Abstract
Microbial resilience is essential for ecosystem stability as environmental conditions fluctuate. Despite resilience emerging from both individual (organism)-level adaptations and community-level processes, integration of these mechanisms remains enigmatic, particularly in arid environments. These extreme ecosystems, spanning over 45% of Earth’s terrestrial surface, provide a natural laboratory for understanding microbial survival under harsh conditions. Here, we use time-resolved multiomics to show that resilience results from dynamic microbial network reorganization enabling the coordination between stochastic processes that maintain community stability, and individual stress responses. Additionally, Thermoproteota emerged as a keystone taxon maintaining nitrogen cycling and fostering cross-feeding networks. Its ecological prominence highlights its central role in arid ecosystems, making it an ideal model organism for understanding microbial adaptation to environmental extremes. Our findings bridge the gap between individual adaptations and community-wide resilience, offering a framework for understanding microbial responses to environmental fluctuations and their implications for ecosystem function.
Similar content being viewed by others
Introduction
Microbial communities are pivotal to ecosystem functions1, yet they face persistent environmental fluctuations across diverse habitats2. These changes can dramatically impact community functioning and stability3, where stability is defined as the tendency of a community to return to its original state after a disturbance4. Stability encompasses both resistance (the ability of a community property to remain unchanged or insensitive to disturbance) and resilience (the capacity to recover after disturbance and return to pre-disturbance conditions)4,5. Recovery typically follows one of four extreme scenarios: full recovery, physiological adaptation, functional redundancy, or complete loss of function with no recovery6. Despite advances, the molecular mechanisms underlying microbial community recovery remain poorly understood, particularly how individual organism strategies integrate with community-level processes to ensure resilience (5,7,8).
Microbial resilience emerges from both individual organisms’ survival strategies and collective community-level coordination5,7,9. At the individual level, adaptations provide the mechanistic basis for survival during extreme fluctuations10,11,12,13. For instance, dormancy allows persistence during unfavorable conditions and rapid reactivation when conditions improve14,15, ensuring long-term population survival16. Upon rewetting, dormant microbes prioritize DNA repair and energy generation15, while others maintain consistent transcription of stress response and nutrient acquisition genes during dry periods17. Additional mechanisms, such as osmolyte accumulation and oxidative stress further protect cellular integrity and enhance persistence18. At the community level, functional redundancy has been shown to buffer ecosystem processes against taxonomic shifts or environmental fluctuations6. While these individual and community-level mechanisms are both well-documented, their integration remains poorly understood.
Recent studies reveal that individual and community responses in arid systems often operate as disconnected processes, functioning at different scales and timescales. For instance, while drought caused >90% of bacterial and archaeal taxa to stop dividing, growing taxa still accounted for 4% of the total community, revealing critical disconnects between individual survival mechanisms and community-level functioning19. Similarly, despite shifts toward more drought-tolerant bacterial groups at the individual organism level, community-level enzyme activities in rhizosphere hotspots remain stable through collaborative relationships rather than compositional changes20. Furthermore, communities in arid and semi-arid ecosystems show little accumulation of osmolytes under stress, with individual energy-saving strategies like dormancy failing to translate to predictable community-level carbon cycling responses21.
These findings highlight fundamental gaps in understanding how individual organisms adaptations integrate with community-level processes, particularly during abrupt environmental transitions22. Advancing understanding requires integrated approaches bridging individual traits and community dynamics. The synthesis of dryland microbiome studies reveals that resilience to desertification emerges not from one scale of organization alone, but from coordinated responses across multiple levels13,23, underscoring the need for multi-omics strategies to unravel how microbial communities maintain function amid intensifying environmental perturbations. Understanding this multi-level response is particularly crucial as climate change intensifies environmental perturbations.
Arid environments offer a unique laboratory for studying microbial resilience. Characterized by extreme temperatures, UV radiation, and variable precipitation22,24, these ecosystems represent critical systems for understanding adaptation25,26. As these regions expand due to climate change27,28, developing integrative frameworks will be essential for predicting ecosystem responses. However, despite the urgent need for such an integrative understanding, few studies have simultaneously characterized both individual microbial adaptations and community-level processes in arid environments during environmental fluctuations.
To address this knowledge gap, we employed an integrated time-resolved multiomics approach analyzing bare soil samples from Arizona’s Saguaro National Park, an iconic desert ecosystem characterized by extreme wet-dry cycles. By tracking individual-level strategies (traits identified at the level of MAGs) alongside community composition and metabolic outputs, we reveal that microbial resilience emerges from a sophisticated balance between community stability and functional adaptability. While stochastic processes maintain stable community composition, organic matter profiles show deterministic shifts reflecting metabolic adaptations. The key mechanism is microbial network reorganization, where interaction patterns shift without major changes in community composition. This allows microbial communities to maintain their taxonomic structure while supporting coordinated individual or population-level responses, directly addressing the integration challenges identified in previous studies. This dynamic is exemplified by ammonia-oxidizing Thermoproteota, whose flexible gene expression enables different network roles while preserving critical ecosystem functions. Our findings advance understanding of microbial resilience in expanding drylands and provide a framework for predicting ecosystem responses to climate change. Our integrative approach comprehensively characterizes both microbial communities and their metabolic outputs, revealing how coordinated responses across multiple organizational levels maintain biogeochemical cycling under extreme conditions, providing insights critical for ecosystem management as arid regions continue to expand globally.
Results
Monsoon events create distinct physicochemical states in arid soils
The North American Monsoon drives shifts in resource availability across desert ecosystems. We captured these transitions with temporal sampling at the Saguaro National Park (Fig. 1A). Soil samples were collected from a depth of 10–15 cm at four sampling sites (biological replicates) across eight time points from May to September 2021, plus one in May 2022 (n = 36, 4 per time point), capturing the monsoon cycle from pre-monsoon drought through post-monsoon recovery (Fig. 1B and Supplementary Fig. 1a). While transitions followed historical monsoon patterns29, 2021 saw some of the highest precipitation in the last decade (Supplementary Fig. 1b), offering a natural experiment to examine microbial responses to extreme environmental transitions.
A Geographic location of soil sampling sites within Saguaro National Park. B Pictures taken at the sampling site during the sampling campaign in May, July, and August of 2021. C Temporal dynamics of environmental and physicochemical factors throughout the 2021 monsoon season. Background indicates whether the sample was taken before, during or after the 2021 monsoon season. D Principal component analysis of the collected samples (n = 36) based on the measured environmental factors. Ellipses indicate whether the sample was taken before, during or after the 2021 monsoon season.
The shift from drought to intense rainfall created distinct temporal patterns in soil, marked by changes in moisture availability, temperature, and potential resource inputs (Fig. 1C, Supplementary Data 1 and Supplementary Fig. 1c). Principal component analysis (PCA) revealed three distinct sample clusters corresponding to pre-monsoon (yellow ellipse), monsoon (blue ellipse), and post-monsoon (red ellipse) periods (Fig. 1D). Post-monsoon samples gradually returned to pre-monsoon conditions, highlighting the cyclical nature of these transitions (Fig. 1C). Moisture content and vegetation density (measured by the Normalized Difference Vegetation Index—NDVI) emerged as primary drivers, explaining 30.47% of observed variance along the first component, while air temperature accounted for 19.66% along the second component. Hierarchical clustering analysis of physicochemical parameters further supported these distinct environmental states between monsoon and non-monsoon periods (Supplementary Fig. 1c).
Community-level responses emerge from local adaptation
To understand microbial responses to monsoon transitions, we employed a multiomics approach that combined microbial taxonomic profiling (using amplicon sequence variants (ASVs) derived from 16S rRNA amplicon sequencing and operational taxonomic units (OTUs) derived from shotgun metagenomics) with organic matter characterization using Fourier-transform ion cyclotron resonance mass spectrometry (FTICR-MS).
Our analysis indicated that microbial community composition remained relatively consistent throughout the monsoon season, despite environmental fluctuations (Fig. 2A, D; Supplementary Fig. 2a, b, h; Supplementary Note 1). Beta diversity analysis based on Bray-Curtis dissimilarity showed that time of collection did not explain significant variation in community structure (PERMANOVA month: R = 0.21, p = 0.947; Fig. 2D and Supplementary Fig. 2b). This compositional consistency is further supported by visually stable phylum-level profiles over time (Fig. 2A) and a high community stability index (Supplementary Fig. 2h). The observed compositional stability reflects resistance to environmental fluctuations, aligning with prior studies suggesting that arid soil microbial communities are adapted to endure dry-wet cycles30 through ecological memory31 and long-term optimization to historical moisture regimes32. However, alpha diversity metrics presented a more dynamic response (Fig. 2F). Unlike previous reports of stable or increased microbial diversity post-precipitation26,33, we observed a temporary decline in microbial diversity followed by recovery. This decline likely results from mechanisms such as differential growth triggered by increased water availability34, formation of anoxic microsites35, and osmotic stress-induced mortality36. Comparison of metagenomics read signatures with other global arid ecosystems revealed that while each ecosystem harbors locally adapted communities, Sonoran Desert more closely resembled those from North American deserts (e.g., Mojave) than geographically distant systems (e.g., Negev) (Fig. 2C), suggesting local adaptation shapes community composition while shared regional species pools support resilience37,38.
A Average relative abundance of the 12 most abundant microbial classes, from OTUs inferred with SingleM, at each sampling time (n = 4). B Average relative abundance of observed elemental compositions based on FTICR-MS data collected in positive mode at each sampling time (n = 4). C Non-metric multidimensional scaling (NMDS) ordination of different metagenomics datasets collected from arid soils based on MASH distances. D Principal coordinate analysis (PCoA) of shotgun metagenomics data (n = 28), based on Bray-Curtis dissimilarities calculated at the SingleM OTU level. E NMDS of FTICR-MS data (n = 28). F Alpha diversity indices (observed richness and Chao1 richness) inferred from SingleM OTUs and FTICR-MS metabolites abundances, respectively. Boxes represent the upper and lower quartiles, the line in each box represents the median and the whiskers represent the maximum and minimum values, no further than 1.5 times the interquartile range; values beyond the whiskers represent outliers and are plotted individually.
FTICR-MS analysis detected 20,971 different masses (m/z) across all samples, with 4175 assigned putative molecular formulas. Unlike lignin-dominated peatlands39,40, these arid soils showed predominance of lipid-like compounds, followed by protein- and lignin-like metabolites (Supplementary Fig. 2d and Supplementary Note 1), which suggests that microbial communities may have a strong influence on soil organic matter composition as reported previously41,42.
Organic matter composition showed significant temporal variation (NMDS: stress = 0.11; PERMANOVA month: R2 = 0.29, p = 0.028, Fig. 2E), particularly in N- and S-containing compounds (e.g., CHON, CHONS, CHONP) (Fig. 2B). During wet periods, osmotic stress-induced cell lysis may have released intracellular metabolites36, and increased substrate accessibility may have enhanced microbial activity43. Though some lipid-like compounds remained stable throughout the monsoon season, amino sugar abundance increased with water availability. These amino sugars, likely derived from microbial components44 may have been accumulated due to osmotic stress-induced mortality35. The subsequent increase in carbohydrates by the end of the monsoon season (September and August) likely reflected post-monsoon vegetation growth and enhanced microbial activity45. Organic matter alpha diversity initially increased at monsoon onset, likely due to precipitation-induced release of labile metabolites46. These metabolites were subsequently consumed, leading to decreased diversity and liability (low nominal oxidation state of carbon - NOSC) by monsoon’s end (Fig. 2F and Supplementary Fig. 2e).
Genomic features supporting resilience in arid soils
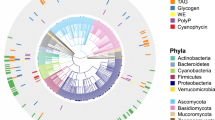
To understand the genomic basis of microbial stability in arid soils, we investigated the mechanisms underlying community-level responses. Our genome-centric approach recovered 282 MAGs (Fig. 3A), including 1 high-quality draft MAG and 281 medium-quality drafts based on the minimum information about metagenome-assembled genomes (MIMAG) standards47 (Supplementary Data 2). Despite representing ~10% of the total sequenced community (per CoverM), they captured ~60% of the sequenced microbial community at the genus level (as inferred by SingleM appraise at 86% sequence identity). This provides a robust foundation for exploring genomic adaptation in arid soils, even though these MAGs represent only a subset of the community and may exclude unassembled functional potential.
A Phylogenetic tree of the 282 recovered metagenome-assembled genomes (MAGs) showing their genome size, completeness, and contamination inferred by CheckM2 as well as their changes in abundance throughout the sampling campaign. B Dot plot showing the distribution of the recovered MAGs based on their GC content and estimated maximum growth rate. C Heatmap showing the presence of stress resistance traits among the different taxa of the recovered MAGs.
Phylogenomic analysis revealed unexpected taxonomic breadth, with MAGs spanning 30 bacterial and one archaeal classes across 18 phyla (Supplementary Fig. 4). While this diversity highlights the complex microbial ecology of arid soils, it also suggests high endemism, only one MAG was classified at species level (average nucleotide identity, ANI > 95%), while most (n = 173) were classified at genus level, indicating potentially unclassified or undercharacterized taxa adapted to arid conditions (Supplementary Data 2). Analysis of the relative abundances of the recovered MAGs in both metagenomics and metatranscriptomics datasets (Supplementary Fig. 5a, b), showed that the Actinomycetota and Thermoproteota are among the most prevalent and active taxa, alongside Tectomicrobia.
