Abstract
Hydrogen bond cooperativity (HBC) plays an important role in stabilizing protein assemblies built by α-helices and β-sheets, the most common secondary structures. However, whether HBC exists in other types of protein secondary structures such as polyproline II (PPII) helices remains unexplored. This is intriguing, since PPII systems as assembling blocks are continuously emerging across multiple fields. Here, using a combination of computational chemistry tools and molecular modeling corroborated by experimental observables, we characterize the distinct H-bonding patterns present in PPII helical bundles and establish that HBC stabilizes intermolecular PPII helices as seen in other protein assemblies such as amyloid fibrils. In addition to cooperative interactions in canonical CO···HN H-bonds, we show that analogous interactions in non-canonical CO···HαCα H-bonds are relevant in Gly-rich PPII bundles, thus compensating for the inability of glycine residues to create hydrophobic cores. Our results provide a mechanistic explanation for the assembly of these bundles.
Similar content being viewed by others
Introduction
The last decade has witnessed remarkable advances in the design of proteins with unnatural folds, functions, and enzymatic activities1. These proteins are built of α-helices and β-sheets, which are the two main classes of secondary structure in globular proteins. However, a third class of protein secondary structure, the polyproline II (PPII) helix, has been rarely utilized in protein design2,3. A triple PPII helix is the basis of collagen, and there are several proteins and protein domains which are composed of bundles of Gly-rich PPII helices4. The abundance of collagen and the diversity of Gly-rich PPII helical bundle proteins underscores the potential usefulness of this structural element as a building block for protein design. In this regard, recent applications have exploited the extended conformation of PPII helices and their assembling potential to develop copolymers with distinct applications in different fields such as biomedicine and biotechnology5. Still, progress is slowed by our limited understanding of the bases of PPII helical bundle assembly and conformational stability. To address this gap in our comprehension, we and others have studied the Hypogastrura harveyi “snow flea” antifreeze protein (HhAFP) as a model system for Gly-rich PPII helical bundles. The structure of the HhAFP consists of a bundle of six PPII helices organized into two sheets oriented antiparallel to each other, each sheet containing three parallel PPII helices6,7. The intramolecular nature of the HhAFP PPII bundle facilitated high-resolution structural and dynamical studies, which showed that despite a high content in glycine residues, and an apparent lack of a stable hydrophobic core6,8, the conformational stability of the intramolecular PPII bundle is similar to typically globular proteins. This observation was explained by a number of distinct contributions such as the two disulfide bonds, canonical backbone CO···HN H-bonds as well as twenty-eight non-canonical backbone CO···HαCα H-bonds in accordance with experimental7,8 and computational reports7, and could be additionally stabilized by n→π* interactions9,10. Moreover, for non-glycine residues the presence of unconventional intramolecular interactions such as N-H···H-C(sp3), π···H-C(sp3) or CO···H-C(sp2) between the main chain and hydrophobic side chains, both aliphatic and aromatic, has been reported in PPII helix-forming peptides11. All these energetic factors may also stabilize other Gly-rich PPII bundles containing a larger number of PPII helices, such as the Granisotoma rainieri “springtail” antifreeze protein (GrAFP)12, the larger isoform of the HhAFP13 or the anaplastic lymphoma kinase (ALK)14 receptor. The PPII bundles assembled by these proteins contain as many as nine, thirteen, and fourteen PPII helices, respectively, as revealed in the crystallographic structures12,14 or homology models13. The unfolded ensemble of the HhAFP short isoform is compact and contains nonlocal electrostatic interactions which might initiate folding15. The cooperative nature of this protein’s folding was established by kinetic studies8 and recent protein engineering studies have revealed that activity increases with the number of PPII helices16. These results are consistent with, but do not prove, hydrogen bond cooperativity (HBC) in PPII helical bundles.
The unexpectedly high conformational stability and the large number of H-bonds that are present in PPII bundles, as seen in the HhAFP fold, are reminiscent of amyloids, which are quasi-infinite β-sheets with networks of aligned H-bonds. In both PPII helices and β-sheet-rich amyloids, the polypeptide chain extends facilitating self-assembly. One critical contribution to the remarkably high conformational stabilities of amyloids are cooperative H-bonding interactions, in which the alignment of CO···HN backbone H-bonding groups leads to increased polarization and strength17,18,19. We, among others, have reported that these HBC effects are also present in Asn and Gln residues, whose amide side chains form strongly hyperpolarized H-bonds20,21,22. Interestingly, in the PPII helices of Gly-rich proteins featuring PPII helical bundles such as the HhAFP, networks of aligned, canonical CO···HN H-bonds which resemble those observed in amyloids are present across the helix-helix interfaces, suggesting that HBC might also occur in these systems. In fact, HBC was previously suggested as the reason why polyglycine exhibited stronger CO···HN H-bonds when grouped in polyglycine II (PGII)-type structures, which do align multiple helices, versus collagen triple helices, which do not23. In addition, a number of aligned non-canonical CO···HαCα H-bonds are also present at the helix-helix interfaces in PPII helical bundles, providing an additional source of stability7,8. These CO···HαCα H-bonds between PPII helices have been long known in proteins like collagen24 and PGII assemblies25,26, and although their cooperative nature has not been established, CO···HC cooperative H-bonding interactions have been reported in model systems, such as in (HFCO)n clusters and other carbonyl compounds27,28, or in theoretical models to study the interactions between water and DMSO29.
In this study, we have investigated the nature of H-bonding interactions in the context of both canonical CO···HN and non-canonical CO···HαCα H-bonds across helix-helix interfaces in Gly-rich PPII helical bundles. Molecular models of PPII-based assemblies based on NMR observables were built to afford a detailed characterization of all H-bonding interactions in these assemblies. By means of four independent approaches: Dispersion-corrected Density Functional Theory (DFT-D), Natural Bonding Orbital (NBO), Quantum Theory of Atoms in Molecules (QTAIM), and Symmetry-Adapted Perturbation Theory (SAPT) calculations, we provide a detailed picture of an energetic kernel that drives PPII bundle assembly. To date, NBO methodology has served to demonstrate cooperative effects between canonical CO···HN H-bonds in amides and peptides describing other types of secondary structures30,31, but not in PPII helices, nor between non-canonical CO···HαCα H-bonds. Regarding QTAIM, whereas both canonical and non-canonical H-bonds of the two most common secondary structures have already been studied32,33, their cooperativity has not and there is no information for PPII helices. In fact, H-bonding cooperativity has only been studied using QTAIM analysis for small molecules34,35. Finally, SAPT allows us to study the nature of non-covalent interactions36, and although HBC has been studied in protein ligands using this methodology, it has not been applied to HBC between the different secondary structures to the best of our knowledge37.
