Abstract
Recent research has shown the gut microbiome’s impact on memory, yet limitations hinder the identification of specific microbes linked to cognitive function. We measured spatial working memory in individual mice before and after fecal microbiota transplantation (FMT) to develop a targeted analysis that identifies memory-associated strains while minimizing host genetic effects. Transplantation of human fecal into C57BL/6 mice yielded varied outcomes: some mice showed significant improvements while others had negligible changes, indicating that these changes are due to differences in FMT colonization. Metagenomic analysis, stratified by memory performance, revealed a positive correlation between the abundance of Akkermansia muciniphila and improved memory. Moreover, administering two A. muciniphila strains, GMB 0476 and GMB 2066, to wild-type mice elevated spatial working memory via BDNF activation. Our findings indicate that specific gut microbes, particularly A. muciniphila, may modulate memory and represent potential targets for therapeutic intervention in cognitive enhancement.
Similar content being viewed by others
Introduction
The human gut microbiome, predominantly comprised of bacteria but also including protozoa, archaea, eukaryotes, and viruses, forms the most intricate microbial community within our body1. Residing symbiotically in the gastrointestinal (GI) tract, these microbes crucially influence a myriad of human phenotypes and diseases, ranging from gastrointestinal disorders like Crohn’s disease and ulcerative colitis2,3, to systemic conditions such as obesity4, diabetes5, and even neurological concerns like depression and neurodegenerative diseases6,7,8,9. As a result, the gut microbiome and its role in human phenotypes and diseases have garnered significant attention in recent times. Despite its paramount importance akin to our genome, efforts to pin down specific gut microbes linked to diseases or phenotypes haven’t progressed as anticipated. The challenge in identifying specific gut microbes linked to diseases and phenotypes at the species level underscores the need for a novel approach to gut microbiome analysis. The vast number of intestinal microbes, comprising around 100 trillion, poses challenges not only in terms of analysis but also in generating various degrees of noisy data that can lead to misleading conclusions10,11,12.
Apart from the complexity of metagenomics data, recent evidence has revealed intrinsic problems with current gut microbiome analysis. The composition of the gut microbiome can vary significantly depending on the location within the GI tract13. Furthermore, the composition of the whole gut microbiome differs substantially from that of the fecal microbiome14. The influence of host genetic backgrounds further complicates the identification of intestinal microbes associated with diseases or phenotypes15,16,17. Host genetic backgrounds have a more significant impact on the gut microbiome than dietary shifts18, making it challenging to establish causal relationships during the analysis of the gut microbiome.
Aging is widely recognized as a major contributor to cognitive decline, including impairments in memory and learning abilities19. This decline has been attributed to several biological mechanisms, such as increased neuroinflammation, oxidative stress, synaptic deterioration, and dysregulation of neurotransmitter systems. In particular, aging leads to structural and functional changes in the hippocampus, a key brain region involved in spatial working memory20. Additionally, age-related alterations in gut microbiota composition have been observed, including a reduction in microbial diversity and beneficial taxa, which may further exacerbate neuroinflammatory processes and cognitive dysfunction21.
Recent studies suggest that the gut microbiome may also play a role in cognitive functions, particularly spatial working memory, a cognitive process critical for navigation and decision-making22. While the influence of genetics, nutrition, and environmental factors on spatial working memory is well documented, the role of the gut microbiome remains underexplored23. Despite growing evidence, direct causal links between specific gut microbes and spatial working memory remain poorly understood. Identifying the precise microbial contributors to cognitive function and their underlying mechanisms is essential for advancing our understanding of the gut–brain axis.
To address this, fecal microbiota transplantation (FMT) has been utilized as a tool to investigate the causal relationship between gut microbes and host phenotypes24. By transferring the gut microbiota from one organism to another, FMT enables researchers to assess how microbial communities influence specific physiological and cognitive traits24. In this study, we used naturally aged C57BL/6 mice as a model to identify gut microbes that may help mitigate age-related cognitive decline, particularly in spatial working memory. Through this approach, we identified Akkermansia muciniphila as a potential key player associated with enhanced spatial working memory. Our findings suggest that Akkermansia muciniphila may counteract these aging-related effects and help support spatial working memory.
Building on these findings, we further investigated whether the administration of Akkermansia muciniphila could enhance spatial working memory. Our results indicate that supplementation with Akkermansia muciniphila improves spatial working memory, providing new insights into the relationship between the gut microbiome and cognitive function. This study underscores the importance of considering gut microbial composition in age-related cognitive decline and highlights potential therapeutic applications of microbiome-based interventions in cognitive health.
Results
FMT of human stool into C57BL/6 mice resulted in diverse effects on spatial working memory
In order to find the gut microbes responsible for regulating the spatial working memory (SWM), a mixture of human gut microbiome was transplanted into naturally aged 12-month-old C57BL/6 mice where their gut microbiome was depleted by administration of a mixture of three broad-spectrum antibiotics and nystatin (Fig. 1a). FMT was then performed for 3 months to ensure that the diversity of the gut microbiota increased and sufficient changes in the host phenotype were observed25,26. We compared changes in SWM in the same mice before and after FMT to evaluate precisely the influence of the gut microbiome on SWM while excluding genetic factors. The differences in SWM of each mouse before FMT reflect the differences in the intelligent characteristics of individual mice. We checked the degree of change in SWM of each mouse after 3 months of FMT to confirm changes in intelligence caused by human intestinal microorganisms, excluding the intelligence of individual mice. Over the three-month experimental period, the changes in SWM could be categorized into three groups: the positive group (PI; n = 10; % Alternation change = 17.6 ± 3.7%, 10 females and 10 males), the ineffective group (II; n = 10; % Alternation change = 0.3 ± 0.9%, 7 females and 3 males), and the negative group (NI; n = 10; % Alternation change = −14.6 ± 1.1%, 6 females and 4 males) (Fig. 1b, c). It should be noted that the original SWM of the PI group (50.2 ± 3.0% at 0 months) was the lowest, while that of the NI group (71.9 ± 1.9% at 0 months) was the highest (Fig. 1d). These differences may result from individual factors, including genetic factors of individual mice. The inverse relationships after FMT, the PI group (67.8 ± 3.1% at 3 months) and NI group (57.3 ± 1.1% at 3 months), signify that the transplanted gut microbiome plays an important role in determining SWM.
Change in mouse spatial working memory following FMT. a Schematic representation of the experimental strategy. b Comparison of spatial working memory in individual mice before and after FMT. c Alteration in spatial working memory in individual mice after FMT. d Correlation between changes in spatial working memory and FMT. The values represent mean ± S.E.M. 0 m, before FMT; 3 m, 3 months after FMT; PI, mouse group with increased spatial working memory after FMT; II, mouse group with unchanged spatial working memory after FMT; NI, mouse group with decreased spatial working memory after FMT. **p < 0.01; ***p < 0.001.
During the experimental period, no significant group differences were observed in the changes in body weight and blood profile (Supplementary Fig. 1). However, it is worth noting that across all three mouse groups, there were general trends observed in the body weight and blood profile. Specifically, body weight, blood glucose levels, LDL-cholesterol levels, and triglyceride levels increased, while HDL-cholesterol levels decreased during the experimental period (Supplementary Fig. 1). These changes reflect the natural ageing process of the mice over the course of the experiment.
