Abstract
Machine learning (ML) and artificial intelligence (AI) techniques are transforming the way chemical reactions are studied today. Datasets from high-throughput experimentation (HTE) are generated to better understand the reaction conditions crucial for outcomes such as yields and selectivities. However, it is often overlooked that datasets from such designed experiments possess a specific structure, which can be captured by a statistical model. Ignoring these data structures when applying ML/AI algorithms can result in misleading conclusions. In contrast, leveraging knowledge about the data-generating process yields reliable, interpretable, and comprehensive insights into reaction mechanisms. A particularly complex dataset is available for the Buchwald-Hartwig amination. Using this dataset, a statistical model for such HTE-generated chemical data is introduced, and a parameter estimation algorithm is developed. Based on the estimated model, new insights into the Buchwald-Hartwig amination are discussed. Our approach is applicable to a wide range of HTE-generated data for chemical reactions and beyond.
Similar content being viewed by others
Introduction
The field of chemistry is currently undergoing a paradigm shift driven by the adoption of data-driven modeling techniques, particularly machine learning (ML) and artificial intelligence (AI) algorithms. These algorithms can potentially transform vast amounts of chemical data into valuable predictions, fundamentally changing the way chemists approach research1,2,3. ML algorithms are increasingly being employed to predict catalyst efficiency4,5, regioselectivity6,7, and chemical properties such as toxicity and electrophilicity8,9,10,11,12. Beyond property prediction, ML/AI methods have been successfully applied in drug discovery13,14, the development of new materials15,16, retrosynthetic analysis17, the optimization of reaction conditions18,19,20,21, and even the creation of synthesis-capable robotic platforms22. Furthermore, advances in modern quantum chemistry and computational techniques now enable the accurate calculation of a wide range of molecular properties. These computed descriptors can be combined with experimental output data to further enhance the predictive capabilities of ML algorithms.
Typically, ML/AI methods require large datasets as input. In many fields today, such datasets are increasingly generated through high-throughput experiments (HTEs). HTE is a scientific approach that combines laboratory automation, optimized experimental design, and the rapid execution of parallel or sequential experiments. It allows for the systematic acquisition of large, reproducible datasets using robotics, miniaturized workflows, and computational data management pipelines. This methodology accelerates discovery, enhances reproducibility, and supports data-driven decision-making. Consequently, HTE has had a transformative impact across diverse scientific and engineering disciplines, including chemistry23,24,25, biology26,27,28, materials science29,30, and other related fields31,32,33.
The goal of any HTE is to investigate a specific variable, referred to as the dependent or target variable or a response, e.g., the yield of a chemical reaction, the success probability of a genetic variant, or ionic conductivity. This is achieved by defining a set of experimental conditions under which this target variable is measured. In essence, an HTE involves systematically obtaining measurements of the target variable across combinations of experimental conditions that are relevant to the study.
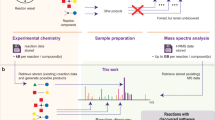
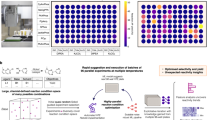
To be more specific, let us consider a dataset related to palladium-catalyzed C-N activation, commonly known as the Buchwald-Hartwig amination (Fig. 1a). Despite its importance in organic synthesis, this reaction is notoriously sensitive and heavily dependent on carefully chosen (often empirically determined) reaction conditions23,34,35,36,37 obtained an impressive Buchwald-Hartwig amination dataset by exploiting HTE reactions, which we will use as a case study in this work. In this HTE,23 selected 22 isoxazole additives (23 were originally considered, but one was excluded from analysis), 15 aromatic halides, 4 palladium catalyst ligands, and 3 bases, as illustrated in Fig. 1b. Each reaction yield in the dataset was subsequently measured under each of the 22 × 15 × 4 × 3 = 3 960 possible combinations of these reaction conditions. In addition to yield values, a set of descriptors of all reaction conditions was provided. Specifically, 19 descriptors were computed for additives, 27 for aryl halides, 10 for bases, and 64 for ligands. Note that these values were not obtained in the HTE but rather chosen as descriptors of selected reaction conditions. Finally,23 applied various ML techniques in order to model and predict obtained yield values in terms of selected descriptors.
The work of23 has received considerable attention and has inspired numerous follow-up studies. By mid-2025, the publication had garnered over 1 100 citations, according to Google Scholar. Various standard ML/AI methods, including gradient boosting, deep forest, neural networks, and k-nearest neighbors, have since been applied in similar contexts, both to the Buchwald-Hartwig amination and to similar datasets from other chemical reactions generated via HTE24,38,39,40,41,42,43.
To identify the most suitable approaches for analyzing any HTE-generated data, it is first essential to understand the type and structure of the data in order to situate it appropriately within a regression framework. In particular, the selection of estimation or prediction methods should take into account not only the type of the target variable but also that of the explanatory features.
For example, in the Buchwald-Hartwig amination dataset, the target variable is the reaction yield, a continuous variable ranging from 0 to 100. This outcome is modeled as a function of four features: additives, aryl halides, bases, and ligands, which collectively determine the reaction performance. In statistical terms, all four features are categorical variables; that is, the values of each feature correspond to discrete, chemically distinct species or conditions with no inherent order. For instance, different additives interact with catalytic intermediates through fundamentally distinct mechanisms, such as coordination, electron donation or withdrawal, steric hindrance, hydrogen bonding, or dispersion interactions, and thus cannot be meaningfully represented on a continuous or ordered scale. Hence, in this case, we are dealing with a regression problem involving a continuous target variable and four categorical features. As will be demonstrated in this work, the data-generating process of such data can be captured by a simple (generalized) linear model, making application of ML/AI techniques superfluous. Moreover, compared to ML/AI approaches, a linear model is fully interpretable, allowing for deeper insight into the underlying reaction mechanisms. In other words, if the data-generated process from an HTE follows a simple parametric model, ML/AI techniques cannot offer any clear advantage, both in terms of modeling and prediction.