Results
Model system to study Gly-Rich PPII bundles
The experimentally resolved 3D structure of the short isoform of the HhAFP consists of six PPII helices, where helices I, III, and V constitute a polar face while helices II, IV, and VI form the non-polar, ice-binding surface (Fig. 1A). The large HhAFP isoform is predicted by AlphaFold2 (AF2) with a similar fold containing thirteen instead of six PPII helices (Fig. 1A), which is in agreement with the model proposed by Mok et al. 13. The GrAFP structure also matches this bundle arrangement (Fig. 1A), with five parallel helices in one face packing antiparallel with a four-helical face12. The dearth of side chains in these Gly-rich sequences underlines the importance of H-bonding for bundle stabilization in these and in other Gly-rich PPII bundles such as in the recently elucidated ALK structure14 (Fig. 1A).
A Experimentally determined structures from the Protein Data Bank (PDB) for HhAFP short isoform (PDB ID: 3BOG), GrAFP (PDB ID: 7JJV) and ALK Gly-rich domain (PDB ID: 7LS0/7LRZ), and predicted structure from AF2 for HhAFP long isoform (AF2 ID: D7PBP2). Roman numerals denote the distinct helices in each protein. The dashed red circle indicates the PPII helix cloned for the computational model. B Gly-rich PPII helical bundle computational model: red spheres represent oxygen; blue, nitrogen; gray, carbon, and white, hydrogen. Shown with a green dashed line are the potential hydrogen bonds that may exist based on the proximity between the hydrogen donor (i.e., N or C) and hydrogen acceptor (i.e., O) atoms. C Scheme utilized for the calculation of the different parameters between PPII helices in the different interfaces. Blue PPII helices give rise to the ab interface; the red ones, to the bc interface, and the green ones, to the cd interface. The interaction energies are probed at these colored interfaces, i.e., where the n-mers are split for the HBS calculations, both DFT-D and SAPT. The rest of the computational parameters are calculated directly between the colored PPII helices through the NBO and QTAIM analyses. For added clarity, the gray eye symbol indicates the interface under study for each case.
In order to investigate the nature of H-bonding interactions across the different PPII bundles described above, we have designed a model system that recreates the interactions seen in the various assemblies. This model relies on replicating the PPII helical stretch spanning residues 31–39 from the central PPII helix in the polar face (i.e., helix III, as depicted in Fig. 1A) of the short HhAFP isoform. These nine residues correspond to three turns of the PPII helix that are properly aligned to form H-bonds with the helices above (helix I) and below (helix V), which lie on the same face and are spaced by 4.7 Å (Fig. 1A). The nine-residue stretch corresponding to residues 31–39 was cloned (i.e., repeated in space) four times to build a five-helical bundle. This leads to five monomeric, nine-residue helices lying within the same plane that are spaced by 4.7 Å from each other. The N- and C-termini of each PPII helix were capped with acetyl and methylamine moieties, respectively, to recreate the sequence continuity beyond the three turns that form the stretch. To isolate the pure PPII architecture from any other driving force for assembly arising from hydrophobic, polar or charged side chains, as well as to generalize the model for all types of Gly-rich PPII bundles, as those shown in Fig. 1A, all non-glycine residues, i.e., H32, N35, and N38, were mutated to glycine. Therefore, each PPII helix will contain 41 heavy atoms, giving rise to a total of 205 for the complete bundle (Fig. 1B). In order to analyze the interactions inherent to these structures, no type of solvent is included in the computational model.
Cooperative bundle assembly of PPII helices and H-Bonding patterns
Interaction energy calculations reveal enhanced H-Bonding strength and cooperativity in growing PPII helical bundles
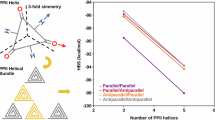
Building on the general model representing Gly-rich PPII bundles elaborated in the previous section, we next sought to characterize the interaction energy between every pair of PPII helices in such an assembly. In this model system, H-bonds drive PPII bundle assembly as there are no nonpolar, charged or aromatic groups which would give rise to hydrophobic, charge-charge or cation-\(\pi\) interactions (Fig. 1B). Thus, the interaction energy between two monomeric helices provides a direct estimation of the H-bonding strength (HBS). On this basis, we employed DFT-D to compute the interaction energies for dimers (a···b), trimers (a···bc and ab···c), tetramers (a···bcd, ab···cd and abc···d) and pentamers (a···bcde, ab···cde, abc···de, abcd···e) of PPII helices, according to the scheme shown in Fig. 1C. Note that for an n-mer system, there are (n-1) interacting interfaces and thus (n-1) HBS possible values. The only interaction that will not be analyzed is abcd···e since, as de is the last interface and e is the last PPII helix, the limit of the model has been reached and it is not possible to study how the addition of monomers affects the different computational parameters. The analysis of the computed HBS values shows that as PPII helices are added to the assembly, the interaction energies between monomers are strengthened. That is, the HBS between helices a and b in a five-helix bundle, abcde, is stronger than in a four-helix bundle, abcd, which is stronger than in a three-helix bundle, abc, which in turn is stronger than in a two-helix bundle ab, (i.e., | HBS(a···b)abcde | > |HBS(a···b)abcd | > |HBS(a···b)abc | > |HBS(a···b)ab | , Fig. 2 and Table 1). In this regard, we recall that we measure the interaction energy between the first two PPII helices (called a and b) as more and more helices are added on top of them, and therefore it is not the total interaction energy of the whole assembly, but rather that between the first two PPII helices what is being quantified. The same holds true for the HBS in the rest of the interfaces as helices are added to the assembly (i.e., | HBS(b···c)abcde | > |HBS(b···c)abcd | > |HBS(b···c)abc| and |HBS(c···d)abcde | > |HBS(c···d)abcd | , Fig. 2 and Table 1). This is proof of cooperative H-bonding interactions or HBC, as seen in amyloids17,18,19,20,21,22. To uncover the source of this cooperativity, we employed various computational methods to analyze the underlying H-bonding Interactions as described in the following sections.
The ab interface is indicated in blue; the bc interface, in red, and the cd interface, in yellow, as the color scheme depicted in Fig. 1C. Note that the HBS values decrease significantly as more PPII helices are added; this is a hallmark of HBC.