FMT altered the composition of the gut microbiome
To determine whether FMT altered the gut microbiome of mice, we first analyzed faecal samples from all mice before and after FMT to determine the changes in microbial composition. As shown in Fig. 2a and Supplementary Table 2, the species richness measured by ACE (203.75 ± 4.391 at 0 months, 337.814 ± 8.126 at 3 months, p < 0.001), Fisher (27.76 ± 0.684 at 0 months, 50.879 ± 1.288 at 3 month, p < 0.001), and Shannon (4.11 ± 0.049 at 0 month, 4.39 ± 0.058 at 3 month, p < 0.001) dramatically improved, indicating that the number of the microbial species constituting the gut microbiome was increased by FMT. Despite the increase in the microbial species of the gut microbiome, the species evenness measured by Simpson (0.966 ± 0.002 at 0 months, 0.96 ± 0.004 at 3 months, p > 0.05), and InvSimpson (31.966 ± 1.8 at 0 months, 32.06 ± 2.881 at 3 months, p > 0.05) indices indicated that the patterns of species distribution were not changed (Fig. 2a and Supplementary Table 2). In accordance with the species richness, phylogenetic classification showed that the microbial compositions dramatically altered and diversified after FMT. The gut microbiome mainly consisted of Bacteroidetes (56.906%) and Firmicutes (39.887%) before FMT (Fig. 2b). However, FMT increased the relative abundance of Verrucomicrobia (9.649%) and decreased the abundance of Bacteroidetes (48.925%) and Firmicutes (37.521%) (Fig. 2b). The microbial distribution at lower classification levels are shown in Supplementary Fig. 2. The analyses of beta diversities (Fig. 2c and Supplementary Fig. 3), phylogenetic trees (Fig. 2d and Supplementary Fig. 4), and co-occurrence network (Fig. 2e and Supplementary Table 3) further confirmed the distinct nature of the gut microbiome before and after FMT. Thus, it was established that FMT effectively replaces the gut microbiota.
a Comparisons of α-diversity indices of the fecal gut microbiome, measured using ACE richness, Fisher, Shannon, Simpson, and Inverse Simpson methods, between before (n = 30) and 3 months after FMT (n = 30) are shown in box plots. b Comparisons of phylogenetic compositions of the fecal gut microbiome at the phylum level between before (n = 30) and 3 months after FMT (n = 30). c The β-diversity plots of the fecal gut microbiome were measured using a nonmetric multidimensional scaling plot before (n = 30) and 3 months after FMT (n = 30). d Maximum-likelihood phylogenetic tree comprising taxa of the fecal gut microbiome before (left, n = 30) and 3 months after FMT (right, n = 30). The circular dendrogram rings represent phylum levels, with corresponding taxa depicted in the inner layer. See Supplementary Fig. 4 for the circular ring (phylum) and inner layer (family) color legends. e Color-coded network graphs of the fecal gut microbiome before (left, n = 30) and 3 months after FMT (right, n = 30), illustrating co-occurrence and mutual exclusion interactions among operational taxonomic units (OTUs). The network was developed using class-level OTUs. Additional network information is provided in Supplementary Table 3. 0m_Feces represents the fecal gut microbiome before FMT. 3m_Feces represents the fecal gut microbiome 3 months after FMT.
The GI gut microbiome analysis elucidated the association of the gut microbiome with spatial working memory
Building on prior findings that FMT influenced gut microbiome composition, we further examined how microbiome changes varied among mice with different SWM performance levels. Specifically, we examined changes in gut microbiota composition according to the degree of SWM change before and after FMT by analyzing fecal samples. As shown in Fig. 3a and Supplementary Table 2, the species richness measured by ACE, Fisher, and Shannon increased after FMT, indicating that the number of microbial species constituting the gut microbiome was increased by FMT in all groups. However, the species evenness assessed by Simpson did not show significant differences before and after FMT across all groups. Comparing alpha-diversity between groups after FMT in fecal samples, neither species richness nor evenness showed significant differences (p > 0.05 in all alpha-diversity index) (Fig. 3a and Supplementary Table 2). In accordance with the species richness, phylogenetic classification showed that the microbial compositions dramatically altered and diversified after FMT. The gut microbiome mainly consisted of Bacteroidetes (55.563% in the PI group, 60.837% in the II group, and 54.319% in the NI group) and Firmicutes (40.437% in the PI group, 36.945% in the II group, and 42.278% in the NI group) before FMT (Fig. 3b, left and Supplementary Table 4). However, FMT caused the proliferation of Verrucomicrobia (10.693% in the PI group, 8.153% in the II group, and 10.100% in the NI group), and thus reduction of the abundances of Bacteroidetes (48.959% in the PI group, 52.217% in the II group, and 45.600% in the NI group) and Firmicutes (35.986% in the PI group, 35.763% in the II group, and 40.814% in the NI group) (Fig. 3b, right and Supplementary Table 4). The microbial distribution at lower classification levels are shown in Supplementary Fig. 5 and Supplementary Tables 5–9. Although changes in the gut microbiome before and after FMT were confirmed in the fecal samples, no correlation was observed in the alpha diversities (p > 0.05) (Fig. 3a and Supplementary Table 2), the beta diversities analyses (p > 0.05) (Fig. 3c, d, Supplementary Figs. 6 and 7, and Supplementary Table 10), phylogenetic trees (Supplementary Fig. 8), and co-occurrence network analysis (Fig. 3e, f and Supplementary Table 3) between groups related to SWM.
a Comparisons of α-diversity indices of the fecal gut microbiome in each group, measured using ACE richness, Fisher, Simpson, and Shannon methods, between before and 3 months after FMT are shown in box plots (n = 10 per group). b Comparisons of phylogenetic compositions of the fecal gut microbiome in each group at the phylum level before FMT (left) and 3 months after FMT (right) (n = 10 per group). c The β-diversity plots of the fecal gut microbiome were measured using nonmetric multidimensional scaling plot before FMT (upper) and 3 months after FMT (lower) in each group (n = 10 per group). d Heatmaps display the unsupervised hierarchical clustering of microbial species’ relative abundance in the fecal gut microbiome of each group before (upper) and 3 months after FMT (lower) (n = 10 per group), with data log-transformed and row-scaled (standardized). The heatmap legend represents the range of row-scaled (standardized) values from log-transformed relative abundance data, with color gradients indicating deviations from the mean abundance of each species. e, f Color-coded network graphs of the fecal gut microbiome in each group before FMT (e) and 3 months after FMT (f) (n = 10 per group), illustrating co-occurrence and mutual exclusion interactions among OTUs. The network was developed using class-level OTUs. Additional network information is provided in Supplementary Table 3.
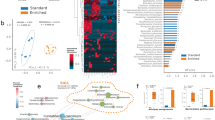
Since fecal gut microbiota does not fully represent the entire gut microbiome14, we further analyzed a possible association between the gut microbiome and SWM by using GI contents. The GI content was obtained by sacrificing the mice after FMT and collecting the contents from the stomach to the end of the colon. As shown in Fig. 4a and Supplementary Table 2, the species richness and distribution pattern of the gut microbiome isolated from the GI contents (GI gut microbiome) were very different from those isolated from feces (fecal gut microbiome). More importantly, there were clearly group differences in the species richness and distribution pattern of the GI gut microbiome unlike in the cases of the fecal gut microbiome. When comparing the alpha-diversity of the GI gut microbiome between the PI and NI groups, significant differences were observed in the ACE (p < 0.05), Fisher (p < 0.05), Simpson (p < 0.01), and Shannon (p < 0.01) indices (Fig. 4a and Supplementary Table 2). In accordance, the actual composition of the gut microbes constituting the gut microbiome was not only different from that of the fecal gut microbiome, but also there was a clear group difference (Fig. 4b, Supplementary Fig. 9 and Supplementary Tables 4–9). The composition of the GI gut microbiome differed significantly among groups according to SWM. At the phylum level, the abundance of Firmicutes was decreased (28.943% in the PI group, 43.796% in the II group, and 49.618% in the NI group), while the abundance of Verrucomicrobia was increased (42.974% in the PI group, 29.403% in the II group, and 16.264% in the NI group), especially in groups with increased SWM. The alpha-diversity analysis confirmed differences between groups according to SWM, while the beta-diversity analysis did not confirm differences between groups (p > 0.05) (Fig. 4c, d, Supplementary Fig. 10, and Supplementary Table 10). However, when compared to fecal samples (p = 0.544), it was observed that there was a slightly greater difference between groups in GI samples (p = 0.266). The phylogenetic trees (Fig. 4e), and co-occurrence network analysis (Fig. 4f and Supplementary Table 3) also confirmed that the gut microbiome of the three groups were differed each other. We believe that analysis of the GI gut microbiome but not the fecal gut microbiome visualized the association between the gut microbiome and SWM because the GI gut microbiome represents more accurately the real gut microbiome than the fecal gut microbiome. This result was perfectly matched with the recent works, which showed that the fecal gut microbiome does not represent the whole gut microbiome14.
