μg/mL; Abcam, cat no. ab2179, IF, 1:2000), BrdU (BD Biosciences, cat. no. 347580, IF, 1:50; Abcam, cat. no. ab6326, IF, 1:100), CREST (Antibodies Inc., cat. no. 15-234-0001, IF, 1:50), Cyclin A (Santa Cruz cat. no. sc-239, WB, 1:500), Cyclin B1 (Santa Cruz cat. no. sc-245, WB, 1:1000), Cyclin E (Santa Cruz cat. no. sc-377100, WB, 1:500), GAL4-TA (Santa Cruz, cat. no. sc-1663, WB, 1:200), γ-H2AX (Abcam, cat. no. ab81299, WB, 1:5000), Histone H2A (Invitrogen, cat. no. MA5-24662, WB, 1:1000), Histone H4 (Santa Cruz, cat. no. sc-25260, WB, 1:200), MCM6 (Santa Cruz, cat. no. sc-393618, WB, 1:200), NCAPD3 (Santa Cruz, cat. no. sc-81597, WB, 1:200), NCAPG (Santa Cruz, cat. no. sc-515297, WB, 1:200), NCAPH (Santa Cruz, cat. no. sc-101013, WB, 1:500; Bethyl cat. no. A300-603A, IP, 5 μg/mL), NCAPH2 (Santa Cruz, cat. no. sc-393333, WB and IF, 1:500 and 1:250; Bethyl cat. no. A302-275A, IP, 5 μg/mL), PCNA (Cell Signaling, cat. no. 2586, WB, 1:5000), p-DRP1 (Cell Signaling, cat. no. 4494S, WB, 1:500), p-Histone H3 (Cell Signaling, cat. no. 9701S, WB, 1:500), RAD51 (Santa Cruz, cat. no. sc-398587, WB, 1:200), RPA2 (Santa Cruz, cat. no. sc-56770, WB, 1:200), SMC2 (Bethyl, cat. no. A300-058A, IF and Immunoprecipitation (IP), 1:1000 and 5 μg/mL; Invitrogen, cat. no. GT4312, WB, 1:1000; Cell Signaling, cat. no. 5329, IP, 5 μg/mL; Abcam, cat. no. ab10412, IF, 1:250) SMC4 (Bethyl, cat. no. A300-063A, WB, 1:1000), TOP2A (Santa Cruz, cat. no. sc-365916, WB, 1:200), TOP3A (Abcam, cat. no. ab108493, WB, 1:1000).
Plasmid Transfection and RNAi
For transient transfections with plasmid DNA, cells at ~70-80% confluency were transfected using Lipofectamine3000 (Invitrogen) diluted in Opti-MEM (Gibco) according to the manufacturer’s instructions. After 72 h, cells were harvested and used for downstream applications. For stable transfections, cells were selected 48 h following transfection with either 750 μg/ml G418 sulfate (Invitrogen) or 1 μg/ml Puromycin (Gibco). Once single-cell colonies reached >100 cells, colonies were collected and grown to confluency. BLM-WMF mutation was confirmed by genomic DNA sequencing and BLM expression by Western blotting. For RNA interference of NCAPH and NCAPH2, cells at ~60-80% confluency were transfected with either NCAPH Silencer Select siRNA (ThermoFisher, siRNA ID. 136898) or NCAPH2 Silencer Select siRNA (ThermoFisher, siRNA ID. 44519) using RNAiMAX (Invitrogen) diluted in Opti-MEM reduced-serum media (Gibco) according to the manufacturer’s instructions and incubated for 72 h prior to downstream application. For the sister chromatid exchange and γ-H2AX elimination assays, two rounds of siRNA were performed. The second transfection was done 48 h following the first transfection and the assays were carried out 24 h following the second transfection. Silencer Select Negative Control No. 1 siRNA (ThermoFisher, cat. no. 4390843) was used as a negative control.
Double-thymidine block
Cells in S phase were obtained as previously described4 with minor modifications. Briefly, cells were arrested at the G1/S boundary with 2 mM thymidine for 24 h, released into drug-free media (14 h for BLM-WT and BLM-WMFmut and 22 h for BLM-KO to account for differences in doubling time identified in Supplementary Fig. S3b,c), shifted to 2 mM thymidine for 16 h, and released into drug-free media for downstream experiments.