Identification of canonical and non-canonical H-Bonding patterns stabilizing Gly-Rich PPII helical bundles
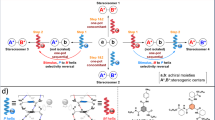
Having established that H-bonding in Gly-rich PPII assembly is cooperative, akin to amyloid assembly, we next ought to characterize which H-bonding interactions are present. We started by identifying the distinct potential stabilizing interactions by visual inspection of our model (Fig. 1B). For a PPII helix composed of three residues i, i + 1 and i + 2, which corresponds to exactly one turn of PPII helix, the carbonyl oxygen of the first residue, i, from one PPII helix interacts with the amide proton HN of the second residue, j + 1, in the next PPII helix to form a canonical CO···HN H-bond (Fig. 3A). In addition to this canonical interaction, we also identified the possible participation of the carbonyl oxygen from residue i in a non-canonical CO···HαCα interaction with the Hα from residue j in the second PPII helix. That is, the carbonyl oxygen from the first residue i in one helix participates in both the formation of a non-canonical H-bond with the Hα from the first residue j in the mating helix, and a canonical H-bond with the HN from the second residue j + 1. This non-canonical CO···HαCα interaction is denoted as a “front” H-bond (since it would be found in the same plane as each face of HhAFP) (Fig. 3A). Furthermore, we advance the existence of two additional non-canonical CO···HαCα H-bonds in Gly-rich PPII bundles that we here have denoted as an “inner” (since it would be found inside the core of HhAFP) and “outer” (since it would be found outside the core of HhAFP) (Fig. 3A). The supposed inner and outer H-bonds involve the two Hα protons of the last (i.e., third) residue in one PPII helix (i.e., residue i + 2). Particularly, in the “inner” H-bond one Hα proton would interact with the carbonyl oxygen from residue j + 2, whereas in the “outer” H-bond the second Hα proton from residue i + 2 would interact with the carbonyl oxygen from the second residue in the mating helix (i.e., j + 1, Fig. 3A). Since the results based on the HBS between PPII helices do not allow one to discern whether cooperativity holds every type of H-bond considered, we resorted to NBO and QTAIM analyses.
A Possible H-bonds/interactions between PPII helices in a 2D PPII helical bundle: CO···HN (purple), CO···HαCα front (yellow), CO···HαCα inner (orange) and CO···HαCα outer (magenta). B Simplified structures for CO···HN (formamide), CO···HαCα front (acetaldehyde), CO···HN + CO···HαCα front (acetamide), CO···HαCα inner (acetaldehyde), CO···HαCα outer (N-methylformamide) and CO···(Hα)2Cα inner+outer (N-(2-oxoethyl)formamide) H-bonds/interactions.
NBO and QTAIM analyses independently corroborate the existence and HBC of N-H···O = C and Cα-Hα···O = C inner H-bonds
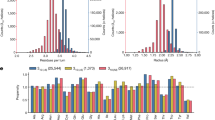
Regarding the NBO analysis (Tables 2 and 3 and Fig. 4), the occupancy of the σ* orbitals of the N-H bonds, q(σ*NH), and the stabilization energy corresponding to the electronic delocalization from not only the lone pairs (n) of the oxygen in those orbitals, but also from the σ and π orbitals of the C = O bonds, E(2)n(O)/σ(CO)/π(CO)→σ*(NH), indicate the progressive strengthening of the CO···HN H-bonds as the PPII helical assembly grows. This electron donation confers partial anionic character on the electron-acceptor species, then enhancing the capacity of the CONH units to act as an electron-donor in the next H-bond, which will be stronger. As for the CO···HαCα inner H-bonds, the q(σ*CαHα) and the E(2)n(O)/σ(CO)/π(CO)→σ*(CαHα) increase as PPII helices are added to the bundle, in a homologous way to what happened with CO···HN H-bonds, also involving the σ(CO) and π(CO) orbitals (Tables 2 and 3 and Fig. 4). These results are supported by the QTAIM analysis (supplementary material).
A Mean occupancy (q) of the corresponding σ*(NH/CαHα) orbitals and B mean stabilization energy (E(2)) of the corresponding n(O)/σ(CO)/π(CO)→σ*(NH/CαHα) electron delocalization for each CO···HN (squares), CO···HαCα front (circles), CO···HαCα inner (triangles) and CO···HαCα outer (inverted triangles) H-bond/interaction between PPII helices. The ab interface for the different systems is indicated in blue; the bc interface, in red, and the cd interface, in green. M06-2X/6-31 + G(d) level of theory.
Front H-bonds exist but show no HBC and outer H-bonds do not exist, according to NBO and QTAIM analysis
For the CO···HαCα front H-bonds, while E(2)n(O)→σ*(CαHα) slightly increases as PPII helices are added to the bundle, the q(σ*CαHα) decreases (Tables 2 and 3 and Fig. 4). This decrease of q(σ*CαHα) can be explained on the basis that the q(σ*NH) follows the opposite trend. Both types of σ* orbitals receive electrons from the same oxygen, so if the amount of electrons reaching one of the two different σ* orbitals increases as the assembly grows, the amount reaching the other σ* orbital will decrease. This phenomenon occurs even if the CONH units, which also give rise to the canonical H-bonds, have more and more electrons available to form the new H-bonds of the next interfaces, which is the reason why E(2)n(O)→σ*(CαHα) slightly increases as PPII helices are added to the assembly, since it depends on q(nO) and not on q(σ*CαHα) (Eq. 3). Finally, for the CO···HαCα outer interactions, the q(σ*CαHα) values remain practically constant as the PPII helical bundle grows and the negligible E(2)n(O)→σ*(CαHα) values do not have a clear tendency to change (Tables 2 and 3 and Fig. 4), suggesting their non-existence and hence non-cooperativity. These results are supported by QTAIM analyses (supplementary material).
Contribution of the different H-bonds to PPII helical bundle stability
The presence of a large number of non-canonical CO···HαCα H-bonds is important for the stability of PPII helical bundles7,8. In order to quantify the extent to which the distinct H-bonds contribute to the interaction energy between PPII helices, we have followed a divide-and-conquer approach by which mimicking molecular fragments resembling all types of H-bonds are subjected to independent HBS calculations38,39,40 (Fig. 3B). Within this framework, each CO···HN canonical H-bond is represented by a pair of formamide molecules. Front and inner non-canonical CO···HαCα H-bonds are mimicked by a pair of acetaldehyde molecules with different spatial orientations. Finally, outer non-canonical CO···HαCα interactions are mimicked using pairs of N-methylformamide molecules. It is necessary to emphasize that the mimetic molecules that give rise to the different interactions maintain the same position, and therefore the same distance, as in the complete PPII helices. In the following subsections, where we employ mimetics rather than full helices, we change the nomenclature from “PPII helices” to “layers of molecules” and “bundle assembly (of PPII helices) growth” for “stack (of layers of molecules) growth”. The term “interface” will now signify the region where mimetic molecules interact, rather than complete PPII helices.