In the broader landscape of HTE-generated data across different scientific fields, all possible combinations of data types can be encountered. While the target variable is typically continuous, the features may be continuous, categorical, or a mixture of both. In settings where both the target and features are continuous, ML/AI methods may offer advantages over simple parametric alternatives in terms of predictive performance, although this typically comes at the expense of interpretability and system-level understanding. Nevertheless, in many disciplines and especially in chemistry, a substantial share of HTE-generated datasets involves categorical features only.
In this work, we focus on HTE-generated datasets with a target variable from the exponential family of distributions and categorical features. Using the Buchwald-Hartwig amination dataset as a case study, we discuss its properties and subsequently propose a simple linear statistical model that accurately captures the corresponding data-generating process. We show that standard estimation techniques are unsuitable for parameter estimation within this model. Subsequently, we develop a tailored estimation method that enables both reliable identification and meaningful interpretation of model parameters, thereby providing deeper insight into the most influential components of the Buchwald-Hartwig amination reaction. Although our approach is demonstrated using a single case study, the proposed method is readily extendable to other datasets characterized by a target variable from the exponential family and categorical features. To substantiate this claim, in the Supplementary Section 1, we provide a link to the Python package that implements our approach for an arbitrary number of features (ranging from two to five), together with the code and analysis of two additional datasets containing three and five categorical features, respectively.
Results
Structures in Buchwald-Hartwig amination data
Let us first exploit the data collected in ref. 23 in detail. There 3 960 values of the yield and each of 120 descriptors are available, which result from measurements under all possible 22 × 15 × 4 × 3 = 3 960 combinations of reaction components. These reaction components will be referred to as additives, halides, ligands and bases, and statistically are categorical variables or factors that take 22, 15, 4, and 3 possible unordered values (known as factor levels), respectively. Additionally, a matrix of 120 descriptors is provided, which contains 19 continuous descriptors of additives, 27 descriptors of halides, 10 descriptors of bases, and 64 descriptors of ligands. First, we discuss the statistical properties of the yield; then, we examine the structures within the matrix of descriptors; and finally, we present a model that adequately captures the data-generating process, allowing for meaningful interpretation.
In a statistical framework, the target variable, in our case, the reaction yield, is typically treated as a random variable and often assumed to follow a normal distribution. However, as the histogram in Fig. 2a clearly shows, the distribution of yields deviates substantially from normality: it has bounded support (restricted to the [0,100] interval, in line with the physical meaning of yield) and is right-skewed (with a greater number of reactions resulting in lower yields). Therefore, we propose modeling the yield using the continuous Bernoulli distribution. Details on this distribution are provided in the Supplementary Section 2. The continuous Bernoulli distribution fits the data reasonably well (see Fig. 2a) and, importantly, belongs to the exponential family.
Let us now examine the matrix of descriptors. It is straightforward to verify that the matrix of 120 descriptors, as provided in ref. 23, has rank 39. In the Supplementary Section 3, we demonstrate that by adding three random descriptors for the additives, the resulting matrix with 123 columns would have rank 41 and can be uniquely linearly transformed into a matrix with dummy (or one-hot) coding, which reflects the underlying experimental design. This implies that the matrix of descriptors contains, up to a linear transformation, only information about combinations of reaction components that lead to the observed yield. Hence, the matrix of descriptors can be ignored.
With this, we arrive at a model in which the target variable follows a continuous Bernoulli distribution, while four categorical variables additives, halides, bases, and ligands serve as explanatory features for this target. This can be formulated in a simple statistical model that adequately captures the data-generating process of the Buchwald-Hartwig amination dataset:
Here \(g:[0,1]\to {\mathbb{R}}\) is the canonical link function of the continuous Bernoulli distribution, which has no closed-form expression but is shown in (Fig. 2b) (see also Supplementary Section 2). Furthermore, \({\mathbb{E}}\) denotes expectation, g−1(μ0) is an overall mean of the yield, αi, i = 1, …, 22 is the contribution (also known as a treatment effect) of the i-th level of factor additives, βj, j = 1, …, 15 is the contribution of the j-th level of the factor halides, γk, k = 1, 2, 3 is the contribution of the k-th level of the factor bases and δl, l = 1, 2, 3, 4 is the contribution of the l-th level of the factor ligands. Further terms denote corresponding interaction effects. For example, (αβ)ij denotes the interaction effect of the i-th level of factor additives and j-th level of the factor halides. The number of parameters in the model (1) is 7 360, and there are 3 400 constraints necessary for identifiability. The full list of these constraints is given in the Supplementary Section 5.
According to the model (1), each parameter describes the effect of the corresponding factor level (e.g., i-th additive) or factor level combination (e.g., i-th additive with j-th halide) on the yield value compared to the baseline μ0. In particular, a positive parameter indicates that the corresponding factor level or factor level combination leads to higher yield values compared to the baseline (interpreted as a stabilizing effect), whereas a negative parameter indicates lower yields relative to the baseline (interpreted as a destabilizing effect).
The interactions between factor levels are particularly valuable from a chemical standpoint. These interactions may reflect cooperative or antagonistic effects on the catalytic cycle. For example, certain i-th additive with j-th base interactions may stabilize or destabilize key intermediates or transition states differently. Statistically and chemically, understanding these interactions allows us to identify conditions that favor reaction selectivity or efficiency. We believe that this modeling approach – integrating statistical design principles with chemical knowledge – provides a robust framework not only for predicting experimental outcomes within the studied domain but also for generating chemically meaningful hypotheses about the underlying mechanisms. This, in turn, can guide future experimentation and mechanistic studies more systematically and efficiently.