Co-immunoprecipitation
Co-immunoprecipitation was performed using two different methods. For the first method98, nuclei were prepared by resuspending cell pellets on ice for 2 min in cytoplasmic lysis buffer [20 mM Tris pH 7.4, 10% Glycerol, 10 mM KCl, 0.2% NP-40, 2 μM EDTA, 1 mM PMSF, and protease inhibitor cocktail (Pierce)]. Nuclei were collected and lysed in nuclear lysis buffer [20 mM Tris pH 7.4, 10 mM KCl, 0.4 M NaCl, 10% glycerol, 1 μM EDTA, 1 mM PMSF, protease inhibitor cocktail (Pierce), phosphatase inhibitor cocktail (Pierce), and 125 U/ml benzonase (MilliporeSigma)] by incubating on ice for 30 min with occasional vortexing. Phosphatase inhibitor cocktail was excluded from phosphatase experiments. 1 mg of nuclear lysate was diluted with low-salt PBS to a final salt concentration of 150 mM prior to pre-clearing. Pre-cleared nuclear lysate was incubated with 30 μL of either Protein A or G agarose beads (Pierce) crosslinked to 5 μg of antibody overnight at 4 °C and washed with nuclear lysis buffer diluted to a final salt concentration of 150 mM with low-salt PBS. For the second method, whole-cell extracts were prepared by resuspending cell pellets in nuclear lysis buffer [20 mM Tris pH 7.4, 10 mM KCl, 0.4 M NaCl, 10% glycerol, 1 μM EDTA, 1 mM PMSF, protease inhibitor cocktail (Pierce), phosphatase inhibitor cocktail (Pierce), and 125 U/ml benzonase (MilliporeSigma)], lysing by Dounce homogenization, and incubating on ice for 30 min with occasional vortexing. 1.5 mg of whole-cell lysate was diluted with low-salt PBS to a volume of 1 ml and a final salt concentration of 150 mM prior to pre-clearing. Pre-cleared lysates were incubated with 5 μg of antibody overnight at 4 °C, incubated for 4 h at 4 °C with 30 μl of Protein G agarose beads, and washed extensively with nuclear lysis buffer. For experiments including lambda protein phosphatase, protein samples were treated with lambda protein phosphatase (New England BioLabs) per manufacturer’s protocol. Briefly, protein sample was treated with 10x NEBuffer for Protein MetalloPhosphatases (PMP) and 10 mM MnCl2 to a total volume of 50 μl. The volume was adjusted based on the volume required for 1.5 mg of protein. 1 μl of lambda protein phosphatase was added to the reaction per 50 μl and incubated at 30 °C for 30 min prior to pre-clearing and co-immunoprecipitation. Inputs are from the same lambda-protein-phosphatase treated lysates used for co-immunoprecipitation.
Liquid chromatography tandem mass spectrometry
As previously described4, BLM co-immunoprecipitates were eluted in 50 μl 0.2 M glycine, pH 2.6, and neutralized with 50 μl Tris, pH 8.0. Eluates were prepared for LC-MS/MS by FASP (Expedeon), followed by trypsin digestion (Promega), desalting and concentration, resuspended in 0.1% formic acid and analyzed on a Q Exactive Plus (Thermo). LC-MS/MS data were analyzed using MaxQuant99 and spectra compared to the UniProt reference proteome data set for Homo sapiens (Proteome ID: UP000005640). The data set was sorted to only include entries with ≤1 identified peptides in eluates from KSVS1452 (BLM-KO). The mass spectrometry proteomics data were deposited with the ProteomeXchange Consortium (http://proteomecentral.proteomexchange.org) via the PRIDE partner repository100 with the dataset identifier PXD018823.
Chromatin biochemical fractionation
Soluble and chromatin-bound fractions were obtained using an established protocol101,102. Briefly, cells were incubated in hypotonic lysis buffer (10 mM HEPES pH 7.9, 10 mM KCl, 1.5 mM, MgCl2, 0.34 M sucrose, 10% glycerol, 0.04% Triton X-100, 1 mM DTT, 1 mM PMSF, protease inhibitors) on ice for 10 min and soluble proteins were separated by centrifugation at 2000 × g for 4 min. The chromatin-enriched pellet was washed with low stringency buffer (3 mM EDTA, 0.2 mM EGTA, and 1 mM DTT) and centrifuged at 1600 × g for 4 min. Chromatin-bound proteins were extracted by incubating the pellet in ice cold RIPA buffer (50 mM Tris–HCl, pH 7.4, 1% NP-40, 0.1% sodium deoxycholate, 150 mM NaCl, 1 mM EDTA, 1 mM PMSF, and protease inhibitors) for 30 min on ice followed by centrifugation at 16000 × g.
Immunofluorescence microscopy
Cells grown directly on poly-L-lysine coated coverslips (Corning) were pre-extracted with 10 mM PIPES pH 6.8, 300 mM Sucrose, 100 mM NaCl, 1 mM MgCl2, 1 mM EGTA, 1 mM DTT, .05% Triton X-100, and protease inhibitor cocktail (Pierce) for 10 min at room temperature. Washed cells were fixed in 100% ice-cold methanol at room temperature for 10 min, blocked with 5% BSA in 0.1% PBS-Tween 20 for 30 min at room temperature, incubated with primary antibodies at 4 °C overnight, washed with PBS, and incubated with highly cross-adsorbed Alexa Fluor-conjugated secondary antibodies (Invitrogen) at room temperature for 1 h. For BLM-coated UFB detection, coverslips were processed using a well-established protocol103. Briefly, coverslips were gently washed with PBS prior to pre-extraction with Buffer A (0.2% Triton X-100, 10 mM PIPES pH 6.8, 1 mM MgCl2, and 10 mM EGTA) for 1 min before adding Buffer B (8% paraformaldehyde, 0.1% Triton X-100, 10 mM PIPES pH 6.8, 1 mM MgCl2, and 10 mM EGTA). Coverslips were incubated in Buffer A/B for 15 min at room temperature before being washed with PBS and permeabilized overnight at 4 °C in PBSAT (3% BSA, 0.5% Triton X-100 diluted in PBS). Coverslips were incubated in anti-BLM (Abcam, cat. no# ab2179, 1:200) diluted in PBSAT overnight at 4 °C, washed with PBSAT, incubated with highly cross-adsorbed Alexa Fluor-488 secondary antibody (Invitrogen) diluted in PBSAT at room temperature for 2 h, and finally washed with PBSAT. Coverslips were mounted to slides with Duolink® in situ mounting medium with DAPI (Sigma). Images were acquired using a Keyence BZ-X710 fluorescence microscope using a Plan-Apochromat λ 60× 1.4/0.13 mm Oil or Plan-Apochromat λ 100× 1.45/0.13 mm Oil objective. Images were analyzed and pseudocolored using ImageJ.