Strengthened canonical N-H···O = C H-bonds versus electron-limited non-canonical Cα-Hα···O = C front interactions in expanding stacks
Regarding canonical H-bonds, the H-bonding strength or HBS among the different interfaces between formamide moieties (Fig. 3B) is strengthened as the stack grows (Fig. 5 and Table 4). These observations are reminiscent of the complete model of PPII bundles, and evince the cooperative reinforcement in CO···HN H-bonds between PPII helices. In the same way, Fig. 5 and Table 4 also show that, when the stack of layers of the acetaldehyde molecules that mimic CO···HαCα front H-bonds (Fig. 3B) grows, the HBS of the different interfaces is strengthened. One could interpret this as evidence of the cooperative reinforcement between non-canonical front H-bonds, which is in conflict with the previous NBO and QTAIM analyses of the full-length system that seem to indicate that they do not show HBC because the carbonyl oxygen’s electron density is shared with the much more favored canonical H-bonds. Perhaps the most intuitive way to visualize the effect that CO···HN H-bonds have on CO···HαCα front H-bonds, as well as to adequately quantify both interactions, is to calculate the HBS corresponding to acetamide molecules since they mimic both H-bonds at the same time (Fig. 3B). When the stack grows, the HBS of the different interfaces is reinforced, and so does the HBS of both the canonical and non-canonical front H-bonds separately, albeit less than the sum of both H-bonds isolated at each interface (Fig. 5 and Table 4). As explained earlier, this situation can be attributed to the limited quantity of electrons that oxygens may donate to the canonical and the non-canonical front H-bonds.
Mean HBS between the molecules that reproduce each CO···HN (formamide, squares), CO···HαCα front (acetaldehyde, circles), CO···HN + CO···HαCα front (acetamide, diamonds, and dashed line), CO···HαCα inner (acetaldehyde, triangles), CO···HαCα outer (N-methylformamide, inverted triangles) and CO···(Hα)2Cα inner+outer (N-(2-oxoethyl)formamide, hourglass and dashed line) H-bond/interaction. As in the Fig. 1C color scheme, the ab interface for the different systems is indicated in blue; the bc interface, in red, and the cd interface, in green.
Strengthened non-canonical Cα-Hα···O = C inner H-bonds versus dubious non-canonical Cα-Hα···O = C outer interactions in expanding stacks
Concerning non-canonical CO···HαCα inner H-bonds, while it is true that the HBS of the different interfaces between pairs of acetaldehyde molecules that mimic these interactions (Fig. 3B) become stronger upon addition of layers to the stack, it does so over a range of energy values that might fall within the uncertainty of the computational level of theory (Fig. 5 and Table 4). The values of the HBS between the N-methylformamide molecules that represent the non-canonical CO···HαCα outer interactions (Fig. 3B) are null within the error of the calculation (Fig. 5 and Table 4). Furthermore, these negligible interaction energies corresponding to the different interfaces do not show any tendency to increase or decrease, nor any evidence of cooperativity therefore, as the stack of mimicking molecules grows. Moreover, it is necessary to take into account that since these two putative non-canonical H-bonds share a common Cα, one interaction cannot be isolated from the other to obtain the HBS that would actually correspond to them. Consequently, we used pairs of N-(2-oxoethyl)formamide molecules to adequately mimic both potential H-bonds together (Fig. 3B), which gave rise to considerable HBS values showing a notable strengthening of the interactions across the different interfaces as the stack grows (Fig. 5 and Table 4). Additional QTAIM and SAPT analyses and results are shown in Supplementary Figs. 1–10 and Supplementary Tables 1–9. These interaction energy values are due almost exclusively to non-canonical inner H-bonds, not to the combination with the apparently non-existent non-canonical outer H-bonds that were initially proposed, as the previous NBO and QTAIM analyses of the full PPII bundle model showed. The findings of this section are in agreement with the NBO and QTAIM analyses of the mimicking systems.
Summing interaction energy contributions in PPII helical bundles provides insight into electron delocalization effects
Since the set of different molecules of acetamide (i.e., CO···HN + HαCα front H-bonds) and N-(2-oxoethyl)formamide (i.e., CO···(Hα)2Cα inner+outer H-bonds/interactions) shown in Fig. 3B are not a bona fide representation of the complete PPII bundle, it would not be expected that the sum of the different contributions to HBS would give the total HBS between full-length PPII helices. Indeed, Table 4 shows that the sum of the the CO···HN and the different CO···HαCα interaction energies produce HBS values initially smaller than the ones obtained when calculating the total HBS between full-length PPII helices shown in Table 1. However, as layers are added to the stack, the HBS values calculated as the sum of contributions exceed the global ones (Tables 4 and 1), demonstrating once again the presence of HBC in PPII helices and suggesting that in complete PPII helices, electron density would be delocalized not only in adjacent helices, but also along the peptide chain itself, generating macrodipoles previously reported for these protein secondary structures41,42.
Summary of interactions
The computational parameters studied here are summarized in Table S10 as the mean variation for each of the proposed H-bonds of the system’s five PPII helices. One should interpret this mean variation as the average of the differences between the computed parameter X for the last and first value for the interfaces ab (Xabcde – Xab), bc (Xabcde – Xabc) and cd (Xabcde – Xabcd). The NBO and QTAIM analyses, as well as the DFT-D calculation of interaction energies (HBS), lead us to suggest the existence and cooperativity of the CO···HN and CO···HαCα inner H-bonds; the existence, but non-cooperativity, of the CO···HαCα front H-bonds, and the non-existence, and therefore non-cooperativity, of the CO···HαCα outer H-bonds. The distance between the H-bond acceptor, the carbonyl oxygen, and the H-bond donor (HN or Hα) is another way to gauge the strengthening and cooperativity, or the lack, of these four types of possible H-bonds. In particular, non-canonical CO···HαCα outer H-bonds, whose existence cannot be established here, feature the longest distances (Table S11 and Fig. S11). Moreover, the shortest distances are seen for the CO···HN and CO···HαCα inner H-bonds and these distances decrease further as the bundle grows (Table S11 and Fig. S11), which is consistent with their HBC. In the case of non-canonical CO···HαCα front H-bonds, intermediate H-bonding distances without any trend to increase or decrease as the assembly grows are obtained (Table S11 and Fig. S11); this is consistent with the lack of HBC seen in the NBO, QTAIM and DFT-D results.
SAPT calculations are somewhat more precise, much more computationally costly, and provide insight into the contributions of the distinct interaction energy components to HBC. Our SAPT results, obtained at a higher level of theory than the rest of the methods, confirm that HBC exists in the cases already detected by DFT-D, NBO and QTAIM calculations, and evidence that it arises mostly from electrostatic effects and not induction or dispersion (Supplementary Tables S5, S6, S7 and Figs. S3, S5, S6, S7, S8, S9, S10). We refer the interested reader to the supplementary material for a specialized, in-depth description of the different interactions analyzed in this study.
