Abstract
Antibiotic tolerance, the ability to survive lethal antibiotics for a prolonged period of time is a rising threat due to its role as a steppingstone towards antibiotic resistance. While tolerance has been recognized as a severe clinical threat, little is known about the mechanisms promoting tolerance. Here, we delineated the physiology of antibiotic tolerance to the classic β-lactam antibiotic, penicillin, to discover the metabolic underpinnings of how tolerant bacteria survive ordinarily lethal antibiotic exposure. We used transcriptomics and metabolomics in the hypertolerant Gram-negative cholera pathogen, Vibrio cholerae, to identify the global regulatory and metabolic response to antibiotic exposure. Key pathways like central carbon metabolism, cell wall synthesis, heat shock, two-component systems, and particularly nucleotide synthesis were significantly altered in response to penicillin. Most notably, nucleotide levels were depleted upon antibiotic exposure, concomitant with upregulation of both purine and pyrimidine synthesis functions. Consistent with a crucial role for nucleotide synthesis in tolerance, we find that targeting nucleotide synthesis synergizes with β-lactam-mediated killing in hypertolerant V. cholerae, Klebsiella pneumoniae and E. coli. These datasets thus reveal new vulnerabilities in tolerant bacteria that can serve as conceptual scaffolds for drug development and for improving antibiotic efficacy.
Similar content being viewed by others
Introduction
Antibiotic resistance has long been recognized as a severe public health threat1. However, antibiotic susceptibility is a spectrum, with multiple steps between full susceptibility and outright resistance. One such stepping-stone towards resistance is antibiotic tolerance, or the ability to survive high doses of a typically bactericidal antibiotic for prolonged periods of time2,3,4,5,6,7. Tolerance has been observed in response to diverse antibiotics, including the β-lactams. β-lactam antibiotics are among our most widely prescribed antibiotics, and rates of antibiotic resistance to these crucial drugs are at an all-time high. What makes this class of antibiotics so effective lies in their target: β-lactams inhibit the penicillin-binding proteins (PBPs), the main enzymes that synthesize the bacterial cell wall (peptidoglycan (PG) layer). This vital element to the bacterial cell envelope regulates cell shape, turgor pressure, and it is unique to prokaryotes, limiting β-lactam off-target toxicity when used in humans. Bacterial cells that are tolerant to these antibiotics effectively shed their peptidoglycan, forming cell wall-deficient spheroplasts, thereby bypassing cell death; this has been observed in diverse and significant clinical pathogens, including Enterobacter cloacae, Klebsiella pneumoniae, Pseudomonas aeruginosa, Haemophilus influenzae, Acinetobacter baumannii, Vibrio cholerae, and in diverse growth conditions, including standard laboratory media, human serum medium, minimal media, and human patients8,9,10,11,12,13. After the antibiotic is removed from the environment, tolerant cells can revert into their normal rod shape and continue to cause an infection. Data from clinical isolates suggest that this phenotype is pervasive in the clinical setting10, and may underlie treatment failure and development of antibiotic resistance5. Spheroplasts are metabolically active, but do not replicate in the presence of an antibiotic. Instead, they rely on stress responses to mitigate the many perturbations associated with cell wall loss10,14,15,16. Stress responses are critical for surviving the perturbations associated with antibiotic exposure, which often causes pleiotropic cellular damage. For example, β-lactam antibiotics kill not only via their primary target (inhibiting cell wall synthesis), but also through increasing metabolic burden via a process termed “futile cycling”17,18. Futile cycling is the constant generation of PG strands that are immediately broken down by endogenous cell wall lytic enzymes called autolysins. In E. coli, this results in poorly-understood perturbances in redox state and enhanced protein synthesis17. How tolerant bacteria withstand this metabolic reprogramming is unknown.
Here, we incorporate transcriptomic and metabolomic data to characterize the global antibiotic tolerance response to penicillin in the Gram-negative pathogen Vibrio cholerae. Vibrio cholerae has emerged as a model pathogen for tolerance research due to its high levels of β-lactam tolerance coupled with the ease of genetic manipulation. Our data suggest that central carbon metabolism is rerouted towards cell wall synthesis (depletion of upper glycolysis intermediates), supporting the notion of ongoing futile cycling, which ultimately results in the depletion of nucleotide precursors (likely via reduction of upper glycolysis shuttling into the pentose phosphate pathway). Consistent with an important role for nucleotide homeostasis in β-lactam tolerance, we show that perturbations of nucleotide homeostasis potentiate PenG-mediated killing. Overall, our data demonstrates the complex metabolic perturbances that tolerant cells must manage, showcasing new potential vulnerabilities that could be exploited as targets for new antimicrobial adjuvants.
Results
Vibrio cholerae’s global response to penicillin exposure
Using the causative agent of cholera disease, Vibrio cholerae, as a model system for investigating tolerance to cell wall-active antibiotics, we first probed global cellular transcript abundance. We treated wild-type N16961 cells with 100 μg/mL PenG (10x MIC)19, and removed samples at 0, 10-, 20-, 40-, 60-, and 90-minutes post treatment. This timeframe captures the full process of spheroplast formation as determined previously19. The utility of our tolerance model is that here, unlike most previous studies (in E. coli), RNAseq was conducted on intact cells treated extensively with high, supra-MIC concentrations of the antibiotic, whereas in non-tolerant bacteria, sub-MIC concentrations or shorter timeframes typically have to be used (Fig. 1A–D). Both GO term and KEGG pathway enrichment revealed differential regulation of cell wall functions, central carbon metabolism, nucleotide homeostasis and cell signaling (Fig. S1), prompting us to focus on these key areas of cellular physiology.
A Cell wall synthesis genes. B Tricarboxylic acid cycle. C Cell signaling pathways. D Heat shock response. See Methods for details. Data presented are genes that were scored as significantly differentially regulated with a cut-off p-value < 0.05. Numbers within the heat map represent raw log2 fold-change.
Genes associated with cell wall biosynthesis were upregulated, with only a few genes repressed (Fig. 1A). This was expected, as penicillin treatment was previously shown to upregulate genes involved in maintaining the cell wall, via activation of the VxrAB cell envelope stress response two-component system14. The nagK (vc1532) gene was one of the few repressed genes, with a marked, 25-fold decrease in expression. NagK is an N-acetylglucosamine (GlcNAc) kinase that converts internal GlcNAc to GlcNAc-6P. Internal GlcNAc is most likely the product of peptidoglycan recycling, for V. cholerae has no other means of synthesizing unphosphorylated GlcNAc20,21. Genes involved in early cell envelope precursor synthesis also exhibited a (albeit less pronounced) reduction in gene expression, such as the uppS (vc2256) and murI (vc0158) genes. UppS is the important undecaprenyl diphosphate synthase whose role is to build the undecaprenyl tether that allows the fundamental PG building block lipid II to traverse the inner membrane into the periplasmic space. MurI functions as a glutamate racemase, converting L-Glu to D-Glu to be incorporated into the growing amino acid side stem of the lipid II molecule22. The downregulation of these genes may reflect a fitness trade-off associated with the generation of their products.