Proximity ligation and SIRF assay
Cells were prepared identically to immunofluorescence microscopy up until blocking. At this stage, the proximity ligation assay was performed using the Duolink In Situ Red Mouse/Rabbit kit (Sigma) following the manufacturer’s instructions. For SIRF69, prior to pre-extraction, cells were labeled with 125 μM EdU (Invitrogen) for 12, 13.5, and 15 min for the BLM-COMP, BLM-WMFmut, and BLM-KO, respectively. The labeling times for the BLM-COMP, BLM-KO, and BLM-WMFmut were determined based on their differences in median replication speed where the BLM-KO is ~28% slower than the BLM-WT and the BLM-WMFmut is ~14% slower. The BLM-KO was labeled for 25% longer and BLM-WMFmut for 12.5% longer than WT for simplicity and to avoid over-labeling. Similar labeling efficiency was confirmed by EdU-EdU SIRF (Supplementary Fig. S5a). Following EdU labeling, Click-iT reaction cocktail was freshly prepared in the order as listed: 2 mM copper sulfate, 9 μM biotin-azide (Invitrogen), 1 μM Alexa Fluor 488-azide (Invitrogen), and 100 mM sodium ascorbate diluted in PBS and added to coverslips and incubated at room temperature for 1 h before proceeding with the proximity ligation assay. Images were acquired using a Keyence BZ-X710 fluorescence microscope using a Plan-Apochromat_λ 60× 1.4/0.13 mm Oil objective. Images were analyzed and pseudocolored using ImageJ.
Mammalian two-hybrid analysis
Full length human BLM and SMC2 cDNAs were cloned into the pVP16 and pM vectors, respectively (Clontech). Fragments and point mutants generated by site-directed mutagenesis in pVP16-BLM and pM-SMC2 were verified by DNA sequencing. Secreted alkaline phosphatase (SEAP) activity expressed from the pG5SEAP reporter vector was determined 72 h post-transfection using the Great EscAPe SEAP Fluorescence Detection Kit (Clontech). All co-transfections were performed in triplicate. Fluorescence was measured using a BioTek Synergy 2 plate reader with Gen5 version 3.09 software.
Flow cytometry
For cell cycle analysis, asynchronous or cells synchronized and released from double-thymidine were harvested, washed twice with sample buffer (0.001% glucose in PBS), resuspended at a concentration of 106 cells/ml, fixed in 70% ethanol at 4 °C overnight, washed twice with sample buffer, and stained with 50 μg/ml propidium iodide (ThermoFisher) in sample buffer with 100 Kunitz units/ml RNase A (Fisher) for 1 h at room temperature. Fluorescence was measured by a BD FACSCantoII flow cytometer and analyzed using FlowJo v.10.10 software (BD Life Sciences, https://www.flowjo.com/solutions/ flowjo/downloads).
Cell doubling time
Cells were seeded at a density of 5000 cells/well of a 6-well plate and counted every day for 12 consecutive days using a Countess II automated cell counter (Invitrogen). Doubling time was calculated in log phase for each cell line tested using the freely available Doubling Time Calculator software (Roth V. 2006 Doubling Time Computing, available from: https://doubling-time.com/compute.php).
DNA fiber labeling and analysis
Single-molecule DNA fiber analysis was performed as previously described104. For replication speed analysis and fork asymmetry, cells at 50-80% confluency were pulsed with 20 μM CldU (Sigma) and 100 μM IdU (Sigma) for 30 min each. To assess fork recovery following replication stress, cells were pulsed with CldU for 30 min, washed, released into media containing 4 mM hydroxyurea (US Biological) for 4 h, washed, and then pulsed with IdU for 45 min. Following the treatments, cells were counted, harvested, and lysed on slides. DNA was then manually stretched by tilting. Slides were fixed for 2 min with 3:1 (vol/vol) methanol-acetic acid, air dried, and stored at -20 °C overnight. DNA was denatured by incubating in 2.5 N HCl for 70 min. Slides were blocked with 10% goat serum diluted in 0.1% PBS-Tween 20 for 1 h, incubated with anti-BrdU (BD Biosciences, Abcam) for 2 h and with anti-mouse Alexa Fluor 488 and anti-rat Alexa Fluor 594 for 1 h at room temperature. Images were acquired using a Keyence BZ-X710 fluorescence microscope using a Plan-Apochromat_λ 60× 1.4/0.13 mm Oil or Plan-Apochromat_λ 100× 1.45/0.13 mm Oil objective. Fiber lengths were analyzed using ImageJ.