Corroboration of the computational model
To further corroborate these findings, we have examined the experimental and computational NMR shift changes produced by H-bond formation. In solution, Gly-rich peptides have a tendency to adopt the extended PPII helix conformation over random coil or other structures43,44,45. Nevertheless, these situations cannot be easily distinguished spectroscopically since the two 1Hα chemical shifts of glycine residues are degenerate. However, we discovered that the two 1Hα of glycine residues in PPII bundles do display different chemical shifts, providing the first set of conformational chemical shifts, Δδ, for Gly-rich PPII bundles7. Experimental conformational shifts for 13Cα and 13CO of Δδ(Cα)exp = −0.59 (±0.65) ppm and Δδ(CO)exp = −0.20 (±1.12) ppm were obtained, while 1Hα nuclei in CO···HαCα H-bonded feature a characteristic conformational shift of Δδ(Hα)exp = −0.59 (±0.32) ppm. In our computational model (Fig. 1B), conformational chemical shifts for these nuclei were predicted to test our model’s ability to serve as a bona fide atomistic representation of PPII bundles. By comparing the corresponding shift for a nuclei in an extended PPII, isolated conformation with respect to that PPII in a bundle, the following DFT-GIAO (Gauge-Including-Atomic-Orbital) theoretical conformational shifts were obtained: Δδ(Cα)theo = −0.59 ppm, Δδ(CO)theo = −0.31 ppm and Δδ(Hα)theo = −0.61 ppm, for H-bonded nuclei. The similarity of the experimental and computational conformational shifts provides additional support for the existence of these H-bonds.
Discussion
The stability of Gly-rich PPII helical bundle domains is puzzling due to the high flexibility of glycine8. The stabilizing action of disulfide bonds46, the hydrophobic effect7, a limited denatured state conformational entropy15, and an important number of non-canonical CαHα···O = C H-bonds7,8 favor these proteins’ folded conformation. Here, we have rationally designed a computational model to scrutinize potential stabilizing contributions in Gly-rich PPII helical bundles that could explain these assemblies’ remarkable stability. Four possible H-bond types between individual helices; namely, canonical NH···O = C H-bonds and three types of putative CαHα···O = C H-bonds called front, inner and outer were investigated.
Based on the DFT-D results and corroborative evidence from the NBO, QTAIM and SAPT calculations, we have established the existence and cooperativity of canonical NH···O = C and inner CαHα···O = C H-bonds, the existence but non-cooperativity of non-canonical front CαHα···O = C H-bonds, and the non-existence and hence non-cooperativity of CαHα···O = C outer H-bonds. Regarding the relative merits of DFT-D versus SAPT calculations, we find that the marginally higher precision of the latter does not justify its much greater computation costs. Considering that many previous studies of amyloids showed cooperativity between canonical NH···O=C H-bonds17,18,19,20,21,22, it is not surprising that these interactions also show cooperativity between PPII helices, as previously proposed23. Furthermore, in both amyloids38 and PPII helical bundles, the NH electron acceptor in canonical H-bonds receives most of the available electron density from the O = C electron donor, which also forms front CαHα···O = C H-bonds. This prevents front CαHα···O = C non-canonical H-bonds from being cooperative. In contrast, inner CαHα···O = C non-canonical H-bonds, which do not have an analogy in amyloids, do exist and show cooperativity and their discovery is a major finding of this study. Our results strongly suggest than CαHα···O = C outer H-bonds do neither exist nor afford stability. This reconciles with experimental evidence where glycine residues engage both Hα in H-bonds only in the collagen-like structures24, but just one Hα in Gly-rich PPII bundles7,8.
While the major contribution to the PPII bundle stability comes from canonical H-bonds, the non-canonical ones also make significant contributions. In the interaction between two helices in our model before experiencing cooperativity, i.e., helices a and b without considering c, d, and e, each CO···HN H-bond represented by two formamide molecules contributes ≈6 kcal/mol (Table 4 and Fig. 5). This increases to ≈7 kcal/mol when also taking into account the extra ≈1 kcal/mol contributed by the CO···HαCα front H-bonds as calculated using a pair of acetamide molecules (Table 4 and Fig. 5). Furthermore, if we additionally consider the ≈3 kcal/mol arising from the CO···HαCα inner interaction, adequately represented by a pair of N-(2-oxoethyl)formamide molecules (Table 4 and Fig. 5), we reach an HBS of ≈10 kcal/mol per PPII helical turn, as opposed to the initial ≈6 kcal/mol corresponding only to the canonical H-bonds. Therefore, the important role of non-canonical H-bonds in the stability of PPII bundles is highlighted.
These results, supported by 1H and 13C NMR chemical shift data, establish canonical NH···O = C and non-canonical CαHα···O = C inner cooperative H-bonds as significant stabilizers of Gly-rich PPII helical bundle domains, especially the long, extensive ones. Indeed, H. harveyi produces two different sized isoforms of its AFP: one with six and with thirteen PPII helices (Fig. 1A)46. It is pertinent that the longer isoform, which is more active than the shorter one, is stable despite having only one disulfide bridge compared to two in the shorter isoform. We advance that HBC would make a larger stabilizing contribution in the case of the longer HhAFP isoform and could help account for its conformational stability. Moreover, a recent investigation of a homologous AFP from G. rainieri found that the insertion of two additional PPII helices augmented biological activity16.
Conclusions
In the past 20 years, several proteins with domains composed of glycine-rich polyproline II helix bundles have been uncovered, but the basis of their conformational stability is still not completely understood47. Here we conclude that the improved comprehension of the basis of PPII helical bundle stability provided by the discovery of HBC could also facilitate the use of Gly-rich PPII helices for protein design. In this regard, our findings raise important questions which should be addressed in future research. First, HN, CαHα and CO dipoles in isolated PPII helices are aligned such that they produce a macrodipole41,42, in analogy to that found in α-helices48. Based on this and previous reports of charge stabilization of macrodipoles in isolated PPII helices41,42, we advance that proteins or protein domains composed of PPII helical bundles may be stabilized by adding complementary charges to the helices’ termini. Secondly, regarding the doubt as to whether the parallel or antiparallel alignment of the PPII helices in bundles is more favorable49, our results suggest the later as it would juxtaposition complementary macrodipoles. Thirdly, our approach paves the way towards future studies of PPII helical networks with H-bond networks extending in multiple angles, which may lead to summation or cancelation of macrodipole effects.