At this point, it is important to emphasize that categorical features naturally limit what can be learned from the model (1). In particular, one cannot predict yield values for unseen levels of factors outside of experimental design. However, one can predict or actually fill in (and this is what23 and follow-up works ultimately do) the missing values within the experimental design. That is, one can perform only a share of 3960 experiments, e.g., 70%, and then predict the yield values for the remaining 30% missing combination of factor levels.
Altogether, model (1) is a (generalized) linear model, and its parameters, once properly estimated, can be used to understand how specific reaction components and their interactions contribute to the yield. For example, one can identify reaction conditions that lead to higher or lower yields or conditions that do not affect the yield at all. This contrasts with black-box ML/AI techniques, which are well-suited for prediction within the experimental design but fail to provide an explanation of the system. Naturally, model (1) can also be used to predict the yield within the experimental design. In the Supplementary Section 7, we compare such predictions with those obtained using several of the most successful ML algorithms, demonstrating that the difference in performance is negligible. We conclude that ML/AI methods cannot bring any advantages over simple linear models if the features are categorical variables.
Estimation results
It turns out that estimating the model (1) requires non-standard approaches. In Section 3 we explain the inherent problems of the model and describe the algorithm we developed.
In the following, we present and analyze the estimated parameters of model (1). The intercept term μ0 in the model (1), which serves as a baseline and represents the transformed overall mean of the yield, was estimated to be − 6.9. Applying the inverse link function g−1( − 6.9) = 0.14, gives an estimator for the expectation of the yield, indicating that the mean yield in this HTE is notably low. The first column of Table 1 reports the estimated main effects. More precisely, the values for 22 additives given in the first column of Table 1 are estimates for parameters αi, i = 1, …, 22 in equation (1). In the same way, 15 halide values are estimates for βj, j = 1, …, 15, 3 base values are estimates for γk, k = 1, 2, 3 and 4 ligand values are estimates for δl, l = 1, …, 4. Fig. 3 provides a complementary visual representation of the estimated main effects, linking the factor level labels to the corresponding chemical structures.
Shown are the estimated coefficients for all levels of additives (A1–A22), halides (H1–H15), bases (B1–B3), and ligands (L1–L4) corresponding to the model (1). The x-axis indicates the magnitude of the main effects (dimensionless), where positive values enhance and negative values reduce yield relative to baseline. Panel colors match the factor coding of Fig. 1 and serve to distinguish component types.
The second column of Table 1 lists the 44 absolute largest two-way interaction effects. For example, an interaction of additive 9 with halide 1 is an estimator for the parameter (αβ)91. Notably, all four-way interaction terms were estimated to be exactly zero, and only a few three-way interactions were found to be non-zero. These remaining three-way interactions are relatively minor in magnitude, especially compared to the main effects and the dominant two-way interactions, suggesting that higher-order interactions play a limited role in this setting. A complete table of all estimated coefficients is available through the link provided in the Supplementary Section 1. In total, 415 parameters were estimated to be non-zero out of all 7 360 parameters of the model (2), reflecting a high degree of sparsity in the fitted model. This sparsity substantially simplifies the interpretation of the estimated model.
The estimation results can also be rationalized from a chemical viewpoint. First, it is chemically intuitive that halides exert the most significant influence on the yield, as the nature of the leaving group (Cl, Br, I) directly affects the oxidative addition step44,45,46, which often involves the rate-determining states (turnover-determining intermediate TDI and turnover-determining transition state TDTS) in Buchwald-Hartwig amination47,48. A better leaving group typically lowers the activation energy, thus significantly enhancing reaction efficiency. However, it is worth noting that, in some cases, iodides have been shown to be less effective counterparts in Pd-catalyzed C-N coupling reactions than other halides due to side reactions. This observation further highlights the intrinsic complexity of the catalytic process under consideration49.
The second most influential factor, additives, often plays subtle but critical roles in catalytic reactions. Chemically, the additives considered in this study (mimicking the behavior of complex substrates) can, among others, stabilize catalytic intermediates or transition states, influence the solubility or aggregation of catalyst complexes, or alter the coordination environment around the palladium center50. However, the modest size of the additive effects observed here (ranging around ± 9 units) aligns with typical scenarios where additives fine-tune rather than fundamentally alter reaction mechanisms51,52,53,54.
Bases and ligands generally have smaller but still meaningful influences, as they primarily facilitate crucial mechanistic steps like deprotonation and oxidative addition/reductive elimination35,55. Thus, these components might be expected to show moderately sized effects, which is precisely reflected in the statistical results obtained. Importantly, the observed strong interactions between halides and bases are highly chemically meaningful. Bases are directly involved in deprotonation steps that become more or less relevant depending on the reactivity of the palladium-halide intermediate47. Hence, the pronounced halide-base interactions are mechanistically justified: the nature of the halide significantly impacts the effectiveness of a base in promoting essential catalytic steps.