Clonogenic survival assay
Cells were seeded at a density of 500 cells/60 mm dish for colony formation, media replaced with fresh media supplemented with varying concentrations of either hydroxyurea (US Biological) or camptothecin (ThermoFisher) for 48 h, washed with PBS, and media replaced with drug-free complete growth media to allow colonies to grow for up to 3 weeks. Colonies were fixed in 3:1 (vol/vol) methanol-acetic acid and stained with 0.5% crystal violet (Fisher) dissolved in 3:1 (vol/vol) methanol-acetic acid solution. Colonies with more than 50 cells were manually scored as survivors using three biological replicates for each cell line. Experiments were performed at least twice with similar results.
Sister chromatid exchange assay
Sister chromatid exchanges were detected in metaphase chromosomes by immunofluorescence using established protocols with minor variation61,105. Briefly, exponentially growing cells were labeled for two cell cycles based on doubling time (siNCAPH2 treated cells were labeled for the same time as BLM-WMFmut) with 10 μM BrdU (Invitrogen) before being treated with 0.1 μg/ml Colcemid (Invitrogen) for 1 h to induce metaphase arrest. Chromosome spreads were prepared by hypotonic treatment with 75 mM KCl for 1 h at 37 °C and then fixed with 3:1 (vol/vol) methanol-acetic acid before being dropped onto glass microscopy slides. To detect BrdU-incorporated DNA, slides were incubated in 2 N HCl for 10 min at room temperature, washed with PBS, blocked, incubated with anti-BrdU (BD Biosciences) at 4 °C overnight, washed with PBS, and incubated with anti-mouse Alex Fluor 488 (Invitrogen) at room temperature for 1 h. Slides were washed with PBS and mounted to coverslips with Duolink® in situ mounting medium with DAPI (Sigma). Images were acquired using a Keyence BZ-X710 fluorescence microscope using a Plan-Apochromat_λ 100× 1.45/0.13 mm Oil objective.
γ-H2AX elimination assay
Exponentially growing cells were exposed to 0.5 μM camptothecin (ThermoFisher) for 1 h, released into fresh media, and harvested at various time points. To detect γ-H2AX, histone extraction from chromatin was performed as previously described98 using 0.5 M HCl, 10% glycerol, 100 mM β-mercaptoethanol and then neutralized using 1.3 M NaOH in 40 mM Tris pH 7.4 with protease inhibitor cocktail (Pierce). Histones were separated on a 16% Tris-Tricine gel prior to Western blotting. Each experiment was performed in triplicate and quantified.
Premature chromosome condensation (PCC) assay
The PCC assay was performed as previously described40 with minor modifications. Untreated cells and those subjected to RNAi were labeled with 10 μM EdU (Invitrogen) for 30 min, washed, treated with 50 nM calyculin A (Sigma) for 1 h to induce chromosome condensation, and chromosome spreads prepared identically to the sister chromatid exchange assay. After spreading, slides were washed, fixed with 100% ice-cold methanol for 10 min at room temperature, washed, and incubated in fresh Click-iT reaction cocktail [2 mM copper sulfate, 10 μM biotin-azide (Invitrogen), 100 mM sodium ascorbate] for 1 h at room temperature. Slides were washed with PBS, incubated with either anti-biotin (Bethyl) or anti-biotin (Jackson Labs) and anti-SMC2 (Bethyl) at 4 °C overnight, washed with PBS, and incubated with anti-mouse Alex Fluor 488 and anti-rabbit Alexa Fluor 555 (Invitrogen) at room temperature for 1 h. Coverslips were washed with PBS and mounted to slides with Duolink® in situ mounting medium with DAPI (Sigma). Images were acquired using a Keyence BZ-X710 fluorescence microscope using a Plan-Apochromat_λ 100× 1.45/0.13 mm Oil objective.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The authors affirm that all data necessary for confirming the conclusions of the article are present within the article, figures, and tables. Cell lines are available upon request. The mass spectrometry proteomics data were previously published4 and are available through the ProteomeXchange Consortium via the PRIDE100 partner repository with the dataset identifier PXD018823. Supplementary Table S1 lists the top hits identified by mass spectrometry proteomics. An example of the gating strategy used to generate Supplementary Fig. S3a can be found in Supplementary Fig. S6. Unprocessed Western blot images can be found in Supplementary Fig. S7–14. Data sets in Excel format are included in Supplementary Data 1 and 2.
References
Ellis, N. A. et al. The Bloom’s syndrome gene product is homologous to RecQ helicases. Cell 83, 655–666 (1995).
German, J. Bloom’s syndrome. I. Genetical and clinical observations in the first twenty-seven patients. Am. J. Hum. Genet. 21, 196–227 (1969).
Cunniff, C., Bassetti, J. A. & Ellis, N. A. Bloom’s Syndrome: clinical spectrum, molecular pathogenesis, and cancer predisposition. Mol. Syndromol. 8, 4–23 (2017).