Methods
Dispersion-corrected Density Functional Theory (DFT-D)
Density Functional Theory (DFT) calculations were carried out with the high-nonlocality M06-2X functional50 that has proven suitable to describe non-covalent interactions and is particularly amenable for extensively H-bonded biological systems21,51,52. As for the basis set, we chose the 6-31 + G(d) as an appropriate balance between accuracy and computational cost for the systems treated here that contain a large number of atoms. Following literature precedents dealing with large biomolecular systems containing multiple H-bonding interactions53,54,55, the D3 dispersion correction scheme56 was applied to all the calculations. Every interaction energy was counterpoise (CP)-corrected following the method proposed by Boys and Bernardi to consider the inherent basis set superposition error57. Finally, it is worth emphasizing that, in order to know the intrinsic energy of the systems under study, all quantum mechanical calculations have been carried out without the inclusion of any implicit or explicit solvent.
The geometry of the PPII helices in the bundle was optimized by keeping the Cαs frozen, but the amide group atoms and hydrogen atoms were allowed to undergo energy minimization to optimize H-bonding interactions and to account for the lack of hydrogens in the PDB structure used as a starting point (PDB ID: 3BOG). The energies of the monomers were computed with the same structure that they present in the optimized bundle to ensure that the interaction energy strictly corresponds to the established H-bonds. In this way, the HBS of XY interface can be calculated using Eq. 1:
For example, the HBS of bc interface for the case of five PPII helices would be obtained through Eq. 2:
For the layers of the mimetic molecules, which maintain the same geometry as the corresponding atoms in the optimized PPII helical bundle, the same process was followed as with the complete PPII helices to calculate the HBS relative to each type of H-bond. The rest of the computational parameters from NBO and QTAIM analyses can be obtained directly at the interface of interest for a given number of PPII helices, while in SAPT calculations the interaction energy is determined without computing the total energy of the monomers or n-mers. Each of the points that appear in the different graphs of this study correspond to the images presented in Fig. 1C.
Natural Bonding Orbital (NBO)
The NBO analysis58 has been performed at the same level of theory, M06-2X/6-31 + G(d), to test the H-bonding nature of the interactions between glycine residues in the Gly-rich PPII bundles studied here. In the context of the NBO methodology, H-bonds are characterized as an electron delocalization from a lone pair (n) of the H-acceptor atom (e.g., an O atom) onto the sigma antibonding (σ*) orbital of the covalent bond between the H-donor atom and the H atom itself (e.g., an N-H or C-H bond). This interaction can be quantitatively described through the stabilization energy, E(2), which is estimated based on the Second Order Perturbation Theory:
where qi is the occupancy of the electron-donor orbital i, Ei and Ej are the energies of i and j, respectively, and Fij is the Fock or Kohn-Sham matrix element between both NBOs. The higher the value of qj and E(2), the greater the i → j electron transfer, leading to a greater stabilization of the molecule.
Quantum Theory of Atoms in Molecules (QTAIM)
Canonical CO···HN and non-canonical CO···HαCα H-bonds in the Gly-rich PPII bundles have been also analyzed using the QTAIM approach59,60 as described in detail in the supplementary material.
Symmetry-Adapted Perturbation Theory (SAPT)
Canonical CO···HN and non-canonical CO···HαCα interactions in the Gly-rich PPII bundles have been further explored using the SAPT methodology61,62 as described in detail in the supplementary material.
Theoretical chemical shift calculations
Nuclear Magnetic Resonance (NMR) shielding tensors of both PPII helices isolated and in bundles were calculated using the gauge-including-atomic-orbital (GIAO) method63,64 at the M062X/6-31 + G(d) computational level too. The corresponding chemical shifts, δ, are obtained as the difference between isotropic chemical shieldings in the nucleus of interest in our model and in the compound used as a reference, σiso and σiso,ref, respectively:
The conformational chemical shifts, Δδ, of a protein are the experimental chemical shifts, δ2, relative to values expected for statistical coil, δ1, which translates into the difference between two different chemical shifts of the same nucleus for which the same reference compound has been used:
Computationally, in DFT-GIAO calculations structures are treated as static instead of dynamic species. Bearing this in mind, and considering that in solution Gly-rich sequences adopt extended PPII helix conformations over statistical coil43,44,45, the theoretical Δδ values have not been calculated using a statistical coil state as the reference. Instead, Δδ values have been obtained as the difference between the σiso of nuclei when the PPII helix is in the model bundle (σiso,2), geometrically optimized as described in the DFT-D calculations section, with respect to the PPII helix in isolation (σiso,1), freezing backbone atoms to maintain the positions seen in the crystallographic structure used as a starting point to create the model system (PDB ID: 3BOG), while 1H nuclei that are absent in this X-ray structure were optimized.
The Quantum Chemistry package used to perform the geometry optimizations, and single point and NMR spectroscopic shielding calculations was Gaussian 1665. The software packages used for the visualization and modeling of the PPII helical-bundles were GaussView 666, Pymol 2.3.067 and Jmol 1468. For the NBO analysis, NBO Version 3.169 was used as implemented in Gaussian 16. The QTAIM analysis was carried out using the Multiwfn 3.8 package70. Finally, SAPT calculations were performed using the Psi4 software71.
Data availability
All relevant data are included in the manuscript main text and supplementary material.
References
Kuhlman, B. & Bradley, P. Advances in protein structure prediction and design. Nat. Rev. Mol. Cell Biol. 20, 681–697 (2019).
Baker, E. G., Bartlett, G. J., Porter Goff, K. L. & Woolfson, D. N. Miniprotein design: past, present, and prospects. Acc. Chem. Res. 50, 2085–2092 (2017).
Shiga, S. et al. Domain-swapping design by polyproline rod insertion. ChemBioChem 20, 2454–2457 (2019).
Mompeán, M., Oroz, J. & Laurents, D. V. Do polyproline II helix associations modulate biomolecular condensates? FEBS Open Bio 11, 2390–2399 (2021).
Lamers, B. A. G. et al. Oligodimethylsiloxane-oligoproline block co-oligomers: the interplay between aggregation and phase segregation in bulk and solution. J. Am. Chem. Soc. 143, 4032–4042 (2021).
Pentelute, B. L. et al. X-Ray structure of snow flea antifreeze protein determined by racemic crystallization of synthetic protein enantiomers. J. Am. Chem. Soc. 130, 9695–9701 (2008).
Treviño, M. Á. et al. The singular NMR fingerprint of a polyproline II helical bundle. J. Am. Chem. Soc. 140, 16988–17000 (2018).
Gates, Z. P. et al. Perplexing cooperative folding and stability of a low-sequence complexity, polyproline 2 protein lacking a hydrophobic core. Proc. Natl Acad. Sci. USA 114, 2241–2246 (2017).
Bartlett, G. J., Newberry, R. W., VanVeller, B., Raines, R. T. & Woolfson, D. N. Interplay of hydrogen bonds and N→π* interactions in proteins. J. Am. Chem. Soc. 135, 18682–18688 (2013).