Most genes involved in peptidoglycan synthesis consistently exhibited an increase in gene expression. Genes like murA, murC, murD, murG, and murJ, which are all involved in synthesizing the lipid II molecule and flipping it across the inner membrane exhibited a 4- to >20-fold increase in expression. This suggests that penicillin exposure, and its associated cell wall-focused damage, triggers the cell to build more PG in order to repair this damage, and/or to prepare for recovery when the antibiotic is removed. The gene coding for aPBP1B, one of the major bifunctional cell wall synthases, mrcB, was also highly upregulated. This result acts as an internal control, since we have previously shown that PBP1b is induced by β-lactam exposure through the two-component system VxrAB16. This makes physiological sense, as aPBPs are the primary PG synthases involved in cell wall damage repair23.
VxrAB is a well-characterized system for detecting cell-wall stress and subsequent β-lactam tolerance in V. cholerae. It is induced by diverse cell wall damage conditions (including β-lactam exposure), and its deletion increases β-lactam susceptibility and drastically reduces tolerance16. This two-component system is localized to the inner membrane, and upon activation by cell wall damage through an unknown signal, the histidine kinase VxrA phosphorylates VxrB to elicit downstream regulatory effects (Fig. S2A). We previously showed that the vxrAB locus itself is induced by PenG, and sought to establish its PenG-dependent regulon14. Conducting the same RNAseq protocol, we thus explored how transcripts changed in a ΔvxrAB mutant background. We then compared ΔvxrAB regulon transcripts to the levels observed in WT strains and found that nearly all VxrAB regulated genes (as determined by our previous ChipSeq experiments14) were differentially regulated by PenG (Fig. S2B). Thus, the VxrAB regulon is activated by PenG exposure.
Central metabolism gene expression changes in response to PenG exposure
Previous studies in other bacteria suggested that TCA cycle genes are induced as a consequence of exposure to β-lactam antibiotics24. Our analysis here revealed that genes involved in central carbon metabolism and the TCA cycle experienced minor dynamic flux over time (Fig. 1B). Some genes, like vc1304 and vc1141, (coding for fumarate hydratase and isocitrate dehydrogenase respectively), were slowly reduced in expression, reaching a 2.8- fold reduction by 90 min of exposure. In contrast, genes coding for the fumarate reductase complex (frdABCD), employed during anaerobic respiration, were modestly (2-fold) increased at 90 minutes. Genes coding for earlier steps in the TCA cycle also displayed an increase in expression, such as vc2084 and vc2085 (~4-fold change increase in early PenG exposure), coding for the succinyl-CoA synthetase complex; genes vc2086 and vc2087, creating 2-oxoglutarate dehydrogenase; and genes sdhCDAB (vc2088-vc2091), which make up the succinate dehydrogenase complex. The upregulation of late-TCA cycle enzymes could indicate a demand for more energy and/or reducing equivalents. The TCA cycle and subsequent respiration creates the most ATP molecules compared to other metabolic pathways, and it seems plausible that the cell attempts to optimize ATP generation given that β-lactam tolerance relies heavily on energy-intensive damage repair pathways14,16. These results could also indicate a resource allocation requirement, in which TCA cycle boosting is needed for the biosynthesis of other metabolites, perhaps amino acids or nucleotide production.
We next focused on cell signaling pathways, including two-component systems (TCS), which are the primary cell signaling means of bacteria. Cell signaling pathways consist of kinases, which respond to diverse signals, and (sometimes through intermediary proteins) associated response-regulators, which activate specific regulons. The kinases are often autoregulated, and activated by antibiotic stress conditions, and we thus focused specifically on V. cholerae’s known TCSs to uncover novel antibiotic-responsive signaling systems (Fig. 1C). Interestingly, the response regulator vc1638, encoding a homolog of PhoP, was induced nearly 16-fold (even higher than the primary cell wall damage response VxrAB). The PhoPQ TCS has been well-characterized in Enterobacterales, where it is induced by antimicrobial peptides, acidic conditions, and magnesium limitation25. We recently found that it is also activated by β-lactams and promotes carbapenem tolerance in Enterobacter cloacae, likely via OM stabilization upon PhoPQ-controlled L-amino-arabinose addition9. Its role in V. cholerae is currently poorly characterized, but the system has been implicated in polymyxin resistance via outer membrane modifications26, alluding to a role similar to PhoPQ in Enterobacterales. The other notable change in two-component systems was a reduction in citAB expression level. This system regulates anaerobic citrate utilization in V. cholerae27 and was strongly downregulated in the presence of PenG. CitAB has been shown to integrate multiple signals, including anaerobiosis, presence of citrate, and glucose levels (through catabolite repression)27, and future work will focus on the role of this two-component system in the response to PenG.
Consistent with historical data28,29,30, we also found many heat shock response31 genes to be significantly upregulated in response to penicillin treatment (Fig. 1D). We previously showed that PenG exposure causes accumulation of oxidized proteins in V. cholerae14, and we propose that heat shock is activated by these likely misfolded proteins.
Nucleotide synthesis pathways are upregulated upon PenG exposure
We noticed a large number of genes related to nucleotide biosynthesis in our transcriptomics dataset. Most of these genes were upregulated in response to penicillin treatment, particularly at early time points (Fig. 2A–C). Both purine and pyrimidine pathways were induced, with the majority of de novo synthesis genes being upregulated in response to PenG. This included the purA-T genes (de novo purine biosynthesis), as well as pyrBEFGH (de novo pyrimidine synthesis) and carAB, whose products convert L-glutamine to carbamoyl phosphate on the way towards pyrimidine biosynthesis32,33 (Fig. 2A, B). In contrast to these largely upregulated synthesis pathways, numerous genes coding for proteins involved in nucleotide homeostasis were markedly downregulated. These included nucleotide salvage pathways like deoD–gsk (coding for purine nucleotide phosphorylase and inosine kinase respectively); cdd, coding for cytidine deaminase; udp, coding for uridine phosphorylase; and UDP sugar hydrolases like ushA (vc2174). Thus, PenG exposure resulted in upregulation of nucleotide synthesis functions, and downregulation of nucleotide “recycling”, perhaps reflecting that a physiological need for preserving nucleotides has been sensed.
A Pyrimidine biosynthesis pathway. B Purine biosynthesis pathway. C genes involved in both nucleotide biosynthetic pathways. See Methods for details. Data presented are all genes that were scored as significantly differentially regulated, with a cut-off p-value < 0.05. Numbers within the heat map represent raw log2 fold-change.
Interestingly, the stringent response gene relA was also upregulated. The RelA enzyme synthesizes (p)ppGpp, a nucleotide-based second messenger with the capabilities to reprogram cell physiology. In other bacteria, RelA plays vital roles in stress survival (e.g., amino acid starvation), virulence and antibiotic tolerance34,35. Indeed, in a special case, Staphyloccus aureus upregulates two ppGpp synthetases RelQ and RelP in response to β-lactam exposure, promoting tolerance36. This prompted us to test whether the stringent response also confers tolerance to V. cholerae spheroplasts. To test the effects of (p)ppGpp on tolerance levels, we created a deletion mutant that lacks production of (p)ppGpp altogether (∆relA∆spoT∆relV, phenotypically confirmed via necessity of external supplementation of casamino acids during growth in minimal medium, Fig. S3A). However, this mutant did not yield any observed defects in antibiotic tolerance, nor resistance (Fig. S3B-D). This suggests that the stringent response does not play a role in PenG tolerance in V. cholerae under these conditions.