Shastri, V. M., Subramanian, V. & Schmidt, K. H. A novel cell-cycle-regulated interaction of the Bloom syndrome helicase BLM with Mcm6 controls replication-linked processes. Nucleic Acids Res. 49, 8699–8713 (2021).
Rao, V. A. et al. Endogenous gamma-H2AX-ATM-Chk2 checkpoint activation in Bloom’s syndrome helicase deficient cells is related to DNA replication arrested forks. Mol. Cancer Res. 5, 713–724 (2007).
Hand, R. & German, J. A retarded rate of DNA chain growth in Bloom’s syndrome. Proc. Natl Acad. Sci. USA 72, 758–762 (1975).
Chaganti, R. S., Schonberg, S. & German, J. A manyfold increase in sister chromatid exchanges in Bloom’s syndrome lymphocytes. Proc. Natl Acad. Sci. USA 71, 4508–4512 (1974).
Davies, S. L., North, P. S. & Hickson, I. D. Role for BLM in replication-fork restart and suppression of origin firing after replicative stress. Nat. Struct. Mol. Biol. 14, 677–679 (2007).
Shastri, V. M. & Schmidt, K. H. Cellular defects caused by hypomorphic variants of the Bloom syndrome helicase gene BLM. Mol. Genet. Genom. Med 4, 106–119 (2016).
Rao, V. A. et al. Phosphorylation of BLM, dissociation from topoisomerase IIIalpha, and colocalization with gamma-H2AX after topoisomerase I-induced replication damage. Mol. Cell Biol. 25, 8925–8937 (2005).
Chang, E. Y. et al. RECQ-like helicases Sgs1 and BLM regulate R-loop-associated genome instability. J. Cell Biol. 216, 3991–4005 (2017).
Xue, C. et al. Single-molecule visualization of human BLM helicase as it acts upon double- and single-stranded DNA substrates. Nucleic Acids Res. 47, 11225–11237 (2019).
Kang, D. et al. Interaction of replication protein A with two acidic peptides from human Bloom syndrome protein. FEBS Lett. 592, 547–558 (2018).
Brosh, R. M. Jr et al. Replication protein A physically interacts with the Bloom’s syndrome protein and stimulates its helicase activity. J. Biol. Chem. 275, 23500–23508 (2000).
Kaur, E., Agrawal, R. & Sengupta, S. Functions of BLM helicase in cells: is it acting like a double-edged sword? Front. Genet. 12, 634789 (2021).
Vindigni, A. & Hickson, I. D. RecQ helicases: multiple structures for multiple functions? HFSP J. 3, 153–164 (2009).
Karow, J. K., Constantinou, A., Li, J. L., West, S. C. & Hickson, I. D. The Bloom’s syndrome gene product promotes branch migration of holliday junctions. Proc. Natl Acad. Sci. USA 97, 6504–6508 (2000).
Wu, L. & Hickson, I. D. The Bloom’s syndrome helicase suppresses crossing over during homologous recombination. Nature 426, 870–874 (2003).
Shorrocks, A. K. et al. The Bloom syndrome complex senses RPA-coated single-stranded DNA to restart stalled replication forks. Nat. Commun. 12, 585 (2021).
Wu, L., Davies, S. L., Levitt, N. C. & Hickson, I. D. Potential role for the BLM helicase in recombinational repair via a conserved interaction with RAD51. J. Biol. Chem. 276, 19375–19381 (2001).
Nimonkar, A. V. et al. BLM-DNA2-RPA-MRN and EXO1-BLM-RPA-MRN constitute two DNA end resection machineries for human DNA break repair. Genes Dev. 25, 350–362 (2011).
Langland, G. et al. The Bloom’s syndrome protein (BLM) interacts with MLH1 but is not required for DNA mismatch repair. J. Biol. Chem. 276, 30031–30035 (2001).
Sharma, S. et al. Stimulation of flap endonuclease-1 by the Bloom’s syndrome protein. J. Biol. Chem. 279, 9847–9856 (2004).
Wang, J., Chen, J. & Gong, Z. TopBP1 controls BLM protein level to maintain genome stability. Mol. Cell 52, 667–678 (2013).
Bhattacharyya, S. et al. Telomerase-associated protein 1, HSP90, and topoisomerase IIalpha associate directly with the BLM helicase in immortalized cells using ALT and modulate its helicase activity using telomeric DNA substrates. J. Biol. Chem. 284, 14966–14977 (2009).
Selak, N. et al. The Bloom’s syndrome helicase (BLM) interacts physically and functionally with p12, the smallest subunit of human DNA polymerase delta. Nucleic Acids Res. 36, 5166–5179 (2008).
Singh, D. K. et al. The human RecQ helicases BLM and RECQL4 cooperate to preserve genome stability. Nucleic Acids Res. 40, 6632–6648 (2012).
von Kobbe, C. et al. Colocalization, physical, and functional interaction between Werner and Bloom syndrome proteins. J. Biol. Chem. 277, 22035–22044 (2002).
Ono, T., Fang, Y., Spector, D. L. & Hirano, T. Spatial and temporal regulation of Condensins I and II in mitotic chromosome assembly in human cells. Mol. Biol. Cell 15, 3296–3308 (2004).