Analysis of carbon fixation pathways revealed diverse metabolic capabilities distributed across the community. The Calvin–Benson–Banshan (CBB) cycle appeared in twelve bacterial genomes from Actinomycetota, Desulfobacterota, and Pseudomonadota, which encoded complete RuBisCO genes (K01601, K01602) and associated pathway components (Supplementary Fig. 6). While some steps of the reductive citrate (rTCA) cycle were widely distributed, only one MAG from Nitrospirota encoded the key enzyme. Similarly, the 3-hydroxypropionate/4-hydroxybutyrate (3HP/4HB) and dicarboxylate/4-hydroxybutyrate (DC/4HB) cycles showed restricted distribution, with key enzymes found only in Thermoproteota, Desulfobacterota, and Tectomicrobia.
Nitrogen cycling functions displayed comparable patterns of specialized and redundant distribution. Nitrification capacity was limited to Thermoproteota, specifically the ammonia-oxidizing archaea (AOA) order Nitrososphaerales. In contrast, denitrification and dissimilatory nitrate reduction were broadly distributed across taxa. Sulfate reduction pathways were mostly limited to assimilatory and dissimilatory functions (see Supplementary Note 2).
The distribution of carbon, nitrogen, and sulfur cycling pathways highlights how metagenomic potential supports environmental adaptation in arid soils. These functions exhibit contrasting patterns, some, like the Calvin–Benson–Banshan cycle and denitrification, show redundancy across multiple phyla, while others are specialized to specific taxa. This functional architecture reflects selective pressures in arid environments, where microbial communities have evolved both redundancy for essential functions and niche specialization. The community maintains genetic infrastructure for metabolic flexibility, enabling rapid pathway activation during environmental shifts. This genomic organization may be a key strategy through which microbial communities maintain critical ecosystem processes under stress5,8,48.
Genomic traits reveal convergent and specialized stress adaptations
At the level of individual MAGs, we investigated how specific traits and stress tolerance mechanisms support microbial survival to the environmental conditions of arid ecosystems. Most recovered MAGs (n = 183) exhibited high GC content and slow predicted growth rates (n = 143, maximum growth rate <0.2) (Fig. 3B), suggesting a common evolutionary response to harsh conditions49. High GC content likely enhances thermotolerance50, while slow growth rates indicate oligotrophic lifestyles51 aligning with previous findings of GC enrichment in arid conditions30 and resource-driven selection against AT base pairs in nutrient-limited soils52. Notably, Thermoproteota demonstrated an alternative strategy having low GC content, which may reflect genome streamlining53, and efficient nitrogen use54. However, while genome streamlining is typically associated with high coding density, our Nitrososphaera MAGs exhibited coding densities ranging from 67.6 to 87.2% (Supplementary Data 2), which are lower than those reported for other Nitrososphaera MAGs (73–85%)55. The lower values in our dataset may reflect incomplete or fragmented genome assemblies, as our MAGs fall within medium-quality thresholds. Alternatively, these patterns could indicate that desert-adapted Nitrososphaera retain a higher proportion of non-coding DNA, potentially related to regulatory flexibility or increased horizontal gene transfer capacity56, which may enhance survival under fluctuating environmental stressors.
Gene expression analysis of stress tolerance traits (Supplementary Data 3), revealed that protein stability mechanisms were widespread, with most taxa encoding and expressing genes for protection, repair, and degradation of denatured and misfolded proteins (Figs. 3C and 4), reflecting adaptation to high temperatures18. These included various heat shock proteins (HSP70, HSP40, HSP60, HSP10, HSP100, HSP24) critical for protein folding and protection57.
Heatmap showing the levels of gene expression, agglomerated at phylum level, at different sampling times for various stress tolerance mechanisms defined in Supplementary Data 7. Gene expression was normalized as Gene length corrected trimmed mean of M-values (geTMM) and transformed into a z-score for each phyla and each stress tolerance mechanism to better represent the changes across the monsoon season.
Osmolyte production and transport showed both common and specialized patterns (Figs. 3C and 4). While trehalose and glutamate production were widespread, glycine betaine biosynthesis was restricted to select groups, and bacteria-exclusive ectoine was found only in Actinomycetota. These osmolytes help manage both high salinity and sudden water influx57.
Oxidative stress responses revealed similar diversity in adaptation strategies. Superoxide dismutase (K04564) showed widespread expression by most MAGs (n = 140) during dry months, particularly expression was higher in Entotheonella (geTMMMay = 478.8; geTMMJuly_1 = 263.47), Rubrobacteria (geTMMMay = 84.73; geTMMJuly_1 = 1.40) and Thermoleophilia (geTMMMay = 72.52; geTMMJuly_1 = 1.86) members (Fig. 4). Different taxa employed specialized catalase systems: heme-based catalases (KatG, KatE) in Acidobacteriota, Actinomycetota, Chloroflexota, Myxococcota and Pseudomonadota (KatG n = 41; KatE n = 8), and manganese catalase (K07217) in Thermoproteota (e.g., Nitrososphaera geTMMMay = 10.63; geTMMJuly_1 = 0.40), Gemmatimonadota and Nitrospirota (n = 41) with the highest expression during dry months (Fig. 4), suggesting specialized strategies for reactive oxygen species management58.
Multifunctional dps proteins (K04047) were present in several phyla (Fig. 3C), providing resistance against oxidative, metal, and thermal stress59, with the highest expression occurring during the dry months for Acidobacteriota, Actinomycetota, Chloroflexota, Gemmatimonadota, Tectomicrobia and Thermoproteota (e.g., Nitrososphaera geTMMMay = 81; geTMMJuly_1 = 3.32), and in the wet months for Pseudomonadota (Fig. 4). Furthermore, the genes for the glyoxylate bypass, suggested to play a role in oxidative stress60, were found encoded in many taxa and expressed during the dry months in Actinomycetota, Chroloflexota (e.g., UBA6077: geTMMMay = 3.46; geTMMJuly_1 = 1.74) and Tectomicrobia (e.g Entotheonella: geTMMMay = 13.88; geTMMJuly_1 = 8.47), and during the wet months in Acidobacteriota (e.g., Terriglobia: geTMMMay = 0.007; geTMMJuly_1 = 0.38), Desulfobacterota, Gemmatimonadota (e.g., Gemmatimonadota: geTMMMay = 0.15; geTMMJuly_1 = 2.78) and Pseudomonadota genomes (n = 48) (Fig. 4).
Other widely distributed stress tolerance mechanisms included the expression of genes relate to DNA repair as most taxa except for Thermoproteota encoded and expressed genes for the RecA (K03553) protein, which catalyzes one of the central steps of the DNA repair and homologous recombination pathway61 as well as polymerases such as polA (K02335) and polIV (K02346, K04479) that may play a role in double-strand DNA break repair62,63. Other strategies related with dormancy and persistence, such as sporulation, exopolysaccharide and lipopolysaccharide production, and polymeric carbon storage are explored in Supplementary Note 3.
Contrasting assembly processes reveal mechanisms of community-level resilience
While our observations demonstrate clear community-level responses to environmental change, the underlying ecological processes maintaining stability remain unclear. To investigate these mechanisms, we analyzed the ecological processes driving both community assembly and metabolite dynamics64,65,66,67. Our analysis revealed contrasting yet complementary mechanisms: stochastic processes predominantly shaped microbiome assemblages, while deterministic processes governed metabolite assemblages (Fig. 5A–C).
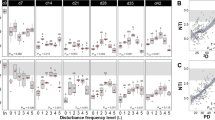
A, B Density plots showing changes in the βNTI of microbial community and metabolome in each month of sampling (n = 32) as well as bar plots indicating the percentage breakdowns of stochastic versus deterministic processes; βNTI was calculated from 16S rRNA ASVs and TWCD metabolites. Assembly processes can be delimited by the red dashed lines: variable selection (βNTI > 2), homogenous selection (βNTI < −2), stochastic assembly (|βNTI| < 2). C Bar plots showing the putative influence of different ecological processes within each month according to the microbial community, bulk metabolites, lipid-, sugar-, and TLC-like metabolites. D Heatmap representing the expression of microbial motility-related genes. Gene length corrected trimmed mean of M-values (geTMM) values for expressed motility-related genes were averaged per each MAG per month; the log10 is represented.
Microbial assembly processes transitioned over time. During the early monsoon (early July), we observed an increase in β-nearest taxon index (βNTI) values > +2, indicating a temporary shift toward deterministic assembly. This coincided with the first major rainfall events (Fig. 1C), suggesting that environmental filtering may have favored rain-responsive taxa. As the monsoon progressed (late July to early August), βNTI values decreased and fell within the stochastic range (|βNTI| < 2), indicating a return to stochastic dynamics that maintained compositional stability.
This transition also aligned with transcriptional changes. Specifically, we detected increased expression of genes associated with microbial motility, such as chemotaxis, flagellar biosynthesis, gliding motility, and pilus systems (Fig. 5D). For example, Myxococcota expressed both gliding and flagellar genes, while Actinobacteria, Chloroflexota, and Desulfobacterota upregulated other mobility pathways. Pseudomonadota showed elevated expression of pilus assembly genes, consistent with twitching motility. These transcriptional patterns suggest that microbial mobility may have supported dispersal and community homogenization, helping to maintain stability under fluctuating moisture conditions.
These shifts in assembly processes may reflect underlying ecological mechanisms that have been proposed in prior studies. For example, the sudden influx of water and nutrients during early monsoon events likely created selection pressures favoring wet-adapted taxa and benefiting tolerant or opportunistic microbial groups9,68. This environmental filtering may have intensified competition between rain-activated and dry-tolerant microbes35, contributing to the transient increase in deterministic assembly that we observed. As rainfall events became more frequent (late July–early August; Fig. 1C), stochastic processes emerged as the dominant force shaping community structure (Fig. 5C). This shift toward stochasticity could be driven by two mechanisms. Increased soil moisture during wetter periods may enhance pore connectivity and facilitate microbial dispersal6, while microbial dormancy during dry periods can buffer against environmental filtering by preserving a reservoir of resilient taxa14,15. These processes may help explain the compositional stability observed during the mid- to late-monsoon period.
In contrast, metabolite assemblages were consistently structured by deterministic processes throughout the monsoon cycle, similar to observations in peatland ecosystems67. βNTI values for metabolites remained outside the stochastic range (|βNTI| > 2), indicating the dominant role of environmental selection and chemical transformations in shaping metabolome composition (Fig. 5A–C).
We observed higher metabolome variability during wet periods, possibly reflecting increased inputs from plant exudates and litter69, as well as microbial byproducts70,71,72 (e.g., osmolytes, secondary metabolites, necromass). Likewise, drier periods were associated with higher relative abundance of sugar- and lipid-like compounds, which may reflect drought-adaptive metabolic strategies. These interpretations align with literature showing that sugar metabolites can function as osmolytes73, and lipid changes often indicate membrane remodeling under water stress74. Abiotic factors, including photodegradation and wind erosion, possibly further shaped organic matter dynamics and metabolome heterogeneity75.
Network restructuring reveals how individual traits shape community responses
To bridge individual adaptations and community-level properties, we analyzed both microbial co-occurrence networks and metabolic interaction networks. Using a random matrix theory (RMT)-based approach to build the co-occurrence networks76, we found that increased precipitation altered network architecture77,78. During dry periods, networks were larger and more modular (higher modularity, transitivity, and node/edge numbers, lower average path distance), whereas wet conditions produced smaller, more hierarchical networks (higher degree centralization, betweenness centralization, and stress centrality) (Fig. 6A and Supplementary Data 4). The shift in network architecture between dry and wet co-occurrence networks represents more than structural reorganization, it reveals fundamental changes in community interaction strategies.
A Co-occurrence networks based on MAGs abundances (TMM) inferred with CoverM for the dry (May and October) and wet (July and August) months. Nodes represent MAGs and are colored based on their Phylum. Node size indicates the degree (number of connections) of each node. Colored ellipses indicate network modules with more than six nodes. B Social networks based on the metabolic interaction potential (MIP) score calculated with SMETANA. Only genome pairs with MIP ≥ 5 were considered for the network. Nodes represent MAGs and are colored based on their Phylum. Node size indicates the degree (number of connections) of each node. C Heatmap showing metabolite donation and uptake patterns between Network Hub microbes and members of other phyla.