From Table 1, one can see another interesting fact: the factor-level interactions of the same halide 1 with two different additives can be dramatically different. Indeed, it is highly negative ( − 32.40) in the case of additive 9, while it is, in contrast, highly positive ( + 27.91) with additive 22. Can this be chemically interpreted? This strongly suggests that these two additives chemically interact in fundamentally different ways with the halide-containing intermediate or a transition state. In Buchwald-Hartwig reactions, the oxidative addition step, where the Pd catalyst inserts into the carbon-halogen bond, is highly sensitive to both electronic and steric effects. Additives can significantly alter this step or subsequent catalytic intermediates. For instance, the highly positive interaction ( + 27.91) indicates that this additive 22 likely stabilizes a crucial catalytic intermediate (such as the Pd(II)-aryl intermediate) formed specifically from halide 1. It may do so through beneficial electronic interactions (electron donation or stabilization through coordination), which enhance reactivity, reduce activation barriers, and thus significantly improve yield. On the other hand, the highly negative interaction ( − 32.40, additive 9) suggests a strong destabilization of the catalytic intermediate or transition state, specifically associated with the same halide 1. This additive might compete with catalytic intermediates for coordination to palladium or otherwise sterically or electronically interfere with essential steps in the catalytic cycle, drastically lowering reaction efficiency. Therefore, the stark contrast between these two interaction values emphasizes how subtle chemical differences between additives can drastically change their roles from strongly beneficial (enhancing catalytic efficiency) to detrimental (interfering with catalytic processes), particularly when combined with specific halides. Such chemical insights highlight the practical importance of carefully choosing additive-halide combinations, as minor structural differences in additives can lead to dramatically different outcomes in reaction efficiency.
Finally, the absolute largest additive–halide interaction coefficients align with a simple mechanistic picture. Additive 22 is strongly positive with halide 1 ( + 27.91) – consistent with a coordinating donor facilitating oxidative addition for an electronically demanding aryl chloride—but negative with halide 2 ( − 9.71) and with reactive halide 10 and halide 13 ( − 6.24, − 2.20), in line with over-stabilization or coordination-site competition when additional assistance is unnecessary. In contrast, additive 9 suppresses halide 1 ( − 32.40) yet is modestly positive with reactive halides (10 and 13: + 2.91, + 2.70). Thus, electron-rich, coordinating additives aid difficult, electron-rich chlorides but can impede already-reactive substrates; weaker or non-coordinating additives show the opposite tendency, providing a general chemical rationale for the observed interaction signs and magnitudes.
Our proposed method is scalable and applicable to a broad range of datasets generated in HTE with target variables from the exponential family of distributions and categorical explanatory features. In the Supplementary Section 1, we provide a link to the analysis of two additional datasets with three and five categorical features, as well as a Python package that enables the analysis of similar datasets.
Methods
In this section, we detail the parameter estimation problem in model (1), which turns out to be a very specific type of generalized linear model. The right-hand side of the model corresponds to the structures used in the analysis of variance (ANOVA) with four factors, where only a single observation is available for each combination of factor levels. However, unlike classical ANOVA models, the response variable in (1) does not follow a normal distribution; instead, it follows a continuous Bernoulli distribution. Model (1) can be rewritten in a matrix form
where Yi is the i-th entry of vector \({{\boldsymbol{Y}}}={({Y}_{1},\ldots ,{Y}_{n})}^{T}\), n = 3 960, which contains all yield values, Zi is the i-th row of a dummy coded matrix \({{\boldsymbol{Z}}}\in {{\mathbb{R}}}^{n\times n}\), corresponding to a four-factor ANOVA experimental design with single replicates (see Supplementary Section 3 for the details on construction of Z), and \({{\boldsymbol{\theta }}}={({\mu }_{0},{\alpha }_{1},\ldots ,{(\alpha \beta \gamma \delta )}_{21,14,2,3})}^{t}\in {{\mathbb{R}}}^{n}\) is the vector of unknown parameters. If Z were a regular matrix satisfying all the assumptions of classical (generalized) linear models, one could proceed with the estimation of θ using iteratively re-weighted least squares and identify the factor levels and their interactions that are particularly influential for the yield in Buchwald-Hartwig amination.
Unfortunately, the matrix Z is not regular. Due to the availability of only a single observation per factor level combination, there are as many parameters as observations. Since it is reasonable to assume that a large portion of the parameters are zero, one might be tempted to apply a classical Lasso algorithm with a response from the exponential family56. However, it can be easily observed that matrix Z does not satisfy the assumptions of the classical Lasso algorithm. In particular, Z has only 16 distinct singular values (see Supplementary Section 4 for proof) and is ill-conditioned, with a condition number of 62.9. As a result, the Lasso estimator would fail to correctly identify the parameter θ, instead setting entries of θ to zero randomly, as discussed in refs. 57 and 58.
The ill-posedness of Z arises from the fact that model (1) is a four-way interactions model: it includes both the main effects of factor levels as well as their interactions, leading to high dependency among the columns of Z. To impose model identifiability and enhance interpretability, it is reasonable to assume that an interaction effect should only be included in the model if the corresponding main effects are non-zero. These constraints are not novel and are referred to as marginality or hierarchically well-formulated in (generalized) linear models59,60 or heredity constraints in designed experiments61. For example, in a model with two-way interactions, the condition that either αi = 0 or βj = 0 implies (αβ)ij = 0 is known as the strong heredity condition. Some statisticians argue that interaction models violating strong heredity are nonsensical. For a more detailed discussion on the statistical reasoning behind heredity conditions, see ref. 62.
Statistically, imposing the heredity conditions makes model (1) more parsimonious and easier to interpret. From a chemical perspective, strong heredity has a clear mechanistic interpretation, particularly in the context of catalytic reactions like Buchwald-Hartwig amination. In chemical terms, interactions between factor levels, such as a specific base and an additive, typically represent cooperative molecular effects. For instance, one component may stabilize or activate an intermediate produced by another, significantly altering the reaction energetics and thus influencing the yield. However, each component must exhibit some intrinsic chemical effect individually for such meaningful chemical interactions to occur. If an additive or base alone has no measurable impact on the reaction outcome, it suggests that the component neither interacts with reaction intermediates nor significantly influences the catalytic pathway. Without individual chemical relevance, it becomes mechanistically implausible that combining two chemically inert components would suddenly produce a notable combined effect. Energetically, reaction outcomes are determined by the stabilization or destabilization of intermediates and transition states. If neither component modifies the reaction’s energy profile individually, there is no plausible molecular mechanism by which their combined presence could dramatically alter the reaction energetics or pathway. Therefore, imposing a strong heredity condition in statistical modeling, where interaction effects are allowed only if the main effects are present, aligns naturally with chemical realism. This ensures that statistically identified interactions correspond to chemically meaningful scenarios, thereby enhancing the interpretability and reliability of the model.