Metabolomics reveal metabolite flux during PenG exposure
To gain an understanding of how transcriptional changes correlate with cellular physiology, we next sought to identify changes in global metabolism associated with PenG exposure. To start with an unbiased approach, we conducted untargeted metabolomics. Following the same growth conditions previously used for the RNAseq experiment, we took samples at two different time points of penicillin treatment, 1 and 3 h. We reasoned that these timepoints would capture metabolite changes associated with transcriptome changes at one timepoint that overlaps with our RNAseq dataset (1 h) vs. a delayed response (3 h). Metabolites were extracted using ice cold methanol, and samples were analyzed using LC-MS (see Methods for details). Four different columns were used (HILIC positive/negative, and C18 positive/negative) to account for charge variations amongst the complete set of metabolites. After analysis of over 1000 metabolites, we arranged the datasets for significant log2 changes, with each dot representing detection on unique columns (Table S1 for full analysis). Displayed here are three subsets of the significant hits in relevant pathways (Fig. 3A–C).
A Metabolites involved in central carbon metabolism, B metabolites involved in bacterial cell wall synthesis, C metabolites involved in nucleotide metabolism. Data represent mean fold change, averaged between the different detection modes using untargeted metabolomics, and error bars represent SD. D Targeted metabolomics validate altered carbon flux. Metabolites were averaged and normalized to the non-treated condition. See Methods for details.
Metabolites associated with cell wall synthesis exhibited a general decrease in abundance, consistent with previous results showing depletion of cell wall precursors in β-lactam-treated cells37,38 due to “futile cycling”18 (Fig. 3A). In contrast, abundance of one cell wall metabolite, N-acetyl-1,6-anhydro-muramic acid, was elevated. This metabolite is the product of peptidoglycan turnover, and confirms that β-lactam exposure elevates cell wall recycling due to enhanced cell wall degradation11,39; it thus serves as an internal control that our untargeted metabolomics revealed informative metabolite changes. Interestingly, one isoform of D-glucosamine-6P increased in abundance, while the other isoform decreased (Table S1). We hypothesize that these two “isoforms” are in fact glucosamine-6P and glucosamine-1P, which we found to be differentially abundant in our targeted experiments (see section on targeted metabolomics).
One major pathway impacted by penicillin treatment was central carbon metabolism, specifically glycolysis. (Fig. 3B). Abundance of metabolites within the entire EMP pathway, from glucose-6-phosphate to pyruvate, decreased upon PenG treatment. Some metabolites were reduced by 4-fold over control while others, like fructose-1,6-bisphosphate and citric acid, experienced an over 8-fold decrease. Pentose phosphate pathway intermediates were also reduced (Fig. S4). These observations are consistent with rerouting of central carbon flux upon PenG exposure.
Lastly, nucleotide metabolites, both purines and pyrimidines and their pathway intermediates, were dramatically depleted in PenG-treated cells, especially the monophosphates (Fig. 3C). Interestingly, ATP levels appeared to be unchanged, which is different from β-lactam exposure in non-tolerant organisms like E. coli and Mycobacteroides abscessus, where ATP levels are at least transiently elevated in response to β-lactams. Elevated ATP has been proposed to be associated with respiratory burst and consequently additional killing by reactive oxygen species17,40,41,42. The depletion of diverse nucleotides we observed in our metabolomics dataset is consistent with the upregulation of nucleotide synthesis genes observed in our RNAseq dataset, as the transcriptional changes likely reflect the cell’s nucleotide starvation response, e.g. through PurR (purines) and other complex regulatory pathways43. We also observed a significant reduction in pentose phosphate pathway intermediates, like ribulose-5-phosphate (Ru5P) and erythrose-4-phosphate (Fig. S4). Our integrated -omics data thus indicate that penicillin treatment may ultimately starve the cell for energy and/or building blocks for diverse biosynthesis pathways.
To further interrogate our untargeted metabolomics results, we conducted targeted metabolomics on cells treated with PenG for 3 hours and quantified select indicator metabolites that reflect cell wall intermediates, glycolysis intermediates, and nucleotides. We subjected the extracted metabolites to LC-MS using a HILIC probe and compared the peaks to known standards. We normalized the average of four biological replicates to the non-treated condition and assembled heatmaps representing altered metabolites (Fig. 3D). First, we quantified the abundance of key glycolytic intermediates glucose-1P, glucose-6P, fructose-6P, fructose-1,6-BP, and pyruvate. After 3 h of penicillin exposure, we observed a reduction in all glycolytic metabolites, but the most extreme change was observed in glucose-6P and fructose-6P, which were reduced by 256-fold and 64-fold, respectively (Fig. 3D). Thus, PenG exposure dramatically alters glycolytic flux in tolerant spheroplasts, as suggested by our untargeted metabolomics. Glucose-6P connects to multiple metabolic pathways, and fructose-6P feeds directly into PG synthesis; the observed reduction in abundance could be indicative of altered flux from glycolysis via F6P (GlmS) into futile PG synthesis. We thus asked whether the depletion had physiological consequences, i.e. whether the reduction in G6P was harmful or protective to tolerant cells. Indeed, in the closely related L-Forms44,45, addition of glucose is detrimental, causing enhanced respiration and concomitant generation of reactive oxygen species46. To test whether spheroplasts exhibited a similar physiology, we supplemented our growth medium with glycolytic intermediates and conducted time-dependent killing (Fig. S5A). Pyruvate, fructose, and glucose at 0.2% did not affect PenG killing, suggesting that despite rapid depletion of intermediates, the ability to conduct glycolysis is neither limiting nor detrimental for V. cholerae spheroplasts.
We also used targeted metabolomics to confirm a reduction in UDP-GlcNAc, as well as other important cell wall precursors, like Diaminopimelic acid (DAP), N-Acetylmuramic acid, and glucosamine-1P (Fig. 3D). This reduction was an expected result due to redirection of carbon flux into futile PG synthesis and is consistent with previous observation in diverse bacteria showing depletion of cell wall precursors upon β-lactam exposure18. Lastly, we confirmed that nucleotides decreased upon penicillin treatment (Fig. 3D). GMP experienced the most pronounced reduction at 2.3-fold. UMP levels also dropped, possibly from the demand for UDP carriers involved in trying to repair the cell wall during penicillin treatment. Thus, our targeted metabolomics confirmed that central carbon metabolism, cell wall precursor levels and nucleotide homeostasis are affected by β-lactam treatment, as indicated by our untargeted metabolomics.
Disruption of nucleotide homeostasis potentiates killing by PenG
Next, we sought to probe the potential for synergistic action with PenG of the pathways uncovered in our combined transcriptomics and metabolomics. To this end, we used chemical inhibitors and supplements designed to inhibit or boost different pathways. After growing cells to exponential phase, we added 100 μg/mL (10 x MIC) PenG for 1 h. We then added the indicated additional compounds and plated for cell viability over time. We found that inhibiting the pentose phosphate pathway (through treatment with 6-aminonicotinamide47) exacerbated the killing effect of penicillin after 24 h (Fig. 5A). The pentose phosphate pathway supplies a key precursor for nucleotide synthesis, PRPP (phosphoribosyl pyrophosphate) (see Fig. 4). Our results therefore indicate that PenG-treated cells are indeed starved for nucleotides, and that proper nucleotide homeostasis is essential for optimal survival of β-lactam exposure. These results add support to our data implicating nucleotide levels as an important aspect of β-lactam susceptibility.