During dry periods, the larger, more modular co-occurrence networks suggest a community-wide strategy for resource conservation. To test whether modules represent metabolically interdependent units, we analyzed metabolite sharing patterns within network modules using our SMETANA-derived metabolic interaction data based on the metabolite interaction potential (MIP) score79 (Supplementary Fig. 7a). Our analysis revealed three key findings: (1) Metabolite sharing is not enriched within modules compared to between modules when considering overall interaction frequency (Supplementary Fig. 7b). (2) Modules show distinct metabolic specialization patterns, with amino acid sharing concentrated in specific modules during dry conditions, while aromatic compounds and carboxylic acids showed a more even distribution across modules (Supplementary Fig. 7a). (3) More metabolites seem to be shared within modules during dry conditions (Supplementary Fig. 7b), with the number of MAGs pairs sharing metabolically important metabolites such as amino acids, carboxylic acids, fatty acids, lipids, and nitrogen cycle-related metabolites decreasing around 38% during wet conditions (Supplementary Fig. 7c). These patterns suggest that network modularity reflects functional specialization of metabolic processes rather than general metabolic complementarity. However, we acknowledge that habitat heterogeneity may also contribute to the observed modularity patterns. This organization stands in stark contrast to conventional understanding of drought responses, where networks typically become fragmented and less connected80,81.
Our findings overturn traditional expectations about water availability’s impact on microbial networks. Unlike other ecosystems where wet conditions promote higher interconnectedness82, dry conditions in our system fostered tighter community organization through modular, interconnected network structures. This tight organization likely reflects the need for efficient resource utilization83, where interdependence enhances environmental resilience. Conversely, wet conditions relaxed these tight resource constraints, allowing for more hierarchical organization with diffuse, opportunistic interactions82,84. These changes in structure suggest microbial interactions and responses play a role in community-level resilience. The higher centrality of wet co-occurrence networks suggests wet environments favor dominant species that quickly capitalize on increased nutrients85, while osmotic stress forces other microbes to regulate metabolism more intensively31. This network restructuring aligns with increased selection processes at monsoon onset (Fig. 5A, B), as water simultaneously drives community structure and metabolic activity changes86,87, while enhancing niche connectivity through dispersal mechanisms6.
Network analysis identified key species bridging individual traits and community function. In dry conditions, an Actinobacteria MAG (DNA-4_bin_15_1) served as a network connector, similar to findings from other deserts, including Namib88, Atacama89, and Sahara90. However, this MAG’s high abundance challenges the assumption that desert community stability relies on rare taxa91, suggesting abundant species can be central to ecosystem function.
Analysis of metabolic interaction potential using SMETANA79 (Fig. 6C) further revealed that dry networks showed more extensive metabolic interactions, indicating increased interdependence and active metabolite exchange83. Moreover, this analysis identified additional potential keystone MAGs (network hubs): Methylomirabilota (bco_bin_52) and Actinomycetota (DNA-4_bin_15_1) in dry conditions, and Acidobacteriota (bco_bin_18_1) during wet periods (Fig. 6B). The Methylomirabilota keystone has roles in greenhouse gas cycling92, soil processes93, and vegetation interactions94. SMETANA analysis in detailed mode (Fig. 6C), revealed that this MAG primarily donates amino acids, as previously reported95, while the Actinomycetota hub absorbs them demonstrating how individual metabolic capabilities shape community-level interactions and eventually resilience through energy-expensive amino acid exchange96.
Analysis of metatranscriptomics data confirmed some of these results, as complete biosynthetic pathways for valine, leucine and isoleucine were found to be present in several MAGs, including Methylomirabilota (Supplementary Fig. 8a), which were expressed through the monsoon season (Supplementary Fig. 8b). Additionally, expression of amino acid transporters was detected in a wide range of phyla (Supplementary Fig. 8c) indicating the ability of these microbes to retrieve amino acids from the environment. These insights into network restructuring show that shifts in microbial interactions and metabolic activity are closely linked to the community’s ability to resist and recover from disturbance. Changes in individual responses, such as the activation of stress mechanisms or the activation and deactivation of biosynthetic pathways, respond to community-wide strategies to either promote the collaboration or competition across community members. This coordination across different ecological organization levels may be key to sustaining ecosystem function under changing precipitation patterns.
Strategic gene regulation by Thermoproteota maintains critical ecosystem processes
To understand how individual taxa maintain specific functions within changing networks, we examined Thermoproteota as a model organism. We selected this phylum because: (1) eight MAGs from the Nitrososphaeraceae family represented the only aerobic ammonia oxidizers in our community, (2) nitrogen availability critically influences arid ecosystem function42,97, and (3) Thermoproteota’s genomic features and dynamic gene expression responses provide a unique model for understanding how individual adaptations support community stability.
Using genome-resolved metatranscriptomics, we tracked Thermoproteota’s gene regulation throughout the monsoon season (Supplementary Data 5). The ammonia oxidation genes amoABC (K10944, K10955, K10946) showed unexpected expression patterns, peaking during dry periods (Fig. 7A), with amoC (K10946) showing highest expression as previously described in ref. 55. This counterintuitive pattern reveals a sophisticated adaptation strategy. During dry periods, when most microbes reduce activity, Thermoproteota maintains elevated transcription of nitrification genes, as previously observed in hyperarid environments98. This activity likely reflects an ability to capitalize on reduced competition for limited nitrogen resources. The strategy is further supported by stress tolerance mechanisms, including specialized protein repair systems, manganese-based catalases for ROS management, and efficient nitrogen utilization. Together, these adaptations suggest that Thermoproteota may contribute to sustaining key ecosystem functions even when other taxa become dormant.
A Heatmap representing the expression of Nitrogen metabolism genes in Thermoproteota MAGs. Reaction-level expression, expressed as gene length corrected trimmed mean of M-values (geTMM) value, indicates that at least 50% of the genes of any of its alternative gene sets were expressed (see “Methods”), and is presented here averaged per MAG and per month. Expression was log10-transformed to improve visualization. B Dotplot showing correlations between amoABC gene complex expression and environmental variables (soil moisture and NDVI). C Heatmap of urease complex and transport gene expression (geTMM).
Environmental correlations further highlight how individual adaptations respond to community-level conditions. A strong negative correlation between amoABC expression and both soil moisture and NDVI (Fig. 7B and Supplementary Data 10) suggests that Thermoproteota sustains high nitrification activity under the driest conditions, potentially outcompeting other microbes for limited nitrogen. Moreover, four Thermoproteota MAGs showed high expression of the urease gene cluster (ureABCDEFG) during wet periods (Fig. 7C), indicating these taxa can metabolize both ammonia and urea as energy sources99. This versatility exemplifies how individual-level adaptations enable persistence across variable conditions, providing ecological advantages in nutrient-limited soils. A detailed overview of gene expression changes related to carbon, nitrogen, sulfur, and other metabolic pathways in Thermoproteota MAGs is provided in Supplementary Note 4.
The lack of reconstructed ammonia-oxidizing bacteria MAGs in our system further highlights how Thermoproteota’s strategy of regulated persistence enables their dominance of nitrification processes. Analysis of the taxonomic profiles of other arid soil datasets showcase the widespread presence of Thermoproteota in arid systems worldwide (Supplementary Fig. 2g). This global distribution underscores their importance in stabilizing nitrogen fluxes, a critical ecosystem function in arid environments that are particularly vulnerable to climate change-induced disruptions42, aligning with previous reports of archaeal dominance in arid soils42 and their broader distribution across oligotrophic environments100,101.
The potential ecological impact of Thermoproteota’s expression patterns extends beyond individual survival to influence community-level processes91. First, elevated transcription of ammonia oxidation genes during dry periods suggests that Thermoproteota may help sustain nitrogen cycling when other taxa reduce activity. Second, its network position shifts from peripheral during wet periods to central during dry periods, demonstrating how individual adaptations influence community organization.Third, its consistent expression of stress tolerance and nitrogen metabolism genes across conditions highlights how specialized taxa may contribute to functional resilience under environmental stress.
Discussion
In this study, we used an integrated time-resolved multiomics approach to explore how microbial communities respond to dramatic environmental shifts. Our results reveal how individual microbial stress responses and community-level adaptations interact to support ecosystem function under fluctuating conditions. Using Sonoran Desert soils as a model, we identified three mechanisms driving microbial resilience: (1) stochastic processes supporting stability, (2) dynamic network reorganization, and (3) coordinated stress responses. We also highlight Thermoproteota as a key taxon in arid soils, with potential as a model for studying adaptation to aridity.
First, regarding microbial network responses, our findings challenge the prevailing view that drought predominantly disrupts microbial networks, as often observed in analysis of microbial co-occurrence networks78,81,102. Instead, we observed larger and more interconnected microbial networks under dry conditions, aligning with evidence that stress can foster tighter microbial associations103. This tightly organized community structure became more diffuse during wet conditions, suggesting dynamic reorganization based on resource availability (Fig. 6A and Supplementary Data 4). These results may reflect unique adaptations of microbial communities in bare soils, where plant buffering effects are limited104,105.
We acknowledge that factors beyond precipitation, including soil microenvironmental heterogeneity, pH gradients, and nutrient patchiness, may also contribute to observed network patterns. Our single-site study design limits our ability to disentangle precipitation effects from other environmental variables, and future multi-site studies would help clarify these factors. However, our temporal sampling within the same location helps control for confounding variables, as major soil properties remain constant while precipitation patterns vary seasonally. The observed increase in network connectivity under drought supports the stress gradient hypothesis106,107,108. During dry periods, certain taxa, such as Thermoproteota and Actinomycetota, identified as network hubs due to being highly connected to other nodes (Zi >2.5) within their own module, as well as to several other modules (Pi >0.62) facilitate functional complementarity and efficient resource sharing among community members. Conversely, wet conditions promoted more centralized networks with increased negative correlations, indicating the rise of dominant microbes and heightened competition84,108. This dynamic network restructuring supports microbial diversity and ecosystem function, as evidenced by stable community composition across monsoon cycles and dominance of stochastic assembly processes (Fig. 5C). These patterns suggest that microbial communities may balance stability and adaptability through interaction strategies under fluctuating environments.
We propose that microbial network reorganization supports community resilience by coordinating responses at multiple biological levels. Tight, modular networks during dry periods enhance resource exchange and foster interdependencies that promote stability under stress79,80. This is consistent with the expression of amino-acid biosynthesis and transport systems across the monsoon season for cross-feeding. In contrast, diffuse and centralized networks during wet periods reflect increased metabolic flexibility and opportunistic interactions, enabling communities to rapidly exploit resource pulses while maintaining core functions13.
Despite limitations in metagenomics-derived network analyses, such as the inability to discern correlations, which may arise from indirect interactions or shared niches109 or that only a subset of the community may be represented, our approach demonstrates the utility of network analysis for inferring ecological functions80,81,110. While assembled MAGs represented only 10% of sequenced reads, as high strain heterogeneity may have prevented us to recover a larger number of MAGs47,111,112, our analysis showed that assembled MAGs captured 86% of the community’s taxonomic diversity. This taxonomic coverage captures key functional and ecological traits. For example, the prominent role of Thermoproteota during dry conditions (Fig. 7A) highlights its contribution to nitrogen cycling, a critical process for maintaining ecosystem function. Its widespread presence in arid environments globally (Supplementary Fig. 2g) indicates its ecological relevance for resilience mechanisms and highlights its potential as a biotechnological target. On the other hand network hubs support the community by facilitating the exchange of metabolically important metabolites such as amino acids or sugars (Fig. 6C). To validate their functional significance, we examined expression patterns of key amino acid biosynthesis and transporter genes in the hub MAGs. Hubs from the dry season network always express these biosynthetic pathways and transporters (Supplementary Fig. 8), with a notable increase following the start of the monsoon: Methylomirabilota (geTMMMay = 2.28, geTMMJuly_1 = 18.66) and Actinomycetota (geTMMMay = 1.22, geTMMJuly_1 = 8.944). In contrast, the wet season network hub has a delayed increase of its expression, peaking in August when the other hubs’ expression was starting to decline: Acidobacteriota (geTMMMay = 0.14, geTMMAugust = 19.24). These coordinated interactions generate reciprocal dependencies that optimize resource utilization, creating dynamic feedback mechanisms where individual metabolic capabilities influence community assembly patterns while established network associations shape individual metabolic investments and gene expression priorities. This bidirectional coupling is supported by widespread expression of amino acid biosynthetic pathways and transport systems across multiple taxa within network modules (Supplementary Fig. 8), demonstrating how individual metabolic capabilities play a role in shaping community structure and, in turn, are shaped by the broader microbial community79.
Complementing these network dynamics, the analysis of organic matter profiles reveals deterministic changes driven by both community-level metabolic adjustments and individual stress responses. Unlike plant-dominated ecosystems39,40, the prevalence of lipid- and protein-like compounds in these soils underscores the strong microbial influence on soil organic matter composition41. During dry periods, the accumulation of osmolytes and stress-related compounds reflects individual microbial adaptations113, while shifts in metabolite profiles during wet periods indicate community-level metabolic reorganization. These changes mirror network restructuring, as metabolite exchange relationships transition between tight and diffuse network arrangements, further illustrating the integration of microbial activities in maintaining ecosystem resilience.