Green, L. C. et al. Contrasting roles of condensin I and condensin II in mitotic chromosome formation. J. Cell Sci. 125, 1591–1604 (2012).
Gibcus, J. H. et al. A pathway for mitotic chromosome formation. Science 359 (2018).
Cutts, E. E. & Vannini, A. Condensin complexes: understanding loop extrusion one conformational change at a time. Biochem. Soc. Trans. 48, 2089–2100 (2020).
Martin, C. A. et al. Mutations in genes encoding condensin complex proteins cause microcephaly through decatenation failure at mitosis. Genes Dev. 30, 2158–2172 (2016).
Heale, J. T. et al. Condensin I interacts with the PARP-1-XRCC1 complex and functions in DNA single-strand break repair. Mol. Cell 21, 837–848 (2006).
Kong, X. et al. Condensin I recruitment to base damage-enriched DNA lesions is modulated by PARP1. PLoS ONE6, e23548 (2011).
Wood, J. L., Liang, Y., Li, K. & Chen, J. Microcephalin/MCPH1 associates with the Condensin II complex to function in homologous recombination repair. J. Biol. Chem. 283, 29586–29592 (2008).
Rosin, L. F., Nguyen, S. C. & Joyce, E. F. Condensin II drives large-scale folding and spatial partitioning of interphase chromosomes in Drosophila nuclei. PLoS Genet. 14, e1007393 (2018).
Wallace, H. A., Klebba, J. E., Kusch, T., Rogers, G. C. & Bosco, G. Condensin II regulates interphase chromatin organization through the Mrg-binding motif of cap-H2. G3 (Bethesda) 5, 803–817 (2015).
Bauer, C. R., Hartl, T. A. & Bosco, G. Condensin II promotes the formation of chromosome territories by inducing axial compaction of polyploid interphase chromosomes. PLoS Genet. 8, e1002873 (2012).
Ono, T., Yamashita, D. & Hirano, T. Condensin II initiates sister chromatid resolution during S phase. J. Cell Biol. 200, 429–441 (2013).
Subramanian, V. et al. Bloom syndrome DNA helicase deficiency is associated with oxidative stress and mitochondrial network changes. Sci. Rep. 11, 2157 (2021).
Ono, T. et al. Differential contributions of condensin I and condensin II to mitotic chromosome architecture in vertebrate cells. Cell 115, 109–121 (2003).
Li, W. et al. Condensin I and II Complexes License Full Estrogen Receptor alpha-Dependent Enhancer Activation. Mol. Cell 59, 188–202 (2015).
Mirzaei, H., Syed, S., Kennedy, J. & Schmidt, K. H. Sgs1 truncations induce genome rearrangements but suppress detrimental effects of BLM overexpression in Saccharomyces cerevisiae. J. Mol. Biol. 405, 877–891 (2011).
Wu, L. et al. The Bloom’s syndrome gene product interacts with topoisomerase III. J. Biol. Chem. 275, 9636–9644 (2000).
Russell, B. et al. Chromosome breakage is regulated by the interaction of the BLM helicase and topoisomerase IIalpha. Cancer Res. 71, 561–571 (2011).
Blackford, A. N. et al. TopBP1 interacts with BLM to maintain genome stability but is dispensable for preventing BLM degradation. Mol. Cell 57, 1133–1141 (2015).
Kaur, S. et al. Chk1-dependent constitutive phosphorylation of BLM helicase at serine 646 decreases after DNA damage. Mol. Cancer Res. 8, 1234–1247 (2010).
Takemoto, A., Kimura, K., Yokoyama, S. & Hanaoka, F. Cell cycle-dependent phosphorylation, nuclear localization, and activation of human condensin. J. Biol. Chem. 279, 4551–4559 (2004).
Hassler, M. et al. Structural Basis of an Asymmetric Condensin ATPase Cycle. Mol. Cell 74, 1175–1188 e9 (2019).
Houlard, M. et al. MCPH1 inhibits Condensin II during interphase by regulating its SMC2-Kleisin interface. Elife 10, e73348 (2021).
Dyson, S., Segura, J., Martinez-Garcia, B., Valdes, A. & Roca, J. Condensin minimizes topoisomerase II-mediated entanglements of DNA in vivo. EMBO J. 40, e105393 (2021).
Cuvier, O. & Hirano, T. A role of topoisomerase II in linking DNA replication to chromosome condensation. J. Cell Biol. 160, 645–655 (2003).
Morao, A. K., Kim, J., Obaji, D., Sun, S. & Ercan, S. Topoisomerases I and II facilitate condensin DC translocation to organize and repress X chromosomes in C. elegans. Mol. Cell 82, 4202–4217 e5 (2022).
Akai, Y. et al. Opposing role of condensin hinge against replication protein A in mitosis and interphase through promoting DNA annealing. Open Biol. 1, 110023 (2011).
Hirano, M. & Hirano, T. Opening closed arms: long-distance activation of SMC ATPase by hinge-DNA interactions. Mol. Cell 21, 175–186 (2006).
Griese, J. J., Witte, G. & Hopfner, K. P. Structure and DNA binding activity of the mouse condensin hinge domain highlight common and diverse features of SMC proteins. Nucleic Acids Res. 38, 3454–3465 (2010).