It turns out that all existing algorithms for parameter estimation in interaction models under (strong) heredity conditions have been developed and implemented specifically for two-way interactions with continuous descriptors and a normal response variable. The main contribution of this work is the extension of the algorithm from63 to handle four-way interactions in ANOVA models with a single observation per factor level combination, where the response variable follows a continuous Bernoulli distribution.
In the following, we give the main ideas of our estimation algorithm; more details are provided in the Supplementary Section 6. As previously discussed, model (1) must be estimated under strong heredity conditions to ensure parameter identifiability. Given that our model includes interactions up to four-factor levels, the following strong heredity conditions should be imposed:
-
(i)
αi = 0 or βj = 0 implies (αβ)ij = 0;
-
(ii)
(αβ)ij = 0 or (αγ)ik = 0 or (βγ)jk = 0 implies (αβγ)ijk = 0;
-
(iii)
(αβγ)ijk = 0 or (αβδ)ijl = 0 or (αγδ)ikl = 0 or (βγδ)jkl = 0 implies (αβγδ)ijkl = 0.
To fix the ideas, let us adopt the matrix form in equation (2), which provides a more convenient representation compared to the model in equation (1). Additionally, we explicitly highlight the hierarchical structure of the matrix Z via representation
where Zi,j denotes the element from the i-th row and j-th column of Z. Each column Zj, j = 2, …, 41 corresponds to a factor level of main effects, that is, Z2, …, Z22 are columns corresponding to levels of factor additives, Z23, …, Z36 capture levels of factor halides, Z37, Z38 belong to factor bases and Z39, Z40, Z41 are ligands. Further columns of Z are built as products of all levels of these four factors, such that only interactions between main effects from different factors are included in the model (no interactions between different factor levels of the same factor are taken). For example, Z42 = Z2Z23 and so on. To exclude products of columns from the same factor, notation f(j) ∈ {1, 2, 3, 4} is introduced, where f(j) = 1 for j = 2, …, 22 (factor additives), f(j) = 2, j = 23, …, 36 (factor halides), f(j) = 3, j = 37, 38 (factor bases) and f(j) = 4, j = 39, 40, 41 (factor ligands). Altogether, ξ describes the contribution of the main effects, while two-way, three-way, and four-way factor level interactions are captured by Θ, Ψ, and Φ, respectively.
Imposing strong heredity conditions translates now to the following constraints
where ρjk is a proportionality constant, corresponding to the interaction of main effects j and k, ζjkl is a proportionality constant, corresponding to the interaction of main effects j, k, l, and τjklm is a proportionality constant, corresponding to the interaction of main effects j, k, l and m.
Consequently, in order to find a solution that satisfies strong heredity conditions, the following penalized negative log-likelihood should be minimized:
where λξ, λρ, λζ and λτ are four tuning parameters that control the amount of regularization and ℓ(ηi, yi) is the log-likelihood of the i-th sample given in the Supplementary Section 6.2.
Algorithm 1
Estimation under strong heredity conditions
Input:Z, Y, \({\widehat{{{\boldsymbol{\mu }}}}}_{0}^{(0)}\), \({\widehat{{{\boldsymbol{\xi }}}}}^{(0)}\), \({\widehat{{{\boldsymbol{\rho }}}}}^{(0)}\), \({\widehat{{{\boldsymbol{\zeta }}}}}^{(0)}\), \({\widehat{{{\boldsymbol{\tau }}}}}^{(0)}\), T, M
Output:\({\widehat{\mu }}_{0}\), \(\widehat{{{\boldsymbol{\xi }}}}\), \(\widehat{{{\boldsymbol{\rho }}}}\), \(\widehat{{{\boldsymbol{\zeta }}}}\), \(\widehat{{{\boldsymbol{\tau }}}}\)
it ← 0;
\({\widehat{\mu }}_{0}\leftarrow {\widehat{\mu }}_{0}^{(0)}\), \(\widehat{{{\boldsymbol{\xi }}}}\leftarrow {\widehat{{{\boldsymbol{\xi }}}}}^{(0)}\), \(\widehat{{{\boldsymbol{\rho }}}}\leftarrow {\widehat{{{\boldsymbol{\rho }}}}}^{(0)}\), \(\widehat{{{\boldsymbol{\zeta }}}}\leftarrow {\widehat{{{\boldsymbol{\zeta }}}}}^{(0)}\), \(\widehat{{{\boldsymbol{\tau }}}}\leftarrow {\widehat{{{\boldsymbol{\tau }}}}}^{(0)}\);
while it < M do
\({Q}_{old}\leftarrow Q({\widehat{\mu }}_{0},\widehat{{{\boldsymbol{\xi }}}},\widehat{{{\boldsymbol{\rho }}}},\widehat{{{\boldsymbol{\zeta }}}},\widehat{{{\boldsymbol{\tau }}}})\);
it ← it + 1;