a Comparisons of α-diversity indices of the GI gut microbiome, measured using ACE richness, Fisher, Simpson, and Shannon methods, 3 months after FMT are shown in box plots (n = 10 per group). b Comparison of phylogenetic compositions of the GI gut microbiome at the phylum level, 3 months after FMT (n = 10 per group). c The β-diversity plot of the GI gut microbiome, measured using a nonmetric multidimensional scaling plot, 3 months after FMT (n = 10 per group). d Heatmap displays the unsupervised hierarchical clustering of microbial species’ relative abundance in the GI gut microbiome of each group 3 months after FMT (n = 10 per group), with data log-transformed and row-scaled (standardized). The heatmap legend represents the range of row-scaled (standardized) values from log-transformed relative abundance data, with color gradients indicating deviations from the mean abundance of each species. e Maximum-likelihood phylogenetic tree comprising taxa of the GI gut microbiome 3 months after FMT in the PI group (left, n = 10), the II group (middle, n = 10), and the NI group (right, n = 10). The circular dendrogram rings represent phylum levels, with corresponding taxa depicted in the inner layer. See Supplementary Fig. 4 for the circular ring (phylum) and inner layer (family) color legends. f Color-coded network graphs of the GI gut microbiome 3 months after FMT in the PI group (left, n = 10), the II group (middle, n = 10), and the NI group (right, n = 10), illustrating co-occurrence and mutual exclusion interactions among OTUs. The network was developed using class-level OTUs. Additional network information is provided in Supplementary Table 3. 3m_GI represents the GI gut microbiome 3 months after FMT.
To determine whether sex affects the relationship between cognitive function and gut microbiota, we conducted analyses by separating the sex of the mice. As shown in Supplementary Fig. 11, alpha-diversity indices did not show significant differences in the fecal gut microbiome after FMT in both males and females but showed striking differences in the GI gut microbiome according to SWM (Supplementary Fig. 11a, b). In addition, in the composition of the gut microbiome at the phylum level, Firmicutes in the GI gut microbiome of the PI group was decreased, and Verrucomicrobia was increased in both males and females (Supplementary Fig. 11c, d). In other words, it was confirmed that there was no difference in the change in SWM according to FMT by sex.
The enrichment of Akkermansia muciniphila in the gut microbiome was positively associated with increased mouse spatial working memory
After identifying the association of the GI gut microbiome with SWM, we were bound to identify the individual gut microbial species associated with SWM. Figure 5 shows the gross association of the gut microbiome with SWM at the phylum level. As expected by the above gut microbiome analyses, there was not any correlation observed in the gut microbiome isolated from fecal samples with SWM. However, unlike the fecal gut microbiome, the GI gut microbiome analysis elucidated an associative relationship with SWM. The abundance of the phylum Verrucomicrobia was clearly correlated with increased SWM in mice compared to other phyla (Fig. 5a, R2 = 0.2241).
Comparisons of the GI gut microbiome and the fecal gut microbiome in the relationship between spatial working memory and gut microbiome at the phylum level. The % Abundance of Verrucomicrobia (a), Bacteroidetes (b), Firmicutes (c), Actinobacteria (d), Proteobacteria (e), Patescibacteria (f), Tenericutes (g), and Cyanobacteria (h) 3 months after FMT are shown in bar plots. The values represent mean ± S.E.M. GI gastrointestinal gut microbiome, Feces fecal gut microbiome.
To confirm the relationship at the species level, the Taxonomizr program was used to obtain the NCBI accession number with the most similar sequence to identify the microbial species, and then the DESeq2 method, which represents the difference in the abundance of intestinal microorganisms as a fold change in the gut microbial community of each group, was used for analysis (Fig. 6a). A total of five bacterial species were found to have significantly different abundance between the PI and NI groups. Among the five bacterial species, two were enriched in the positive group (Fig. 6b, c), while three were enriched in the negative group (Fig. 6d–f). Although the five bacteria showed an association with the SWM of mice, Akkermansia muciniphila was the only bacterial species realistically associated with SWM if considering the abundance of the bacteria in the gut microbiome. A. muciniphila was not only abundantly present in the gut microbiome of the mice groups so as to affect the phenotype of its host but also linearly correlated with the increase of SWM in an abundance-dependent manner.
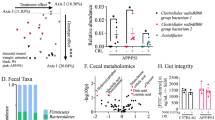
a Log2-fold change in abundance of most abundantly present species in the GI gut microbiome of the PI and NI groups (n = 10 per group) analyzed by DESeq2 differential abundance analysis. Each point represents a species comparison between two experimental groups. b, c Comparison of the GI gut microbiome and fecal gut microbiome of two species with higher % abundance in the PI group than in the NI group. Akkermansia muciniphila (b) and Olsenella profusa (c) are shown in the line graph. d–f Comparison of the GI gut microbiome and fecal gut microbiome of three species with higher % abundance in the NI group than in the PI group. Lactobacillus johnsonii (d), Turicibacter sanguinis (e), and Kineothrix alysoides (f) are shown in the line graph. The values represent mean ± S.E.M.
A. muciniphila was not quantified in human stool samples. However, previous studies have shown that the abundance of A. muciniphila increases with age in humans27,28,29,30, whereas it decreases in mice31,32,33. Our results also confirmed that A. muciniphila was almost absent in the gut microbiota of 12-month-old mice before FMT, and increased after FMT (Fig. 2b, Verrucomicrobia). Therefore, it is difficult to view it as a restoration of the original gut microbiota of mice, but rather as a microorganism established due to FMT.
Akkermansia muciniphila increased the mouse spatial working memory by increasing the expression level of BDNF in the hippocampus
In the FMT model, we observed that despite an increase in A. muciniphila, SWM remained unchanged in the II group and declined in the NI group. This suggests that the presence of A. muciniphila is not sufficient to enhance cognitive function when introduced within a complex microbial community. Given that FMT introduces a diverse microbial population, it is possible that bacterial species with opposing effects on cognition were also transferred, potentially offsetting the benefits of A. muciniphila.
To clarify the specific role of A. muciniphila, we conducted additional experiments where candidate strains were administered individually. These experiments were designed to isolate the effects of A. muciniphila without interference from other microbial species. Since it is known that cognitive function decreases with aging in mice34, we examined whether ingestion of A. muciniphila suppresses the decline in cognitive function due to aging. The two strains of A. muciniphila, GMB 0476 and GMB 2066, were orally administrated into aged mice expected to have cognitive decline, establishing a de novo microbiome for 3 months. As shown in Fig. 7a, feeding of A. muciniphila gradually increased the SWM, and there was a clear group difference after 3 months of the experimental period. The positive effects of both strains further validate the effect of A. muciniphila on the increase of the SWM. Also, it is interesting to note that A. muciniphila GMB 2066 (an increase from 61.8% to 66.8%) showed a better effect than A. muciniphila GMB 0476 (an increase from 61.8% to 65.7%).
a Effects of A. muciniphila administration on the cognitive ability by the y-maze spontaneous alternation test (Control group: n = 15 at 1 month point, n = 14 at 2 month point, n = 13 at 3 month point; GMB 0476 group: n = 17; GMB 2066: n = 18). b Images of mouse brain tissues stained with H&E (Control group: n = 5, GMB 0476 group: n = 5, GMB 2066: n = 5). c–e Effects of A. muciniphila administration on the BDNF and NEUN expression in the brain hippocampus stained with IF (c). The fluorescence intensity of NEUN (d) and BDNF (e) are shown in bar plots (Control group: n = 5, GMB 0476 group: n = 5, GMB 2066: n = 5). f Effect of A. muciniphila administration on the BDNF expression in the brain confirmed by western blot (Control group: n = 5, GMB 0476 group: n = 5, GMB 2066: n = 5). Scale bars in (b) and (c), 100 μm. The values represent mean ± S.E.M. *p < 0.05; **p < 0.01.