Major metabolites are presented with lines indicating connecting reactions. From central metabolism (gray) to the pentose phosphate pathway (green), the metabolites then get funneled into either the purine (red) or pyrimidine (orange) pathways. Relevant amino acid branch points are listed in blue, and genetic material is denoted in purple and pink.
Seeking to unravel the connection between nucleotides and tolerance further, we also challenged V. cholerae with both PenG and trimethoprim (trim), which blocks the conversion of dihydrofolate to tetrahydrofolate (ultimately resulting in purine and thymine starvation), and measured survival (Fig. 5B). After 24 h, trim by itself (50 µg/mL, 60 x MIC) reduced viability by 10-fold, while PenG caused the typically observed 50-fold killing (at 100 µg/mL, 10 x MIC). In combination, however, we observed a 105-fold reduction in viability, demonstrating synergistic killing between trim and PenG.
A CFU/mL were calculated over time with and without PenG, and/or 6-aminonicotinamide (6-AN, 100 μg/mL). B CFU/mL were calculated over time with and without PenG, and/or trimethoprim (Trim, 50 µg/mL, 60 x MIC). C Cells were treated as described for B), but with or without addition of nucleosides (NS, 2X) D CFU/mL were calculated over time with and without PenG, and/or fosfomycin (Fosfo, 100 µg/mL, 4 x MIC). A, B, and D are means of at least six independent biological replicates. C represents the means of 3 biological replicates. Error bars represent standard error. Statistical significance determined by unpaired t-test with Welch’s Corrections (* = p < 0.05, ** = p < 0.01).
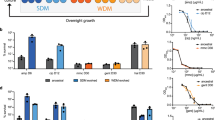
We reasoned that if nucleotide pools are limiting in PenG-treated cells, external addition of nucleotides should rescue PenG-mediated lethality. We tested a variety of nucleotides, as well as nucleotide analogs with documented inhibitory effects48,49,50,51. These supplementations did not cause any observable changes to PenG tolerance (Fig. S5B, C). Many bacteria import nucleosides at a higher rate than nucleotides33,52, and we thus used a nucleoside mix, with and without PenG, and plated for CFU/mL. Interestingly, this supplementation did not have any observable effects on tolerance either (Fig. 5C, Fig. S5D, E). Lastly, we turned to our synergy experiment with trimethoprim. Under these conditions, nucleoside supplementation reduced synergistic killing (Fig. 5C), but did not restore viability observed with PenG single treatment. In aggregate, these experiments suggest that i) nucleosides are imported by V. cholerae during antibiotic exposure, ii) nucleoside levels are not limiting for spheroplasts survival per se (perhaps due to the observed upregulation of de novo synthesis pathways) and iii) nucleosides can become limiting in spheroplasts when their synthesis is additionally suppressed. We next tested whether perturbations of nucleotide homeostasis could potentiate β-lactams against other Gram-negative pathogens as well. To this end, we turned to two hypertolerant clinical isolates of Enterobacterales, the Klebsiella pneumoniae strain TS1 and Escherichia coli bloodstream isolate WCM006. In a separate study10, we found both isolates to exhibit high tolerance to the carbapenem antibiotic meropenem, while being fully susceptible (MIC < 2 µg/mL). We thus treated both isolates with meropenem (10 µg/mL), trimethoprim (30 µg/mL), or both in the more host-like growth medium Brain-Heart Infusion broth. In both bacteria, trimethoprim dramatically potentiated meropenem-mediated killing (Fig. 6), demonstrating strong synergy in killing (Bliss Independence Model). Addition of nucleoside mix to these bacteria resulted in a reduction of trim-mero combinatorial killing (albeit statistically significant only in K. pneumoniae; perhaps due to a higher proficiency in NS uptake during stress conditions) (Fig. S5E). Thus, it is likely that futile cycling-mediated depletion of nucleotide precursors may represent a conserved physiological state among tolerant spheroplasts.
CFU/mL were calculated over time with and without meropenem (Mero, 10 µg/mL), and/or trimethoprim (Trim, 30 µg/mL). Data are means +/- standard deviation of four independent biological replicates. “Additive” line demonstrates expected results (based on Bliss Independence Model calculations) if the two antibiotics were additive instead of synergistic.
In an attempt to genetically manipulate the purine biosynthesis pathway, we next deleted the nucleotide starvation regulator PurR. We did not observe any defects in PenG tolerance nor resistance (Fig. S6A-B). This was surprising, as other studies have found altered antibiotic efficacy of a purR deletion mutant in the presence of β-lactam antibiotics51,53,54,55. It is possible that V. cholerae’s regulation of nucleotide starvation does not mimic what has been established in well-studied model organisms, and further work is needed to uncover the regulatory effects of PurR in this system.
Lastly, we also reasoned that the depletion of PG precursors should make tolerant cells particularly susceptible to inhibition of PG precursor synthesis, e.g., by sabotaging post-antibiotic recovery. We thus exposed cells to a combination of PenG (100 µg/mL, 10 x MIC) and Fosfomycin (100 µg/mL, 4 x MIC). Fosfomycin inhibits MurA, which catalyzes an early step in PG precursor synthesis. Interestingly, fosfomycin by itself, despite ultimately causing cell wall degradation like PenG19, did not result in any reduction in viability. This perhaps reflects that PenG does indeed cause death through more than just cell wall degradation, i.e. the additional killing is due to futile cycling and other metabolic demands. However, in combination, PenG and fosfomycin caused a dramatic, 105-fold reduction in plating efficiency, demonstrating strong synergy between PenG and precursor synthesis inhibition (Fig. 5C). We conclude that PG precursors are limiting in tolerant cells (see proposed model in Fig. 7).
Upon β-lactam exposure, the nucleotide pool is placed under stress, from processes like transcription, translation, and cell wall repair. This causes de novo biosynthesis of nucleotides to be activated. Inhibiting the pentose phosphate pathway (PPP), which feeds de novo biosynthesis, through the exposure of 6-aminonicotinamide, the cell suffers more from penicillin exposure.
Discussion
Antibiotic tolerance is an understudied contributor to antibiotic treatment failure. The mechanisms and physiology of tolerance have only begun to be unraveled. In this work, we dissected the alterations in gene expression and metabolic flux associated with antibiotic tolerance. Using a multi-omics tool set and a hypertolerant model organism, we identified large genetic and metabolic perturbations in multiple key categories including cell wall biosynthesis, heat shock response, the TCA cycle, and nucleotide metabolism. Interestingly, with the exception of nucleotide homeostasis, pathways that were differentially regulated transcriptionally did not generally show a corresponding metabolomic signature and vice versa. It is possible that both transcriptomic and metabolic fluxes are too individually dynamic to capture overlap in our experimental system; additionally, however, the metabolome is also altered by diverse inputs (not just gene expression), which may make it difficult to uncover connections between the two. Similarly, the lack of mutant phenotypes (e.g., ∆ppGpp) likely reflects a dynamic situation where not all responses actually contribute to survival. At any rate, pathways that are altered in tolerant cells (and causal for tolerance) could serve as targets for future exploration of antimicrobial adjuvants, which we have begun in this study by showing that disturbing nucleotide homeostasis strongly synergizes with a β-lactam antibiotic against tolerant V. cholerae.