Update \({\widehat{{{\boldsymbol{\mu }}}}}_{0}\): \({\widehat{{{\boldsymbol{\mu }}}}}_{0}\leftarrow \arg {\min }_{{\mu }_{0}}Q({\mu }_{0},\widehat{{{\boldsymbol{\xi }}}},\widehat{{{\boldsymbol{\rho }}}},\widehat{{{\boldsymbol{\zeta }}}},\widehat{{{\boldsymbol{\tau }}}})\);
Update each component of \(\widehat{{{\boldsymbol{\xi }}}}\): \({\widehat{\xi }}_{j}\leftarrow \arg \mathop{\min }_{{\xi }_{j}}Q\left(\right.{\widehat{\mu }}_{0},{\xi }_{j},{\widehat{{{\boldsymbol{\xi }}}}}_{-j},\widehat{{{\boldsymbol{\rho }}}},\widehat{{{\boldsymbol{\zeta }}}},\widehat{{{\boldsymbol{\tau }}}}\left)\right.\);
Update each component of \(\widehat{{{\boldsymbol{\rho }}}}\): \({\widehat{{{\boldsymbol{\rho }}}}}_{jk}\leftarrow \arg \mathop{\min }_{{\rho }_{jk}}Q \left(\right.{\widehat{\mu }}_{0},\widehat{{{\boldsymbol{\xi }}}},{\rho }_{jk},{\widehat{{{\boldsymbol{\rho }}}}}_{-jk}, \widehat{{{\boldsymbol{\zeta }}}},\widehat{{{\boldsymbol{\tau }}}}\left)\right.\);
Update each component of \(\widehat{{{\boldsymbol{\zeta }}}}\): \({\widehat{\zeta }}_{jkl}\leftarrow \arg \mathop{\min }_{{\zeta }_{jkl}}Q\left(\right.{\widehat{\mu }}_{0},\widehat{{{\boldsymbol{\xi }}}},\widehat{{{\boldsymbol{\rho }}}},{\zeta }_{jkl},{\widehat{{{\boldsymbol{\zeta }}}}}_{-jkl},\widehat{{{\boldsymbol{\tau }}}}\left(\right.\);
Update \(\widehat{{{\boldsymbol{\tau }}}}\): \(\widehat{\tau }\leftarrow \arg \mathop{\min }_{{{\boldsymbol{\tau }}}}Q({\widehat{\mu }}_{0},\widehat{{{\boldsymbol{\xi }}}},\widehat{{{\boldsymbol{\rho }}}},\widehat{{{\boldsymbol{\zeta }}}},{{\boldsymbol{\tau }}})\);
\({Q}_{new}\leftarrow Q({\widehat{\mu }}_{0},\widehat{{{\boldsymbol{\xi }}}},\widehat{{{\boldsymbol{\rho }}}},\widehat{{{\boldsymbol{\zeta }}}},\widehat{{{\boldsymbol{\tau }}}})\);
if \(| {Q}_{old}-{Q}_{new}| \le T\cdot | {Q}_{old}|\) then
return \({\widehat{\mu }}_{0}\), \(\widehat{{{\boldsymbol{\xi }}}}\), \(\widehat{{{\boldsymbol{\rho }}}}\), \(\widehat{{{\boldsymbol{\zeta }}}}\), \(\widehat{{{\boldsymbol{\tau }}}}\);
else
Continue;
end if
end while
The minimization problem in equation (3) does not have a closed-form solution but can be solved using a coordinate descent approach. The algorithm takes as input the data matrix Z, the response vector Y, a convergence tolerance level T, a maximum number of iterations M, and sensible initializations for the model parameters: μ0, ξ, ρ, ζ, and τ.
Initializing with estimated main effects and setting all other parameters to zero has been found to be both robust and simple; further details can be found in the Supplementary Section 6. The main steps of the algorithm are outlined in Algorithm 1. In the algorithm description, v−j denotes a vector v without the j-th main effect, v−jk is a vector v without the component corresponding to the interaction between j-th main effect and k-th main effect, and v−jkl lacks the component of the three-way interaction between j-th, k-th and l-th main effects.
Data availability
The data used in ref. 23, as well as data for two more examples discussed in the Supplementary Section 1, are publicly available and can also be downloaded at GitHub: https://doi.org/10.5281/zenodo.17748952.
Code availability
The Python code that implements our algorithm and provides the data analysis is available on GitHub: https://doi.org/10.5281/zenodo.17748952
References
Butler, K. T., Davies, D. W., Cartwright, H., Isayev, O. & Walsh, A. Machine learning for molecular and materials science. Nature 559, 547–555 (2018).
Williams, W. L. et al. The evolution of data-driven modeling in organic chemistry. ACS Cent. Sci. 7, 1622–1637 (2021).
Mater, A. C. & Coote, M. L. Deep learning in chemistry. J. Chem. Inf. Model. 59, 2545–2559 (2019).
Takahashi, K. & Miyazato, I. Rapid estimation of activation energy in heterogeneous catalytic reactions via machine learning. J. Comput. Chem. 39, 2405–2408 (2018).
Yang, W., Fidelis, T. T. & Sun, W.-H. Prediction of catalytic activities of bis (imino) pyridine metal complexes by machine learning. J. Comput. Chem. 41, 1064–1067 (2020).
Borghini, A., Crotti, P., Pietra, D., Favero, L. & Bianucci, A. M. Chemical reactivity predictions: Use of data mining techniques for analyzing regioselective azidolysis of epoxides. J. Comput. Chem. 31, 2612–2619 (2010).
Ishioka, S. et al. Unveiling gas-phase oxidative coupling of methane via data analysis. J. Comput. Chem. 42, 1447–1451 (2021).
Mayr, A., Klambauer, G., Unterthiner, T. & Hochreiter, S. Toxicity prediction using deep learning front. Environ. Sci. 3, 1–15 (2016).