Previous studies have shown that brain-derived neurotrophic factor (BDNF), one of the major brain neurotrophic factors, is associated with neural plasticity and short- and long-term memory35,36,37,38. As a result of observing the brains of the mice, histological examination of the hippocampal area did not show significant differences among the experimental mice (Fig. 7b). However, the expression levels of brain-derived neurotrophic factor (BDNF) in the hippocampus were much more highly expressed in the A. muciniphila-feeding groups (Fig. 7c–f). Especially in accordance with the performance results of the memory test (Fig. 7a), BDNF in the A. muciniphila GMB 2066 feeding group (fluorescence intensity 10.8) was more highly expressed than that of the A. muciniphila GMB 0476 feeding group (fluorescence intensity 6.7) (Fig. 7c–f). This coincidence of the memory test result and the BDNF expression levels clearly indicates that A. muciniphila increased the mouse SWM by increasing the expression level of BDNF in the hippocampus. In addition to the improvement in SWM, the feeding of A. muciniphila resulted in enhanced blood profiles, including reductions in triglycerides (TG), blood glucose, and LDL-cholesterol levels, along with an increase in HDL-cholesterol (Supplementary Fig. 12). However, feeding of A. muciniphila does not improve immune response (Supplementary Fig. 13) and physical performance (Supplementary Fig. 14). The results from these targeted interventions of A. muciniphila provide stronger evidence for the direct impact of A. muciniphila on cognitive function.
Discussion
Recent extensive studies on the gut microbiome have highlighted its crucial role in a wide array of human diseases and phenotypes39,40,41,42,43,44,45,46,47,48,49,50. Intriguingly, some research suggests that the gut microbiome might even influence neurological diseases and neuronal functions through the gut-brain axis, pointing to a potential link between the gut microbiome and intelligence6,8,9,51,52. Many studies have showcased the impact of the gut microbiome on memory performance in experimental animals53,54,55,56,57. A recent study revealed that pasteurized A. muciniphila treatment have beneficial effects on anxiety-like behavior and memory defects in the C. rodentium infection model58. However, the mechanisms of A. muciniphila are still unclear. This study, which demonstrated an improvement in mouse memory via BDNF activation after feeding A. muciniphila, provides the first mechanistic evidence for the role of specific gut microbes in memory enhancement. BDNF has been implicated in neuroplasticity to form new neural connections59,60. Given that neuroplasticity is a foundational mechanism for learning and memory61,62, it is noteworthy to mention that A. muciniphila enhances mouse spatial working memory by elevating BDNF expression levels (Fig. 7). Furthermore, the improvement of spatial working memory was consistently observed with two independent A. muciniphila strains (GMB 0476 and GMB 2066), which further strengthens the validation of A. muciniphila’s effect on memory improvement (Fig. 7).
In addition to the memory-improving effect observed in this study, several previous works consistently demonstrated that A. muciniphila lowers blood glucose levels, LDL-cholesterol levels, and TG while increasing HDL-cholesterol levels63,64,65. It also shows promise in improving liver steatosis, reducing inflammation, and enhancing responsiveness to cancer immunotherapies66,67,68. The blood profile analysis conducted in this study also revealed improvements, including lowered levels of LDL-cholesterol, TG, and blood glucose, as well as increased HDL-cholesterol levels (Supplementary Fig. 12). However, these improvements were not as significant as those reported in previous studies. Past animal experiments assessing the effects of A. muciniphila on metabolic diseases typically employed diets high in fat and glucose69,70,71,72. In contrast, a normal diet was used in our study to evaluate A. muciniphila’s impact on spatial working memory. Despite the normal diet, the beneficial effects of A. muciniphila on blood profile improvement were evident, consistent with earlier research63,64,65.
While the gut microbiome significantly affects human intelligence, including memory, specific gut microbes associated with memory remain unidentified due to current limitations in gut microbiome analysis. These limitations stem from the host’s genetic background, which plays a critical role in determining diseases or phenotypes, and the fact that the fecal gut microbiome does not represent the entire gut microbiome of the host. In this study, we mitigated the effects of the host genetic background on memory by comparing the same individual mouse before and after FMT. As the genetic background of an individual mouse remains constant, any change in spatial working memory post-FMT can be attributed solely to the gut microbiome. In the analysis of the gut microbiome of mice before and after FMT, we were able to more accurately assess the impact of the gut microbiome on spatial working memory by analyzing not only fecal samples but also the entire gastrointestinal microbiome. With this methodology, we identified A. muciniphila as a gut microbe that enhances spatial working memory in mice (Figs. 1–6) and validated these findings through a proof-of-concept animal experiment (Fig. 7). While this study employed the spontaneous alternation Y-maze to assess cognitive improvement, a comprehensive evaluation of cognitive function requires additional tests, such as those measuring long-term memory and other cognitive domains. Further investigations are needed to explore the underlying mechanisms of A. muciniphila effects on cognition.
Despite these promising findings, certain limitations should be acknowledged. This study was conducted in a controlled laboratory environment using a specific mouse model, which may not fully replicate the complexity of human gut microbiome interactions and cognitive processes. Additionally, individual variability in response suggests that biological diversity, dietary factors, and environmental influences may contribute to the efficacy of A. muciniphila. Another limitation is that we cannot determine whether the differences in efficacy between the two A. muciniphila strains are due to the functional effects of the strains themselves or differences in their colonization levels. Future research should investigate the cognitive benefits associated with varying degrees of colonization while incorporating diverse study populations, extended observation periods, and mechanistic analyses to further elucidate the role of A. muciniphila in cognitive function and neuroprotection.
Methods
Study design and animal experiment
Thirty 12-month-old female/male C57BL/6 mice were purchased from the Animal Facility of Aging Science (KJ309) in Korea Basic Science Institute (Gwangju, Republic of Korea) and individually housed in a specific pathogen-free (SPF) facility with sterilized food and water ad libitum. Animals were maintained in a 12-h light/12-h dark cycle at 22 ± 1 °C and 40–50% humidity. Microbiological monitoring in a specific-pathogen-free (SPF) facility was performed using a Laser scanning confocal microscope (Leica TCS SP5 AOBS/Tandem, KJ302) at the Honam Regional Center of Korea Basic Science Institute (KBSI).
To investigate the influence of human gut microbes on SWM in mice, fecal samples were collected from ten healthy volunteers aged 20–40 years. The fecal samples were mixed in JM medium (Alpha Biochemicals Co., Jeonju, Republic of Korea) to create a composite fecal mixture containing a variety of human gut microbes73. After 1 week of acclimation, mice were weighed, and individual fecal and blood samples were collected. Subsequently, all mice were given water containing three types of antibiotics (1 g/L ampicillin, 0.5 g/L kanamycin, and 0.5 g/L cefoxitin; Sigma-Aldrich, MA, USA) and antifungal (0.5 g/L nystatin; Sigma-Aldrich, MA, USA) for 2 days to deplete their endogenous microbiota. Following microbiota depletion, mice were orally administered the human fecal mixture (20 μL) via oral gavage using a 22 G oral gavage needle, twice a week for 3 months to transplant human gut microbes into the mice. While FMT has the potential to introduce harmful factors, no adverse health effects were observed in this study. Mice were closely monitored for signs of distress, weight loss, or other health deterioration, but no unexpected adverse events occurred. Before and three months after FMT, mice were weighed, and individual fecal and blood samples were collected. Y-maze spontaneous alternation tests were conducted both before and three months after the administration of the human fecal mixture in a behavior analysis room set up separately from the rearing room. By comparing the Y-maze records before and after FMT, the subjects were divided into groups with increased SWM (PI, n = 10), unchanged SWM (II, n = 10), and decreased SWM (NI, n = 10).