Davalos, A. R. & Campisi, J. Bloom syndrome cells undergo p53-dependent apoptosis and delayed assembly of BRCA1 and NBS1 repair complexes at stalled replication forks. J. Cell Biol. 162, 1197–1209 (2003).
Sidorova, J. M., Kehrli, K., Mao, F. & Monnat, R. Jr Distinct functions of human RECQ helicases WRN and BLM in replication fork recovery and progression after hydroxyurea-induced stalling. DNA Repair (Amst.) 12, 128–139 (2013).
Wu, L. & Hickson, I. D. Role of the BLM helicase in replication fork management. DNA Repair (Amst.) 6, 936–944 (2007).
Waisertreiger, I., Popovich, K., Block, M., Anderson, K. R. & Barlow, J. H. Visualizing locus-specific sister chromatid exchange reveals differential patterns of replication stress-induced fragile site breakage. Oncogene 39, 1260–1272 (2020).
Frorath, B., Schmidt-Preuss, U., Siemers, U., Zollner, M. & Rudiger, H. W. Heterozygous carriers for Bloom syndrome exhibit a spontaneously increased micronucleus formation in cultured fibroblasts. Hum. Genet. 67, 52–55 (1984).
Chan, K. L., North, P. S. & Hickson, I. D. BLM is required for faithful chromosome segregation and its localization defines a class of ultrafine anaphase bridges. Embo J. 26, 3397–3409 (2007).
Rosin, M. P. & German, J. Evidence for chromosome instability in vivo in Bloom syndrome: increased numbers of micronuclei in exfoliated cells. Hum. Genet. 71, 187–191 (1985).
Khan, T. N. et al. Mutations in NCAPG2 Cause a Severe Neurodevelopmental Syndrome that Expands the Phenotypic Spectrum of Condensinopathies. Am. J. Hum. Genet. 104, 94–111 (2019).
Krupina, K., Goginashvili, A. & Cleveland, D. W. Causes and consequences of micronuclei. Curr. Opin. Cell Biol. 70, 91–99 (2021).
Degrassi, F. et al. CREST staining of micronuclei from free-living rodents to detect environmental contamination in situ. Mutagenesis 14, 391–396 (1999).
Vazquez-Diez, C., Yamagata, K., Trivedi, S., Haverfield, J. & FitzHarris, G. Micronucleus formation causes perpetual unilateral chromosome inheritance in mouse embryos. Proc. Natl Acad. Sci. USA 113, 626–631 (2016).
Roy, S. & Schlacher, K. SIRF: A Single-cell Assay for in situ Protein Interaction with Nascent DNA Replication Forks. Bio Protoc. 9, e3377 (2019).
Higashi, T. L. & Uhlmann, F. SMC complexes: Lifting the lid on loop extrusion. Curr. Opin. Cell Biol. 74, 13–22 (2022).
Xu, X. & Yanagida, M. Suppressor screening reveals common kleisin-hinge interaction in condensin and cohesin, but different modes of regulation. Proc. Natl Acad. Sci. USA 116, 10889–10898 (2019).
Lee, B. G. et al. Cryo-EM structures of holo condensin reveal a subunit flip-flop mechanism. Nat. Struct. Mol. Biol. 27, 743–751 (2020).
Janissen, R., Barth, R., Davidson, I. F., Peters, J. M. & Dekker, C. All eukaryotic SMC proteins induce a twist of -0.6 at each DNA loop extrusion step. Sci. Adv. 10, eadt1832 (2024).
Dekker, C., Haering, C. H., Peters, J. M. & Rowland, B. D. How do molecular motors fold the genome? Science 382, 646–648 (2023).
Pradhan, B. et al. SMC complexes can traverse physical roadblocks bigger than their ring size. Cell Rep. 41, 111491 (2022).
Woods, C. G., Bond, J. & Enard, W. Autosomal recessive primary microcephaly (MCPH): a review of clinical, molecular, and evolutionary findings. Am. J. Hum. Genet. 76, 717–728 (2005).
Langer, K., Cunniff, C. M. & Kucine, N. Bloom Syndrome. In GeneReviews® (eds Adam, M. P. et al.) (University of Washington, 1993).
Gonenc, I. I. et al. Single-cell transcription profiles in Bloom syndrome patients link BLM deficiency with altered condensin complex expression signatures. Hum. Mol. Genet. 31, 2185–2193 (2022).
Saharia, A. et al. FEN1 ensures telomere stability by facilitating replication fork re-initiation. J. Biol. Chem. 285, 27057–27066 (2010).
Petermann, E., Orta, M. L., Issaeva, N., Schultz, N. & Helleday, T. Hydroxyurea-stalled replication forks become progressively inactivated and require two different RAD51-mediated pathways for restart and repair. Mol. Cell 37, 492–502 (2010).
Shintomi, K. & Hirano, T. The relative ratio of condensin I to II determines chromosome shapes. Genes Dev. 25, 1464–1469 (2011).
Baxter, J. et al. Positive supercoiling of mitotic DNA drives decatenation by topoisomerase II in eukaryotes. Science 331, 1328–1332 (2011).