A key unanswered question about spheroplast-mediated tolerance is how these bacteria mitigate the dramatic damage sustained due to loss of their cell wall. It has become clear in recent years that antibiotics cause disturbances in cellular physiology that go beyond the simple direct consequences of target inhibition. Indeed, we and others have previously measured a marked accumulation of reactive oxygen species in response to β-lactam exposure, likely due to dysregulation of the electron transport chain, though the degree to which this contributes to killing is unclear (and evidently minimal in tolerant spheroplast)14,56,57. Additionally, data from E. coli and Streptococcus pneumoniae demonstrate that β-lactams induce “futile cycling”, i.e., the constant generation and degradation of uncrosslinked PG strands, which ultimately results in poorly-understood energy and redox imbalances, thereby putatively contributing to cell death and/or growth arrest17,18. However, spheroplasts do not appreciably die over extended time periods, indicating that they possess efficient repair processes that mitigate both ROS production and negative consequences of futile cycling. Our data shed more light on how this might be accomplished. For example, one of the immediate consequences of ROS accumulation is the production of oxidized (e.g., likely dysfunctional) proteins12. In our transcriptomics dataset we show that cells respond to this by highly upregulating heat shock proteins, which might help remove or refold non-functional proteins. Our metabolomics experiment shows clear signatures of futile cycling, i.e. depletion of cell wall precursors and concomitant rerouting of central carbon metabolism away from glycolysis. At the same time, ATP homeostasis does not appear to be disturbed, indicating that spheroplasts are perfectly capable of buffering the energy demand that should be associated with changed metabolism. The observation that nucleotides are depleted in spheroplasts, concomitant with upregulation of nucleotide synthesis pathways can be interpreted as an indirect consequence of futile cycling. Rerouting of central carbon metabolism into futile cell wall synthesis could both deplete precursors for the pentose phosphate pathway (which ultimately generates nucleotide precursors) and also increase the use of nucleotides involved in activation of precursors, e.g. UDP-GlcNAc. The upregulation of stress responses (like the VxrAB cell envelope stress system) can be expected to put additional demand on ATP and GTP due to enhanced anabolic reactions. It is interesting in this context that the triphosphate forms of nucleotides (ATP and GTP) themselves do not seem to be affected. This may indicate that nucleotide diphosphate kinases have low Kd values and can operate optimally even in dramatically reduced substrate concentrations, explaining perhaps why this energy demand does not by itself have lethal consequences in tolerant spheroplasts.
On a broader note, our data uncover that spheroplasts appear to be in a physiological state that is distinct from susceptible bacteria or even the closely related L-Forms44,45. For example, we found that the ability to generate ppGpp (a well-characterized facilitator of tolerance in persisters58) does not contribute to tolerance in spheroplasts. In contrast to L-form survival46 and E. coli resistance to killing59,60, ROS levels do not seem to be limiting for spheroplast survival, and addition of glucose does not kill these cells. Also, unlike E. coli treated with β-lactams, we do not observe clear increases in ATP levels, protein synthesis or accumulation of glycolytic intermediates and cell wall precursors (in fact we observed the opposite). However, the connection between nucleotide homeostasis and β-lactam susceptibility is not without precedent, suggesting broad conservation: In S. aureus, mutations in the pentose phosphate pathway reduce oxacillin-induced lysis, though this is thought to be due to surface alterations and/or slow growth61,62. In B. subtilis, nucleotide homeostasis is tied to cefuroxime resistance via mutations in RNA polymerase63. Overall, a somewhat inconclusive body of work has emerged on the role of nucleotide synthesis on tolerance, persistence, and resistance51. This inconclusiveness might be caused by differential effects that the underlying physiology has on various forms of tolerance vs. various forms of resistance. Our results here now shed light onto the complex and interconnected response to β-lactam antibiotics in a hypertolerant bacterium. Our data also highlight novel points of vulnerability to devise new avenues for antibiotic adjuvants.
Methods
Strain construction
The wild-type strain used in this study is V. cholerae El Tor strain N1696164, and the ΔvxrAB strain was constructed in a prior study16. Deletion constructs were made using pTOX65. In short, 500 bp upstream and downstream of the gene were PCR amplified using the relevant primers listed in Table S2. The fragments were then simultaneously ligated and inserted into the EcoRV-digested vector using Gibson Assembly and heat transformed into DH5α competent E. coli cells followed by selection on LB agar with 0.2% glucose (w/v) and 20 μg/mL chloramphenicol. Plasmids were extracted using Omega Miniprep kit and verified via Sanger Sequencing. Plasmids were then heat transformed into conjugative MFDλpir and conjugated into WT N16961 following the steps outlined below.
Conjugation into WT N16961 went as follows: 100 μL of donor and recipient were mixed together, spotted on LB agar with 60 μM DAP for MFD, and left to incubate for 4 h at 37 °C. The inoculum was then scraped and spread on LB agar with 100 μg/mL streptomycin and 20 μg/mL chloramphenicol and 0.2% glucose (w/v) (pTOX) and incubated overnight at 30 °C. Transconjugants were single-colony purified and then spread onto counter-selection plates: minimal media agar with 2% rhamnose (w/v) (pTOX). Plates were incubated overnight at 30 °C. Colonies were screened for the correct deletion using primers in Table S2. All strains were additionally verified via whole genome sequencing.
The relV gene was deleted in a ∆relA∆spoT background (gift from the Waldor lab) using the allele exchange plasmid pCVD442. In brief, homology regions upstream and downstream of relV were amplified using primers TDP1112/1113 and TDP 1114/1115 and cloned into SmaI-digested pCVD442 using isothermal assembly. Successful clones (sequence verified) were conjugated into the ∆relA∆spoT strain using the donor strain SM10. Post- conjugation, transconjugant single crossover colonies were purified on strep (200 μg/mL)/carbenicillin (100 μg/mL). Single colonies were then spread on LB + strep + 10%sucrose, followed by incubation at 30 °C overnight. Successful mutants were verified via colony PCR (using primers TDP 1110/1111).
Bacterial Growth Conditions
V. cholerae was grown in Luria-Bertani (also called lysogeny broth) (LB) medium (for a 1 L bottle, 10 g Casein peptone, 5 g yeast extract, 10 g NaCl, and 12 g agar, unless otherwise indicated. All premade from Fischer Bioreagents) at 30 °C. 200 μg/mL of streptomycin was also added (N16961 is streptomycin resistant). M9 minimal medium was made at by mixing 15 g agar, 200 mL 5x M9 salts in 1 L of sterile water (1 L contains 35 g Na2HPO4•7H2O, 15 g KH2PO4, 2.5 g NaCl, 5 g NH4Cl), 0.5 mL 1 M MgSO4, 0.1 mL 1 M CaCl2, 1 mL FeCl3/citric acid, 10 mL glucose (20% w/v). All samples were grown overnight at 30 °C and experimented at 37 °C. Klebsiella pneumoniae and Escherichia coli were grown in Brain-Heart Infusion broth (BHI, beef heart infusion 5 g/L, calf brain infusion 12.5 g/L, disodium hydrogen phosphate, 2.5 g/L, D-glucose, 2 g/L, peptone, 10 g/L, NaCl, 5 g/L).