For efficacy evaluation experiments, 21-month-old C57BL/6 mice (female, n = 50) were purchased from the Animal Facility of Aging Science at the Korea Basic Science Institute (Gwangju, Republic of Korea). The mice were divided into three groups and housed in cages by group (maximum of five mice per cage): Control (n = 15), GMB 0476 (feeding A. muciniphila GMB 0476, n = 17), and GMB 2066 (feeding A. muciniphila GMB 2066, n = 18). The gut microbiomes of mice were depleted by the daily administration of antibiotic/antifungal mixtures used above for 2 days. The two strains of A. muciniphila GMB 0476 and GMB 2066 were obtained from Gut Microbe Bank (https://www.gmbank.org), and cultured by using JM medium at 37 °C anaerobic condition. A. muciniphila was orally administered to mice daily to establish a de novo microbiome for 3 months.
The primary outcome measure used to determine the sample size was SWM performance. Sample size estimation was based on expected differences in SWM performance between groups, as assessed by the Y-maze test. Based on previous studies and a priori power analysis (80% power, α = 0.05), we calculated that 15 mice per group were required to detect an expected size of 0.8. Additionally, to account for potential dropouts due to unforeseen toxic reactions or the attrition of aged mice, we allocated an additional 2–3 mice per group. Mice were assigned to experimental groups using stratified randomization based on body weight to ensure similar average body weight across groups.
Body weight and behavior tests were performed monthly. However, no specific measures were taken to control potential confounders, such as the order of treatments and measurements or animal/cage location. Humane endpoints were established to minimize unnecessary suffering. Mice were monitored weekly for severe weight loss, significant lethargy, or abnormal behavior. If these conditions were observed, they were excluded from the study and euthanized according to ethical guidelines. Additionally, any mice that died from non-experimental causes before the study endpoint were excluded from the final analysis. These exclusion criteria were established a priori. In the control group, the number of mice decreased due to natural death, with 14 mice remaining one month after the start of the experiments, and 13 mice remaining two and three months later. Group allocation information was not blinded at any stage of the experiment. Researchers conducting the experiment, assessing outcomes, and analyzing data were aware of the group assignments.
At the end of the experiment, mice were euthanized under deep anesthesia with 3% isoflurane inhalation to minimize pain and distress. Once complete anesthesia was confirmed, blood was collected via cardiac puncture, and the brain was extracted for further analysis. All procedures were performed following ethical guidelines to ensure humane treatment.
Y-maze spontaneous alternation test
To assess spatial working memory, the Y-maze spontaneous alternation test was employed. The maze consisted of three black opaque Perspex arms (A, B, and C), each measuring 21 cm × 7 cm × 15.5 cm and containing a visual cue. Each mouse was placed individually in arm A and allowed to explore freely for 8 min. The sequence of arm entries was recorded. Spontaneous alternation, indicative of spatial working memory, was defined as entering all three arms consecutively, with overlapping triplets. The percentage of alternations was calculated as the number of actual alternations divided by the maximum number of alternations (the total number of arm entries − 2)74.
Analyses of biochemical parameters
The blood glucose level was determined using glucose test strips and a handheld blood glucose meter (Accu-Chek Active; Roche Diagnostic GmbH, Mannheim, Germany).
The serum levels of total cholesterol (TCHO), triglyceride (TG), and high-density lipoprotein cholesterol (HDL-CHO) were determined by enzymatic methods using commercial assay kits (Asan Pharmaceutical Co., Seoul, Republic of Korea) as described previously75,76. In brief, the low-density lipoprotein cholesterol (LDL-CHO) levels were calculated using Friedewald’s equation [(LDL-CHO) = (TCHO) − ((HDL-CHO) − (TG)/5)].
DNA extraction and 16S rDNA gene sequencing
To isolate genomic DNA from bacterial samples, a previously established phenol-chloroform isoamyl alcohol extraction method was employed77,78. Briefly, samples were suspended in lysis buffer (200 mM NaCl, 200 mM Tris-HCl (pH 8.0), 20 mM EDTA) and subjected to bead-beating for cell disruption. Following this, genomic DNA was partitioned from the lysate into the aqueous phase using a mixture of Phenol:Chloroform:Isoamylalcohol. The DNA was subsequently precipitated with 3 M sodium acetate and isopropanol, rinsed with 70% ethanol to remove impurities, and finally re-suspended in TE buffer (10 mM Tris-HCl (pH 8.0), 1 mM EDTA) for further analysis.
Metagenome sequencing analyses of the gut microbiome DNA were processed and sequenced by a commercial company, ebiogen, Inc. (Republic of Korea). They employed the Illumina 16S Metagenomic Sequencing Library protocols. Briefly, this involved amplifying the 16S rRNA gene V3–V4 region using specific primers for each sample. To add multiplexing indices and Illumina sequencing adapters, a subsequent limited-cycle amplification step was performed. The amplified libraries were then normalized and pooled using the PicoGreen, and size-verified using the Agilent TapeStation DNA ScreenTape D1000 system (Agilent Technologies, Santa Clara, CA, USA). Finally, paired-end sequencing was performed on the Illumina MiSeq platform (Illumina, San Diego, CA, USA). Sequence data was subsequently processed using DADA2 software (version 1.10.1) to account for potential amplicon errors79.
Data and statistical analyses
Statistical analyses were performed using established protocols as previously described77,78,80. Briefly, the QIIME 2 platform was used for data processing and analysis. Following denoising and quality control, the taxonomic classification of sequencing data was performed using the Q2-Feature classifier. Alpha-diversity metrics were calculated using the “q2-diversity” plugin with appropriate sampling depth. Normalized OUT data was then imported into R (version 3.6.1) using the phyloseq package (version 1.28.0) for visualization of alpha (within-group) and beta (between-group) diversity. Alpha-diversity comparisons were performed using the Kruskal–Wallis rank sum test. For beta-diversity analysis, Bray-Curtis dissimilarity matrices were generated from log-transformed OTU data using the vegan package in R. Non-metric multidimensional scaling (NMDS) was then performed to visualize community dissimilarities. The ADONIS test (vegan package) with 999 permutations was used to assess the statistical significance of factors influencing Bray–Curtis distances. Finally, unsupervised filtering with a 5% abundance threshold identified the most abundant taxa for heatmap generation using average linkage hierarchical clustering and Bray–Curtis distance metrics.
Construction of phylogenetic tree and co-occurrence network
Phylogenetic trees and co-occurrence networks were constructed for each experimental group following previously established methods77,78. To build the phylogenetic trees, taxa that could not be classified to the species level were assigned a new classification based on their NCBI accession numbers using the R package Taxonomizr (0.5.3)81. Subsequently, 16S rRNA sequences were aligned using default ClustalW parameters. These alignments served as the basis for constructing maximum-likelihood phylogenetic trees in MEGAX with 500 bootstrap replicates. Finally, all phylogenetic trees were visualized using the iTOL online platform.
To investigate microbial co-occurrence patterns associated with spatial working memory changes in mice, co-abundance networks were constructed using a permutation-renormalization-bootstrap approach. Unnormalized abundance data for each experimental group was uploaded separately to CoNet, a Java Cytoscape plugin. CoNet employed multiple ensemble correlation methods to identify significant co-occurrences of operational taxonomic units (OTUs) across samples. OTUs present in less than three samples were excluded from the analysis. The final network was generated by combining five similarity measures (Spearman and Pearson correlations, mutual information score, Bray–Curtis dissimilarity, and Kullback–Leibler Dissimilarity) calculated by CoNet. P-values from these measures were merged using Brown’s method and subsequently adjusted for multiple comparisons with the Benjamini–Hochberg correction (adjusted P-value < 0.05). Edges in the network represent significant co-abundance relationships between OTUs identified by at least two of the five similarity measures. Finally, the igraph package in R was used to visualize the co-occurrence network structure. This package employs the Louvain algorithm to identify network communities, optimizing the modularity score within each network.