Ryu, S., Kwon, Y. & Kim, W. Y. A bayesian graph convolutional network for reliable prediction of molecular properties with uncertainty quantification. Chem. Sci. 10, 8438–8446 (2019).
Druchok, M., Yarish, D., Gurbych, O. & Maksymenko, M. Toward efficient generation, correction, and properties control of unique drug-like structures. J. Comput. Chem. 42, 746–760 (2021).
Hoffmann, G. et al. Predicting experimental electrophilicities from quantum and topological descriptors: a machine learning approach. J. Comput. Chem. 41, 2124–2136 (2020).
Subramanian, G., Ramsundar, B., Pande, V. & Denny, R. A. Computational modeling of β-secretase 1 (bace-1) inhibitors using ligand based approaches. J. Chem. Inf. Model. 56, 1936–1949 (2016).
Burbidge, R., Trotter, M., Buxton, B. & Holden, S. Drug design by machine learning: support vector machines for pharmaceutical data analysis. Comput. Chem. 26, 5–14 (2001).
Ekins, S. The next era: deep learning in pharmaceutical research. Pharm. Res. 33, 2594–2603 (2016).
Sun, W. et al. Machine learning–assisted molecular design and efficiency prediction for high-performance organic photovoltaic materials. Sci. Adv. 5, eaay4275 (2019).
Xue, D. et al. Accelerated search for materials with targeted properties by adaptive design. Nat. Commun. 7, 1–9 (2016).
Segler, M. H., Preuss, M. & Waller, M. P. Planning chemical syntheses with deep neural networks and symbolic ai. Nature 555, 604–610 (2018).
Bédard, A.-C. et al. Reconfigurable system for automated optimization of diverse chemical reactions. Science 361, 1220–1225 (2018).
Gao, H. et al. Using machine learning to predict suitable conditions for organic reactions. ACS Cent. Sci. 4, 1465–1476 (2018).
Genheden, S. et al. Prediction of the chemical context for buchwald-hartwig coupling reactions. Mol. Inform. 41, 2100294 (2022).
Li, J. & Eastgate, M. D. Making better decisions during synthetic route design: leveraging prediction to achieve greenness-by-design. React. Chem. Eng. 4, 1595–1607 (2019).
Granda, J. M., Donina, L., Dragone, V., Long, D.-L. & Cronin, L. Controlling an organic synthesis robot with machine learning to search for new reactivity. Nature 559, 377–381 (2018).
Ahneman, D. T., Estrada, J. G., Lin, S., Dreher, S. D. & Doyle, A. G. Predicting reaction performance in C–N cross-coupling using machine learning. Science 360, 186–190 (2018).
Nielsen, M. K., Ahneman, D. T., Riera, O. & Doyle, A. G. Deoxyfluorination with sulfonyl fluorides: Navigating reaction space with machine learning. J. Am. Chem. Soc. 140, 5004–5008 (2018).
Schwaller, P., Vaucher, A., Laino, T. & Reymond, J.-L. Prediction of chemical reaction yields using deep learning. Mach. Learn. Sci. Technol. 2, 015016 (2021).
Georgakopoulos-Soares, I., Chan, C. S. Y., Ahituv, N. & Hemberg, M. High-throughput techniques enable advances in the roles of DNA and RNA secondary structures in transcriptional and post-transcriptional gene regulation. Genome Biol. 23, 159 (2022).
Dunham, A. S., Beltrao, P. & AlQuraishi, M. High-throughput deep learning variant effect prediction with sequence unet. Genome Biol. 24, 110 (2023).
Soon, W. W., Hariharan, M. & Snyder, M. P. High-throughput sequencing for biology and medicine. Mol. Syst. Biol. 9, 640 (2013).
Toyama, R. et al. High-throughput materials exploration system for the anomalous hall effect using combinatorial experiments and machine learning. npj Comput. Mater. 11, 269 (2025).
Greeley, J., Jaramillo, T. F., Bonde, J., Chorkendorff, I. & Nørskov, J. K. Computational high-throughput screening of electrocatalytic materials for hydrogen evolution. Nat. Mater. 5, 909–913 (2006).
Holewa, P. et al. High-throughput quantum photonic devices emitting indistinguishable photons in the telecom C-band. Nat. Commun. 15, 3358 (2024).
He, B. et al. High-throughput screening platform for solid electrolytes combining hierarchical ion-transport prediction algorithms. Sci. Data 7, 151 (2020).
Yamankurt, G. et al. Exploration of the nanomedicine-design space with high-throughput screening and machine learning. Nat. Biomed. Eng. 3, 318–327 (2019).
Surry, D. S. & Buchwald, S. L. Biaryl phosphane ligands in palladium-catalyzed amination. Angew. Chem. Int. Ed. 47, 6338–6361 (2008).
Hartwig, J. F. Evolution of a fourth generation catalyst for the amination and thioetherification of aryl halides. Acc. Chem. Res. 41, 1534–1544 (2008).
Ruiz-Castillo, P. & Buchwald, S. L. Applications of palladium-catalyzed c–n cross-coupling reactions. Chem. Rev. 116, 12564–12649 (2016).
Heravi, M. M., Kheilkordi, Z., Zadsirjan, V., Heydari, M. & Malmir, M. Buchwald-hartwig reaction: An overview. J. Organomet. Chem. 861, 17–104 (2018).
Dong, J., Peng, L., Yang, X., Zhang, Z. & Zhang, P. Xgboost-based intelligence yield prediction and reaction factors analysis of amination reaction. J. Comput. Chem. 43, 289–302 (2022).