Newberry, R. W. & Raines, R. T. The N→π* interaction. Acc. Chem. Res. 50, 1838–1846 (2017).
Zhai, L., Otani, Y. & Ohwada, T. Uncovering the networks of topological neighborhoods in β-strand and amyloid β-sheet structures. Sci. Rep. 9, 10737 (2019).
Scholl, C. L., Tsuda, S., Graham, L. A. & Davies, P. L. Crystal waters on the nine polyproline type II helical bundle springtail antifreeze protein from Granisotoma rainieri match the ice lattice. FEBS J. 288, 4332–4347 (2021).
Mok, Y.-F. et al. Structural basis for the superior activity of the large isoform of snow flea antifreeze protein. Biochemistry 49, 2593–2603 (2010).
Li, T. et al. Structural basis for ligand reception by anaplastic lymphoma kinase. Nature 600, 148–152 (2021).
Treviño, M. Á. et al. Insight into polyproline II helical bundle stability in an antifreeze protein denatured state. Biophys. J. 121, 4560–4568 (2022).
Scholl, C. L. & Davies, P. L. Protein engineering of antifreeze proteins reveals that their activity scales with the area of the ice-binding site. FEBS Lett. 597, 538–546 (2023).
Viswanathan, R., Asensio, A. & Dannenberg, J. J. Cooperative hydrogen-bonding in models of antiparallel β-sheets. J. Phys. Chem. A 108, 9205–9212 (2004).
Tsemekhman, K., Goldschmidt, L., Eisenberg, D. & Baker, D. Cooperative hydrogen bonding in amyloid formation. Protein Sci. 16, 761–764 (2007).
Mompeán, M. et al. Structural evidence of amyloid fibril formation in the putative aggregation domain of TDP-43. J. Phys. Chem. Lett. 6, 2608–2615 (2015).
Plumley, J. A. & Dannenberg, J. J. The importance of hydrogen bonding between the glutamine side chains to the formation of amyloid VQIVYK parallel β-sheets: an ONIOM DFT/AM1 study. J. Am. Chem. Soc. 132, 1758–1759 (2010).
Mompeán, M., Nogales, A., Ezquerra, T. A. & Laurents, D. V. Complex system assembly underlies a two-tiered model of highly delocalized electrons. J. Phys. Chem. Lett. 7, 1859–1864 (2016).
Mompeán, M., Baralle, M., Buratti, E. & Laurents, D. V. An amyloid-like pathological conformation of TDP-43 is stabilized by hypercooperative hydrogen bonds. Front. Mol. Neurosci. 9, 125 (2016).
Plumley, J. A., Tsai, M. & Dannenberg, J. J. Aggregation of capped hexaglycine strands into hydrogen-bonding motifs representative of pleated and rippled β-sheets, collagen, and polyglycine I and II crystal structures. a density functional theory study. J. Phys. Chem. B 115, 1562–1570 (2011).
Bella, J. & Berman, H. M. Crystallographic evidence for Cα–H···O = C hydrogen bonds in a collagen triple helix. J. Mol. Biol. 264, 734–742 (1996).
Ramachandran, G. N., Sasisekharan, V. & Ramakrishnan, C. Molecular structure of polyglycine II. Biochim. Biophys. Acta 112, 168–170 (1966).
Krimm, S. & Bandekar, J. Vibrational spectroscopy and conformation of peptides, polypeptides, and proteins. Adv. Protein Chem. 38, 181–364 (1986).
Kar, T. & Scheiner, S. Comparison of cooperativity in CH···O and OH···O hydrogen bonds. J. Phys. Chem. A 108, 9161–9168 (2004).
Blatchford, M. A., Raveendran, P. & Wallen, S. L. Spectroscopic studies of model carbonyl compounds in CO2: evidence for cooperative C − H···O interactions. J. Phys. Chem. A 107, 10311–10323 (2003).
Li, Q., An, X., Gong, B. & Cheng, J. Spectroscopic and theoretical evidence for the cooperativity between red-shift hydrogen bond and blue-shift hydrogen bond in DMSO aqueous solutions. Spectrochim. Acta A. Mol. Biomol. Spectrosc. 69, 211–215 (2008).
Esrafili, M. D., Behzadi, H. & Hadipour, N. L. Theoretical study of N–H· · ·O hydrogen bonding properties and cooperativity effects in linear acetamide clusters. Theor. Chem. Acc. 121, 135–146 (2008).
Ludwig, R. Cooperative hydrogen bonding in amides and peptides. J. Mol. Liq. 84, 65–75 (2000).
Parthasarathi, R., Raman, S. S., Subramanian, V. & Ramasami, T. Bader’s electron density analysis of hydrogen bonding in secondary structural elements of protein. J. Phys. Chem. A 111, 7141–7148 (2007).
Arnold, W. D. & Oldfield, E. The chemical nature of hydrogen bonding in proteins via NMR: J-Couplings, chemical shifts, and AIM theory. J. Am. Chem. Soc. 122, 12835–12841 (2000).
Alkorta, I. et al. Cooperativity in multiple unusual weak bonds. Theor. Chem. Acc. 126, 1–14 (2010).
Venkataramanan, N. S. & Suvitha, A. Nature of bonding and cooperativity in linear DMSO clusters: a DFT, AIM and NCI analysis. J. Mol. Graph. Model. 81, 50–59 (2018).
Sánchez-Sanz, G., Trujillo, C., Alkorta, I. & Elguero, J. Understanding regium bonds and their competition with hydrogen bonds in Au2:HX complexes. ChemPhysChem 20, 1572–1580 (2019).
Jezierska, A. & Panek, J. J. Cooperativity of hydrogen bonding network in microsolvated biotin, the ligand of avidin class proteins. J. Mol. Model. 25, 361 (2019).
Scheiner, S. Contributions of NH···O and CH···O hydrogen bonds to the stability of β-sheets in proteins. J. Phys. Chem. B 110, 18670–18679 (2006).
Scheiner, S. Theoretical analysis of the contributions made by CH••OH bonds to protein structure. Curr. Org. Chem. 14, 106–128 (2010).
Scheiner, S. Dissection of the factors affecting formation of a CH ∙ ∙ ∙ O H-bond. A case study. Crystals 5, 327–345 (2015).
Kuemin, M., Schweizer, S., Ochsenfeld, C. & Wennemers, H. Effects of terminal functional groups on the stability of the polyproline II structure: a combined experimental and theoretical study. J. Am. Chem. Soc. 131, 15474–15482 (2009).
Huang, K.-Y. & Horng, J.-C. Impacts of the terminal charged residues on polyproline conformation. J. Phys. Chem. B 123, 138–147 (2019).