At the individual organism level, our analysis revealed shared and specialized survival strategies among taxa, suggesting strong selective pressures for traits that support both individual survival and community function. Actinomycetota employed multiple adaptations including high GC, chaperone proteins114, spore formation115, osmolyte production116, and versatile metabolic strategies117. Similarly, Thermoproteota maintained function through stress response mechanisms including chaperones, DNA-repair proteins, and stress-related proteins like katG, katE, dps, and superoxide dismutases (Fig. 7A and Supplementary Fig. 6). The consistency of stress response mechanisms across diverse taxa suggests strong selective pressure for maintaining both individual survival and community function118. The widespread distribution of heat shock proteins, osmolyte production pathways, and oxidative stress responses indicates that these mechanisms represent essential adaptations to arid conditions14,102. These diverse survival strategies underscore the critical role of functional redundancy and metabolic versatility in maintaining community stability, demonstrating how alternative strategies can achieve similar community-level outcomes14,15. These individual MAG-level adaptations form the building blocks of network-mediated resilience, enabling communities to buffer environmental perturbations and sustain key ecosystem processes.
Together, these findings across multiple organizational levels provide a framework for understanding microbial resilience in arid environments. Stochastic processes support stable community structures, while dynamic network reorganization and MAG-level stress responses facilitate adaptation to extreme conditions (Fig. 8). The central role of Thermoproteota in nitrogen cycling suggests its potential as an ecological indicator of resilience. These strategies point to common mechanisms of community adaptation and resilience potentially broadly applicable across arid ecosystems, and potentially extend to non-arid environments facing environmental stress.
Resilience emerges from interactions between individual-level mechanisms (genomic traits, metabolic functions, gene regulation) and community-level processes (composition, assembly). These interactions drive network restructuring, enabling resource sharing and protection during dry periods, and resource exploitation during wet periods. Individual-level adaptations include high GC content, DNA repair mechanisms, and osmolyte production, while community-level responses involve both co-occurrence and metabolic interaction networks.
While these multi-level resilience mechanisms currently enable functional persistence through monsoon cycles, they also suggest vulnerabilities to future climate change. Increased aridity, prolonged droughts, or disruptions to stochastic processes may exceed the adaptive capacity of these microbial communities, leading to cascading effects and destabilization31,82,119,120. To safeguard critical ecosystem functions, future efforts should prioritize functional redundancy within microbial communities, conserving microbial diversity hotspots, and monitoring network reorganization thresholds. Additionally, integrating predictive modeling with multiomics datasets could facilitate early detection of resilience breakdowns under changing climate scenarios. Tracking key taxa such as Thermoproteota will be essential, providing critical data on how microbial communities navigate environmental extremes and informing strategies to safeguard ecosystem functionality.
Methods
Soil collection
Soil samples were collected at the Saguaro National Park West (Permit #: SAGU-2021-SCI-0006), Tucson, Arizona (32° 15 ‘05.4“N, 111° 09’ 40.2“W) from a depth of 10–15 cm using ethanol-sterilized shovels. Sample collection occurred across 9 time points spanning pre-monsoon (May 2021, 2022), monsoon (June–August 2021), and post-monsoon (September-October 2021) seasons, with 4 biological replicates (i.e., 4 sampling sites per month) per time point (total n = 36). Samples were transported on ice to the laboratory, and sequentially sieved through 4000 and 2000 µm Fieldmaster® sieves (Science First, Yulee, FL). Samples were stored at −80 °C until analysis.
The sampling site is located within the Sonoran Desert in the Southwestern USA. A region that experiences extreme temperatures exceeding 40 °C, with maximum temperatures reaching 48 °C121. It has an annual precipitation average of 200 mm (1980–2016), with regional variations across the desert landscape122. The area’s distinctive bimodal precipitation pattern includes winter frontal system rainfall and summer monsoon storms123, with a characteristic dry period in late spring.
Soil physicochemical properties
For soil moisture, around 5 g of soil were placed in 15 ml Eppendorf tubes (3 replicates per sample) and lyophilized on a FreeZone 2.5 l Benchtop Freeze Dryer (Labconco, Kansas City, MO) for 72 h. Moisture content (%) was calculated as: [(initial weight − lyophilized weight)/initial weight] × 100. For pH measurements, 2 g of soil were mixed with 6 ml double-distilled water, vortexed for 15 min, centrifuged for 3 min, and incubated at room temperature for 30 min. pH was measured twice per sample using a benchtop Orion Star A211 pH meter (Thermo Scientific, Waltham, MA).
Total carbon (TC) and nitrogen (TN) were measured for all samples (n = 36) using a ECS 4010 Elemental combustion analyzer (Costech Analytical Technologies, Valencia, California) by the Arizona Laboratory for Emerging Contaminants (ALEC) at the University of Arizona. Major cations (K, Ca, Na, Mg) and trace metals (V, Cr, Fe, Co, As, Se) were quantified using an Agilent 7700 inductively coupled plasma mass spectrometer (ICP-MS) (Agilent, Santa Clara, CA) as described before124 with the exception that the US EPA Method 3051A was used for soil acid digestion. All measurements are available in Supplementary Data 1.
Daily precipitation data, soil and air temperature measurements were retrieved from the U.S. Climate Reference Network Data website (https://www.ncei.noaa.gov/pub/data/uscrn/). Precipitation data of each sampling point was calculated as the aggregated precipitation of the 4 days previous to the sampling. Normalized difference vegetation index (NDVI) values were retrieved from the Terra Moderate Resolution Imaging Spectroradiometer (MODIS) Vegetation Indices (MOD13Q1) Version 6.1125 dataset. As the NDVI values are measured on a 16-day basis, the NDVI of each sampling point was determined by interpolation using a LOESS function fitted using data collected between 05/09/2021 and 05/28/2022.
For multivariate analysis of the environmental variables, variables were selected to avoid collinearity by calculating the variance inflation factor, using the performance package126, and removing those with variance inflation factor larger than 10. Remaining environmental variables were then standardized by z-score with the deconstand function (method = standardize) of the vegan package. Standardized values were used to produce a hierarchical clustering of the samples using Manhattan distances. Additionally, a PCA was also performed with this data to better visualize the differences between samples.
FTICR-MS sample preparation and data preprocessing
Ultra-high resolution soil organic matter profiling was conducted using Direct Infusion Fourier Transform Ion Cyclotron Resonance Mass Spectrometry (FTICR-MS). Soil metabolites were extracted through sequential water and methanol extraction. A total of 2.5 g of each 2021 soil sample (n = 32) was mixed with 5 ml of ddH2O, followed by two rounds of vortexing and 2-h sonication. Samples were then centrifuged at 5000 rpm for 5 min. The supernatant was collected and stored at −20 °C. The remaining soil mix underwent two rounds extra of extraction with the same protocol using 5 ml of HPLC-grade methanol instead of ddH2O. Combined water and methanol extracts were purified using solid phase extraction (SPE) to remove contaminants127. Extracts were eluted using 1.5 mL of HPLC methanol, and the final purified extracts were shipped to the Pacific Northwest National Laboratory (PNNL) for mass spectrometry analysis.
High-resolution mass spectra data was collected with a 12-Tesla Bruker SolariX FTICR mass spectrometer (Bruker, SolariX, Billerica, MA) located at the Environmental Molecular Sciences Laboratory at PNNL in Richland, WA. Samples were directly infused into the instrument using a custom automated direct infusion cart that performed two offline blanks between each sample. The FTICR-MS was outfitted with a standard electrospray ionization (ESI) source, and data was acquired in negative and positive mode with the needle voltage set to +4.2 kV and −4.2 kV, respectively. Data was collected at 4 MW, and the ion accumulation time (IAT) was optimized for each polarity. In negative mode the IAT was set to 0.08 s and data was measured from 98.28 to 2000 m/z with a resolution of 300 K at 333.1118 m/z. In positive mode the IAT was set to 0.08 s and the data were measured from 147.42 to 1400 m/z with a resolution of 380 K at 385.1176 m/z. One hundred forty-four scans were co-added for each sample and internally calibrated using OM homologous series separated by 14 Da (–CH2 groups). The mass measurement accuracy was typically within 1 ppm for singly charged ions across a broad m/z range (100–900 m/z). Bruker Data Analysis (version 4.2) was used to convert raw spectra to a list of m/z values by applying FTMS peak picker module with a signal-to-noise ratio (S/N) threshold of 7 and absolute intensity threshold of 100. Chemical formulae were then assigned using Formularity128, with parameter: S/N > 7, and mass measurement error <0.5 ppm, allowing only for the presence of C, H, O, N, S, and P. We present results from positive mode analysis only, which revealed the strongest shifts in metabolite composition.
Thermodynamic indexes for the FTICR molecular formulae, chemodiversity indexes and putative biochemical transformations were calculated using MetaboDirect (version 1.0.7)129 keeping only peaks that were present in at least 4 samples and with the sum normalization method. Ordination analysis based on FTICR-MS data was done based on Jaccard distances using a presence/absence matrix of FTICR-MS peaks. The number of nitrogen and sulfur-containing transformations were inferred based on the assigned transformation names (Supplementary Data 6).
DNA extraction
DNA was extracted from 250 mg soil samples using the DNeasy PowerSoil Pro Kit (Qiagen, Hilden, Germany) according to manufacturer protocols. DNA quantification was performed using the Qubit DNA high-sensitivity kit (ThermoFisher Scientific, Waltham, MA). Separate DNA extractions were conducted for 16S amplicon sequencing and shotgun metagenomics. The University of Arizona Genetics Core (RRID: SCR_012429) performed all library preparation and sequencing data generation.
16S rRNA amplicon sequencing
Amplicon sequencing was conducted on samples collected from May to October 2021 (n = 32). The V3-V4 regions of the 16S rRNA gene were amplified using previously designed primers130 (Forward: 5’-CCTACGGGNGGCWGCAG-3’ and Reverse: 5’-GACTACHVGGGTATCTAATCC-3’), with Illumina™ overhang adapters at their 5′ ends. Amplicon amplification was performed using KAPA HiFi HotStart Ready Mix (Roche Sequencing Solutions, Pleasanton, CA) following manufacturer protocols. A secondary PCR utilized Nextera XT v2 Index kits sets A-D (Illumina, San Diego, CA). Amplicon purification employed HighPrep PCR magnetic beads (MagBio, Gaithersburg, MD) at 0.8X and 1.2X volumes for first and second PCR products, respectively. Libraries were quantified using QuantiFlor dsDNA System (Promega, Madison, WI) on a BioTek FLX 800 plate reader (BioTek, Charlotte, VT). Size determination of pooled amplicons used an AATI Fragment Analyzer with HS NGS Fragment kit (Agilent, Santa Clara, CA). Pools were diluted to <20 nM and quantified using a Roche LightCycler 480-II with KAPA Illumina Library Quantification Kit (Roche Sequencing Solutions, Pleasanton, CA). Sequencing was performed on an Illumina MiSeq platform using the v3-600 cycle kit (Illumina, San Diego, CA), generating 250 bp paired-end reads.
Metagenomics sequencing
Metagenomic libraries were prepared from samples collected in 2021 during May (Sampling time point 1; n = 4), July (Sampling time points 3, 4 and 5; n = 12), August (Sampling time point 6; n = 4), October (Sampling time point 8; n = 4), and May 2022 (Sampling time point 9; n = 4). Quality control involved genomic DNA (gDNA) quantification using QuantiFlor dsDNA System on a BioTek FLX 800 plate reader, and size and integrity assessment using AATI Fragment Analyzer with HS NGS Genomic DNA kit. Genomic DNA (350–550 ng) was sheared to 200 bp using standard conditions on an S2 sonicator (Covaris, Woburn, MA). Library preparation utilized NEBNext Ultra II DNA Library Prep Kit for Illumina (New England Biolabs, Ipswich, MA) with NEBNext Multiplex Oligos for Illumina UDI primer pairs (New England Biolabs, Ipswich, MA), employing 3 PCR enrichment cycles. Libraries were purified using 0.8X volume HighPrep PCR magnetic beads, quantified with QuantiFlor dsDNA System on a BioTek FLX 800 plate reader, and sized using AATI Fragment Analyzer with HS NGS Fragment kit. Size-adjusted libraries were quantified using Roche LightCycler 480-II with KAPA Illumina Library Quantification Kit before equal pooling. Initial metagenomic sequencing generated 2 × 150 bp paired-end reads using one full lane of a NovaSeq 6000 S4 flow cell v1.5 (300 cycles) on the NovaSeq 6000 platform (Illumina, San Diego, CA). A second sequencing round was performed for six selected samples (one per time point) using another full lane of a NovaSeq 6000 S4 flow cell v1.5 (300 cycles).
RNA extraction and sequencing
Metatranscriptomic sequencing was conducted on the same samples used for metagenomic analysis (n = 28). RNA was extracted from 2 g soil samples using RNeasy PowerSoil Total RNA kit (Qiagen, Hilden, Germany) following manufacturer's protocols. DNA removal was performed using DNase I (New England Biolabs, Ipswich, MA), followed by additional purification using RNA Clean Kit T2030 (New England Biolabs, Ipswich, MA). Initial RNA quantification employed Qubit RNA high sensitivity kit (ThermoFisher Scientific, Waltham, MA), with RNA integrity assessed via 2100 Bioanalyzer (Agilent, Santa Clara, CA) before submission to the University of Arizona Genetics Core (RRID: SCR_012429) for library preparation and sequencing.