Mu, X., Dong, J., Peng, L. & Yang, X. Deep forest-based intelligent yield predicting of Buchwald-Hartwig coupling reaction. MATCH Commun. Math. Comput. Chem. 88, 5–27 (2022).
Zhao, Y. et al. An optimized deep convolutional neural network for yield prediction of Buchwald-Hartwig amination. Chem. Phys. 550, 111296 (2021).
Wang, L. et al. Chemcnet: An Explainable Integrated Model for Intelligent Analyzing Chemistry Synthesis Reactions. MATCH Commun. Math. Comput. Chem. 91, 41–78 (2024).
Zuranski, A. M., Martinez Alvarado, J. I., Shields, B. J. & Doyle, A. G. Predicting reaction yields via supervised learning. Acc. Chem. Res. 54, 1856–1865 (2021).
Zuranski, A. M., Gandhi, S. S. & Doyle, A. G. A machine learning approach to model interaction effects: Development and application to alcohol deoxyfluorination. J. Am. Chem. Soc. 145, 7898–7909 (2023).
Farina, V. High-turnover palladium catalysts in cross-coupling and heck chemistry: A critical overview. Adv. Synth. Catal. 346, 1553–1582 (2004).
Hartwig, J. F. Electronic effects on reductive elimination to form carbon-carbon and carbon-heteroatom bonds from palladium(II) complexes. Inorg. Chem. 46, 1936–1947 (2007).
Negishi, E. Magical power of transition metals: Past, present, and future (Nobel lecture). Angew. Chem. Int. Ed. 50, 6738–6764 (2011).
Alcázar-Román, L. M. & Hartwig, J. F. Mechanism of aryl chloride amination: Base-induced oxidative addition. J. Am. Chem. Soc. 123, 12905–12906 (2001).
Tian, J., Wang, G., Qi, Z.-H. & Ma, J. Ligand effects of BrettPhos and RuPhos on rate-limiting steps in Buchwald-Hartwig amination reaction due to the modulation of steric hindrance and electronic structure. ACS Omega 5, 21385–21391 (2020).
Fors, B. P., Davis, N. R. & Buchwald, S. L. An efficient process for Pd-catalyzed C–N cross-coupling reactions of aryl iodides: Insight into controlling factors. J. Am. Chem. Soc. 131, 5766–5768 (2009).
Li, D. et al. Buchwald-Hartwig amination of coordinating heterocycles enabled by large-but-flexible Pd-BIAN-NHC catalysts. Chem. – A Eur. J. 28, e202103341 (2022).
Shen, Q., Shekhar, S., Stambuli, J. P. & Hartwig, J. F. Highly reactive, general, and long-lived catalysts for coupling heteroaryl and aryl chlorides with primary nitrogen nucleophiles. Angew. Chem. Int. Ed. 44, 1371–1375 (2005).
Maiti, D., Fors, B. P., Henderson, J. L., Nakamura, Y. & Buchwald, S. L. Palladium-catalyzed coupling of functionalized primary and secondary amines with aryl and heteroaryl halides: Two ligands suffice in most cases. Chem. Sci. 2, 57–68 (2011).
Ruiz-Castillo, P. & Buchwald, S. L. Applications of palladium-catalyzed C-N cross-coupling reactions. Chem. Rev. 116, 12564–12649 (2016).
Surry, D. S. & Buchwald, S. L. Dialkylbiaryl phosphines in Pd-catalyzed amination: A user’s guide. Chem. Sci. 2, 27–50 (2011).
Dorel, R., Grugel, C. P. & Haydl, A. M. The Buchwald–Hartwig amination after 25 years. Angew. Chem. Int. Ed. 58, 17118–17129 (2019).
van de Geer, S. A. High-dimensional generalized linear models and the lasso. The Ann. Stat. 36, 614–645 (2008).
Zhao, P. & Yu, B. On Model Selection Consistency of Lasso. J. Mach. Learn. Res. 7, 2541–2563 (2006).
Wang, H., Lengerich, B. J., Aragam, B. & Xing, E. P. Precision lasso: AccOunting For Correlations And Linear Dependencies In High-dimensional Genomic Data. Bioinformatics 35, 1181–1187 (2018).
McCullagh, P. & Nelder, J. A.Generalized Linear Models (Chapman & Hall / CRC, London, 1989).
Peixoto, J. L. Hierarchical variable selection in polynomial regression models. Am. Stat. 41, 311–313 (1987).
Hamada, M. & Wu, C. F. J. Analysis of designed experiments with complex aliasing. J. Qual. Technol. 24, 130–137 (1992).
Bien, J., Taylor, J. & Tibshirani, R. A lasso for hierarchical interactions. The Ann. Stat. 41, 1111–1141 (2013).
Choi, N. H., Li, W. & Zhu, J. Variable selection with the strong heredity constraint and its oracle property. J. Am. Stat. Assoc. 105, 354–364 (2010).
Acknowledgements
T.K. is grateful to Prof. Efstathia Bura for helpful discussions. B.M. thanks Prof. Leticia González and Prof. Nuno Maulide for their generous support, fruitful discussions, and resources. The support of the University of Vienna’s Data Science research network is also acknowledged. Open access funding provided by the University of Vienna.
Author information
Authors and Affiliations
Contributions
T.K. formulated the model for the data and suggested the algorithm for estimation. R.A.M. programmed the algorithm in R and Python and performed data analysis for all datasets. G.F. contributed to the theoretical analysis of the model and the implementation of the algorithm. B.M. provided a chemical interpretation of the results.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Chemistry thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Krivobokova, T., Morariu, RA., Finocchio, G. et al. Modelling and estimation of chemical reaction yields from high-throughput experiments. Commun Chem 9, 61 (2026). https://doi.org/10.1038/s42004-025-01866-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42004-025-01866-8