Cell culture condition for RNA-Seq
Wild-type and mutant strains were from an isolated single colony and grown in LB medium containing 200 µg/mL streptomycin overnight at 30 °C. 2 mL of the culture was then added to 250 mL LB + 200 µg/mL streptomycin in a 1 L flask and left to grow until mid- to late exponential phase (OD600 = 0.4 - 0.5). 30 mL of exponential phase cultures were pelleted after treatment of (+) / (-) 100 µg/mL PenG as indicated time points (0, 10, 20, 40, 60, 90 min). RNA was isolated by using TRIzol reagent (Ambion) and DNA was removed using the Turbo DNase kit (Ambion). RNA samples were pooled as pool 1 (first set of biological triplicates and technical replicate 1) and pool 2 (second set of biological triplicates and technical replicate 2). Following cDNA synthesis, samples were subjected to high-throughput sequencing. The RNAseq data was first uploaded to the Galaxy web platform (https://usegalaxy.org) for quality control and trimming of the raw sequencing reads. Next the trimmed reads were aligned to the Vibrio cholerae N16961 reference genome (from NCBI: GCF_000006745.1) and htseq-count was used to calculate the number of reads mapping to each feature. The count files were then used in R to perform differential expression analysis using DESeq2. The resulting log2FoldChange values were then used in GraphPad Prism 9 to generate heatmaps for visual representation of the data. GO and KEGG pathway analysis was conducted using the tools available at https://bioinformatics.sdstate.edu/go/
Cell culture conditions for metabolomics
Three or four biological replicates were grown in LB overnight at 30 °C (for untargeted and targeted respectively). 2 mL of culture was added to 250 mL of LB in a 1 L flask and incubated 1.5 h at 37 °C to OD600 = 0.4-0.6. (+) / (-) 100 μg/mL PenG was added to the flasks and left to incubate an additional 1 or 3 h. After treatment, 2 mL of sample were taken per condition and pelleted at 7000 rpm for 2 min. The supernatant was removed, and the cell material pellet was flash frozen with liquid nitrogen. 200 μL of cold 80% methanol was added to the pellets. Pellets were stored at -80 °C.
Untargeted metabolomics were conducted by the Cornell BRC facility for preparation and analysis. Samples were vortexed for 5 min on vortex mixer (1800 rpm/5 minutes) until the pellet disintegrated and was mixed thoroughly in extraction solvent. Cells were then sonicated (in Ultrasonic Bath) for 20 min. Cells were then vortexed for 10 seconds to evenly resuspend immediately followed by removal of a normalized amount (based on number of proteins in each sample). Then, 80% Methanol was added to make final vol = 600 μl in each sample. The samples were then incubated 60 min at 4 °C, followed by centrifugation at 16,000 g/10 mins/4°C. The supernatant was removed (vol = 250 μl X2) into separate 1.5 ml Eppendorf tubes (2 aliquots -one for each for HILIC & C18 analysis), dried in speed vac and stored at -20 °C until ready to analyze. The dried extract was reconstituted in 200 μL ACN 20%, 0.1% FA (30 μL) for C18 and ACN 20% for HILIC. The solution was then sonicated for about 5 min and then transferred into LCMS vial, 10 μl for QC for each group; the rest was kept at -20 °C. For assessing the LCMS machine performance, two QA samples containing 15 standard compounds were run before and after real samples respectively. For normalizing the chromatograms, a global Quality Control sample containing all groups was run twice at the beginning of the sequence, at the end, and every 6 samples, and for compound identification, each of three group QCs (0, 1, 3 h) was run twice by MS/MS. Global Quality Control sample is a pool of all the samples. Group QC sample is a pool of all 6 samples within the time group.
Compound discoverer was used for untargeted metabolomics analysis, comparing the m/z in the samples with different databases: Chemspider, bioCyc, HMDB, SMDPD, Lipidmaps, Mzcloud. C18 LC-MS summary: 589 and 441 annotated metabolites were identified following CD search and filtering for positive ion and negative ion mode respectively. After removing the background and redundancy, 554 and 413 unique metabolites were annotated, in which 318 and 218 metabolites were identified by MS/MS for preferred ion. HILIC LC-MS summary: 787 and 505 annotated metabolites were identified following CD search and filtering for positive ion and negative ion mode respectively. After removing the background and redundancy, 760 and 502 unique metabolites were annotated, in which 696 and 469 metabolites were identified by MS/MS for preferred ion. Presented are those with a > 2-fold change and an adjusted p-value < 0.05. Note: Same name of metabolites with retention times >0.3 min were manually labeled by original name iso, for which more than 2 isoforms labeled by iso1…4 are considered as unique isoforms. For targeted metabolomics, HILIC protocol and analysis was conducted following66. Heatmaps were generated using Prism, with averaged peak heights normalized to the control ((-) PenG) condition.
Synergy Killing Assay
WT Vibrio cholerae cultures were grown overnight in 5 mL LB broth at 30°C. The following morning, they were diluted 1:100 into fresh LB and grown at 37 °C for 1.5 h. (+) / (-) 100 μg/mL PenG was then added for 1 hour. Other chemicals were then added at the following concentrations: 100 μg/mL 6-aminonicotinamide (6-AN); 100 μg/mL fosfomycin (fosfo); 50 μg/mL trimethoprim (trim) (others designated in figure legends). The nucleotide mix was purchased from MilliporeSigma™ EmbryoMax™ Nucleosides (100X) and contained nucleotides at the following concentrations; Cytidine: 0.73 g/L; Guanosine: 0.85 g/L; Uridine: 0.73 g/L; Adenosine: 0.8 g/L; Thymidine: 0.24 g/L. We used a 2x final concentration (50-fold dilution of stock) for our assays. After chemical addition, 100 μL samples were taken at 0, 1-, 3-, 6- (for some of the experiments), and 24-hour timepoints and serially diluted in 1xPBS. 5 μL spots were plated on LB agar (or M9 minimal media supplemented with 0.2% glucose) and incubated overnight at 30 °C (37 °C for M9). CFU/mL were counted the following day. For K. pneumoniae and E. coli, killing experiments were conducted as previously reported: Overnights in BHI were diluted 10-fold into fresh, prewarmed BHI medium supplemented with meropenem (10 µg/mL), trimethoprim (50 µg/mL), the combination, or vehicle controls (water for meropenem; DMSO for trimethoprim). Cells were then incubated standing at 37 C for the indicated timeframe, and CFU/mL determined as described above. To quantify synergy, Bliss Independence Model calculations were conducted. We calculated Eadditive = KA + KB-(KA * KB) where K = 1 – [survival fraction] (CFU/mL after exposure/initial CFU/mL) of antibiotic A (meropenem) and B (trimethoprim). Synergy is established if Eadditive < KA*KB. The “additive” line in killing graphs indicates the expected values if effects were additive rather than synergistic, calculated as [survival fractionA] * [survival fractionB] * initial CFU/mL.
Data Availability
RNAseq data have been deposited under Bioproject accession number PRJNA1338477.
Code availability
Not applicable
References
Ventola, C. L. The antibiotic resistance crisis. P T 40, 277–283 (2015).
Dörr, T. Understanding tolerance to cell wall–active antibiotics. Ann. N. Y. Acad. Sci. 1496, 35–58 (2021).