Prior to library preparation, secondary RNA quantification was performed using Qubit Fluorometer 2.0 with Qubit Broad Range RNA Quantification Kit (ThermoFisher Scientific, Waltham, MA), and RNA quality was evaluated using AATI Fragment Analyzer with 15 nt RNA Kit. RNA ribodepletion was conducted on 50–500 ng input RNA using NEBnext rRNA Depletion Kit (Bacteria) (New England Biolabs, Ipswich, MA). Libraries were prepared using NEBNext Ultra II Directional RNA Library Prep Kit for Illumina (New England Biolabs, Ipswich, MA) and NEBnext Multiplex UDI Primer Pairs for Illumina (New England Biolabs, Ipswich, MA), with 9–14 PCR enrichment cycles. Double purification of libraries used 0.8X volume HighPrep PCR magnetic beads. RNA library quantification and sequencing followed the same protocols described for DNA libraries.
Comparison with other datasets
Metagenomics read signatures were compared with shotgun metagenomics reads available in public databases from other arid ecosystems across the world including the Negev Desert (PRJEB36534)14 and (PRJNA657906)131, Sinai Desert (PRJNA810925)132, Xilingol League (PRJNA916612)133, Mojave Desert (PRJNA620793)134, Cuatro Cienagas (PRJNA857298)135, as well as our same study ecosystem, the Sonoran Desert (PRJNA664514)136, using MASH distances137. Microbial taxonomic profiles of all datasets were inferred with SingleM (v0.18.2)138.
Microbial community profiling and analysis
Microbial community profiles based on ASVs were generated using 16S rRNA gene amplicon sequencing reads with the DADA2 (v1.26) pipeline139. Briefly, adapter trimming was performed using cutadapt140, trimmed reads were then filtered and quality trimmed to a minimum length of 50 and maximum expected error of 2. Error rates were learned directly from processed reads using DADA2 sample inference algorithm to infer sequence variants. Fully denoised sequences were generated by merging ASVs from forward and reverse reads followed by chimeric sequence removal. Taxonomy was assigned using a naive Bayesian classifier141 with Genome Taxonomy Database (GTDB) version r220142.
To account for 16S rRNA gene copy number variation effects on community analysis143, metagenomic-based taxonomic profiling was also performed using SingleM (v0.18.2)138 on metagenomic raw reads following tool authors’ recommendations. Alpha diversity indexes were calculated as the average value of the indexes obtained from each marker gene. For other downstream analyses we used the rplP gene (encoding ribosomal protein L16 L10e), due to its effectiveness in distinguishing between closely and distantly related genomes144.
Statistical analyses were conducted in R (v4.3.2)145. The 16S and SingleM datasets were rarefied to 25,000 and 800 reads, respectively, for alpha and beta diversity calculations. Other analyses using these datasets were made using the unrarefied values. Observed and estimated richness (Shannon diversity index and Chao1) were calculated with the phyloseq R package146, while beta diversity analysis was done with Bray Curtis distances calculated with the vegan package147. Principal coordinates analysis (PCoA) was performed using phyloseq, with PERMANOVA testing for differences between sampling time points.
For phylogenetic analysis of 16S rRNA data, multiple sequence alignment was done using the DECIPHER package (version 2.26.0)148. A maximum-likelihood phylogenetic tree was constructed using FastTree (version 2.1.1)149, with a generalized time-reversible model and gamma option for branch length rescaling and Gamma20-based likelihood computation. The tree was midpoint-rooted using the phytools package (version 2.3.0)150. Compositional stability was quantified by calculating a multi-order compositional stability index103, between each pair of samples collected from the same subsite at two adjacent time points.
Metagenome processing and assembly
Sequencing generated 44–180 million read pairs during the first round and 416–588 million read pairs during the second round per sample. Read quality assessment was performed using FastQC (v0.11.9)151. Raw reads underwent preprocessing using the RQCFilter pipeline (BBtools version 38.90)152. Cleaned reads were assembled with MEGAHIT (v1.2.9)153 with the “--presets meta-large” option as recommended for high-biodiversity soil metagenomes. To optimize metagenome assembled genome (MAG) recovery, eight metagenome assemblies were generated using two strategies: (1) Co-assembly of samples from the same sampling point (6 assemblies: May, July_2, July_3, August, October, May_2022) and (2) Co-assembly of samples from the same sequencing run (2 assemblies: one with all samples from the first sequencing run (28 samples), other with all samples from the second sequencing run (6 samples), see Metagenomics sequencing section). Assemblies were then filtered to retain only contigs larger than 2000 bp. In total we retrieved between 0.3 and 1.5 million contigs from each assembly with a N50 between 3200 and 3500 bp. Read mapping to the filtered assemblies using CoverM (version 0.7.0)154, with flags -p minimap2-sr -m trimmed_mean --min-read-percent-identity 0.95 --min-read-aligned-percent 0.75, indicated they accounted for 29.74–42.89% of the reads of each sample.
Recovery of metagenome assembled genomes
For MAG recovery, filtered contigs of each assembly were binned separately using CONCOCT (v1.0.0)155, MetaBAT2 (v2.12.1)156, and MaxBin2 (v2.2.6)157 with default options. The resulting bin sets were refined using the bin refinement module from metaWRAP (v1.3.0)158, with parameters -c 50 and -x 10. Refined bins were manually curated using anvi’o (version 8)159, based on sequence coverage and the presence of single-copy core genes identified with anvi-run-hmms. Bin quality and completeness were checked using CheckM2 (v1.0.2)160. Only bins with more than 50% completeness and less than 10% contamination (medium quality draft genomes) were retained and used for the analysis resulting in a set of 661 bins. Bins generated from the different assemblies were dereplicated at 95% average nucleotide identity similarity (species-level161) using galah (v0.4.0)162 resulting in 282 dereplicated MAGs. All downstream analyses using both the metagenomics and metratranscriptomics data refer exclusively to this dereplicated MAG set. Read mapping using CoverM, with the same parameters as before, revealed that the dereplicated MAGs accounted for 3.40–9.11% of the reads from each sample, comparable to other metagenomics studies of soil microbial communities (2.7–22.4%)111. This resulted in mean coverage depths for each MAG ranging from 0.9X to 100X per sample, with over 50% of MAGs exceeding 3X average coverage across all samples. This showcases how the multilayered co-assembly strategy supported confident genome binning, robust taxonomic classification, and functional potential profiling that would not have been possible otherwise. To determine how much of the microbial communities was represented by the recovered MAGs, we compared the taxonomic profiles of the reads, the assemblies, and the recovered MAGs using SingleM appraise at 86% sequence identity (genus level) and the –imperfect flag.
Dereplicated MAGs were taxonomically classified using the Genome Database Taxonomy toolkit (GTDB-tk) (v2.4.0)163 and the R09-RS220 database. Phylogenetic trees for bacterial and archaeal species were built separately using the de novo workflow of GTDB-tk based on 120 marker genes for bacteria and 53 marker genes for archaea. Trees were rooted using the p__Patescibacteria and p__Altiarcheota for the bacteria and the archaeal tree respectively. Visualization of the trees was performed using the ggtree164 and ggtreeExtra165 R packages.
Functional annotation of the dereplicated MAGs was performed using the automated DRAM pipeline166. Additional databases used in the same DRAM run included the NCycDB167, SCycDB168, and PCycDB169. KEGG annotations were integrated with annotations from NCycDB, SCycDB, and PCycDB based on KEGG Orthology (KO) numbers. This integration enabled differentiation between closely related orthologs from KEGG databases (e.g., K19044 encoding either amoA or pmoA) using metabolism-specific database results.
MAG stress tolerance trait identification was based on literature-derived definitions, KEGG database annotations170, and modified microtrait definitions171 as detailed in Supplementary Data 7. Traits were considered present in a MAG when at least 50% of trait-defining genes were detected. Functional pathway analysis for carbon, nitrogen, and sulfur metabolism in MAGs was conducted at both reaction (Supplementary Data 8) and pathway levels (Supplementary Data 9). Reaction and pathway definitions, including key enzymes, were sourced from the KEGG database and literature review. A reaction was considered present in a MAG when at least 50% of the required genes were detected. Similarly, pathways were considered present when a MAG contained at least 50% of pathway reactions plus required key enzymes.
Maximal growth rate for each of the MAGs was predicted based on genome-wide codon usage statistics using the gRodon2172 R package. Briefly, minimal doubling time was predicted with the predictGrowth function using mode=partial to account for the medium-quality draft nature of the dereplicated MAGs. Maximal growth rate was calculated as 1/minimal doubling time.
Metatranscriptomics processing
The quality of metatranscriptomics reads was evaluated with FastQC151. Raw reads were processed with the RQCFilter pipeline (BBtools version 38.90)152 to trim adapters and filter contaminants (parameters: removehuman = t removecat = t, removedog = t removemouse = t lambda = t). Further filtering of risobomal RNA was performed using SortMeRNA (version 4.3.6)173. Bowtie2 (v2.5.4)174 was used to map the filtered and trimmed reads against the dereplicated MAGs. The resulting SAM files were sorted using samtools175. Mapped reads counts were obtained at the transcript level and summarized based on gene_id (parameters: -t CDS -g gene_id) using featureCounts (version 2.0.6)176. Only transcripts with read counts higher than 5 and present in at least 3 samples were kept for downstream analysis. Read counts were normalized following the gene length corrected trimmed mean of M-values (geTMM) method177.
A MAG was considered to express a metabolic pathway based on KEGG definitions as follows:
-
1.
A reaction was considered expressed if at least 50% of the genes of any of its alternative gene sets were expressed (KOs forming a gene set are defined by plus sign in the definition, alternative gene sets are separated by commas).
-
2.
A module was considered expressed if at least 50% of its reactions were expressed. This 50% cutoff was selected to account for the incompleteness of MAGs, which are often neither fully assembled nor circularized, resulting in partial metabolic pathways.
Microbial and metabolite community assembly analysis
Microbial phylogenetic signal
Microbial community assembly shifts during the monsoon season were analyzed using ecological null modeling64,65, based on the assumption that phylogenetically related ASVs share ecological similarities (phylogenetic signal)65. Phylogenetic signal analysis encompassed 19 environmental variables (detailed in Supplementary Data 1), using the between-ASV difference in environmental optima and between-ASV phylogenetic distance.
Environmental optima were calculated using abundance-weighted mean values for each environmental variable using the analog package (version 0.17.6)178. Between-ASV environmental optima differences were computed using Euclidean distance for all environmental variables. Between-OTU phylogenetic distances were derived from the microbial phylogenetic tree using the adephylo package (version 1.1.16)179. Phylogenetic signal assessment was done using a Mantel correlogram based on a Pearson’s correlation coefficient across 50 distance classes with 999 permutations, applying progressive Holm-Bonferroni correction for multiple testing.
Metabolite dendrogram construction
To analyze monsoon-mediated shifts in metabolite assembly processes in arid soils, we applied ecological null modeling methods originally developed for microbial data64,65, and then adapted for the analysis of metabolites66,67. This framework allows us to infer whether shifts in community composition (microbial or metabolite) are primarily shaped by consistent environmental filters, random processes, or spatial dispersal dynamics, key to understanding how monsoon regimes modulate soil biogeochemistry in arid ecosystems.
Metabolites were categorized into three groups based on molecular class assignments180: group 1: lipid-like, group 2: carbohydrate-, protein- and amino sugar- like compounds referred to as sugars and group 3: tannin-, lignin- and condensed hydrocarbon-like compounds referred to as TLC.
Relational dendrograms were constructed following the methodology described by ref. 66, for both bulk metabolite data and the three metabolite pools using three approaches (1) metabolite molecular characteristics (MCD) based on metabolite structural properties (e.g., elemental composition, double-bond equivalents, modified aromaticity index, and Kendrick’s defect), (2) metabolite potential biochemical transformations (TD) based on putative chemical conversions derived from ultrahigh mass resolution differences between metabolites enabled with FTICR MS data, and (3) metabolite transformation-weighted characteristics (TWCD) integrating both MCD and TD dendrograms. Dendrogram construction utilized binary presence/absence values to eliminate charge competition bias in abundance estimates66.
Microbial and metabolite β-diversity analysis and ecological null modeling
Ecological processes influencing microbial and metabolome assemblages during monsoon periods were estimated using βNTI and Raup-Crick Bray-Curtis index (RCBC), analyzing the microbial phylogenetic tree and TWCD metabolite-derived dendrogram as previously described64,65,66. Briefly, microbial and metabolite β-mean nearest taxon distance (βMNTD) was calculated using the comdist function from the picante package (v1.8.2) and compared to a null distribution generated from 1000 randomizations. The βNTI was then computed by standardizing the observed βMNTD against the null expectation, following the approach described in ref. 64.
Following this approach, |βNTI| > 2 indicates predominance of deterministic processes, such as environmental abiotic factors and biotic interactions influence microbial community turnover and differences in production and degradation rates of metabolites. Meanwhile, |βNTI| < 2 indicates that stochastic processes such as random processes of birth, death, colonization, and speciation of microbial communities181 cause changes in species diversity and composition. Similarly, metabolite shifts caused by physical forces or vector movements are considered as stochastic changes66. Moreover, when βNTI > 2, variable selection indicates that divergent environmental factors drive high compositional turnover between a pair of communities. Conversely, when βNTI < −2, homogeneous selection indicates consistent selective pressures from stable environmental conditions are the primary cause of low compositional turnover between local communities64.