Charbin, A., Bouchoux, C. & Uhlmann, F. Condensin aids sister chromatid decatenation by topoisomerase II. Nucleic Acids Res. 42, 340–348 (2014).
Yankiwski, V., Marciniak, R. A., Guarente, L. & Neff, N. F. Nuclear structure in normal and Bloom syndrome cells. Proc. Natl Acad. Sci. USA 97, 5214–5219 (2000).
Addis Jones, O., Tiwari, A., Olukoga, T., Herbert, A. & Chan, K. L. PLK1 facilitates chromosome biorientation by suppressing centromere disintegration driven by BLM-mediated unwinding and spindle pulling. Nat. Commun. 10, 2861 (2019).
Rouzeau, S. et al. Bloom’s syndrome and PICH helicases cooperate with topoisomerase IIalpha in centromere disjunction before anaphase. PLoS ONE7, e33905 (2012).
Stephens, A. D., Haase, J., Vicci, L., Taylor, R. M. 2nd & Bloom, K. Cohesin, condensin, and the intramolecular centromere loop together generate the mitotic chromatin spring. J. Cell Biol. 193, 1167–1180 (2011).
Sen Gupta, A. et al. Defining a core configuration for human centromeres during mitosis. Nat. Commun. 14, 7947 (2023).
Wallace, H. A., Rana, V., Nguyen, H. Q. & Bosco, G. Condensin II subunit NCAPH2 associates with shelterin protein TRF1 and is required for telomere stability. J. Cell Physiol. 234, 20755–20768 (2019).
Barefield, C. & Karlseder, J. The BLM helicase contributes to telomere maintenance through processing of late-replicating intermediate structures. Nucleic Acids Res. 40, 7358–7367 (2012).
Lu, H. et al. RECQL4 Promotes DNA End Resection in Repair of DNA Double-Strand Breaks. Cell Rep. 16, 161–173 (2016).
Daniloski, Z., Bisht, K. K., McStay, B. & Smith, S. Resolution of human ribosomal DNA occurs in anaphase, dependent on tankyrase 1, condensin II, and topoisomerase IIalpha. Genes Dev. 33, 276–281 (2019).
Hashash, N., Johnson, A. L. & Cha, R. S. Topoisomerase II- and condensin-dependent breakage of MEC1ATR-sensitive fragile sites occurs independently of spindle tension, anaphase, or cytokinesis. PLoS Genet. 8, e1002978 (2012).
Baxter, J. & Aragon, L. A model for chromosome condensation based on the interplay between condensin and topoisomerase II. Trends Genet 28, 110–117 (2012).
Coelho, P. A., Queiroz-Machado, J. & Sunkel, C. E. Condensin-dependent localisation of topoisomerase II to an axial chromosomal structure is required for sister chromatid resolution during mitosis. J. Cell Sci. 116, 4763–4776 (2003).
Samejima, K. et al. Mitotic chromosomes are compacted laterally by KIF4 and condensin and axially by topoisomerase IIalpha. J. Cell Biol. 199, 755–770 (2012).
Hirano, T. Condensins: universal organizers of chromosomes with diverse functions. Genes Dev. 26, 1659–1678 (2012).
Rios-Doria, J. et al. Ectopic expression of histone H2AX mutants reveals a role for its post-translational modifications. Cancer Biol. Ther. 8, 422–434 (2009).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Vizcaino, J. A. et al. The PRoteomics IDEntifications (PRIDE) database and associated tools: status in 2013. Nucleic Acids Res. 41, D1063–D1069 (2013).
Sakwe, A. M., Nguyen, T., Athanasopoulos, V., Shire, K. & Frappier, L. Identification and characterization of a novel component of the human minichromosome maintenance complex. Mol. Cell Biol. 27, 3044–3055 (2007).
Ritzi, M. et al. Complex protein-DNA dynamics at the latent origin of DNA replication of Epstein-Barr virus. J. Cell Sci. 116, 3971–3984 (2003).
Bizard, A. H., Nielsen, C. F. & Hickson, I. D. Detection of Ultrafine Anaphase Bridges. Methods Mol. Biol. 1672, 495–508 (2018).
Couch, F. B. et al. ATR phosphorylates SMARCAL1 to prevent replication fork collapse. Genes Dev. 27, 1610–1623 (2013).
Perry, P. & Wolff, S. New Giemsa method for the differential staining of sister chromatids. Nature 251, 156–158 (1974).
Lee, B. G., Rhodes, J. & Lowe, J. Clamping of DNA shuts the condensin neck gate. Proc. Natl Acad. Sci. USA 119, e2120006119 (2022).
Acknowledgements
The study was supported by National Institutes of Health grant R01GM139296 to K.H.S.
Author information
Authors and Affiliations
Contributions
B.R., G.K., and V.S. conducted the experiments; B.R. and K.H.S. designed the study, analyzed and interpreted data; and B.R and K.H.S. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editors: Tiago Dantas and Mengtan Xing.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Rodemoyer, B., Kariyawasam, G., Subramanian, V. et al. Condensin II interacts with BLM helicase in S phase to maintain genome stability. Commun Biol 8, 492 (2025). https://doi.org/10.1038/s42003-025-07916-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-025-07916-0