Deventer, A. T., Stevens, C. E., Stewart, A. & Hobbs, J. K. Antibiotic tolerance among clinical isolates: mechanisms, detection, prevalence, and significance. Clin. Microbiol. Rev. 0, e00106–e00124 (2024).
Deter, H. S., Hossain, T. & Butzin, N. C. Antibiotic tolerance is associated with a broad and complex transcriptional response in E. coli. Sci. Rep. 11, 6112 (2021).
Levin-Reisman, I. et al. Antibiotic tolerance facilitates the evolution of resistance. Science 355, 826–830 (2017).
Windels, E. M., Michiels, J. E., Van den Bergh, B., Fauvart, M. & Michiels, J. Antibiotics: combatting tolerance to stop resistance. mBio 10, e02095–19 (2019).
Santi, I., Manfredi, P., Maffei, E., Egli, A. & Jenal, U. Evolution of antibiotic tolerance shapes resistance development in chronic pseudomonas aeruginosa infections. mBio 12, e03482–20 (2021).
Cross, T. et al. Spheroplast-mediated carbapenem tolerance in gram-negative pathogens. Antimicrobial Agents Chemother. 63, e00756-19 (2019).
Murtha, A. N. et al. High-level carbapenem tolerance requires antibiotic-induced outer membrane modifications. PLOS Pathog. 18, e1010307 (2022).
Cross, T. et al. Prevalence and mechanisms of high-level carbapenem antibiotic tolerance in clinical isolates of Klebsiella pneumoniae. Preprint at https://doi.org/10.1101/2025.02.19.639047 (2025).
Islam, N. et al. Peptidoglycan recycling promotes outer membrane integrity and carbapenem tolerance in acinetobacter baumannii. mBio 13, e0100122 (2022).
Monahan, L. G. et al. Rapid conversion of Pseudomonas aeruginosa to a spherical cell morphotype facilitates tolerance to carbapenems and penicillins but increases susceptibility to antimicrobial peptides. Antimicrob. Agents Chemother. 58, 1956–1962 (2014).
Bottone, E. J., Brandman, Z. & Schneierson, S. S. Spheroplasts of Haemophilus influenzae Induced by Cell Wall-Active Antibiotics and Their Effect upon the Interpretation of Susceptibility Tests. Antimicrobial Agents Chemother. 9, 327–333 (1976).
Shin, J. et al. A multifaceted cellular damage repair and prevention pathway promotes high-level tolerance to β-lactam antibiotics. EMBO Rep. 22, e51790 (2021).
Weaver, A. I. et al. Genetic Determinants of Penicillin Tolerance in Vibrio cholerae. Antimicrob Agents Chemother 62, (2018).
Dörr, T. et al. A cell wall damage response mediated by a sensor kinase/response regulator pair enables beta-lactam tolerance. PNAS 113, 404–409 (2016).
Lobritz, M. A. et al. Increased energy demand from anabolic-catabolic processes drives β-lactam antibiotic lethality. Cell Chem. Biol. 29, 276–286.e4 (2022).
Cho, H., Uehara, T. & Bernhardt, T. G. Beta-Lactam Antibiotics Induce a Lethal Malfunctioning of the Bacterial Cell Wall Synthesis Machinery. Cell 159, 1300–1311 (2014).
Dörr, T., Davis, B. M. & Waldor, M. K. Endopeptidase-Mediated Beta Lactam Tolerance. PLOS Pathog. 11, e1004850 (2015).
Mayer, C. et al. Bacteria’s different ways to recycle their own cell wall. Int. J. Med. Microbiol. 309, 151326 (2019).
Park, J. T. & Uehara, T. How Bacteria Consume Their Own Exoskeletons (Turnover and Recycling of Cell Wall Peptidoglycan). Microbiol. Mol. Biol. Rev. 72, 211–227 (2008).
Egan, A. J. F., Errington, J. & Vollmer, W. Regulation of peptidoglycan synthesis and remodelling. Nat. Rev. Microbiol. 18, 1–15, https://doi.org/10.1038/s41579-020-0366-3 (2020).
Vigouroux, A. et al. Class-A penicillin binding proteins do not contribute to cell shape but repair cell-wall defects. eLife 9, e51998 (2020).
Bie, L. et al. Comparative Analysis of Transcriptomic Response of Escherichia coli K-12 MG1655 to nine representative classes of antibiotics. Microbiol. Spectr 11, e00317 (2023).
Groisman, E. A., Duprey, A. & Choi, J. How the PhoP/PhoQ System Controls Virulence and Mg2+ homeostasis: lessons in signal transduction, pathogenesis, physiology, and evolution. Microbiol. Mol. Biol. Rev. 85, e0017620 (2021).
Matson, J. S., Livny, J. & DiRita, V. J. A putative Vibrio cholerae two-component system controls a conserved periplasmic protein in response to the antimicrobial peptide polymyxin B. PLOS ONE 12, e0186199 (2017).
Liu, M. et al. CitAB Two-Component System-Regulated Citrate Utilization Contributes to Vibrio cholerae Competitiveness with the Gut Microbiota. Infect. Immun. 87, e00746–18 (2019).
Lazaretti, W. Y. et al. Upregulation of the clpB gene in response to heat shock and beta-lactam antibiotics in Acinetobacter baumannii. Mol. Biol. Rep. 47, 1499–1505 (2020).
Poole, K. Bacterial stress responses as determinants of antimicrobial resistance. J. Antimicrob. Chemother. 67, 2069–2089 (2012).
Powell, J. K. & Young, K. D. Lysis of Escherichia coli by beta-lactams which bind penicillin-binding proteins 1a and 1b: inhibition by heat shock proteins. J. Bacteriol. 173, 4021–4026 (1991).
Slamti, L., Livny, J. & Waldor, M. K. Global Gene Expression and Phenotypic Analysis of a Vibrio cholerae rpoH Deletion Mutant. J. Bacteriol. 189, 351–362 (2007).
Piette, J. et al. DNA sequence of the carA gene and the control region of carAB: tandem promoters, respectively controlled by arginine and the pyrimidines, regulate the synthesis of carbamoyl-phosphate synthetase in Escherichia coli K-12. Proc. Natl. Acad. Sci. USA 81, 4134–4138 (1984).
Goncheva, M. I., Chin, D. & Heinrichs, D. E. Nucleotide biosynthesis: the base of bacterial pathogenesis. Trends Microbiol. 30, 793–804 (2022).
Sinha, A. K. & Winther, K. S. The RelA hydrolase domain acts as a molecular switch for (p)ppGpp synthesis. Commun. Biol. 4, 1–10 (2021).
Urwin, L., Savva, O. & Corrigan, R. M. Microbial Primer: what is the stringent response and how does it allow bacteria to survive stress? Microbiology 170, 001483 (2024).
Geiger, T., Kästle, B., Gratani, F. L., Goerke, C. & Wolz, C. Two Small (p)ppGpp synthases in staphylococcus aureus mediate tolerance against cell envelope stress conditions. J. Bacteriol. 196, 894–902 (2014).
Kohlrausch, U. & Höltje, J. V. Analysis of murein and murein precursors during antibiotic-induced lysis of Escherichia coli. J. Bacteriol. 173, 3425–3431 (1991).