Furthermore, the RCBC turnover index was used to determine the influence of stochastic ecological processes. Briefly, the observed presence/absence-based Bray-Curtis values from pairwise community comparisons were calculated and compared to a null expectation (generated through 1000 randomizations). Deviations of observed values from the null comparisons were then normalized between +1 and −1 to produce the RCBC metric. An RCBC > 0.95 with |βNTI| < 2 indicates that higher-than-expected compositional differences between communities (or metabolomes) are primarily due to dispersal limitation, allowing ecological drift. In contrast, an RCBC < −0.95 with |βNTI| < 2 suggests that lower-than-expected compositional differences are driven mainly by homogenizing dispersal. Finally, when |RCBC | <0.95 and |βNTI| < 2, compositional turnover between communities (or metabolomes) is not predominantly influenced by selection, dispersal, or ecological drift, a condition referred to as undominated65.
Co-occurrence network analysis
MAG interactions were analyzed through co-occurrence microbial networks using a RMT-based approach76. Networks were constructed using the Molecular Ecological Network Analysis pipeline (MENA)76,182 website (http://ieg4.rccc.ou.edu/mena/login.cgi). Two separate networks were constructed, one using data from sampling points from two dry months (May and October) and another with data from two wet sampling points (July and August). Metagenome-assembled genomes abundances expressed as trimmed means of M-values (TMM) were obtained using CoverM. Separate abundance tables for each network were uploaded and processed using the MENA pipeline. Briefly, a similarity matrix was built using Spearman correlations of features present in at least half of the samples. An association threshold was then determined using the RMT approach182. Network topological parameters were calculated with the MENA pipeline. Module determination employed a greedy modularity optimization algorithm183. Node topological roles were assigned based on within-module connectivity (Zi) and participation coefficient (Pi)76. Networks were visualized using the igraph184 and ggnetwork185 R packages.
Metabolic interaction network
Genome scale metabolic (GEM) draft models were reconstructed for each MAG using CarveMe186, with distinct templates for bacterial and archaeal MAGs. Models were constructed without gap filling to avoid false-positive cross-feeding metabolic interaction predictions187.
To better understand metabolic interactions among the microorganisms in each of the co-occurrence networks, metabolic interactions were assessed using SMETANA79. SMETANA was run in global mode for each possible pair of MAGs within each of the two co-occurrence networks to calculate a metabolic interaction potential (MIP) score between each pair. Only pairs with MIP scores ≥5 were considered highly interacting and were used for building the metabolic interaction network188. The cross-feeding potential between pairs of highly interacting MAGs was then investigated using the detailed mode of SMETANA, keeping only those metabolites with a SMETANA score higher or equal than 0.1, as well as those related with nitrogen and sulfur metabolism. Node topological roles were assigned based on within-module connectivity (Zi) and participation coefficient (Pi)76. Networks were visualized using the igraph and ggnetwork R packages.
Data availability
All environmental measurements are available as part of Supplementary Data 1. Raw metagenomics, metatranscriptomics, and amplicon sequencing data is available at the NCBI database (Bioproject Accession: PRJNA1193034, https://www.ncbi.nlm.nih.gov/bioproject/?term=PRJNA1193034). All supplementary datasets mentioned in the manuscript, as well as the processing report of FTICR-MS data, assembled MAGs sequences and their annotations are available in the OSF repository with https://doi.org/10.17605/OSF.IO/WR6Z4.
Code availability
Code for all statistical analyses in this manuscript are available in its GitHub repository: https://github.com/tfaily-lab/arid_dual_resilience_paper.
Change history
25 September 2025
In the version of this article initially published, a link was missing in the data availability section (https://doi.org/10.17605/OSF.IO/WR6Z4), which is now amended in the HTML and PDF versions of the article.
References
Philippot, L., Chenu, C., Kappler, A., Rillig, M. C. & Fierer, N. The interplay between microbial communities and soil properties. Nat. Rev. Microbiol. 22, 226–239 (2024).
Nguyen, J., Lara-Gutiérrez, J. & Stocker, R. Environmental fluctuations and their effects on microbial communities, populations and individuals. FEMS Microbiol. Rev. 45, fuaa068 (2021).
Zhou, J. et al. Stochasticity, succession, and environmental perturbations in a fluidic ecosystem. Proc. Natl. Acad. Sci. USA 111, E836–E845 (2014).
de Vries, F. T. & Shade, A. Controls on soil microbial community stability under climate change. Front. Microbiol. 4, 265 (2013).
Allison, S. D. & Martiny, J. B. H. Colloquium paper: resistance, resilience, and redundancy in microbial communities. Proc. Natl. Acad. Sci. USA 105, 11512–11519 (2008).
Philippot, L., Griffiths, B. S. & Langenheder, S. Microbial community resilience across ecosystems and multiple disturbances. Microbiol. Mol. Biol. Rev. 85, e00026-20 (2021).
Shade, A. et al. Fundamentals of microbial community resistance and resilience. Front. Microbiol. 3, 417 (2012).
Louca, S. et al. Function and functional redundancy in microbial systems. Nat. Ecol. Evol. 2, 936–943 (2018).
Schimel, J., Balser, T. C. & Wallenstein, M. Microbial stress-response physiology and its implications for ecosystem function. Ecology 88, 1386–1394 (2007).
Barnard, R. L., Osborne, C. & Firestone, M. Responses of soil bacterial and fungal communities to extreme desiccation and rewetting. ISME J. 7, 2229–2241 (2013).
Schimel, J. P. Life in dry soils: effects of drought on soil microbial communities and processes. Annu. Rev. Ecol. Evol. Syst. 49, 409–432 (2018).
Ouyang, Y. & Li, X. Recent research progress on soil microbial responses to drying–rewetting cycles. Sheng Tai Xue Bao 33, 1–6 (2013).
Bardgett, R. D. & Caruso, T. Soil microbial community responses to climate extremes: resistance, resilience and transitions to alternative states. Philos. Trans. R. Soc. Lond. B Biol. Sci. 375, 20190112 (2020).
Meier, D. V., Imminger, S., Gillor, O. & Woebken, D. Distribution of mixotrophy and desiccation survival mechanisms across microbial genomes in an arid biological soil crust community. mSystems 6, e00786-20 (2021).
Imminger, S. et al. Survival and rapid resuscitation permit limited productivity in desert microbial communities. Nat. Commun. 15, 1–17 (2024).
Shoemaker, W. R. & Lennon, J. T. Evolution with a seed bank: The population genetic consequences of microbial dormancy. Evol. Appl. 11, 60–75 (2018).
León-Sobrino, C., Ramond, J.-B., Maggs-Kölling, G. & Cowan, D. A. Nutrient acquisition, rather than stress response over diel cycles, drives microbial transcription in a hyper-arid Namib Desert soil. Front. Microbiol. 10, 1054 (2019).
Coclet, C., Cowan, D. & Lebre, P. H. Survival under Stress: Microbial Adaptation in Hot Desert Soils. in Microbiology of Hot Deserts (eds. Ramond, J.-B. & Cowan, D. A.) 293–317 (Springer International Publishing, 2022).
Metze, D. et al. Microbial growth under drought is confined to distinct taxa and modified by potential future climate conditions. Nat. Commun. 14, 5895 (2023).
Zhang, X. et al. Resistance of microbial community and its functional sensitivity in the rhizosphere hotspots to drought. Soil Biol. Biochem. 161, 108360 (2021).
Malik, A. A. & Bouskill, N. J. Drought impacts on microbial trait distribution and feedback to soil carbon cycling. Funct. Ecol. 36, 1442–1456 (2022).
Collins, S. L. et al. A multiscale, hierarchical model of pulse dynamics in arid-land ecosystems. Annu. Rev. Ecol. Evol. Syst. 45, 397–419 (2014).
Coleine, C. et al. Dryland microbiomes reveal community adaptations to desertification and climate change. ISME J. 18, wrae056 (2024).
Maestre, F. T. et al. Structure and functioning of dryland ecosystems in a changing world. Annu. Rev. Ecol. Evol. Syst. 47, 215–237 (2016).
Jiang, H. et al. Seasonal dynamics of soil microbiome in response to dry-wet alternation along the Jinsha River Dry-hot Valley. BMC Microbiol. 24, 496 (2024).
Naidoo, Y., Valverde, A., Pierneef, R. E. & Cowan, D. A. Differences in precipitation regime shape microbial community composition and functional potential in Namib Desert soils. Microb. Ecol. 83, 689–701 (2022).
Overpeck, J. T. & Udall, B. Climate change and the aridification of North America. Proc. Natl. Acad. Sci. USA 117, 11856–11858 (2020).
Burrell, A. L., Evans, J. P. & De Kauwe, M. G. Anthropogenic climate change has driven over 5 million km2 of drylands towards desertification. Nat. Commun. 11, 3853 (2020).
Adams, D. K. & Comrie, A. C. The North American monsoon. Bull. Am. Meteorol. Soc. 78, 2197–2214 (1997).
Li, C. et al. The adjustment of life history strategies drives the ecological adaptations of soil microbiota to aridity. Mol. Ecol. 31, 2920–2934 (2022).
Evans, S. E. & Wallenstein, M. D. Soil microbial community response to drying and rewetting stress: does historical precipitation regime matter? Biogeochemistry 109, 101–116 (2012).
Wallenstein, M. D. & Hall, E. K. A trait-based framework for predicting when and where microbial adaptation to climate change will affect ecosystem functioning. Biogeochemistry 109, 35–47 (2012).
Wu, K., Xu, W. & Yang, W. Effects of precipitation changes on soil bacterial community composition and diversity in the Junggar desert of Xinjiang, China. PeerJ 8, e8433 (2020).
Vásquez-Dean, J., Maza, F., Morel, I., Pulgar, R. & González, M. Microbial communities from arid environments on a global scale. A systematic review. Biol. Res. 53, 29 (2020).
Šťovíček, A., Kim, M., Or, D. & Gillor, O. Microbial community response to hydration-desiccation cycles in desert soil. Sci. Rep. 7, 45735 (2017).
Blazewicz, S. J., Schwartz, E. & Firestone, M. K. Growth and death of bacteria and fungi underlie rainfall-induced carbon dioxide pulses from seasonally dried soil. Ecology 95, 1162–1172 (2014).
Lindström, E. S. & Langenheder, S. Local and regional factors influencing bacterial community assembly: Bacterial community assembly. Environ. Microbiol. Rep. 4, 1–9 (2012).
Nemergut, D. R. et al. Patterns and processes of microbial community assembly. Microbiol. Mol. Biol. Rev. 77, 342–356 (2013).
Wilson, R. M. et al. Soil metabolome response to whole-ecosystem warming at the spruce and peatland responses under changing environments experiment. Proc. Natl. Acad. Sci. USA 118, e2004192118 (2021).
AminiTabrizi, R. et al. Controls on soil organic matter degradation and subsequent greenhouse gas emissions across a permafrost thaw gradient in Northern Sweden. Front. Earth Sci. 8, 557961 (2020).
Tfaily, M. M., Hess, N. J., Koyama, A. & Evans, R. D. Elevated [CO2] changes soil organic matter composition and substrate diversity in an arid ecosystem. Geoderma 330, 1–8 (2018).
Ramond, J.-B., Jordaan, K., Díez, B., Heinzelmann, S. M. & Cowan, D. A. Microbial biogeochemical cycling of nitrogen in arid ecosystems. Microbiol. Mol. Biol. Rev. 86, e00109–e00121 (2022).
Moyano, F. E. et al. The moisture response of soil heterotrophic respiration: interaction with soil properties. Biogeosciences 9, 1173–1182 (2012).
Yu, Z. et al. Molecular insights into the transformation of dissolved organic matter during hyperthermophilic composting using ESI FT-ICR MS. Bioresour. Technol. 292, 122007 (2019).
Li, H. et al. Responses of soil bacterial communities to nitrogen deposition and precipitation increment are closely linked with aboveground community variation. Microb. Ecol. 71, 974–989 (2016).
Chen, Q., Niu, B., Hu, Y., Luo, T. & Zhang, G. Warming and increased precipitation indirectly affect the composition and turnover of labile-fraction soil organic matter by directly affecting vegetation and microorganisms. Sci. Total Environ. 714, 136787 (2020).
Bowers, R. M. et al. Minimum information about a single amplified genome (MISAG) and a metagenome-assembled genome (MIMAG) of bacteria and archaea. Nat. Biotechnol. 35, 725–731 (2017).
Biggs, C. R. et al. Does functional redundancy affect ecological stability and resilience? A review and meta-analysis. Ecosphere 11, e03184 (2020).
Liu, L. et al. Microbial diversity and adaptive strategies in the Mars-like Qaidam Basin, North Tibetan Plateau, China. Environ. Microbiol. Rep. 14, 873–885 (2022).
Musto, H. et al. Correlations between genomic GC levels and optimal growth temperatures in prokaryotes. FEBS Lett. 573, 73–77 (2004).