Differential abundance
For species-level analysis, we used the Taxonomizr package (version 0.5.3) in R to assign species names based on NCBI accession numbers81. Subsequently, following established protocols77,78, differential abundance analysis was performed using DESeq2 software (version 1.24.0)82,83. This analysis compared the relative abundance of taxa in the gut microbiome between spatial working memory groups. Taxa with a presence in less than 0.5% of samples were excluded from the analysis to ensure robust comparisons.
H & E staining
The histological analysis protocol was followed as previously described84. For mouse anesthesia, Open the abdominal cavity and fully expose the heart and lungs. A needle is inserted at the end of the left ventricle of the heart, and a saline solution is injected for 10 min. After 10 min, a 10% formalin solution is injected with a needle at the same location. The mouse brain was harvested and fixed in a 10% formalin solution. The brain tissue was embedded in paraffin and sliced at 5 µm thickness. After staining the nuclei with Harris hematoxylin solution (Sigma-Aldrich, Catalog#HHS32) for 5 min and washing with water. The cytoplasm was stained with Eosin solution (Sigma-Aldrich, Catalog#HT110116) for 3 min. And dehydrated by immersion in 80% ethanol three times, 95% ethanol three times, and 100% ethanol three times. The stained tissue was sealed using a mounting agent (Vector Laboratories, Catalog#H-5000-60) and cover glass. Images were captured under a light and fluorescence slide scanner (400X, KFBio, Catalog#KF-FL-005).
Immunohistochemistry
For immunostaining, brain tissues from mice at 12 weeks of the experimental intervention were embedded in paraffin and cut into 5 µm thick sections on a microtome machine. The tissue sections were deparaffinized with xylene and rehydrated with a descending series of diluted ethanol (100%, 95%, 70%, and 50%) and H2O. Antigens in the tissue sections were retrieved with 10 mM sodium citrate buffer, pH 6.0 for 25 min at 85 °C, subsequently rinsed in PBS and then incubated in 0.3% H2O2 in PBS to quench endogenous peroxidase. After that, the tissue section was rinsed with PBS for incubation with 5% normal goat serum in PBS containing 0.025% Triton X-100 at RT for 30 min to block nonspecific binding. Sections were then blocked with 5% BSA for 30 min and incubated with primary (BDNF Polyclonal Antibody, rabbit and Anti-NeuN Antibody, clone A60, mouse) antibody (1:200; Invitrogen, Catalog#PA585730 and Sigma-Aldrich, Catalog#MAB377) overnight at 4 °C. The sections were incubated with secondary (Goat anti-Rabbit IgG (H + L) Cross-adsorbed secondary antibody Alexa Fluor 594 and Goat anti-mouse IgG (H + L) highly Cross-adsorbed secondary antibody, Alexa Fluor 488) antibody (1:200; Invitrogen, Catalog#A-11012 and Catalog#A-11029) for 1 h. Nuclei were stained with DAPI (Thermo Fisher Scientific, Catalog#62248) for 15 min. Slides were mounted with an antifade mounting medium solution (Vector Laboratories, Catalog#H-1400-10). Images were captured under a light and fluorescence slide scanner (400X, KFBio, Catalog#KF-FL-005).
Western blotting
For the western blot assay, brain tissue was homogenized and proteins were extracted using lysis buffer (Thermo Fisher, USA, Cat #89900), and additionally, a protein inhibitor cocktail was added (Abbkine Scientific Co., China, Cat #BMP100). Tissue lysate supernatants were subjected to SDS-PAGE Gel Electrophoresis, and the separated proteins were transferred to a nitrocellulose membrane. The membrane was then incubated with primary antibodies specific to the protein of interest, BDNF (Thermo Fisher, USA, Cat #PA585730) and beta-actin (Abcam, UK, Cat #ab8227), followed by incubation with a secondary antibody (Goat Anti-Rabbit IgG H&L (HRP): Abcam, UK, Cat #ab6721) conjugated to an enzyme for detection. The protein of interest was visualized using an imaging system (iBright CL750 Imaging system, Invitrogen, USA, Cat #A44116).
Enzyme-linked immunosorbent assay
The mouse blood serum was used to measure the levels of three cytokines, IL-6, IL-1β, and TNF-α using ELISA kits from MyBioSource (Cat #MBS175960, #MBS175967, and #MBS175787). The kit manuals typically provide detailed instructions for each step of the ELISA protocol, including sample preparation, incubation times and temperatures, washing steps, detection reagents, and data analysis.
Motor function test
After the consumption of two strains of A. muciniphila, the motor abilities were evaluated through the Pole test, Wire suspension test73, and the Rotarod test78, which measured the locomotor ability of the hind limbs. In the Pole test, the time taken for the mouse’s head to descend completely from the top to the floor was measured using a straight wooden pole with a diameter of 1 cm and a height of 60 cm. This measurement was used to assess the muscle strength of the mouse. The Wire test involved placing the mouse on a 120 cm wire and measuring the time it could hold on before falling off. This method was used to evaluate the muscle endurance of the mouse. The Rotarod test involved placing the mouse on a rotating rod with a diameter of 3 cm and setting it to rotate at 30 rpm. The time it took for the mouse to fall off the rotating rod was measured. This method was used to evaluate the locomotor ability and motor skills of the mouse’s hind limbs. All motor ability tests were conducted with a time limit of 300 s, and the evaluations were performed at monthly intervals.
Statistics and reproducibility
All statistical analyses are reported as the mean ± standard error of the mean (SEM). Statistical comparisons were conducted using Student’s t-test or one-way ANOVA, as appropriate. The differences in the relative abundance of bacterial populations were analyzed using the Mann–Whitney sum rank tests in R software. Significance was declared at P < 0.05 with Benjamini and Hochberg’s adjustment. The normality of the data was assessed using the Shapiro–Wilk test. The homogeneity of variance was tested using Levene’s test. If the assumptions for parametric tests were violated, non-parametric alternatives such as the Mann–Whitney U test or Kruskal–Wallis test were used instead.
The sample size was determined based on prior studies and a priori power analysis (80% power, α = 0.05), which estimated that at least 15 mice per group were needed to detect an expected effect size of 0.8. To account for potential attrition due to aging-related mortality or other unexpected factors, an additional 2–3 mice were included in each group. Biological replicates were defined as individual mice within each experimental group, with sample sizes ranging from 15 to 18 per group. Measurements were taken from each mouse separately, ensuring independent observations. No technical replicates were included in the statistical analysis.
Ethics approval
Human fecal samples for FMT were collected from healthy participants who provided written informed consent. This study was approved by the Institutional Review Board (IRB) of Jeonbuk National University Hospital (Approval No. CHU_KOREAN_FOOD_2-2_2010) and conducted in accordance with the Declaration of Helsinki. All ethical regulations relevant to human research participants were followed.
Animal experiments were approved by the Institutional Animal Care and Use Committee (IACUC) of the Korea Basic Science Institute (Approval No. KBSI-IACUC-23-12) and conducted following the ARRIVE guidelines. We have complied with all relevant ethical regulations for animal use. A detailed study protocol, including research objectives, design, and analysis plan, was prepared before the study began but was not registered in a public database.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The 16s rRNA sequencing data were deposited in the NCBI SRA database under the BioProject accession number: PRJNA123409085. The taxonomic profiles processed from this for analysis were deposited in Figshare (https://doi.org/10.6084/m9.figshare.22123937.v2)86. All numerical source data underlying the graphs and charts presented in the figure were deposited in Figshare (https://doi.org/10.6084/m9.figshare.28594700.v1)87. Uncropped western blot images were provided in Supplementary Fig. 15.
References
Turnbaugh, P. J. et al. The human microbiome project. Nature 449, 804–810 (2007).
Pascal, V. et al. A microbial signature for Crohn’s disease. Gut 66, 813–822 (2017).