Uehara, T. & Park, J. T. Growth of Escherichia coli: significance of peptidoglycan degradation during elongation and septation. J. Bacteriol. 190, 3914–3922 (2008).
Dik, D. A., Fisher, J. F. & Mobashery, S. Cell-wall recycling of the gram-negative bacteria and the nexus to antibiotic resistance. Chem. Rev. 118, 5952–5984 (2018).
Lindman, M. & Dick, T. Bedaquiline Eliminates Bactericidal Activity of β-Lactams against Mycobacterium abscessus. Antimicrobial Agents and Chemotherapy 63, https://doi.org/10.1128/aac.00827-19 (2019).
Shetty, A. & Dick, T. Mycobacterial cell wall synthesis inhibitors cause lethal ATP burst. Front. Microbiol. 9, 01898 (2018).
Dwyer, D. J. et al. Antibiotics induce redox-related physiological alterations as part of their lethality. Proc. Natl. Acad. Sci. USA 111, E2100–E2109 (2014).
Turnbough, C. L. & Switzer, R. L. Regulation of pyrimidine biosynthetic gene expression in bacteria: repression without repressors. Microbiol. Mol. Biol. Rev. 72, 266–300 (2008).
Errington, J. Cell wall-deficient, L-form bacteria in the 21st century: a personal perspective. Biochem Soc. Trans. 45, 287–295 (2017).
Mickiewicz, K. M. et al. Possible role of L-form switching in recurrent urinary tract infection. Nat. Commun. 10, 4379 (2019).
Kawai, Y. et al. Crucial role for central carbon metabolism in the bacterial L-form switch and killing by β-lactam antibiotics. Nat. Microbiol. 4, 1716–1726 (2019).
Rashida, Z. & Laxman, S. The pentose phosphate pathway and organization of metabolic networks enabling growth programs. Curr. Opin. Syst. Biol. 28, 100390 (2021).
Niazy, A. A., Alrashed, M. M., Lambarte, R. N. A. & Niazy, A. A. 5-Fluorouracil inhibits bacterial growth and reduces biofilm in addition to having synergetic effects with gentamicin against pseudomonas aeruginosa. Microorganisms 12, 2257 (2024).
Zhang, M. et al. Deciphering the antibacterial mechanisms of 5-fluorouracil in escherichia coli through biochemical and transcriptomic Analyses. Antibiotics 13, 528 (2024).
Brown, N. C., Wisseman, C. L. & Matsushita, T. Inhibition of Bacterial DNA Replication by 6-(p-Hydroxyphenylazo)-uracil. Nat. N. Biol. 237, 72–74 (1972).
Stevens, C. E., Deventer, A. T. & Hobbs, J. K. The impact of bacterial purine metabolism on antibiotic efficacy. npj Antimicrob. Resist 3, 69 (2025).
Gumpenberger, T. et al. Nucleoside uptake in Vibrio cholerae and its role in the transition fitness from host to environment. Mol. Microbiol. 99, 470–483 (2016).
Nolan, A. C. et al. Purine Nucleosides Interfere with c-di-AMP Levels and Act as Adjuvants To Re-Sensitize MRSA To β-Lactam Antibiotics. mBio 14, e02478–22 (2022).
Sause, W. E. et al. The purine biosynthesis regulator PurR moonlights as a virulence regulator in Staphylococcus aureus. Proc. Natl. Acad. Sci. 116, 13563–13572 (2019).
Xiong, Y. Q. et al. The purine biosynthesis repressor, purr, contributes to vancomycin susceptibility of methicillin-resistant staphylococcus aureus in experimental endocarditis. J. Infect. Dis. 229, 1648–1657 (2024).
Keren, I., Wu, Y., Inocencio, J., Mulcahy, L. R. & Lewis, K. Killing by bactericidal antibiotics does not depend on reactive oxygen species. Science 339, 1213–1216 (2013).
Liu, Y. & Imlay, J. A. Cell death from antibiotics without the involvement of reactive oxygen species. Science 339, 1210–1213 (2013).
Fung, D. K. et al. A shared alarmone–GTP switch controls persister formation in bacteria. Nat. Microbiol 10, 1617–1629 (2025).
Kohanski, M. A., Dwyer, D. J., Hayete, B., Lawrence, C. A. & Collins, J. J. A common mechanism of cellular death induced by bactericidal antibiotics. Cell 130, 797–810 (2007).
Thorsing, M., Bentin, T., Givskov, M., Tolker-Nielsen, T. & Goltermann, L. The bactericidal activity of β-lactam antibiotics is increased by metabolizable sugar species. Microbiol. (Read.) 161, 1999–2007 (2015).
Stevens, C. E. et al. Staphylococcus aureus COL: an atypical model Strain of MRSA That Exhibits Slow Growth and Antibiotic Tolerance due to a Mutation in PRPP Synthetase. Mol Microbiol https://doi.org/10.1111/mmi.70000 (2025).
Zeden, M. S. et al. Metabolic reprogramming and altered cell envelope characteristics in a pentose phosphate pathway mutant increases MRSA resistance to β-lactam antibiotics. PLOS Pathog. 19, e1011536 (2023).
Patel, Y. & Helmann, J. D. A mutation in RNA polymerase imparts resistance to β-lactams by preventing dysregulation of amino acid and nucleotide metabolism. Cell Rep. 44, 115268 (2025).
Heidelberg, J. F. et al. DNA sequence of both chromosomes of the cholera pathogen Vibrio cholerae. Nature 406, 477–483 (2000).
Lazarus, J. E. et al. A new suite of allelic-exchange vectors for the scarless modification of proteobacterial genomes. Appl. Environ. Microbiol. 85, e00990-19 (2019).
Planck, K. A. & Rhee, K. Metabolomics of Mycobacterium tuberculosisMycobacterium tuberculosis (M. tuberculosis). in Mycobacteria Protocols (eds Parish, T. & Kumar, A.) 579–593 (Springer US, New York, NY, 2021). https://doi.org/10.1007/978-1-0716-1460-0_25.
Acknowledgements
MRK was supported by a training grant from the National Institutes of Health under award T32AI145821. Tolerance research in the Dörr lab was supported by NIH R01 AI143704. The Rhee lab is funded by NIH R25 AI140472. This work was additionally partially supported by a seed grant from the Cornell University Biotechnology Resource Center (BRC) for experiments conducted in the BRC Proteomics and Metabolomics facility (RRID:SCR_021743). We thank Lars Westblade (Weill Cornell Medicine) for the clinical isolates of K. pneumoniae and E. coli. KR and TD acknowledge support from the Cornell Center for Antimicrobial Resistance Research and Education (CCARRE).
Author information
Authors and Affiliations
Contributions
M.R.K. designed the study, conducted experiments, and analyzed the data. M.I.K. conducted experiments, and contributed to data analysis. A.S. contributed reagents, conducted experiments, and analyzed data. U.B. and J.H.S. conducted experiments. K.R. contributed reagents and analyzed data. T.D. designed the study, conducted experiments, analyzed data, and acquired funding.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Keller, M.R., Kazi, M.I., Saleh, A. et al. Global response to antibiotic exposure reveals a critical role for nucleotide metabolism in high-level β-lactam tolerance. npj Antimicrob Resist 4, 11 (2026). https://doi.org/10.1038/s44259-026-00183-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s44259-026-00183-x