Fierer, N. et al. Cross-biome metagenomic analyses of soil microbial communities and their functional attributes. Proc. Natl. Acad. Sci. USA 109, 21390–21395 (2012).
Chuckran, P. F. et al. Edaphic controls on genome size and GC content of bacteria in soil microbial communities. Soil Biol. Biochem. 178, 108935 (2023).
Giovannoni, S. J., Cameron Thrash, J. & Temperton, B. Implications of streamlining theory for microbial ecology. ISME J. 8, 1553–1565 (2014).
Liu, Q., Chen, Y. & Xu, X.-W. Genomic insight into strategy, interaction and evolution of nitrifiers in metabolizing key labile-dissolved organic nitrogen in different environmental niches. Front. Microbiol. 14, 1273211 (2023).
Bei, Q. et al. Metabolic potential of Nitrososphaera-associated clades. ISME J. 18, wrae086 (2024).
Van Etten, J., Stephens, T. G. & Bhattacharya, D. Genetic transfer in action: Uncovering DNA flow in an extremophilic microbial community. Environ. Microbiol. 27, e70048 (2025).
Ranawat, P. & Rawat, S. Stress response physiology of thermophiles. Arch. Microbiol. 199, 391–414 (2017).
Sun, D. et al. KatG and KatE confer Acinetobacter resistance to hydrogen peroxide but sensitize bacteria to killing by phagocytic respiratory burst. Life Sci. 148, 31–40 (2016).
Karas, V. O., Westerlaken, I. & Meyer, A. S. The DNA-binding protein from starved cells (DPs) utilizes dual functions to defend cells against multiple stresses. J. Bacteriol. 197, 3206–3215 (2015).
Ahn, S., Jung, J., Jang, I.-A., Madsen, E. L. & Park, W. Role of glyoxylate shunt in oxidative stress response. J. Biol. Chem. 291, 11928–11938 (2016).
Kuzminov, A. Recombinational repair of DNA damage in Escherichia coli and Bacteriophage λ. Microbiol. Mol. Biol. Rev. 63, 751–813 (1999).
Zahradka, K. et al. Reassembly of shattered chromosomes in Deinococcus radiodurans. Nature 443, 569–573 (2006).
Leem, S. H., Ropp, P. A. & Sugino, A. The yeast Saccharomyces cerevisiae DNA polymerase IV: possible involvement in double strand break DNA repair. Nucleic Acids Res. 22, 3011–3017 (1994).
Stegen, J. C., Lin, X., Fredrickson, J. K. & Konopka, A. E. Estimating and mapping ecological processes influencing microbial community assembly. Front. Microbiol. 6, 370 (2015).
Stegen, J. C., Lin, X., Konopka, A. E. & Fredrickson, J. K. Stochastic and deterministic assembly processes in subsurface microbial communities. ISME J. 6, 1653–1664 (2012).
Danczak, R. E. et al. Using metacommunity ecology to understand environmental metabolomes. Nat. Commun. 11, 6369 (2020).
Freire-Zapata, V. et al. Microbiome-metabolite linkages drive greenhouse gas dynamics over a permafrost thaw gradient. Nat. Microbiol. 1, 17 (2024).
Wang, X.-B. et al. A drying-rewetting cycle imposes more important shifts on soil microbial communities than does reduced precipitation. mSystems 7, e0024722 (2022).
Plaza, C., Gascó, G., Méndez, A. M., Zaccone, C. & Maestre, F. T. Soil organic matter in dryland ecosystems. in The Future of Soil Carbon 39–70 (Elsevier, 2018).
Kim, M. & Or, D. Individual-based model of microbial life on hydrated rough soil surfaces. PLoS ONE 11, e0147394 (2016).
Warren, C. R. Response of osmolytes in soil to drying and rewetting. Soil Biol. Biochem. 70, 22–32 (2014).
Halverson, L. J., Jones, T. M. & Firestone, M. K. Release of intracellular solutes by four soil bacteria exposed to dilution stress. Soil Sci. Soc. Am. J. 64, 1630–1637 (2000).
Taravati, A. et al. Various effects of sugar and polyols on the protein structure and function: role as osmolyte on protein stability. World Appl. Sci. J. 2, 353–362 (2007).
Couvillion, S. P. et al. Rapid remodeling of the soil lipidome in response to a drying-rewetting event. Microbiome 11, 34 (2023).
Lal, R. Carbon sequestration in dryland ecosystems. Environ. Manag. 33, 528–544 (2004).
Deng, Y. et al. Molecular ecological network analyses. BMC Bioinform. 13, 113 (2012).
Wang, X. et al. Decreased soil multifunctionality is associated with altered microbial network properties under precipitation reduction in a semiarid grassland. Imeta 2, e106 (2023).
Wang, S., Wang, X., Han, X. & Deng, Y. Higher precipitation strengthens the microbial interactions in semi-arid grassland soils. Glob. Ecol. Biogeogr. 27, 570–580 (2018).
Zelezniak, A. et al. Metabolic dependencies drive species co-occurrence in diverse microbial communities. Proc. Natl. Acad. Sci. USA 112, 6449–6454 (2015).
de Vries, F. T. et al. Soil bacterial networks are less stable under drought than fungal networks. Nat. Commun. 9, 3033 (2018).
Gao, C. et al. Co-occurrence networks reveal more complexity than community composition in resistance and resilience of microbial communities. Nat. Commun. 13, 3867 (2022).
Evans, S. E. & Wallenstein, M. D. Climate change alters ecological strategies of soil bacteria. Ecol. Lett. 17, 155–164 (2014).
Leung, P. M. et al. Energetic basis of microbial growth and persistence in desert ecosystems. mSystems 5, e00495-19 (2020).
Hu, Y. et al. Seasonal patterns of soil microbial community response to warming and increased precipitation in a semiarid steppe. Appl. Soil Ecol. 182, 104712 (2023).
Liu, D. et al. Response of microbial communities and their metabolic functions to Drying−Rewetting stress in a temperate forest soil. Microorganisms 7, 129 (2019).
Nielsen, U. N. & Ball, B. A. Impacts of altered precipitation regimes on soil communities and biogeochemistry in arid and semi-arid ecosystems. Glob. Change Biol. 21, 1407–1421 (2015).
Armstrong, A. et al. Temporal dynamics of hot desert microbial communities reveal structural and functional responses to water input. Sci. Rep. 6, 34434 (2016).
Marasco, R. et al. Rhizosheath microbial community assembly of sympatric desert speargrasses is independent of the plant host. Microbiome 6, 215 (2018).
Mandakovic, D. et al. Structure and co-occurrence patterns in microbial communities under acute environmental stress reveal ecological factors fostering resilience. Sci. Rep. 8, 5875 (2018).
Mosqueira, M. J. et al. Consistent bacterial selection by date palm root system across heterogeneous desert oasis agroecosystems. Sci. Rep. 9, 4033 (2019).
Ramond, J.-B. & Cowan, D. A. Microbial ecology of hot desert soils. in Ecological Studies 89–110 (Springer International Publishing, 2022).
Kpalari, D. F. et al. Soil bacterial community and greenhouse gas emissions as responded to the coupled application of nitrogen fertilizer and microbial decomposing inoculants in wheat (Triticum aestivum L.) seedling stage under different water regimes. Agronomy 13, 2950 (2023).
Yang, Q. et al. Erosion and deposition significantly affect the microbial diversity, co-occurrence network, and multifunctionality in agricultural soils of Northeast China. J. Soils Sediments 24, 888–900 (2023).
He, J. et al. Distinct composition patterns of bacterial and fungal communities and biogeochemical cycling genes depend on the vegetation type in arid soil. Appl. Soil Ecol. 191, 105064 (2023).
Li, J., Liu, T., McIlroy, S. J., Tyson, G. W. & Guo, J. Phylogenetic and metabolic diversity of microbial communities performing anaerobic ammonium and methane oxidations under different nitrogen loadings. ISME Commun. 3, 39 (2023).
Mee, M. T., Collins, J. J., Church, G. M. & Wang, H. H. Syntrophic exchange in synthetic microbial communities. Proc. Natl. Acad. Sci. USA 111, E2149–E2156 (2014).
Shen, J.-P., Xu, Z. & He, J.-Z. Frontiers in the microbial processes of ammonia oxidation in soils and sediments. J. Soils Sediments 14, 1023–1029 (2014).
León-Sobrino, C. et al. Temporal dynamics of microbial transcription in wetted hyperarid desert soils. FEMS Microbiol. Ecol. 100, fiae009 (2024).
Lu, L. & Jia, Z. Urease gene-containing Archaea dominate autotrophic ammonia oxidation in two acid soils: urea-linked archaeal ammonia oxidation in acid soil. Environ. Microbiol. 15, 1795–1809 (2013).
Hatzenpichler, R. Diversity, physiology, and niche differentiation of ammonia-oxidizing archaea. Appl. Environ. Microbiol. 78, 7501–7510 (2012).
Zhao, R., Wang, H., Yang, H., Yun, Y. & Barton, H. A. Ammonia-oxidizing Archaea dominate ammonia-oxidizing communities within alkaline cave sediments. Geomicrobiol. J. 34, 511–523 (2017).
Neilson, J. W. et al. Significant impacts of increasing aridity on the arid soil microbiome. mSystems 2, e00195-16 (2017).
Yuan, M. M. et al. Climate warming enhances microbial network complexity and stability. Nat. Clim. Change 11, 343–348 (2021).
Maestre, F. T. et al. Research needs on the biodiversity-ecosystem functioning relationship in drylands. npj Biodivers. 3, 12 (2024).
Gao, S. et al. Effects of temperature increase and nitrogen addition on the early litter decomposition in permafrost peatlands. Catena 209, 105801 (2022).
Bertness, M. D. & Callaway, R. Positive interactions in communities. Trends Ecol. Evol. 9, 191–193 (1994).
Hammarlund, S. P. & Harcombe, W. R. Refining the stress gradient hypothesis in a microbial community. Proc. Natl. Acad. Sci. USA 116, 15760–15762 (2019).
Hernandez, D. J., David, A. S., Menges, E. S., Searcy, C. A. & Afkhami, M. E. Environmental stress destabilizes microbial networks. ISME J. 15, 1722–1734 (2021).
Goberna, M. et al. Incorporating phylogenetic metrics to microbial co-occurrence networks based on amplicon sequences to discern community assembly processes. Mol. Ecol. Resour. 19, 1552–1564 (2019).
Jia, M. et al. Effects of precipitation change and nitrogen addition on the composition, diversity, and molecular ecological network of soil bacterial communities in a desert steppe. PLoS ONE 16, e0248194 (2021).
Xu, L. et al. Genome-resolved metagenomics reveals role of iron metabolism in drought-induced rhizosphere microbiome dynamics. Nat. Commun. 12, 3209 (2021).
Dong, X. et al. Metagenomic views of microbial communities in sand sediments associated with coral reefs. Microb. Ecol. 85, 465–477 (2023).
Honeker, L. K. et al. Elucidating drought-tolerance mechanisms in plant roots through 1H NMR metabolomics in parallel with MALDI-MS, and NanoSIMS imaging techniques. Environ. Sci. Technol. 56, 2021–2032 (2022).
Shivlata, L. & Satyanarayana, T. Thermophilic and alkaliphilic Actinobacteria: biology and potential applications. Front. Microbiol. 6, 1014 (2015).
Hamedi, J., Poorinmohammad, N. & Papiran, R. Growth and life cycle of actinobacteria. in Biology and Biotechnology of Actinobacteria (eds. Wink, J., Mohammadipanah, F. & Hamedi, J.) 29–50 (Springer International Publishing, 2017).
Bouskill, N. J. et al. Belowground response to drought in a tropical forest soil. I. changes in microbial functional potential and metabolism. Front. Microbiol. 7, 525 (2016).
Ramin, K. I. & Allison, S. D. Bacterial tradeoffs in growth rate and extracellular enzymes. Front. Microbiol. 10, 2956 (2019).
Pointing, S. B. & Belnap, J. Microbial colonization and controls in dryland systems. Nat. Rev. Microbiol. 10, 551–562 (2012).
Veach, A. M. & Zeglin, L. H. Historical drought affects microbial population dynamics and activity during soil drying and re-wet. Microb. Ecol. 79, 662–674 (2020).
Ouyang, Y. & Li, X. Effect of repeated drying-rewetting cycles on soil extracellular enzyme activities and microbial community composition in arid and semi-arid ecosystems. Eur. J. Soil Biol. 98, 103187 (2020).
Weiss, J. L. & Overpeck, J. T. Is the Sonoran desert losing its cool? Glob. Chang. Biol. 11, 2065–2077 (2005).
Biederman, J. A. et al. Shrubland carbon sink depends upon winter water availability in the warm deserts of North America. Agric. For. Meteorol. 249, 407–419 (2018).
Douglas, M., Maddox, R., Howard, K. & Reyes, S. The Mexican monsoon. J. Clim. 6, 1665–1677 (1993).
AminiTabrizi, R., Dontsova, K., Graf Grachet, N. & Tfaily, M. M. Elevated temperatures drive abiotic and biotic degradation of organic matter in a peat bog under oxic conditions. Sci. Total Environ. 804, 150045 (2022).