Shen, Z. H. et al. Relationship between intestinal microbiota and ulcerative colitis: Mechanisms and clinical application of probiotics and fecal microbiota transplantation. World J. Gastroenterol. 24, 5–14 (2018).
Turnbaugh, P. J. et al. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 444, 1027–1031 (2006).
Han, H. et al. Gut Microbiota and type 1 diabetes. Int. J. Mol. Sci. 19, 995 (2018).
Zhu, F., Tu, H. & Chen, T. The microbiota-gut-brain axis in depression: the potential pathophysiological mechanisms and microbiota combined antidepression effect. Nutrients 14, 2081 (2022).
Padhi, P. et al. Mechanistic insights into gut microbiome dysbiosis-mediated neuroimmune dysregulation and protein misfolding and clearance in the pathogenesis of chronic neurodegenerative disorders. Front. Neurosci. 16, 836605 (2022).
Tan, A. H., Lim, S. Y. & Lang, A. E. The microbiome-gut-brain axis in Parkinson disease—from basic research to the clinic. Nat. Rev. Neurol. 18, 476–495 (2022).
Escobar, Y. H., O’Piela, D., Wold, L. E. & Mackos, A. R. Influence of the microbiota-gut-brain axis on cognition in Alzheimer’s Disease. J. Alzheimer’s. Dis.87, 17–31 (2022).
Gonzalez, A. & Knight, R. Advancing analytical algorithms and pipelines for billions of microbial sequences. Curr. Opin. Biotechnol. 23, 64–71 (2012).
Bailén, M., Bressa, C., Larrosa, M. & González-Soltero, R. Bioinformatic strategies to address limitations of 16rRNA short-read amplicons from different sequencing platforms. J. Microbiol. Methods 169, 105811 (2020).
Pan, A. Y. Statistical analysis of microbiome data: The challenge of sparsity. Curr. Opin. Endocr. Metab. Res. 19, 35–40 (2021).
Lkhagva, E. et al. The regional diversity of gut microbiome along the GI tract of male C57BL/6 mice. BMC Microbiol. 21, 44 (2021).
Ahn, J. S. et al. Fecal microbiome does not represent whole gut microbiome. Cell. Microbiol. 2023, 6868417 (2023).
Blekhman, R. et al. Host genetic variation impacts microbiome composition across human body sites. Genome Biol. 16, 191 (2015).
Lim, M. Y. et al. The effect of heritability and host genetics on the gut microbiota and metabolic syndrome. Gut 66, 1031–1038 (2017).
Cahana, I. & Iraqi, F. A. Impact of host genetics on gut microbiome: take-home lessons from human and mouse studies. Anim. Models Exp. Med. 3, 229–236 (2020).
Lkhagva, E., Chung, H. J., Ahn, J. S. & Hong, S. T. Host factors affect the gut microbiome more significantly than diet shift. Microorganisms 9, 2520 (2021).
Deary, I. J. et al. Age-associated cognitive decline. Br. Med. Bull. 92, 135–152 (2009).
Lee, J. & Kim, H. J. Normal aging induces changes in the brain and neurodegeneration progress: review of the structural, biochemical, metabolic, cellular, and molecular changes. Front. Aging Neurosci. 14, 931536 (2022).
Wu, M. L. et al. Age-related cognitive decline is associated with microbiota-gut-brain axis disorders and neuroinflammation in mice. Behav. Brain Res. 402, 113125 (2021).
Kossowska, M. et al. The interplay between gut microbiota and cognitive functioning in the healthy aging population: a systematic review. Nutrients 16, 852 (2024).
Karlsgodt, K. H. et al. Genetic influence on the working memory circuitry: behavior, structure, function and extensions to illness. Behav. Brain Res. 225, 610–622 (2011).
Lee, P., Yacyshyn, B. R. & Yacyshyn, M. B. Gut microbiota and obesity: an opportunity to alter obesity through faecal microbiota transplant (FMT). Diab. Obes. Metab. 21, 479–490 (2019).
Basson, A. R. et al. Autologous fecal microbiota transplantation for the treatment of inflammatory bowel disease. Transl. Res. 226, 1–11 (2020).
Yadegar, A. et al. Fecal microbiota transplantation: current challenges and future landscapes. Clin. Microbiol. Rev. 37, e0006022 (2024).
Bäckhed, F. et al. Dynamics and stabilization of the human gut microbiome during the first year of life. Cell Host Microbe 17, 690–703 (2015).
Guo, M. et al. Developmental differences in the intestinal microbiota of Chinese 1-year-old infants and 4-year-old children. Sci. Rep. 10, 19470 (2020).
Kong, F. et al. Gut microbiota signatures of longevity. Curr. Biol. 26, R832–R833 (2016).
Biagi, E. et al. Gut microbiota and extreme longevity. Curr. Biol. 26, 1480–1485 (2016).
Alam, M. S. et al. Aging-induced dysbiosis of gut microbiota as a risk factor for increased Listeria monocytogenes infection. Front. Immunol. 12, 672353 (2021).
van der Lugt, B. et al. Integrative analysis of gut microbiota composition, host colonic gene expression and intraluminal metabolites in aging C57BL/6J mice. Aging 10, 930–950 (2018).
Zhang, X. et al. Age-related compositional changes and correlations of gut microbiome, serum metabolome, and immune factor in rats. GeroScience 43, 709–725 (2021).
Clifford, K. P. et al. Brain structure and working memory adaptations associated with maturation and aging in mice. Front. Aging Neurosci. 15, 1195748 (2023).
Poo, M. M. Neurotrophins as synaptic modulators. Nat. Rev. Neurosci. 2, 24–32 (2001).
Tyler, W. J., Alonso, M., Bramham, C. R. & Pozzo-Miller, L. D. From acquisition to consolidation: on the role of brain-derived neurotrophic factor signaling in hippocampal-dependent learning. Learn Mem. 9, 224–237 (2002).
Bekinschtein, P. et al. BDNF is essential to promote persistence of long-term memory storage. Proc. Natl. Acad. Sci. USA 105, 2711–2716 (2008).
Li, B. et al. Impaired spatial working memory and decreased frontal cortex BDNF protein level in dopamine transporter knockout mice. Eur. J. Pharm. 628, 104–107 (2010).
Chung, H. J., Nguyen, T. T. B., Kim, H. J. & Hong, S. T. Gut microbiota as a missing link between nutrients and traits of human. Front. Microbiol. 9, 1510 (2018).
Belkaid, Y. & Hand, T. W. Role of the microbiota in immunity and inflammation. Cell 157, 121–141 (2014).
Honda, K. & Littman, D. R. The microbiota in adaptive immune homeostasis and disease. Nature 535, 75–84 (2016).
Morgan, X. C. et al. Dysfunction of the intestinal microbiome in inflammatory bowel disease and treatment. Genome Biol. 13, R79 (2012).
Sheflin, A. M., Whitney, A. K. & Weir, T. L. Cancer-promoting effects of microbial dysbiosis. Curr. Oncol. Rep. 16, (2014). 406.
Zou, R. et al. Dysbiosis of gut fungal microbiota in children with autism spectrum disorders. J. Autism Dev. Disord. 51, 267–275 (2021).
Liu, C. et al. Understanding the gut microbiota and sarcopenia: a systematic review. J. Cachexia Sarcopenia Muscle 12, 1393–1407 (2021).
Russell, S. L. et al. Early life antibiotic-driven changes in microbiota enhance susceptibility to allergic asthma. EMBO Rep. 13, 440–447 (2012).
Karlsson, F. H. et al. Symptomatic atherosclerosis is associated with an altered gut metagenome. Nat. Commun. 3, 1245 (2012).
Liu, J., Lkhagva, E., Chung, H. J., Kim, H. J. & Hong, S. T. The pharmabiotic approach to treat hyperammonemia. Nutrients 10, 140 (2018).
Nguyen, T. T. B., Chung, H. J., Kim, H. J. & Hong, S. T. Establishment of an ideal gut microbiota to boost healthy growth of neonates. Crit. Rev. Microbiol. 45, 118–129 (2019).