Ohnishi, S., Kamikubo, H., Onitsuka, M., Kataoka, M. & Shortle, D. Conformational preference of polyglycine in solution to elongated structure. J. Am. Chem. Soc. 128, 16338–16344 (2006).
Bykov, S. & Asher, S. Raman studies of solution polyglycine conformations. J. Phys. Chem. B 114, 6636–6641 (2010).
Mompeán, M. et al. Glycine rich segments adopt polyproline II helices: implications for biomolecular condensate formation. Arch. Biochem. Biophys. 704, 108867 (2021).
Graham, L. A. & Davies, P. L. Glycine-rich antifreeze proteins from snow fleas. Science 310, 461–461 (2005).
Rodríguez, C. S. & Laurents, D. V. Architectonic principles of polyproline II helix bundle protein domains. Arch. Biochem. Biophys. 756, 109981 (2024).
Wada, A. The Alpha-Helix as an Electric Macro-Dipole. Adv. Biophys. 9, 1–63 (1976).
Crick, F. H. C. & Rich, A. Structure of polyglycine II. Nature 176, 780–781 (1955).
Zhao, Y. & Truhlar, D. G. The M06 suite of density functionals for main group thermochemistry, thermochemical kinetics, noncovalent interactions, excited states, and transition elements: two new functionals and systematic testing of four M06-class functionals and 12 other functionals. Theor. Chem. Acc. 120, 215–241 (2008).
Mompeán, M. et al. Molecular mechanism of the inhibition of TDP-43 amyloidogenesis by QBP1. Arch. Biochem. Biophys. 675, 108113 (2019).
Martín-Pintado, N. et al. Dramatic effect of furanose C2′ substitution on structure and stability: directing the folding of the human telomeric quadruplex with a single fluorine atom. J. Am. Chem. Soc. 135, 5344–5347 (2013).
Arey, J. S., Aeberhard, P. C., Lin, I.-C. & Rothlisberger, U. Hydrogen bonding described using dispersion-corrected density functional theory. J. Phys. Chem. B 113, 4726–4732 (2009).
DiLabio, G. A., Johnson, E. R. & Otero-de-la-Roza, A. Performance of conventional and dispersion-corrected density-functional theory methods for hydrogen bonding interaction energies. Phys. Chem. Chem. Phys. 15, 12821–12828 (2013).
Boese, A. D. Density functional theory and hydrogen bonds: are we there yet? ChemPhysChem 16, 978–985 (2015).
Grimme, S., Antony, J., Ehrlich, S. & Krieg, H. A consistent and accurate ab initio parametrization of Density Functional Dispersion Correction (DFT-D) for the 94 Elements H-Pu. J. Chem. Phys. 132, 154104 (2010).
Boys, S. F. & Bernardi, F. The calculation of small molecular interactions by the differences of separate total energies. Some procedures with reduced errors. Mol. Phys. 19, 553–566 (1970).
Reed, A. E., Curtiss, L. A. & Weinhold, F. Intermolecular interactions from a natural bond orbital, donor-acceptor viewpoint. Chem. Rev. 88, 899–926 (1988).
Bader, R. F. W. Atoms in Molecules. Acc. Chem. Res. 18, 9–15 (1985).
Bader, R. F. W. Atoms in Molecules: A Quantum Theory (Clarendon Press, 1990).
Jeziorski, B., Moszynski, R. & Szalewicz, K. Perturbation theory approach to intermolecular potential energy surfaces of van der Waals complexes. Chem. Rev. 94, 1887–1930 (1994).
Hohenstein, E. G. & Sherrill, C. D. Density fitting of intramonomer correlation effects in symmetry-adapted perturbation theory. J. Chem. Phys. 133, 014101 (2010).
Ditchfield, R. Self-consistent perturbation theory of diamagnetism. Mol. Phys. 27, 789–807 (1974).
Wolinski, K., Hinton, J. F. & Pulay, P. Efficient implementation of the gauge-independent atomic orbital method for NMR chemical shift calculations. J. Am. Chem. Soc. 112, 8251–8260 (1990).
Frisch, M. J. et al. Gaussian 16, Rev. A.01 (Wallingford, CT, 2016).
Dennington, R., Keith, T. A. & Millam, J. M. GaussView, version 6.0.16 (Semichem Inc., Shawnee Mission, KS, USA, 2016).
DeLano, W. L. et al. Pymol: an open-source molecular graphics tool. CCP4 Newsl. Protein Crystallogr. 40, 82–92 (2002).
Jmol Development Team. (2016).
Glendening, E. D., Reed, A. E., Carpenter, J. E., Weinhold, F. NBO Version 3.1 (Gaussian Inc., Pittsburgh, 2003).
Lu, T. & Chen, F. Multiwfn: a multifunctional wavefunction analyzer. J. Comput. Chem. 33, 580–592 (2012).
Smith, D. G. A. et al. PSI4 1.4: open-source software for high-throughput quantum chemistry. J. Chem. Phys. 152, 184108 (2020).
Acknowledgements
This work has been funded by grants PID2019-109306RB-I00 & PID2022-137806OB-I00 (D.V.L.) and PID2020-113907RA-I00 (M.M.) from MCIN/AEI/10.13039/501100011033. Co-funded by the European Union (ERC, 101042403 – BiFOLDOME). Views and opinions expressed are however those of the authors only and do not necessarily reflect those of the European Union or the European Research Council. Neither the European Union nor the granting authority can be held responsible for them. The authors also gratefully acknowledge the SGAI-CSIC for its assistance and help while using the DRAGO Supercomputer.
Author information
Authors and Affiliations
Contributions
Funding acquisition: D.V.L. and M.M.; Conceptionalization: R.L.S., D.V.L., and M.M.; Data acquisition and analysis: R.L.S. and M.M.; Data Interpretation: R.L.S. and M.M.; Manuscript preparation: R.L.S., D.V.L., and M.M.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Chemistry thanks Konstantin Röder and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
López-Sánchez, R., Laurents, D.V. & Mompeán, M. Hydrogen bonding patterns and cooperativity in polyproline II helical bundles. Commun Chem 7, 191 (2024). https://doi.org/10.1038/s42004-024-01268-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42004-024-01268-2
This article is cited by
-
ACSM5 c.1273 C> A Polymorphism Promotes Malignant Progression of Lung Adenocarcinoma in Xuanwei Region
Applied Biochemistry and Biotechnology (2026)
-
Orientation affects hydrogen bonding cooperativity in polyproline II helical bundles
Communications Chemistry (2025)
-
Hydrogen atom dynamics in double-well hydrogen bond potentials: a case study of O–H...O bonds of potassium dihydrogen phosphate, tris-potassium hydrogen bisulphate and urea–phosphoric acid crystals
Bulletin of Materials Science (2025)