Didan, K. MODIS/Terra vegetation indices 16-day L3 global 250m SIN Grid V061 [Data set]. https://doi.org/10.5067/MODIS/MOD13Q1.061 (2021).
Lüdecke, D., Ben-Shachar, M., Patil, I., Waggoner, P. & Makowski, D. Performance: an R package for assessment, comparison and testing of statistical models. J. Open Source Softw. 6, 3139 (2021).
Dittmar, T., Koch, B., Hertkorn, N. & Kattner, G. A simple and efficient method for the solid-phase extraction of dissolved organic matter (SPE-DOM) from seawater: SPE-DOM from seawater. Limnol. Oceanogr. Methods 6, 230–235 (2008).
Tolić, N. et al. Formularity: Software for automated formula assignment of natural and other organic matter from ultrahigh-resolution mass spectra. Anal. Chem. 89, 12659–12665 (2017).
Ayala-Ortiz, C. et al. MetaboDirect: an analytical pipeline for the processing of FT-ICR MS-based metabolomic data. Microbiome 11, 28 (2023).
Klindworth, A. et al. Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res. 41, e1 (2013).
Bay, S. K. et al. Chemosynthetic and photosynthetic bacteria contribute differentially to primary production across a steep desert aridity gradient. ISME J. 15, 3339–3356 (2021).
Lian, W.-H. et al. Culturomics- and metagenomics-based insights into the microbial community and function of rhizosphere soils in Sinai desert farming systems. Environ. Microbiome 18, 4 (2023).
Zhao, M. et al. Asymmetric responses of abundance and diversity of N-cycling genes to altered precipitation in arid grasslands. Funct. Ecol. 37, 2953–2966 (2023).
Nguyen, T. M. et al. Whole community shotgun metagenomes of two biological soil crust types from the Mojave Desert. Microbiol. Resour. Announc. 13, e0098023 (2024).
Barrón-Sandoval, A. et al. Functional significance of microbial diversity in arid soils: biological soil crusts and nitrogen fixation as a model system. FEMS Microbiol. Ecol. 99, fiad009 (2023).
Chen, Y., Neilson, J. W., Kushwaha, P., Maier, R. M. & Barberán, A. Life-history strategies of soil microbial communities in an arid ecosystem. ISME J. 15, 649–657 (2021).
Ondov, B. D. et al. Mash: fast genome and metagenome distance estimation using MinHash. Genome Biol. 17, 132 (2016).
Woodcroft, B. J. et al. Comprehensive taxonomic identification of microbial species in metagenomic data using SingleM and Sandpiper. Nat. Biotechnol. https://doi.org/10.1038/s41587-025-02738-1 (2025).
Callahan, B. J. et al. DADA2: high-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 17, 10 (2011).
Wang, Q., Garrity, G. M., Tiedje, J. M. & Cole, J. R. Naïve Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl. Environ. Microbiol. 73, 5261–5267 (2007).
Parks, D. H. et al. GTDB: an ongoing census of bacterial and archaeal diversity through a phylogenetically consistent, rank normalized and complete genome-based taxonomy. Nucleic Acids Res. 50, D785–D794 (2022).
Větrovský, T. & Baldrian, P. The variability of the 16S rRNA gene in bacterial genomes and its consequences for bacterial community analyses. PLoS ONE 8, e57923 (2013).
Lan, Y., Rosen, G. & Hershberg, R. Marker genes that are less conserved in their sequences are useful for predicting genome-wide similarity levels between closely related prokaryotic strains. Microbiome 4, 18 (2016).
R Core Team. R: A Language and Environment for Statistical Computing. (R Foundation for Statistical Computing, Vienna, Austria, 2023).
McMurdie, P. J. & Holmes, S. phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 8, e61217 (2013).
Oksanen, J. et al. Vegan: Community Ecology Package. https://github.com/vegandevs/vegan (2024).
Wright, E. Using DECIPHER v2.0 to analyze big biological sequence data in R. R J. 8, 352 (2016).
Price, M. N., Dehal, P. S. & Arkin, A. P. FastTree 2–approximately maximum-likelihood trees for large alignments. PLoS ONE 5, e9490 (2010).
Revell, L. J. phytools 2.0: an updated R ecosystem for phylogenetic comparative methods (and other things). PeerJ 12, e16505 (2024).
Andrews, S. Babraham Bioinformatics - FastQC a quality control tool for high throughput sequence data. FastQC https://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (2010).
Bushnell, B. BBMap. SourceForge https://sourceforge.net/projects/bbmap/ (2021).
Li, D. et al. MEGAHIT v1.0: A fast and scalable metagenome assembler driven by advanced methodologies and community practices. Methods 102, 3–11 (2016).
Aroney, S. T. N. et al. CoverM: read coverage calculator for metagenomics. https://doi.org/10.5281/zenodo.10531253 (2024).
Alneberg, J. et al. Binning metagenomic contigs by coverage and composition. Nat. Methods 11, 1144–1146 (2014).
Kang, D. D. et al. MetaBAT 2: an adaptive binning algorithm for robust and efficient genome reconstruction from metagenome assemblies. PeerJ 7, e7359 (2019).
Wu, Y.-W., Simmons, B. A. & Singer, S. W. MaxBin 2.0: an automated binning algorithm to recover genomes from multiple metagenomic datasets. Bioinformatics 32, 605–607 (2016).
Uritskiy, G. V., DiRuggiero, J. & Taylor, J. MetaWRAP-a flexible pipeline for genome-resolved metagenomic data analysis. Microbiome 6, 158 (2018).
Eren, A. M. et al. Anvi’o: an advanced analysis and visualization platform for ‘omics data. PeerJ 3, e1319 (2015).
Chklovski, A., Parks, D. H., Woodcroft, B. J. & Tyson, G. W. CheckM2: a rapid, scalable and accurate tool for assessing microbial genome quality using machine learning. Nat. Methods 20, 1203–1212 (2023).
Jain, C., Rodriguez-R, L. M., Phillippy, A. M., Konstantinidis, K. T. & Aluru, S. High throughput ANI analysis of 90K prokaryotic genomes reveals clear species boundaries. Nat. Commun. 9, 5114 (2018).
Woodcroft, B. J. Galah: More Scalable Dereplication for Metagenome Assembled Genomes (Github, 2024).
Chaumeil, P.-A., Mussig, A. J., Hugenholtz, P. & Parks, D. H. GTDB-Tk v2: memory friendly classification with the genome taxonomy database. Bioinformatics 38, 5315–5316 (2022).
Yu, G., Smith, D. K., Zhu, H., Guan, Y. & Lam, T. T.-Y. Ggtree: an R package for visualization and annotation of phylogenetic trees with their covariates and other associated data. Methods Ecol. Evol. 8, 28–36 (2017).
Xu, S. et al. GgtreeExtra: compact visualization of richly annotated phylogenetic data. Mol. Biol. Evol. 38, 4039–4042 (2021).
Shaffer, M. et al. DRAM for distilling microbial metabolism to automate the curation of microbiome function. Nucleic Acids Res. 48, 8883–8900 (2020).
Tu, Q., Lin, L., Cheng, L., Deng, Y. & He, Z. NCycDB: a curated integrative database for fast and accurate metagenomic profiling of nitrogen cycling genes. Bioinformatics 35, 1040–1048 (2019).
Yu, X. et al. SCycDB: a curated functional gene database for metagenomic profiling of sulphur cycling pathways. Mol. Ecol. Resour. 21, 924–940 (2021).
Zeng, J. et al. PCycDB: a comprehensive and accurate database for fast analysis of phosphorus cycling genes. Microbiome 10, 101 (2022).
Kanehisa, M. The KEGG database. Novartis Found. Symp. 247, 91–101 (2002).
Karaoz, U. & Brodie, E. L. MicroTrait: a toolset for a trait-based representation of microbial genomes. Front. Bioinform. 2, 918853 (2022).
Weissman, J. L., Hou, S. & Fuhrman, J. A. Estimating maximal microbial growth rates from cultures, metagenomes, and single cells via codon usage patterns. Proc. Natl. Acad. Sci. USA 118, e2016810118 (2021).
Kopylova, E., Noé, L. & Touzet, H. SortMeRNA: fast and accurate filtering of ribosomal RNAs in metatranscriptomic data. Bioinformatics 28, 3211–3217 (2012).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, H. et al. The sequence alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Smid, M. et al. Gene length corrected trimmed mean of M-values (GeTMM) processing of RNA-seq data performs similarly in intersample analyses while improving intrasample comparisons. BMC Bioinform. 19, 236 (2018).
Simpson, G. L. Analogue methods in palaeoecology: using the analogue Package. J. Stat. Softw. 22, 1–29 (2007).
Jombart, T., Balloux, F. & Dray, S. Adephylo: new tools for investigating the phylogenetic signal in biological traits. Bioinformatics 26, 1907–1909 (2010).
Tfaily, M. M. et al. Sequential extraction protocol for organic matter from soils and sediments using high resolution mass spectrometry. Anal. Chim. Acta 972, 54–61 (2017).
Chase, J. M. Drought mediates the importance of stochastic community assembly. Proc. Natl. Acad. Sci. USA 104, 17430–17434 (2007).
Zhou, J. et al. Functional molecular ecological networks. MBio 1, 10–1128 (2010).
Newman, M. E. J. Modularity and community structure in networks. Proc. Natl. Acad. Sci. USA 103, 8577–8582 (2006).
Csardi, G. & Nepusz, T. The igraph software package for complex network research. Complex Syst. 1695, 1–9 (2005).
Briatte, F. ggnetwork: geometries to plot networks with ‘ggplot2’. https://CRAN.R-project.org/package=ggnetwork (2024).
Machado, D., Andrejev, S., Tramontano, M. & Patil, K. R. Fast automated reconstruction of genome-scale metabolic models for microbial species and communities. Nucleic Acids Res. 46, 7542–7553 (2018).
Giordano, N. et al. Genome-scale community modelling reveals conserved metabolic cross-feedings in epipelagic bacterioplankton communities. Nat. Commun. 15, 2721 (2024).
Du, H. et al. Microbial active functional modules derived from network analysis and metabolic interactions decipher the complex microbiome assembly in mangrove sediments. Microbiome 10, 224 (2022).
Acknowledgements
This study was supported in part by the University of Arizona (UA) UA Research, Innovation and Impact (RII) Core Facilities Pilot Program and UA Start-Up funds awarded to PI M. Tfaily. We also acknowledge future support from the U.S. National Science Foundation under Award No. 2519761, which will fund the continuation of this work starting November 2025 (NSF IOS, Organismal Response to Climate Change, NSF 25-504). We thank the Environmental Molecular Sciences Laboratory (EMSL), a DOE Office of Science User Facility sponsored by the Office of Biological and Environmental Research and operated under Contract DE-AC05-76RL01830 and more specifically Rosalie Chu for their help in generating the FTICR-MS data. This includes EMSL project 60352 awarded to PI M. Tfaily. We thank the Arizona Genetics Core, University of Arizona, Tucson, AZ (Facility RRID:SCR_012429, https://azgc.arizona.edu/) for their help in generating all the next generation sequencing data. We thank the ALEC, University of Arizona, Tucson, AZ, for their help with the analysis of total carbon, total nitrogen, and metallic ions. Bioinformatics analysis in this publication used High Performance Computing (HPC) resources supported by the University of Arizona TRIF, UITS, and Research, Innovation, and Impact (RII) and maintained by the UArizona Research Technologies department. Additionally, this work used Bridges-2 at Pittsburgh Supercomputing Center through allocation BIO220147 from the Advanced Cyberinfrastructure Coordination Ecosystem: Services & Support (ACCESS) program, which is supported by U.S. National Science Foundation grants #2138259, #2138286, #2138307, #2137603, and #2138296. Soil samples were collected under permit SAGU-2021-SCI-0006 issued by the U.S. National Park Service for research conducted in Saguaro National Park West, Tucson, Arizona (32°15′05.4″N, 111°09′40.2″W). We thank the National Park Service and Saguaro National Park staff for their support and access to field sites.
Author information
Authors and Affiliations
Contributions
C.A.-O., V.F.-Z., and M.M.T. conceptualized, designed and supervised the study. C.A.-O., V.F.-Z,. and M.M.T. coordinated sampling efforts. C.A.-O. and V.F.-Z. collected samples. C.A.-O. and V.F.-Z. processed the samples. C.A.-O. and V.F.-Z. analyzed and visualized the data. C.A.-O., V.F.-Z., and M.M.T. wrote the initial draft of the manuscript. All authors contributed to manuscript revisions and approved the final version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Earth & Environment thanks Ziting Wang and Malte Herold for their contribution to the peer review of this work. Primary handling editors: Kate Buckeridge and Somaparna Ghosh. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ayala-Ortiz, C., Freire-Zapata, V. & Tfaily, M.M. Stochastic assembly and metabolic network reorganization drive microbial resilience in arid soils. Commun Earth Environ 6, 647 (2025). https://doi.org/10.1038/s43247-025-02637-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s43247-025-02637-y