Nguyen, T. T. B., Jin, Y. Y., Chung, H. J. & Hong, S. T. Pharmabiotics as an emerging medication for metabolic syndrome and its related diseases. Molecules (Basel, Switz.) 22, 1795 (2017).
Li, Y. et al. Gut metagenomic characteristics of ADHD reveal low Bacteroides ovatus-associated host cognitive impairment. Gut Microbes 14, 2125747 (2022).
Srikantha, P. & Mohajeri, M. H. The possible role of the microbiota-gut-brain-axis in autism spectrum disorder. Int. J. Mol. Sci. 20, 2115 (2019).
Silva, V. et al. The impact of the gut microbiome on memory and sleep in Drosophila. J. Exp. Biol. 224, jeb233619 (2021).
Kubinyi, E., Bel Rhali, S., Sándor, S., Szabó, A. & Felföldi, T. Gut microbiome composition is associated with age and memory performance in pet dogs. Animals 10, 1488 (2020).
Li, L. et al. Gut microbiome drives individual memory variation in bumblebees. Nat. Commun. 12, 6588 (2021).
Zheng, H. et al. Depletion of acetate-producing bacteria from the gut microbiota facilitates cognitive impairment through the gut-brain neural mechanism in diabetic mice. Microbiome 9, 145 (2021).
D’Amato, A. et al. Faecal microbiota transplant from aged donor mice affects spatial learning and memory via modulating hippocampal synaptic plasticity- and neurotransmission-related proteins in young recipients. Microbiome 8, 140 (2020).
Meynier, M. et al. Pasteurized akkermansia muciniphila improves irritable bowel syndrome-like symptoms and related behavioral disorders in mice. Gut Microbes 16, 2298026 (2024).
Yang, T. et al. The role of BDNF on neural plasticity in depression. Front. Cell. Neurosci. 14, 82 (2020).
Berton, O. et al. Essential role of BDNF in the mesolimbic dopamine pathway in social defeat stress. Science 311, 864–868 (2006).
Machado, S. et al. Aprendizaje y memoria implícita: mecanismos y neuroplasticidad [Learning and implicit memory: mechanisms and neuroplasticity]. Rev. Neurol. 46, 543–549 (2008).
Mateos-Aparicio, P. & Rodríguez-Moreno, A. The impact of studying brain plasticity. Front. Cell. Neurosci. 13, 66 (2019).
Lin, X. Q. et al. Akkermansia muciniphila suppresses high-fat diet-induced obesity and related metabolic disorders in beagles. Molecules. 27, 6074 (2022).
Qu, L. et al. Improvement in Zebrafish with diabetes and Alzheimer’s Disease Treated with Pasteurized Akkermansia muciniphila. Microbiol. Spectr. 11, e0084923 (2023).
Shen, J. et al. Low-density lipoprotein receptor signaling mediates the triglyceride-lowering action of Akkermansia muciniphila in genetic-induced hyperlipidemia. Arteriosclerosis Thrombosis Vasc. Biol. 36, 1448–1456 (2016).
Li, T. et al. Akkermansia muciniphila suppressing nonalcoholic steatohepatitis associated tumorigenesis through CXCR6+ natural killer T cells. Front. Immunol. 13, 1047570 (2022).
Raftar, S. K. A. et al. The anti-inflammatory effects of Akkermansia muciniphila and its derivates in HFD/CCL4-induced murine model of liver injury. Sci. Rep. 12, 2453 (2022).
Fan, S. et al. Akkermansia muciniphila: a potential booster to improve the effectiveness of cancer immunotherapy. J. Cancer Res. Clin. Oncol. https://doi.org/10.1007/s00432-023-05199-8 (2023).
Org, E. et al. Genetic and environmental control of host-gut microbiota interactions. Genome Res. 25, 1558–1569 (2015).
Shin, J. et al. Elucidation of Akkermansia muciniphila probiotic traits driven by mucin depletion. Front. Microbiol. 10, 1137 (2019).
Depommier, C. et al. Pasteurized Akkermansia muciniphila increases whole-body energy expenditure and fecal energy excretion in diet-induced obese mice. Gut Microbes 11, 1231–1245 (2020).
Yoon, H. S. et al. Akkermansia muciniphila secretes a glucagon-like peptide-1-inducing protein that improves glucose homeostasis and ameliorates metabolic disease in mice. Nat. Microbiol. 6, 563–573 (2021).
Ahn, J. S. et al. Identification of the intestinal microbes associated with locomotion. Int. J. Mol. Sci. 24, 11392 (2023).
Kraeuter, A. K., Guest, P. C. & Sarnyai, Z. The Y-maze for assessment of spatial working and reference memory in mice. Methods Mol. Biol.1916, 105–111 (2019).
Chung, H. J., Sharma, S. P., Kim, H. J., Baek, S. H. & Hong, S. T. The resveratrol-enriched rice DJ526 boosts motor coordination and physical strength. Sci. Rep. 6, 23958 (2016).
Baek, S. H. et al. Treatment of obesity with the resveratrol-enriched rice DJ-526. Sci. Rep. 4, 3879 (2014).
Zhai, C. et al. Comparative analysis of original and replaced gut microbiomes within same individuals identified the intestinal microbes associated with weight gaining. Microorganisms 10, 1062 (2022).
Ahn, J. S. et al. Identification of muscle strength-related gut microbes through human fecal microbiome transplantation. Int. J. Mol. Sci. 25, 662 (2024).
Callahan, B. J. et al. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
Callahan, B. J., Sankaran, K., Fukuyama, J. A., McMurdie, P. J. & Holmes, S. P. Bioconductor Workflow for Microbiome Data Analysis: from raw reads to community analyses. F1000Research 5, 1492 (2016).
Sherrill-Mix, S. taxonomizr: Functions to Work with NCBI Accessions and Taxonomy. R Package Version 0.5. Available online https://CRAN.R-project.org/package=taxonomizr.
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Paulson, J. N., Stine, O. C., Bravo, H. C. & Pop, M. Differential abundance analysis for microbial marker-gene surveys. Nat. Methods 10, 1200–1202 (2013).
Chung, H. J., Lee, H. K., Kwon, K. B., Kim, H. J. & Hong, S. T. Transferrin as a thermosensitizer in radiofrequency hyperthermia for cancer treatment. Sci. Rep. 8, 13505 (2018).
Ahn, J. S., Han, E. J., & Chung, H. J. Comparision of gut microbiome related to SWM change before and 3 month after FMT. NCBI SRA Database (PRJNA1234090) (2025).
Hong, S. T., Chung, H. J. & Ahn, J. S. (2023). Intelligent GI, Feces figshare. figshare https://doi.org/10.6084/m9.figshare.22123937.v2 (2023).
Ahn, J. S. Increasing Spatial Working Memory in Mice with Akkermansia muciniphila - Numerical source data. figshare https://doi.org/10.6084/m9.figshare.28594700.v1 (2025).
Acknowledgements
This research was funded by the Korea Basic Science Institute (KBSI) grant C512230. This research was also supported by the Bio&Medical Technology Development Program of the National Research Foundation (NRF), funded by the Korean government (MSIT) (No. RS-2023-00224099).
Author information
Authors and Affiliations
Contributions
S.T.H., J.S.A., and H.J.C. designed the project and experiments; J.S.A. and S.K. analyzed results; J.S.A. and S.K. performed the experimental works; J.S.A., S.K., E.J.H., H.J.C., and S.T.H. wrote the paper. All authors read and approved the final paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editors: Joao Valente. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ahn, JS., Kim, S., Han, EJ. et al. Increasing spatial working memory in mice with Akkermansia muciniphila. Commun Biol 8, 546 (2025). https://doi.org/10.1038/s42003-025-07975-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-025-07975-3
This article is cited by
-
Discovery of intestinal microorganisms that affect the improvement of muscle strength
Scientific Reports (2025)